Part I
THE ORIGINS OF SPECIES BARRIERS
Mayr was well known for his championship of the biological species concept and for asserting a predominant role for the geographic separation of populations in the diversification process that gives rise to separate species. The genetic version of this perspective is, to a good approximation, the Dobzhansky–Muller model of divergence, in which genes that have been the site of adaptive fixations within separate populations may also be the site of negative epistatic interactions in species hybrids and cause inviability or sterility when hybridization occurs (Dobzhansky, 1936; Muller, 1940; Orr, 1995). But what are these adaptations that accumulate within separate populations and give rise to reproductive barriers? This question is addressed by Allen Orr in “The Genetic Basis of Reproductive Isolation: Insights from Drosophila” (Chapter 2), and he explains what we know from the growing handful of cases in which the actual genes that contribute to low hybrid fitness have been isolated. The most striking commonality to emerge from these studies is that these genes have extraordinarily rapid rates of adaptive amino acid replacement. This finding makes sense, for if we suppose that some fraction of amino acid replacements are prone to negatively epistatic interactions when placed in a hybrid background, then those genes that have the highest rates of amino acid replacement will also tend to be those that cause these types of Dobzhansky–Muller incompatibilities.
This discovery of rapidly evolving genes that contribute to reproductive barriers also necessarily focuses our attention on the kinds of phenotypes and genes that are particularly prone to evolve rapidly. Classically,
much of the discussion on rapid population divergence tended to focus on the kinds of environmental or geographic circumstances that might promote rapid evolution. However, in recent years, the attention has shifted to situations where intraspecies and intragenomic conflicts can lead to rapid evolution of genes (Rice, 1998). Such conflicts arise whenever natural selection favors alleles that are penetrant under some circumstances (such as in one sex), even though those same alleles may reduce other components of fitness that are manifest in other contexts (such as in the other sex). Genomic conflicts can lead to tit-for-tat, or arms-race, evolution between groups of genes within the same genome. In “Inter-Locus Antagonistic Coevolution as an Engine of Speciation: Assessment with Hemiclonal Analysis” (Chapter 3), William Rice et al. explore this issue directly by developing a model evolutionary system with Drosophila melanogaster. In this system, individual haploid chromosome complements (hemiclones) are drawn, using genetic tricks that are possible with Drosophila, from a longstanding laboratory population. Once isolated, each hemiclone can be measured, by replicating in combination with other hemiclones, for its net effect upon fitness of particular phenotypes. The approach allows a careful assessment of the selection gradient and additive genetic variance for traits that enhance fitness in males but reduce fitness in their female mates.
The findings that rapid evolution of genes, including that caused by genomic conflict, can lead to the formation of reproductive barriers notwithstanding, there remains the question of how much gene flow can be tolerated between diverging populations if speciation is to occur (Wright, 1940). Not even populations with many rapidly evolving genes can be expected to become reproductively isolated from other populations if gene flow rates are high. With this point in mind, Francisco Ayala and Mario Coluzzi, in “Chromosome Speciation: Humans, Drosophila, and Mosquitoes” (Chapter 4), explore models in which recombination suppressors, such as chromosomal inversions, can enhance the opportunity for adaptive divergence in the face of gene flow between parapatric populations (Coluzzi, 1982; Noor et al., 2001; Rieseberg, 2001). These models are more plausible than those in which chromosomal inversions enable divergence by causing low hybrid fitness, and they are supported particularly by recent evidence from Drosophila and Anopheles.
Regardless of the rates and roles of genetic changes that contribute to divergence, there remain very large questions about the phenotypic manifestation of these changes. Mary Jane West-Eberhard, in “Developmental Plasticity and the Origin of Species Differences” (Chapter 5), addresses these questions and stresses that phenotypes may derive from genes in ways that are highly contingent upon other genes and upon environmental circumstances. Thus, single new alleles or new environmental circum-
stances alone, in the absence of genetic changes, may trigger large changes in the phenotype. Furthermore, if genetic accommodation is common, such as by the Baldwin effect (Baldwin, 1896; Simpson, 1953), then phenotypic variation that arises primarily by environmental causes may play a driving role in divergence.
REFERENCES
Baldwin, J. M. (1896) A new factor in evolution. Am. Nat. 30, 441–451.
Coluzzi, M. (1982) Spatial distribution of chromosomal inversions and speciation in anopheline mosquitoes. In Mechanisms of Speciation, ed. Barigozzi, C. (Liss, New York), pp. 143–153.
Dobzhansky, T. (1936) Studies on hybrid sterility. II. Localization of sterility factors in Drosophila pseudoobscura hybrids. Genetics 21, 113–135.
Muller, H. J. (1940) Bearings of the Drosophila work on systematics. In The New Systematics, ed. Huxley, J. (Clarendon, Oxford), pp. 185–268.
Noor, M. A., Grams, K. L., Bertucci, L. A. & Reiland, J. (2001) Chromosomal inversions and the reproductive isolation of species. Proc. Natl. Acad. Sci. USA 98, 12084–12088.
Orr, H. A. (1995) The population-genetics of speciation—The evolution of hybrid incompatibilities. Genetics 139, 1805–1813.
Rice, W. R. (1998) Intergenomic conflict, interlocus antagonistic coevolution, and the evolution of reproductive isolation. In Endless Forms: Species and Speciation, eds. Howard, D. J. & Berlocher, S. H. (Oxford Univ. Press, New York), pp. 261–270.
Rieseberg, L. H. (2001) Chromosomal rearrangements and speciation. Trends Ecol. Evol. 16, 351–358.
Simpson, G. G. (1953) The Baldwin effect. Evolution 7, 110–117.
Wright, S. (1940) Breeding structure of populations in relation to speciation. Am. Nat. 74, 232–248.
2
The Genetic Basis of Reproductive Isolation: Insights from Drosophila
H. ALLEN ORR*
Recent studies of the genetics of speciation in Drosophila have focused on two problems: (i) identifying and characterizing the genes that cause reproductive isolation, and (ii) determining the evolutionary forces that drove the divergence of these “speciation genes.” Here, I review this work. I conclude that speciation genes correspond to ordinary loci having normal functions within species. These genes fall into several functional classes, although a role in transcriptional regulation could prove particularly common. More important, speciation genes are typically very rapidly evolving, and this divergence is often driven by positive Darwinian selection. Finally, I review recent work in Drosophila pseudoobscura on the possible role of meiotic drive in the evolution of the genes that cause postzygotic isolation.
Ernst Mayr made at least three contributions that are fundamental to the genetic study of speciation. The first, and surely most important, was his codification of what we mean by species and thus by speciation. In his book, Systematics and the Origin of Species, Mayr (1942) famously argued that species are best defined by the Biological Species Concept: species are groups of actually or potentially interbreeding natural populations that are reproductively isolated from other such groups. Reproductive isolation is thus the sine qua non of good species.
Mayr’s second contribution, elaborated in his book, Animal Species
and Evolution (1963), was his argument that reproductive isolation usually evolves in allopatry. This has, of course, been an enormously controversial claim, and I will not enter into it in detail. Suffice it to say that the weight of evidence, both biogeographic and comparative, now strongly suggests that Mayr was correct. Although sympatric speciation may well occur, reproductive isolation appears usually to evolve in allopatry (reviewed in Coyne and Orr, 2004, chapters 3 and 4).
Mayr’s third key claim, introduced in his 1942 book but elaborated in Mayr (1954) and later in Animal Species and Evolution, was that genetic drift plays a critical role in speciation. According to Mayr, large populations suffer a sort of evolutionary inertia: the conservative forces of gene flow and epistasis prevent, or at least render unlikely, the evolution of novel morphologies or new coadapted gene complexes in large, geographically widespread species. Such evolution is, he suggested, far more likely in island or peripheral populations founded by a few individuals. Genetic drift in such populations may allow rapid evolution to new fitness peaks, permitting the evolution of reproductive isolation between ancestral and founder populations. Such “founder effect” models of speciation proved extraordinarily popular, representing, as Provine (1989) noted, “the favored explanation for at least island speciation since 1954.” Despite this popularity, it is difficult to point to unambiguous evidence for founder effect speciation, and the idea has grown controversial. I return to this issue below.
The research program that Mayr helped to formulate, that to understand the origin of species one must understand the origin of reproductive isolation, has experienced a renaissance over the last two decades. Evolutionists have performed an impressive number of new and careful studies of the biogeography, ecology, and genetics of speciation, and we now understand a good deal about the evolution of reproductive isolation (Coyne and Orr, 2004). We can, for instance, describe the rate at which this reproductive isolation evolves (Fig. 2.1), and the ecological factors that drive speciation, at least in some cases (Schluter, 2000). We can also point to plausible examples in which the process of reinforcement may have completed speciation (Coyne and Orr, 2004; Noor, 1999; Servedio and Noor, 2003). Finally, we know a fair amount about the number and location of the genes that cause reproductive isolation (at least in its postzygotic form), and the causes of patterns like Haldane’s rule (Coyne and Orr, 2004; Laurie, 1997; Orr, 1997).
Given this progress, it is natural to ask: What now are the major outstanding problems in speciation? On the genetical front (I am not competent to address any other), the answers seem clear. We must (i) find the genes that cause reproductive isolation and (ii) identify the evolutionary forces that drove their divergence.
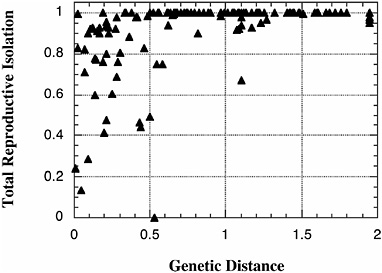
FIGURE 2.1 The rate of increase in “total” reproductive isolation with genetic distance in Drosophila. Postzygotic isolation is measured such that a value of 0 means no reproductive isolation (either pre- or postzygotic) and a value of 1 means complete reproductive isolation. Genetic distance is measured by Nei’s genetic distance, which increases approximately linearly with time over the values shown. Data include both allopatric and sympatric species pairs. Although the measure of total reproductive isolation shown includes aspects of both pre-and postzygotic isolation, it is imperfect and generally excludes forms of reproductive isolation (e.g., ecological isolation) that are not readily measured in the laboratory. Data are based on Coyne and Orr (1989, 1997). [Reproduced with permission from Coyne and Orr, 1997 (Copyright 1997, International Journal of Organic Evolution).]
THE PROBLEM
Biologists who do not specialize in speciation are invariably surprised to learn that evolutionists have identified very few genes causing reproductive isolation. The reason for this slow progress is, however, simple. If species are taxa that are reproductively isolated, a genetics of speciation must, almost by definition, be a genetics where such a thing is not possible, between organisms that do not exchange genes. The consequence of this methodological dilemma is that evolutionists have been unable to address a large set of fundamental questions about the genes that cause reproductive isolation, so-called speciation genes. (This perhaps unfortunate term, which is now entrenched in the literature, refers to any locus that causes reproductive isolation, whether in F1 or later-generation hybrids, and whether the gene was among the first to cause isolation or not.)
TABLE 2.1 Speciation Genes Causing Postzygotic Reproductive Isolation That Have Been Identified at the DNA Sequence Level
|
Gene |
Taxon |
Hybrid Phenotype |
Gene Type |
References |
|
Xmrk-2 |
Xiphophorus |
Inviability |
Receptor tyrosine kinase |
Wittbrodt et al. (1989), Schartl et al. (1999), Malitschek et al. (1995) |
|
OdsH |
Drosophila |
Sterility |
Transcription factor |
Ting et al. (1998), Wu and Ting (2004), Sun et al. (2004) |
|
Hmr |
Drosophila |
Inviability |
Transcription factor |
Barbash et al. (2003, 2004) |
|
Nup96 |
Drosophila |
Inviability |
Nucleoporin |
Presgraves et al. (2003) |
These questions include: Are speciation genes “ordinary” genes that have normal functions within species? If so, do speciation genes fall into one or at least a few functional classes? Are substitutions in speciation genes concentrated in coding or regulatory sequences? And do speciation genes diverge by natural selection or genetic drift? This last question is perhaps the most important: if we can identify speciation genes at the level of DNA sequences, we should be able to bring to bear a powerful set of molecular population genetic tools (e.g., McDonald–Kreitman and Hudson–Kreitman–Aguade tests) on the Mayrian question of the relative roles of deterministic vs. stochastic forces in the origin of species.
The methodological dilemma that plagues speciation genetics is sufficiently serious that, until recently, evolutionists could point to only a single gene that causes postzygotic reproductive isolation (Table 2.1). (I focus on postzygotic isolation because it has been the subject of most genetic analysis, both in general and in my own laboratory; see, however, Metz and Palumbi, 1996; Palumbi, 1992; Vacquier, 1998; Swanson and Vacquier, 2002, for discussion of genes causing prezygotic isolation.) It had long been known that crosses between certain fish species of the genus Xiphophorus, e.g., between the platyfish and swordtail, result in hybrid inviability. In particular, backcross hybrids between these species often develop malignant melanomas and die (Schartl et al., 1994). Half a
century of genetic study has revealed that this hybrid lethality results from a two-locus “Dobzhansky–Muller” incompatibility: one locus on the X chromosome of the platyfish is incompatible with another (apparently single) locus on an autosome of the swordtail. Melanoma results when these loci are brought together in a hybrid genome. Molecular genetic analyses allowed identification and characterization of the X-linked partner in this incompatibility: Xmrk-2. Xmrk-2 encodes a novel receptor tyrosine kinase, which, while found as a duplicate gene on the platyfish X, is absent from the swordtail X (Wittbrodt et al., 1989; Schartl et al., 1999; Malitschek et al., 1995). Xmrk-2 appears to be misexpressed in species hybrids, causing tumor formation (Malitschek et al., 1995).
Xmrk-2 was subjected to intense genetic analysis because of its role in cancer. There is little reason to believe, however, that postzygotic isolation usually involves malignancies. The recent renaissance in the genetics of speciation has therefore featured a determined effort to find additional speciation genes regardless of the form of hybrid inviability or sterility involved. One cannot, after all, draw robust conclusions about the identities or evolutionary histories of the genes causing reproductive isolation from a sample of one. Fortunately, these recent efforts, by a number of laboratories, have largely succeeded, and we now know the identities of several additional speciation genes. I briefly review this work. I then turn to recent efforts to characterize the evolutionary forces that drive the divergence of such genes. As we will see, these forces may sometimes assume a surprising form.
THE SEARCH FOR SPECIATION GENES
The general methodological dilemma facing the genetics of speciation involves a number of particularly unfortunate special cases. Perhaps worst of all, one of our most sophisticated genetic model systems, Drosophila melanogaster, forms only inviable or sterile hybrids when crossed to any other species. As a result, D. melanogaster has, until recently, proved nearly useless in the study of speciation.
Drosophila geneticists have taken two approaches to sidestep this problem. The first step involves study of non-melanogaster fly species, wherein at least some of the tools first developed in D. melanogaster may be available (e.g., germ-line transformation, or P element insertions). This approach allowed identification of a gene that causes hybrid male sterility. Coyne and Charlesworth (1986) mapped a locus causing sterility of Drosophila simulans–Drosophila mauritiana hybrid males to a small region of the D. mauritiana X chromosome. Subsequent work (Perez and Wu, 1995; Perez et al., 1993) refined this mapping, showing that at least two hybrid sterility factors reside in the region, one of which reduces hybrid fertility
by half when introgressed alone onto a D. simulans genetic background. Ting et al. (1998) identified this locus, Odysseus-Homeobox gene (OdsH), at the molecular level. As its name indicates, OdsH is an X-linked homeobox gene and thus a presumed transcription factor (Ting et al., 1998; Wu and Ting, 2004). Recent transformation experiments appear to confirm that OdsH causes partial male sterility among backcross (although not F1) hybrids (Sun et al., 2004). Sterility may reflect misexpression of OdsH in hybrid testes (Wu and Ting, 2004). OdsH evolved rapidly between D. simulans and D. mauritiana, and this evolution was almost certainly driven by positive Darwinian selection: nonsynonymous substitutions greatly outnumber synonymous ones in the lineage leading to D. mauritiana (Ting et al., 1998).
The second approach to identifying speciation genes has taken more direct advantage of D. melanogaster. Although any hybrids formed between this species and its closest relatives are completely sterile, this fact does not preclude its use in the search for hybrid inviability genes. Several workers (Barbash et al., 2000; Hutter and Ashburner, 1987; Hutter et al., 1990; Orr and Irving, 2000) used tools from D. melanogaster (e.g., chromosomal duplications and deletions, loss-of-function mutations, and germline transformation) to map and characterize an X-linked gene, Hybrid male rescue (Hmr), that causes the inviability of certain F1 hybrids between D. melanogaster and its sister species, e.g., D. simulans. This work culminated in the recent identification by Barbash et al. (2003) of Hmr at the sequence level: Hmr encodes a transcriptional regulator of or related to the MYB family. It seems likely that Hmr’s presumed normal function in gene regulation is disrupted in species hybrids, causing inviability (Barbash et al., 2003). Very recent work reveals that Hmr is also rapidly evolving, and that this evolution was also driven by positive Darwinian selection (Barbash et al., 2004).
My laboratory has focused on deficiency mapping to locate and identify speciation genes. The critical point is that such mapping, which involves the introduction of cytologically defined chromosomal deletions from D. melanogaster into species hybrids, can be performed within F1 hybrids, whose sterility is therefore irrelevant. Daven Presgraves (2003) used a large set of deficiencies from D. melanogaster to screen for autosomal genes from D. simulans that cause hybrid inviability when present with an X chromosome from D. melanogaster. The essence of this approach is that it allows detection and mapping of recessive, and thus, normally masked, speciation genes: if a chromosomal region includes a recessive hybrid inviability gene(s) from D. simulans, control hybrids that carry an undeleted (balancer) chromosome from D. melanogaster should be viable, whereas experimental hybrids that carry a deleted chromosome (in the relevant region) from D. melanogaster should be inviable.
Presgraves (2003) singly introduced >200 deficiencies into D. melanogaster–D. simulans species hybrids. He found that 20 deficiencies significantly reduced hybrid fitness, and that 10 of these deficiencies were hybrid lethal, i.e., uncovered genes from D. simulans that cause essentially complete inviability on a partly D. melanogaster genetic background. Further crosses using attached-X chromosome manipulations revealed that these D. simulans autosomal regions interact with the D. melanogaster X chromosome to cause hybrid inviability (Presgraves, 2003). This work implies that D. melanogaster and D. simulans, although diverged for only ≈2.5 million years (Hey and Kliman, 1993; Li, 1997), are separated by many genes causing hybrid inviability. Indeed, it appears that ≈10% of all viability-essential genes have diverged functionally between these species to such an extent that a gene from one species kills hybrids if made homozygous (hemizygous) in the genetic background of the other species (Presgraves, 2003).
In subsequent work, Presgraves et al. (2003) dissected a small deficiency region known to cause the inviability of D. melanogaster–D. simulans hybrids. A combination of fine-scale deletion mapping and complementation testing (Fig. 2.2) revealed that the hybrid inviability effect of this third chromosomal region mapped to a single complementation group, which was ultimately shown to correspond to Nucleoporin96 (Nup96). This locus interacts with another (presently unknown) locus on the D. melanogaster X chromosome to cause hybrid inviability. Nup96 encodes a component of the nuclear pore complex, a large structure that perforates the nuclear membrane of all eukaryotic cells and that regulates the trafficking of proteins and RNA in and out of the nucleus. Although the ≈30 nucleoporin proteins that constitute the nuclear pore complex are typically conserved evolutionarily, Nup96 evolved rapidly between D. melanogaster and D. simulans. Molecular population genetic analyses further reveal that this divergence was driven by positive Darwinian selection in both species lineages. Because neither species shows evidence of recent adaptive sweeps (e.g., depressed nucleotide diversity or skewed frequency spectra), this adaptive evolution likely occurred early in the split of the two species (Presgraves et al., 2003).
In very recent work, my laboratory has turned to the attempt to identify and characterize speciation genes that cause hybrid sterility, not inviability. The chief reason is that it is now clear that hybrid sterility, and, especially hybrid male sterility, evolves faster than hybrid inviability in several groups of animals, including Drosophila (reviewed in Coyne and Orr, 2004). Hybrid male sterility may thus represent a more important phenotype than hybrid inviability in the earliest stages of speciation. Our current efforts focus on the so-called 4-sim hybrid sterility system: Muller and Pontecorvo (1942) and Pontecorvo (1943) showed long ago that D.
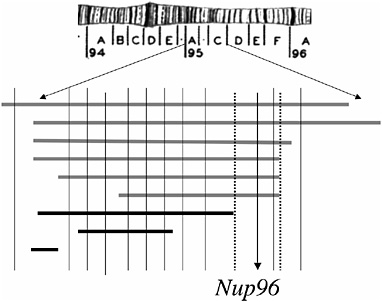
FIGURE 2.2 Deficiency and complementation mapping of a hybrid inviability gene. The region shown corresponds to cytological region 95 of the third chromosome of D. melanogaster. Bars shown in gray below the chromosome uncover recessive hybrid inviability in D. melanogaster–D. simulans hybrids and those shown in black do not. Vertical lines represent known viability-essential complementation groups in the region. Hybrid inviability maps to a single locus, Nup96. Data are based on Presgraves et al., 2003. [Reproduced with permission from Presgraves et al., 2003 (Copyright 2003).]
melanogaster males that are homozygous for the “dot” fourth chromosome from D. simulans are sterile; sterility results from sperm immotility. Heterozygous 4-sim males and all genotypes of females are fertile. Deficiency mapping shows that hybrid male sterility localizes to a very small region of this very small chromosome (Orr, 1992; the fourth includes only ≈70 loci [Adams et al., 2000]). It seems likely therefore that 4-sim hybrid sterility reflects the action of a single locus that interacts with other loci elsewhere in the D. melanogaster genome. My laboratory has recently further narrowed the location of this putative hybrid sterility gene and has begun to test the few remaining candidate loci in the region for their possible role in hybrid sterility (J. P. Masly, personal communication).
In summary, the above genetic studies have finally allowed evolutionists to examine the factors that cause postzygotic isolation. It is clear that these factors correspond to ordinary genes, having normal functions within species: no evidence has been uncovered for the idea that speciation involves novel processes like the mass mobilization of transposable
elements in hybrids (Engels and Preston, 1979; Kidwell, 1983; Rose and Doolittle, 1983). It is also clear that speciation genes do not fall into a single functional class. Although two of the above genes (OdsH and Hmr) play a role in transcriptional regulation, and this occurrence could well prove common, others do not. Nup96, for example, encodes a structural protein. Similarly, it is clear that the divergence of noncoding regulatory sequences is not the invariable cause of postzygotic isolation: complementation tests with multiple mapped loss-of-function mutations show that Nup96’s effect on hybrid inviability is due to divergence of the Np96 protein itself (Presgraves et al., 2003). Also, the genes causing postzygotic isolation are sometimes members of duplicate gene families (Xrmk-2 and OdsH), but are sometimes single-copy genes (Hmr and Nup96). Although there is good evidence that the genes causing postzygotic isolation are on average partially recessive, as predicted by the dominance theory of Haldane’s rule (reviewed in Coyne and Orr, 2004), it also clear that speciation genes can differ dramatically in dominance. Nup96 and Hmr, for instance, act mostly recessively in hybrids, whereas the (currently unknown) gene(s) from D. simulans with which Hmr interacts to cause hybrid inviability appears quite dominant (Hutter et al., 1990). Recessive speciation genes can obviously contribute only to the sterility or inviability of F2, or backcross, not F1, hybrids (unless, of course, the gene is X-linked). Whereas any given recessive gene may well cause less reproductive isolation than any given dominant one, recessive speciation genes appear more common than dominant; not surprisingly, then, later-generation hybrid problems often appear before F1 problems (reviewed in Coyne and Orr, 2004, chapters 7 and 8).
Although the study of speciation genes remains in its infancy, two patterns do, so far at least, appear to characterize their evolution. First, speciation genes are rapidly evolving. Second, these genes often evolve by positive Darwinian selection (Barbash et al., 2003; Presgraves et al., 2003).
SPECIATION GENES: INTRAGENOMIC CONFLICT?
While the above work shows that positive selection plays a key role in the divergence of speciation genes, it says nothing about the nature of this selection. Departures from neutrality as revealed by, e.g., McDonald–Kreitman tests, are equally consistent with sexual selection, natural selection to abiotic factors, or natural selection to biotic factors. Surprisingly, recent work suggests that a particular form of the last hypothesis, one involving adaptation not to other organisms but to intragenomic events, may play a role in speciation. In particular, it now appears that meiotic drive may sometimes drive the evolution of hybrid sterility genes.
This idea was first suggested in the early 1990s by Frank (1991) and
Hurst and Pomiankowski (1991). These authors argued that mutations that cause meiotic drive may often become suppressed within species (because drive at one locus causes selection at other loci for suppression of drive), but can become reexpressed in species hybrids: if the mutations that suppress drive are less than completely dominant, they may fail to suppress segregation distortion in heterozygous hybrids. Frank (1991) and Hurst and Pomiankowski (1991) further speculated that such drive might sometimes cause the sterility of hybrids. Other variations on the meiotic drive theory of postzygotic isolation have appeared recently (Henikoff and Malik, 2002; Henikoff et al., 2001; Tao and Hartl, 2003; Tao et al., 2001).
Although early experimental work found no evidence of meiotic drive in species hybrids (Coyne and Orr, 1993; Johnson and Wu, 1992), more recent work has uncovered such evidence (Cazemajor et al., 1997; Mercot et al., 1995; Montchamp-Moreau and Joly, 1997; Tao et al., 2001). In the most impressive of these efforts, Tao et al. (2001) showed that a <80-kb region of the third chromosome of D. mauritiana causes meiotic drive in an otherwise D. simulans genetic background. Remarkably, this same small region also causes hybrid male sterility, leading Tao et al. (2001) to conclude that the same gene, too much yin (tmy), likely causes both hybrid segregation distortion and hybrid sterility; tmy has not yet been identified at the DNA sequence level.
Recently, my laboratory has discovered strong segregation distortion in hybrids between two closely related subspecies of D. pseudoobscura. When females of the Bogota subspecies are crossed to males of the USA subspecies, fertile F1 females and sterile F1 males result. Although these males have traditionally been described as completely sterile, we have discovered that they are, in fact, weakly fertile. Surprisingly, these F1 males produce nearly all daughters (≈95%) when crossed to any genotype of females (pure Bogota, pure USA, and hybrid F1 sisters) (Orr and Irving, 2005). X-linked genetic markers reveal that this sex ratio distortion is not due to somatic sexual transformation (i.e., phenotypic females are genetic females) and egg-to-adult viability studies strongly suggest that it is not due to the inviability of sons (Orr and Irving, 2005). Instead, hybrid F1 males appear to suffer severe segregation distortion. Although we do not yet know the proximate mechanism of segregation distortion (e.g., classical meiotic drive in the male germ line, immotility of Y-bearing sperm in the female reproductive tract, postfertilization failure of pronuclei fusion, etc.), we have completed a preliminary genetic analysis of this hybrid segregation distortion.
Our results reveal that the genes that normally suppress segregation distortion within the Bogota subspecies are autosomal (and perhaps also Y-linked) (Orr and Irving, 2005). More important, the genes that cause hybrid segregation distortion reside in several regions of the Bogota X
FIGURE 2.3 Regions of the D. pseudoobscura X chromosome implicated in hybrid sterility between the Bogota and USA subspecies (Orr and Irving, 2001). Recent genetic analysis of segregation distortion in the same hybridization implicates approximately the same chromosomal regions (Orr and Irving, 2005). The markers shown are all visible mutations. Kosambi-corrected map positions are provided. [Reproduced with permission from Orr and Irving, 2001 (Copyright 2001, Genetics Society of America).]
chromosome. Remarkably, these regions show essentially complete epistasis: little or no distortion occurs unless genes from both the left (XL) and right (XR) arms of the X derive from Bogota. These results are strikingly similar to those characterizing hybrid male sterility between the same subspecies (Orr, 1989; Orr and Irving, 2001; see Fig. 2.3). More suggestive still, the severity of hybrid sterility and the extent of offspring sex ratio distortion are strongly correlated across individual backcross hybrid males (Fig. 2.4). Future high-resolution introgression analysis
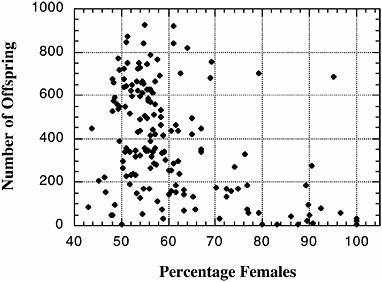
FIGURE 2.4 Correlation between D. pseudoobscura Bogota-USA hybrid fertility (as measured by offspring production) and hybrid segregation distortion (as measured by offspring sex ratio). Each data point reflects the offspring of a single recombinant backcross generation male. Data are based on Orr and Irving (2005). [Reproduced with permission from Orr and Irving, 2005 (Copyright 2005, Genetics Society of America).]
within a small but critical region of XR will let us determine whether the genes that cause hybrid sterility can be separated meiotically from those that cause hybrid segregation distortion is warranted (N. Phadnis, personal communication).
It is clear, then, that hybrid males between these young subspecies suffer both sterility and segregation distortion, and that these two forms of hybrid dysfunction have similar genetic bases. We cannot, therefore, reject the possibility that at least some of the same genes cause both phenomena, as first suggested by Frank (1991) and Hurst and Pomiankowski (1991). More generally, we cannot reject the possibility that arms races between selfish genetic factors like those that cause meiotic drive contribute to the evolution of postzygotic reproductive isolation.
CONCLUSIONS
Molecular evolutionary analyses of speciation genes show that these loci are rapidly evolving, and that this evolution is often driven by positive Darwinian selection. Although the sample of genes characterized thus far by various laboratories remains small, and concentrated in the genus Drosophila, I suspect that these patterns may prove general, although likely not universal. Our recent work, along with that of several other groups, also suggests that the selection underlying the evolution of speciation genes may sometimes assume a surprising form, response to intragenomic conflicts, perhaps involving meiotic drive.
In summary, it would appear that two of Mayr’s three seminal contributions to the study of speciation were correct, or, at the least, extremely productive. First, the entire research program of the genetics of speciation over the last half-century arose out of Mayr’s Biological Species Concept. Whatever one’s views on the philosophical strengths or weaknesses of this concept, it has, as a practical matter, given rise to an extraordinarily rich research program, one that has led to a number of substantive discoveries. In a phrase, the modern genetics of speciation is a genetics of reproductive isolation. Second, there can be little doubt that this reproductive isolation typically, if not always, evolves in allopatry (Coyne and Orr, 2004; Mayr, 1963). Finally, however, recent work provides no evidence for a crucial role of genetic drift in speciation. Instead, both ecological studies (Schluter, 2000) and the genetical studies reviewed here point to an important role for positive Darwinian selection in the evolution of reproductive isolation. Although the above data do not exclude all varieties of drift-based models of speciation (founder-effect theories do, after all, feature a role for positive selection), such models seem at present unparsimonious (see also Coyne, 1994; Coyne and Orr, 2004).
It remains a testament to Mayr’s vast influence, however, that all of
the questions asked above, whatever their answers, were first raised or emphasized by Mayr nearly half a century ago. It seems doubtful that a modern genetics of speciation in anything like its present form would have arisen without his fundamental contributions.
ACKNOWLEDGMENTS
I thank F. Ayala, W. Fitch, and J. Hey for organizing the symposium in which this work was presented. I also thank J. Hey and an anonymous reviewer for helpful comments on an earlier draft of this paper. This work was supported by National Institutes of Health Grant GM519k32.
REFERENCES
Adams, M. D., Celniker, S. E., Holt, R. A., Evans, C. A., Gocayne, J. D., Amanatides, P. G., Scherer, S. E., Li, P. W., Hoskins, R. A., Galle, R. F., et al. (2000) The genome sequence of Drosophila melanogaster. Science 287, 2185–2195.
Barbash, D. A., Roote, J. & Ashburner, M. (2000) The Drosophila melanogaster Hybrid male rescue gene causes inviability in male and female species hybrids. Genetics 154, 1747–1771.
Barbash, D. A., Siino, D. F., Tarone, A. M. & Roote, J. (2003) A rapidly evolving MYB-related protein causes species isolation in Drosophila. Proc. Natl. Acad. Sci. USA 100, 5302–5307.
Barbash, D. A., Awadalla, P. & Tarone, A. M. (2004) Functional divergence caused by ancient positive selection of a Drosophila hybrid incompatibility locus. PLoS 2, 839–848.
Cazemajor, M., Landre, C. & Montchamp-Moreau, C. (1997) Genetics 147, 635–642.
Coyne, J. A. (1994) Ernst Mayr on the origin of species. Evolution (Lawrence, Kans.) 48, 19–30.
Coyne, J. A. & Charlesworth, B. (1986) Location of an X-linked factor causing male sterility in hybrids of Drosophila simulans and D. mauritiana. Heredity 57, 243–246.
Coyne, J. A. & Orr, H. A. (1989) Patterns of speciation in Drosophila. Evolution (Lawrence, Kans.) 43, 362–381.
Coyne, J. A. & Orr, H. A. (1993) Further evidence against meiotic-drive models of hybrid sterility . Evolution (Lawrence, Kans.) 47, 685–687.
Coyne, J. A. & Orr, H. A. (1997) “Patterns of speciation in Drosophila” revisited. Evolution (Lawrence, Kans.) 51, 295–303.
Coyne, J. A. & Orr, H. A. (2004) Speciation (Sinauer, Sunderland, MA).
Engels, W. R. & Preston, C. R. (1979) Hybrid dysgenesis in Drosophila melanogaster: The biology of male and female sterility. Genetics 92, 161–174.
Frank, S. H. (1991) Divergence of meiotic drive-suppressors as an explanation for sex-biased hybrid sterility and inviability. Evolution (Lawrence, Kans.) 45, 262–267.
Henikoff, S. & Malik, H. S. (2002) Selfish drivers. Science 417, 227.
Henikoff, S., Ahmad, K. & Malik, H. S. (2001) The centromere paradox: Stable inheritance with rapidly evolving DNA. Science 293, 1098–1102.
Hey, J. & Kliman, R. M. (1993) Population genetics and phylogenetics of DNA sequence variation at multiple loci within the Drosophila melanogaster species complex. Mol. Biol. Evol. 10, 804–822.
Hurst, L. D. & Pomiankowski, A. (1991) Causes of sex ratio bias may account for unisexual sterility in hybrids: A new explanation of Haldane’s rule and related phenomena. Genetics 128, 841–858.
Hutter, P. & Ashburner, M. (1987) Genetic rescue of inviable hybrids between Drosophila melanogaster and its sibling species. Nature 327, 331–333.
Hutter, P., Roote, J. & Ashburner, M. (1990) A genetic basis for the inviability of hybrids between sibling species of Drosophila. Genetics 124, 909–920.
Johnson, N. A. & Wu, C.-I. (1992) An empirical test of the meiotic drive models of hybrid sterility: Sex ratio data from hybrids between Drosophila simulans and Drosophila sechellia. Genetics 130, 507–511.
Kidwell, M. G. (1983) Intraspecific hybrid sterility. In The Genetics and Biology of Drosophila, eds. Ashburner, M., Carson, H. L., & Thompson, J. J. N. (Academic, London), Vol. 3c, pp. 125–154.
Laurie, C. C. (1997) The weaker sex is heterogametic: 75 years of Haldane’s rule. Genetics 147, 937–951.
Li, W.-H. (1997) Molecular Evolution (Sinauer, Sunderland, MA).
Malitschek, B., Fornzler, D. & Schartl, M. (1995) Melanoma formation in Xiphophorus: a model system for the role of receptor tyrosine kinases in tumorigenesis. BioEssays 17, 1017–1023.
Mayr, E. (1942) Systematics and the Origin of Species (Columbia Univ. Press, New York).
Mayr, E. (1954) Change of genetic environment and evolution. In Evolution as a Process, eds. Huxley, J., Hardy, A. C. & Ford, E. B. (Allen & Unwin, London), pp. 157–180.
Mayr, E. (1963) Animal Species and Evolution (Belknap Press of Harvard Univ. Press, Cambridge, MA).
Mercot, H., Atlan, A., Jacques, M. & Montchamp-Moreau, C. (1995) Sex-ratio distortion in Drosophila simulans: Co-occurrence of a meiotic drive and a suppressor of drive. J. Evol. Biol. 8, 283–300.
Metz, E. C. & Palumbi, S. R. (1996) Positive selection and sequence rearrangements generate extensive polymorphism in the gamete recognition protein bindin. Mol. Biol. Evol. 13, 397–406.
Montchamp-Moreau, C. & Joly, D. (1997) The sex-ratio trait in Drosophila simulans: Genetic analysis of distortion and suppression. Heredity 79, 24–30.
Muller, H. J. & Pontecorvo, G. (1942) Recessive genes causing interspecific sterility and other disharmonies between Drosophila melanogaster and simulans. Genetics 27, 157.
Noor, M. (1999) Reinforcement and other consequences of sympatry. Heredity 83, 503–508.
Orr, H. A. (1989) Complex epistasis and the genetic basis of hybrid sterility in the Drosophila pseudoobscura Bogota-USA hybridization. Evolution (Lawrence, Kans.) 43, 180–189.
Orr, H. A. (1992) Mapping and characterization of a “speciation gene” in Drosophila. Genet. Res. 59, 73–80.
Orr, H. A. (1997) Haldane’s rule. Annu. Rev. Ecol. Syst. 28, 195–218.
Orr, H. A. & Irving, S. (2000) Genetic analysis of the Hybrid male rescue locus of Drosophila. Genetics 155, 225–231.
Orr, H. A. & Irving, S. (2001) Segregation distortion in hybrids between the Bogota and USA subspecies of Drosophila pseudoobscura. Genetics 158, 1089–1100.
Orr, H. A. & Irving, S. (2005) Abnormal spermiogenesis is associated with the X-linked sex-ratio trait in Drosophila simulans. Genetics 169, 671–682.
Palumbi, S. R. (1992) Marine speciation on a small planet. Trends Ecol. Evol. 7, 114–118.
Perez, D. E. & Wu, C.-I. (1995) Further characterization of the Odysseus locus of hybrid sterility in Drosophila: One gene is not enough. Genetics 140, 201–206.
Perez, D. E., Wu, C.-I., Johnson, N. A. & Wu, M.-L. (1993) Genetics of reproductive isolation in the Drosophila simulans clade: DNA-marker assisted mapping and characterization of a hybrid-male sterility gene, Odysseus (Ods). Genetics 134, 261–275.
Pontecorvo, G. (1943) Hybrid sterility in artificially produced recombinants between Drosophila melanogaster and D. simulans. Proc. R. Soc. Edinburgh Sect. B 61, 385–397.
Presgraves, D. C. (2003) A fine-scale genetic analysis of hybrid incompatibilities in Drosophila. Genetics 163, 955–972.
Presgraves, D. C., Balagopalan, L., Abmayr, S. M. & Orr, H. A. (2003) Adaptive evolution drives divergence of a hybrid inviability gene between two species of Drosophila. Nature 423, 715–719.
Provine, W. B. (1989) Founder effects and genetic revolutions in microevolution and speciation:a historical perspective. In Genetics, Speciation, and the Founder Principle, eds. Giddings, L. V., Kaneshiro, K. Y. & Anderson, W. W. (Oxford Univ. Press, New York), pp. 43–76.
Rose, M. & Doolittle, W. F. (1983) Molecular biological mechanisms of speciation. Science 220, 157–162.
Schartl, A., Dimitrijevic, N. & Schartl, M. (1994) Evolutionary origin and molecular biology of the melanoma-inducing oncogene of Xiphophorus. Pigm. Cell. Res. 7, 428–432.
Schartl, M., Hornung, U., Gutbrod, H., Volff, J.-N. & Wittbrodt, J. (1999) Melanoma loss-of-function mutants in Xiphophorus caused by Xmrk-oncogene deletion and gene disruption by a transposable element. Genetics 153, 1385–1394.
Schluter, D. (2000) The Ecology of Adaptive Radiation (Oxford Univ. Press, Oxford).
Servedio, M. R. & Noor, M. A. F. (2003) The role of reinforcement in speciation: Theory and data. Annu. Rev. Ecol. Syst. 34, 339–364.
Sun, S., Ting, C.-T. & Wu, C.-I. (2004) The normal function of a speciation gene, Odysseus, and its hybrid sterility effect. Science 305, 81–83.
Swanson, W. J. & Vacquier, V. (2002) Rapid evolution of reproductive proteins. Nat. Rev. Genet. 3, 137–144.
Tao, Y. & Hartl, D. L. (2003) Genetic dissection of hybrid incompatibilities between Drosophila simulans and Drosophila mauritiana. III. Heterogeneous accumulation of hybrid incompatibilities, degree of dominance and implications for Haldane’s rule. Evolution (Lawrence, Kans.) 57, 2580–2598.
Tao, Y., Hartl, D. L. & Laurie, C. C. (2001) Sex-ratio distortion associated with reproductive isolation in Drosophila. Proc. Natl. Acad. Sci. USA 98, 13183–13188.
Ting, C.-T., Tsaur, S.-C., Wu, M.-L. & Wu, C.-I. (1998) A rapidly evolving homeobox at the site of a hybrid sterility gene. Science 282, 1501–1504.
Vacquier, V. D. (1998) Evolution of gamete recognition proteins. Science 281, 1995–1998.
Wittbrodt, J., Adam, D., Malitschek, B., Maueler, W., Raulf, F., Telling, A., Robertson, S. M. & Schartl, M. (1989) Novel putative receptor tyrosine kinase encoded by the melanoma-inducing Tu locus in Xiphophorus. Nature 341, 415–421.
Wu, C.-I. & Ting, C.-T. (2004) Genes and speciation. Nat. Rev. Genet. 5, 114–122.



















