Bioinformatics: Emerging Opportunities and Emerging Gaps1
Paula E.Stephan and Grant Black
Georgia State University
INTRODUCTION
A typical gene lab can produce 100 terabytes of information a year, the equivalent of 1 million encyclopedias.2 Few biologists have the computational skills needed to fully explore such an astonishing amount of data; nor do they have the skills to explore the exploding amount of data being generated from clinical trials. The immense amount of data that are available, and the knowledge that this is but the tip of the data iceberg means that researchers must increasing-
ly rely on an interdisciplinary approach to not only succeed in but to just proceed with research. It also means that individuals are needed who can work in the emerging field of bioinformatics, combining skills of computer science with a knowledge of biology.
This paper examines this emerging field. We begin by discussing demand for individuals who can work in the field. We then summarize the number of individuals in the pipeline who are currently being trained in the field. The indication that demand is strong and the pipeline sparsely populated leads us to ask why the response of higher education has been sluggish. We close by suggesting possible solutions to address the problem of sluggish response.
DEMAND FOR INDIVIDUALS IN BIOINFOMATICS
By all accounts the field of bioinformatics/computational biology is booming. The scientific press stresses the high salaries paid to new hires ($65,000 for persons with top master’s training; $90,000 or more for Ph.D.s) and the intensity with which headhunters seek out possible candidates.3 Universities complain that their students are “grabbed” before they are able to complete their degrees and that their faculty and students are lured to industry, creating the concern that the bioinformatics field is “eating its seed.”4
Here we use a two-part methodology to investigate demand: we analyze position advertisements in Science as well as summarize data collected from a survey of programs concerning their placements of students. Figure 1 presents job openings in bioinformatics and computational biology by month for a two-year period as measured by counting position announcements in Science. Given the methodology, the numbers reported are a lower bound.5 In 1996, 209 posi-
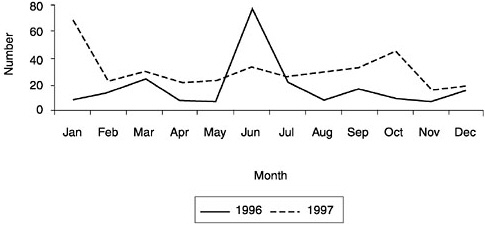
FIGURE 1 Job openings in bioinformatics and computional biology from Science ads, 1996 & 1997.
tions were advertised; in 1997 this had increased by 69 percent to 354.6 These counts include two special advertising supplements focused on biotechnology, one in June 1996 the other in July 1997.7 Both supplements were dominated by ads from SmithKline.8 In a typical month (ignoring special supplements) the journal averaged 12 position announcements in 1996. This had more than doubled by 1997, rising to 25.
Table 1 organizes the information in terms of type of entity placing the ad, rather than number of position announcements. Three categories are listed: firms, universities, and other not-for-profits, including government.9 We see that the number of entities placing ads grew from 70 to 118 between 1996 and 1997, representing a growth of 68 percent. In both years the majority—about 63 percent of entities placing ads—were firms, and the number of firms placing ads grew by 70 percent.10 In addition to large firms, such as Bristol-Myers Squibb, Eli Lilly, SmithKline-Beecham, Pfizer, Merck, Abbott, Bayer, and Monsanto, a
|
6 |
This finding of growth is consistent with Wendy Yee’s report that from 1995–1996 the number of ads related to bioinformatices tripled. See “The Top Five Career Trends of 1996: Informatics Anything,” http:\www.nextwave.org/server-java/SAM/pastloop/trend2.htm. |
|
7 |
There was also a special supplement in July 1996, apparently an addendum to the June supplement. |
|
8 |
In the 1996 supplement SmithKline said that they wanted to about double their staff of 30. In the 1997 supplement they again said they wanted to about double their staff, this time reported at 40, suggesting that SmithKline has plans to grow and is experiencing difficulty filling positions in bioinformatics/computational biology. |
|
9 |
Note that institutes that are affiliated with universities, such as Whitehead, are here counted as “other not for profit.” |
|
10 |
Note: these are firm, not enterprise, counts. |
TABLE 1 Number of Distinct Entities Advertising Positions in Science
|
|
Year |
|
||
|
Sector |
1996 (share) |
1997 (share) |
Number of distinct position announcements in 1996 and 1997 |
Growth between 1996 and 1997 (%) |
|
Firms |
44 (62.8) |
75 (63.6) |
90 |
70.4 |
|
Not-for-profit universities |
17 (24.3) |
22 (18.6) |
36 |
29.4 |
|
Other not for profit |
9 (12.9) |
21 (17.8) |
27 |
133 |
|
Total |
70 |
118 |
153 |
68.6 |
number of smaller biotech firms, such as Regeneron, Immunex, and Zenez, have position announcements. A substantial number of firms—29 to be exact—placed ads in both 1996 and 1997.
In contrast, only 36 universities ran position announcements during the period, and the growth rate for university ads was slightly less than 30 percent. Universities placing ads included the University of California at Los Angeles (UCLA), the University of California at Irvine (UC Irvine), the University of California at San Francisco (UCSF), the University of Pennsylvania, the University of Southern California (USC), and California Institute of Technology. Only three universities placed ads in both years. A number of not-for-profit entities (such as the Centers for Disease Control) also ran position announcements and the number of ads from this sector more than doubled.
Based on the position announcements, jobs in computational biology range from entry-level data analysts and programmers to senior-level scientists and research directors.11 Lower-level positions that are more directly computer-oriented call for as little as an undergraduate science degree, and some state no degree requirements. The majority of positions call for a doctorate degree in either a science (preferably molecular biology) or computer science with considerable programming or bioinformatics experience, although a number of positions explicitly advertise for individuals with a master’s degree.
The second part of the methodology to study demand involved using data collected from a survey of programs concerning the placement of students. The
data are for the 12-month period March 1998 to March 1999. Of the 21 identified formal and informal programs, 16 supplied data and eight institutions/programs reported students taking jobs during the period. All but one indicated that students completed their degrees prior to employment, suggesting that earlier media reports that students are commonly recruited before completing programs do not hold for most trainees in the field.
Fifty-three individuals were reportedly placed: three at the undergraduate level, 23 at the master’s level, 13 at the doctorate level and 14 with postdoctorate training. Nine students who graduated during the period chose to continue their education and training. Five of these went to a postdoc position either from a Ph.D. program or from a previous postdoc appointment. Three moved on to graduate programs, and one went to medical school. Specific placement information was ascertained for 42 of the 53 hires. Seven of the 53 placements are at an academic institution, and only one position is at a government-sponsored institution. The remaining identified placements are in the private sector and include four graduates who established their own firms.
Salary data for a subset of the hires are presented in Table 2.12 Salary ranges are reported for 71 percent of the hires during the period. The greatest lack of information is at the postdoctorate and doctorate levels. As expected, salaries for the most part climb as the level of training rises, starting in the $40,000–$50,000 range for BAs and reaching over $100,000 for one post doc. But there are exceptions. For example, two of the three undergraduates who were placed received salaries between $50,000 and $60,000. This is higher than that earned by seven of the masters students, although ten of the 19 master’s students for whom we have salary information earn more than $60,000. One masters student received a starting salary of over $100,000. Reported salaries for five hires at the doctorate level are over $70,000. One is between $80,000 and $90,000; another is over $100,000. Three postdocs received placements with a salary between $80,000 to $90,000. One was placed at a salary of over $100,000. One institution reported that one or more master’s students received a signing bonus.
Our surveys of ads and programs lead us to conclude that (1) demand is strong and growing but small relative to other areas, (2) demand is driven in large part by industry, (3) salaries are high relative to other areas in the life sciences, and (4) the majority of jobs being advertised are not being filled by graduates of programs. This suggests that a number of the jobs being advertised either remain unfilled or are filled by individuals coming from outside the field. The head of Merck’s computational biology program, for example, is a physicist.
TABLE 2 Salary Ranges by Training Level, January 1998 to March 1999
|
Training Level |
Salary Range $ (number of hires) |
|
Undergraduate |
40,001–50,000 (1) 50,001–60,000 (2) |
|
Masters |
40,001–50,000 (7) 50,001–60,000 (2) 60,001–70,000 (1) over 70,000 (8) over 100,000 (1) unknown (4) |
|
Doctorate |
60,001–70,000 (1) over 70,000 (5) 80,001–90,000 (1) over 100,000 (1) unknown (5) |
|
Postdoctoratea |
80,001–90,000 (3) over 100,000 (1) unknown (5) |
|
aExcludes placements for 1997 trainees listed in the W.M.Keck Center’s 1997 Annual Report. |
|
PIPELINE
Table 3 summarizes student enrollment in training programs by degree level as of March 1999. The data come from the same survey from which the salary data were collected. Of the 21 identified programs, information was gathered for 16. Information concerning type of support is also provided. We see from the table that doctorate programs dominate enrollment in terms of the number of
TABLE 3 Characteristics of Formal Training Programs as of March 1999a
|
|
Undergraduate |
Masters |
Doctorate |
Postdoctorate |
|
Total Number of Programs |
3 |
5 |
9 |
7 |
|
Number of Programs with Internal Supportb |
1 |
1 |
3 |
0 |
|
Number of Programs with External Support |
2 |
3 |
8 |
9 |
|
Enrollmentsc |
23 |
35 |
86 |
~25 |
|
Note: Placement information is included in the respective programs for several Ph.D. students and postdocs trained at the University of California-Santa Cruz. The nature of the reported data broadens the time period of placements, starting as early as 1995 and ending in May 1999; thus, the reporting period extends beyond the January 1998 to March 1999 period. Placement information is included for five 1997 postdocs listed in the W.M.Keck Center’s 1997 Annual Report and, thus, in the case of the postdocs, the reporting period extends before January 1998. Due to the lack of exact counts reported, the precise completion rate cannot be determined. aBaylor College of Medicine, Boston University, Northwestern University, Rutgers University, Stanford Universtiy, University of California-Santa Cruz, University of Pennsylvania, University of Washington, W.M.Keck Center for Computational Biology. The Keck Center includes Baylor College of Medicine, Rice University and the University of Houston. Note that Baylor also has a program that is independent of the Keck Center. This program is counted separately here. bBased only on institutions responding to the question regarding internal sources of funding; several institutions did not respond to this question. cIncludes counts of students in degree programs in the Department of Computer Sciences at the University of California-Davis and the University of California-Santa Cruz; there are no formal bioinformatics/computational biology programs at UC-Davis and no formal undergraduate program at UC-Santa Cruz. |
||||
training programs and the number of students (86). The largest program is at Stanford, followed by Rutgers, the W.M.Keck Center program, and Baylor College of Medicine. The smallest enrollment in the spring of 1999 was at the bachelor’s level, with the University of Pennsylvania having by far the largest program. Thirty-five students were enrolled in the various masters programs and approximately 25 in formal postdoc programs at the time of the survey. Enrollment in master’s programs was expected to more than double in the fall of 1999 when new programs at George Mason University, the Georgia Institute of Technology, and Boston University came on line.
THE GAP
This strong demand in bioinformatics and the creation of new programs comes at a time when the prospects of young life scientists look less than promising. Despite the “hot” reputation of the life sciences (and in part because of its
hot reputation), a large and increasing number of early-career life scientists are unable to find the type of job that will permit them to become independent researchers and establish their own lab. Sufficient concern over the career outcomes of young life scientists exists to have warranted the establishment by the National Research Council of the Committee on Dimensions, Causes, and Implications of Recent Trends in the Careers of Life Scientists. The committee issued its report in September 1998, stating that “the imbalance between the number of life-science Ph.D.’s being produced and the availability of positions that permit them to become independent investigators concerns the committee.” The committee concludes that “Intense competition for jobs has created a ‘crisis of expectation’ among young life scientists. Much of this imbalance is the result of the professional structure of the life sciences research enterprise where “the important work of conducting experiments rests almost entirely on the shoulders of graduate students and postdoctral fellows.” Recommendations of the committee included restraint of the rate of growth of the number of graduate students in the life sciences.13
Is it not contradictory that the committee concluded that a “crisis of expectations exists for young life scientists” at a time when demand is strong and growing in the field of bioinformatics? Why are there but nine doctoral programs in the United States in computational biology,14 while there are approximately 194 programs in biochemistry and molecular biology and over 100 in molecular and general genetics?15 More generally, the contrast of the two fields leads one to ask if the structure of the U.S. science enterprise leads to missed opportunities in emerging fields, particularly when the demand is heavily centered in industry.
Here we examine four interrelated explanations of this gap: The four are (1) the low incentive for individual faculty to establish such programs and attract students in the area, (2) an educational system that responds differently when demand is driven by industry as opposed to when demand is driven by universities and research labs, (3) the interdisciplinary nature of the field creates disincentives to the establishment of programs, and (4) the quick fix—turning life scientists into computational biologists—is not possible, given the skills and quantitative abilities of individuals in the life sciences, nor is the incentive present for computer scientists to opt for additional training in the life sciences.
Lack of Incentive
The research structure that has evolved in academe in the U.S. means that faculty are extremely responsive to research funding opportunities since it is external grants that provide resources to purchase equipment and support graduate students and postdocs—the collaborators who are absolutely essential to the lab of the principal investigator (PI). Furthermore, at many medical schools in the U.S., funding not only supports the collaborators, it also supports the PI, which means that the PI can only retain his or her academic appointment as long as the PI has funding to cover the cost of the lab and the PI’s salary.16
This suggests that an effective way to alter the educational mix of graduate students is to alter the amount of research funds directed to an area and thus provide the incentive for faculty to recruit students into the field. To what degree has this occurred in computational biology/bioinformatics.
The evidence, which is difficult to assemble due to its fragmented nature, suggests that funding agencies have only begun to do this and still not to a major extent.17 Instead, funders have placed their targeted computational eggs in the training basket. NSF has provided training funds through its Computational Biology Activities and the Sloan and Keck foundations have targeted funds to the training of individuals in bioinformatics; non-targeted training funds have also come from NIH. While such a strategy may be best in the long run, in the short run training grant initiatives may be ignored by many faculty. This is because training grants signal collective bodies. Research grants signal individuals. It is difficult to get academic units composed of competitive PIs focused on where the next grant will come from to engage in the collective response required to succeed in creating new programs. Faculty are much more attuned to thinking about individual-investigator-initiated grants. And, in the past, little funding has been targeted by federal agencies at research in computational biology. For example, at a time when NIH supported more than 25,000 active research grants a year, only 96 R01s listed the key words “computational and biology” and only 11 R29s listed these key words. A similar statement can be made with regard to NSF CAREER grants. Of the approximately 400 active grants in 1996, only six appear to be directly related to the area of computational biology. Moreover, while many of the training grants were targeted at bioinformatics, most of these research awards were not targeted specifically to the area.18
Educational System Responds More Slowly to Demand Driven by Industry as Opposed to Demand Driven by Research Opportunities
As indicated in the introduction, demand for computational biologists is substantially driven by industry, which sees genetic data as “the major driving force” in drug discovery.19 SmithKline Beecham is a case in point. Their June 7, 1996, full-page ad in Science reports that they had 30 individuals working in the area, with plans to double that number by 1997 (p. 1527). Six months later, they ran another ad in Science, again saying that they had plans to double their staff, this time reported to be at 40. Moreover, SmithKline has aggressively hired established researchers from academe and the non-profit research sector. In 1995 they succeeded in attracting David Searls away from the University of Pennsylvania, and shortly after Searls’s arrival they hired James Fickett from the Los Alamos National Laboratory, Randall Smith from Baylor College of Medicine, and Chris Rawlings from the Imperial Cancer Research Fund in London.20
Does it make a difference that the demand is industry-driven, as opposed to driven by academe? We are inclined to say yes for two reasons. First, every time industry hires a faculty member it means that there is one less professor to train future computational biologists. Thus, while the practice of recruiting faculty from academe provides a ready source of knowledge, and hence spillovers from academe to industry, the practice—where replacement is difficult—impairs academe’s capacity to continue the training initiatives it has already begun.21 The Baylor program reportedly experienced difficulty when Randall Smith left to join SmithKline, and, while the program at the University of Pennsylvania survived despite Searls’ departure, the remaining faculty were stretched as a result.
Second, academic departments in the life sciences are arguably not as responsive to demand driven by industry as are departments in engineering and computer sciences, which have long had a tradition of placing a sizeable number
of their graduates in industry. Few life science Ph.D.s head directly to industry upon completing their Ph.D.s.22 The reason stems from the fact that it is research funding—much more than the availability of jobs for graduates—that drives the size of academic programs in the life sciences. This is because research funding provides ready support for Ph.D. students in the life sciences and funding for the postdoctorate positions that recent Ph.D.s (and not-so-recent Ph.D.s) hold with such proclivity. In the biotech world of the late twentieth century, life science departments have found a ready supply of aspiring students who are willing to commit eight to ten years of their life to becoming life scientists so that they can have a shot at becoming a PI to continue working on the frontiers of knowledge. And, while it is viewed as both honorable and profitable for established faculty to work with industry, the profession would appear to still stigmatize the individual whose early career goal is to work in industry.
The Interdisciplinary Nature of Computational Biology Creates Disincentives to the Establishment of Programs
Bioinformatics requires training in computer and information science, mathematics, and the life sciences. Coordination among these three fields can often be an institutional nightmare since it involves not only cooperation across department lines but also across colleges. The department of computer sciences is often located in a college of engineering, while mathematics and life science departments are generally located in the college of arts and sciences. The situation is further complicated by the fact that universities that have medical schools often have an additional department of life sciences in the medical school.
The problems in working across department lines are difficult enough when departments are within the same college. They are compounded when departments are in different colleges or universities. For example, how are students to be advised? How are courses to be numbered and shared? How are contributions to be valued across college/university lines? And these are the simple questions. The harder questions concern which department/college will get “credit” for the new field. How will resources be shared? Who will get the new positions if individuals trained in computational biology are hired? Who will evaluate individuals’ promotion and tenure?23
The fields also differ in terms of career goals and opportunities for students. Michael Ashburner, director of research at the European Bioinformatics Institute, argues that more resources should be funnelled into master’s programs to provide uniform, specialized training since almost all those involved in bioinformatics come from another field (Gavaghan 1997). Yet, terminal master’s programs have historically been unpopular in the life sciences, in part because the ready supply of Ph.D. students and postdocs provided the needed assistance in the lab and in part because the field often stigmatized those with a terminal master’s degree.24 This stands in marked contrast to the fields of engineering and computer science where, a master’s education is looked favorably upon and employment is found (and encouraged) in industry.25
The Lack of a Quick Fix
A plausible fix to the “shortage” of individuals in computational biology is to turn young life scientists into computational biologists—or to take those with degrees in mathematics or computer information systems and augment their skills. Indeed, without a proactive strategy, this is what an economist would predict would occur. The number of postdoctoral grants offered in the area by Sloan, NSF, and Burroughs Welcome suggests that they have adopted such a strategy.
Several reasons, however, lead us to suspect that this strategy is less effective than might originally appear at face value. First, at the doctoral level the market for individuals trained in computer science appears to be sufficiently strong to retain computer scientists in that field. Table 4 reports the 1995 median annual salary of recent Ph.D.s employed full time in the broad areas of computer and information sciences and life and related sciences. The subcategory of biological and health sciences is also included. Three cohorts are identified: 1993– 94 graduates, 1990–92 graduates, and 1985–89 graduates. The large difference between salaries in computer and information sciences and salaries in life sci-
TABLE 4 Median Annual Salary of FTE Recent Ph.D. Graduates, 1995
|
|
1993–94 Graduates |
1990–92 Graduates |
1985–89 Graduates |
|
Computer and Information Sciences |
$54,000 |
$61,000 |
$65,000 |
|
Life and Related Sciences |
$30,400 |
$40,000 |
$52,000 |
|
Biological and Health Sciences |
$30,000 |
$38,600 |
$52,000 |
|
SOURCE: NSF/SRS, Characteristics of Doctoral Scientists and Engineers in the United States: 1995, 1997a. |
|||
ence for those who have been out for one to two years reflects the fact that a majority of individuals in the life sciences hold postdoctoral positions upon graduating. The difference narrows as the life scientists move out of these positions, but a substantial differential of 25 percent exists for those who have been out six to ten years. This suggests that job prospects in computer science are sufficiently strong to preclude computer scientists from seeking additional formal training in the biological sciences. The quick-fix strategy is more attractive to those trained in the biological sciences where the market, as we have already indicated, is considerably weaker.
Table 4 suggests that life scientists may have the incentive to seek additional training to become computational biologists. Do they have the background and aptitude to transform themselves into computational biologists? The response to the University of Pennsylvania’s training initiative in computational biology was remarkable. Over 200 individuals applied for the two postdoctoral positions. Yet, according to the faculty member who directs the program at the University of Pennsylvania, less than a handful qualified for the program precisely because the applicants had so little background in mathematics/statistics. There is reason to believe that this lack of quantitative background is generic to those with Ph.D.s in biology—not specific to the applicants to the University of Pennsylvania program. An examination of the requirements of five highly rated biology departments demonstrates that none have formal mathematical requirements for entry into their graduate programs; only a handful of graduate courses have a mathematics prerequisite up to introductory calculus.26
It is not just that life scientists lack training in math and statistics. A credible argument can be made that the typical life scientist lacks interest and excep-
TABLE 5 GRE Scores by Intended Field of Graduate Study, for Seniors and Nonenrolled College Graduates, 1993–96
|
Intended Graduate Field of Study |
|
Mean Score |
Percent of Test-takers with Score above 700 |
Percent of Test-takers with Score of 800 |
|
Biological Sciences |
Verbal |
501 |
3.6 |
0.1 |
|
Quantitative |
595 |
20.7 |
1.1 |
|
|
Health and Medical Sciences |
Verbal |
449 |
0.7 |
0.0 |
|
Quantitative |
515 |
5.8 |
0.1 |
|
|
Computer and Information Sciences |
Verbal |
483 |
5.4 |
0.2 |
|
Quantitative |
672 |
52.2 |
5.6 |
|
|
Mathematical Sciences |
Verbal |
502 |
6.5 |
0.2 |
|
Quantitative |
698 |
60.6 |
8.8 |
|
|
SOURCE: 1997–98 Guide to the Use of Scores, Educational Testing Service, 1997b |
||||
tional aptitude in these areas. This is somewhat borne out by data supplied by the Graduate Records Exam. Table 5 presents data on the scores of GRE test-takers from 1993–96 by their intended field of graduate study. The data indicate that individuals intending to pursue graduate study in the biological and health sciences test substantially lower in the quantitative area than those intending to study computer and mathematical sciences.27 While the mean quantitative score for biological sciences was 595, the score for computer sciences was 672—a difference of 77 points. Moreover, less than 21 percent of test-takers intending to enter the biological sciences achieved a score above 700 compared to 52 percent in computer and information sciences.
CONCLUSION AND RECOMMNENDATIONS FOR WAYS TO ENCOURAGE THE DEVELOPMENT OF NEW PROGRAMS
This paper explores four reasons why the current educational system appears to be sluggish in responding to the increased demand for individuals trained in computational biology. The first and second reasons are interrelated. Specifically, we argue that the size and direction of Ph.D. programs in the life sciences are more responsive to signals embedded in funding opportunities for faculty research than to the signals provided by the job market for graduates. While this may appear perverse, it is the logical consequence of a research regime that places great emphasis on having doctoral students and postdoctoral students in the lab and can persist as long as there is an adequate supply of applicants. Such a supply has been forthcoming in the United States in recent years because of (1) the “hot” reputation of biotechnology; (2) the availability of immigrant scientists and (3) the ready supply of postdoctoral positions that permit graduate schools to provide placement for graduates.
The third reason for the sluggish response relates to the interdisciplinary nature of bioinformatics. Given the fields involved (mathematics, computer science, and biology), collaboration typically requires working across college lines within a university. While this is not impossible, the bureaucracy and incentive structure of academe act to discourage cooperation across disciplines. Finally, we have argued that there is no “quick fix.” Individuals trained in computer science have few economic incentives to change their stripes by acquiring additional training in biology. And, if they did, the response would be far from quick since they would require a substantial amount of training in biology. In contrast, the loose labor market for young life scientists means that the incentive is there for life scientists to augment their skills and become computational biologists. But for life scientists the path may be difficult since many lack both the mathematical training and the inclination to become successful computational biologists.
We end by proposing four possible ways to address the apparent shortage of individuals trained in bioinformatics. First, it is important to foster ways for faculty to interact across disciplines and institutional boundaries both within universities and within urban areas. Providing space for interdisciplinary efforts could encourage such interaction; the “if you build it they will come and they will talk” argument. Second, provide funds for targeted research in the area and continue to provide funds for training awards and institutional awards. While training grants can affect outcomes in the long run, there is nothing like targeted research awards to get the attention of faculty in the short run. Third, provide information to students on career outcomes in bioinformatics. This involves making faculty aware of the career outcomes of their students so that faculty can help prospective students make well-informed decisions. Finally, in an emerging field such as bioinformatics, it is important to recruit early in the pipeline in order to attract the “right” kind of mind. Perhaps it is not surprising that the University of Pennsylvania has decided that the best way to meet the demand for computational biologists is to “grow their own,” offering undergraduate and master’s programs in computational biology in order to attract the “right” kind of mind and integrate the curriculum at an early stage.28
REFERENCES
Educational Testing Service. 1997a. 1997–98 Guide to the Use of Scores. Princeton, New Jersey: Educational Testing Service.
Educational Testing Service. 1997b. Sex, Race, Ethnicity, and Performance on the GRE General Test. Princeton, New Jersey: Educational Testing Service.
Gavaghan, Helen. 1997. “Running to Catch Up in Europe.” Nature, 389:420–422.
Gershon, Diane. 1997. “Bioinformatics in a Post-genomics Age.” Nature. 389:417–418.
Goldberger, Marvin L., Brendan A.Maher, and Pamela E.Flattau (eds.). 1995. Research-Doctorate Programs in the United States: Continuity and Change. Washington, D.C.: National Academy Press.
Hicks, D. and J.S.Katz. 1996. “Science Policy for a Highly Collaborative Science System.” Science and Public Policy. 23:39–44.
Hollingsworth, Rogers. 1995. “Major Discoveries and Biomedical Research Organizations: Perspectives on Interdisciplinarity, Nurturing Leadership, and Integrated Structure and Cultures,” prepared exclusively for Interdisciplinarity Project and to be published by University of Toronto Press.
Kaiser, Jocelyn (ed.). 1998. “Hopkins’s Genetic Database to Close.” Science. 279:645.
Katz, J.S., D.Hicks, M.Sharp, and B.R.Martin. 1995. The Changing Shape of British Science. Brighton: Science Policy Research Unit at the University of Sussex.
Malakoff, David. 1999. “NIH Urged to Fund Centers to Merge Computing and Biology.” Science. (June 11, 1999): 1742.
Marshall, Eliot. 1996. “Hot Property: Biologists Who Compute.” Science. June 21:1730–32.
Marshall, Eliot. 1996. “Demand Outstrips Supply.” Science. June 21:1731.
National Research Council. 1995. Research Doctorate Programs in the United States: Continuity and Change. M.Goldberger, B.Maher, and P.Ebert, editors. Washington, D.C.: National Academy Press.
National Research Council. 1998. Trends in the Early Careers of Life Scientists. Committee on Dimensions, Causes, and Implications of Recent Trends in the Careers of Life Scientists. Washington, D.C.: National Academy Press.
National Science Foundation. 1997. Characteristics of Doctoral Scientists and Engineers in the United States: 1995, Detailed Statistical Tables. Arlington, VA: National Science Foundation.
National Science Foundation. 1997. Characteristics of Recent Science and Engineering Graduates: 1995, Detailed Statistical Tables . Arlington, VA: National Science Foundation.
National Science Foundation. 1996. Science and Engineering Doctorate Awards: 1996, Detailed Statistical Tables. Arlington, VA: National Science Foundation.
Stephan, Paula and Grant Black. 1999. “Bioinformatics: Does the U.S. System Lead to Missed Opportunities in Emerging Fields? A Case Study.” Science and Public Policy, (December): 1–15.
Stephan, Paula and Grant Black. 1999. “Hiring Patterns Experienced by Students Enrolled in Bionformatics/Computational Biology Programs.” Report to the Alfred P.Sloan Foundation. May.
Wickware, Potter. 1997. “Choices and Challenges.” Nature. 389:420.
Yap, Ting K., Frieder Ophir, and Robert L.Mantino. 1996. High Performance Computational Methods for Biological Sequence Analysis, Boston: Kluwer Academic Publishers.
Yee, Wendy. “The Top Five Career Trends of 1996: Informatics Anything,” http:\www.nextwave.org/server-java/SAM/pastloop/trend2.htm.

















