Colloquium
Suramin inhibits initiation of defense signaling by systemin, chitosan, and a β-glucan elicitor in suspension-cultured Lycopersicon peruvianum cells
Johannes Stratmann, Justin Scheer, and Clarence A. Ryan*
Institute of Biological Chemistry, Washington State University, Pullman, WA 99164-6340
Systemin-mediated defense signaling in tomato (Lycopersicon esculentum) plants is analogous to the cytokine-mediated inflammatory response in animals. Herein, we report that the initiation of defense signaling in suspension-cultured cells of Lycopersicon peruvianum by the peptide systemin, as well as by chitosan and β-glucan elicitor from Phytophtora megasperma, is inhibited by the polysulfonated naphtylurea compound suramin, a known inhibitor of cytokine and growth factor receptor interactions in animal cells. Using a radioreceptor assay, we show that suramin interfered with the binding of the systemin analog 125I-Tyr-2,Ala-15-systemin to the systemin receptor with an IC50of 160 µM. Additionally, labeling of the systemin receptor with a photoaffinity analog of systemin was inhibited in the presence of suramin. Receptormediated tyrosine phosphorylation of a 48-kDa mitogen-activated protein kinase and alkalinization of the medium of suspensioncultured cells in response to systemin and carbohydrate elicitors were also inhibited by suramin. The inhibition of medium alkalinization by suramin was reversible in the presence of high concentrations of systemin and carbohydrate elicitors. Calyculin A and erythrosin B, intracellular inhibitors of phosphatases and plasma membrane proton ATPases, respectively, both induce medium alkalinization, but neither response was inhibited by suramin. The polysulfonated compound heparin did not inhibit systemininduced medium alkalinization. NF 007, a suramin derivative, induced medium alkalinization, indicating that neither NF 007 nor heparin interact with elicitor receptors like suramin. The data indicate that cell-surface receptors in plants show some common structural features with animal cytokine and growth factor receptors that can interact with suramin to interfere with ligand binding.
S ystemin is an 18-amino acid wound signal in plants that is released during herbivore attacks or other mechanical damage (1, 2). Over 20 defense-related proteins are synthesized in response to systemin, including components of the signal transduction pathway and several antinutritional proteins (3, 4). Recently, a 160-kDa systemin-binding protein was identified in plasma membranes of suspension-cultured cells of Lycopersicon peruvianum whose properties are characteristic of a systemin receptor. The protein exhibited a single, specific, high-affinity binding site for systemin (Kd = 0.15 nM). Competition for systemin binding by systemin analogs corresponded with their biological activities, indicating that this protein represents the physiological receptor for systemin (5, 6).
Several intracellular signaling events occur within a few minutes after systemin perception, including increases in proton and Ca2+ concentrations (7, 8, 9 and 10), efflux of potassium ions (10), transcription of a calmodulin gene (11), and increases in activities of a 48-kDa mitogen-activated protein kinase (MAPK; refs.5 and 12) and a phospholipase A2 (13).
In addition to wounding, defense genes can be activated by supplying solutions of systemin and oligosaccharide elicitors through stems of excised plants. Cumulative evidence indicates that systemin and oligosaccharide elicitors activate the same signaling pathway as does wounding (14, 15 and 16), but they employ unique receptors.
Systemin and several pathogen elicitors induce medium alkalinization in suspension-cultured tomato cells (10, 17, 18). The change of the proton gradient across the membrane in response to systemin is mediated through the inhibition of a plasma membrane H+-ATPase leading to an influx of protons accompanied by an efflux of potassium ions (9, 10). The alkalinization response seems to be an essential event leading to defense gene activation (9).
The wound signaling pathway in tomato plants is analogous to the inflammatory response of animals initiated by infections or severe trauma and mediated by tumor necrosis factor-α and other cytokines (3, 4, 19, 20). In animal cells, interactions of growth factors with their receptors are inhibited by suramin, a polysulfonated naphtylurea compound, 8,8′-[carbonylbis[imino-3,1-phenylenecarbonylimino(4-methyl-3,1-phenylene)carbonyl-imino]]bis-1,3,5-naphtalenetrisulfonic acid hexasodium salt. Suramin interferes with the binding of several polypeptide signals, including epidermal growth factor (21, 22 and 23), platelet-derived growth factor (21, 22, 24, 25), basic fibroblast growth factor (26, 27), transforming growth factor-β (22), interleukins (28, 29), and tumor necrosis factor-α (30).
We report herein that suramin potently inhibits binding of the polypeptide defense signal systemin to the systemin receptor in tomato cells. Suramin also inhibits rapid receptor-mediated cellular responses induced by systemin and oligosaccharide elicitors.
Materials and Methods
Plant Material. Tomato plants (Lycopersicon esculentum cv. Castlemart) were grown for 14–17 days under 17 h of light (>300 µE·m−2·s−1) at 28°C and 7 h of darkness at 17°C.
Proteinase Inhibitor Assay. Proteinase inhibitors were quantified by radial immunodiffusion as described (31, 32). Systemin was supplied to excised plants through their cut stems, and leaves were wounded as described (12). Thereafter, the plants were
This paper was presented at the National Academy of Sciences colloquium “Virulence and Defense in Host–Pathogen Interactions: Common Features Between Plants and Animals, ” held December 9–11, 1999, at the Arnold and Mabel Beckman Center in Irvine, CA.
Abbreviations: MBP, myelin basic protein; MAPK, mitogen-activated protein kinase; pmg elicitor, β-glucan elicitor from Phytophtora megasperma f. subsp. glycinea.
|
* |
To whom reprint requests should be addressed. |
placed in sealed Plexiglas boxes as described and incubated under constant light at >300 µE·m−2·s−1 for 24 h at 28°C (12).
Suspension-Cultured Cells. Suspension-cultured cells of L. peruvianum (cells kindly provided by A. Schaller; ref.9) were cultivated under constant light as described (6). Cells were subcultured weekly and used for experiments 4–8 days after subculturing. For the experiments, 1.5 ml of cell suspension cultures were transferred to multiwell plates that were vigorously shaken on an orbital shaker under ambient room light and temperature conditions. After an ≈1-h equilibration period, the pH of the medium had reached a starting pH of 4.8 ± 0.2.
Alkalinization Assay. Test compounds were supplied to 1.5 ml of cell suspension cultures at the times indicated in the text. The pH of the medium was measured with a pH meter (Orion, Model EA940; Beverly, MA) with a semimicro combination electrode (Orion 8103BN, Beverly, MA). Addition of systemin or oligosaccharide elicitors to the cultured cells causes an alkalinization of the medium within 1–4 min. Over the next 6–15 min, the pH increases by about 0.6–1.2 pH units, remains at the high pH for 5 –10 min, and thereafter decreases. The pH in untreated controls consistently increased less than 0.3 pH units over the course of an experiment. Changes in the pH of the medium are indicated as “∆ pH” above the controls. Aqueous solutions of systemin and chitosan were prepared as described (12). The β-glucan elicitor from Phytophtora megasperma f. subsp. glycinea (pmg elicitor; partially purified according to Ayers et al., ref.33) was a generous gift of E. Farmer (University of Lausanne, Lausanne, Switzerland).
Myelin Basic Protein (MBP) Kinase Assay. At the times indicated, cells were mixed with extraction buffer (12) and frozen in liquid nitrogen. Extracts were obtained by sonicating the cells twice for 20 s (Model 300 Sonic Dismembrator, microtip; Fisher) and subsequent centrifugation at 30,000 × g. Supernatants were analyzed for MBP kinase activity by an in-gel kinase assay with MBP (Sigma) as an artificial substrate as described (12).
Immunocomplex Kinase Assay. Phosphotyrosine-containing proteins in cell extracts were immunocomplexed with the monoclonal phosphotyrosine-specific antibody PY20 coupled to Sepharose 4B (Stratagene) and assayed for MBP kinase activity as described (34).
Radioreceptor Assay. The binding of the systemin derivative 125I-Tyr-2,Ala-15-systemin to cell-surface binding sites was performed as described (6). Suramin was applied 1–10 min before addition of 125I-Tyr-2,Ala-15-systemin.
Photoaffinity Labeling. Photoaffinity labeling of the systemin receptor was performed in the presence or absence of competing systemin (200-fold excess; 150 pmol) or suramin (1 mM). Preparation of the analog 125I-N-(4-[p-azidosalicylamido]butyl)-3′(2′-Cys-3, Ala-15-systemindithiol)propionamide and photoaffinity labeling were performed as described (6). After photocrosslinking, microsomal membranes were prepared by adding the cells to 250 µl of extraction buffer containing 0.5 M D-sorbitol, 50 mM Mops, 10 mM Tris·HCl, 10 mM EGTA, 2.5 mM potassium metabisulfite, 2.0 mM salicylhydroxamic acid, 1 mM PMSF, plant protease inhibitor mixture (diluted 1/100; Sigma), and 1% β-mercaptoethanol, pH 7.6 (KOH). The membranes were extracted by sonication (Model 300 Sonic Dismembrator) for 1 min on ice. After sonication, 1 ml of extraction buffer was added, and the extract was vortexed and centrifuged at 12,000 × g for 5 min at room temperature. A 1-ml volume of the supernatant was removed and added to 9 ml of extraction buffer. The 10-ml extracts were ultracentrifuged for 1 h at 100,000 × g, after which the supernatant was removed and the pellets were gently rinsed with 2 ml of 10 mM Tris/1% sodium-dodecylsulfate, pH 8.0. The pellets were resuspended in 100 µl of the same buffer, and 30 µl was analyzed by SDS/7.5% PAGE and subsequent phosphorimaging of the dried gel.
Table 1. EC50s for medium alkalinization in response to elicitors
|
EC50 |
||
|
Elicitor |
−SUR |
+SUR |
|
Systemin, pg/ml |
360 |
9,600 |
|
Chitosan, ng/ml |
2 |
200 |
|
pmg elicitor, µg/ml |
22 |
250 |
|
EC50, elicitor concentration for half-maximal medium alkalinization ofsuspension-cultured tomato cells in the absence (−SUR) and presence(+SUR) of suramin. EC50s were calculated based on Fig. 1. |
||
Results
Proteinase Inhibitor Synthesis in Response to Wounding and Systemin Is Inhibited by Suramin. Suramin had been shown to interfere with binding of several animal polypeptide cytokines and growth factors to their plasma-membrane receptors (see the introduction). Because the 18-amino acid wound signal systemin is perceived by a membrane-bound receptor, we investigated whether suramin would inhibit the systemin-induced defense response in 14-day-old tomato plants. Supplying young excised plants simultaneously with 2.5 nM systemin and 700 µM suramin inhibited systemin-induced synthesis of serine proteinase inhibitors I and II in leaves by 95 ± 5% and 75 ± 18%, respectively. Synthesis of the two inhibitors in leaves of excised plants in response to wounding was 100% inhibited when the plants had been pretreated for 1 h with 700 µM suramin, a level consistent with its inhibition of animal growth factor–cell-surface receptor interactions.
Alkalinization of L. peruvianum Cell Medium in Response to Systemin, Chitosan, and pmg Elicitor Is Inhibited by Suramin. To study suramin effects on early cellular responses related to the synthesis of defense-related proteins, suspension-cultured cells of L. peruvianum were used. Previous studies had shown that elicitors of plant defense responses cause a rapid potassium ion efflux and proton influx leading to alkalinization of the culture medium of suspension-cultured cells through inhibition of the plasma membrane H+-ATPase (9, 10, 35). The pH of the medium of cultured tomato cells had been shown to increase about 1 pH unit within 10 min in response to systemin and to remain at the higher pH for about 25 min (10). The EC50 for alkalinization of tomato suspension-cultured cells induced by systemin is shown in Table 1, compared with the alkalinization of the medium by the oligosaccharide elicitors chitosan (a glucosamine oligomer) and pmg elicitor (a β-glucan mixture derived from mycelial walls of P. megasperma f. subsp. glycinea; ref.33). The EC50 for the alkalinization response induced by pmg elicitor varied considerably from the EC50s of systemin and chitosan, but the pmg elicitor was purified only partially, whereas systemin and chitosan had been highly purified.
The effect of suramin on alkalinization of the medium in response to increasing concentrations of systemin, chitosan, and pmg elicitor was assayed by pretreating the cells with a constant suramin concentration of 700 µM for 10 min before the application of elicitors (Fig. 1). The alkalinization response to lower elicitor concentrations was inhibited by suramin, resulting in an increase in the EC50s (Table 1). In the presence of high concentrations of elicitor, the suramin inhibition was reversed (Fig. 1). The reversible inhibition of the medium alkalinization in response to all three elicitors suggested that suramin competes with elicitors for extracellular elicitor-binding sites.
With each elicitor, the alkalinization response was inhibited
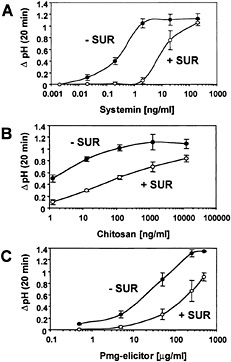
Fig. 1. Systemin-, chitosan-, and pmg elicitor-induced medium alkalinization is inhibited by suramin. Cells of L. peruvianum were left untreated (●, − SUR) or treated for 10 min with 700 µM suramin (○, + SUR). Subsequently, systemin (A), chitosan (B), or pmg elicitor (C) was supplied at the concentrations indicated. The pH of the medium was measured 20 min after addition of elicitors. The increase in medium pH (∆ pH) is indicated as the difference between the pH at time 0 and at 20 min.
severely by suramin in a concentration-dependent manner (Fig. 2). In these experiments, the data reported are for the time of maximal response for each elicitor in the absence of suramin: 7 min for systemin, 10 min for chitosan, and 20 min for pmg elicitor. The suramin concentrations for IC50 of systemin-, chitosan-, and pmg elicitor-induced medium alkalinization were similar (90 µM, 100 µM, and 130 µM, respectively). However, a pronounced difference between systemin and the carbohydrate elicitors was observed with regard to their maximal inhibition by suramin. Although a complete inhibition of the systemin response was reached at 350 µM suramin, the same suramin concentration resulted in only 70% inhibition of the response to both carbohydrate elicitors. The maximum inhibition (80%) of the alkalinization response caused by chitosan and pmg elicitor occurred at 1.5 mM suramin.
Suramin Interferes with Binding of Systemin to Its Cell-Surface Receptor. The reversible inhibition of the elicitor-induced alkalinization response suggested that suramin interfered with the binding of elicitors to their receptors, similar to its effect on animal growth factor receptors. To determine whether suramin was interacting with the systemin receptor, we used the radio-receptor assay and photoaffinity labeling assay that led to the original identification of the receptor (6). Fig. 3 shows that suramin interfered in a concentration-dependent manner with the specific binding of an iodinated bioactive systemin analog, 125I-Tyr-2,Ala-15-systemin, to cell-surface binding sites. The IC50 (160 µM) of suramin for systemin binding to the receptor is closely correlated to the IC50 for inhibition of medium alkalinization (90 µM; Fig. 3).
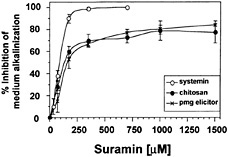
Fig. 2. Inhibition by suramin of medium alkalinization in response to systemin, chitosan, and pmg elicitor is concentration dependent. Cells of L. peruvianum were treated for 10 min with suramin at the concentrations indicated. The cells were then supplied with 1 nM systemin (○), 125 ng/ml chitosan (●), or 50 µg/ml pmg elicitor ( × ), and the pH change of the growth medium was determined after 7 min (systemin), 10 min (chitosan), or 20 min (pmg elicitor). The pH change is expressed as percentage of inhibition of medium alkalinization in the absence of suramin.
Suramin at a concentration of 1 mM also completely blocked the reaction of the systemin photoaffinity analog, 125I-N-(4-[p-azidosalicylamido]butyl)-3′(2′-Cys-3, Ala-15-systemindithiol)-propionamide, with the 160-kDa systemin receptor (Fig. 4).
Suramin Inhibits Tyrosine Phosphorylation of MBP Kinase in Response to Systemin, Chitosan, and pmg Elicitor. A rapid intracellular response to systemin in both suspension-cultured cells and in tomato plants is the activation of a 48-kDa MBP/MAPK (5, 12). In both assay systems, the activation by systemin could be blocked by the systemin analog Ala-17-systemin in which threonine at position 17 of the systemin sequence is replaced by an alanine residue (5, 12). Ala-17-systemin had also been shown to antagonize systemin binding to its receptor (5, 6). Thus, the activation of MBP kinase by systemin is likely to be mediated through the systemin receptor. Fig. 5A shows that a 48-kDa MBP kinase is activated not only by systemin but also by chitosan and pmg elicitor in suspension-cultured cells of L. peruvianum. All three responses were strongly inhibited in the presence of 1 mM suramin. The kinase activation was associated with tyrosine phosphorylation of the MBP kinase, which is a characteristic feature of MAPKs. Accordingly, in the presence of suramin, no tyrosine phosphorylation of the 48-kDa MBP kinase induced by systemin and carbohydrate elicitors was detectable (Fig. 5B).
Specific Structural Features Determine the Inhibitory Activity of Suramin. The inhibiting effect of suramin on responses triggered by different elicitor classes seems to be due to an unspecific binding of suramin to a diverse array of cell-surface components. Because suramin is a highly negatively charged molecule, the negative charges likely participate in ionic interactions with cell-surface components. To determine whether structurally similar molecules would function like suramin, heparin, a gly-
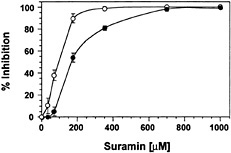
Fig. 3. Suramin blocks binding of 125I-Tyr-2,Ala-15-systemin to its cell-surface binding site. Cells of L. peruvianum were treated for ≤1 min with suramin at the concentrations indicated. The cells were then supplied with 1 nM 125I-Tyr-2,Ala-15-systemin with or without a 200-fold excess of unlabeled systemin. After 7 min, the cells were washed, and the specific binding of systemin to the cells was determined by subtracting total binding from unspecific binding (●). The data are expressed as percentage of inhibition of specific binding in the absence of suramin. For comparison, the effect of suramin on medium alkalinization in response to systemin is shown (○).
cosaminoglycan of the animal extracellular matrix that possesses sulfonic acid groups similar to those of suramin, and a suramin derivative, NF 007, were assayed for the inhibition of elicitor responses. Suspension-cultured cells were treated with heparin at a concentration of 1,000 µg/ml and were treated 20 min later with systemin, chitosan, or pmg elicitor or left untreated. Heparin alone did not induce the alkalinization response, nor did it affect the systemin-induced response. However, heparin did inhibit medium alkalinization moderately (≈25%) in response to carbohydrate elicitors (Table 2).
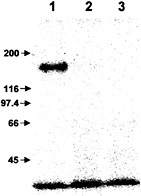
Fig. 4. Suramin interferes with photoaffinity labeling of the systemin receptor. 125I-N-(4-[p-azidosalicylamido]butyl)-3′(2′-Cys-3,Ala-15-systemindithiol)propionamide was incubated with suspension-cultured cells in the absence (lane 1) or presence (lane 2) of 150 pmol unlabeled systemin or in the presence of 1 mM suramin (lane 3). After photoactivation of the crosslinker, microsomal membranes were prepared, and membrane proteins were separated by SDS/PAGE. The radiolabeled receptor protein was visualized by phosphorimaging. Numbers and arrows at left indicate the position and molecular mass (in kDa) of protein standards.
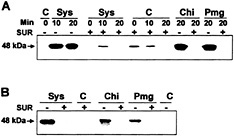
Fig. 5. Systemin, chitosan, and pmg elicitor activation of 48-kDa MBP kinase is inhibited by suramin. (A) MBP kinase activity. Cells of L. peruvianum were pretreated for 5 min with 1 mM suramin (SUR +) or left untreated (SUR −). They were then supplied with 3.3 nM systemin (Sys), 0.5 µg/ml chitosan (Chi), 50 µg/ml pmg elicitor (Pmg), or water (C). At the times indicated after addition of the elicitors, cells were frozen, extracted, and assayed for MBP kinase activity with an in-gel kinase assay. (B) Immunocomplex kinase assay. Phosphotyrosine-containing proteins in the extracts of samples (20 min) shown in A were complexed with a monoclonal phosphotyrosine-specific antibody coupled to Sepharose 4B and washed. Immunocomplexes were then analyzed for MBP kinase activity with an in-gel kinase assay.
The suramin-derivative NF 007 [8-(3-nitrobenzamido)-naphtalenetrisulfonic acid], at a concentration of 670 µg/ml, induced medium alkalinization (0.46 ± 0.03 pH units within 10 min after application) and was not studied further as a possible inhibitor of systemin action.
Suramin Does Not Block Medium Alkalinization Induced by Calyculin A and Erythrosin B. Two inhibitors of enzymes involved in intracellular signaling events in plants and animals, calyculin A (Cal), a protein phosphatase inhibitor, and erythrosin B, a membranepermeable inhibitor of the PM H+-ATPase, are known to induce medium alkalinization of suspension-cultured tomato cells rapidly. These compounds are presumed to enter the plant cells where they inhibit specific enzymes that are involved in regulating the signaling cascade that leads to inhibition of plasma-membrane H+-ATPase activity and medium alkalinization (9). No inhibition of the medium alkalinization induced by either compound was detected at a suramin concentration of 1,000 µg/ml (Fig. 6).
Discussion
The signaling pathway for the activation of defensive genes of tomato leaves in response to herbivorous insects and pathogens shows striking analogies to the animal inflammatory response signaling pathway ( Fig. 7; ref.3). For example, systemin is released in tomato leaves in response to herbivore attacks or other severe wounding; it is processed from a larger precursor by proteolytic cleavage (36); and its action is mediated by a cell-surface membrane-bound receptor (5, 6). The signaling pathways for systemin and proinflammatory factors like tumor necrosis factor-α include signaling steps such as activation of MAPKs, release of calcium from intracellular stores, and the activation of a phospholipase A2 (19, 20, 37). In animal cells, these events result in the release of arachidonic acid (20:4) from membranes and the production of prostaglandins. In tomato leaves, the activation of a phospholipase A2 releases linolenic acid (18:3) from membranes leading to the production of jasmonic acid, a prostaglandin analog (refs.3, 13, 38, and 39; Fig. 7).
Many animal growth factor receptor interactions and subsequent cellular responses can be inhibited by suramin, a polyanionic naphtylurea compound (see the introduction). This fact prompted us to determine whether suramin would also inhibit systemin receptor interactions. An extracellular inhibitor of
Table 2. Inhibition of medium alkalinization in response to elicitors in the presence of suramin or heparin
|
Percentage of inhibition of medium alkalinization |
||
|
Elicitor |
+Suramin |
+Heparin |
|
Systemin |
100 ± 0 |
1 ± 1 |
|
Chitosan |
73 ± 5 |
28 ± 8 |
|
pmg elicitor |
75 ± 5 |
23 ± 6 |
|
Cells of L. peruvianum were left untreated or supplied for 10 min with 1,000 µg/ml ( = 700 µM) suramin or 1,000 µg/ml heparin and then with 1 nM systemin, 50 µg/ml pmg elicitor, or 125 ng/ml chitosan. The pH change of the growthmedium was determined 20 min after addition of elicitors and expressedas percentage of inhibition of medium alkalinization. The responseto the elicitors alone was referred to as 0% inhibition. |
||
receptor-mediated signaling in plants might also provide a tool to characterize more thoroughly the interaction of other defense signals, such as oligosaccharide elicitors, with their cell-surface receptors. An understanding of the inhibitory mechanism could help to clarify the possible relationships between polypeptide-regulated signaling systems in plants and animals.
We initially found that suramin blocked both the wound- and systemin-induced synthesis of proteinase inhibitor proteins in leaves of excised tomato plants (see Results). Based on the known suramin effects on animal cells, this finding suggested that suramin may be interfering with the perception of the defense signal systemin. To characterize the effects of suramin on systemin receptor-mediated defense responses, we used suspension-cultured cells of L. peruvianum. These cells responded to systemin, chitosan, and pmg elicitor with an initiation of a rapid influx of protons, resulting in a pronounced medium alkalinization. This response was inhibited by suramin (Fig. 1 and Fig. 2). The half-maximal suramin concentrations (IC50s) for the inhibition of medium alkalinization in response to systemin and the two oligosaccharide elicitors were similar. However, although suramin inhibited the systemin-induced alkalinization 100%, it inhibited the carbohydrate elicitor induction by only about 80% (Fig. 2). This disparity apparently reflects the differences among the specific receptors. The IC50s of suramin calculated for inhibition of the alkalinization response induced by all three elicitors are very similar to its IC50s for inhibition of animal cytokine and growth factor binding to the respective receptors (21, 23, 25).
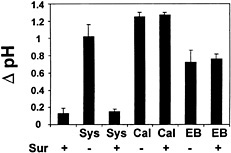
Fig. 6. Effect of suramin on medium alkalinization induced by systemin, calyculin A, and erythrosin B. Cells of L. peruvianum were either left untreated (Sur −) or treated with 1,000 µM suramin (Sur +) for 5 min. Thereafter, 3.3 nM systemin (Sys), 2 µM calyculin A (Cal), or 20 µM erythrosin B (EB) was added to the cells. The pH of the medium was measured 10 min after addition of elicitors, and the increase in medium pH (∆ pH) was determined as described in Fig. 2.
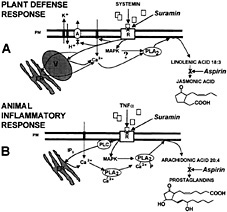
Fig. 7. Analogies between animal and plant defense-signaling pathways. ( A) Early signaling events of the tomato wound response (47). (B) Early signaling events of the animal inflammatory response (20). A, plasma membrane proton ATPase; ER, endoplasmic reticulum; IP 3, inositoltriphosphate; PLA2, phospholipase A2; PLC, phospholipase C; PM, plasma membrane; R, receptor; TNFα, tumor necrosis factor-α; V, vacuole.
The inhibition of the elicitor-induced medium alkalinization by suramin was reversible (Fig. 1). This result strongly indicated that the suramin effect is extracellular and that suramin interacts with elicitor binding sites in a reversible manner. To provide further evidence that suramin may compete with binding of systemin to the systemin receptor on the cell surface, we measured the specific binding of an iodinated bioactive systemin derivative to its receptor in the presence and absence of suramin. In a radioreceptor assay, the specific binding of iodinated systemin to its receptor was inhibited by suramin in a concentration-dependent manner (Fig. 3). The IC50 for the inhibition of specific systemin binding (160 µM) was near the IC50 for inhibition of the alkalinization response (90 µM). Furthermore, suramin totally prevented the labeling of the systemin receptor by a systemin-photoaffinity analog (Fig. 4). This result provides strong evidence that suramin inhibits systemin signaling by interfering with the binding of systemin to the systemin receptor. Cochromatography of the iodinated systemin derivative with suramin revealed that suramin does not bind to systemin (data not shown). Suramin most likely directly binds to the systemin receptor, thereby preventing its activation by systemin. The reversible inhibition by suramin of both systemin-induced and carbohydrate elicitor-induced medium alkalinization (Fig. 1) suggested that suramin may be interfering with the respective receptors in a manner similar to its effect on the binding of animal cytokines and growth factors to their respective receptors.
Another early signaling event in response to systemin and carbohydrate elicitors is the activation of a 48-kDa MBP kinase (Fig. 5). The MBP kinase activation was associated with tyrosine phosphorylation, indicating that this kinase belongs to the MAPK family of protein kinases (Fig. 5). MAPKs participate in various animal and plant stress-induced signaling pathways (19, 40). It had been shown previously that the activation by systemin of a 48-kDa MAPK in leaves of tomato plants and in cells of L.
peruvianum is a systemin receptor-mediated process (5, 12). The MAPK activation in the cells of L. peruvianum in response to systemin, chitosan, and pmg elicitor was inhibited by suramin (Fig. 5), thereby supporting the role of suramin as an inhibitor of receptor function.
Heparin, a glycosaminoglycan of the animal extracellular matrix with sulfonic acid groups such as found in suramin, and suramin both were reported to inhibit binding of 125I-basic fibroblast growth factor to its binding sites (27). However, heparin had no effect on the medium alkalinization of suspension-cultured tomato cells in response to systemin (Table 2). Thus, the negative charges alone cannot account for the suramin effect on the systemin receptor. The alkalinization of cell medium by carbohydrate elicitors was, however, partially inhibited by heparin (Table 2). This inhibition may reflect different structural features of the different binding sites that can interact with suramin and heparin. NF 007, 8-(3-Nitrobenzamido)-naphtalenetrisulfonic acid, another sulfonic acid-containing molecule, is a derivative of suramin. Unlike suramin, NF 007 interacts with cells of L. peruvianum as an inducer rather than an inhibitor of medium alkalinization. Thus, common structural features of suramin and NF 007 might be important for binding to an elicitor binding site, but additional determinants are required for suramin to act as an inhibitor. These properties together with the low IC50s of suramin observed for the inhibition of the alkalinization response, induced by all three elicitors, suggest that suramin may recognize some characteristic structural features that might be common among the different elicitor receptors. The identity of sites involved in suramin interactions with the membrane-associated systemin and elicitor receptors could provide more specific information concerning the possible structural relationship between plant receptors and perhaps between plant and animal receptors as well.
Based on its polyanionic hydrophilic character, it has been assumed that suramin is not able to pass the membrane barrier and enter the cell where it can interfere with intracellular signaling processes, e.g., through inhibition of tyrosine phosphatases, G proteins, and DNA polymerases (41, 42 and 43). However, in some animal cell systems, suramin seems to enter cells via endocytosis (44). Because endocytotic mechanisms are also known in plants (45), it was important to determine whether suramin can pass the plasma membrane barrier and interfere with intracellular signaling processes leading to the alkalinization response. Calyculin A, a protein phosphatase 1 and 2a inhibitor, and erythrosin B, a membrane-permeable inhibitor of the plasma membrane H+-ATPase, both activate the alkalinization response in suspension-cultured tomato cells (9, 46). The roles of these inhibitors in activating the defense response is not known, but it is likely that they inhibit enzymes that are negative regulators of the response (9). Suramin did not inhibit calyculin A- or erythrosin B-induced alkalinization responses (Fig. 6). These data indicated that suramin acts upstream of the targets of the inhibitors, supporting its role in interfering extracellularly with binding of systemin and oligosaccharide elicitors to their receptors.
Taken together, the results of this study further support a role for the receptors of systemin, chitosan, and pmg elicitor in activating the intracellular events leading to defense gene activation in response to herbivores and pathogens. Suramin provides a tool to investigate receptor-mediated defense responses in plant cells. It may prove to be useful in elucidating the early events in plant defense signaling and in exploring the analogies between signaling pathways of plants and animals further (Fig. 7).
We thank Sue Vogtman and Tom Koehler for growing and maintaining plants. This research was supported in part by Washington State University College of Agriculture and Home Economics (Project No. 1791), the National Science Foundation (Grant No. IBN 9601099), and the United States Department of Agriculture Competitive Grants Program (Grant No. 9801502).
1. Pearce, G., Strydom, D., Johnson, S. & Ryan, C. A. ( 1991) Science 253, 895–898.
2. Schaller, A. & Ryan, C. A. ( 1996) BioEssays 18, 27–33.
3. Bergey, D. R., Howe, G. & Ryan, C. A. ( 1996) Proc. Natl. Acad. Sci. USA 93, 12053–12058.
4. Ryan, C. A. & Pearce. G. ( 1998) Annu. Rev. Cell Dev. Biol. 14, 1–17.
5. Meindl, T., Boller, T. & Felix, G. ( 1998) Plant Cell 10, 1561–1570.
6. Scheer, J. & Ryan, C. A. ( 1999) Plant Cell 11, 1525–1535.
7. Moyen, C. & Johannes, E. ( 1996) Plant Cell Environ. 19, 464–470.
8. Moyen, C., Hammond-Kosack, K. E., Jones, J., Knight, M. R. & Johannes, E. ( 1998) Plant Cell Environ. 21, 1101–1111.
9. Schaller, A. & Oecking, C. ( 1999) Plant Cell 11, 263–272.
10. Felix, G. & Boller, T. ( 1995) Plant J. 7, 381–389.
11. Bergey, D. R. & Ryan, C. A. ( 1999) Plant Mol. Biol. 40, 815–823.
12. Stratmann, J. W. & Ryan, C. A. ( 1997) Proc. Natl. Acad. Sci. USA 94, 11085–11089.
13. Narváez-Vásquez, J., Florin-Christensen, J. & Ryan, C. A. ( 1999) Plant Cell 11, 2249–2260.
14. Walker-Simmons, M. & Ryan, C. A. ( 1984) Plant Physiol. 76, 787–790.
15. Doares, S. H., Syrovets, T., Weiler, E. W. & Ryan, C. A. ( 1995) Proc. Natl. Acad. Sci. USA 92, 4095–4098.
16. Howe, G. A., Lightner, J., Browse, J. & Ryan, C. A. ( 1996) Plant Cell 8, 2067–2077.
17. Felix, G., Regenass, M. & Boller, T. ( 1993) Plant J. 4, 307–316.
18. Felix, G., Grosskopf, D. G., Regenass, M. & Boller, T. ( 1991) Proc. Natl. Acad. Sci. USA 88, 8831–8834.
19. Kyriakis, J. M. & Avruch, J. ( 1996) J. Biol. Chem. 271, 24313–24316.
20. Lin, L.-L., Wartmann, M., Lin, A. Y., Knopf, J. F., Seth, A. & Davis, R. J. ( 1993) Cell 72, 269–278.
21. Betsholtz, C., Johnsson, A., Heldin, C.-H. & Westermark, B. ( 1986) Proc. Natl. Acad. Sci. USA 83, 6440–6444.
22. Coffey, R. J., Leof, E. B., Shipley, G. D. & Moses, H. L. ( 1987) J. Cell. Physiol. 132, 143–148.
23. Kopp, R. & Pfeiffer, A. ( 1990) Cancer Res. 50, 6490–6496.
24. Williams, L. T., Tremble, P. M., Lavin, M. F. & Sunday, M. E. ( 1984) J. Biol. Chem. 259, 5287–5294.
25. Hosang, M. ( 1985) J. Cell. Biochem. 29, 265–273.
26. Roghani, M., Mohammadi, M, Schlessinger, J. & Moscatelli, D. ( 1996) J. Biol. Chem. 271, 31154–31159.
27. Bono, F., Rigon, P., Lamarche, I., Savi, P., Salel, V. & Herbert, J. M. ( 1997) Biochem. J. 326, 661–668.
28. Leland, P., Obiri, N., Aggarwal, B. B. & Puri, R. K. ( 1995) Oncol. Res. 7, 227–235.
29. Mills, G. B., Zhang, N., May, C., Hill, M. & Chung, A. ( 1990) Cancer Res. 50, 3036–3042.
30. Alzani, R., Corti, A., Grazioli, L., Cozzi, E., Ghezzi, P. & Marcucci, F. ( 1993) J. Biol. Chem. 268, 12526–12529.
31. Ryan, C. A. ( 1967) Anal. Biochem. 19, 434–440.
32. Trautman, R., Cowan, K. M. & Wagner, G. G. ( 1971) Immunochemistry 8, 901–916.
33. Ayers, A. R., Ebel, J., Valent, B. & Albersheim, P. ( 1976) Plant Physiol. 57, 760–776.
34. Stratmann, J. W., Stelmach, B., Weiler, E. & Ryan, C. A. ( 2000) Photochem. Photobiol. 71, 116–123.
35. Mathieu, Y., Guern, J., Spiro, M. D., O'Neill, M. A., Kates, K., Darvill, A. G. & Albersheim, P. ( 1998) Plant J. 16, 305–311.
36. McGurl, B., Pearce, G., Orozco-Cardenas, M. L. & Ryan, C. A. ( 1992) Science 255, 1570–1573.
37. Leslie, C. C. ( 1997) J. Biol. Chem. 272, 16709–16712.
38. Hamberg, M. & Gardner, H. W. ( 1992) Biochim. Biophys. Acta 1165, 1–18.
39. Farmer, E. E. & Ryan, C. A. ( 1992) Plant Cell 4, 129–134.
40. Hirt, H. ( 1997) Trends Plant Sci. 2, 11–15.
41. Zhang, Y.-L., Keng, Y.-F., Zhao, Y., Wu, L. & Zhang, Z.-Y. ( 1998) J. Biol. Chem. 273, 12281–12287.
42. Hohenegger, M., Waldhoer, M., Beindl, W., Böing, B., Kreimeyer, A., Nickel, P., Nanoff, C. & Freissmuth, M. ( 1998) Proc. Natl. Acad. Sci. USA 95, 346–351.
43. Spigelman, Z., Dowers, A., Kennedy, S., DiSorbo, D., O'Brien, M., Barr, R. & McCaffrey, R. ( 1987) Cancer Res. 47, 4694–4698.
44. Huang, S. S. & Huang, J. S. ( 1988) J. Biol. Chem. 263, 12608–12618.
45. Battey, N. H., James, N. C., Greenland, A. J. & Brownlee, C. ( 1999) Plant Cell 11, 643–659.
46. Felix, G., Regenass, M., Spanu, P. & Boller, T. ( 1994) Proc. Natl. Acad. Sci. USA 91, 952–956.
47. Ryan, C. A. ( 2000) Biochem. Biophys. Acta 1477, 112–121.






