Part II
DISCERNING RECENT DIVERGENCE
One of Mayr’s achievements that is not always counted is that he made the difficult questions on evolutionary divergence seem accessible. By laying out clear scenarios for a seemingly intractable process in which evolutionary factors interact with the geographic circumstances of populations to cause divergence, he fueled the interest and enthusiasm of generations of evolutionary biologists. The modern fruits of this enthusiasm are the many studies that detail reconstruction of recent cases where evolution has given rise to new species. The progress, and rapid pace of progress, in this field is clearly shown in the paper by Scott Edwards et al., “Speciation in Birds: Genes, Geography, and Sexual Selection” (Chapter 6), which outlines how present-day studies on speciation in birds are gaining the genetic and theoretical sophistication that had formerly only been associated with the model Drosophila systems. Topics such as the role of sexual selection and the frequency of sympatric speciation are now being addressed genetically in a number of avian systems.
Among the most intricate of speciation puzzles are those involving obligate mutualistic relationships. If one species of a mutualistic assemblage diverges into two, must the other species of the assemblage follow along? This is the question that arises for figs and their pollinating wasps, and it is addressed by Carlos Machado, Nancy Robbins, Tom Gilbert, and Allen Herre in “Critical Review of Host Specificity and Its Coevolutionary Implications in the Fig/Fig-Wasp Mutualism” (Chapter 7). Each of the 750 or so species of fig depends upon fig wasps for pollination; the
wasps, in turn, require the ovaries of the figs as oviposition sites (Wiebes, 1979). Strong reciprocal species specificity suggests that when individual fig species or individual wasp species undergo speciation, they do so in tandem with their mutualistic partner (cospeciation). But Machado et al.’s phylogenetic and population genetic study (Chapter 7) shows that the history has not been this straightforward and that host switches by wasps, and possibly species hybridization by figs, have created partly independent phylogenetic histories of figs and wasps.
In Mayr’s world view, new species arise under allopatry, and, after that, as divergence accrues, the geographic ranges of related species may later come to overlap. In this way, related but divergent species may be sympatric, in contrast to most closely related species, which are expected to have disjunct, allopatric distributions. This sequence of events was outlined explicitly by Mayr in a 1954 paper on the biogeography of sea urchins (Mayr, 1954). Stephen Palumbi and Harilaos Lessios, in “Evolutionary Animation: How Do Molecular Phylogenies Compare to Mayr’s Reconstruction of Speciation Patterns in the Sea?” (Chapter 8), have returned to this same Echinoid system and reconsidered Mayr’s synthesis using DNA sequence data. They find that although the pattern described by Mayr still largely applies, rapidly evolving gamete recognition proteins play a strong role in reproductive isolation. In contrast, Mayr had envisioned the evolution of reproductive isolation by a more genomewide steady accumulation of substitutions.
For many biologists, the question of whether geographic separation is strictly necessary for speciation (i.e., the question of whether sympatric or parapatric speciation occurs) comes into sharpest focus with the case of Rhagoletis pomonella. This is the apple maggot fly that has diverged into two host races (apple and hawthorne), apparently under geographic sympatry and aided by the different fruiting times of the two hosts (Filchak et al., 2000). Mayr’s former student Guy Bush discovered the history of sympatric divergence in Rhagoletis, and it has long been a standard component of the debates on the prevalence of sympatric speciation. Now we learn from Guy Bush’s former student Jeffrey Feder and his colleagues, in “Mayr, Dobzhansky, and Bush and the Complexities of Sympatric Speciation in Rhagoletis” (Chapter 9), that the sympatric divergence that occurred within U.S. populations may have been facilitated by genetic variation that came in by means of gene flow from largely separated populations in Mexico.
The question of sympatric speciation has also been much discussed in the context of the highly speciose cichlid fishes from the great African lakes: Victoria, Malawi, and Tanganyika (Mayr, 1984). Particularly in the cases of Lakes Malawi and Victoria, which are relatively young, it is a wonder how hundreds of species could form within confined bodies of water within <1 million years. Yong-Jin Won, Arjun Sivasundar, Yong
Wang, and Jody Hey, in “On the Origin of Lake Malawi Cichlid Species: A Population Genetic Analysis of Divergence” (Chapter 10), take a close look at a group of rock-dwelling species from Lake Malawi. To gain resolution, they used a new type of genetic marker that includes a microsatellite and linked sequence and a new Bayesian method for fitting complex models of divergence (Hey et al., 2004). The results suggest that some of these species have formed within the past few thousands years and that gene exchange is ongoing between species at some loci.
REFERENCES
Filchak, K. E., Roethele, J. B. & Feder, J. L. (2000) Natural selection and sympatric divergence in the apple maggot Rhagoletis pomonella. Nature 407, 739–742.
Hey, J., Won, Y.-J., Sivasundar, A., Nielsen, R. & Markert, J. A. (2004) Using nuclear haplotypes with microsatellites to study gene flow between recently separated Cichlid species. Mol. Ecol. 13, 909–919.
Mayr, E. (1954) Geographic speciation in tropical Echinoids. Evolution 8, 1–18.
Mayr, E. (1984) Evolution of fish species flocks: A commentary. In Evolution of Fish Species Flocks, eds. Echelle, A. A. & Kornfield, I. (Univ. of Maine Press, Orono, ME), pp. 3–11.
Wiebes, J. T. (1979) Co-evolution of figs and their insect pollinators. Annu. Rev. Ecol. Syst. 10, 1–12.
6
Speciation in Birds: Genes, Geography, and Sexual Selection
SCOTT V. EDWARDS,*§ SARAH B. KINGAN,*§ JENNIFER D. CALKINS,†§ CHRISTOPHER N. BALAKRISHNAN,‡§ W. BRYAN JENNINGS,*§ WILLIE J. SWANSON,†§ AND MICHAEL D. SORENSON‡§
Molecular studies of speciation in birds over the last three decades have been dominated by a focus on the geography, ecology, and timing of speciation, a tradition traceable to Mayr’s Systematics and the Origin of Species. However, in the recent years, interest in the behavioral and molecular mechanisms of speciation in birds has increased, building in part on the older traditions and observations from domesticated species. The result is that many of the same mechanisms proffered for model lineages such as Drosophila—mechanisms such as genetic incompatibilities, reinforcement, and sexual selection—are now being seriously entertained for birds, albeit with much lower resolution. The recent completion of a draft sequence of the chicken genome, and an abundance of single nucleotide polymorphisms on the autosomes and sex chromosomes, will dramatically accelerate research on the molecular mechanisms of avian speciation over the next few years. The challenge for ornithologists is now to inform well studied examples of speciation in nature with increased molecular resolution—to clone speciation genes if they
exist—and thereby evaluate the relative roles of extrinsic, intrinsic, deterministic, and stochastic causes for avian diversification.
Presumably, it was only a distant dream of mid-century participants in the Modern Synthesis that one day genetic analysis would come to dominate so completely the analysis of speciation in a clade such as birds. That genetics would be a useful tool in the analysis of speciation in model organisms had been evident since the early days of Drosophila genetics. Yet Mayr’s Systematics and the Origin of Species (1942), published 11 years before the discovery of the structure of DNA, was a treasure-trove of speciation stories not of logistically tractable species with easily sampled and manipulated populations; rather, this book focused on speciation stories from the distant South Pacific, on what were, even for ornithologists, virtually inaccessible taxa with ranges straddling some of the most remote and challenging habitats of the planet. The allure of the exotic continues for ornithologists: Mayr and Diamond have recently undertaken a complete taxonomic and biogeographic revision of the birds of the Solomon Islands (2001), and detailed molecular phylogeographic tests of several speciation stories in this assemblage are finally underway (Filardi, 2003; Filardi and Smith, 2005; Smith, 2003). Indeed, the role of molecular techniques introduced to ornithology with the first allozyme surveys of avian populations nearly 35 years ago has been primarily to inform the geography and timing of speciation, thereby emphasizing extrinsic aspects of the speciation process: species delimitation, allopatric speciation, ecological divergence, bottlenecks, and the role of the Pleistocene (Avise, 2000; Barrowclough, 1983). Those mechanisms of avian speciation described by Mayr in terms of internal factors—for example, speciation resulting from so-called “genetic revolutions” (Mayr, 1963)—were often vague and, in the case of genetic revolutions, have been largely discredited (Barton and Charlesworth, 1987).
In the last 10 years, however, there has been a renewed interest in the behavioral, cognitive, and even molecular mechanisms of speciation in birds. This renaissance, spearheaded largely by recent reviews by Price (Irwin and Price, 1999; Price, 1998, 2002; Price and Bouvier, 2002), builds in part on the ancient tradition of avian husbandry and domestication, and in part on theoretical models suggesting a role for diverse behavioral factors in bird speciation, including sexual selection, sexual imprinting, learning, reinforcement, and genetic incompatibilities. Although biogeographic analyses still largely support allopatric speciation models (Coyne and Price, 2000), recent years have also witnessed the first serious attempts to document sympatric speciation in birds (Grant and Grant, 1979;
Sorenson et al., 2003), and the frequency of cases of sympatric speciation and divergence due to hybrid incompatibilities or reinforcement is an open question (Coyne and Orr, 2004). Now, a draft of the complete chicken genome and >2.8 million chicken SNPs have been determined roughly 60 years after Mayr’s landmark book (International Chicken Genome Sequencing Consortium, 2004; International Chicken Polymorphism Map Consortium, 2004). This treasure-trove of genes and genetic markers will no doubt spur rapid advances in both the geography and genetics of speciation in birds. This article reviews recent studies of extrinsic and intrinsic aspects of speciation in birds, focusing specifically on systematic and mechanistic issues that challenge the universality of the allopatric speciation paradigm.
GENE TREES, SPECIES DELIMITATION, AND PHYLOGEOGRAPHIC PATTERNS
Genetic data are serving an ever-increasingly important role in the delimitation of species, yet considerable controversy remains over which criteria to apply to this age-old problem (Avise and Ball, 1990; Coyne and Orr, 2004; Cracraft, 1983; Hey, 2001; Sites and Marshall, 2003; Wiens and Penkrot, 2002). Indeed, determining which of myriad species delimitation methods and species definitions is most appropriate for one’s focal taxa remains one of the paramount challenges in systematics, with important consequences for evolutionary biologists as well as for conservation biologists (Crandall et al., 2000; Moritz, 2002; Sites and Crandall, 1997). The fact that many avian sister taxa occur in allopatry—particularly in Gondwanan continents such as South America and Australia (Bates et al., 1998; Cracraft, 1991)—makes interpretation of biogeographic histories more straightforward but can also make attempts at species delimitation particularly challenging. It has long been recognized that the biological species concept (BSC), with its emphasis on reproductive isolation, is inapplicable in many allopatric situations because there is no opportunity to test for reproductive isolation, rendering the concept arbitrary (Zink and McKitrick, 1995). Species concepts emphasizing genetic clustering of forms can be equally arbitrary (reviewed in Irwin and Price, 1999). Diagnosibility—the ability to delimit and identify distinguishing character states for a given collection of individuals or taxa, usually but not always in a phylogenetic context—has been proffered as a general consideration when delimiting species (Cracraft, 1983). Although diagnosibility is sometimes construed as being equivalent to “fixed” character or genetic differences between taxa, alternate fixations are not a requirement for diagnosibility. The rise in sophisticated statistical genetic algorithms and large-scale multilocus analyses of variation in birds and other taxa con-
firm that it is trivial to diagnose species, subspecies, or even populations, even in the presence of abundant shared polymorphisms; a recent study of native and introduced house finch populations in North America was able to readily diagnose populations that had been separated for 50–100 years (Wang et al., 2003), even in the absence of fixed genetic or morphological differences. A naïve evaluation of the genetic patterns in this study would have reasonably inferred the existence of three diagnosable “species” of native and introduced House Finches within the continental United States and Hawaii. A similar ease of diagnosibility using large-scale multilocus approaches is observed for geographic populations of humans (Tishkoff and Kidd, 2004). Thus, diagnosibility will become highly problematic as the resolving power of multilocus approaches increases; the specter of statistical significance without biological significance will be perennial.
The criterion of monophyly advocated by proponents of the phylogenetic species concept (PSC) also suffers from arbitrariness, particularly given the disproportionate emphasis on mtDNA. It is not surprising that a number of workers have advocated the use of mtDNA in species delimitation, given its rapid attainment of reciprocal monophyly (relative to nuclear genes) and frequent ability to diagnose populations (Moore, 1994; Moritz, 1994, 2002; Wiens and Penkrot, 2002). Reciprocal monophyly of mtDNA is attractive because it is an objective criterion that can be applied to any animal system. However, use of mtDNA alone to delimit species has been criticized, because the complete organismal history is not captured and because many other loci in the genome may not exhibit reciprocal monophyly (Hudson and Coyne, 2002; Sites and Crandall, 1997; Wiens and Penkrot, 2002).
Another gene-tree-based criterion for species delimitation calls for finding reciprocal monophyly among a large majority (≈95%) of sampled nuclear genes (Avise and Ball, 1990; Baum and Shaw, 1995). Because nuclear DNA (nDNA) achieves reciprocal monophyly much slower than mtDNA, this criterion is considered conservative (Hudson and Coyne, 2002; Sites and Crandall, 1997). However, to date, and certainly at the time this species concept was put forward, there have been no avian data sets consisting of multiple independent gene trees with which to test the utility of this approach. A recent study of speciation in three Australian grassfinches (Poephila) using 30 independent nuclear loci provides a test of this concept. One of the taxa (Poephila cincta) examined is phenotypically and geographically very distinct from the other two, and its species status has never been questioned (Schodde and Mason, 1999); the two allopatric western lineages of long-tailed finches (Poephila acuticauda/ hecki) are distinguished by fewer characters but are nonetheless diagnosable morphologically and have been designated phylogenetic species (Cracraft, 1986). Coalescent estimates of population divergence times sug-
gest a split of ≈0.5 million years ago for the two western lineages, and >0.7 million years ago for the basal split with cincta. Given the dynamics of nuclear genes under reasonable assumptions of ancestral population sizes for birds, it is therefore not surprising that considerable heterogeneity and conflict among the gene trees in Poephila was observed, even with regard to alleles sampled from the divergent cincta lineage (Fig. 6.1a) (Nei, 1987; Wakeley and Hey, 1997). There is reason to believe that these conflicts are the result of incomplete lineage sorting, rather than of hybridization or gene flow (Avise, 2000). Application of the criterion of genealogical concordance among ≈95% of sampled gene trees would result in lumping all three taxa together despite their considerable temporal and phenotypic divergence. We suggest that concordance among multiple nuclear genes will rarely be achieved among avian lineages that are considered good species by multiple other criteria; as in Drosophila (Ting et al., 2000), if the characters leading to diagnosibility diverge by natural selection, they may outpace the well known but slow progression of the neutral nuclear genome from paraphyly to polyphyly to reciprocal monophyly (Neigel and Avise, 1986).
Although the incidence of reciprocal monophyly among genes and organisms may vary considerably, it is instructive to examine further the efficacy of this criterion in delimiting species of Poephila. The probability that a gene tree will match the species tree in a three-species scenario of divergence and isolation without gene flow has been known for some time (Hudson, 1992; Nei, 1987) and is determined by T, the ratio of time elapsed between speciation events to the ancestral effective population size (Fig. 6.1b). T must therefore equal or exceed 2.6 if there is to be significant concordance between the gene trees and species tree. The multilocus Poephila data suggests a value for T of 0.4, implying a substantial ancestral population size relative to divergence time, also the likely cause for the lack of concordance among gene trees. The probability of incongruence between gene and species trees for the Poephila loci are not strictly equivalent to the genealogical concordance among loci described by Avise and Ball (1990), because in the Poephila study only a single allele was sampled per species; thus, some of the loci exhibiting congruence with the species tree (Topology 1, Fig. 6.1a) may in fact exhibit genealogical incongruence, manifested as incomplete lineage sorting, upon sampling of further alleles. Nonetheless, these data, the first substantial sampling of multiple gene trees for an avian species, suggest that species arising rapidly or having ancestors with large effective population sizes will not satisfy the concordant genealogies criterion even though they are reasonable species under the BSC or morphological PSC. For avian species, finding any nuclear gene that has achieved reciprocal monophyly, whether by
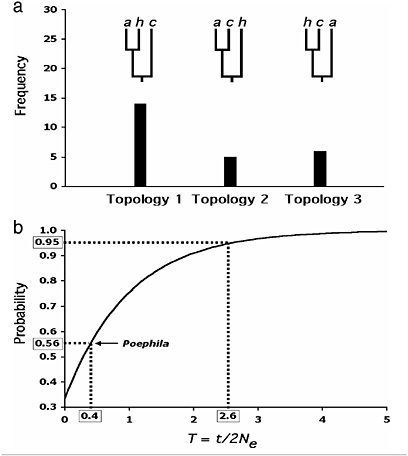
FIGURE 6.1 Conflicts between gene and species trees in Poephila finches, and their implications. (a) Frequency distribution of all three possible gene trees encountered in a survey of nuclear DNA sequence variation among three species of Australian grassfinches (Poephila). Branch tips are labeled as follows: a, P. acuticauda; h, P. hecki; c, P. cincta. P. cincta vs. P. acuticauda/hecki is a well established divergence across the Carpentarian barrier in northern Australia and is supported by numerous species-level phenotypic differences; P. hecki vs. acuticauda are allopatric but represent a more recent taxonomic split which is regarded by some (Cracraft, 1986) as phylogenetic species. In the genetic survey, a single allele was sampled from each of the three taxa for thirty presumably unlinked loci. Twenty-five gene trees exhibiting all three possible topologies were unambiguously reconstructed. Gene trees for five loci could not be resolved. Topology 1 reflects the presumed species tree. (b) Unconditional probabilities of a gene tree being congruent with the species tree over a range of values for T (after Hudson, 1992). The curve is based on the equation for three species [P(congruent gene tree) = 1 − 2/3e−T (Hudson, 1992; Nei, 1987), where T is equal to t/2Ne, t is defined as the time between speciation events (in generations), and 2Ne is twice the size of the effective population size of the basal ancestor. When T = 0, the topology of a gene tree of three sampled alleles is expected to be random with respect to the species tree (i.e., probability of 0.33). Note that T must be at least as large as 2.6 for a 0.95 probability of congruence, yet the empirically derived probability for Poephila finches of 0.56 results in a T value of only 0.4. The Poephila tree congruence probability is based on the fact that 14 of 25 independent gene trees matched the presumed species tree observed by W.B.J. and S.V.E. (unpublished work).
neutral or selective means (akin to finding a fixed diagnostic phenotypic trait), may be a reasonable criterion for delimiting species.
It is certain that molecular approaches will continue to play an important role in species delimitation. However, the battle over species delimitation between the nuclear and mitochondrial genomes, and between the BSC and PSC, will have no victor. Nuclear gene trees will not provide enough phylogenetic resolution to satisfy avian systematists. Furthermore, the high levels of recombination detected in the first surveys of avian SNP variation appear prohibitive for standard phylogenetic analysis (Edwards and Dillon, 2004; International Chicken Polymorphism Map Consortium, 2004). On the other hand, in our view, maternally inherited mtDNA can never capture enough of a species history to delimit species on its own. Although mtDNA will frequently deliver clean phylogeographic breaks within avian species, these breaks need not have their origin in reproductive isolation (Irwin, 2002) and may in some cases be driven by natural selection (Hudson and Turelli, 2003). These same species showing clean mitochondrial breaks will frequently look very messy with regard to nuclear gene splits, as decades of allozyme analyses have already confirmed. We suggest that, despite its disproportionate contribution to revealing phylogeographic patterns and its ability to reflect cessation of female gene flow more rapidly than nuclear genes, mtDNA should not have priority over nuclear genes in avian species delimitation. Ours is a generation of avian systematists raised primarily on single locus analyses of avian phylogeny and divergence. But nuclear genes can and should be interrogated in questions of avian taxonomy, even if the interpretation of nuclear histories and the contrast with mtDNA histories will be challenging.
The Role of Song in Allopatric and Sympatric Speciation
As suggested by Mayr (1942), divergence of populations in allopatry appears to be the dominant mode of speciation in birds. Molecular phylogenies have provided new opportunities for testing alternative geographic models, and avian sister taxa generally meet the expectation of having allopatric distributions, whether because of vicariance or dispersal (Drovetski et al., 2004; Pereira and Baker, 2004; Voelker, 1999; Zink et al., 2000). Current distributions, however, do not necessarily reflect the geo-
graphic context of speciation given the potential for dispersal and range expansion in birds (Chesser and Zink, 1994). When expectations are derived from a model incorporating random changes in geographic range, phylogenetic data suggest allopatric speciation as the predominant mode in several avian groups, with sympatry due to range changes after speciation (Barraclough and Vogler, 2000). Greater asymmetry in range size for recently evolved species also implies a role for peripatric speciation (Barraclough and Vogler, 2000), as suggested in other recent avian studies (Drovetski and Ronquist, 2003; Johnson and Cicero, 2002; Omland, 1997). The lability of geographic ranges, however, ultimately limits the power of phylogenetic approaches to distinguish between alternative geographic models of speciation (Losos and Glor, 2004).
Greater insight into avian speciation is perhaps gained by focusing on the processes of population divergence and mechanisms of reproductive isolation, particularly in closely related taxa and/or diverging populations. Both ecological and sexual selection may contribute to rapid morphological and behavioral divergence in allopatric or parapatric populations (Rasner et al., 2004; Uy and Borgia, 2000; Yeh, 2004). Whether these changes lead to speciation, however, depends on the evolution of reproductive isolation before or after secondary contact. Avian species typically retain hybrid viability and fertility for millions of years after speciation, reflecting a general lack of intrinsic isolating mechanisms among closely related species (Price and Bouvier, 2002). Although ecological and/ or sexual selection against hybrids may help to maintain species boundaries, reproductive isolation in birds will often depend on prezygotic mechanisms. Thus, divergence in characters involved in mate choice, such as song, plumage, and behavioral displays, likely play a central role in avian speciation. The role of song is particularly interesting given the multiple factors influencing vocal evolution and the potential for rapid change through learning and cultural evolution.
Gradual divergence of song in allopatric populations may result in reproductive isolation upon secondary contact. Although generally difficult to observe, this process is captured in present day populations of the greenish warbler (Phylloscopus trochiloides) complex. The range of this Old World species forms a narrow ring around the southern margin of the Tibetan plateau with eastern and western populations extending northward, expanding longitudinally, and meeting in Siberia. Genetic composition and songs change gradually through the nearly continuous ring of intergrading populations, but eastern and western populations in the north are reproductively isolated because of differences in song (Irwin et al., 2001b). Parallel sexual selection for increased song complexity in northern latitudes has apparently resulted in the stochastic divergence of songs in eastern and western populations (Irwin, 2000). The gradual intergrada-
tion through intermediate populations in this ring species are analogous to the gradual changes that might occur over time in geographically isolated populations diverging in a similar manner (Irwin et al., 2001a).
Songs may diverge as a direct result of habitat-dependent selection or indirectly as a consequence of morphological adaptations, such as those related to foraging. It has long been recognized that different types of vocalizations vary in their quality as signals in different habitats, and recent studies suggest a role for habitat-dependent selection in population divergence and reproductive isolation. Two subspecies of song sparrows (Melospiza melodia) differ both in song characteristics and preferred habitat, with Melospiza melodia hermanii occupying more dense vegetation than Melospiza melodia fallax and singing a lower frequency song with more widely spaced elements, a pattern consistent with acoustic adaptation (Patten et al., 2004). Playback experiments further indicate that both males and females show greater response to homotypic songs, suggesting a role for song in reproductive isolation and the consequent development of significant genetic differentiation between the subspecies (Patten et al., 2004). Similar habitat-dependent vocal divergence accompanies morphological differentiation in the little greenbul (Andropodus virens) and may promote population differentiation across ecological gradients (Slabbekoorn and Smith, 2002b).
Ecological selection on other characters also may result in correlated vocal evolution that contributes to prezygotic reproductive isolation. In Darwin’s finches, bill morphology and vocal characteristics are correlated because of physical constraints on sound production, perhaps contributing to the diversification of these species (Podos, 2001). In contrast, the black-bellied seedcracker (Pyrenestes ostrinus), which shows a similar pattern of divergent selection on bill morphology, shows no effect of bill size on vocal performance, contributing to the conclusion that bill size variation in this species reflects only intraspecific niche polymorphism and not incipient speciation (Slabbekoorn and Smith, 2000; Smith, 1993).
As a learned behavior in many birds, song is subject to rapid cultural evolution in which stochastic innovations or errors in copying are spread as individuals learn songs from their parents and/or neighbors (Grant and Grant, 1996; Payne, 1996). Song learning may also increase the rate at which genetic predispositions to learn or prefer certain songs evolve in allopatry (Lachlan and Servedio, 2004). Song learning, however, may sometimes inhibit reproductive isolation upon secondary contact if there has been only minimal divergence in the capacity to learn particular songs and/or the morphological structures affecting sound production (Slabbekoorn and Smith, 2002a). A well known example of reproductive character displacement involves the Ficedula flycatchers in which a sexually selected male plumage trait shows greater divergence in sympatry
than in allopatry (Saetre et al., 1997). Results were mixed, however, in a recent analysis of vocal divergence in this system (Haavie et al., 2004). In sympatry, songs of pied flycatchers Ficedula hypoleuca have converged on those of collared flycatchers Ficedula albicollis because of heterospecific copying and singing of mixed songs. Although collared flycatcher songs in the zone of contact have diverged away from pied flycatcher songs typical of allopatric populations, the net effect of these changes is greater song similarity in sympatry.
Allopatric divergence of songs among suboscines and other birds in which differences in song are genetically determined may evolve more slowly but should also contribute to reproductive isolation. In a large comparative analysis of antbirds (Thamnophilidae), Seddon (2005) found evidence of vocal divergence both as a correlated effect of morphological evolution and as a response to habitat-dependent selection on signal transmission. In addition, among trios of closely related species, sympatric forms exhibited striking vocal divergence in comparison to allopatric taxa, providing strong evidence for reproductive character displacement and the role of song in reproductive isolation. Despite divergence of song in allopatry, individuals of different species may recognize each other as potential mates upon secondary contact, leading to hybridization. If song differences are genetically determined, hybrid offspring may have intermediate songs (de Kort et al., 2002).
Given divergence in vocalizations, the development of song preferences through sexual imprinting may contribute to reproductive isolation even without genetic evolution of female preferences (Irwin and Price, 1999; Price, 1998). In Darwin’s finches, for example, cultural inheritance clearly plays a greater role than bill morphology in determining songs and song preferences (Grant and Grant, 1996, 1997) and is critical in promoting reproductive isolation after secondary contact of populations that have diverged in feeding adaptations in allopatry (Grant and Grant, 2002). Ecological selection against hybrid individuals also helped maintain species boundaries, at least before changes associated with El Niño events in the 1980s (Grant and Grant, 1998). In the past 20 years, however, increased fitness of hybrids has resulted in substantial genetic introgression from Geospiza fortis to Geospiza scandens on Daphne Major (Grant et al., 2004). Although learned songs and song preferences are strong determinants of pair formation in these species, reproductive isolation is imperfect because of constraints on mate choice imposed by asymmetries in population size and operational sex ratios as well as infrequent cases of individuals misimprinting on heterospecific songs (Grant and Grant, 1996, 1997, 1998).
Song learning and sexual imprinting explain the recent diversification of brood parasitic indigobirds (genus Vidua), the best and perhaps
only example of sympatric speciation in birds (Fig. 6.2). Male indigobirds include mimicry of host song in a repertoire that also includes speciesspecific indigobird vocalizations learned from other male indigobirds mimicking the same host (Payne et al., 1998). Likewise, female preferences for both male song and host nests result from imprinting on the host (Payne et al., 2000). Thus, speciation in indigobirds begins with reproductive isolation as a consequence of host colonization and only then proceeds to divergence in other characters, including host-specific mimicry of mouth markings and colors by indigobird chicks. Indigobird species within a region show a pattern of incomplete but significant genetic differentiation (Sorenson et al., 2003) but also genetic continuity across intermediate spatial scales (Sefc et al., 2005), a pattern consistent with recent sympatric speciation and current reproductive isolation (Via, 2001). As in Darwin’s finches, song learning likely plays a role in hybridization between indigobird species. When females parasitize the host of another indigobird species (Payne et al., 1993), their offspring are likely to hybridize in the subsequent generation because they have imprinted on the songs of the alternate host (Payne and Sorenson, 2004). The frequency of misimprinting and hybridization, however, appears to be lower in indigobirds than among the Geospiza ground finches (Grant et al., 2004; Payne et al., 1993).
In birds generally, the importance of prezygotic isolating mechanisms may allow for rapid speciation, whereas the slow development of intrinsic postzygotic isolation will facilitate continuing hybridization. Closely related taxa may therefore be strongly differentiated at only a small number of loci influenced by divergent ecological or sexual selection. Loci “that can be shown to cause some degree of ecological, sexual or postmating isolation between young, or even nascent, species” are good candidates for speciation genes (Wu and Ting, 2004). Finding such genes and understanding the genetics of avian speciation are challenging but increasingly realistic objectives as genomic resources and molecular methods continue to evolve.
Sex Chromosomes and Avian Speciation
The architects of the Modern Synthesis laid the foundation for a body of work that has resulted in two “rules” of speciation that directly implicate sex chromosomes: Haldane’s rule and the large X(Z)-effect (Coyne and Orr, 1989). Haldane’s rule is the preferential sterility or inviability of the heterogametic sex in hybrid crosses, when a sex-biased fitness loss in hybrids occurs. This phenomenon is found across diverse taxa, including butterflies and birds in which the female is heterogametic (ZW) (Haldane, 1922). Although the X chromosome was experimentally implicated in
Haldane’s rule and hybrid male sterility in Drosophila decades ago (Dobzansky, 1936), it was not until recently that this large effect of the hemizygous sex chromosome was documented for birds, using genetic data from natural hybrid zones and domesticated species (Price, 2002; Saetre et al., 2003). In this section, we review empirical and theoretical work that explores these two rules of speciation in birds.
The phenomenon of Haldane’s rule describes patterns of postzygotic incompatibilities in hybrids and is likely caused by negative epistatic interactions between loci derived from divergent parental genomes (Coyne and Orr, 2004). Heterogametic hybrids are more severely affected by these interactions because, unlike the homogametic sex, they fully express recessive sex-linked genes. Interestingly, avian and Lepidopteran F1 hybrid females may suffer from an additional source of negative epistasis between parental genomes, namely that between the maternally derived mitochondria or cytoplasm and the paternally derived Z chromosomes (Presgraves, 2002). There is debate over the extent to which Haldane’s rule is driven by interactions among sometimes rapidly diverging sex chromosomes per se, or whether it is the peculiar dominance patterns exhibited by the hemizygous sex chromosomes that underlie the rule. Support for Haldane’s rule is excellent in birds based on experimental studies of hybrid fitness in ducks (Tubaro and Lijtmaer, 2002), pigeons and doves (Lijtmaer et al., 2003), and many other avian taxa (Price and Bouvier, 2002).
Price and Bouvier (2002) characterized patterns of postzygotic incompatibilities in birds using published data from 254 hybrid crosses and found that the order in which incompatibilities accumulate with increasing species divergence differs between birds and other taxa, a pattern that informs the causes of Haldane’s rule in birds (Fig. 6.3). In Drosophila, male sterility appears at early stages of divergence, followed in turn by male inviability, then female sterility, and finally female inviability. By contrast, avian incompatibilities accumulate in the following order: female sterility, male sterility, female inviability, and male inviability (Price and Bouvier, 2002). Thus, in birds, homogametic (male) sterility evolves at earlier stages than does homogametic (female) sterility in Drosophila (Coyne and Orr, 1989). The appearance of homogametic (male) sterility before heterogametic (female) inviability in birds may reflect a general trend, regardless of sex-chromosome system, of the rapid evolution of male reproductive genes via sexual selection, resulting in high divergence between species at these loci (Wu and Davis, 1993). However, the rapid evolution of male reproductive genes via sexual selection—so-called “faster-male evolution” or the “sexual selection model” of Wu and Davis (1993)—works in opposition to Haldane’s rule in birds because this particular force will negatively affect the homogametic sex (males), not the
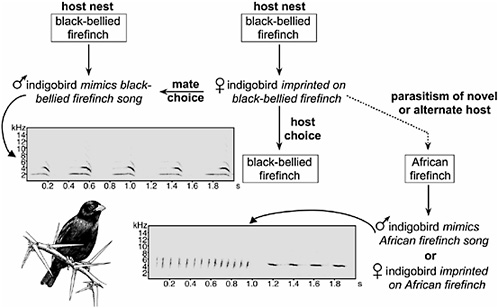
FIGURE 6.2 Behavioral imprinting maintains host specificity and genetic cohesion of indigobird species but also provides a mechanism for rapid speciation when new hosts are colonized. Male indigobirds mimic the songs of their hosts, whereas females use song cues to choose both their mates and the nests they parasitize. Rarely, females lay in the nest of a novel or alternate host. The resulting offspring imprint on the novel host and are therefore reproductively isolated from their parent population. Indigobird drawing by Karen Klitz.
heterogametic sex (females), in ZW species. By contrast, sexual selection accentuates the pattern for sterility in XY species, such as Drosophila, perhaps explaining the preponderance of cases of hybrid male sterility in the genus (Coyne and Orr, 2004).
Support for sexual selection as a modulator of Haldane’s rule in birds was first detected in the relative proportions of cases of Haldane’s rule for sterility and inviability (Kirkpatrick and Hall, 2004). Specifically, the majority of cases of Haldane’s rule in XY species involve heterogametic sterility, whereas Wu and Davis (1993) initially found as many cases for heterogametic inviability as for sterility in birds and other ZW species. This pattern is expected when traits expressed only in males are evolving rapidly because it results in male hybrid sterility in early stages of divergence, resulting in fewer cases of Haldane’s rule for sterility. However, in the larger avian data set of Price and Bouvier (2002), five times as many cases of Haldane’s rule involve sterility as inviability, a pattern that does not support sexual selection as a modulator of Haldane’s rule. Direct evidence for a role for sexually selected genes in avian speciation, such as an examination of the molecular evolution of avian reproductive genes (see next section), is a necessary but not sufficient condition for their role Haldane’s rule in birds. Additionally, the domain of such sexually selected genes needs to be determined: Do they include sexually selected traits such as plumage and song that are not directly involved in the physical production of gametes? Because males and females of many bird species exhibit correlated evolution of plumage and other sexually selected traits, the specific expression patterns of genes related to both reproduction and morphology need to be investigated to determine how such traits will influence Haldane’s rule.
The rate of evolution of sex chromosomes has important implications for their role in avian speciation. In general, when rapidly evolving loci diverge between species, incompatibilities between these and other loci can arise when parental genomes come together in hybrids. In the same way that rapid evolution of sexual traits in males can produce incompatibilities in hybrids at loci involved in male sexual traits—whether autosomal or sex-linked—a faster rate of evolution of the sex chromosomes, combined with effects of dominance (Coyne and Orr, 2004), is one hypothesis for the large X(Z)-effect in low-fitness hybrids. So-called “faster-X(Z) evolution” owing to selection on favorable mutations on the hemizygous X(Z) when they are partially recessive, manifested as more rapid substitution rates on the X chromosome than on autosomes, was demonstrated theoretically by Charlesworth et al. (1987). More recently, Kirkpatrick and Hall (2004) modeled the relative rates of evolution of sex chromosomes and autosomes while accounting for interactions between mode of inheritance and the intrinsically higher mutation rate in males
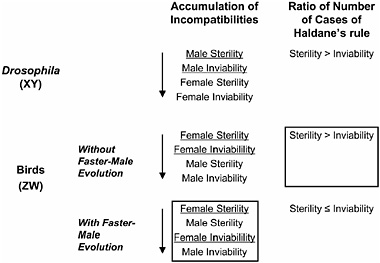
FIGURE 6.3 Summary of Haldane’s rule in birds. Predictions are based on the presence or absence of faster-male evolution (sexual selection hypothesis), which changes the order of accumulation of incompatibilities in WZ species. The predicted order of accumulation of incompatibilities for the case without sexual selection is based on the pattern in Drosophila. The data observed by Price and Bouvier (2002) are boxed. The heterogametic sex is underlined.
known as “male-driven evolution.” Male-driven evolution (not to be confused with faster-male evolution) has been documented in birds, mammals, and plants, most likely because of a greater number of cell divisions in the male than in the female germ line (Hurst and Ellegren, 1998); critically, such studies have documented faster neutral evolution on the avian Z but not faster adaptive, presumably nonsynonymous evolution, as predicted by Charlesworth et al. (1987). Values of α, the ratio of male to female mutation rates, have been estimated from sequence data and range from ≈1.8 to ≈6.4 in both birds and mammals (reviewed in Wu and Ting, 2004). The avian Z chromosome is expected to have a higher neutral mutation rate than the autosomes because it passes through the male germ line twice as often as the female germ line, whereas avian autosomes spend equal time in both germ lines. In contrast to predictions of hemizygous sex chromosome evolution in mammals, where mutations must be strongly recessive for the X to evolve faster than autosomes (Charlesworth et al., 1987), the higher mutational flux on the Z of birds is predicted to produce faster rates of Z chromosome evolution relative to autosomes even when mutations are strongly dominant (Kirkpatrick and Hall, 2004). Perhaps not surprisingly, support for fast-X evolution is inconsistent (Betancourt et al., 2002; Countermann et al., 2004), whereas
evidence of fast-Z evolution is more compelling (Axelsson et al., 2003; Bartosch-Harlid et al., 2003; Carmichael et al., 2000; Ellegren and Fridolffson, 1997; Kahn and Quinn, 1999). Sequence data flowing from the chicken genome project have already facilitated confirmation of faster-Z evolution in birds, both at the intra- and interspecific levels (Axelsson et al., 2003).
The first empirical inquiries into the role of sex chromosomes in the speciation process of natural avian groups have focused on the well studied system provided by hybridization in Old World Ficedula flycatchers. Saetre et al. (2003) found that Z-linked single nucleotide polymorphism (SNP) markers in F. hypoleuca and F. albicollis showed little evidence for introgression, whereas substantial introgression was documented for autosomal SNPs. The sex chromosomes had a large effect on the fertility of hybrids: Among birds with heterozygous sex chromosomes (one from each parental species), 7 of 7 females were sterile, as opposed to 3 of 11 males; a pattern that is consistent with Haldane’s rule, although whether interactions among Z loci or between the Z and W or autosomes is the cause remains unclear. The possibility of linkage of loci involved in pre-and postzygotic isolation in this system (Saetre et al., 2003) motivated the development of a novel model of reinforcement (Servedio and Saetre, 2003). The model focused on the evolution of linkages among a male trait locus, a female preference locus (collectively referred to as “mating loci”), and two postzygotic incompatibility loci. Consistent with previous theory, prezygotic isolation is reinforced through tight linkage between the mating loci and incompatibility loci. As incompatibility loci diverge between the two populations, causing a decrease in the fitness of hybrids, the frequency of assortative mating increases, thereby reducing the occurrence of interspecific matings. The tight linkage between the mating loci and the incompatibility loci creates a positive feedback loop because the frequency of linked incompatibility loci increases in tandem with the loci causing population-specific mating as a result of genetic hitchhiking. The positive feedback loop is enhanced when the linkage group occurs on a sex chromosome for two reasons. First, if as assumed in the model crossing over does not occur between sex chromosomes in the heterogametic sex, hitchhiking is enhanced. (This lack of crossing over does not strictly represent the case in nature because, as in mammals, many birds possess a pseudoautosomal region of varying size in which crossing over does in fact occur.) Second, recessive mutations are more exposed to positive selection in the heterogametic sex and may rapidly sweep through the population. The model reveals a number of interesting features about the dynamics of the Ficedula hybrid zone (Saetre et al., 2003). However, it is unclear how well the Ficedula hybrid zone represents the diversity of avian hybrid zones. Many avian hybrids show little evidence of fitness
loss (Grant and Grant, 1992) and may even enjoy an ecological advantage over parental species (Good et al., 2000), conditions that do not favor reinforcement of prezygotic isolation.
Studies in birds and other taxa indicate that sex chromosomes may disproportionately harbor genes related to sex and reproduction (Reinhold, 1998; Wang et al., 2001). Literature on a variety of domesticated avian species suggests that 22% of traits that distinguish breeds, including likely targets of sexual selection in natural populations such as plumage and vocalizations, are sex-linked (Price, 2002), an excess when one considers that the Z chromosome comprises ≈2.7% of the chicken genome, by our estimate using data from the chicken genome project (International Chicken Polymorphism Map Consortium, 2004). Avian speciation is commonly demonstrated to involve prezygotic isolation in traits such as song or display (Grant and Grant, 1997; see previous section), making the genes that underlie these traits promising candidate speciation genes (Wu and Ting, 2004). The dawn of the genomic era for ornithologists provides exciting opportunities to study the genomic composition of avian sex chromosomes and will allow a better understanding of the complex interaction between their gene content, gene expression patterns, and rate of evolution in the context of speciation.
CRYPTIC MATE CHOICE AND CONFLICT: A ROLE FOR REPRODUCTIVE PROTEINS IN AVIAN SPECIATION?
Birds provide an abundance of examples of intense sexual selection through cryptic female choice and sperm competition (Birkhead and Møller, 1992). The dramatic evolutionary consequences of this competition have been documented at the molecular level in mammals and invertebrates, through genes collectively known as reproductive proteins. Rapid evolution of reproductive proteins has been documented in a diverse array of taxonomic groups ranging from humans to corn and is thought to be a component of the speciation process (Swanson and Vacquier, 2002). However, to our knowledge there are no examples of this process from birds. One potential protein is the female-specific gene HNTW, which, although its function is unknown, shows adaptive evolution (Ceplitis and Ellegren, 2004). With the recent draft release of the chicken genome (International Chicken Genome Sequencing Consortium, 2004) we can expect that the molecular evolution of avian reproductive genes will come under intense scrutiny, a development that is particularly exciting because birds offer unique opportunities to study the selective forces driving the rapid evolution of reproductive proteins.
Reproductive genes can be split into two classes. First, there are gamete recognition proteins on the surface of gametes that are directly in-
volved in sperm–egg interaction. The classic example of a rapidly evolving gamete recognition protein is the abalone sperm protein lysin, perhaps the most rapidly diverging protein yet discovered (Lee and Vacquier, 1992). Lysin acts to dissolve the egg vitelline envelope, a process that demonstrates species specificity. In mammals it has been demonstrated that sperm and egg molecules are among the most diverse found within the genome, with a minimum of 10 reproductive genes showing evidence of adaptive evolution (Swanson et al., 2003b). One such gene is the mammalian egg coat protein ZP3. The region within this protein undergoing adaptive evolution corresponds to experimentally determined binding sites (Swanson et al., 2001a), suggesting that the rapid evolution relates to fertilization.
The second class of reproductive proteins exhibiting rapid evolutionary change are not directly involved in surface recognition of the gametes. These include components of seminal fluid (Kingan et al., 2003), pheromones and protamines. In Drosophila seminal fluid, an estimated 10% of the genes show the signatures of adaptive evolution (Swanson et al., 2003a). Many of these genes, called accessory gland proteins (ACPs) act to manipulate female reproductive behavior, thus increasing male fitness (Wolfner, 1997). In primates, seminogellin II (SEM2), a major component of seminal fluid, shows rapid adaptive evolution. This protein is involved in copulatory plug formation in rodents, and in primates its rate of evolution shows a correlation with mating system (Dorus et al., 2004).
The recent comparison of predicted genes in chicken genome with the human genome supports the pattern of divergence found in reproductive proteins across taxa (International Chicken Genome Sequencing Consortium, 2004). Genes implicated in reproduction appear less conserved between chicken and human than genes involved in typical “housekeeping” functions. For example, among genes classified into 10 different tissue specificities, those expressed in the testis showed the most divergence: 65% sequence conservation compared with the mean of 75% across all genes. Many of these genes, such as ZP3, have orthologues in birds and would be important candidates for targets of natural selection. Other potential target genes in birds include seminal fluid proteins. The prediction of natural selection on such genes in other species can be inferred from reproductive observations. Adkins-Reagan (1999) documented a viscous mucoprotein produced by Japanese quail (Corturnix japonica) thought to increase the probability of fertilization when a hard-shelled egg is in the uterus. The origin of such viscosity must have a basis in protein evolution, although the target loci have not yet been identified.
Cryptic female choice, sperm competition, and sexual conflict are three nonexclusive hypotheses for the forces driving the rapid evolution of these proteins (Swanson and Vacquier, 2002). Cryptic female choice of
reproductive proteins involves the “preference” of male proteins on the surface of the sperm or in the seminal fluid by egg coat proteins, egg proteins, or proteins in the female reproductive tract, examples of which come mostly from invertebrates (Palumbi, 1999). The ability of many birds to store sperm provides a ready mechanism for cryptic female choice. Sperm competition involves the direct competition between sperm of different males providing a potent source selection acting to improve sperm motility (Birkhean and Møller, 1992). Sexual conflict occurs when the reproductive goals of the sexes differ and is thought to drive rapid coevolutionary arms races in reproductive proteins at the molecular level (Gavrilits, 2000; Gowaty, 1996; Rice, 1996). Testing the various hypotheses for the rapid evolution of reproductive proteins is particularly promising in birds. In no other taxonomic group is so much known about the diversity of mating systems and the natural history of female preferences driving trait differences (Parker and Burley, 1998). Combining evolutionary data on reproductive proteins with predictions of sperm competition and mate choice from behavioral studies, as has been attempted with the SEM2 locus in primates (Dorus et al., 2004), is a promising avenue of research in birds. Furthermore, reproductive traits unique to birds, such as physiological polyspermy, the ability of multiple sperm to penetrate the egg without rendering it inviable, permit testing of specific hypothesis underlying rapid adaptive protein evolution. Also, variation across avian species in particular traits such as the presence and type of intromittent organs, from penises to cloacal protuberances to the absence of any intromittent organ, allows for hypothesis-testing that would be difficult in taxonomic groups without such variation. Conversely, the study of reproductive protein evolution in birds might help clarify the roles of these traits on reproductive evolution and reinforcement. Avian mating systems are thought to play an important role in the speciation process (Jennions and Petrie, 2000), and reproductive proteins provide a convenient link between these two arenas.
CONCLUSION
Rich natural histories, diverse biogeographies, and complex character traits and mating systems have made birds central to the formulation of many speciation theories. Now, these and other ideas are more amenable to direct testing with the increased molecular access provided by the chicken genome and by new genomic technologies and resources. These new tools will increase the resolving power of both phylogeographic analysis of speciation and of interactions among diverging genomes. Largescale EST surveys, bacterial artificial chromosome (BAC) libraries, and other genetic resources for dimorphic species such as zebra finches,
turkeys, and Japanese quail are available, and high-resolution linkage maps will soon follow. We can expect the information from these model avian species to inform the analysis of speciation within their respective clades and beyond (Edwards et al., 2005). Linking of these genetic analyses with predictions from theory and application to natural populations will make for exciting times to come in avian speciation studies.
ACKNOWLEDGMENTS
We thank K. De Quieroz, J. Hey, H. A. Orr, C. Filardi, and J. Wakeley for helpful discussion and R. Payne, G. P. Saeter, T. Price, M. Servedio, and J. Feder for very constructive comments on the manuscript. This work was supported in part by National Science Foundation grants (to S.V.E. and M.D.S.). W.J.S. was supported by National Institutes of Health Grant HD42563-01A1 and National Science Foundation Grant DEB-0410112.
REFERENCES
Adkins-Reagan, E. (1999) Foam produced by male coturnix quail what is its function? Auk 116, 184–193.
Avise, J. C. (2000) Phylogeography: The History and Formation of Species (Harvard Univ. Press, Cambridge, MA).
Avise, J. C. & Ball, R. M. (1990) Principles of genealogical concordance in species concepts and biological taxonomy. Oxf. Surv. Evol. Biol. 7, 45–67.
Axelsson, E., Smith, N. G., Sundstrom, H., Berlin, S. & Ellegren, H. (2004) Male-biased mutation rate and divergence in autosomal, Z-linked and W-linked introns of chicken and turkey. Mol. Biol. Evol. 21, 1538–1547.
Barraclough, T. G. & Vogler, A. P. (2000) Detecting the geographical pattern of speciation from species-level phylogenies. Am. Nat. 155, 419–434.
Barrowclough, G. F. (1983) In Perspectives in Ornithology: Essays Presented for the Centennial of the American Ornithologists’ Union, eds. Brush, A. H. & Clark, G. A., Jr. (Cambridge Univ. Press, New York), pp. 223–261.
Barton, N. H. & Charlesworth, B. (1987) Genetic revolutions, founder effects and speciation. Annu. Rev. Ecol. Syst. 15, 133–164.
Bartosch-Harlid, A., Berlin, S., Smith, N. G., Moller, A. P. & Ellegren, H. (2003) Life history and the male mutation bias . Evolution (Lawrence, Kans.) 57, 2398–2406.
Bates, J. M., Hackett, S. J. & Cracraft, J. (1998) Area-relationships in the Neotropical lowlands: An hypothesis based on raw distributions of Passerine birds. J. Biogeogr. 25, 783–793.
Baum, D. & Shaw, K. L. (1995) In Experimental and Molecular Approaches to Plant Biosystematics, eds. Hoch, P. C. & Stephenson, A. C. (Missouri Botanical Garden, St. Louis), pp. 289–303.
Betancourt, A. J., Presgraves, D. C. & Swanson, W. J. (2002) A test for faster X evolution in Drosophila. Mol. Biol. Evol. 1816–1819.
Birkhead, T. R. & Møller, A. P. (1992) Sperm Competition in Birds; Evolutionary Causes and Consequences (Academic, London).
Carmichael, A. N., Fridolfsson, A. K., Halverson, J. & Ellegren, H. (2000) Male-biased mutation rates revealed from Z and W chromosome-linked ATP synthase alpha-subunit (ATP5A1) sequences in birds. J. Mol. Evol. 50, 443–447.
Ceplitis, H. & Ellegren, H. (2004) Adaptive molecular evolution of HINTW, a female-specific gene in birds. Mol. Biol. Evol. 21, 249–254.
Charlesworth, B., Coyne, J. A. & Barton, N. H. (1987) The relative rates of evolution of sex-chromosomes and autosomes. Am. Nat. 130, 113–146.
Chesser, R. T. & Zink, R. M. (1994) Modes of speciation in birds—A test of Lynchs method. Evolution (Lawrence, Kans.) 48, 490–497.
Countermann, B. A., Ortiz-Barrientos, D. & Noor, M. A. F. (2004) Using comparative genomic data to test for fast-X evolution. Evolution (Lawrence, Kans.) 656–660.
Coyne, J. A. & Orr, H. A. (1989) In Speciation and Its Consequences, eds. Otte, D. & Endler, J. A. (Sinauer, Sunderland, MA), pp. 180–207.
Coyne, J. A. & Orr, H. A. (2004) Speciation (Sinauer, Sunderland, MA).
Coyne, J. A. & Price, T. D. (2000) Little evidence for sympatric speciation in island birds. Evolution (Lawrence, Kans.) 54, 2166–2171.
Cracraft, J. (1983) Species concepts and speciation analysis. Curr. Ornithol. 1, 159–187.
Cracraft, J. (1986) Origin and evolution of continental biotas: speciation and historical congruence within the Australian avifauna. Evolution (Lawrence, Kans.) 40, 977–996.
Cracraft, J. (1991) Patterns of diversification within continental biotas: Hierarchical congruence among the areas of endemism of Australian vertebrates. Aust. Syst. Bot. 4, 211–227.
Crandall, K. A., Bininda-Emonds, O. R. P., Mace, G. M. & Wayne, R. K. (2000) Considering evolutionary processes in conservation biology. Trends Ecol. Evol. 15, 290–295.
de Kort, S. R., den Hartog, P. M. & ten Cate, C. (2002) Diverge or merge? The effect of sympatric occurrence on the territorial vocalizations of the vinaceous dove Streptopelia vinacea and the ring-necked dove S. capicola. J. Avian Biol. 33, 150–158.
Dobzansky, T. (1936) Studies on hybrid sterility. II. Localization of sterility factors in Drosophila pseudoobscura hybrids. Genetics 21.
Dorus, S., Evans, P. D., Wyckoff, G. J., Choi, S. S. & Lahn, B. T. (2004) Rate of molecular evolution of the seminal protein gene SEMG2 correlates with levels of female promiscuity . Nat. Genet. 36, 1326–1329.
Drovetski, S. V. & Ronquist, F. (2003) Plio-Pleistocene climatic oscilations, Holarctic biogeography and speciation in an avian subfamily. J. Biogeogr. 30, 1173–1181.
Drovetski, S. V., Zink, R. M., Rohwer, S., Fadeev, I. V., Nesterov, E. V., Karagodin, I., Koblik, E. A. & Redkin, Y. A. (2004) Complex biogeographic history of a Holarctic passerine. Proc. R. Soc. London Ser B 271, 545–551.
Edwards, S. V. & Dillon, M. (2004) Hitchhiking and recombination in birds: Evidence from Mhc-linked and unlinked loci in red-winged blackbirds (Agelaius phoeniceus). Genet. Res. 84, 175–192.
Edwards, S. V., Jennings, W. B. & Shedlock, A. M. (2005) Phylogenetics of modern birds in the era of genomics. Proc. R. Soc. London Ser. B, in press.
Ellegren, H. & Fridolfsson, A. K. (1997) Male-driven evolution of DNA sequences in birds. Nat. Genet. 17, 182–184.
Filardi, C. E. (2003). Ph.D. dissertation (Univ. of Washington, Seattle).
Filardi, C. E. & Smith, C. E. (2005) Molecular phylogenetics of monarch flycatchers (genus Monarcha) with emphasis on Solomon Island endemics. Mol. Phylogenet. Evol., in press.
Gavrilets, S. (2000) Rapid evolution of reproductive barriers driven by sexual conflict. Nature 403, 886–889.
Good, T. P., Ellis, J. C., Annett, C. A. & Pierotti, R. (2000) Bounded hybrid superiority in an avian hybrid zone: Effects of mate, diet, and habitat choice. Evolution (Lawrence, Kans.) 54, 1774–1783.
Gowaty, P. A. (1996) In Partnerships in Birds, ed. Black, J. M. (Oxford Univ. Press, Oxford), pp. 21–52.
Grant, B. R. & Grant, P. R. (1979) Darwin’s finches: Population variation and sympatric speciation. Proc. Natl. Acad. Sci. USA 76, 2359–2363.
Grant, B. R. & Grant, P. R. (1996) Cultural inheritance of song and its role in the evolution of Darwin’s finches. Evolution (Lawrence, Kans.) 50, 2471–2487.
Grant, B. R. & Grant, P. R. (1998) In Endless Forms: Species and Speciation, eds. Howard, D. J. & Berlocher, S. H. (Oxford Univ. Press, Oxford).
Grant, B. R. & Grant, P. R. (2002) Simulating secondary contact in allopatric speciation: An empirical test of premating isolation. Biol. J. Linn. Soc. 76, 545–556.
Grant, P. R. & Grant, B. R. (1992) Hybridization of bird species. Science 256, 193–197.
Grant, P. R. & Grant, B. R. (1997) Hybridization, sexual imprinting, and mate choice. Am. Nat. 149, 1–28.
Grant, P. R., Grant, B. R., Markert, J. A., Keller, L. F. & Petren, K. (2004) Convergent evolution of Darwin’s finches caused by introgressive hybridization and selection. Evolution (Lawrence, Kans.) 58, 1588–1599.
Haavie, J., Borge, T., Bures, S., Garamszegi, L. Z., Lampe, H. M., Moreno, J., Qvarnstrom, A., Torok, J. & Saetre, G. P. (2004) Flycatcher song in allopatry and sympatry—Convergence, divergence and reinforcement. J. Evol. Biol. 17, 227–237.
Haldane, J. B. S. (1922) Sex ratio and unisexual sterility in animal hybrids. J. Genet. 12, 101–109.
Hey, J. (2001) Genes, Categories, and Species: The Evolutionary and Cognitive Causes of the Species Problem (Oxford Univ. Press, New York).
Hudson, R. R. (1992) Gene trees, species trees, and the segregation of ancestral alleles. Genetics 131, 509–513.
Hudson, R. R. & Coyne, J. A. (2002) Mathematical consequences of the genealogical species concept. Evolution (Lawrence, Kans.) 56, 1557–1565.
Hudson, R. R. & Turelli, M. (2003) Stochasticity overrules the “three-times rule”: Genetic drift, genetic draft, and coalescence times for nuclear loci versus mitochondrial DNA. Evolution (Lawrence, Kans.) 57, 182–190.
Hurst, L. D. & Ellegren, H. (1998) Sex biases in the mutation rate. Trends Genet. 14, 446–452.
International Chicken Genome Sequencing Consortium. (2004) Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432, 695–716.
International Chicken Polymorphism Map Consortium. (2004) A genetic variation map for chicken with 2.8 million single-nucleotide polymorphisms. Nature 432, 717–722.
Irwin, D. E. (2000) Song variation in an avian ring species. Evolution (Lawrence, Kans.) 54, 998–1010.
Irwin, D. E. (2002) Phylogeographic breaks without geographic barriers to gene flow. Evolution (Lawrence, Kans.) 56, 2383–2394.
Irwin, D. E. & Price, T. D. (1999) Sexual imprinting, learning and speciation. Heredity 82, 347–354.
Irwin, D. E., Bensch, S. & Price, T. D. (2001a) Speciation in a ring. Nature 409, 333–337.
Irwin, D. E., Irwin, J. H. & Price, T. D. (2001b) Ring species as bridges between microevolution and speciation. Genetica 112, 223–243.
Jennions, M. D. & Petrie, M. (2000) Why do females mate multiply? A review of the genetic benefits. Biol. Rev. 75, 21–64.
Johnson, N. K. & Cicero, C. (2002) The role of ecologic diversification in sibling speciation of Empidonax flycatchers (Tyrannidae): Multigene evidence from mtDNA. Mol. Ecol. 11, 2065–2081.
Kahn, N. W. & Quinn, T. W. (1999) Male-driven evolution among Eoaves? A test of the replicative division hypothesis in a heterogametic female (ZW) system. J. Mol. Evol. 49, 750–759.
Kingan, S. B., Tatar, M. & Rand, D. M. (2003) Reduced polymorphism in the Chimpanzee semen coagulating protein semenogelin I. J. Mol. Evol. 57, 159–169.
Kirkpatrick, M. & Hall, D. W. (2004) Male-biased mutation, sex linkage, and the rate of adaptive evolution. Evolution (Lawrence, Kans.) 58, 437–440.
Lachlan, R. F. & Servedio, M. R. (2004) Song learning accelerates allopatric speciation. Evolution (Lawrence, Kans.) 58, 2049–2063.
Lee, Y. H. & Vacquier, V. D. (1992) The divergence of species-specific abalone sperm lysins is promoted by positive Darwinian selection. Biol. Bull. (Woods Hole, MA) 182, 97–104.
Lijtmaer, D. A., Mahler, B. & Tubaro, P. L. (2003) Hybridization and postzygotic isolation patterns in pigeons and doves . Evolution (Lawrence, Kans.) 57, 1411–1418.
Losos, J. B. & Glor, R. E. (2003) Phylogenetic comparative methods and the geography of speciation. Trends Ecol. Evol. 18, 220–227.
Mayr, E. (1942) Systematics and the Origin of Species (Columbia Univ. Press, New York).
Mayr, E. (1963) Animal Species and Evolution (Harvard Univ. Press, Cambridge, MA).
Mayr, E. & Diamond, J. (2001) The Birds of Northern Melanesia: Speciation Ecology, and Biogeography (Oxford Univ. Press, New York).
Moore, W. S. (1995) Inferring phylogenies from mtDNA variation: mitochondrial-gene trees versus nuclear-gene trees. Evolution (Lawrence, Kans.) 49, 718–726.
Moritz, C. (1994) Applications of mitochondrial DNA analysis on conservation: A critical review. Mol. Ecol. 3, 401–411.
Moritz, C. (2002) Strategies to protect biological diversity and the evolutionary processes that sustain it. Syst. Biol. 51, 238–254.
Nei, M. (1987) Molecular Evolutionary Genetics (Columbia Univ. Press, New York).
Neigel, J. E. & Avise, J. C. (1986) In Evolutionary Processes and Theory, eds. Karlin, S. & Nevo, E. (Academic, New York), pp. 515–534.
Omland, K. E. (1997) Examining two standard assumptions of ancestral reconstructions: Repeated loss of dichromatism in dabbling ducks (Anatini). Evolution (Lawrence, Kans.) 51, 1636–1646.
Palumbi, S. R. (1999) All males are not created equal: Fertility differences depend on gamete recognition polymorphisms in sea urchins. Proc. Natl. Acad. Sci. USA 96, 12632–12637.
Parker, P. G. & Burley, N. T. (1998) Avian Reproductive Tactics: Female and Male Perspectives (Am. Ornithol. Union, Washington, DC).
Patten, M. A., Rotenberry, J. T. & Zuk, M. (2004) Habitat selection, acoustic adaptation, and the evolution of reproductive isolation. Evolution (Lawrence, Kans.) 58, 2144–2155.
Payne, R. B. (1996) In Ecology and Evolution of Acoustic Communication in Birds, eds. Kroodsma, D. E. & Miller, E. H. (Cornell Univ. Press, Ithaca, NY), pp. 198–220.
Payne, R. B. & Sorenson, M. D. (2004) Behavioral and genetic identification of a hybrid Vidua: Maternal origin and mate choice in a brood-parasitic finch. Auk 121, 156–161.
Payne, R. B., Payne, L. L., Nhlane, M. E. D. & Hustler, K. (1993) Species status and distribution of the parasitic indigo-birds Vidua in east and southern Africa. Proc. VIII Pan-Afr. Ornithol. Congr. 40–52.
Payne, R. B., Payne, L. L. & Woods, J. L. (1998) Song learning in brood parasitic indigobirds Vidua chalybeata: Song mimicry of the host species. Anim. Behav. 55, 1537–1553.
Payne, R. B., Payne, L. L., Woods, J. L. & Sorenson, M. D. (2000) Imprinting and the origin of parasite-host species associations in brood-parasitic indigobirds, Vidua chalybeata. Anim. Behav. 59, 69–81.
Pereira, S. L. & Baker, A. J. (2004) Vicariant speciation of curassows (Aves, Cracidae): A hypothesis based on mitochondrial DNA phylogeny. Auk 121, 682–694.
Podos, J. (2001) Correlated evolution of morphology and vocal signal structure in Darwin’s finches. Nature 409, 185–188.
Presgraves, D. C. (2002) Patterns of postzygotic isolation in Lepidoptera. Evolution (Lawrence, Kans.) 56, 1168–1183.
Price, T. D. (1998) Sexual selection and natural selection in bird speciation. Philos. Trans. R. Soc. London B 353, 251–260.
Price, T. D. (2002) Domesticated birds as a model for the genetics of speciation via sexual selection. Genetica 116, 311–327.
Price, T. D. & Bouvier, M. M. (2002) The evolution of F1 postzygotic incompatibilities in birds. Evolution (Lawrence, Kans.) 56 2083–2089.
Rasner, C. A., Yeh, P., Eggert, L. S., Hunt, K. E., Woodruff, D. S. & Price, T. D. (2004) Genetic and morphological evolution following a founder event in the dark-eyed junco, Junco hyemalis thurberi. Mol. Ecol. 13, 671–681.
Reinhold, K. (1998) Sex linkage among genes controlling sexually selected traits. Behav. Ecol. Sociobiol. 44, 1–7.
Rice, W. R. (1996) Sexually antagonistic male adaptation triggered by experimental arrest of female evolution. Nature 381, 232–234.
Saetre, G. P., Moum, T., Bures, S., Kral, M., Adamjan, M. & Moreno, J. (1997) A sexually selected character displacement in flycatchers reinforces premating isolation. Nature 387, 589–592.
Saetre, G.-P., Borge, T., Lindroos, K., Haavie, J., Sheldon, B. C., Primmer, C. & Syvänen, A.-C. (2003) Sex chromosome evolution and speciation in Ficedula flycatchers. Proc. R. Soc. London Ser. B 270, 53–59.
Schodde, R. & Mason, I. J. (1999) The Directory of Australian Birds (CSIRO, Canberra, Australia).
Seddon, N. (2005) Ecological adaptation and species recognition drives vocal evolution in Neotropical suboscine birds. Evolution (Lawrence, Kans.) 59, 97–112.
Sefc, K. M., Payne, R. B. & Sorenson, M. D. (2005) Genetic continuity of brood parasitic indigobird species. Mol. Ecol., in press.
Servedio, M. R. & Sætre, G. P. (2003) Speciation as a positive feedback loop between postzygotic and prezygotic barriers to gene flow. Proc. R. Soc. London Ser. B 270, 1473–1479.
Sites, J. W. & Crandall, K. A. (1997) Testing species boundaries in biodiversity studies. Conserv. Biol. 11, 1289–1297.
Sites, J. W. & Marshall, J. C. (2003) Delimiting species: A Renaissance issue in systematic biology. Trends Ecol. Evol. 18, 462–470.
Slabbekoorn, H. & Smith, T. B. (2000) Does bill size polymorphism affect courtship song characteristics in the African finch Pyrenestes ostrinus? Biol. J. Linn. Soc. 71, 737–753.
Slabbekoorn, H. & Smith, T. B. (2002a) Bird song, ecology and speciation. Philos. Trans. R. Soc. London Ser. B 357, 493–503.
Slabbekoorn, H. & Smith, T. B. (2002b) Habitat-dependent song divergence in the little greenbul: An analysis of environmental selection pressures on acoustic signals. Evolution (Lawrence, Kans.) 56, 1849–1858.
Smith, C. E. (2003) Ph.D. dissertation (Univ. of Washington, Seattle).
Smith, T. B. (1993) Disruptive selection and the genetic-basis of bill size polymorphism in the African finch Pyrenestes. Nature 363, 618–620.
Sorenson, M. D., Sefc, K. M. & Payne, R. B. (2003) Speciation by host switch in brood parasitic indigobirds. Nature 424, 928–931.
Swanson, W. J. & Vacquier, V. D. (2002) Rapid evolution of reproductive proteins. Nat. Rev. Genet. 3, 137–144.
Swanson, W. J., Clark, A. G., Waldrip-Dail, H. M., Wolfner, M. F. & Aquadro, C. F. (2001a) Evolutionary EST analysis identifies rapidly evolving male reproductive proteins in Drosophila. Proc. Natl. Acad. Sci. USA 98, 7375–7379.
Swanson, W. J., Yang, Z., Wolfner, M. F. & Aquadro, C. F. (2001b) Positive Darwinian selection drives the evolution of several female reproductive proteins in mammals. Proc. Natl. Acad. Sci. USA 98, 2509–2514.
Swanson, W. J., Nielsen, R. & Yang, Q. (2003) Pervasive adaptive evolution in mammalian fertilization proteins. Mol. Biol. Evol. 20, 18–20.
Ting, C. T., Tsaur, S. C. & Wu, C. I. (2000) The phylogeny of closely related species as revealed by the genealogy of a speciation gene, Odysseus . Proc. Natl. Acad. Sci. USA 97, 5313–5316.
Tishkoff, S. A. & Kidd, K. K. (2004) Implications of biogeography of human populations for “race” and medicine. Nat. Genet. 36, S21–S27.
Tubaro, P. L. & Lijtmaer, D. A. (2002) Hybridization patterns and the evolution of reproductive isolation in ducks. Biol. J. Linn. Soc. 77, 193–200.
Uy, J. A. C. & Borgia, G. (2000) Sexual selection drives rapid divergence in bowerbird display traits. Evolution (Lawrence, Kans.) 54, 273–278.
Via, S. (2001) Sympatric speciation in animals: The ugly duckling grows up. Trends Ecol. Evol. 16, 381–390.
Voelker, G. (1999) Dispersal, vicariance, and clocks: Historical biogeography and speciation in a cosmopolitan passerine genus (Anthus : motacillidae). Evolution (Lawrence, Kans.) 53, 1536–1552.
Wakeley, J. & Hey, J. (1997) Estimating ancestral population parameters. Genetics 145, 847–855.
Wang, P. J., McCarrey, J. R., Yang, F. & Page, D. C. (2001) An abundance of X-linked genes expressed in spermatogonia. Nat. Genet. 27, 422–426.
Wang, Z., Hill, G. E., Baker, A. J. & Edwards, S. V. (2003) Reconciling actual and inferred population histories in the house finch (Carpodacus mexicanus) by AFLP analysis. Evolution (Lawrence, Kans.) 57, 2852–2864.
Wiens, J. J. & Penkrot, T. A. (2002) Delimiting species using DNA and morphological variation and discordant species limits in spiny lizards (Sceloporus). Syst. Biol. 51, 69–91.
Wolfner, M. F. (1997) Tokens of love: Functions and regulation of Drosophila male accessory gland products. Insect Biochem. Mol. Biol. 27, 179–192.
Wu, C. I. & Davis, A. W. (1993) Evolution of postmating reproductive isolation: The composite nature of Haldane’s rule and its genetic bases. Am. Nat. 142, 187–212.
Wu, C. I. & Ting, C. T. (2004) Genes and speciation. Nat. Rev. Genet. 5, 114–122.
Yeh, P. J. (2004) Rapid evolution of a sexually selected trait following population establishment in a novel habitat. Evolution (Lawrence, Kans.) 58, 166–174.
Zink, R. M. & McKitrick, M. C. (1995) The debate over species concepts and its implications for ornithology. Auk 112, 701–719.
Zink, R. M., Blackwell-Rago, R. C. & Ronquist, F. (2000) The shifting roles of dispersal and vicariance in biogeography. Proc. R. Soc. London Ser. B 267, 497–503.





























