4
Chromosome Speciation: Humans, Drosophila, and Mosquitoes
FRANCISCO J. AYALA* AND MARIO COLUZZI†
Chromosome rearrangements (such as inversions, fusions, and fissions) may play significant roles in the speciation between parapatric (contiguous) or partly sympatric (geographically overlapping) populations. According to the “hybrid-dysfunction” model, speciation occurs because hybrids with heterozygous chromosome rearrangements produce dysfunctional gametes and thus have low reproductive fitness. Natural selection will, therefore, promote mutations that reduce the probability of intercrossing between populations carrying different rearrangements and thus promote their reproductive isolation. This model encounters a disabling difficulty: namely, how to account for the spread in a population of a chromosome rearrangement after it first arises as a mutation in a single individual. The “suppressed-recombination” model of speciation points out that chromosome rearrangements act as a genetic filter between populations. Mutations associated with the rearranged chromosomes cannot flow from one to another population, whereas genetic exchange will freely occur between colinear chromosomes. Mutations adaptive to local conditions will, therefore, accumulate differentially in the protected chromosome regions so that parapatric or partially sympatric populations will genetically differentiate, eventually evolving into
different species. The speciation model of suppressed recombination has recently been tested by gene and DNA sequence comparisons between humans and chimpanzees, between Drosophila species, and between species related to Anopheles gambiae, the vector of malignant malaria in Africa.
The process of evolution is continuous through time but yields in space discontinuous groups of organisms. The continuity of the process links the myriad living organisms with the last universal common ancestor, from which all living organisms descend. Organisms evolve differences because of the haphazard mutation process, adaptation to different environmental circumstances, interaction with other organisms, constraints imposed by the organisms’ past evolutionary history, and the like. The discontinuities are encompassed in the Linnean system of classification, which is hierarchical, with gradually more inclusive categories: species, genus, family, order, and so on.
“Species” is a category of classification, the most basic, within which are placed groups of organisms designated by specific names such as Homo sapiens, Mus musculus, Drosophila melanogaster, or Quercus borealis. But species have a biological reality that is lacking in more inclusive groups of organisms. In sexually reproducing organisms, individual members of a species are able to interbreed and thus share in a common gene pool. Collectively, there is variation among the members of a species, but there is also continuity in space and time. Species are evolutionary units. Because of these properties, some philosophers have affirmed that species, but not more inclusive groups of organisms, are metaphysical individuals. According to Hull (1977), “species fit as naturally into the idealized category of spatiotemporally localized individuals as do particular organisms” (Ghiselin, 1974).
Dobzhansky (1935a,b) pointed out in 1935 this double biological reality of the concept of species: (i) as a category of classification, just like genus, family, and other categories, a logical construct pragmatically necessary for organizing the enormous diversity of the living world and (ii) as a category with “an attribute peculiar only to itself” (1935b), because a species is a natural entity, a collectivity that has biological continuity defined, in sexually reproducing organisms, by the capacity to interbreed among individuals of the same species and their incapacity to interbreed with individuals of other species. The biological species concept, as it came to be known, defines species precisely by these two attributes: ability to interbreed within the species and reproductive isolation from other species. The evolutionary process of speciation, by which one species splits into two, is equivalent to the evolutionary emergence of reproductive isolation.
Dobzhansky (1935a) described a species as “a group of individuals fully fertile inter se, but barred from interbreeding with other similar groups by its physiological properties.” But Dobzhansky was aware that complex concepts such as species cannot satisfactorily be defined by a particular set of words encompassed in a single sentence. He did not place the definition cited above in quotes and provided, in the same and other writings, additional definitions that pointed toward other species characteristics such as their being temporary instantiations of the evolutionary process: “Considered dynamically, the species represents that stage of evolutionary divergence at which the once actually or potentially interbreeding array of forms becomes segregated into two or more separate arrays which are physiologically incapable of interbreeding” (1935a). This definition is quoted by Dobzhansky (1937b) in the first edition of his classic Genetics and the Origin of Species, in a chapter significantly titled “Species as Natural Units.” He adds: “Species is a stage in a process, not a static unit” (1937b). As he had earlier pointed out: “The fundamental importance of this stage is due to the fact that it is only the development of the isolating mechanisms that makes possible the coexistence in the same geographic area of different discrete groups of organisms…. This, in turn, opens the possibility for the organisms dwelling together to become adapted to different places in the general economy of nature” (1935a). Species are natural units that evolve and adapt autonomously. Dobzhansky noted that “a stage must exist in the process of evolutionary divergence at which an originally panmictic population becomes split into two or more populations that interbreed with each other no longer…. The emphasis should be placed, however, not on the absence of actual interbreeding between the different form complexes, but rather on the presence of physiological mechanisms making interbreeding difficult or impossible” (1935a). In Genetics and the Origin of Species (1937b), a full chapter is dedicated to “isolating mechanisms,” a label that he had first proposed in Dobzhansky (1937a).
For nearly four decades, Dobzhansky continued to use similarly worded definitions of species, with the phrases “reproductive community” or “Mendelian population” sometimes replacing or added to his earlier “group of organisms” or “array of forms.” In his last major work, Genetics of the Evolutionary Process (1970), Dobzhansky writes: “the process of species formation, in contrast to race formation, involves the development of reproductive isolating mechanisms. An ancestral species is transformed into two or more derived species when an array of interbreeding Mendelian populations becomes segregated into two or more reproductively isolated arrays. Species are, accordingly, systems of populations; the gene exchange between these systems is limited or prevented in nature by a reproductive isolating mechanism or perhaps by a combi-
nation of several such mechanisms. In short, a species is the most inclusive Mendelian population.” Moreover, he again asserts: “Species is not only a category of classification, but also a form of supraindividual biological integration” (1970).
In Systematics and the Origin of Species, Mayr (1942) commended Dobzhansky for identifying interbreeding and reproductive isolation as the distinguishing features of the species concept and proposed a short definition: “Species are groups of actually or potentially interbreeding natural populations which are reproductively isolated from other such groups.” He has used, identically or with some word differences, this definition in later writings. Indeed, Mayr is generally perceived as the leading exponent of the biological species concept and the most successful investigator of the application of this concept to a great variety of species and species groups throughout the animal world, as several papers in this collection bear witness.
Mayr repeatedly wrote that species are real and not merely human constructs that are convenient for organizing biological diversity, as some taxonomists, as well as nominalist philosophers, would claim. He supported the claim by Ghiselin (1974), Hull (1977), and others that species are metaphysical individuals, once this language was introduced in the evolutionary literature (Mayr, 1976, 1987). The integration of its gene pool provides the necessary cohesion for any particular species taxon to be considered an “individual.” The integration of gene pools, in turn, derives from the two dimensions incorporated in the definition of species, the ability of its members to interbreed, and their reproductive isolation from other species.
MODELS OF CHROMOSOMAL SPECIATION
Changes in chromosome number or structure may contribute to speciation. Polyploidy, the multiplication of the chromosome complement, may yield a new species in a single generation, reproductively isolated from its ancestral species. For example, a tetraploid plant crossed with a diploid ancestor produces sterile hybrid progeny. Polyploidy is more common among angiosperms than among gymnosperms. Nearly 50% of all existing angiosperm species are estimated to have arisen by ancient polyploidy, more of them by allopolyploidy (doubling of the chromosome complement in a hybrid between two previously existing species) than by autopolyploidy (multiplication of the chromosome complement of a single species). Polyploidy is also common among ferns. Some important cultivated plants are polyploids, such as wheat, oat, tobacco, potato, banana, strawberry, sugar cane, and coffee. Polyploidy is less common in animals; polyploidy species occur among hermaphrodites, such as earth-
worms and planarians, or in species with parthenogenetic females, such as some shrimps, sow bugs, moths, and, more commonly, beetles, as well as some fish and salamanders.
Chromosome rearrangements, such as Robertsonian fusions and fissions, translocations, and inversions, may play a role in speciation. There are a number of models proposing that chromosomal rearrangements accelerate genic diversification between populations and, therefore, facilitate speciation. We will consider two classes of models, which we will call the “hybrid-dysfunction” and “suppressed-recombination” models of speciation.
Hybrid-dysfunction models claim that recombination between rearranged chromosomes generates gametes with some chromosomal segments deleted and others duplicated, thereby creating a partial reproductive barrier because the heterokaryotypic hybrid exhibits reduced reproductive fitness, also called “underdominance.” Under these conditions, natural selection will, in both populations, favor mutations that reduce the probability of intercrossing and will eventually lead to complete reproductive isolation. The great multiplication of species of flightless Australian grasshoppers of the subfamily Morabinae can largely be attributed, according to White (1968, 1978), to underdominance in hybrids between populations with different chromosome rearrangements. A chromosomal rearrangement may first become established in a small local colony, either at the periphery of the distribution area of the ancestral species or inside it, by random drift. The colony may expand within a certain area and there displace the ancestral form if its members display high fitness in that area. The low fitness of the hybrids will keep the two populations separate and facilitate the evolution of prezygotic isolating mechanisms, which will inhibit the formation of hybrids. White (1968, 1978) refers to this model of speciation as “stasipatric,” because it largely occurs in situ, rather than allopatrically, yet it differs from models of sympatric speciation that do not attribute a major role for chromosome rearrangements.
The hybrid dysfunction model of speciation encounters the following disabling difficulty (Hey, 2003; Machado et al., 2002; Navarro and Barton, 2003a,b; Noor et al., 2001b; Rieseberg, 2001; Spirito, 2000). A chromosome rearrangement will first appear in the population as a mutation in a single individual. This individual will be able to mate only with individuals without the mutation. If hybrids have reduced fitness, the chromosome mutation will be selected against and eliminated from the population. The hybrid dysfunction model is unlikely to have much general validity, precisely because it seems so unlikely that a chromosome rearrangement that reduces the fitness of heterozygotes will be at all established within its ancestral population, although this may occasionally occur by random
drift, particularly if hybrid underdominance is only slight (in which case natural selection for reproductive isolation will be weak as well).
A speciation model of suppressed recombination was proposed by Coluzzi (1982) in his account of multiple speciation events within the species complex related to Anopheles gambiae, the main vectors in Africa for the transmission of malignant malaria, caused by the protozoan Plasmodium falciparum. The World Health Organization estimates that there are annually 300–500 million cases of malaria and >1 million deaths, mostly in sub-Saharan Africa, and the most of those who die are children. Seven species have been identifiedwithin the A. gambiae complex and have arisen, according to Coluzzi (1982; Coluzzi et al., 2002), within the last 5,000 years.
Suppressed-recombination models of speciation have recently been proposed by Rieseberg (2001) to account for speciation in wild sunflowers (Riesberg et al., 1995); by Noor et al. (2001a,b) and Machado et al. (2002), using evidence from the closely related Drosophila persimilis, Drosophila pseudoobscura, and D. p. bogotana; and by Navarro and Barton (2003a,b), who have mathematically modeled the process and supported the model’s predictions by comparing genomic DNA sequences between humans and chimpanzees.
We will successively examine the human–chimpanzee and Drosophila evidence advanced in support of the model and then return to speciation in the A. gambiae complex.
HUMAN SPECIATION
Genomic studies have confirmed that a substantial number of chromosomal rearrangements have occurred between humans and chimpanzees. In particular, nine chromosomes (nos. 1, 4, 5, 9, 12, 15, 16, 17, and 18) exhibit pericentric inversions between humans and chimpanzees, and human chromosome 2 represents a fusion of two acrocentric chromosomes present in chimpanzees (chromosomes 12 and 13) and other great apes (chromosomes 11 and 12 in gorillas and orangutans) (de Boer and Seuánez, 1982; Yunis and Prakash, 1982). If these chromosomal rearrangements occurred early in the divergence between ancestral populations of chimps and humans, they “would facilitate genic divergence during the time when the diverging populations are in parapatry, i.e., have limited gene flow” (Navarro and Barton, 2003b). The hypothesis proposes that alleles favored in one or the other population will be trapped at the chromosomal barrier and thus would cause the two populations to diverge genically as they adapt to their distinct prevailing environmental conditions. Accumulation of incompatibilities would gradually result in reproductive isolation and speciation. In regions not protected by chromosome
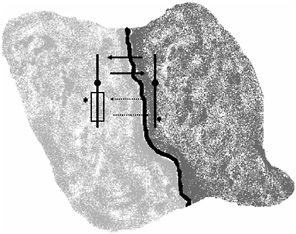
FIGURE 4.1 Two populations share a common boundary where hybridization occurs. Shown are two metacentric chromosomes that differ by an inversion (box) and incompatible alleles at two loci (*). Gene flow can readily occur along regions not linked to the inverted region (solid arrows) but is severely inhibited in regions linked to the inversion (dotted arrows). Natural selection favors the evolution of reproductive isolation between the populations by accumulation of incompatible alleles along the chromosome regions protected from recombination by the inversions. Figure was modified from Hey (2003).
rearrangements, allelic differences that might arise would not readily accumulate, because genetic recombination would tend to diffuse them between the populations (Fig. 4.1). This hypothesis can be tested, according to Navarro and Barton (2003b), by comparing genic differentiation between humans and chimps for different chromosome regions.
According to evolutionary theory, the rate of nonsynonymous nucleotide substitution per nonsynonymous site (KA) is generally expected to be much lower than the rate of synonymous substitution per synonymous site (KS), because random amino acid changes are usually deleterious, whereas synonymous changes are likely to be neutral or nearly so (Kimura, 1983). Thus, the expectation is KA ≪ KS, except when positive selection is involved favoring particular amino acid replacements, in which case KA will increase and may even become larger than KS. KA/KS ratios close to or >1 indicate positive selection.
Navarro and Barton (2003b) have investigated nucleotide sequences that exhibit nucleotide differences between chimps and humans in 115 genes, about evenly distributed between rearranged chromosomes (59 genes) and colinear chromosomes (56 genes) (Table 4.1). Of the 26 genes with KA/KS ratios ≤1, 20 (76.9%) are located on rearranged chromosomes,
TABLE 4.1 Rate of Gene Evolution for Rearranged and Colinear Chromosomes in Humans Versus Chimpanzees
|
KA/KS |
Rearranged Chromosomes |
Colinear Chromosomes |
Total |
|
Number of genes (all) |
59 |
56 |
115 |
|
>1 |
20 |
06 |
026 |
|
<1 |
39 |
50 |
089 |
|
Number of genes (KS > 0) |
54 |
54 |
108 |
|
>1 |
15 |
04 |
019 |
|
<1 |
39 |
50 |
089 |
|
KA/KS |
00.84 |
00.37 |
000.61 |
|
Data are from Navarro and Barton (2003b). KA and KS are rates of evolution for nonsynonymous and synonymous nucleotides, respectively. |
|||
and only 6 are in colinear chromosomes. Of 89 chromosomes with KA/KS ratios >1, 39 (43.8%) are on rearranged chromosomes, and 50 are on colinear chromosomes. The average KA/KS ratio for rearranged chromosomes (0.84) is more than double the value for colinear chromosomes (0.37). (These ratios do not include seven genes for which KS = 0, which would have given a ratio of infinity: five genes in rearranged chromosomes and two in colinear chromosomes.)
The KA/KS ratios and related results are consistent with the hypothesis tested, the suppressed-recombination model of speciation, although Navarro and Barton (2003b) also discussed other factors that may have been involved. The 2-fold difference between the two ratios is surprising, because it is so large. As these researchers note, and as also interpreted by Rieseberg and Livingstone (2003), a 2-fold difference could be explained, under the hypothesis, only if the chromosomal rearrangements have been barriers in parapatry and have exchanged genes through hybridization no less than half the time of divergence between the human and chimpanzee lineages. The human and chimpanzee lineages diverged 7–8 million years ago. Sahelanthropus tchadensis and Orrorin tugenensis are the oldest known taxa in the human lineage. They lived 6–7 million years ago and were prevailingly bipedal, at least when they were on the ground. Ardipithecus ramidus, Australopithecus anamensis, Australopithecus afarensis, and Kenyanthropus platyops count among the species of the human lineage that lived between 3.5 million and 5.5 million years ago, although not all of them may have been direct ancestors of Homo sapiens. These taxa were prevailingly or exclusively bipedal. For example, the pelvis of Lucy, a well-known specimen of A. afarensis, dated ≈3.5 million years ago, is very similar in configuration and proportions to a modern human pelvis and,
in any case, drastically different from that of a chimpanzee or any other modern ape. Given these and other anatomical incompatibilities, the inference that human and chimp ancestors may have hybridized for 3–4 million years after the first divergence of their lineages seems, prima facie, unlikely.
The inferences made by Navarro and Barton (2003b) encounter other theoretical difficulties. The model of suppressed recombination predicts that once incompatible alleles start to accumulate, the process will accelerate so that an increased fraction of allele substitutions in one of the species would be incompatible with the other species and speciation would rapidly occur (Hey, 2003; Orr and Turelli, 2001). Therefore, the period during which genetic differences accumulate preferentially in the rearranged chromosomes should not last very long. Moreover, millions of years have elapsed of separate evolution between the human and chimpanzee lineages, which should have largely erased the signal predicted by the model: namely, the expected greater genic differentiation between the rearranged chromosomes than between the colinear chromosomes, because this differentiation would have occurred so long ago, and other processes would have largely contributed to the current genetic differentiation between the two lineages. One additional difficulty is the implied assumption that the chromosomal rearrangements that differentiate humans from chimps and other apes happened all in the human lineage, early and within a short time.
Several ensuing empirical investigations have brought into question the inferences of Navarro and Barton (2003b). Lu et al. (2003) have estimated KA, KS, and KA/KS in 85 genes, about equally distributed between rearranged chromosomes (42 genes) and colinear chromosomes (43 genes). Between humans and chimpanzees, the KA/KS ratio is 1.41 times greater (0.820 versus 0.581) for rearranged chromosomes than for colinear chromosomes (Table 4.2), consistent with the result of Navarro and Barton (2003b), although the difference is smaller than the >2-fold (2.2) increase observed by these researchers. Further, as a control, they have compared the same 85 genes between humans and other primate species, namely, 21 genes from either orangutan or gibbon, plus 64 genes from Old World monkeys. Remarkably, the KA/KS ratios are also 1.41 times greater for rearranged chromosomes than for colinear chromosomes in humans (0.623 versus 0.443) (Table 4.2). This result is consistent with the hypothesis that positive selection may have more intensely promoted genic evolution in the rearranged chromosomes but not with Navarro and Barton’s (2003b) hypothesis that the high KA/KS ratios are associated with speciation between humans and chimpanzees according to the suppressed-recombination model. Lu et al. (2003) suggest that the higher KA/KS ratios observed in rearranged chromosomes may simply reflect a bias in the distribution of rapidly evolving
TABLE 4.2 Rate of Gene Evolution for Rearranged (R) and Collinear (C) Chromosomes in Humans Versus Chimpanzees or Other Primates
|
Rate of Evolution |
Rearranged |
Colinear |
R/C |
|
Human: chimpanzee (all genes, 109) |
|||
|
KA/KS |
0.780 (54) |
0.483 (55) |
1.61 |
|
KA |
0.007 (54) |
0.005 (55) |
1.40 |
|
KS |
0.016 (54) |
0.017 (55) |
0.94 |
|
Human: chimpanzee (overlapping genes, 85) |
|||
|
KA/KS |
0.820 (42) |
0.581 (43) |
1.41 |
|
KA |
0.007 (42) |
0.006 (43) |
1.16 |
|
KS |
0.016 (42) |
0.014 (43) |
1.14 |
|
Human: outgroup primates (overlapping genes, 85) |
|||
|
KA/KS |
0.623 (42) |
0.443 (43) |
1.41 |
|
KA |
0.031 (42) |
0.024 (43) |
1.30 |
|
KS |
0.061 (42) |
0.059 (43) |
1.03 |
|
Data are from Lu et al., 2003. KA and KS are as defined in Table 1. Number of genes is given in parentheses. |
|||
genes among chromosomes. Genes in the rearranged chromosomes may happen to be evolving faster than genes in colinear chromosomes. Faster evolution is known to be the case for certain genes; for example, genes encoding glycophorins and protamines, which happen to be located on rearranged chromosomes, are known to be rapidly evolving in all higher primates (Lu et al., 2003).
Zhang et al. (2004) have examined nucleotide differentiation between human and chimpanzee DNA sequences amounting to 4,108,833 nucleotides: 1,831,676 nucleotides distributed among seven rearranged chromosomes, and 2,277,157 nucleotides distributed among six colinear chromosomes. The average nucleotide divergence per chromosome is 1.20 ± 0.14% for the rearranged chromosomes and larger (1.40 ± 0.08%) for the colinear chromosomes (Table 4.3). This finding is inconsistent with Navarro and Barton’s (2003b) hypothesis; indeed, the rate of nucleotide evolution is faster in the colinear chromosomes than in the rearranged
TABLE 4.3 Nucleotide Sequence Divergence Between Humans and Chimpanzees
|
Chromosomes |
Number of Nucleotides |
Percent Divergence |
|
Rearranged chromosomes |
1,831,676 |
1.20 ± 0.14 |
|
Colinear chromosomes |
2,277,157 |
1.40 ± 0.08 |
|
Total |
4,108,833 |
1.29 ± 0.09 |
|
Data are from Zhang et al. (2004). |
||
TABLE 4.4 Nonsynonymous (N) and Synonymous (S) Nucleotide Divergence Between Humans and Chimpanzees
|
Chromosomes |
Number of Genes |
Number of Nucleotides |
N |
S |
N/S |
dN/dS |
|
Rearranged |
568 |
202,455 |
288 |
596 |
0.483 |
0.236 ± 0.028 |
|
Colinear |
512 |
184,728 |
288 |
532 |
0.541 |
0.239 ± 0.019 |
|
Total |
1,080 |
387,183 |
576 |
1,128 |
0.510 |
0.237 ± 0.016 |
|
Data are from Zhang et al. (2004). dN and dS are the rates of nonsynonymous and synonymous substitution, respectively. |
||||||
chromosomes. The nucleotide sequences analyzed may have largely involved noncoding DNA, thus evolving mostly neutrally. But the suppressed recombination model predicts that the rate of molecular evolution, neutral or not, should be greater in rearranged chromosomes than in colinear chromosomes.
A comparison of the number of nonsynonymous (n) versus synonymous substitutions between humans and chimpanzees by using cDNA sequences and thus involving protein coding genes has also been performed by Zhang et al. (2004). The total number of nucleotides examined is approximately the same for the 10 rearranged chromosomes (202,455 nucleotides) as for the 13 colinear chromosomes (184,728 nucleotides), and so is the number of genes in each class (568 versus 512 genes in rearranged versus colinear chromosomes). The ratio of nonsynonymous to synonymous substitutions is 0.483 in rearranged chromosomes, and the ratio is 0.541 in colinear chromosomes (Table 4.4). This result is contrary to the one observed by Navarro and Barton (2003b) and is inconsistent with their hypothesis of an accelerated rate of evolution in the rearranged chromosomes. An additional analysis involving 304 gene sequences of the ratio of nonsynonymous to synonymous rates of evolution yields a higher proportion of genes under positive selection in colinear chromosomes (29.5% of genes with ratios >1) than in rearranged chromosomes (15.5% of genes with ratios >1) (Table 4.5).
Cáceres et al. (2003) have importantly discovered that a large majority (≈90%) of 169 genes with a great variety of functions exhibit higher expression levels in the cerebral cortex of humans than in the cerebral cortex of chimpanzees, which suggests elevated levels of neuronal activity in the human brain. Marquès-Bonet et al. (2004) have distributed the genes analyzed by Cáceres et al. (2003) into those present in colinear and rearranged chromosomes, observing that the average proportional increase of expression in humans is 1.463 ± 0.014 (mean ± SE) in colinear chromosomes but 1.543 ± 0.019 in rearranged chromosomes, a difference that is statisti-
TABLE 4.5 Number of Genes Under Positive Selection (dN/dS > 1) in Rearranged (R) Versus Colinear (C) Human Chromosomes
|
dN/dS |
Rearranged |
Colinear |
R/C |
|
Total genes |
148 |
156 |
304 |
|
>1 |
23 (15.5) |
46 (29.5) |
0.500 |
|
<1 |
125 (84.5) |
110 (71.5) |
1.136 |
|
Data are from Zhang et al. (2004). dN and dS are as defined in Table 4.4. Percentages are given in parentheses. |
|||
cally significant by a permutation test. They interpret these results as supporting their theory of chromosomal speciation by suppressed recombination. However, Zhang et al. (2004) have analyzed the same data and similarly observed increased expression in rearranged chromosomes (80 genes) relative to colinear chromosomes (72 genes), but neither the difference nor the average of the differences between individual chromosomes is statistically significant. Table 4.6 shows that genes in rearranged chromosomes 4, 5, and 9, as well as in colinear chromosome 22, have human expression levels >1.6 times higher than in chimpanzees, but in none of these chromosomes is the expression level significantly higher than the average for all genes of 1.503 (1.463 and 1.543 for colinear and rearranged chromosomes, respectively). The importance of Cáceres et al. (2003) is the higher expression levels in the human cerebral cortex, but gene expression increases may have happened at various times. The evidence shown in Table 4.6 does not warrant the conclusion that gene expression increases in the human cortex were distinctly promoted in rearranged chromosomes at the time when the human lineage first separated from the chimpanzee lineage some 7–8 million years ago. Brain size in the human lineage notably increased starting with Homo habilis, ≈2 million years ago. It may very well have been the case that gene expression increases in the cerebral cortex were, at least in good part, associated with the increase, by about a factor of 3, that happened in the human lineage during the last 2 million years.
Any increased genic differentiation that may have preferentially occurred in rearranged chromosomes during the original divergence of humans and chimps would likely be undetectable after millions of years of further divergent evolution in the two lineages. Among the apes, there are favorable populations for testing the suppressed-recombination model of speciation, such as the eight Bornean subspecies or races of orangutans that are parapatric in their distribution, with borderlines formed by rivers [the eight subspecies have been given formal taxonomic names (von
TABLE 4.6 Average Proportional Increase in Gene Expression in the Cerebral Cortex of Humans Compared with Chimpanzees
|
Chromosomes |
Colinear |
Rearranged |
|
1 |
|
1.55 ± 0.04 |
|
2 |
1.45 ± 0.03 |
|
|
3 |
1.46 ± 0.03 |
|
|
4 |
|
1.62 ± 0.08 |
|
5 |
1.67 ± 0.08 |
|
|
6 |
1.43 ± 0.03 |
|
|
7 |
1.51 ± 0.05 |
|
|
8 |
1.41 ± 0.04 |
|
|
9 |
|
1.66 ± 0.10 |
|
10 |
1.38 ± 0.03 |
|
|
11 |
1.42 ± 0.03 |
|
|
12 |
|
1.43 ± 0.04 |
|
13 |
1.45 ± 0.05 |
|
|
14 |
1.50 ± 0.06 |
|
|
15 |
|
1.50 ± 0.07 |
|
16 |
1.56 ± 0.07 |
|
|
17 |
1.53 ± 0.05 |
|
|
18 |
1.54 ± 0.09 |
|
|
19 |
1.53 ± 0.05 |
|
|
20 |
1.41 ± 0.04 |
|
|
21 |
1.48 ± 0.07 |
|
|
22 |
1.63 ± 0.10 |
|
|
Total |
1.463 ± 0.014 |
1.543 ± 0.019 |
|
Data are from Marquès-Bonet et al. (2004). |
||
Koenigswald, 1982)]. Pericentric inversions, such as those between humans and chimpanzees, are common among the great apes, including those in chromosomes 2 and 3 that differentiate the Sumatran and Borneo subspecies of orangutans; polymorphic pericentric inversions occur within each one of the two subspecies, for instance, in chromosome 9 (de Boer and Seuánez, 1982).
SPECIATION IN DROSOPHILA
D. pseudoobscura is a widely distributed Nearctic species common in temperate forests throughout the western third of North America, extending from British Columbia to Guatemala and from the Pacific to the Great Plains. A geographically isolated subspecies, D. p. bogotana, lives in the altiplano near Bogota, Colombia. More narrowly distributed than D. pseudoobscura is its sibling species, D. persimilis, common in the temperate forests of the American northwest, from British Columbia to southern
California. Throughout this territory, the two species are sympatric and abundant, but D. persimilis tends to be more common in cooler, moister environments. D. persimilis and D. pseudoobscura are reproductively isolated by a cascade of reproductive isolating mechanisms that include sexual isolation, hybrid male sterility, hybrid male courtship dysfunction, and hybrid backcross inviability. Hybrids have been found in nature at extremely low frequencies (Dobzhansky, 1973; Powell, 1997), but they evince that there is a possibility for gene introgression across the species by means of backcrosses of hybrid females to the parental species. Interspecific hybrid females are fertile, even though males are sterile.
The nuclear genome of the species is distributed in five large chromosome arms and one very small chromosome. The X chromosome is metacentric, incorporating two chromosome arms. The two species differ by fixed large paracentric inversions on the left arm of chromosome X and on chromosome 2 (Anderson et al., 1977; Moore and Taylor, 1986). The genes associated with isolating mechanisms between D. pseudoobscura and D. persimilis are largely located on these chromosome inversions, with no evidence of major barriers decreasing the potential for gene flow across the other major chromosome arms, just as would be expected according to the suppressed-recombination model of speciation (Noor et al., 2001a,b). Fixed inversion polymorphisms inhibiting recombination would facilitate genic differentiation along the inverted segments, where genes promoting reproductive isolation between the species would have gradually accumulated. Consistent with this interpretation is the observation that the reproductive isolation factors between the allopatric pair, D. persimilis and D. p. bogotana, are located in the colinear regions of their chromosomes rather than on the inverted segments (Brown et al., 2004), demonstrating that the protection against recombination provided by chromosomal inversions has played no particular role in promoting reproductive isolation between these two populations, which are not geographically contiguous or overlapping.
The divergence between D. pseudoobscura and D. persimilis is estimated, by molecular and other information, to have occurred some 500,000 years ago; the separation between the two subspecies D. p. pseudoobscura and D. p. bogotana is dated ≈150,000 years ago, when a propagule of the species somehow colonized the altiplano near Bogota (Brown et al., 2004; Wang et al., 1997). Machado et al. (2002) have examined DNA sequence variation at 11 loci in a large sample of strains geographically representative of the three taxa. Their results are consistent with gene flow between the largely sympatric D. pseudoobscura and D. persimilis but not between the allopatric D. p. pseudoobscura and D. p. bogotana. The evidence indicates that most of the gene flow is ancient rather than recent. Most importantly, as predicted by the suppressed-recombination model
of speciation, genomic regions associated with reproductive isolation between D. pseudoobscura and D. persimilis show less evidence of gene flow than regions not so associated.
SPECIATION IN MALARIA’S MOSQUITOES
Fig. 4.2 shows the geographic distribution of seven species of the A. gambiae complex, and Fig. 4.3 shows their phylogeny. Epidemiologically most important are A. gambiae and Anopheles arabiensis. Where the geographic distributions of these two species overlap, there is competitive exclusion between them, with A. gambiae prevailing in the rain forests and other messic habitats and A. arabiensis prevailing in xeric habitats. The seven species are siblings that are morphologically nearly indistinguishable, but they differ in genetic and ecological attributes, including breeding sites, as well as egg configuration and some subtle morphological traits. Chromosome rearrangements are common, so that the species are identified by their chromosome configurations, primarily inversion polymorphisms of the second and third chromosomes, which are particularly extensive in A. gambiae and A. arabiensis and are distinctive for each species (Coluzzi and Bradley, 1999; Coluzzi et al., 2002).
A. arabiensis is considered the most likely ancestral species, which may have originated in the Middle East and reached Africa through the Arabian peninsula. Two sources of evidence support A. arabiensis as the ancestral species: It is the only member of the complex present in the Horn of Africa and in the Arabian peninsula, and it exhibits a fixed second-chromosome arrangement (labeled 2La) (Coluzzi et al., 2002), which is thought to be ancestral because it is also present in other species groups such as the Anopheles subpictus complex, where it is fixed in at least one of the siblings. Various sources of evidence indicate that A. arabiensis was originally zoophilic and exophilic but acquired anthropophily and domesticity secondarily in Sudan and West Africa, the regions where this species exhibits its most extensive chromosome polymorphisms. A. arabiensis may have first dispersed in East Africa starting >6,000 years ago and soon reached Madagascar, where it remains zoophilic and exophilic, having failed there to adapt to human environments perhaps because low human density has not provided suitable selective pressure for this adaptation. In East Africa, where human density is higher, A. arabiensis gradually became anthropophilic, although never to the extent of A. gambiae.
Chromosome inversion patterns indicate that A. arabiensis gave rise to Anopheles quadriannulatus, from which it notably differs by three X-chromosome inversions, where factors are located that account for the reproductive isolation between the two species and which has retained the ancestral condition of being zoophilic and exophilic. A. quadriannulatus
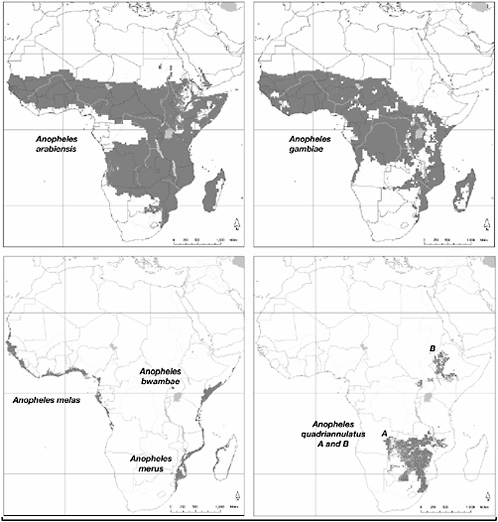
FIGURE 4.2 Geographic distribution of A. gambiae and six other closely related species. A. arabiensis, descended from a Pyretophorus species from the Arabian peninsula, is the likely ancestral species of the complex (see Fig. 4.3). Originally, A. arabiensis was zoophilic and exophilic, but it became anthropophilic and domestic by gradual adaptation to the human environment in Sudan and western Africa. A. quadriannulatus A and A. quadriannulatus B retain the original zoophily and exophily of their ancestral homonymous species, which also gave rise to A. bwambae and A. melas, and to A. gambiae, the most effective vector of malignant human malaria. A. gambiae and its strong anthropophily evolved <4000 B.P. with human invasion of the rain forest and introduction of slash-and-burn agriculture. In western Africa, A. gambiae is well represented in the Sahel region, extending up to 18° N, also the northern limit of A. arabiensis. In the Sudan, A. arabiensis, but not A. gambiae, is found along the river Nile upwards to the Egyptian border. Genetic data indicate that A. merus descends from A. gambiae and became adapted to breed in brackish, tide-dependent pools independently of A. melas.
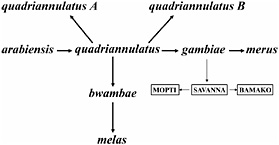
FIGURE 4.3 Phylogeny of seven species and three incipient species related to A. gambiae. The likely ancestral species is A. arabiensis, which differs from A. quadriannulatus by three X-chromosome inversions and differs from A. gambiae by two additional X-chromosome inversions. Reproductive factors among these species are primarily located on the X chromosome. Several incipient species can be recognized that are related to A. arabiensis, A. melas, or A. gambiae. Three incipient species related to A. gambiae, labeled Mopti, Savanna, and Bamako, are shown.
has split into two species, A. quadriannulatus A in southern Africa and A. quadriannulatus B in Ethiopia, which have homosequential chromosomes (although two polymorphic inversions are present in species A). These two allopatric species represent relics of the ancestral species, which genically diverged from each other after their geographic distribution became discontinuous. Two lineages originated from A. quadriannulatus: one giving rise to Anopheles bwambae, with a restricted geographic presence in northeast Uganda, and Anopheles melas, with a narrow distribution along the western coast of Africa, and a second, far more important lineage giving rise to A. gambiae.
The origins of A. gambiae can be traced to the late Neolithic, <4000 B.P. A. gambiae exhibits the primitive chromosome arrangement 2R (used within the complex as the standard of reference), which is adapted to the African rain forest, where, nevertheless, A. gambiae can only breed in environments modified by human agriculture, given that the larvae are “eliophilic,” requiring sunlight for breeding (Coluzzi et al., 2002). Agriculture was introduced in Africa ≈8000 B.P., imported from Mesopotamia into the lower Nile valley, but the forest remained for a long time impenetrable, without any traces of agricultural activity up to 4000 years B.P. Extensive penetration of the forest began ≈3000 B.P., made possible by climate change and the temporary “savannization” of much of the central African rain forest, a process which began ≈2800 B.P. and lasted ≈5 centuries (Willis et al., 2004). When the forest belt regained its original range of distribution, ≈2300 B.P., it was invaded by Bantu agriculturalists who adopted “slash-and-burn” agricultural techniques. Increase in rainfall and
the return of the forest were determinant factors for the spreading of the tze-tze fly (Glossina), vector of the lethal animal trypanosomiasis (Trypanosoma brucei), which decimated cattle, thus promoting the adaptation of A. quadriannulatus to humans, who became an easy host for the blood meal and, more importantly, caused the opening of the forest canopy. These conditions promoted strong selection for anthropophily and domesticity, which facilitated the evolution of A. gambiae (Coluzzi, 1982; Willis et al., 2004). There was a consequent increase in the rate of human infection by P. falciparum, which in turn led to strong selection for highly virulent strains, such as are the current forms of this parasite. Genetic evidence indicates that the expansion of malignant P. falciparum in Africa and throughout the world tropics occurred only within the last few thousand years (Hartl, 2004; Rich et al., 1998; Tishkoff et al., 2001).
A. gambiae differs from its ancestral A. quadriannulatus by two X-chromosome inversions, where factors responsible for their reproductive isolation are located (as is the case for the three X-chromosome inversions fixed between A. arabiensis and A. quadriannulatus; see above). A. gambiae has a nearly continentwide distribution in sub-Saharan African and is highly anthropophilic, like A. arabiensis, with which it accounts for most of the transmission in Africa of P. falciparum (but surpasses it). Further speciation is occurring within A. gambiae (as well as within A. arabiensis and A. melas). In southern Mali and northern Guinea, there are three chromosomally distinct populations of A. gambiae that are partially sympatric or parapatric and manifest enough assortative mating to qualify as incipient species. These populations are named “Savanna,” “Mopti,” and “Bamako” (Fig. 4.3). Chromosome-rearrangement evidence indicates that A. gambiae is the ancestor of A. merus, a species adapted to breed in tide-dependent pools.
A suppressed-recombination model of speciation was proposed by Coluzzi (1982) to account for the speciation patterns elucidated in the A. gambiae complex of species, particularly the successive adaptation of new species to environments quite different from the xeric conditions to which A. arabiensis was originally suited, such new environments as the rain forest or brackish waters. In outline, the essential components of the model are as follows (Coluzzi, 1982). A chromosomally monomorphic population colonizes a suitable environment and expands in numbers. The population will encounter adaptively marginal conditions, whether at the periphery of its distribution or at different times as a consequence of the seasons or other climatic and ecological oscillations. Mutations that increase adaptation to these marginal conditions will be favored wherever such conditions exist, yielding new “ecotypes” [i.e., “intraspecific groups having distinctive characters that result from the selective pressure of the local environment” (Lincoln et al., 1998)]. As the prevalence of the marginal
conditions oscillates, so will the subpopulations adapted to them, resulting in population flushes and crashes. Breeding between individuals from a subpopulation with those of the central population will tend to diffuse the alleles that are distinctively adapted to the marginal conditions, except for alleles that may have been captured within a chromosome rearrangement. Interbreeding between the central and the marginal population in parapatric zones of contact will thus homogenize their genetic makeup, except for the alleles protected by the chromosome rearrangements, where new adaptive alleles will accumulate, including those that will promote reproductive isolation. Reproductive isolation will gradually evolve, yielding incipient speciation and eventually full species.
Coluzzi (1982) explored the variety of environmental and genetic conditions that may yield various possible outcomes, the most significant of which are (i) speciation, as just described, (ii) extinction of the distinctive genotypes adapted to the marginal conditions, and (iii) incorporation of the rearranged chromosomes into the main population as an adaptive polymorphism, particularly when the heterozygotes exhibit overdominance. Speciation would facilitate adaptation to the originally marginal conditions, leading eventually to full exploitation of new environments or ecological niches. The fixed X-chromosome inversions that differentiate A. arabiensis, A. quadriannulatus, and A. gambiae include reproductive isolation factors between these species and thus would have been instrumental in the speciation process, according to the suppressed-recombination model. Fixed and polymorphic inversions in other chromosomes, particularly on the right arm of chromosome 2, characterize genetically distinct subpopulations that have adapted to different regions and niches. Some are associated with incipient speciation processes that still prevail within this young species complex (see Fig. 4.2 and Coluzzi et al., 2002).
The suppressed-recombination model of chromosome speciation predicts that sympatric sister species will be more different with respect to fixed chromosome rearrangements than allopatric sister species. Such is the state of affairs prevailing among the sibling species of the A. gambiae complex. Fixed rearrangements occur between sympatric sister species, but not between the two allopatric species A. quadriannulatus A and A. quadriannulatus B, relics of a widely distributed species that genically diverged allopatrically after their geographic distribution became discontinuous.
The suppressed-recombination speciation model also predicts that while gene exchange persists between the diverging populations, genes protected by the rearrangements will accumulate allelic differences faster than genes in the colinear chromosomes, where gene flow occurs between populations. The recent origin of seven species of the A. gambiae complex and the continuing processes of incipient speciation throughout the com-
plex provide an ideal system to test this prediction. Hybrid females are fertile between all of the species pairs of the complex. Therefore, the potential for gene flow persists up to the present, although hybrids are rarely detected in nature (0.02–0.76%) (Temu, 1997; Touré et al., 1998). In any case, the increased genic differentiation between the rearranged chromosomes that would have taken place during the speciation process should be detectable, given that it has occurred within the last few thousand years, rather than hundreds of thousands (as between D. pseudoobscura and D. persimilis) or millions of years (as between humans and chimpanzees) ago.
The currently available evidence is limited but consistent with the model. An investigation of DNA sequence variation in four gene loci (one from each of chromosomes X and 2 and two from chromosome 3) sampled from multiple specimens shows considerable similarity of gene polymorphisms and even haplotypes between the two autosomal chromosomes of A. gambiae and A. arabiensis (Besansky et al., 2003), which share polymorphic inversions. At the X chromosome locus, however, there are fixed nucleotide differences and greater overall nucleotide differentiation between A. gambiae and A. arabiensis. These results are consistent with the suppressed-recombination model because the X chromosomes of the two species are fixed for different inversions, where reproductive factors are located.
The shared nucleotide polymorphisms in the autosomal chromosomes may, however, be ancestral rather than originated by recent gene flow between the two species. An investigation of the ND5 locus of mitochondrial DNA shows that ancestral haplotypes persist in the two species, A. gambiae and A. arabiensis (Donnelly et al., 2004). Nevertheless, comparison of allopatric and sympatric populations suggests locale-specific unidirectional introgression from A. arabiensis into A. gambiae. Indeed, the acquisition by A. gambiae of alleles from the more arid-adapted A. arabiensis may have contributed to its spread and ecological dominance (Besansky et al., 2003). A more definitive test of the suppressed-recombination model of chromosomal speciation would call for estimation of the KA/KS ratio in 20 or more genes distributed among the three chromosomes of these two species, as well as among the other five species and additional incipient species.
It may be worth adding that among the nearly 500 known members of the genus Anopheles, there are no fewer than 170 cryptic taxa belonging to 30 complexes of closely related species (Harbach, 2004). Most siblings are outcomes of recent speciation processes detected by paracentric inversions, mostly involving the X chromosome, as well as ribosomal DNA sequences. The chromosome rearrangements act as mechanisms of crossover suppression for reorganized regulatory units of gene expression.
These recently evolved complexes of Anopheles taxa are splendid materials for testing models of speciation mediated by chromosome rearrangements.
REFERENCES
Anderson, W. W., Ayala, F. J. & Michod, R. E. (1977) Chromosomal and allozymic diagnosis of three species of Drosophila: Drosophila pseudoobscura, D. persimilis, and D. miranda. J. Hered. 68, 70–74.
Besansky, N. J., Krzywinski, J., Lhemann, T., Simard, F., Kern, M., Mukabayire, O., Fontenille, D., Touré, Y. & Sagnon, N. F. (2003) Semipermeable species boundaries between Anopheles gambiae and Anopheles arabiensis: Evidence from multilocus DNA sequence variation. Proc. Natl. Acad. Sci. USA 100, 10818–10823.
Brown, K. M., Burk, L. M., Henagan, L. M. & Noor, M. A. F. (2004) A test of the chromosomal rearrangement model of speciation in Drosophila pseudoobscura. Evolution 58, 1856–1860.
Cáceres, M., Lachuer, J., Zapala, M. A., Redmond, J. C., Kudo, L., Geschwind, D. H., Lockhart, D. J., Preuss, T. M. & Barlow, C. (2003) Elevated gene expression levels distinguish human from non-human primate brains. Proc. Natl. Acad. Sci. USA 100, 13030–13035.
Coluzzi, M. (1982) Spatial distribution of chromosomal inversions and speciation in anopheline mosquitoes. In Mechanisms of Speciation (Liss, New York), pp. 143–153.
Coluzzi, M. & Bradley, D., eds. (1999) The Malaria Challenge After One Hundred Years of Malariology (Lombardo Editore, Rome). Parassitologia 41(1–3).
Coluzzi, M., Sabatini, A., della Torre, A., Di Deco, M. A. & Tetrarca, V. (2002) A polytene chromosome analysis of the Anopheles gambiae species complex. Science 298, 1415–1418.
de Boer, L. E. M. & Seuánez, H. N. (1982) The chromosomes of the oran utan and their relevance to the conservation of the species. In The Orang Utan: Its Biology and Conservation, ed. de Boer, L. E. M. (W. Junk, The Hague, The Netherlands), pp. 135–170.
Dobzhansky, T. (1935a) A critique of the species concept in biology. Philos. Sci. 2, 344–355.
Dobzhansky, T. (1935b) Drosophila Miranda, a new species. Genetics 20, 377–391.
Dobzhansky, T. (1937a) Genetic nature of species differences. Am. Nat. 71, 404–420.
Dobzhansky, T. (1937b) Genetics and the Origin of Species (Columbia Univ. Press, New York).
Dobzhansky, T. (1970) Genetics of the Evolutionary Process (Columbia Univ. Press, New York).
Dobzhansky, T. (1973) Is there gene exchange between Drosophila pseudoobscura and Drosophila persimilis in their natural habitats? Am. Nat. 107, 312–314.
Donnelly, M. J., Pinto, J., Girod, R., Besansky, N. J. & Lehmann, T. (2004) Revisiting the role of introgression vs shared ancestral polymorphisms as key processes shaping genetic diversity in the recently separated sibling species of the Anopheles gambiae complex. Heredity 92, 61–68.
Ghiselin, M. (1974) A radical solution to the species problem. Syst. Zool. 23, 536–544.
Harbach, R. E. (2004) The classification of genus Anopheles (Diptera : Culicidae): A working hypothesis of phylogenetic relationships. Bull. Entomol. Res. 94, 537–553.
Hartl, D. L. (2004) The origin of malaria: Mixed messages from genetic diversity. Nat. Rev. Microbiol. 2, 15–22.
Hey, J. (2003) Speciation and inversions: Chimps and humans. BioEssays 25, 825–828.
Hull, D. L. (1977) The ontological status of species as evolutionary units. In Foundational Problems in Special Sciences, eds. Butts, R. & Hintikka, J. (Kluwer, Dordrecht, The Netherlands), pp. 91–102.
Kimura, M. (1983) The Neutral Theory of Molecular Evolution (Cambridge Univ. Press, Cambridge, U.K.).
Lincoln, R. J., Boxshall, G. A. & Clark, P. F. (1998) A Dictionary of Ecology, Evolution, and Systematics (Cambridge Univ. Press, Cambridge, U.K.), 2nd Ed.
Lu, J., Li, W. H. & Wu, C. I. (2003) Comment on “Chromosomal speciation and molecular divergence-accelerated evolution in rearranged chromosomes.” Science 302, 988.
Machado, C. A., Kliman, R. M., Markert, J. A. & Hey, J. (2002) Inferring the history of speciation from multilocus DNA sequence data: The case of Drosophila pseudoobscura and close relatives. Mol. Biol. Evol. 19, 472–488.
Marquès-Bonet, T., Cáceres, M., Bertranpetit, J., Preuss, T. M., Thomas, J. W. & Navarro, A. (2004) Chromosomal rearrangements and the genomic distribution of gene-expression divergence in humans and chimpanzees. Trends Genet. 20, 524–529.
Mayr, E. (1942) Systematics and the Origin of Species (Columbia Univ. Press, New York).
Mayr, E. (1976) Is the species a class or an individual? Syst. Zool. 25, 192.
Mayr, E. (1987) The ontological status of species: Scientific progress and philosophical terminology. Biol. Philos. 2, 146–166.
Moore, B. C. & Taylor, C. E. (1986) Drosophila of Southern California. 3. Gene arrangements of Drosophila persimilis. J. Hered. 77, 313–323.
Navarro, A. & Barton, N. H. (2003a) Accumulating postzygotic isolation genes in parapatry: A new twist on chromosomal speciation. Evolution 57, 447–459.
Navarro, A. & Barton, N. H. (2003b) Chromosomal speciation and molecular divergence—Accelerated evolution in rearranged chromosomes. Science 300, 321–324.
Noor, M. A. F., Grams, K. L., Bertucci, L. A., Almendarez, Y., Reiland, J. & Smith, K. R. (2001a) The genetics of reproductive isolation and the potential for gene exchange between Drosophila pseudoobscura and D. persimilis via backcross hybrid males. Evolution 55, 512–521.
Noor, M. A. F., Grams, K. L., Bertucci, L. A. & Reiland, J. (2001b) Chromosomal inversions and the reproductive isolation of species. Proc. Natl. Acad. Sci. USA 98, 12084–12088.
Orr, H. A. & Turelli, M. (2001) The evolution of postzygotic isolation: Accumulating Dobzhansky–Muller incompatibilities. Evolution 55, 1085–1094.
Powell, J. R. (1997) Progress and Prospects in Evolutionary Biology: The Drosophila Model (Oxford Univ. Press, New York).
Rich, S. M., Licht, M. C., Hudson, R. R. & Ayala, F. J. (1998) Malaria’s Eve: Evidence of a recent population bottleneck throughout the world populations of Plasmodium falciparum. Proc. Natl. Acad. Sci. USA 95, 4425–4430.
Rieseberg, L. H. (2001) Chromosomal rearrangements and speciation. Trends Ecol. Evol. 16, 351–358.
Rieseberg, L. H. & Livingstone, K. (2003) Evolution—Chromosomal speciation in primates. Science 300, 267–268.
Rieseberg, L. H., Vanfossen, C. & Desrochers, A. M. (1995) Hybrid speciation accompanied by genomic reorganization in wild sunflowers. Nature 375, 313–316.
Spirito, F. (2000) The role of chromosomal change in speciation. In Endless Forms, eds. Howard, D. J. & Berlocher, S. H. (Oxford Univ. Press, New York), pp. 320–329.
Temu, E. A. (1997) Detection of hybrids in natural populations of the Anopheles gambiae complex by the rDNA-based, PCR method. Ann. Trop. Med. Parasitol. 91, 963–965.
Tishkoff, S. A., Varkonyi, R., Cahinhinan, N., Abbes, S., Argyropoulos, G., Destro-Bisol, G., Drousiotou, A., Dangerfield, B., Lefranc, G., Loiselet, J., et al. (2001) Haplotype diversity and linkage disequilibrium at human G6PD: Recent origin of alleles that confer malarial resistance. Science 293, 455–462.
Touré, Y. T., Petrarca, V., Traore, S. F., Coulibaly, A., Maiga, H. M., Sankare, O., Sow, M., DiDecco, M. A. & Coluzzi, M. (1998) The distribution and inversion polymorphism of chromosomally recognized taxa of the Anopheles gambiae complex in Mali, West Africa. Parassitologia (Rome) 40, 477–511.
von Koenigswald, G. H. R. (1982) Distribution and evolution of the orang utan, Pongo pygmaeus (Hoppius). In The Orang Utan: Its Biology and Conservation, ed. de Boer, L. E. M. (W. Junk, The Hague, The Netherlands), pp. 1–15.
Wang, R. L., Wakeley, J. & Hey, J. (1997) Gene flow and natural selection in the origin of Drosophila pseudoobscura and close relatives. Genetics 147, 1091–1106.
White, M. J. D. (1968) Models of speciation. Science 159, 1065–1070.
White, M. J. D. (1978) Modes of Speciation (Freeman, San Francisco).
Willis, K. J., Gillson, L. & Brncic, T. M. (2004) How “virgin” is virgin rainforest? Science 304, 402–403.
Yunis, J. J. & Prakash, O. (1982) The origin of man—A chromosomal pictorial legacy. Science 215, 1525–1530.
Zhang, J., Wang, X. & Podlaha, O. (2004) Testing the chromosomal speciation hypothesis for humans and chimpanzees. Genome Res. 14, 845–851.























