2
Detection of Potential Biohazards
Central to the topic of quarantining samples and certifying them for release from quarantine is the problem of detecting life in them, that is, determining if life is present or has recently been present. The problem is compounded by the possibility of life forms that function and reproduce in a manner outside the range of our experience with terrestrial life. Detection of evidence of past life in returned martian samples is discussed in Chapter 3; discovery of fossilized life in the samples would profoundly influence the search for extant life.
BASIC ASSUMPTIONS AND MEASUREMENTS
A search for evidence of life, assumed to be carbon-based and microbial,1 can take advantage of many recent advances in molecular biology and organic geochemistry. Analytical and investigative techniques used in these areas of research are advancing rapidly, and many new methods are likely to be in place by the time samples are returned from Mars. A series of tests can be envisioned that will provide evidence of viable or recently dead organisms, detect chemical fossils or probable biological molecules (biomarkers), and, at the same time, quantify contamination by terrestrial microbes and organic compounds. To avoid misinterpretation of “false positives,” it will be essential to detect and identify terrestrial contaminants, both microbial and organic, on the outbound spacecraft (particularly on surfaces that may come into contact with the returned samples) as well as in all areas within the quarantine facility in which returned samples are exposed to the environment (where the nature and amount of contamination should be carefully monitored during sample processing). However, although these techniques are effective for identifying terrestrial life or its products, signals arising from martian organisms or their products might differ substantially from those of their terrestrial counterparts.
Measurement of the total concentration of organic carbon2 in each martian subsample will provide an invaluable baseline. Samples in which organic carbon is below 10–12 gC/g are unlikely to contain microorganisms. Samples containing detectable amounts of organic carbon will be worthy of intensive—and cautious—study.
Some analytical techniques can utilize samples as small as 100 micrograms and have carbon-detection limits of about 10 nanograms (roughly equivalent to 3 million bacterial cells of minimum size).3 With more specific information about the material in the samples, lower detection limits are possible.4 Improvements can be confidently foreseen.5,6 Plausibly, the amount of sample required might be reduced to 10 micrograms while preserving sensitivity at the parts-per-million level. The resulting ability to detect 10–11 gC would probably represent the ultimate attainable sensitivity for organic carbon. Thus purely chemical tests could not equal the sensitivity of an optimally targeted bioassay, which might have the capability of detecting only a few viable cells containing in total 10–13 gC or less. However, the chemical analysis is general. It does not require provision of the specific nutrients and conditions a given organism might require for growth. Moreover, its efficacy would be enhanced because it would detect not only viable cells but also organic substrates being utilized by cells or debris resulting from the accumulation of dead cells. In spite of its limitations, analysis for organic carbon is probably the best available technique for screening and surveying samples.
DETECTING LIFE
The attempt to detect actual life, extant or recent, is clearly different from a search for the chemical traces left by life or for identifiable fossils (the topic of Chapter 3). Whether by molecular biological techniques or by cultivation in the laboratory, detection of terrestrial microorganisms is more difficult than is commonly appreciated. For example, the polymerase chain reaction (PCR—see the section “Detection by Analysis of Chemical and Isotopic Structures” below), a powerful molecular biological technique, is capable of detecting minute amounts of genetic material but, because of its extreme sensitivity, is also easily subject to error-causing contamination. Moreover, although positive results from techniques such as PCR sometimes provide evidence that biological activity may have occurred in a sample, cultivation is still the only means of demonstrating viability unequivocally. Even so, cultivation unfortunately is not a comprehensive solution to the problem of detection, because many terrestrial organisms that are clearly “alive” have never been successfully cultivated in the laboratory.7,8
Further, although it can be argued that existing technology (and likely future developments) will allow the detection of organic compounds and their identification even if they should be novel compounds unique to Mars, existing life detection methods are not so easily adapted to finding Mars life if the latter employs novel and unique biomolecules. Nevertheless, the principles of genetic and metabolic analysis of geological and environmental samples will be extremely useful in efforts to identify and quantify the magnitude and sources of any terrestrial contamination in martian samples, and they may serve for detecting Mars life if it is sufficiently similar to life on Earth.
Potential Biohazards: Organisms, Viruses, Viroids, Prions
Before discussing techniques for life detection, it is important to explain why researchers will pay essentially all their attention to organisms, largely ignoring viruses, viroids, prions, or other possible biohazards. As the following paragraphs make clear, only organisms are sufficiently self-reliant to proliferate in a completely alien world: The other agents require host organisms and also have extremely specific requirements with respect to their hosts.
As used here, the term organisms refers to all forms of cellular life, including bacteria, archaea, eukaryotic microorganisms, plants, and animals. All organisms possess their own genetic material, deoxyribonucleic acid (DNA). In addition, all organisms possess (1) machinery for duplicating the DNA, the process called DNA replication; (2) machinery for expressing genes by copying parts of the DNA into a related molecule called ribonucleic acid (RNA), the process called transcription; and (3) machinery for reading some RNA molecules as instructions for the synthesis of specific proteins, the process called translation. On Earth, all organisms perform these processes, and, in the case of transcription and translation, the machineries are so highly conserved that they are commonly used to detect and identify organisms. Parts of the translational machinery, the small subunit ribosomal RNA (SSU rRNA), or its gene (frequently called the rDNA), are particularly easily detected in all characterized organisms and have been examined for more than 30,000 different cases.9
Some organisms can synthesize everything they need from a relative handful of inorganic compounds (e.g., water, carbon dioxide, ammonia, sulfate, and some trace minerals).10 Plants and cyanobacteria are examples of such primary producers (autotrophs). Other organisms (heterotrophs) cannot make new organic matter and so must consume organic compounds. The nutritional requirements of these organisms can range from the very simple (e.g., only one organic compound needed) to the very complex (perhaps 100 different compounds needed).11 Such fastidious organisms are very specialized, and each type is usually restricted to a very few specialized environments.
Because most biologically important compounds are available only when produced by organisms, inevitably the organisms with the most complex nutritional requirements require an association with a host organism or community of organisms than can supply their needs. Some are pathogenic or predatory, while others are symbiotic or scavengers. If appropriate hosts (alive or dead) are not available, such organisms will eventually die.
By contrast, an organism that is nutritionally more flexible can often be found in a variety of different environments. These organisms can pose at least two threats. First, some can act as opportunistic pathogens, taking advantage of the nutritional richness of a host organism when it is available and thereby endangering the host, but also being capable of living independent of a host as well. Second, some as alien organisms might irreversibly damage, or even take over, an ecosystem, as has occurred when species of plants or animals have accidentally or intentionally been released into new environments. Although many die out naturally, others overwhelm the new ecosystem in which they lack natural enemies, or might otherwise destroy a system that is not adapted to their effects.
Viruses are obligate parasites, deriving from their host organisms not just important chemical compounds but also protein-synthesizing machinery. Although viruses possess their own genetic material (DNA or RNA), they always rely on their host for its translational machinery, and sometimes also for its genome-replicating and genome-transcribing apparatus. Thus, no matter what raw molecules are provided to a virus, it cannot, in the absence of an appropriate host organism, make any new protein and hence cannot make more viruses. This absolute need for co-opting cellular machinery leads to substantial specialization of viruses in terms of the hosts that they can successfully infect. Thus, although viruses can be very successful among and destructive of the terrestrial organisms with which they have coevolved, it is extremely unlikely that viruses for any truly novel form of life—one that is not based in the same genetic materials and exactly the same relationships between gene sequence and amino acid sequence in proteins (the translation table that we call the genetic code)—would be able to replicate on Earth.12 This means that we need not speculate about a particularly resistant strain of virus that defies the biological principles of terrestrial life.
|
9 |
Woese, C.R. 1987. Bacterial evolution. Microbiol. Rev. 51:221-271; available online at <http://www.cme.msu.edu/RDP/html/index.html>. See also Maidak, B.L., Cole, J.R., Lilburn, T.G., Parker, C.T., Jr., Saxman, P.R., Stredwick, J.M., Garrity, G.M., Li, B., Olsen, G.J., Pramanik, S., Schmidt, T.M., and Tiedje, J.M. 2000. The RDP (Ribosomal Database Project) continues. Nucleic Acids Res. 28:173-174; available online at <http://www.cme.msu.edu/RDP/html/index.html>. |
|
10 |
Brock, T.D., and Madigan, M.T. 1991. Biology of Microorganisms, 6th Edition. Prentice-Hall, Englewood Cliffs, N.J., pp. 121-127. |
|
11 |
Brock and Madigan, 1991; see footnote 10 above. |
|
12 |
Space Studies Board, National Research Council. 1998. Evaluating the Biological Potential in Samples Returned from Planetary Satellites and Small Solar System Bodies. National Academy Press, Washington, D.C. |
Viroids are in many respects akin to viruses, but they are simpler: They are nothing but naked genetic material (RNA), relying entirely on host machinery for replication, transcription, and translation.13 Viroids have been found only in plants, and any given type of viroid has a very limited range of hosts. Also, because viroids do not possess any molecules or structures to facilitate their entry into a host, they depend on wounding of a plant, for example by an insect, to gain entry. All the considerations that limit speculation about exotic viruses apply also to viroids. With respect to inactivation, the most significant differences between viruses and viroids are that viroids as naked RNA are more susceptible to chemical degradation but, because they have less genetic material, viroids also present a smaller target for inactivation by gamma irradiation.
Prions are infectious proteins.14 They have no genetic material. Instead they reflect an inappropriate folding of a normal cellular protein, or part of one. In essence, a prion causes the misfolding of a normal protein, making that protein behave as a prion, creating a chain reaction of misfolding. Because a prion has no genetic material, the host organism must already produce a protein that can be directed into the incorrect configuration by the prion. In all known cases, the range of organisms that prions can infect is limited, because corresponding proteins must exist in each host and must be sufficiently conserved in structure to allow for both normal folding and prion-induced folding. Transmission of prions between individuals appears to be very limited, with the exception of extreme cases such as those in which diseased brains have been added to animal feed, or extracts of diseased brains from one animal have been injected into the cranium of another animal. Overall, although the possibility cannot be excluded that a molecule of extraterrestrial origin could act as a prion, leading to the pathological refolding of a terrestrial protein, there is no reason to consider such a possibility as more likely than a random terrestrial molecule acting as a prion. These two factors, low contagiousness and the low likelihood that an alien molecule would specifically interact with a terrestrial protein, make prions of little concern.
In summary, two categories of biological threat could be posed by samples returned from Mars. The first is that the samples might contain biological materials similar to those found on Earth. In this case, all four types of infectious agents—organisms, viruses, viroids, and prions—would be of concern, but then the means of sterilizing the samples are also well understood. The second is that the samples might possibly contain a life form significantly different from terrestrial life. In this case, returned viruses and viroids could not replicate in terrestrial organisms, because such agents would rely on a gene expression system different from that of organisms on Earth. Alien organisms might, however, find a niche in which they could proliferate. At the present state of technology, it would be difficult or impossible to fully evaluate the potential impacts of alien life forms on complex terrestrial ecosystems. Researchers anticipate technological advances, in particular advances associated with the monitoring of complex gene expression patterns, that may provide tools for detecting environmental perturbations. However, the necessary first step toward evaluation of how possible alien life forms might affect complex terrestrial ecosystems or human health will be the ability to detect the presence of microbial life in samples returned to Earth.
There are two additional scientifically pragmatic reasons for focusing life detection efforts on organisms rather than viral parasites or infectious proteins. First, because they constitute the vast bulk of the biomass in terrestrial ecosystems (since all of the other agents must be produced by organisms), organisms or their traces might be easier to detect than other agents. Second, demonstrating that one has found an alien virus, viroid, or prion without a known host organism on which to grow it is an essentially futile exercise.
Detection by Cultivation
The traditional method for detecting viable organisms is cultivation. If growth on an otherwise sterile medium is observed, the result is definitive. No known abiological inoculum will produce increasing numbers of cells (a result that can be traced back to Pasteur’s refutation of spontaneous generation). Verification of the occurrence of growth, not chemical precipitation, is frequently carried out initially by optical microscopy, and it can be confirmed by electron microscopic examination.15 Growing samples can be repeatedly diluted and regrown (although
there are caveats). Despite published reports of “growth” that was almost certainly chemical rather than biological,16 such results are extremely rare and are generally negated by the use of appropriate experimental controls.17 In addition to uninoculated cultures, inocula subjected to sterilizing treatments can be used as controls in experiments designed to check for positive growth.18 By far the most significant source of false positive indications of growth is contaminated cultures. Contamination is a substantial concern for all aspects of sample analysis, but contamination with viable organisms presents the added complication that the organisms can replicate and their significance thereby increase.19
The more significant problem with the use of culturing for life detection is that of false negatives, leading to failure to detect the existence of viable organisms that defy cultivation.20 In spite of great advances in available methods, several lines of evidence indicate that the majority of organisms on Earth (whether measured by cell count, cell mass, or genetic diversity) have not been successfully cultivated in the laboratory. In the worst case, there is a complete lack of recordable growth; less extreme is the case in which organisms must be incubated for months before there is an observable increase in cell density. Studies of some organisms in their natural environment suggest that slow growth is intrinsic to the organism, and not just a result of failure to optimize the culture conditions.21,22
Thus cultivation-based studies will not provide high confidence that a sample is devoid of viable organisms. Nevertheless, the intrinsic sensitivity of the method for organisms that do grow successfully, and the historical role of cultivation in detecting contamination (by screening for common, easy-to-grow terrestrial organisms), mean that culture-based studies are important and cannot be bypassed.
Efforts to test the martian samples by cultivation will be limited primarily by logistics. Attempts at cultivation must be performed with unsterilized samples and hence must be carried out within the quarantine facility. Although a variety of growth conditions can be tested without specialized equipment, testing becomes more difficult when the samples require special environmental conditions (temperature, pressure, unusual atmospheric composition, and so on). Cultivation can provide very sensitive assays for terrestrial contamination and conceivably for some forms of alien life but cannot prove the absence of extraterrestrial life. The possibility of false negatives suggests that this approach should be limited to a circumscribed set of carefully chosen growth conditions.
Detection by Microscopic Examination
In surveys of terrestrial samples, some of the commonest methods of screening for cells are direct microscopic examination, epifluorescence microscopy, and electron microscopy.23 In direct microscopy, many types of large or complex cells are quite evident, but small simple cells cannot be unambiguously identified. If putative cells are organized spatially in a “microcolony,” there is increased reason to believe that they are of biological origin. If samples are stained with a DNA-binding fluorescent dye, cells become easier to see. However, a test that relies on extraterrestrial life being DNA-based is less robust than one that tests merely for cellular organization. For assessing cellular structure (as opposed to abiotic structures), electron microscopy provides additional resolution and hence more information. Although having a capability for basic optical microscopy within the quarantine facility is important, electron microscopy and most other detailed examinations should be carried out on sterilized samples outside the facility (Chapter 6)—assuming, of course, that the sterilization procedure used does not
destroy morphological evidence of life. How various methods of sterilization affect microscopic morphological evidence of life needs to be established.
Viability cannot be assessed by the methods mentioned above. Most staining procedures and methods of preparing samples for electron microscopy kill any cells that were alive. Although there are some indirect tests of viability (or, more correctly, metabolic activity) based on dye exclusion, or dye uptake,24 none are generally applicable to diverse (let alone extraterrestrial) samples, and interpretation of the results may be confused by staining of abiological particles.
Detection by Analysis of Chemical and Isotopic Structures
Some of the most general and robust methods for detecting extant or recent life are based on detecting the presence of specific molecules, or chirality25 or isotopic anomalies in molecules. In all of these cases, viability, or even cellular structure, is not required. These criteria are also discussed below in the context of fossil evidence of long-dead organisms.
Molecules whose presence can be taken as evidence of life are proteins, DNA, RNA, straight-chain fatty acids, and a variety of other lipids. One common and sensitive method for detection of terrestrial life (viable or recently dead) is the use of the polymerase chain reaction to amplify fragments of DNA encoding ribosomal RNA (rRNA). The process is described in Box 2.1 and is sketched in Figure 2.1.1. Ribosomal RNA is a common target of PCR because it is universal among terrestrial organisms, and there is an extensive database of the diverse sequences that have been obtained.26 Although, as noted above, it is not realistic to depend on DNA and rRNA genes being present in extraterrestrial life, their presence does provide a sensitive test of terrestrial contamination (even more sensitive than growth, in that it is not limited to organisms that researchers know how to cultivate).
The analysis of relatively large or complex molecules (whole genes) is motivated by attempts to avoid false positives caused by abiotic organic molecules. However, this approach is limited because it is modeled on the detection of molecules produced by terrestrial organisms. Alternatively, even without prior knowledge of the specific molecules involved, evidence for the biological origin of organic compounds can be based on the chiral enrichment of the molecules present. Although it is possible for abiotic processes to create anomalies, naturally occurring departures from random chirality in demonstrably abiotic samples are rare at best. If there are gross enantiomeric excesses, then net chirality can be measured with little preconceived notion of the identities of the molecules present. However, prior knowledge of the molecules to be screened can help optimize the sensitivity of the assays.
Stable isotopes are almost inevitably fractionated by biological systems. The compounds affected include the cellular constituents and the products of the cellular energy-producing reactions. Bulk samples can be analyzed for average composition, or an ion microprobe can probe localized features (resulting in greater sensitivity and much lower throughput, or speed of analysis, since many such features must be analyzed). The test is not definitive, however (see Box 3.2 on carbon isotopes in Chapter 3).
Terrestrial life also has an elemental composition that is far from equilibrated with its environment. The degree of disequilibrium can be assessed either by probing the bulk composition of a sample, or by using tomographic techniques to produce a spatially resolved map of abundances, with a much decreased analytical throughput.
|
24 |
Mason, D.J., Lopez-Amoros, R., Allman, R., Stark, J.M., and Lloyd, D. 1995. The ability of membrane potential dyes and calcafluor white to distinguish between viable and non-viable bacteria. J. Appl. Bacteriol. 78:309-315. |
|
25 |
Chirality refers to the fact that asymmetric molecules can occur as right- and left-handed variants. Biological processes tend to favor one variant or the other, while abiotic processes produce them in equal numbers. |
|
26 |
Maidak, B.L., Cole, J.R., Lilburn, T.G., Parker, C.T., Jr., Saxman, P.R., Stredwick, J.M., Garrity, G.M., Li, B., Olsen, G.J., Pramanik, S., Schmidt, T.M., and Tiedje, J.M. 2000. The RDP (Ribosomal Database Project) continues. Nucleic Acids Res. 28:173-174; available online at <http://www.cme.msu.edu/RDP/html/index.html>. |
|
Box 2.1 The Polymerase Chain Reaction Before a cell can divide, it must make a precise copy of all of its genes, so that each daughter cell inherits all of the original genes. Called DNA replication (DNA is the material of which genes are made), this process is initiated by an enzyme called DNA polymerase. The polymerase chain reaction (PCR) is a technique for amplifying (making many copies of) a defined portion of a DNA molecule by using the same biochemical reactions that cells use to copy their genes before they divide.1 With DNA polymerase as the catalyst, it is possible to convert one copy of a DNA molecule into two copies identical to the first. If this process is repeated, the two copies become four copies. A third cycle results in eight copies, and the pattern can be rapidly continued until there are billions of copies of the original, making the DNA relatively easy to detect in the laboratory. The process is called a chain reaction because the product of each reaction is more molecules that can serve as the substrates (templates) from which more reactions can occur. How PCR works is outlined below and diagrammed in Figure 2.1.1.
|
Relative Sensitivity and Throughput of Detection Techniques
The sensitivities of the various techniques outlined above are not discussed in detail here. To facilitate comparison, Table 2.1 indicates some of the properties of the techniques, characterizing them qualitatively or in terms of broad ranges, since it is frequently possible to improve one parameter at the expense of another.
Analyses based on cultivation and observation of growth can take advantage of amplification to detect as little as a single viable organism (of the right type) in a sample. In addition, this method can screen a sample of virtually any mass without an increase in the complexity of or time required for the test, so it has a high throughput. The
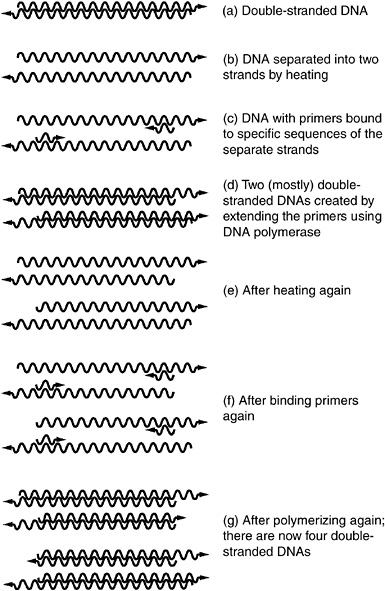 FIGURE 2.1.1 Schematic representation of the successive splitting and copying of nucleic acid molecules that constitute the polymerase chain reaction. |
disadvantage of this technique is that growth is often slow and there are a limited number of organisms that can be detected.
Analyses based on PCR can also take advantage of amplification to detect as little as a single intact rRNA gene in a sample. In addition, PCR can be used to screen samples whose mass ranges from nanograms to a gram without an increase in the complexity of or time required for the test, so it also has a high throughput. Although the use of PCR with rRNA genes will enable detection of virtually any terrestrial organism, the disadvantage of this technique is that it is unlikely to detect novel life forms that originated independently of terrestrial life. In spite
TABLE 2.1 Properties of Some Techniques for Detecting Viable and Recently Dead Cells
|
Technique |
Sensitivity |
Specificity for Life Detection |
Generalitya |
Time per Sample |
Time per Gramb |
|
Cultivation to test for growthc |
Very high: 1-10 cells |
Very high |
Requires cell type that grows in culture |
Days to months |
Days to months |
|
Test for metabolic activity |
Moderate: 103-106 cells |
Moderate to high |
Specific metabolism, but any biology |
Minutes to days |
Minutes to days |
|
Microscopic examinationd |
Very high: 1-100 cells |
Moderate |
High |
Minutes to hours |
Days to months |
|
PCRe |
Very high: 1-10 cells |
Very high |
Requires rRNA genef |
Minutes to hours |
Minutes to hours |
|
Chemical structure analysisg |
Moderate to low: 106-1015 molecules; 103-109 cells |
High |
Moderate |
Minutes to days |
Minutes to days |
|
Chirality analysis |
Low: >1010 molecules; 106-109 cells |
Very high |
Very high |
Minutes to days |
Minutes to days |
|
Isotope stability analysis |
Very high to moderate: 105-108 cells |
High |
Very high |
Minutes to hours |
Hours to months, depending onsensitivity |
|
Elemental composition analysis |
High to low: 100-108 cells |
Moderate |
Very high |
Minutes to hours |
Hours to months, depending on sensitivity |
|
aBased on sensitivity for detection of extraterrestrial life. bTime per gram exceeds time per sample in the cases where studying a gram of material requires analysis of many samples within the gram. cThat is, an increase in biomass. dObservation at the micrometer level. eValues assigned assume high DNA-extraction efficiencies and optimized PCR conditions. fOther highly conserved genes might also be targeted. gBiomolecular structures, e.g., lipids, macromolecules. |
|||||
of these limitations, the potentially high sensitivity and high throughput of both cultivation and PCR make them uniquely valuable for testing returned martian samples.
Although the ability to detect as little as one cell by culturing or amplification by PCR (albeit with significantly less than 100% reliability) is appealing, this virtue is tempered by two facts. First, lack of detection is not a reliable indicator that the sample tested contained no viable organisms. Under any conditions chosen, most known organisms will not respond to culturing.27 And although PCR is a more general method of detecting viable or recently dead terrestrial organisms than culturing, it too is capable of producing false negatives.
Another limitation of both culturing and PCR is that only a relatively small portion of a returned sample can be subjected to these detection techniques because doing so greatly compromises the samples for all subsequent
TABLE 2.2 Time Scalesa for Optical Scanning
scientific analyses, especially nonbiological analyses. The absence of detectable life in a small subsample can only place bounds on the abundance of life in the remaining material. If the density of organisms were, for example, 300 per complete sample, then the expected number in a 1% sample would be 3 and the probability of detecting none would be approximately 0.05, according to Poisson statistics. That is, if organisms detectable only at a low level were present, there would be a significant chance of missing them. If the density of organisms were 30 per complete sample, then the probability of detecting none in a 1% sample would be about 0.74, i.e., quite high. In essence, it would be possible to say only that viable organisms were not abundant in the sample, not that the rest of the sample was free of life.
Although it might be possible to achieve comparable sensitivity with microscopic searches for cell morphology, studies of stable isotopes, or analyses of elemental distributions, and although such studies might have the advantage of being able to detect novel life forms, none of these methods has a very high throughput. If the surfaces of sand-sized grains were scanned at the rate of 1,000 µm2 per second, then to cover the surfaces of 1 gram of grains would require more than 3 weeks for 1-mm-diameter spherical sand particles, and more than 8 months for 0.1-mm particles (Table 2.2). Although it might be possible to examine samples more rapidly than this, a price would be paid in increased errors.
Life: Indigenous Versus Contaminant
A critical part of life detection in martian samples will be the ability to assess whether the source is extraterrestrial or terrestrial. In the case of extant or recent life, researchers expect to find intact molecules, and possibly cell growth. Short of cultured growth, almost certainly the best test for terrestrial contamination is the use of PCR amplification of rRNA genes. For reliability, more than one portion of the molecule (and possibly more than one gene) should be targeted for amplification. If there is more than a trace of terrestrial or terrestrial-like life, this approach should detect it.
If PCR were to detect rRNA genes in a sample, it would mean that the life would be traceable to a common origin with terrestrial life (based on the assumption that independent origin of exactly the same genetic material and machinery for gene expression is unimaginably improbable). Such a finding could indicate either of two things: (1) recent contamination of the sample or (2) an ancient common heritage of life on Mars and Earth. Because of the broad rRNA-based sampling of life that has been conducted on Earth,28 it is likely that recent
|
28 |
Maidak, B.L., Cole, J.R., Lilburn, T.G., Parker, C.T., Jr., Saxman, P.R., Stredwick, J.M., Garrity, G.M., Li, B., Olsen, G.J., Pramanik, S., Schmidt, T.M., and Tiedje, J.M. 2000. The RDP (Ribosomal Database Project) continues. Nucleic Acids Res. 28:173-174; available online at <http://www.cme.msu.edu/RDP/html/index.html>. |
contamination could be quickly recognized because it would yield rRNA sequences similar to specific, present-day lineages on Earth. In addition, if a novel rRNA gene sequence (one that is not represented in current databases) were found, it would be possible to use primer sequences directed for this novel sequence in the PCR protocol to screen rapidly and systematically for the same sequence in other terrestrial environments. Specific attention could be focused on screening sources of materials brought into the quarantine facility. If the recovered sequences were only distantly related to known terrestrial sequences, then the possibility would have to be taken seriously that they were linked by an ancient transfer event between the planets (i.e., panspermia). More complex histories would obviously be possible for an amplified sample that produced multiple sequences, but phylogenetic analyses could be used to check consistency with alternative hypotheses regarding origins and transport.
Life detected from the presence of life-associated chemical reactions would be very difficult to assess in terms of terrestrial or extraterrestrial origin. Depending on the reaction tested, the signal might also be of abiotic origin.29,30 In the event of suggestive reactions, it would be important to seek other cellular or molecular evidence.
If chemical signatures of life were found, they could be compared to terrestrial biomolecules. A match would again raise questions as to whether the chemical compounds were similar because of recent contamination or because they had an ancient common origin. However, because the use of specific chemical compounds to detect life is a relatively insensitive technique, only abundant contamination would be detectable—an abundance that would also facilitate the identification of its source.
Other types of tests for evidence for life can be even more difficult to evaluate in this regard. Many combinations of observations are possible, so it is not productive to attempt an exhaustive analysis.











