Colloquium
Phenotypic variation and intracellular parasitism by Histoplasma capsulatum
Silke Kügler*, Tricia Schurtz Sebghati*, Linda Groppe Eissenberg*, and William E. Goldman†
Department of Molecular Microbiology, Washington University School of Medicine, St. Louis, MO 63110
The success of Histoplasma capsulatum as an intracellular pathogen depends completely on successful conversion of the saprophytic mycelial (mold) form of this fungus to a parasitic yeast form. It is therefore not surprising that yeast phase-specific genes and gene products are proving to be important for survival and proliferation of H. capsulatum within macrophages. In this study, we have focused on the role and regulation of two yeast-specific characteristics: α-(1,3)-glucan, a cell wall polysaccharide modulated by cell-density (quorum) sensing, and a secreted calcium-binding protein (CBP) that is essential for pathogenicity.
H istoplasma capsulatum is the best studied of the dimorphic fungal pathogens, all of which undergo a reversible morphological variation that is tightly linked to their lifestyle and pathogenesis. The normal home for these fungi is the soil, where they exist in a mycelial (mold) form. Humans and other mammals inhale aerosolized conidia and hyphal fragments, prompting a dramatic conversion of the mycelial form to budding yeasts. The phenotypic variation between mycelial and yeast forms corresponds to a complete switch in lifestyle, from a saprophytic soil-dwelling fungus to a parasitic form that is highly adapted for growth at higher temperature (37°C) and for avoidance of host defense mechanisms. In fact, H. capsulatum thrives within the normally harsh environment of the phagolysosomes of macrophages (1), the primary host cell type. In this regard, this yeast is unique from other intracellular pathogens in its ability to prevent the acidification of the phagolysosome (2), presumably limiting the antifungal effectiveness of this compartment.
Little else is known about how H. capsulatum survives and proliferates inside macrophages, but clues to these strategies have come from studying characteristics that are specific to the yeast form. Because the yeast cell is functionally dedicated to intracellular parasitism, it is likely that identification and characterization of genes that are exclusively expressed in the yeast phase will lead to a better understanding of H. capsulatum virulence. Our laboratory has focused on two yeast phase-specific characteristics: an unusual cell wall polysaccharide that is intimately linked to strain-specific pathogenicity and a secreted calcium-binding protein (CBP) that has become, to our knowledge, the first formally defined virulence determinant of H. capsulatum.
Modulation of α-(1,3)-Glucan in the Cell Wall
Many virulent strains of H. capsulatum possess α-(1,3)-glucan in the cell wall of the yeast form, although it is absent in the mycelial form. This polysaccharide is likewise present in the yeast phase of two other pathogenic dimorphic fungi, Paracoccidiodes brasiliensis (3) and Blastomyces dermatitidis (4). In each of these species, spontaneous variants that have lost their α-(1,3)-glucan have also lost virulence (3, 5, 6). Whether this polysaccharide directly influences virulence is unknown; its absence may simply alter cell wall architecture, permeability, or secretion. Regardless, its presence correlates with strain-specific virulence in a mouse model of infection and influences the dynamics of intracellular survival and proliferation (5, 7).
Strains of H. capsulatum regulate α-(1,3)-glucan production such that it seems to be constitutively synthesized when the yeasts are proliferating inside macrophages. This regulation is in sharp contrast to the modulation of this phenotype during broth culture growth. When yeasts from a dense culture were washed and then diluted to a low density in standard growth medium, only an estimated 30% of the yeasts had α-(1,3)-glucan detectable in their cell walls 24 h later (Fig. 1). The percentage remains low until the culture begins to enter stationary phase, when α-(1,3)-glucan is once again detectable in the wall of virtually every yeast cell.
This phenotypic variation is a consistent feature of broth culture growth, with one important exception. When yeasts from a dense culture were diluted into medium that contained filtrate from a stationary phase culture, most yeasts remained positive for α-(1,3)-glucan 24 h later (data not shown). They tested positive even when the filtrate comprised as little as 4% of the new medium, negating the possibility that nutrient deprivation was responsible for blocking this modulation. These results suggest that H. capsulatum yeast cells release a factor that, when present in sufficient concentration, promotes α-(1,3)-glucan incorporation into the cell wall.
Growth-dependent modulation of α-(1,3)-glucan production resembles the “quorum-sensing” phenomenon seen in many bacteria (8, 9 and 10). In those systems, microbes release an “autoinducer” at a constant rate, such that its concentration in the external environment is directly proportional to the cell density. Eventually, a sensor molecule detects a critical concentration of the autoinducer and signals a transcriptionally regulated phenotypic variation. Among bacteria, the autoinducer is usually an acyl-homoserine lactone ( 10) or a peptide (9), small mediators that are diffusible across membranes. Our initial characterizations suggest that the cell density autoinducer released by H. capsulatum is a larger molecule (molecular weight > 6,000, based on dialysis experiments) and therefore unlikely to diffuse across membranes. It is tempting to speculate about a regulatory role for this system during intracellular parasitism; a high intraphagosomal concentration of the autoinducer could signal α-(1,3)-glucan synthesis to remain active, similar to the regulation in vitro when broth cultures become dense. Histoplasma may therefore be the first characterized example of an organism that uses a quorum-sensing mechanism to detect that it is within a host cell compartment.
Calcium and Intracellular Parasitism
Another Histoplasma yeast phase-specific phenotype is the production of CBP, a secreted calcium-binding protein. Se-
This paper was presented at the National Academy of Sciences colloquium “Virulence and Defense in Host–Pathogen Interactions: Common Features Between Plants and Animals, ” held December 9–11, 1999, at the Arnold and Mabel Beckman Center in Irvine, CA.
|
* |
S.K., T.S.S., and L.G.E. contributed equally to this work. |
|
† |
To whom reprint requests should be addressed. E-mail: goldman@borcim.wustl.edu. |
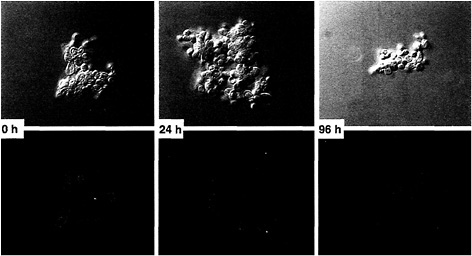
Fig. 1. Growth-dependent modulation of α-(1,3)-glucan in H. capsulatum yeast cell walls. H. capsulatum G186AR yeasts were washed and used as an inoculum (Left) for a culture initiated at low density (2 × 106 cells per ml). At 24 h (Center) and 96 h (Right) after inoculation, an aliquot of yeasts was removed and monitored for α-(1,3)-glucan by comparing immunofluorescence (Upper) and differential interference contrast (Lower) images (magnification ×1,700). Murine monoclonal IgM antibody MOPC104E was used to detect α-(1,3)-glucan, and FITC-tagged goat anti-murine IgM was used as a fluorescent secondary antibody.
cretion of CBP is correlated not only with yeast/mycelial morphology but also with dependence on calcium for growth; H. capsulatum in the yeast phase is capable of growing in the presence of high levels of EGTA, whereas mycelial growth in limiting calcium is inhibited ( 11). This dependence may reflect a general problem faced by intracellular pathogens, because both Salmonella and Toxoplasma have specific responses to low calcium conditions that are duplicated during host cell parasitism ( 12, 13). Still, defining the role of CBP in Histoplasma virulence will depend on more than correlative and circumstantial evidence, and our efforts have focused on developing a formal molecular genetic proof.
In many fungi, the ability to evaluate protein function by gene disruption strategies is complicated by “illegitimate” recombination events. Simple allelic replacement is therefore difficult to detect among the high background of ectopic insertions, and marker-based selection strategies become complicated by duplications, rearrangements, and deletions that can accompany the desired recombination event. For molecular genetics in H. capsulatum, we have developed a range of strategies that are based on transformation with linear telomeric plasmids (14, 15). This genetic system has been adapted most recently as an allelic replacement tool, allowing us to disrupt the CBP1 locus in H. capsulatum and test its role in calcium acquisition and virulence.
To disrupt CBP1 in a virulent strain of H. capsulatum (G186AR), a linear telomeric plasmid was designed with several unique features to enrich for homologous recombination events. Most importantly, inverted telomeric repeats at each end of the linear plasmid help maintain the plasmid extrachromosomally and nearly eliminate ectopic integration events. For selection and disruption, the linear plasmid contains a URA5 gene, and the CBP1 gene has an internal fragment replaced by a hygromycin resistance marker (hph). In addition, over 5 kilobases of flanking DNA (upstream and downstream of the CBP1 coding sequence) was included to increase the frequency of the desired double crossover event. This construct was transformed into a uracil auxotroph of H. capsulatum G186AR, and transformants were selected initially as uracil prototrophs. Cultures were then grown in the presence of 5-fluoroorotic acid (selecting against URA5 on the plasmid vector) and hygromycin (selecting for retention of the disrupted CBP1 gene). After this two-step selection strategy, cbp1-null mutants were isolated at high frequency, and their genotypes were confirmed by PCR, Southern analysis, and protein gels (data not shown).
Two cbp1-null isolates were first tested for their ability to grow in the presence of EGTA. The growth of these knockout strains was inhibited in medium containing EGTA at a concentration as low as 150 µM (Fig. 2). (Normal growth medium used for Histoplasma contains 300 µM calcium.) In contrast, wild-type H. capsulatum strains were able to grow in medium containing greater than 1 mM EGTA. CBP1 expression was restored in one of the disrupted strains by transformation with another telomeric plasmid carrying wild-type CBP1. Complementing the knockout with CBP1 restores growth in calciumlimited medium (Fig. 2).
To determine the virulence of the CBP knockout strain, we evaluated its interaction with a macrophage-like cell line (P388D1 cells). The ability of yeast strains to kill P388D1 cells has been correlated previously with virulence as measured in a mouse model of histoplasmosis (7). When tested in this in vitro virulence model, the cbp1-null strain could be seen inside of the macrophages but was unable to destroy the macrophage monolayer. When the knockout strain was complemented with CBP1 in trans, the macrophages were killed as efficiently as they were by the wild-type virulent strain (Fig. 3).
These results describe the first successful disruption of a virulence determinant in H. capsulatum. The actual function of CBP in vivo still needs to be elucidated. CBP may act in a siderophore-like manner, binding calcium ions and providing
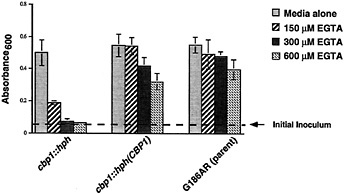
Fig. 2. CBP and calcium dependency of H. capsulatum growth. Cultures of H. capsulatum were grown for 4 days in normal medium or in medium with EGTA added to chelate calcium. Cumulative growth was measured (by absorbance at 600 nm) for the cbp1-null strain (cbp1::hph), the trans-complemented strain cbp1::hph(CBP1), and the wild-type parental strain G186AR. Each bar represents the mean for triplicate samples, with error bars indicating standard deviations from the mean. Growth inhibition by EGTA could be reversed by adding equimolar calcium chloride to the cultures (data not shown).
yeasts with calcium needed for growth. Alternatively, CBP may bind calcium to modulate the phagolysosomal environment such that yeast proliferation is possible.
Regulation of CBP
CBP1 is regulated at the level of transcription, and we have shown previously by Northern blot analysis that mRNA is present in the yeast phase but not detectable in the mycelial phase (16). Further studies with reverse transcription–PCR demonstrated that CBP1 continues to be expressed while H. capsulatum yeasts grow inside P388D1 cells. This expression is consistent with experiments with mice inoculated with H. capsulatum; splenocytes harvested later from these mice respond to purified CBP in proliferation assays, implying that CBP is secreted during infection of mammalian hosts (17). All of these regulatory studies have used either mycelial or yeast cultures of H. capsulatum, but there has been no means to examine carefully the developmental programming of CBP1 expression during the transition between these two forms. For this purpose, we have developed a reporter system with a synthetic gene encoding the green fluorescent protein (gfp). A promoter fusion was constructed, consisting of 1.3 kilobases of the 5′ untranslated region of CBP1, including the start codon ATG, fused to the gfp gene (0.7 kilobases). By using the telomeric plasmid system, constructs containing different modified gfp sequences were tested for fluorescence in H. capsulatum. One human codon-optimized version of gfp with the amino acid exchanges F64L, S65T, and H231L (18) yielded brightly fluorescent yeast cells. As expected, mycelial cells showed only very weak or no fluorescence, consistent with the CBP phenotype and with Northern blot analysis of CBP1 expression.
Fig. 3. Role of CBP in parasitism of macrophages. P388D1 macrophage-like cells were inoculated with strains of H. capsulatum at a multiplicity of one yeast per five macrophages and then cocultured for 5 days as described (7). (A) cbp1-null strain cbp1::hph; (B) trans-complemented strain cbp1::hph(CBP1); and (C) wild-type parental strain G186AR (magnification × 800).
To follow the regulation of CBP1 during the transition from yeast cells to mycelia, a similar plasmid construct was randomly integrated into the chromosome to stabilize gfp copy number from cell to cell. Expression of gfp was monitored by fluorescence microscopy after placing a yeast culture at 25°C to trigger transition to the mycelial form (Fig. 4). During the first 24 h, the yeast cells and their initial hyphal extensions remained brightly fluorescent. By the next day, fluorescence of the yeast cells was reduced, and the longer hyphae had almost no detectable fluorescence. At 72 h after the temperature shift, hardly any fluorescence was visible, although the residual fluorescence was localized only to yeast cells. These results demonstrate that CBP1 is not simply temperature regulated, because fluorescence from preexisting CBP-GFP would have been obviously reduced at 24 h (particularly in the hyphal extensions). Instead, it seems that CBP1 is developmentally
Fig. 4. Regulation of CBP1 during transition from yeast to mycelial forms. H. capsulatum strain G186ASura5 carrying the CBP1 promoter-gfp fusion was grown as yeast at 37°C and then transferred to 25°C at time 0 h. (A) Differential interference contrast microscopy, (B) Fluorescence microscopy (magnification ×1,200).
down-regulated as the morphological conversion to the mycelial form progresses.
Histoplasma as a Model System
Among the fungi, only a few primary pathogens cause systemic mycoses, and among those, H. capsulatum has received the most research attention in terms of biology, biochemistry, and molecular genetics. Historically, the mycelia–yeast transition and the ability of yeasts to destroy macrophages have been the areas of primary focus. With the development of molecular genetic tools such as freely replicating plasmids, reporter genes, and gene disruption strategies, it is now possible to probe this organism's fascinating biology with genetic precision and functional proof. It is also likely that many of the lessons learned from H. capsulatum will be applicable to the other dimorphic fungal pathogens, most of which are closely related and cause similar clinical syndromes. The existence of a morphological form dedicated to parasitism provides a highly visible target in the hunt for genes and regulatory mechanisms that are most likely to be involved in fungal pathogenesis.
This work was supported by Public Health Service Grants AI25584 (to W.E.G.), AI07172 (to Washington University), and HL07317 (to Washington University). W.E.G. is a recipient of the Burroughs–Wellcome Fund Scholar Award in Molecular Pathogenic Mycology.
1. Eissenberg, L. G., Schlesinger, P. H. & Goldman, W. E. ( 1988) J. Leukocyte Biol. 43, 483–491.
2. Eissenberg, L. G., Goldman, W. E. & Schlesinger, P. H. ( 1993) J. Exp. Med. 177, 1605–1611.
3. San-Blas, G., San-Blas, F. & Serrano, L. E. ( 1977) Infect. Immun. 15, 343–346.
4. Kanetsuna, F. & Carbonell, L. M. ( 1971) J. Bacteriol. 106, 946–948.
5. Klimpel, K. R. & Goldman, W. E. ( 1987) Infect. Immun. 55, 528–533.
6. Hogan, L. & Klein, B. ( 1994) Infect. Immun. 62, 3543–3546.
7. Eissenberg, L. G., West, J. L., Woods, J. P. & Goldman, W. E. ( 1991) Infect. Immun. 59, 1639–1646.
8. Greenberg, E. P. ( 1997) Am. Soc. Microbiol. News 63, 371–377.
9. Dunny, G. M. & Leonard, B. A. B. ( 1997) Annu. Rev. Microbiol. 51, 527–564.
10. Fuqua, C, Winans, S. C. & Greenberg, E. P. ( 1996) Annu. Rev. Microbiol. 50, 727–751.
11. Batanghari, J. W. & Goldman, W. E. ( 1997) Infect. Immun. 65, 5257–5261.
12. García Véscovi, E., Soncini, F. C. & Groisman, E. A. ( 1996) Cell 84, 165–174.
13. Sibley, L. D., Weidner, E. & Krahenbuhl, J. L. ( 1985) Nature ( London) 315, 416–419.
14. Woods, J. P. & Goldman, W. E. ( 1992) Mol. Microbiol. 6, 3603–3610.
15. Woods, J. P. & Goldman, W. E. ( 1993) J. Bacteriol. 175, 636–641.
16. Patel, J. B., Batanghari, J. W. & Goldman, W. E. ( 1998) J. Bacteriol. 180, 1786–1792.
17. Batanghari, J. W., Deepe, G. S., Jr., Di Cera, E. & Goldman, W. E. ( 1998) Mol. Microbiol. 27, 531–539.
18. Haas, J., Park, E.-C. & Seed, B. ( 1996) Curr. Biol. 6, 315–324.





