1
Framing the Issue
In these early years of the 21st century, scientific discovery and understanding are playing an important and growing role in meeting the challenges—environmental, human health, economic—facing societies everywhere. At the forefront are advances in biology. Indeed, it is reasonable to say we are entering the Age of Biology, paralleling in many ways the Age of Physics in the first half of the 20th century.1
For many thousands of years, humans have been manipulating plant and animal stocks—first by accident and later selectively—to meet changing societal and environmental needs. But the discovery of the structure of DNA in 1953, followed by the invention of DNA recombinant technology two decades later, paved the way for the powerful potential to manipulate genes directly and in such a way that the “nature” of an organism can be altered with precision in a single generation. In 2001, scientists finished the initial draft of the human genome sequence, representing a shift in the way biology is studied and opening a portal to vast post-genomic possibilities—from RNA interference (RNAi) therapeutics to DNA nanotechnology. This rapid pace of technological growth in the life sciences research enterprise reflects a revolutionary change in the way people interact with biological systems and a growing capacity to manipulate such systems. Such advancing technologies offer great promise for improving the quality of human life: promoting health, preventing disease, and ensuring adequate food and even the possibility of new energy sources. However, as with all technological advances, there is a potential dark side, the ability for these technologies to be used, either purposefully or negligently, in ways that cause harm to humans. Devising optimal approaches for preventing this has been the overarching aim of this committee.
This chapter provides an overview of recent growth in the life sciences and its associated technologies—with an emphasis on the rapid and shifting nature of this growth. It defines key terms that are used through-
out this report and explores the broad-based nature of the threat posed by the rapid, unpredictable growth, and widespread dissemination of life sciences knowledge and associated technologies. This overview takes into account contemporary understanding of how naturally emerging pathogens cause disease and recently developed technologies that have opened up novel approaches to engineer potentially more harmful agents from both pathogenic and nonpathogenic microbes or viruses. In reviewing this material, the committee developed a heightened awareness of the tremendous potential of the benefits to be derived from the advancement of knowledge and technological growth in the life sciences. At the same time, committee members came to appreciate the magnitude of what hangs in the balance should society fail to address the potential for these technologies to be exploited to cause harm or, by overreacting and imposing unduly restrictive measures on activities in the life sciences, unwittingly muzzle the ability of the life sciences to contribute to future human good.
COMMITTEE CHARGE AND PROCESS
As discussed above and in more detail throughout the report, life sciences knowledge, materials, and technologies are advancing with tremendous speed, making it possible to identify and manipulate features of living systems in ways never before possible. On a daily basis and in laboratories around the world, biomedical researchers are using sophisticated technologies to manipulate microorganisms in an effort to understand how microbes cause disease and to develop better preventative and therapeutic measures against infectious disease. Plant biologists are applying similar tools in their studies of crops and other plants in an effort to improve agricultural yield and explore the potential for the use of plants as inexpensive platforms for vaccine, antibody, and other product manufacturing. Similar efforts are underway with animal husbandry. Scientists and engineers in many fields are relying on continuing advances in the life sciences to identify pharmaceuticals for the treatment of cancer and other chronic diseases, develop environmental remediation technologies, improve biodefense capabilities, and create new materials.
Moreover, other fields not traditionally viewed as biotechnologies—such as materials science, information technology, and nanotechnology—are converging with biotechnology in unforeseen ways and thereby enabling the development of previously unimaginable technological applications. It is undeniable that this new knowledge and these advancing technologies hold enormous potential to improve public health and agriculture, strengthen national economies, and close the development gap between resource-rich and resource-poor countries. However, as with all scientific revolutions, there is a potential dark side to the advancing
power and global spread of these and other technologies. Every major new technology has been used for hostile purposes, and many experts believe it is naive to think that the extraordinary growth in the life sciences and its associated technologies might not be similarly exploited for malevolent purposes.2
This is true despite formal prohibitions against the use of biological weapons and even though, since antiquity, humans have reviled the use of disease for hostile purposes. In its most recent unclassified report on the future global landscape, the National Intelligence Council argued that, although most future (i.e., over the course of the next 15 years) terrorist attacks are expected to involve conventional weapons, a bioterrorist attack will likely occur by 2020.3 Official U.S. statements continue to cite around a dozen countries that are believed to have or to be pursuing biological weapons capabilities.4
The threat of bioterrorism, coupled with the global spread of expertise in biotechnology and biological manufacturing processes, raises concerns about how this advancing technological prowess could enable the creation and production of new biological weapons and agents of biological terrorism possessing unique and dangerous but largely unpredictable characteristics. The Committee on Advances in Technology and the Prevention of Their Application to Next Generation Biowarfare Threats, an ad hoc committee of the National Research Council and the Institute of Medicine, was constituted to examine current trends and future objectives of research in the life sciences, as well as technologies convergent with the life sciences enterprise from other disciplines, such as materials science and nanotechnology, that may enable the development of a new generation of biological threats over the next five to ten years, with the aim of identifying ways to anticipate, identify, and mitigate these dangers.
As part of its study, the committee convened a workshop in September 2004 at the Instituto Nacional de Salud Pública (National Institute of Public Health) in Cuernavaca, Mexico. The purpose of this information gathering workshop was to sample global perspectives on the current advancing technology landscape. Experts from different fields and from around the world presented their diverse outlooks on advancing technologies and forces that drive technological progress; local and regional capacities for life sciences research, development, and application (both beneficial and nefarious); national perceptions and awareness of the risks associated with advancing technologies; and strategic measures that have been taken or could or should be taken to address and manage the potential misapplication of technology(ies) for malevolent purposes. The results of this workshop helped inform the committee as it developed this report.
The committee was charged to:
-
Examine current scientific trends and the likely trajectory of future research activities in public health, life sciences, and biomedical and materials science that contain applications relevant to the development of “next generation” agents of biological origin 5 to 10 years into the future.
-
Evaluate the potential for hostile uses of research advances in genetic engineering and biotechnology that will make biological agents more potent or damaging. Included in this evaluation will be the degree to which the integration of multiple advancing technologies over the next 5 to 10 years could result in a synergistic effect.
-
Identify the current and potential future capabilities that could enable the ability of individuals, organizations, or countries to identify, acquire, master, and independently advance these technologies for both beneficial and hostile purposes.
-
Identify and recommend the knowledge and tools that will be needed by the national security, biomedical science, and public health communities to anticipate, prevent, recognize, mitigate, and respond to the destructive potential associated with advancing technologies.
In interpreting its charge the committee sought to examine current trends and future objectives of research in public health and the life and biomedical sciences that contain applications relevant to the development of new types of biological weapons or agents of bioterrorism, with a focus on five to ten years into the future. It is recognized that the global technology landscape is shifting so dramatically and rapidly that any attempt by the committee to devise a formal risk assessment of the future threat horizon exploiting dual-use technologies by state actors, non-state actors, or individuals could be an exercise in futility. Given that within just the past few years the global scientific community has already witnessed the unexpected emergence of some remarkable new technologies, such as RNA interference and nanobiotechnology, biological threats in the next five to ten years could extend well beyond those that can be predicted today.
Rather than a formal risk assessment, the committee has proposed a conceptual framework for how to think about the nature of the future threat landscape. Indeed, as the world becomes more competent and sophisticated in the biological sciences, it is vitally important that the national security, public health, and biomedical science communities have the necessary knowledge and tools to address the present and future applications of advances in the life sciences.
This report is part of a larger body of work that the National Academies has undertaken in recent years on science and security and the contributions that science and technology could make to countering terror-
ism, beginning with Scientific Communication and National Security in 1982 and continuing with Chemical and Biological Terrorism: Research and Development to Improve Civilian Medical Responses (1999), Firepower in the Lab: Automation in the Fight Against Infectious Diseases and Bioterrorism (2001), Making the Nation Safer: The Role of Science and Technology in Countering Terrorism (2002), Biological Threats and Terrorism: Assessing the Science and Response Capabilities (2002), and Countering Agricultural Terrorism (2002). Most recently, and of particular relevance to this report, is the National Research Council report Biotechnology Research in an Age of Terrorism (2004). The principal difference between that report and the present report is that the former revolves around issues pertaining to research oversight and the flow of scientific knowledge, with a focus on the United States, whereas this report adopts a more global perspective and broadly considers the use and applications of such knowledge.
EMERGING TECHNOLOGIES IN THE LIFE SCIENCES
Heralded by Science magazine as the 2002 “Breakthrough of the Year,”5 RNA interference (RNAi) has emerged as a promising therapeutic approach for the treatment of a wide range of diseases, including cancer.6 Yet just a year before it earned its breakthrough title, RNAi was met with doubt and criticism.7 RNAi therapy involves using small interfering RNA molecules (siRNAs) to cleave and destroy sequence-specific RNA and, in so doing, silence endogenous genes that participate in the pathway of human disease. The technology is expected to prove particularly valuable in cases where the targeted RNA encodes genes and protein products with activities that cannot be modulated today by conventional drugs. Several recent experiments indicate that investigators are well on their way to overcoming the clinical challenges of delivering effective RNAi therapy.8 In October 2004, Acuity Pharmaceuticals (Philadelphia, PA) announced that it was beginning a Phase I clinical trial of an investigational drug known as Cand5, making Cand5 the first RNAi therapeutic to enter clinical trial. Cand5 is an siRNA that turns off the expression of proteins contributing to vision loss in patients with age-related macular degeneration.
In addition to its therapeutic applications, RNAi has emerged as a key basic research tool for use in functional genomics; by blocking the expression of a particular gene, one can create a phenotype that yields clues about the function of that gene. RNAi technology is forecast to grow at an annual average rate of just over 30 percent between 2003 and 2010.9 Although European and U.S.-based companies currently dominate the market (i.e., there are about 50 U.S. and European companies active in the RNAi market, most of their revenues coming from RNAi reagents and
research tools),10 this may change over the course of the next several years as Asian companies begin specializing in RNAi applications.
Touted alongside RNAi in Massachusetts Institute of Technology’s (MIT) 2004 Technology Review as one of the top 10 emerging technologies that “will change your world,” synthetic biology is the assemblage of gene networks—or circuits (i.e., analogous to silicon circuits)—that can guide the construction of novel, synthetic proteins and direct cells to perform assigned tasks.11 By assembling genes into circuits that direct cells to perform assigned tasks, synthetic biologists have taken genetic engineering to a level so profoundly different from recombinant technology that, in an October 2004 Nature news article, the latter was referred to as “old hat.”12 DNA synthesis applications are now largely limited to places like the MIT’s Independent Activities Period (IAP) course, where students design DNA circuitry, send their designs via the Internet to Blue Heron Biotechnology, Inc. (Bothell, WA), and then introduce the resulting synthetic DNA molecules into E. coli strains.13 Because it is in its early growth phase, the future industrial potential of synthetic biology is unclear.14 Meanwhile, research scientists are using the technology to design unique genomes and test novel hypotheses and models.
In just five years, nanotechnology has catapulted from being a specialty of a handful of physicists and chemists to a worldwide scientific and industrial enterprise.15 The U.S. government estimates that the nanotech economy will be worth $1 trillion by 2012, and the White House recently requested $1 billion for fiscal 2006 to develop nanotechnology (up from $442 million in 2001). In April 2005, the National Academies Keck Futures Initiative announced that it had awarded a total of $1 million to 14 interdisciplinary research projects in nanoscience and nanotechnology. The awards, which are considered seed money to allow recipients to develop research approaches and position themselves competitively for other project funding, will be used for a variety of projects ranging from an examination of the interactions of nanoparticles with biosystems to the development of a new approach for capturing solar energy.
Nanoparticles are already being used in a variety of commercial products, like sunscreen, paint, inkjet paper, stain-resistant trousers, and highly durable engine parts.16 Some industry analysts predict that by lowering drug toxicity and the cost of treatment (among other benefits), nanotechnology-enabled drug delivery systems will probably be among the first biomedical markets to evolve and to provide significant business revenue opportunities.17 For example, Elan Corporation (Dublin, Ireland) has developed a proprietary technology known as NanoCrystal, which transforms poorly water-soluble drugs into nanometer-sized particles that can be used to create any of a variety of more soluble common dosage forms for both parenteral and oral administration. There are sev-
eral NanoCrystal-based therapeutics already on the market or in development.18
Nanobiotechnology—also known as DNA nanotechnology—refers to the convergence of nanotechnology with molecular biology.19 In fact, most of the examples in the preceding paragraph fall within its domain. Nanobiotechnology and nanobiotech start-up companies constitute nearly 50 percent of the venture capital invested in nanotechnology.20 Scientists are increasingly reporting discoveries with implications for potential applications of nanobiotechnology. For example, in January 2005, in a paper published in Physical Review Letters, researchers from the University of California, Los Angeles, described a nanoscale mechanism for externally controlling protein function, a technological advance that could ultimately lead to a generation of targeted “smart” drugs that are active only when certain DNA is present or a certain gene is expressed.21 In February 2005, in a paper published in the Proceedings of the National Academy of Sciences, Northwestern University researchers described a nanoparticle-based assay for detecting the onset of Alzheimer’s disease.22 Also in February 2005, an Illinois-based company, Nanosphere, Inc., announced plans to expand and market the application of the same assay to a variety of other diseases, including cancer.23
While new tools, like RNAi therapeutics and nano-based drug delivery are emerging, already proven tools such as the polymerase chain reaction (PCR) and DNA sequencing, are becoming more versatile, more affordable, and faster. For example, real-time, or quantitative PCR (qPCR), which is arguably one of the fastest growing PCR technologies, allows users to quantitatively monitor the amplification process as copies of DNA accumulate (unlike “traditional” PCR, which provides only an end product, a “yes/no” answer, and a qualitative measure of the abundance of the target material).24 In 2004, the least expensive qPCR thermocycler on the market was listed in the mid-$20,000 range. In spring 2005, Bio-Rad Laboratories (Hercules, CA) launched a “personal” qPCR machine that sells for about $16,500 and is one of the smallest machines on the market (i.e., in terms of size and the number of samples it can accommodate).
Moreover, it should not be forgotten that PCR itself was not widely anticipated before its arrival on the scene.25 And it is instructive to remember how it developed, first as a relatively straightforward concept in which DNA synthesis was recycled through a series of cyclic thermal manipulations.26 This resulted in a doubling of the product each thermal cycle with an exponential amplification of the product over many thermal cycles of annealing, extension, and denaturation, with the DNA polymerase enzyme being destroyed during the denaturation step. However, it was not until a thermally-resistant DNA polymerase was isolated from nature that the process became widely available and widely utilized.
Now PCR is as indispensable a “tool” for many 21st-century biologists as a microscope was to a 19th-century microbe hunter. Its impact on accelerating the velocity of life sciences research is readily appreciated by anyone in the field, as most biotechnologists today would have difficulty accomplishing their aims without this technique. Its importance overall to the life sciences is reflected in the relatively unusual actions of the Norwegian Nobel Committee, conferring its award on the inventor of PCR, Kary Mullis, only a few years after the technique was first reported. Parallels to the thinking that went into PCR are seen today in an unrelated field—the investigation of spongiform encephalopathies, like “mad cow disease,” where an analogous cycling technique has been reported recently for in vitro amplification of prions, putative infectious agents that lack genes (i.e., DNA or RNA) and that consist of a protein with “infectious” capacity to initiate misfolding of similar proteins.27 This series of events in the development of PCR recapitulates a theme in the life sciences: the sudden arrival of a new technique, followed by its technological exploitation, further refinement, and subsequent extension to other related fields. Similar scenarios have accompanied the discovery of restriction endonucleases and the development of recombinant DNA, and are unfolding now with RNAi technology or recently described multiplex DNA synthesis capabilities.
The speed of DNA sequencing, DNA synthesis, and protein structural analysis—each a different measure of biotechnological power—has increased practically exponentially over the past 15 years.28 Indeed, progress in the life sciences, rather than being “linear,” is often marked by periodic and unpredictable major breakthroughs in our understanding of the living world that consequently radically transforms the growth and development of advances in disparate disciplines.29 At present, the 10 plant and animal genomes and the approximately 100 microbial genomes that are sequenced every year are done so, largely, at a small number of factory-like DNA sequencing centers. It has been estimated that if technological developments continue to improve the efficiency of DNA sequencing as they have up to this point, by 2010 a single lab worker will be able to sequence (or synthesize) about 1010 bases in one day (there are 3 × 109 bases in the human genome).30
The future of DNA synthesis is likely to follow a similarly rapid trajectory, with scientists being able to synthesize complete microbial genomes by 2010 if not sooner.31 In December 2004 Harvard University’s George Church and colleagues published an article in Nature describing a new microchip-based technology for the multiplex synthesis of long oligonucleotides.32 The researchers used the new technology to synthesize all 21 genes that encode proteins of the E. coli 30S ribosomal subunit. This technological advance is coupled with falling prices. In 2000, sequence
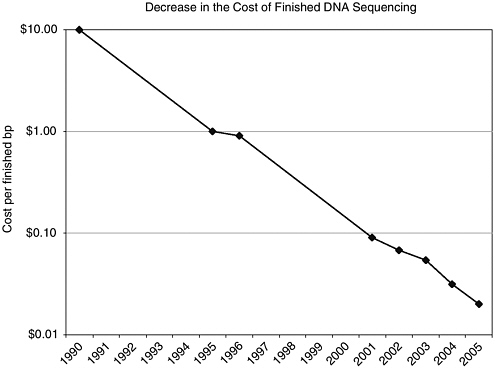
FIGURE 1-1 The plunging cost of DNA sequencing has opened new applications in science and medicine.
SOURCE: Reprinted with permission. Service, RF. 2006. Gene sequencing: The race for the $1,000 genome. Science 311(5767):1544-1546. Available online at www.sciencemag.org/cgi/content/full/311/5767/1544.
assembly cost about $10 to $12 per base pair. By the beginning of 2005, the cost had dropped to about $2 per base pair (e.g., Blue Heron offers a special price of $1.60 for new customers33), and it is expected to fall to 1 cent per base pair within the next couple of years34 (see Figure 1-1).
This has had real and practical consequences. For example, when the first successful autonomously replicating RNA replicons for hepatitis C virus were described by the Bartenschlager laboratory in 1999,35 several other groups immediately synthesized the entire ~7,000 nucleotide-long complementary DNA sequence of this RNA so as to be able to access this technology. De novo chemical synthesis was judged to be a more rapid, or less expensive means to acquire the technology than working through Materials Transfer Agreements, etc., with those who first described the replicons. The DNA synthetic “muscle” for this was readily available on a
contract basis, even five years ago. Such an exercise would be trivial today, however, given recent advances in DNA synthetic capacities.36
Similar predictions about feasibility, rapidity, and affordability can be made for the structural analysis of proteins and other biologically important molecules. It is not unreasonable to expect that, before long, scientists will develop and have access to computer programs that simulate in detail the molecular processes in cells, so that the interaction of cells with pathogenic microbes and molecules can be fully anticipated and understood.
Notable Features of Technological Growth in the Life Sciences
Technological growth in the life sciences is characterized by several notable features. These are critically important to recognize if a reasonable estimate is to be made of what is or is not possible in predicting its future.
First, as described above, progress in biology has been marked repeatedly by successive serendipitous discoveries and applications that over time have lead to the widespread adoption of new technologies with independent scientific and economic impacts. Indeed, the rapid growth of bio- and other relevant technologies over the past 30 years has been driven by two processes working together: a quantitative increase in performance coupled with a decrease in the cost of existing technologies (such as template independent DNA synthesis) and instruments, as explained in the previous section, and sudden and occasionally dramatic qualitative changes (paradigm shifts) resulting from unanticipated new inventions, unexpected discoveries, and insights, all of which may be significantly enhanced by the occurrence of unforeseen, historically significant events that impact significantly on human society and its everyday concerns. In addition to recombinant DNA technology (which sparked the biotech revolution back in the 1970s), prominent new inventions and discoveries in recent history include PCR (i.e., which originated in the mid-1980s as described above), the transfer of nuclei from cell to cell (i.e., cloning, also known as somatic cell nuclear transfer, or SCNT), the advent of RNAi technology (as described above), and the introduction of new techniques for parallel DNA synthesis capable of greatly accelerating the rate at which genes can be created de novo. New inventions and discoveries like these are a precondition for the rapid growth of technology. They result in the capacity to reduce the development costs associated with new and potentially very useful products, such as the recombinant hepatitis B vaccine, one of the early “fruits” of the recombinant DNA era, or to genetically engineer crops with intrinsic resistance to pests.
Equally important, however, are both public and political support for
these efforts. Such support can, in turn, drive the availability of government or venture capital funding required to fuel the advancement of research and development activities in the life sciences. Current levels of government support in the life sciences can be attributed in part to unforeseen historical events, such as the political decision to declare a “war on cancer” in the 1970s, the occurrence of the HIV/AIDS pandemic in the 1980s, and the 2001 anthrax mailings, which, in part, contributed to the current “war on terror.” On the other hand, the public perception of risks can readily derail the expansion of this technology, as evidenced by the impact of the “green” movement in Europe on the acceptance of genetically engineered crops by the public.
This constantly changing and rapidly growing global technological landscape, marked as it is by the seemingly stochastic arrival of new paradigm-shifting concepts, makes it extremely difficult, if not impossible, to predict specific future trends. Just a year before it earned its “Breakthrough of the Year” title by Science magazine,37 RNAi was met with doubt and criticism. Self-assembling nano-devices, such as the DNAzyme (a device that can bind and cleave RNA molecules one by one) developed in 2004 by Purdue University researcher Chengde Mao, were unimaginable just a couple of years ago.38 About the only thing one can predict is that the life sciences will continue to advance quickly, in a variety of directions, and that new and previously unanticipated paradigm shifts are very likely to occur in the future.
Second, as difficult as it is to predict what kind of technological or scientific breakthroughs might occur next, it is practically impossible to know where in the world these breakthroughs might happen. As discussed in greater detail in Chapter 2 of this report and in an earlier workshop summary report from this committee, a number of countries around the world are investing heavily in life sciences technologies.39 Indeed, several countries that are not commonly viewed as being technologically sophisticated, or that have not been considered technologically savvy in the past, are making remarkable progress in biotechnology and are well-positioned to become regional or global leaders in the near future. Importantly, the rapid global dispersion of life sciences materials, knowledge, and technologies is not limited to technologies with proven therapeutic and market value. While India is currently strong in generic and bulk biopharmaceutical manufacturing, several factors, including its growing technological expertise and its 2005 accession to the World Trade Organization, are contributing to its greater capacity for innovation and research and development of novel products. South Korea is rapidly gaining global prominence for its breakthrough contributions to stem cell research, although some of these “breakthroughs” are now in dispute.40 Meanwhile, Singapore has identified biotechnology as a central pillar of its fu-
ture economy. Biotechnology is no longer the restricted playing field of a few privileged nations, but is truly a global enterprise.41
Third, the number of known biologically active molecules, and potential genetically engineered organisms, that could cause harm to humans through inadvertent use, inappropriate use, or as a result of purely malevolent intent such as in the development of a weaponizable biological or chemical agent, is increasing rapidly. This stands in sharp contrast to the still relatively small number of nuclear materials that could potentially be used for malign intent. This is evident in the increasing pace of research activity in the life sciences, as reflected in the number of biotech drug approvals (i.e., as opposed to large pharma drug approvals), which grew from fewer than 5 in 1982 (and none in 1983) to more than 30 in 2000.42 According to the Biotechnology Industry Organization (BIO), there are approximately 370 biotech drug products and vaccines currently in clinical trials targeting more than 200 diseases.43 This growing number of potential and approved drugs is due in part to a fundamental shift in the drug discovery process.
New technologies—genomics, microarrays, proteomics, structural biology, combinatorial chemistry, toxicogenomics, and database mining—allow drug developers to identify likely molecular targets early in the discovery process and then screen large numbers of compounds that bind to and affect the targets. Moreover, purely “in silico” screening approaches are becoming more common. There has thus been a shift in drug discovery methodology from pure empiricism to more rationally based drug design. In addition, new methods for synthesizing chemical libraries have led to the aggregate generation of several hundred million new potential ligands, while the same discovery process has identified thousands of potentially toxic compounds each year.44 Until now, most of the databases produced by these efforts have been proprietary and jealously guarded by the companies that generated them.45 However, the Chemical Genomics Center network, recently established by the National Institutes of Health (NIH), will make this type of information much more accessible (see Box 1-1). Without any knowledge of the underlying biological mechanism, it will be possible to mine this vast chemical database to unearth structural relationships between desirable targets and the chemical compounds known to interact with them. While originally conceived as leading ultimately to a “roadmap” of the functions of the myriad proteins expressed by the human genome, it is also possible that this novel program could become a “roadmap” to new generations of very efficient poisons. Thus, as will become evident throughout this report, almost any effort to advance knowledge in the life sciences, such as the NIH Roadmap (Box 1-1), brings with it the potential for malevolent use as well as beneficial impact.
Finally, one can imagine a future where, as biotechnology continues to change radically, rather than becoming big and centralized, the life sciences and related applications may become increasingly domesticated and accessible. An example of this is the recent pet store appearance of genetically modified tropical fish with new and brilliant colors (see Box 1-2). The fish were developed as a commercial product by the Taikong Corporation of Taiwan; they first appeared on the commercial market in Taiwan, Japan, Hong Kong, and Malaysia in 2003. More importantly, just as computer technology was transformed over the course of a few decades to a point where computers were small enough and cheap enough to be used in homes (e.g., to prepare income tax returns or homework) and then to a point where computer games and toys became a dominant feature of children’s lives, biotechnology may similarly be transformed.46
Definitions
The life sciences are defined broadly in this report to include any field of science that is leading to or has the potential to lead to an enhanced understanding of living organisms, especially human life. These sciences include, for example, branches of mathematics and computational science, as these are now being applied in efforts to effectively model a wide variety of biological systems, or materials science, as it is applied to the manipulation of biological systems. Here “associated technology” refers to the development and application of tools, machines, materials, and processes based on knowledge derived within or applied to the life sciences: genetic engineering, synthetic biology, aerosol technology, combinatorial chemistry, and nanotechnology are just a few of these technologies. The future problem in some respects resembles the current threat; for every biological threat agent there must be a means for delivery to the target (aerosols are usually thought to be the most efficient mode for agents that are not contagious). The efficient delivery of medicines by aerosol (insulin) or nanotechnology is intensely relevant “enablig technologies” to future problems, even if they are not inherently “biology.” In addition, the future holds threats that are so different from the classical biothreats that they fall into entirely new categories and will require highly innovative ways to detect and counter them. The aim of this report, therefore, is to be as inclusive as possible in looking at technologies—including those not traditionally viewed as biotechnologies—but that may lead to the creation and production of new biological weapons and biological warfare threats.
The terms “weapon” and “bioweapon” are also used broadly, and include any biological agent or biologically active molecule or other entity47 that is used or developed and/or stockpiled for use in an effort to cause harm to humans, plants, or animals. In some cases a bioweapon
|
BOX 1-1 Even seemingly “benign” or solely beneficial activities such as those envisioned by the achievements of the National Institutes of Health (NIH) Roadmap could serve as a new source of potential dual-use information—that is, information that can be used inappropriately or for malicious purposes as well as for the beneficial intent for which it is designed. The NIH Roadmap is designed to identify major opportunities and gaps in biomedical research that no single NIH institute could tackle alone but that, by relying on an interdisciplinary research approach, would make the biggest impact on medical research progress in the coming century. It is highlighted here simply to point out that the potential dual-use nature of the information to be generated may not be fully appreciated. The challenge is to devise a strategy for allowing this necessary and beneficial research to move forward while preventing inappropriate or malicious use. There are three NIH Roadmap “themes.” The first, “New Pathways to Discovery,” among other goals, addresses the need to know more precisely the combination of molecular events that lead to disease and involves establishing a library of chemical molecules for use in identifying potential targets for new therapies and other purposes. The second, “Research Teams of the Future,” involves stimulating new ways of combining skills and disciplines in the physical and biological sciences, for example, by encouraging public-private partnerships and encouraging investigators to conduct research that is high-risk but also high-payoff. The third, “Re-engineering the Clinical Research Enterprise,” addresses the need for new partnerships and networks between and among the scientific and clinical communities in order to better translate research discoveries into drugs, treatments, and preventative methods. Each of these themes comprises several or more groups and initiatives, the details of which can be viewed on the NIH Roadmap Web site: |
may be purposefully modified to enhance its ability to be delivered or to otherwise cause harm. However, it is not necessary for a biological agent to be specifically “weaponized” for it to be used as a weapon, as, for example, a routine culture of a bacterial pathogen might simply be added to food or drinking water.
The term “dual-use” refers to the capacity or potential for biological agents, information, materials and supplies, or technologies to be used for either harmful or peaceful purposes. This definition needs to be distinguished from a common use of the term within defense circles, wherein “dual-use” implies a potential military application for a civilian technology.
|
nihroadmap.nih.gov/overview.asp. One example of an initiative that may eventually lead to potent new dual-use information is the Chemical Genomics Center, established in June 2004 by the Molecular Libraries and Imaging Implementation Group, as part of the New Pathways to Discovery theme. This center will be part of a consortium of chemical genomics screening centers to be located across the country whose purpose will be to identify small molecule inhibitors of every important human cellular protein or signaling pathway. Part of the rationale for the chemical genomics initiative(s) is that, in contrast to researchers in the pharmaceutical industry, many academic and government scientists do not have easy access to large libraries of small molecules (i.e., organic chemical compounds that are smaller than proteins and that can be used as tools to modulate gene function). The database will give academic and government researchers an opportunity to identify useful biological targets and thereby contribute more vigorously to the early stages of drug development. With plans to screen more than 100,000 small-molecule compounds within its first year of operation, one of the goals of the Chemical Genomics Center network is to explore the areas of the human genome for which small molecule chemical probes have yet to be identified. Data generated by the network will be deposited in a comprehensive database of chemical structures (and their biological activities). The database, known as PubChem, will be freely available to the entire scientific community. In addition to screening and probe data, it will list compound information from the scientific literature. Should this come to pass, it will offer enormous opportunities for industry and academic scientists alike to pursue novel “drugable” targets in a search for small molecule inhibitors of certain pathways that could offer substantial clinical benefit. However, the availability of information and reagents that enable one to disrupt critical human physiological systems has profound implications for the nature of the future biological and chemical threat spectrum. |
As used in this report, the terms “inappropriate use” and “malicious use” or “malevolent use” capture a continuum of potentially dangerous activities that are largely distinguished on the basis of intent. Thus, (1) the use of technology without the intent to cause harm but with unanticipated dual-use consequences, including experiments or other activities conducted with inadequate oversight or without an awareness of the consequences of certain outcomes would be considered inappropriate use, while, (2) the deliberate use of technology for the creation, development, production, or deployment of biological weapons is considered malevolent or malicious use, with malevolent indicating the intent to cause death or
|
BOX 1-2 The GloFish™, a fluorescent red zebrafish sold as a novel pet, has become the first transgenic animal sold to U.S. consumers. Its sale has produced regulatory controversies, a lawsuit, and profits for its proponent, Yorktown Technologies (Austin, TX). With the market plan calling for sales in a widening number of countries, continuing controversy seems likely. What Is a GloFish™? The GloFish™ is a trademarked transgenic zebrafish (Danio rerio) expressing a red fluorescent protein from a sea anemone under the transcriptional control of the promoter from the myosin light peptide 2 gene of zebrafish.a Produced and patented by a group at the National University of Singapore,b exclusive rights for international marketing were purchased by Yorktown Technologies in 2002. Yorktown produces GloFish™ through contracts with 5-D Tropical (Plant City, FL) and Segrest Farms (Gibsonton, FL) and began marketing the fish in the United States in December 2003. Issues Posed The prospect of commercial sales of GloFish™ has raised a number of issues. Among them was the issue of whether GloFish™ pose an environmental hazard. Zebrafish, a tropical species native to south Asia, are sensitive to low temperatures. Despite decades of production and use in the United States, zebrafish have not established self-sustaining populations in this country. Laboratory tests showed that viability, reproductive success, and temperature tolerance of transgenics were equal to or somewhat less than those of the wild type.c While preliminary, results supported the expectation that the modification would not increase invasiveness and that environmental risk was small. Commercialization of the GloFish™ in the United States poses regulatory uncertainty because existing biotechnology policy bases oversight on use of the product. Sales of ornamental fishes are not federally regulated. The Food and Drug Administration asserts jurisdiction over genetically modified animals using the New Animal Drug Application process.d After a brief internal review and interagency consultation, the FDA’s Center for Veterinary Medicine determined that “because tropical aquarium fish are not used for food purposes, they pose no threat to the food supply. There is no evidence that these genetically engineered zebra fish pose any more threat to the environment than their unmodified counterparts which have |
|
long been widely sold in the United States. In the absence of any clear risk to the public health, the FDA finds no reason to regulate these particular fish.”e Alan Blake, CEO of Yorktown Technologies, also made contact with the U.S. Department of Agriculture, the U.S. Fish and Wildlife Service, and the Environmental Protection Agency, which expressed no regulatory concerns regarding GloFish.™ Future Prospects Future prospects for the GloFish™ include marketing additional color lines in a wider range of markets. Not only red, but also green and yellow fluorescent proteins have been introduced into stable transgenic lines, yielding green, yellow, and orange fish. Commercialization of fluorescent zebrafish has gone forward in several countries and is stymied in others. Fluorescent green zebrafish developed in Taiwan have been sold in Taiwan, Malaysia, and Hong Kong. Singapore confiscated attempted imports of the fish. Despite this, Yorktown Technologies is considering other markets, including parts of Asia and Latin America. Extensive information requirements suggest that GloFish™ will not be marketed in Canada or the European Union in the near future. Despite these regulatory challenges, according to Blake, “The GloFish™ venture is a profitable one, and the company looks forward to continuing to provide a safe and enjoyable product for many years to come.”f
|
serious injury, and malicious a lesser degree of intended damage. These latter terms do not include the deliberate use of technology to create potentially harmful materials or other disease-causing agents for defensive research purposes in the absence of any intent to cause harm (e.g., the equivalent of computer hacking48). The committee recognizes, however, the controversial and troubling aspects of such approaches, and their potential to add to, as well as potentially help mitigate, the threat of bioterrorism or biological warfare.
The term “bioterrorist” refers to individuals or groups, usually nonstate actors, that develop and/or use biological agents with the intent to cause harm. On the other hand, “biological warfare” refers to the intentional use of such weapons by state actors, regardless of whether they are deployed against civilian or military targets, or on either a large or small scale.
Finally, the term “biosecurity” is used to refer to security against the inadvertent, inappropriate, or intentional malicious or malevolent use of potentially dangerous biological agents or biotechnology, including the development, production, stockpiling, or use of biological weapons as well as natural outbreaks of newly emergent and epidemic diseases. Although it is not used as often as it is in other settings, to refer to a situation where adequate food and basic health are assured,49 there may be significant overlap in measures that guarantee “biosecurity” in either sense.
20TH CENTURY GERM-BASED BIOWARFARE
History has demonstrated that research in biology, even when conducted without any military application in mind, may still contribute to the production of biological weapons. Indeed, people figured out how to intentionally spread illnesses long before naturalists discovered that germs cause disease.50 But it was only after the discovery of the germ theory of disease in the late 19th century that infectious diseases were seriously considered, on a continual basis, as tools of war. Biologists were able for the first time to identify, isolate, and culture disease-causing microbes under controlled conditions and use them to intentionally induce disease in a “naïve” host.
For example, one of the first attempts to use anthrax during warfare was in World War I, when the Germans reportedly attempted to ship horses and cattle inoculated with B. anthracis (as well as Burkholderia mallei, the bacterium that causes glanders in livestock) from U.S. ports to Allies.51 In 1917, German spies were caught allegedly trying to spread B. anthracis among reindeer herds in northern Norway, near the Russian border.52 These charges were confirmed when anthrax-laced sugar cubes, obtained from a Swedish-German-Finnish aristocrat arrested as a German
agent in 1917, were recently found to still be viable after being stored in the archives of a Norwegian museum for the past 80 years.53
During World War II, every major combatant had a biological weapons program in place (including the United Kingdom, the United States, Canada, France, the Soviet Union, Germany, and Japan).54 The U.S. offensive biological weapons program originated in 1942, at Camp Detrick, in Frederick, Maryland. Its focus during WWII was on biological warfare research on the causative agents of anthrax, botulism, and many other human, animal, and plant pathogens.55 The Special Projects Division of the Army Chemical Warfare Service, which was primarily responsible for carrying out the program, had at its peak approximately 3,900 personnel, including about 2,800 Army and 1,000 Navy personnel and 100 civilians. Although Camp Detrick remained the parent research and pilot plant center, field-testing facilities were established in 1943 and 1944 in Mississippi and Utah, respectively, and production plants were constructed in Indiana and Pine Bluff, Arkansas in 1944. After WWII, until the program was dismantled in 1969, it developed and perfected offensive weapons capabilities for the Department of the Army and certain weapons capabilities for the Air Force, Navy, and the Central Intelligence Agency, using a variety of human, animal, and plant pathogens. All work was conducted under the strictest secrecy.56
Japan’s secret biological warfare program, Imperial Unit 731 (hereinafter Unit 731), which was officially known as the Army Anti-Epidemic Prevention and Water Supply Unit, studied, cultured, and developed a large number of biological agents, including B. anthracis and C. perfringens, which reportedly were used on prisoners of war.57 There were at least four operational units of Japan’s secret biological warfare complex: Unit 731, located in Ping Fan; Unit 100 in Changchun; Unit 9420 in Singapore; and Unit Ei 1644 in Nanking. There is also some evidence that the Japanese had an “epidemic prevention center”—a euphemism for biological weapons research on tropical diseases—in Rangoon, Burma. Each unit had 10 to 15 individual facilities located within and outside mainland China.58 During the Sino-Japanese War (1937-1945), Japan repeatedly attacked China with the plague-causing bacterium Yersinia pestis, reportedly targeting over eleven cities. At least 700 Chinese reportedly died from plague alone, although the actual morbidity and mortality associated with Unit 731’s germ warfare “experiments” against Chinese nationals and others is likely to be several orders of magnitude higher.59
Even after the Biological and Toxin Weapons Convention (BWC) was opened for signature in 1972, the Soviet Union retained and expanded an extensive secret biological weapons program that involved tens of thousands of workers—the largest biological weapons complex ever created. On April 3, 1979, an accidental release of anthrax was believed to have
occurred at the Soviet Institute of Microbiology and Virology in Sverdlovsk. A reported 67 people died from inhalation anthrax, and another 33, perhaps more, were reported to have been infected with B. anthracis, the causative agent of the disease.60 For years the Soviet government officially maintained the cover story that the outbreak was gastrointestinal anthrax and was due to ingestion of contaminated beef, denying that the incident had anything to do with an accidental release of anthrax from an upwind military research facility. In 1992, the Russian press reported that President Boris Yeltsin officially acknowledged that the 1979 incident had in fact been an accidental airborne release of anthrax spores from a military research facility, although many Russian scientists continued to steadfastly deny the occurrence of such an accident. The incident reinforced U.S. suspicions that the Soviets had a biological weapons program, despite having signed the BWC in 1972 and upon ratification, making the statement: “The Soviet Union does not possess any bacteriological agents and toxins, weapons, equipment, or means of delivery.” The quantity of spores released at Sverdlovsk was recently estimated at less than one gram, but the basis for this estimate is speculative.61
The nature and extent of the former Soviet Union’s biological weapons program became known to western governments after Vlademir Pasechnik and then Ken Alibek, chief and deputy chief (respectively) of Biopreparat, defected to the United Kingdom62 and the United States in the late 1980s and early 1990s.63 Biopreparat, an ostensibly state-owned pharmaceutical organization was, in reality, carrying out a secret offensive and defensive biological weapons program that operated from 1972 until at least 1992.64 It was the most sophisticated biological weapons program in the world, and its size and scope were enormous. By the early 1990s, more than 60,000 people were involved in the research, development, and production of biological agents for use in weapons, and the complex had the capability to stockpile hundreds of tons of material containing anthrax spores and dozens of tons of material containing other pathogens, including smallpox and plague agents.65 Many state programs were involved in various aspects of this effort. The Ministry of Defense and its research facilities, of course, played a central role in setting requirements for the program and in program implementation. Components of the Ministry of Health and the Ministry of Agriculture and selected institutes of the Soviet Academy of Sciences were also involved. The KGB developed the capability to deliver biological weapons through clandestine systems.66 These activities were carried out despite Soviet assurance set forth in international agreements not to develop a biological weapons capability and open declarations to the United Nations that it was not developing such capacity.
South Africa’s clandestine program, Project Coast, a chemical and bio-
logical weapons program that existed from the 1980s until 1993-1994, when the government announced that it was dismantling all weapons of mass destruction programs, serves as another example of a recent state-level, clandestine bioweapons program. The extent of Project Coast was not publicly known until 1998-1999, when the Truth and Reconciliation Commission offered immunity to many scientists in exchange for disclosure of their involvement with the project.67 The now-transparent history serves as a dramatic example of how science can be subverted to undermine entire communities and how scientists can be persuaded to participate in a clandestine state-level biological weapons program. At the time of the project, research conducted in the national interest was considered the most important research in the country.68 As recommended by the international community, the South African government has attempted to keep many experts in this area employed under its watch rather than have them take their expertise elsewhere.69
Beating Nature: Is It Possible to Engineer a “Better” Pathogen?
The rapid, unpredictable, and widespread growth of the life sciences and biotechnology has raised concerns that, while such growth benefits national development and enriches the quality of life for millions of people worldwide, it also creates new opportunities for inappropriate or malicious use. The question then becomes, what type of biological agent, or bioweapon, poses the greater threat, and do human-engineered bioweapons pose a greater or lesser threat than naturally emerging infectious disease agents?
Natural Threats
It has been argued by some that nature serves as the most potent reservoir of biological threats to humans, animals, and plants and as a source of biothreat diversity. This issue deserves further discussion here, given the relevance of the counter-arguments (synthetic or engineered agents may be as potent or more potent—at least in the short-term—than “natural pathogens”) that are based on the potential impact of advancing technologies. This argument posits that deliberate efforts to create novel biological threat agents will not succeed in constructing agents more (or even, as) potentially harmful than those that have or can arise through natural means, because of the broad spectrum of natural mechanisms that give rise to biological diversity and the competitive and selective pressures brought to bear on these natural agents. In considering this argument, however, it is important to consider the principles underlying “pathogenicity” and to recognize that the capacity to injure humans does not, by
TABLE 1-1 Cases and Deaths of Emerging Infectious Diseases in the Past and Present
|
Historic Pandemicsa |
Causative Agent |
Cases |
Deaths |
|
Justinian Plague, 6th Century (First recorded outbreak of bubonic plague) |
bacterium Yersinia pestis |
142 million (based on an estimated 70% mortality rate) |
~100 million |
|
The “Black Death” |
bacterium Yersinia pestis |
|
25 million |
|
China Plague (or “Third Pandemic”), 1896-1930 |
bacterium Yersinia pestis |
30 million |
12 million |
|
Spanish Flu 1918-1919 |
influenza A virus |
200 million |
50 million - 100 million |
|
Current Pandemicsb |
Causative Agent |
Cases |
Deaths |
|
Malaria |
Plasmodium parasites |
300 million to 500 million per year |
1.5 million to 2 million per year |
|
Tuberculosis |
Mycobacterium tuberculosis |
8 million to 10 million per year |
2 million per year |
|
Hepatitis C |
Hepatitis C virus (HCV) |
~170 million (cumulative) |
10,000 per year (U.S. only) |
|
HIV/AIDS |
Human Immunodeficiency Virus-1 (HIV-1) |
more than 60 million (cumulative) |
more than 20 million (cumulative) |
itself, provide any virus, bacterium, or other infectious agent with a selective survival advantage. Rather, injury or “disease” occurs as an incidental effect of mechanisms evolved by the infectious agent to promote its multiplication and long-term survival. To illustrate the devastation that natural biological agents can cause, Table 1-1 provides a snapshot of cases and deaths of emerging infectious diseases in the past and present. In addition to those listed, many other infectious diseases have emerged, reemerged, or developed drug resistance over the past couple of decades and across the globe; every hour an estimated 1,500 people die from an infectious disease.70
Influenza virus is considered by many to be the greatest natural infec-
|
Recent and Current Outbreaksc |
Causative Agent |
Cases |
Deaths |
|
Marburg hemorrhagic fever (in Angola; as of May 17, 2005) |
Marburg virus |
337 |
311 |
|
Avian influenza (in Asia and Eurasia) beginning January 2003; as of April 18, 2006) |
H5N1 Influenza A virus |
194 |
109 |
|
Meningococcal disease (in Burkina Faso, from January 1 to April 20, 2003) |
N. meningitidis |
7,146 |
1,058 |
|
Severe Acute Respiratory Syndrome (Worldwide, from November 1, 2002, to July 31, 2003) |
SARS-associated coronavirus (SARS-CoV) |
8,096 |
774 |
|
aInformation on historic pandemics is adapted from “Killer Diseases through Time,” The Scientist 17(11), 2003:16; updated information from Institute of Medicine. 2005. The Threat of Pandemic Influenza: Are We Ready? Washington, DC: The National Academies Press; Osterholm, M. 2005. Preparing for the Next Pandemic. Foreign Affairs, 84, (4): 24-37; McNeill, W. 1998. Plagues and Peoples. Anchor Book published by Doubleday Press: New York. bInstitute of Medicine. 2003. Microbial Threats to Health: Emergence, Detection, and Response. Washington, DC: The National Academies Press. cInformation on recent outbreaks is from the World Health Organization. |
|||
tious disease threat faced by the world today. However, it is but one example of a potentially devastating natural threat. The magnitude of the threat posed by influenza reflects several different features of the virus: its ability to be readily transmitted among humans, to cause significant tissue injury, and to circumvent preexisting immunity within a population by the rapid acquisition of novel surface antigens that are not recognized by antibodies elicited by prior influenza infections. This latter feature of influenza illustrates one of several natural mechanisms by which viruses and microbes create genetic diversity in their populations. However, influenza virus does this in an ongoing fashion and at a dizzying pace, at times making fantastic genetic leaps. Many scientists consider an
influenza pandemic—one that could conceivably kill tens or even hundreds of millions of people worldwide—to be imminent.71 With the spread of the highly pathogenic avian influenza (H5N1) in Asia and Eurasia, politicians and the general public have recently begun to realize the danger.72
Sometimes referred to as a “continually emerging” infectious disease agent (as opposed to an emerging one), influenza viruses cause epidemics annually, in part aided by a phenomenon known as “antigenic drift.” Antigenic drift reflects the fact that the virus constantly accumulates genetic mutations (errors in its genome) and that over time, this eventually results in significant antigenic changes in its surface proteins that lessen their ability to be recognized by virus-neutralizing antibodies prevailing in the host population. Less often, a more dramatic change in the antigenic structure of the virus takes place through a process of reassortment of its segmented genome. This occurs through mixing of gene segments from different influenza viruses co-infecting the same host, producing a new influenza strain with a different complement of gene segments. Reassortment between avian and human influenza viruses is thought to occur in intermediate hosts, such as swine, and can lead to the appearance of a novel human virus with potentially heightened virulence as well as complete resistance to preexisting immunity (“antigenic shift”).73 Such an event is thought to have led to the emergence of a pandemic strain of influenza virus three times in the past century: in 1918 (“Spanish” influenza, H1N1); in 1957 (“Asian” influenza, H2N2); and in 1968 (“Hong Kong” influenza, H3N2).74 The 1918 influenza A pandemic (H1N1), which may have claimed as many as 50 million to 100 million lives worldwide in less than a year, ranks as one of the worst disasters in human history.75 Isolated cases and small outbreaks of disease due to highly pathogenic avian influenza have become more frequent over the past decade. The current epizootic of H5N1 avian influenza in Asia and Eurasia is unprecedented in its scale, geographic distribution, and economic loss. Tens of millions of birds have died of influenza, and hundreds of millions more have been culled to protect humans.76 According to information provided by the World Health Organization, between January 2003 and April 21, 2006, there were 204 confirmed human cases and 113 deaths of avian influenza A (H5N1), spread across 9 countries: Viet Nam (93 cases, 42 deaths), Thailand (22 cases, 14 deaths), Cambodia (6 cases, 6 deaths), China (17 cases, 12 deaths), Indonesia (32 cases, 24 deaths), Turkey (12 cases, 4 deaths), and Iraq (2 cases, 2 deaths).77 Evidence suggests that the currently circulating H5N1 virus has accumulated mutations that have made it increasingly infectious and deadly in multiple bird species, as well as in mammals.78 Thus far there has been little evidence of human-to-human transmission of the H5N1 virus, but
many experts are concerned that the virus may need to accumulate only a limited number of mutations in order for it to acquire the ability to be efficiently spread between people.
The influenza virus genome is composed of RNA, placing it among a group of human pathogens that evolve relatively quickly, even within a single host. Other RNA viruses, such as HIV and hepatitis C, are also particularly prone to this behavior. These agents generate significant degrees of genetic variability and appear to have “sampled” (or to be sampling) a large proportion of, if not all possible, gene sequence possibilities (“sequence space”) as they replicate and spread in host populations. This is due to the lack of proofreading capabilities in the polymerases responsible for amplification or copying their RNA genomes, a feature that distinguishes these RNA-dependent RNA polymerases and reverse transcriptases from the DNA polymerases that copy bacterial or protozoan genomes. This lack of proofreading allows for rapid generation of sequence diversity and, coupled with very efficient replication schema (a typical, chronically infected human, produces about 1012 new hepatitis C virus particles per day and a typical HIV-infected person about 10-fold less), promotes a process of accelerated “natural selection” that optimizes the ability of the virus to sustain a successful interaction with the host, which generally means to be able to multiply and spread to the next host.
Despite the fact that their genomes are copied with much greater fidelity, bacteria are also capable of generating genetic diversity by sharing mobile genetic elements, such as plasmids, or by receiving exogenous genes via bacterial virus infection. In fact, genes that confer virulence on a bacterium tend to be carried on mobile genetic elements.79 These “accessory” genes and functional potential allow an organism to compete more successfully in their interactions with a host and are dispensable to those microbes that either choose a different (exogenous) habitat or temporarily adopt a nonpathogenic lifestyle in the host. Complex regulatory systems recognize cues indicative of the host environment and modulate expression of virulence-associated genes accordingly. Some DNA viruses, such as the herpesviruses, appear to have “picked up” genes from their hosts, probably through a process of DNA recombination, and have modified these for their own purposes, thus increasing their genetic diversity and potential for survival.
Given the clear capability of at least some microbes and viruses to evolve quickly, acquire new genes, and alter their behavior, it might seem reasonable that over hundreds of thousands of years all conceivable biological agents have been “built” and “tested” and that the agents seen today are the most “successful” of these. Thus, is there any reason to think that it might be possible to artifically create a more successful biological agent? Possibly not, but it is important to understand that “successful” in
this context means the most able to survive within, on, or near human populations over time. “Success” does not necessarily equate with virulence, or pathogenicity, the ability to cause disease or injury.
The Evolution of Pathogenicity: What Does It Take to Cause Disease?
Early views of pathogenicity and virulence were based on the assumption that these characteristics were intrinsic properties of microorganisms, although it was recognized that pathogenicity was neither invariant nor absolute.80 Over the course of the last century, as increasing numbers of viral and microbial pathogens were identified and the pathogenesis of multiple infectious diseases was characterized, the complexity and individuality of host-pathogen relationships became evident, while the general definitions of pathogenicity and virulence became increasingly qualified and cumbersome. Viral pathogenicity reflects two fundamental features of a viral infection: the ability of the virus to cause direct injury to tissue (i.e., its cytopathogenicity) and the amount of injury associated with either an effective or ineffective immune response to the presence of the virus. More subtle forms of pathogenicity also arise, such as when a virus such as a papillomavirus causes malignant transformation of a cell or when a differentiated cellular function (i.e., insulin secretion or T-helper cell function) is lost along with the targeted destruction of a special, differentiated cell type. Bacterial, fungal, and multicellular parasites are somewhat different in that they can survive on host mucosal surfaces or skin, and do not necessarily require invasion of a cell to multiply and survive. Pathogenicity may be equated in many, but not all, bacterial pathogens with the inherent ability to cross host cell barriers—a property conferred by the expression of virulence factors, many of which are encoded by discrete DNA segments known as pathogenicity islands. On the other hand, from the host’s perspective, disease occurs only when the presence of a microbe (whether protozoan, bacterial, or viral) results in damage—whether that damage is actually mediated by the pathogen itself or by the host’s immune response to it.81
Virulence—defined broadly as the ability of an infectious agent to cause disease in a host—is a relatively rare trait even among those microbes capable of survival within a host, such as humans. All but a tiny fraction of the microbes that have thus far been found on the planet are incapable of replication under human physiologic conditions. Of those that spend a significant fraction of their existence in a human or other mammalian host, most are on or near a mucosal surface, or on the skin, in competition with a wide variety of other microbial strains and species. The human body has been estimated to contain approximately 1014 cells, 90 percent of which are microbial!82 Some human endogenous sites, such
as the skin and mucosal surfaces, have evolved with a robust microbial community in attendance. Successful colonization of these sites by so-called commensals does not precipitate a strong enough response to result in damage. In fact, most microbes that reside in the gut—such as Lactobacillus spp. and Bacteroides spp.—may actually serve a protective, not pathogenic, role.83
Rather than producing overt illness in their hosts, the vast majority of microbes establish themselves as persistent colonists: either low-impact parasites (organisms that cause asymptomatic infections), commensals (organisms that “eat from the same table,” deriving benefit without harming their hosts), or symbionts (microbes that benefit their hosts).84 These states, while separate, represent a section of a continuum—one that extends to pathogenicity and disease—which is occupied by various microbial species at various times depending on environmental, genetic, and host factors.85 Persistent colonization of a host by a microbe is rarely a random event; such coexistence depends on a relationship between host and microbe that can be characterized as a stable equilibrium.86 Pathogenic microbes acquire genes that enable them to exploit their hosts, but they generally have evolved to do so in ways that allow both the hosts and pathogens to persist. In certain cases (e.g., when microbes cause persistent, asymptomatic infection), this equilibrium can be disrupted by physiological or genetic changes in either the host or microbe, shifting the relationship toward pathogenesis and resulting in illness and possibly death for the host.87 The selective forces controlling evolution of the microbe are determined by its survival on the planet, not necessarily what it does to its host. However, there are often no direct positive benefits derived by the microbe per se in causing disease or killing its host.88
Of the several thousand species estimated to inhabit the body, only a handful are capable of causing disease on a routine basis, while only a modest additional number are capable of causing disease when host defenses become impaired. Those that regularly cause disease in unimpaired hosts employ a strategy for replication and survival that involves colonization of a highly protected anatomic site that is usually off-limits to microbes; the strategy includes mechanisms for resisting or subverting host defenses. The net result of this strategy and the ensuing host response is damage to the host and disease. Among the different viral and bacterial species that routinely cause human disease, there are multiple and diverse strategies for gaining access to the appropriate habitat, adhering to the relevant receptors, overcoming host defenses, replicating and/or persisting.89 Sometimes the strategy involves a long-term association with the host and more subtle disruptions of host physiology. Hepatitis C virus is a prime example, causing a persistent infection of the liver that typically remains asymptomatic for decades and causes significant disease
(cirrhosis or liver cancer) only in a minority of infected persons. Another is Helicobacter pylori, which infects half the world’s population but causes gastric disease in only one out of five carriers. H. pylori is an example of a potentially pathogenic (parasitic) microbe that more often assumes the role of commensal or symbiont.90
Recent research indicates that viral as well as bacterial pathogens that infect or colonize animals share broadly common strategies with those that infect plants.91 Both can express proteins that mimic, suppress, or modulate host cell-signaling pathways and enhance pathogen fitness, and both are recognized by similarly sophisticated host surveillance systems. Striking architectural similarities between surface appendages of plant and animal pathogenic bacteria suggest common mechanisms of infection, while structural differences—most notably the presence versus absence of a cell wall—reflect the profound differences between plant and animal cells. Studies of “interkingdom” pathogens, such as Pseudomonas aeruginosa, which can infect both humans and plants, such as Arabidopsis thaliana, reveal common features that permit a wide host range.92
Pathogenic bacteria are relatively restricted in their phylogenetic distribution across the bacterial domain. To date, only 7 of the more than 80 divisions of bacteria contain well-recognized pathogens, and within these 7 divisions the distribution of pathogens is focal. Clearly, some microbes are inherently capable of adapting to life within or on humans, while others are not. Notably, there is not a single known organism within the domain Archaea that is capable on its own of causing disease in humans (see Figure 1-2).93
A few methanogens are common inhabitants of the human intestinal tract, and they have recently become strongly implicated in the common gum disease chronic periodontitis, where their role is believed to be indirect, as partners in syntrophic relationships with other bacteria.94 However, the true spectrum of archaeal associations with disease and archaeal virulence mechanisms has barely been explored, in part because of the difficulty in detecting and characterizing these organisms. Similarly, in recent years there has been a growing awareness that viral agents also inhabit and replicate robustly in humans in the absence of disease expression, such as the DNA TT viruses95 that persistently infect the majority of some well-studied human populations, or GB virus C, a distantly-related, non-hepatotropic cousin of the RNA hepatitis C virus that thus far is not recognized to cause any specific disease.
In discussing the known diversity of pathogens and the considerable microbial community diversity that has yet to be characterized, it is important to recognize that the pressures that have guided the evolution of these specialized microbes over long periods of time impose a number of critical constraints. All organisms, including pathogens, have been se-
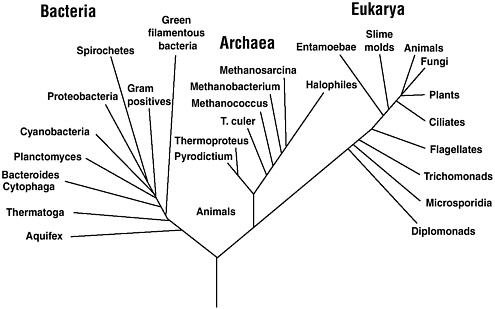
FIGURE 1-2 Archaea represents one of the three domains of life, the other two being Bacteria and Eukarya.
SOURCE: Todar, K. 2004. Major Groups of Prokaryotes. University of Wisconsin-Madison Department of Bacteriology. Available online at www.bact.wisc.edu/Bact303/MajorGroupsOfProkaryotes [accessed January 4, 2006].
lected for their ability to persist and survive on the planet. There is, after all, a fitness cost associated with being too virulent—not just for the host but for the parasite as well. If a pathogen kills or inflicts too much damage on its host, it may impact its own survivability and transmissibility, and could end up effectively committing suicide. Most successful pathogens maintain an evolutionary middle ground with respect to the amount of damage they exact on their host; to survive in the privileged anatomic niche they have chosen and to be transmitted to a new susceptible host, they may need to inflict some degree of injury but not so much that they hinder the fitness of their host as an optimal partner in attaining these goals: pathogen survival, persistence, and transmission.
An interesting and potentially serious anomaly is provided by those infectious agents, such as some arthropod-borne viruses like West Nile virus, that infect humans “accidentally.” For such pathogens, human infection is not a necessary part of its essential life cycle, as for example West Nile virus usually cycles between avian and mosquito species with only occasional forays into mammals such as humans. When this happens, disease (tissue injury) can be catastrophic, as the host-pathogen
interaction has not been tempered by evolution. It is interesting that most pathogens on the category A select agent list fall into this category, among them anthrax, plague, tularemia, and the viral hemorrhagic fevers.
Recent studies have suggested that some natural human pathogens are not nearly as virulent as they could be. Genes associated with “hypervirulence” have been identified: Some encode products that down-modulate virulence (mutations that lead to depressed virulence); other genes that are found to be inactive or missing in some pathogens encode factors that might enhance virulence in some hosts.96 Hyper-virulence is thought to reflect an exaggerated form of behavior that might be deleterious to the microbe over long periods of time, and a behavior against which natural selective forces tend to act; however, over shorter periods of time, or in more restricted portions of the natural environment, these behaviors or capabilities might not be so detrimental or even relevant. For example, in extant strains of disease-causing Salmonella enterica, the product of the pcgL gene, a D-Ala-D-Ala dipeptidase, facilitates growth in nutrient-poor conditions, presumably found outside the human host.97 Naturally occurring disruptions of this gene in some clinical strains cause the organism to be hyper-virulent, but at the cost of impaired survival in the external environment. Some pathogens occasionally increase their virulence to overcome a different type of disadvantage they face in the environment, such as poor vector competence.98
Thus, the setting and conditions under which nature judges the “success” of a pathogen may limit our appreciation for the kinds of virulence properties that might be possible in a biological agent and cause us to arrive at false conclusions concerning our ability to create new pathogenic agents. The overall survival strategy of a pathogen may involve adaptation to an external environment and transmission among hosts over thousands of years; the result is often attenuation of virulence for the human host.99 Nature rewards long-term survival of an organism whereas, long-term survival is not an important requirement for an organism to be capable of causing disease and disrupting human populations in the short-term (e.g., months, years). This is particularly so if the infectious agent is aided and abetted in its production and distribution by a motivated and knowledgeable actor. Such reflections suggest that the spectrum of virulence, both in terms of severity and phylogenetic diversity, observed among natural pathogens may not reflect the spectrum that is possible and that might be achieved, albeit for only short periods of time, if one were to explore additional realms of genetic diversity and genomic arrangement (see Chapter 3, “sampling biological diversity”). Thus, Mother Nature may not be “the mother of all terrorists” after all, and it is reasonable to anticipate that humans are capable of engineering infectious agents with virulence equal to or perhaps far worse than any observed naturally.
The Importance of the Host Response
Our growing understanding of host-microbial interactions has led to an increased awareness of the host as a key component and determinant of the host-microbe outcome. While this greater awareness and understanding of host defense may lead to new strategies for the recognition, prevention, treatment, and prediction of outcome of microbial disease, it also broadens the knowledge base from which bioterrorists could design new forms of biological weapons that disrupt host homeostatic systems,100 with diverse and potentially devastating consequences.
Irrespective of whether caused by a virus or bacterium, infectious diseases typically result either from direct tissue injury caused by an infectious agent or from the host’s response to it. How, then, does the host differentiate between pathogenic and benign microbes? Evidence has accumulated, over the past decade, to suggest that the host’s “innate” immune system recognizes the presence of molecular “danger signals” in the form of molecular patterns that are primarily associated with most microorganisms and viruses, regardless of whether it is a pathogen or commensal.101 For example, in mammals, considerable research on the epithelial cells lining the gut and other mucosal surfaces indicates that this cell layer serves as a key interchange in a signaling network that transmits signals between microbes, and adjacent and underlying immune and inflammatory cells.102 Commensal microorganisms constantly stimulate pattern recognition receptors (e.g., Toll-like receptors (TLR)) at a low level and produce stereotyped responses that are protective for the gut mucosa.103 Thus, in addition to providing a mechanical barrier between the host and its environment, mucosal epithelial cells sense the makeup of the intestinal microflora and provide signals to the host that affect the growth, development, and function of nearby cells, including the activation of inflammatory and immune responses. Macrophages and other cells of the immune system continuously patrol the inner tissues of the body, expressing similar molecular pattern receptors and constantly on the search for similar stimuli where they should not be. Parenchymal cells also express Toll-like receptors and, while much less well studied, are likely to contribute to this constant watch. The distinction between host responses to pathogens and commensals is thus likely to relate to the location, as well as the timing, duration, and intensity of the stimuli that initiate these stereotyped responses. The complexion of the responses signaled by the activation of these molecular pattern receptors can be modified through the action of secreted virulence factors (e.g., toxins) or in the case of many viruses by direct interference with the intracellular signaling pathways. Either an inappropriate dampening or induction of signaling by the host’s pattern recognition receptors, or the provocation of signaling responses
in cells that do not typically encounter microbes, can produce damage and disease.
The ability to measure host responses in terms of genome-wide fluctuations in gene transcript or protein abundance, using DNA microarrays or mass spectroscopy, respectively, has raised the possibility of new approaches for early diagnosis and outcome prediction in infectious diseases. Patterns of host transcript or protein abundance may reveal the nature of the causative factor in microbial disease and help classify infected hosts based on future clinical course, as well as help elucidate disease mechanisms.
Comprehensive analyses of host responses will also help define signal transduction pathways and the regulatory mechanisms elicited by biological agents and may lead to new therapeutic approaches for controlling inflammatory and immune responses, as well as for regulating the growth and development of epithelial and other cell types. For example, recent findings indicate that Crohn’s disease—a chronic inflammatory bowel disease—results from mutations in a recently recognized mammalian microbe-associated molecular pattern (MAMP) receptor known as Nod2, which recognizes as ligand a bacterial-derived muramyl peptide.104 These mutations appear to result in defective regulation of the MAMP receptor’s response to either commensal or pathogenic bacteria, with resultant aberrant downstream signaling.
Thus elucidation of the wiring diagram of the host and its programs for responding to pathogens, as well as detailed descriptions of the varied strategies used by pathogens for disruption of these programs, will be critically important for the design of improved diagnostics, therapeutics, and preventives. However, it will also provide new and potent opportunities for the would-be malefactor. Deliberate efforts to disrupt host response systems could target critical nodes in the host cell wiring diagram directly. Importantly, the effectors (i.e., weapons) might not necessarily be traditional “threat agents.”105
Advancing Technologies Will Alter the Future Threat Spectrum
Although this report is concerned with the evolution of science and technology capabilities over the next 5 to 10 years with implications for next-generation threats, it is clear that today’s capabilities in the life sciences and related technologies may have already changed the nature of the biothreat “space.” In a 1996 Department of Defense (DOD) report, it was argued that advances in biotechnology and genetic engineering had provided the means to modify agents in very specific ways and had consequently facilitated the development of a new generation of biological warfare agents that could potentially be more dangerous than classical
agents. The report identified five “potential types of novel biological agents”—all microorganisms: benign microorganisms that have been genetically altered to produce a toxin, venom, or bioregulator; engineered microorganisms that are resistant to antibiotics, standard vaccines, and therapeutics; microorganisms with enhanced aerosol and environmental stability; immunologically-altered microorganisms able to defeat standard identification, detection, and diagnostic methods; and combinations of any of the above four types coupled with improved delivery systems.106 In a 1997 study, six imagined future bioweapon constructs were described, conveying an even stronger sense of the broad range of possibilities that could be created by advances in life science technologies: designer genes, viruses, and other life forms (e.g., organisms or life forms designed to be drug-resistant); designer diseases; binary biological weapons (i.e., one involving infection with one agent that has little initial pathogenic consequence until a subsequent co-infection from a second organism, or an environmental trigger, activates the pathogenic aspect of the original infection); gene therapy as a weapon; stealth viruses (i.e., cryptic viral infections); and host-swapping (zoonotic) diseases.107 This dynamic biological “threat space”—past, present, and future—is illustrated in Figure 1-3. This timeline depicts the relative threat level presented by traditional
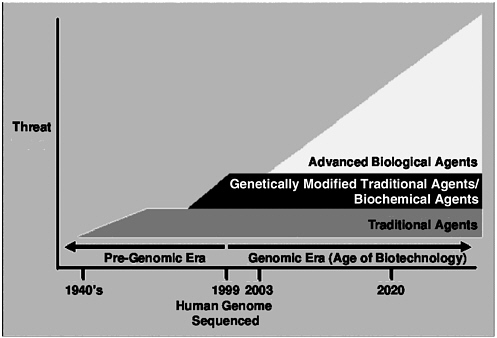
FIGURE 1-3 Timeline describing impact of biotechnology on biological warfare threat. Reprinted with permission from BioSecurity and Bioterrorism: Biodefense Strategy, Practice, and Science 1(3), 2003.
agents (e.g., naturally occurring bacterial or viral agents), genetically modified traditional agents (e.g., antibiotic-resistant bacteria), and advanced biological agents (novel biological weapons agents created by using biotechnological applications).
Returning to influenza as an illustrative example, advances in technology have led to the possibility that, even if a new lethal influenza A virus does not emerge in nature in the near future, one could be artificially generated through reverse genetic engineering (see Chapter 3 for a description of the technology).108 Although not possible until recently with negative-strand RNA viruses, in October 2004 researchers at the University of Wisconsin used reverse genetic engineering techniques to partially reconstruct the highly virulent strain of influenza responsible for the 1918-1919 pandemic;109 and the following year the complete sequence and characterization of the 1918-1919 influenza A virus was reconstructed.110 Although the knowledge, facilities, and ingenuity to carry out this sort of experiment are beyond the abilities of most non-experts at this time, this situation is likely to change over the next 5 to 10 years.111
Some experts argue that bioregulators, which are small, biologically active organic compounds, may pose a more serious dual-use risk than had been previously perceived, particularly as improved targeted delivery technologies have made the potential dissemination of these compounds much more feasible than in the past. This shift in the perceived magnitude of the risk posed by bioregulators highlights the fact that the materials, equipment, and technology necessary for disseminating and delivering biological agents to their intended recipient(s) may be equally, if not more, important than the agents themselves in determining the risk they pose. However, growing understanding of how these macromolecules function and interact in mammalian systems has also increased their dual-use potential.112
The immune, neurological, and neuroendocrine systems are particularly vulnerable to bioregulator modification. In fact, the capacity to develop bioweapons that can be aimed at the interaction of the immune and neuroendocrine systems again points to a shift in focus from the agents to, in this case, how a range of agents can be exploited (or created) to affect the human body in targeted, insidious ways. In addition to bioregulators, large numbers of potentially toxic compounds are being generated, and identified as such, as a result of high-throughput drug discovery efforts, reflecting important changes in the drug discovery process and the expanded application of new combinatorial chemistry and other high-throughput technologies. Small interfering RNAs, described in detail earlier in this chapter, also fall into this category of biologically active small molecules.
The growing concern regarding bioregulators and other agents of biological origin does not diminish the importance of naturally-occurring or
engineered pathogenic organisms. But it does mandate the need to adopt a broader perspective in assessing the threat, focusing not on a narrow list of pathogens, but on a much wider spectrum that includes biologically active chemical agents. Bioregulators are not currently included in the select agents list, which comprises those agents currently considered to be the greatest biosecurity threats.113 The threat spectrum is broad and evolving—in some ways predictably, in other ways unexpectedly. The viruses, microbes, and toxins listed as select agents are just one aspect of the continually changing, complex threat landscape.114,115 In the future, genetic engineering and other technologies may lead to the development of pathogenic organisms with unique and unpredictable characteristics, and biochemical compounds that target homeostatic and physiological processes will likely pose a greater threat than they do now.
The Development and Use of Biological Weapons
Human history seems to suggest that, as technology advances, malevolent use is the rule, not the exception. As Matthew Meselson observed, “Every major technology—metallurgy, explosives, internal combustion, aviation, electronics, nuclear energy—has been intensively exploited, not only for peaceful purposes, but also hostile ones.”116 The rapidly growing and constantly shifting global knowledge base and technology landscape may lessen delivery and other challenges to developing bioweapons capabilities—at both the state and individual actor levels. With respect to the former, based on unclassified U.S. government information, the media, foreign governments, and academic reports, there are about a dozen countries with suspected, likely, or probable biological weapons capabilities.117 Although biological weapons are outlawed by the 1972 BWC,118 the prohibitions of the BWC do not extend to basic and applied research nor to the actions of individuals—only state parties. In the 30 years since the BWC came into effect, the acquisition and engineering of biological agents has become easier and cheaper for both state and nonstate actors, and for individuals. State actors that have not been involved with biological weapons in the past (e.g., because such weapons have not been considered accessible, or particularly useful) may start developing new bioweapons programs, either secretly or under the cover of biodefense research programs. Likewise, the technologies and tools required to develop bioweapons capabilities are becoming increasingly accessible and affordable to individuals.
National security experts and even biologists themselves are concerned that rogue scientists could create new biological weapons—such as deadly viruses that lack natural foes. They also worry about innocent mistakes: organisms that could potentially create havoc if allowed to re-
produce outside the lab. In July 2000, the topic of Scientific American magazine’s “The Amateur Scientist” was “PCR at Home.” Gazing into the not too distant future, it seems likely that not only will the fruits of biotechnology—whether recombinant vaccines or GloFish™—become increasingly common features of everyday life, but the technology itself, like computer technology, will become increasingly accessible to interested amateurs. If and when biotechnology becomes as user-friendly as computer technology is currently, plant and animal breeding hobbyists and professionals, as well as gardeners, pet owners, and “bio-Unabombers”119 will likely access and use these technologies for good or evil. Sooner or later, it is reasonable to expect the appearance of “bio-hackers,”120,121 mirroring the computer hackers that repeatedly cause mischief today through the creation of a succession of more and more sophisticated computer “viruses.”
Since the mid-1990s, when the specter of bioterrorism emerged, only a small number of credible incidents of bioterrorism have been experienced—such as the mailing of anthrax spores through the U.S. Postal Service in the fall of 2001. Additionally, a few criminal events have occurred that have involved the use of disease-causing microorganisms or biologically-derived toxins, as can be seen in Table 1-2. The data presented below exclude accidental releases, such as happened in Sverdlovsk, USSR, in April 1979, resulting in the documented deaths of 67 people from inhalational anthrax as well as purposeful, experimental releases, such as the
TABLE 1-2 Authenticated Acts of Biological Warfare or Terrorism Directed Against People, 1940 to 2004
|
Date |
Location |
Details of Episode |
Key Information Sources |
|
1940-1941 |
China: Hangzhou and Nanjing |
Japanese aircraft drop packages containing fleas infected with Yersinia pestis. There are reports of several other such episodes later. |
Recent testimony in a Tokyo court by one of the aircraft pilots. |
|
1957-1963 |
Brazil: Mato Grosso |
Introduction of smallpox, influenza, tuberculosis, and measles into Indian tribal populations via contaminated gifts and mestizos in furtherance of large-scale land takeovers. |
The Figueiredo report (1968), which led to the indictment of 134 employees of the Government Service for the Protection of Indians |
|
Date |
Location |
Details of Episode |
Key Information Sources |
|
1981 |
United Kingdom: Chemical Defence Establishment (CDE) Porton Down, Wiltshire |
“Dark Harvest Commandos” deposit outside a defense research facility a parcel of soil containing anthrax bacteria taken from a former bioweapons proving ground. |
The perpetrators’ own account plus CDE’s soil analysis confirming presence of B. anthracis at less than 10 org/gram. |
|
1984 |
United States: The Dalles, Oregon |
Rajneeshee cultists seek to influence local elections by infecting voters with salmonellosis by contaminating neighborhood restaurants. |
Subsequent medical investigation of 751 sick persons. |
|
1989 |
Namibia: near Windhoek |
Covert operation by a South African government agency, the Civil Cooperation Bureau (CCB), to contaminate the water supply of a refugee camp with cholera bacteria. |
Perpetrator’s testimony during recent trial of Brig. Dr. Wouter Basson. |
|
1990-1993 |
Japan: Tokyo |
Aum Shinrikyo cultists, prior to their 1994-1995 sarin attacks, had sprayed biological agents, including anthrax bacteria, against several U.S. and other facilities in and around the city, but with no discernible effect. |
Confessions and other information contained in leaked police reports. |
|
2001 |
United States |
U.S. Postal Service used by still-unknown perpetrator(s) to spread anthrax spores contained in letters addressed to individuals in the media and the U.S. Senate: 22 people catch anthrax in its cutaneous or inhalational forms: 5 die; 10,286 receive postexposure prophylaxis; and many more have their daily lives disrupted. |
Medical investigations coordinated through the Centers for Disease Control and Prevention. |
|
SOURCE: Compiled from published and documentary sources held in the Sussex Harvard Information Bank, University of Sussex, United Kingdom. |
|||
“hot agent” field trials conducted by the United Kingdom, the United States, and others. Episodes involving non-harmful simulants, such as the biological hoaxes whose proliferation accelerated in 1997, are excluded from the table, as are the myriad recent instances of false alarms, in which substances initially suspected of being harmful were subsequently found not to be so.
However, with the increasing availability of and accessibility to pertinent knowledge and technology, it is not at all unreasonable to anticipate that biological threats will be increasingly sought, threatened, and used for warfare, terrorism, and criminal purposes and by increasingly less sophisticated and resourced individuals, groups, or nations. One can envision that the ratio of credible threats and actual attacks relative to hoaxes could actually increase substantially in the years to come.
Biological Weapons Are Fundamentally Different from Other “Weapons of Mass Destruction”
Clear thinking about threats posed by and appropriate responses to biological, chemical, and nuclear weapons is often confused by the vocabulary commonly used in discussing these threats. In particular, analysts, policymakers, journalists, and even scientists often refer casually to weapons of mass destruction, or WMD, as if biological, chemical, and nuclear weapons were each merely variants of some common type of weapon. But, in fact, these weapons differ greatly in their proliferation potential, the challenges they pose for deterrence, and the effectiveness of defensive measures (each of these areas of difference are discussed in detail below).122 Moreover, the WMD label fails to capture the disparate future trajectories of the technologies underlying biological, chemical, and nuclear weapons, so it will likely become ever more misleading over time. Box 1-3 provides the United Nations’ definition of weapons of mass destruction. Some communities, such as the DOD, have tried to reconcile the “terminology problem” by introducing yet another term “weapons of mass effect.”123
This does not mean there is no common ground or any lessons to be applied from experience dealing with one class of weapons to the others. Moreover, there is no question that from a political and strategic point of view these weapons are often strongly coupled, for example, in the connection between biological or chemical weapons and state doctrines regarding “no first use” of nuclear weapons. Of note, one striking similarity between nuclear technology and biotechnology is their dual-use nature. Nuclear technology was seen from the beginning as offering peaceful benefits, which today include research reactors, power reactors, and radioisotope production. The range of benefits is not nearly as great as the media
|
BOX 1-3 In 1948, the United Nations defined “weapons of mass destruction” as “atomic explosive weapons, radio-active material weapons, lethal chemical and biological weapons, and any weapons developed in the future which have characteristics comparable in destructive effect to those of the atomic bomb.” SOURCE: United Nations Security Council, Commission for Conventional Armaments. 1948. Resolution Adopted by the Commisssion at Its Thirteenth Meeting, 12 August 1948, and a Second Progress Report of the Commisssion. S/C.3/32/Rev.1; August 12:2. |
hype predicted back in the mid-1950s when President Eisenhower presented his Atoms for Peace program to the General Assembly of Nations (e.g., automobiles that would run for a year on vitamin-sized nuclear pellets).124 Yet despite this similarity, there is a demarcation between weapons-related and nonweapons-related nuclear science and technology. From the beginning, the civilian uses of nuclear energy were cordoned off from weapons developments through a large investment in security classification, international diplomacy, and a discourse that insists that nuclear weapons are special. Quite the opposite is true of the life sciences.
Clear thinking on the issue must proceed from an understanding of the significant differences among these weapons. Although there are lessons to be learned from the history of and our experience with nuclear weapons technology, many of the differences between the nuclear and biological realms are too great to adopt a similar mix of nonproliferation, deterrence, and defense. Effective strategies for anticipating, identifying, and mitigating the dangers associated with advancing and emerging life sciences technologies demand a clear understanding of the varied and unique nature of the biological threat spectrum.
Proliferation Potential
One of the most pronounced differences between biological weapons and other weapons of mass destruction is the proliferation potential of the former. Imagine a line that begins with nuclear weapons at the left, continues through chemical, radiological and biological weapons, and terminates with cyber “weapons” on the right. Moving from left to right along
TABLE 1-3 Characteristcis of Fissile Materials and Pathogens
|
Fissile Materials |
Biological Pathogens |
|
Do not exist in nature |
Generally found in nature |
|
Nonliving, synthetic |
Living, replicative |
|
Difficult and costly to produce |
Easy and cheap to produce |
|
Not diverse: plutonium and highly enriched uranium are the only fissile materials used in nuclear weapons |
Highly diverse: more than 20 pathogens are suitable for biological warfare |
|
Can be inventoried and tracked in a quantitative manner |
Because pathogens reproduce, inventory control is unreliable |
|
Can be detected at a distance from the emission of ionizing radiation |
Cannot be detected at a distance with available technologies |
|
Weapons-grade fissile materials are stored at a limited number of military sites |
Pathogens are present in many types of facilities and at multiple locations within a facility |
|
Few nonmilitary applications (such as research reactors, thermoelectric generators, and production of radioisotopes) |
Many legitimate applications in biomedical research and the pharmaceutical/biotechnology industry |
|
SOURCE: Tucker, J.B. 2003. Preventing the misuse of pathogens: The need for global biosecurity standards. Arms Control Today, June. |
|
this continuum, an effective nonproliferation regime becomes increasingly difficult. To make these differences clear, it is useful to begin with a comparison of the characteristics of fissile materials and pathogens, as depicted in Table 1-3.
Nuclear nonproliferation policy seeks to limit the number of states with nuclear weapons and to keep such weapons out of the hands of nonstate groups. The Nuclear Nonproliferation Treaty (NPT) established a near-global verification regime, carried out by the International Atomic Energy Agency (IAEA), intended to prevent the diversion of fissile material from civilian use to weapons programs. The regime employs inspections, audits, and surveillance cameras and instrumentation, monitoring some 1,100 facilities and installations worldwide. Despite important shortcomings, the NPT inspection regime has been largely successful, in part because the facilities needed to produce weapons-usable uranium or plutonium are, for the most part, big and hard to hide.125 As the NPT verification regime has faced new challenges, the inspection regime has been modified in response so as to continue playing an important and credible nonproliferation role. The regime has lagged behind technical develop-
ments, but the pace of technical evolution has been slow enough that the regime has, with delay, still been useful.
The roughly forty nations of the Nuclear Suppliers Group (NSG) further pursue nonproliferation objectives by adhering to consensus guidelines for nuclear and nuclear-related dual-use exports. These guidelines are intended to supplement the NPT by controlling the transfer of listed items without hindering legitimate international nuclear cooperation. United Nations Security Council Resolution 1540 aims to extend many of these nonproliferation measures to all states, not just members of the NSG. Through Cooperative Threat Reduction, the United States and other members of the Group of Eight industrialized nations (G-8) act to impede the theft of nuclear material or the transfer of personnel with nuclear weapons-relevant knowledge from the former Soviet Union and other states. Diplomatic pressure and security guarantees have also played crucial roles, and intelligence has been vital throughout.
Despite its challenges, including difficulties created by uncooperative nations that may possess capable delivery systems, the nuclear nonproliferation regime has been reasonably successful in part because the production of nuclear weapons-usable plutonium or uranium has substantial technical requirements (reactors or enrichment plants, respectively) that create conspicuous bottlenecks for any would-be weapons program. Declared facilities that need to be monitored are few, and undeclared illegal facilities are at risk of discovery by intelligence or other means. Moreover, the special nuclear materials used in nuclear weapons do not occur naturally. Any detection of such materials outside of legitimate, declared facilities would automatically be a matter of grave concern. The theft of nuclear weapons or weapons-grade nuclear material may be the only way to avoid these bottlenecks, making the prevention of nuclear theft an extremely high priority.
At the far right end on the continuum of unconventional weapons are cyber weapons. At this extreme, a traditional nonproliferation regime seems nearly insurmountable, if not impossible. Exports of certain high-end computers, components, and codes are certainly controlled, but these neither prevent cyber attacks nor reduce the use of these technologies for illicit or malicious purposes. Such attacks may in principle be launched from any of over 100 million computers all over the world that already have access to the Internet—and that number is growing rapidly. Implementing a nonproliferation on-site verification regime analogous to that used to monitor nuclear facilities would therefore seemingly require unannounced inspections or the monitoring of hundreds of millions of residences and businesses. Cybersecurity poses a reductio ad absurdum for a traditional inspection regime.
Chemical, biological, and radiological weapons fall between the
nuclear and cyber extremes of the continuum of unconventional weapons described above in the context of nonproliferation control regimes. The Organization for the Prohibition of Chemical Weapons (OPCW), established under the Chemical Weapons Convention (CWC) oversees an international verification regime for chemical weapons that is more challenging than that for nuclear weapons because of the larger number of relevant facilities and dual-use materials. An entire industrial sector and well over 5,000 inspectable facilities must be monitored by the OPCW. Nonetheless, under the CWC, chemical weapons stocks have been declared and verified, the destruction of these stocks has begun, even if fitfully, and hundreds of inspections of dual-use chemical plants have taken place. For example, Albania, India, the Republic of Korea, Libya, Russia, and the United States have declared over 70,000 metric tons of chemical agents. Except for Albania, these six countries, along with Bosnia and Herzegovina, China, France, Iran, Japan, Serbia and Montenegro, and the United Kingdom have declared post-1945 chemical weapons production facilities (Japan’s declaration referred to facilities that had belonged to the Aum Shinrikyo). Completion of destruction of the declared stocks falls due in 2012. Once the disarmament phase is complete, the dual-use monitoring component will dominate verification, permitting increased attention to the challenges of new science and technology. The CWC regime is further supplemented by the Australia Group of nations that, analogously to the NSG, establishes consensus guidelines restricting the export of weapons-relevant materials.
Biological weapons pose greater challenges to a nonproliferation regime than do chemical weapons, so they lie closer to the cyber extreme along this continuum. There are some formal analogies to the nuclear and chemical cases, but the analogies only go so far. For example, while the maintenance of inventories of critical materials is of considerable use in monitoring the proliferation of nuclear weapons and, while less so, still of use in monitoring the proliferation of chemical weapons, there is little basis for such efforts in monitoring or preventing the proliferation of biological weapons. Due to the inherent replicative properties of bacteria and viruses, large stocks of biological agents can readily be produced from very small quantities of stolen agent, so small that their absence would not be missed from even the most carefully inventoried collections. Moreover, DNA synthesis technology is the equivalent of a matter compiler for genetic material—the transition from “matter” to “information” pushes the control of biology closer to the cyber extreme.
The 1925 Geneva Protocol forbade the use of biological and chemical weapons,126 and the BWC, which entered into force in 1975, forbids the development, production, and stockpiling of biological weapons. Article I of the treaty reads:
Each State Party to this Convention undertakes never in any circumstances to develop, produce, stockpile or otherwise acquire or retain: (1) Microbial or other biological agents, or toxins whatever their origin or method of production, of types and in quantities that have no justification for prophylactic, protective or other peaceful purposes; (2) Weapons, equipment or means of delivery designed to use such agents or toxins for hostile purposes or in armed conflict.
Like the CWC, but unlike the NPT, the BWC prohibits an entire class of weapons for every country in the world, and in that sense is perhaps better characterized as an arms control treaty than a “nonproliferation” treaty. But unlike the NPT and CWC, the BWC established no inspection and monitoring regime, and there is no agency analogous to the IAEA or OPCW, respectively, that is responsible for technical monitoring of compliance with the treaty. Monitoring compliance with this treaty is especially problematic since, under the BWC, ambiguities may arise over what level of effort is consistent with “prophylactic, protective or other peaceful purposes” and, to a considerable extent, intent becomes an extremely important issue. It may prove very difficult to determine what levels of defensive efforts are legitimate and which serve as camouflage for an illicit offensive program.
After the discovery of the Iraqi and Soviet offensive biological weapons programs in the early 1990s, an international effort began in 1994 to negotiate a compliance protocol to the BWC. The protocol would have required all countries to declare certain facilities, and it would have established on-site visits for facilities meeting certain criteria and the possibility of targeted investigations in the event of suspicions regarding either particular facilities or disease outbreaks. An Organization for the Prohibition of Biological Weapons, analogous to the OPCW, was to have been created. In July 2001, however, the United States rejected the draft protocol, stating that the BWC was unverifiable; that the proposed protocol would threaten both private-sector proprietary information and U.S. government biodefense secrets; that it would threaten existing export controls; and that it risked giving a false stamp of legitimacy to violators of the BWC by failing to detect their violations but nevertheless declaring them to be in compliance with the BWC.127
There are other nonproliferation measures in place that are reminiscent of those established for the BWC, NPT, and CWC. For example, the Australia Group adopts consensus national export controls to impede the transfer of biological agents and technology where possible. Nevertheless, the biological nonproliferation regime faces intrinsically greater challenges than does its nuclear or even chemical counterpart, because many of the relevant materials, technologies, and knowledge are far more widespread in the biological case. As this report documents, these technologies
are rapidly becoming “democratized” in that they are far more available and accessible to individuals as well as states and will become ever more so in the years to come.
Deterrence and Dissuasion
Deterrence through the threat of retaliation was a central nuclear weapons strategy during the Cold War. Deterring any form of terrorism may prove difficult, since some terrorist groups may be unconcerned about retaliation or may hope to remain unidentified. This conclusion may hold regardless of the type of weapon being used. However, biological weapons pose a unique challenge, as some infections may incubate without symptoms for days or even weeks, making it especially difficult to trace an attack back to its perpetrators.
Biological attacks may also be dissuaded through means other than the threat of retaliation (dissuasion by deterrence), such as the force of law. In the initiatives now being pressed forward nationally and internationally to criminalize the activities of individuals (i.e., activities that the BWC and the CWC prohibit at the state level), dissuasion may grow from the possibility of responsible individuals being held accountable in courts of law, provided intelligence and nonproliferation create a significant probability of detection and assuming, too, the existence of determined and empowered prosecutors.128 Dissuasion of a different kind may also reside in the obloquy of detection that is made public, and the political or other consequences that could follow from it.
Defense
So far, biological weapons have been based on agents that also cause naturally occurring disease outbreaks. This aspect of these weapons has no good analog in the realm of nuclear weapons and only partial ones in the realm of chemical weapons. Many of the same tools that address natural disease threats will be needed to respond to an attack using biological weapons, or to prevent such an attack from succeeding, and this is likely to remain true even in a future case involving a genetically engineered pathogen. In the biological case, therefore, there is the opportunity to ensure that many of the steps required to improve biodefense will benefit public health even if major acts of bioterrorism never occur. “Defense” in the case of biological security means, above all, improvements in domestic and international disease surveillance and response and strengthened public health systems. The recent evolution of biomedical defense programs has largely focused on detection and vaccine development. This “dual-use” aspect of much of biodefense is very different from the nuclear
case, though it bears some similarity to the chemical case where preparation for accidental chemical spills or explosions is relevant to consequence management for terrorism as well.
The “Arms Race” Metaphor and the Difficult Issue of Secrecy
The “arms race” metaphor must be used with caution; it too is in danger of calling up misleading analogies to the Cold War nuclear arms race that are not relevant to the biological case. We are transitioning from the classical biowarfare/bioterrorism mode in which naturally occurring agents are the threat weapons. They may have minor modifications such as induction of antibiotic resistance, but generally most countermeasures will be expected to be active, and the list of candidate agents is only debated in a small group of organisms. In this situation, production of vaccines and countermeasures is not inherently destabilizing provided transparency and interchange are maintained. (It is worth noting that we are far from solving defenses against these agents at this time.) However, in tomorrow’s world of proliferating threats that may be quite different from the 20th century agents and indeed may not even exist at this time, there will be very different considerations. The primary driver of offensive capabilities in that biological arms race is not mainly a particular adversary but rather the ongoing global advance of biotechnology and microbiological and biomedical research. That is, protective measures are in a race with the malevolent application of potentially beneficial basic research, rather than primarily against technologies being developed in weapons programs of other countries.129
However, there is a legitimate concern that defensive research undertaken in one country’s program could be misperceived as offensive, or potentially offensive, in character and drive other nations to pursue offensive research as well. For example, would it be legal and wise to have classified biodefense research activities produce modified pathogens that no adversaries are believed to have yet created in order to ostensibly understand and more robustly defend against them? On the one hand, defense against such potential threats that we can already anticipate would seem to prompt an answer of “yes.” On the other hand, such classified research risks making the responsible government a driver of the biological arms race, as other nations may misperceive such research as offensive.
Yet “open” biodefense research risks providing information to individuals or groups with malicious or malevolent intent. Strategic decisions must be made about what, if any, biological weapons research will be conducted in the name of biodefense, how much of this research will be classified, and how that program will be publicly described or even per-
formed under a regime of transparency. This too is very different from the nuclear weapons case and the NPT, under which it is legally permissible, at least for the time being, for the United States and certain other NPT states to stockpile and conduct research in nuclear weapons. Under the BWC, no nation may develop or stockpile biological weapons. However, the situation is complicated by the reality that a vast reservoir of technical information that could be used to develop bioweapons capabilities is freely available in the public domain, unlike the smaller number of closely held secrets associated with nuclear weapons technology. Moreover, much of biotechnology, unlike nuclear technology, is increasingly accessible even to minimally trained individuals.
The risks associated with open versus closed biodefense research activities highlight the difficult questions surrounding the issue of secrecy and whether there is certain biological information that should be kept secret and, if so, where the boundary that defines this information lies and what circumstances dictate a need for secrecy. Other questions about secrecy pertain to the role of “sensitive” information and whether information should only be categorized as either classified or not, and the challenges associated with imposing a global secrecy structure.130
The Need to Strike a Balance: Benefits of Technological Growth
Much of this report focuses on the potential misuse of technology. Indeed, the rationale for conducting this study was based on the growing threat of misuse posed by dual-use advances in science and technology. Importantly, there are also tremendous benefits to be gained from the very same scientific and technological advances. The purpose of this section is to enumerate some of these benefits. A more detailed discussion of these benefits, particularly for the developing world, may be found in an earlier workshop summary report from this committee, An International Perspective on Advancing Technologies and Strategies for Managing Dual-Use Risks: A Workshop Report.131
There is no question that many populations have benefited greatly from advances in biotechnology and applications of related technologies (e.g., nanotechnology and informatics) to biomedicine and agriculture. Health biotechnology holds out promises for improved nutrition, a cleaner environment, a longer and healthier lifespan, and cures for many once-formidable diseases. Even older technologies, such as classic vaccine technology, have enabled the eradication or reduction of many once-dreaded diseases, such as smallpox, polio, diphtheria, tetanus, and whooping cough. Newer reverse genetic technologies for RNA viruses may make possible rapid, rational development of vaccines against newly recognized pathogens, such as SARS, avian influenza, Nipah, and many others for
which current technologies are too slow, hit-or-miss, or too resource intensive for timely vaccine development.
In the developing world, broader application of biotechnology may make it economically feasible for resource-limited countries to produce inexpensive vaccines to protect their own populations against emerging infections that most afflict them. However, the potential applications of life sciences technologies extend far beyond more affordable vaccines. A technology foresight study conducted by the University of Toronto Joint Centre for Bioethics (JCB) identified the ten biotechnology-related developments that are likely to improve human health in developing countries within the next five to ten years:132 molecular diagnostics, recombinant vaccines, drug and vaccine delivery systems, bioremediation, sequencing pathogen genomes, female-controlled STI protection, bioinformatics, enriched genetically modified crops, recombinant drugs, and combinatorial chemistry.
In addition to improved health, world agriculture stands to benefit greatly from new discoveries in the life sciences and growing technological capabilities. Many staple crop plants represent virtual monocultures—that is, specific strains that were selected and propagated to give high yield under certain specific conditions.133 Biotechnology could enable agriculturalists to develop plants and livestock that are more resistant to disease, pests, or harsh environmental conditions. Such modified agricultural stock could be both an assurance for the future—against the possibility of a disease wiping out a major part of the food supply, as happened in the Irish potato famine of the mid-19th century—and a boon to resource-poor farmers in developing countries.
Importantly, while posing risks, U.S. biodefense efforts stand to benefit as well from global cooperation in the spread of biotechnology. Efforts to expand disease surveillance, improve detection and diagnostic capabilities, and develop new vaccines and therapeutics—all of which are crucial for a rapid, effective response in the event of either a deliberately introduced or a naturally occurring biological attack—will continue to rely on excellent scientific research and technological growth. Much of the latter in particular relies, in turn, on international collaboration, a theme explored in depth in Chapter 2. It has been argued that domestic regulations designed to strengthen biodefense but that either indirectly and/or directly restrict international collaboration may, ironically, “have an unintended negative influence on the current U.S. effort to develop enhanced biodefense capabilities, because it creates incentives to shift away from international collaboration in biodefense research and production.”134 This is particularly alarming with respect to vaccine research, in which international biotechnology companies play a vital role. Of the six category A biological threats on the U.S. Department of Health
and Human Services’ (DHHS) select agents list (i.e., anthrax, botulinum toxin, plague, smallpox, tularemia, and viral hemorraghic fevers), vaccines for all but botulinum toxin are currently being developed in cooperation with international biotechnology companies located in Canada, Japan, Russia, and throughout Europe. Global cooperation, including the training of foreign nationals in the United States, also provides opportunities to introduce foreign scientists to the concept of dual-use risk and the need to pursue a culture of awareness and responsibility in the global scientific community.
The Dual-Use Dilemma
It can be expected that tension between the potential beneficial and malevolent uses of technology will increase in the future, as science and technology increasingly empower users to manipulate the materials and processes of life itself. The same reverse genetic technologies that can be used to develop new vaccines against RNA viruses could also be used to construct modified viruses, including possibly viruses that express heterologous virulence factors that result in more lethal disease. The Fink report summarized the details and implications of three recent examples of “contentious research” in the life sciences—experiments that resulted in the creation of new infectious agents or knowledge with dual-use potential: the 2001 ectromelia virus (mousepox) experiment, in which Australian researchers engineered a recombinant virus that expressed the mouse interleukin-4 (IL-4) gene and, in so doing, inadvertently created a lethal virus that kills mice genetically resistant or recently vaccinated against mousepox;135 the 2002 announcement that researchers from the State University of New York, Stony Brook, had artificially synthesized a virulent poliovirus from scratch, using mail-order segments of DNA and a viral genome map freely available on the Internet;136 and a 2002 study on the difference in virulence between the Variola major virus, which causes smallpox and has a 30 to 40 percent mortality rate, and the vaccinia virus, which is used to vaccinate humans against smallpox and does not cause disease in immunocompetent persons.137
The following serve as additional illustrative examples of scientific publications that pose dual-use dilemmas (see also Figure 1-4):
-
Publication of the complete DNA sequences of human pathogens. This information, which is widely available on public databases, could potentially facilitate the development and production of novel biological weapons agents. According to one assessment: “The ever-expanding microbial genome databases now provide a parts list of all potential genes involved in pathogenicity and virulence, adhesion and colonization of
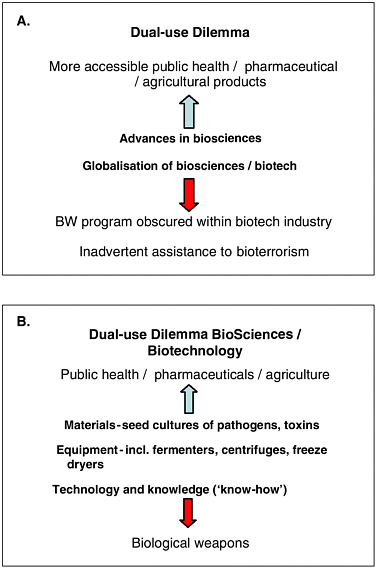
FIGURE 1-4 A and B The Dual-Use Dilemma.
SOURCE: B. Mathews, presentation at committee workshop, Cuernavaca, Mexico.
-
host cells, immune-response evasion and antibiotic resistance, from which to pick and choose the most lethal combinations.”138
-
A method for the construction of “fusion toxins,” derived from two distinct toxins, for the purpose of killing cancer cells.139 This technique might be redirected to develop novel toxins that could target the normal cells of almost any tissue when introduced into a human host.
-
Development of a genetically engineered strain of Bacillus anthracis containing an inserted gene for a foreign toxin, potentially rendering the agent resistant to the existing anthrax vaccine.140
-
Development of “stealth” viruses that can evade the human immune system and serve as molecular vectors to introduce curative genes into patients with inherited diseases, or alternatively express unwanted proteins such as toxins.141
-
Publication of molecular details of two highly virulent strains of influenza, the 1997 Hong Kong flu142 and the 1918 Spanish flu.143 Other labs are trying to sequence and publish all of the genes in the 1918 influenza strain,144 which killed 20 million to 40 million people, so as to be prepared should it reemerge.145 Such work may help control influenza, but it also makes it easier to recreate the highly pathogenic 1918 virus for malevolent purposes.
-
Genetic engineering of the tobacco plant to produce subunits of cholera toxin, making it theoretically possible to produce large quantities of this toxin cheaply and relatively easily, for good or bad.146
-
Efforts to do away with insulin injections for diabetes by developing new technologies for delivering drugs by aerosol spray.147
-
The deliberate release of botulinum toxin into the U.S. milk supply could poison several hundred thousand individuals, but early detection measures, such as in-process testing, could eliminate this threat. Researchers have explored the effects of a deliberate release of botulinum bacteria at various points in the milk supply chain, including raw milk silos and tanker delivery trucks, with varied assumptions for key variables.148 This work helps biodefense and public health workers pinpoint vulnerabilities so they can be appropriately addressed, but it also could help direct a terrorist to the most vulnerable points in the milk supply.
In the past, dual-use concerns have focused on pathogens and on the challenges associated with controlling dangerous pathogens. As already emphasized, this committee’s deliberations have indicated that the problem will be far broader and more profound in the future. For example, advances in neurobiology may make it possible to manipulate behavior and thought processes, while gene therapy and gene expression technologies just now coming to fruition will make it possible to activate endogenous molecules in the body—with possibly wide-ranging and devasting effects. Advances in synthetic biology and nanotechnology will offer similar rich opportunities for dual use. Nanodevices that may be used to unplug blocked arteries could instead be employed to interfere with circulatory function. Advanced drug delivery technologies and pharmacogenomics knowledge could be used to develop and deliver
with greater efficiency new bioweapons, perhaps even selectively targeting certain racial or ethnic groups.
COMMITTEE PROCESS
In considering the rapid and unpredictable advance of life sciences technologies, it became apparent to the committee that the possibility of inappropriate or malevolent use could never be completely eliminated without fundamentally undermining the vitality of the scientific enterprise and endangering the tremendous benefits this enterprise brings. However, we can also learn from past experiences. For example, recombinant DNA was a new enabling technology some 30 years ago, and the possibilities of this powerful new technology led to widespread concern in both the scientific and the political communities. The issue was discussed at the now-famous Asilomar Conference of 1975, when scientists gathered to discuss the safety of manipulating DNA from different species and when many of the safeguards now in place were originally developed.149 As scientists developed more experience with the technology, many of the original stringent safeguards could gradually be softened. Similarly, this committee has considered how possibilities for inappropriate and malicious use could be greatly reduced or mitigated, as discussed in greater detail in Chapter 4.
In creating the ad hoc Committee on Advances in Technology and the Prevention of Their Application to Next Generation Biowarfare Threats, the National Research Council (the operating arm of The National Academies) and the Institute of Medicine selected committee members representing a broad spectrum of backgrounds, expertise, and interests. Areas of expertise included public health, molecular and cellular biology, biophysics, clinical medicine, drug and vaccine discovery, national security and law enforcement, bioethics, and sociology (see Appendix C for biographical information of committee members). In addition, the committee relied on the expertise and advice of representatives of the executive branch of the U.S. government, international governmental and nongovernmental organizations, national governmental and nongovernmental technical and policy experts, and educators and private consultants. Information available from the open literature and materials submitted by experts were reviewed and considered during the committee’s deliberations (see Appendix B).
REPORT ROADMAP
Chapter 2 of this report reviews the current global dispersion of tools and technologies used in the life sciences enterprise both domestically
and globally. This global dispersion is being driven by a multitude of economic, social, and political forces. Chapter 3 provides an overview and perspective on the breadth and types of technologies that will—directly or indirectly—have an impact on how the life sciences enterprise will evolve in the near-term future. Finally, Chapter 4 presents the committee’s conclusions and recommendations about the ways in which the adoption of a “web of prevention” approach might enhance our collective abilities to mitigate or minimize the negative consequences of inadvertent, inappropriate or purposeful malevolent applications of any of these technologies in the decades to come.
ENDNOTES
|
1 |
Cracraft, J. 2004. Editorial: A new AIBS for the age of biology. BioScience (November). Available online at www.aibs.org/bioscience-editorials/editorial_2004_11.html [accessed January 4, 2006]. |
|
2 |
Meselson, M. 1999. The problem of biological weapons. Presentation given to the 1818th Slated Meeting of the American Academy of Arts and Sciences, Cambridge, MA, January 13. |
|
3 |
National Intelligence Council. 2004. Mapping the Global Future, Report of the National Intelligence Council’s 2020 Project. Available online at www.cia.gov/nic/NIC_globaltrend2020.html#contents [accessed May 3, 2005]. |
|
4 |
National Research Council. 2004. Biotechnology Research in an Age of Terrorism. Washington, DC: The National Academies Press. |
|
5 |
Couzin, J. 2002. Breakthrough of the year: small RNAs make big splash. Science 298(5602):2296-2297. Available online at http://www.sciencemag.org/cgi/reprint/298/5602/2296.pdf [accessed January 4, 2006]. |
|
6 |
Cancer Bulletin, National Cancer Institute. 2005. “Cancer biologists are using RNAi to do everything from investigating individual genes to running high-throughput screens for new drugs to developing therapeutics,” says Dr. Natasha Caplen, head of the Gene Silencing Section in NCI’s Center for Cancer Research. Available online at www.nci.nih.gov/ncicancerbulletin/NCI_Cancer_Bulletin_032905/page4 [accessed January 4, 2006]. |
|
7 |
10 emerging technologies that will change your world. Technology Review (February 2004). Available online at www.lib.demokritos.gr/InTheNews/emerging0204.htm [accessed January 4, 2006]. |
|
8 |
Scherr, M. et al. 2003. Inhibition of GM-CSF receptor function by stable RNA interference in a NOD/SCID mouse hematopoietic stem cell transplantation model. Oligonucleotides 13:353-363; Song, E. et al. 2003. RNA Interference targeting Fas protects mice from fulminant hepatitis. Nature Medicine 9(3):347-351; Soutschek, J. et al. 2004. Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 432(7014):173-178. |
|
9 |
Frost & Sullivan. 2004:B349. |
|
10 |
Ibid. |
|
11 |
10 emerging technologies that will change your world Technology Review |
|
|
(February 2004). Available online at www.lib.demokritos.gr/InTheNews/emerging0204.htm [accessed January 4, 2006]. |
|
12 |
Ball, P. 2004. Synthetic biology: Starting from scratch. Nature 431 (7009): 624-626. Available online at www.nature.com/nature/journal/v431/n7009/pdf/431624a.pdf [accessed January 4, 2006]. |
|
13 |
Morton, O. 2005. Life, reinvented. Wired 13.01 (January). Available online at www.wired.com/wired/archive/13.01/mit.html [accessed January 4, 2006]. |
|
14 |
June 29 /PRNewswire/—Synthetic Genomics, Inc., a new company that will develop and commercialize synthetic biology, was launched today. J. Craig Venter, Ph.D., is the founder, chairman, and chief executive officer of the company. Synthetic Genomics, Inc. was founded in 2005 to develop and commercialize synthetic biology. The company is applying recent scientific advances, including newly discovered genetic sequences of novel photosynthetic and metabolic pathways, to execute various life functions within a synthetically devised organism. These breakthroughs present unprecedented opportunities that could restructure and revolutionize industries including energy, industrial organic compounds, pharmaceuticals, CO2 sequestration, fine chemicals, and environmental remediation. Synthetic Genomics, Inc.’s initial focus will be on ethanol and hydrogen production. The company will engineer modular “cassette” based systems to execute specific functions using reprogrammed cells as bio-factories. After leveraging enormous archives of genomic sequence data, the company will integrate novel processes to design, build, and test desired outputs from synthetic organisms. openwetware.mit.edu/index.php?title=Craig_Venter’s_Synthetic_Genomics_plans_to_%22program_cells%22_to_produce_hydrogen [accessed January 4, 2006]. |
|
15 |
Service, R.F. 2004. Nanotechnology grows up. Science 304 (5678): 732-1734. Available online at www.sciencemag.org/cgi/reprint/304/5678/1732.pdf [accessed January 4, 2006]. |
|
16 |
Chang, K. 2005. Tiny is beautiful: translating ‘Nano’ into practical. New York Times (February 22); N. Munro. 2005. How fast can nanotechnology go? National Journal 37(8). |
|
17 |
Moradi, M. 2005. Six opportunities in nano-enabled drug delivery systems. NanoMarkets, LC (February 23). |
|
18 |
The first was Wyeth’s first solid-dose formulation of the immunosuppressant Rapamune® (sirolimus), which received marketing approval from the U.S. Food and Drug Adminstration (FDA) in August, 2000. Rapamune, which was developed to reduce organ rejection in patients who receive kidney transplants, had previously been available only as an oral solution which required refrigeration and mixing with water or orange juice prior to administration. The NanoCrystal-based tablet allows for more convenient storage and administration. As a more recent example of the utilization of this technology, Johnson & Johnson Pharmaceutical Research & Development, L.L.C., is currently conducting a Phase III clinical study of a NanoCrystal-based formulation of the drug paliperidone palmitate for use in patients with schizophrenia. |
|
19 |
Fortina, P. et al. 2005. Nanobiotechnology: The promise and reality of new approaches to molecular recognition. Trends in Biotechnology 23(4):168-173; C.M. Niemeyer and C.A. Mirkin, eds. 2004. Nanobiotechnology: Concepts, Applications and Perspectives, (Wiley). |
|
20 |
Paull, R. et al. 2003. Investing in nanotechnology. Nature Biotechnology 21(10):1144-1147. |
|
21 |
Choi, B. et al. 2005. Artificial allosteric control of maltose binding protein. Physical Review Letters 94(3):038103. |
|
22 |
Georganopoulou, D.G. et al. 2005. Nanoparticle-based detection in cerebral spinal fluid of a soluble pathogenic biomarker for Alzheimer’s disease. Proceedings of the National Academy of Sciences 102 (7):2273-2276. |
|
23 |
www.nanosphere-inc.com/3_media/1_pr/020105.html [accessed February 23, 2005]. |
|
24 |
First described in the mid-1980s, PCR has become the workhorse of biological laboratories worldwide. Researchers and clinicians use the technology to multiply, or copy, specific regions of genomes for use in various types of downstream analyses (e.g., to detect the presence of a specific DNA sequence). |
|
25 |
Mullis, K. 1990. The unusual origin of the polymerase chain reaction. Scientific American 262(4):56-61 and 64-65. |
|
26 |
Saiki R.K., et al. 1985. Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230(4732):1350-1354; Saiki R.K., et al. 1986. Analysis of enzymatically amplified beta-globin and HLA-DO alpha DNA with allele-specific oligonucleotide probes. Nature 324(6093):163-166. |
|
27 |
Castilla, J. et al. 2005. In vitro generation of infectious scrapie prions. Cell 121(2):195-206. |
|
28 |
Carlson, R. 2003. The pace and proliferation of biological techniques. Biosecurity and Bioterrorism: Biodefense Strategy, Practice, and Science 1(3):1-12. |
|
29 |
Eldredge, N. and S.J. Gould. 1972. Punctuated equilibria: An alternative to phyletic gradualism. Models in Paleobiology. Throughout most of the last century, researchers developing the synthetic theory of evolution primarily focused on microevolution, which is slight genetic change over a few generations in a population. Beginning in the early 1970s, this model was challenged by Stephen J. Gould, Niles Eldredge, and other leading paleontologists. They asserted that there is sufficient fossil evidence to show that some species remained essentially the same for millions of years and then underwent short periods of very rapid, major change. Gould suggested that a more accurate model in such species lines would be punctuated equilibrium. |
|
30 |
Carlson, R. 2003. The pace and proliferation of biological techniques. Biosecurity and Bioterrorism: Biodefense Strategy, Practice, and Science 1(3):1-12. |
|
31 |
Craig Venter, briefing to the NSABB, Bethesda, MD on July 1, 2005. Reviewer “J” states that “According to John Mulligan, CEO of Blue Heron Biotechnology, his company has already received a proposal to synthesize a complete bacterial genome. This is technologically feasible at Blue Heron today, which means that the year 2010 may be far too conservative.” |
|
32 |
Tian, J. et al. 2004. Accurate multiplex gene synthesis from programmable DNA microchips. Nature 432(7020):1050-1054. Available online at www.nature.com/nature/journal/v432/n7020/pdf/nature03151.pdf. [accessed January 4, 2006]. |
|
33 |
www.blueheronbio.com/ [accessed January 14, 2005]. |
|
34 |
In the next 5 years the net price for long fragments of chemically synthesized DNA seems *very* unlikely to (i.e., will not) drop below $0.10 per base pair. The |
|
|
$0.01 per base pair number might become possible for the synthesis process itself, but the synthesis number does not include ancillary costs for essential things like handling of intermediate and final materials, sequence verification, and so on. |
|
35 |
Lohmann, V. et al. 1999. Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science 285(5424):110-113. Available online at www.sciencemag.org/cgi/reprint/285/5424/110.pdf [accessed January 4, 2006]. |
|
36 |
Tian, J. et al. 2004. Accurate multiplex gene synthesis from programmable DNA microchips. Nature 432(7020):1050-1054. Available online at www.nature.com/nature/journal/v432/n7020/pdf/nature03151.pdf [accessed January 4, 2006]. |
|
37 |
Couzin, J. 2002. Breakthrough of the year: small RNAs make big splash. Science 298(5602):2296-2297. Available online at http://www.sciencemag.org/cgi/reprint/298/5602/2296.pdf [accessed January 4, 2006]; 10 emerging technologies that will change your world,” Technology Review (February 2004). Available online at www.lib.demokritos.gr/InTheNews/emerging0204.htm [accessed January 4, 2006]. |
|
38 |
Chen, Y. and C. Mao. 2004. Putting a brake on an autonomous DNA nanomotor. Journal of the American Chemical Society 126:8626-8627; Institute of Medicine/National Research Council. 2005. An International Perspective on Advancing Technologies and Strategies for Managing Dual-Use Risks. Washington, DC: The National Academies Press: 49-52. |
|
39 |
Institute of Medicine/National Research Council. 2005. An International Perspective on Advancing Technologies and Strategies for Managing Dual-Use Risks. Washington, DC: The National Academies Press. |
|
40 |
Normile, D., G. Vogel, and C. Holden. 2005. Stem cells: Cloning researcher says work is flawed but claims results stand.” Science 310(5756):1886-1887. Available online at www.sciencemag.org/cgi/reprint/310/5756/1886.pdf [accessed January 4, 2006]. |
|
41 |
Institute of Medicine/National Research Council. 2005. An International Perspective on Advancing Technologies and Strategies for Managing Dual-Use Risks. Washington, DC: The National Academies Press. |
|
42 |
Berg, C. et al. 2002. The evolution of biotech. Nature Reviews. Drug Discovery 1(1):845-846. |
|
43 |
|
|
44 |
Wheelis, M. 2002. Biotechnology and biochemical weapons. The Nonproliferation Review 9(1):48-53. Available online at cns.miis.edu/pubs/npr/vol09/91/91whee.htm [accessed January 4, 2006]. |
|
45 |
Of course, there might be an equivalent concern raised by companies holding this information. |
|
46 |
Carlson, S. 2000. The Amateur Scientist: PCR at Home. Scientific American (July). |
|
47 |
The definition of a bioweapon, while meant to be inclusive, does not extend to nuclear weapons or devices. |
|
48 |
It should be noted that in a precedent that is quite informative for attempts described later in the report to introduce codes of ethics and conduct for biological scientists, computer “hacking” is coming to stand for activities that are remotely done to computers and networks without the consent of those who own and oper- |
|
|
ate those machines and networks, regardless of motivation. The Information Technology community is working to develop an ethic that this is not acceptable, even if there is no malice involved. Some such activity may be playful, or pranks, or done without malice; other hacking causes massive damage without any real intent to do so, and still other such activity is intended to, and succeeds at, causing real damage. But all are illegitimate. |
|
49 |
“Human security” means to protect the vital core of all human lives in ways that enhance human freedoms and human fulfillment … [by] creating political, social, environmental, economic, military and cultural systems that together give people the building blocks of survival, livelihood and dignity. From United Nations Commission on Human Security. 2003. Human Security—Now. Available at www.humansecurity-chs.org/finalreport/English/FinalReport.pdf [accessed February 27, 2006]. |
|
50 |
One of the earliest recorded instances of biological warfare occurred in 600 BC, when the Athenian leader Solon used the noxious roots of the Helleborus plant to poison the water supply in the city of Kirrha. Later, the Greeks and Romans may have used human and animal corpses to poison drinking water wells. And Alexander the Great is thought to have catapulted dead bodies over the walls of besieged cities, possibly as a means of spreading disease and inciting terror among the urban inhabitants. A related technique, used in the Middle Ages, was to deliberately leave dead human or animal corpses behind, in areas that would be occupied shortly by invading troops; catapults were used as well. For further details about these and other later examples of germ-based warfare, including allegations that U.S. government agents deliberately infected the Plains Indians in the 1800s by trading with the Indians smallpox-laden blankets, see National Research Council. 2004. Biotechnology Research in an Age of Terrorism. Washington, DC: The National Academies Press: 34-35. |
|
51 |
Stockholm International Peace Research Institute (SIPRI). 1971. The Rise of CB Weapons. Vol 1. In: The Problem of Chemical and Biological Warfare. New York: Humanities Press. |
|
52 |
Wheelis, M. 1999. Biological sabotage in World War I. in Geissler E. and J.E. Van Courtland Moon, eds. 1999. Biological and Toxin Weapons: Research, Development and Use from the Middle Ages to 1945. SIPRI Chemical and Biological Warfare Studies 18 London: Oxford University Press; 52. |
|
53 |
Redmond, C. et al. 1998. Deadly relic of the great war. Nature 393(6687):747-748. |
|
54 |
Geissler, E., and J.E. Van Courtland Moon, eds. 1999. Biological and Toxin Weapons: Research, Development and Use from the Middle Ages to 1945. SIPRI Chemical and Biological Warfare Studies 18 London: Oxford University Press. |
|
55 |
Bernstein, B. 1988. America’s biological warfare program in the Second World War. Journal of Strategic Studies 11(September): 292-317, especially 304 and 308-310. In addition to Bacillus anthracis and Clostridium botulinum, pathogens studied at Camp Detrick included the causative agents of: glanders; brucellosis; tularemia; melioidosis; plague; smallpox; psittacosis; coccidiomycosis; a variety of plant pathogens including the causative agents for rice blast; rice brown spot disease; late blight of potato; and cereal stem rust. Animal and avian pathogens studied included rinderpest virus, Newcastle disease virus, and fowl plague virus. The |
|
|
Problem of Chemical and Biological Warfare, SIPRI I, London: Oxford University Press, 1971:122. See also Cochrane, R.C. 1947. Biological Warfare Research in the United States. In History of the Chemical Warfare Service in World War II (1 July 1940-15 August 1945), Vol. II (declassified). Historical Section, Office of Chief, Chemical Corps. |
|
56 |
U.S. Department of the Army. 1977. U.S. Army Activity in the U.S. Biological Warfare Programs I. (unclassified) February 23:1-3. |
|
57 |
See Williams, P. and D. Wallace. 1989. Unit 731: The Japanese Army’s Secret of Secrets. London: Hodder and Stoughton: 280-281; and Harris, S.H. 1994. Factories of Death: Japanese Biological Warfare, 1932-45, and the American Cover-Up. London: Routledge. |
|
58 |
Ibid. |
|
59 |
At least 3,000 people, including Chinese civilians, Russians, Mongolians and Koreans, died in the experiments between 1939 and 1945, Chinese state media have said. Outside the site, more than 200,000 Chinese were killed by biological weapons produced by Unit 731, they said. (Reuters, July 18, 2005) |
|
60 |
Guilleman, J. 2001. Anthrax: The Investigation of a Deadly Outbreak. Berkeley and Los Angeles, CA: University of California Press. |
|
61 |
Meselson, M. et al. 1994. The Sverdlovsk anthrax outbreak of 1979. Science 266(5188):1202-1208; Meselson, M. 2001. Note regarding source strength. The ASA Newsletter 87:1, 10-12. |
|
62 |
Dimitri Vladimir Pasechnik was a Soviet microbiologist whose defection to Britain in 1989 disclosed the fact that Moscow’s germ warfare programme was 10 times greater than previously feared in the West. See portal.telegraph.co.uk/news/main.jhtml?view=DETAILS&grid=&targetRule=5&xml=/news/2001/11/29/db2903.xml. |
|
63 |
Kelly, D.C. The trilateral agreement: Lessons for biological weapons verification. In Finlay, T., and O. Meier, eds. 2002. Verification Yearbook 2002. London: VERTIC: 93-109; Domaradskij, I.V. and W. Orent. 2003. Biowarrior: Inside the Soviet/Russian Biological War Machine. Amherst, NY: Prometheus Books. |
|
64 |
Alibek, K. and S. Handelman. 1999. Biohazard: The Chilling True Story of the Largest Covert Biological Weapons Program in the World—Told from the Inside by the Man Who Ran It. New York: Random House. |
|
65 |
For personnel numbers, see Leitenberg, M. 1993. The Conversion of Biological Warfare research and Development Facilities to Peaceful Uses. in Control of Dual-Use Threat Agents: The Vaccines for Peace Programme, SIPRI Chemical and Biological Warfare Series, 15 London, Oxford University Press. For the environmental impacts associated with biological weapons field testing, see Choffnes, E. 2001. Germs on the Loose. The Bulletin of the Atomic Scientists 57(March/April):57-61; Alibek, K. and S. Handelsman. 1999. Biohazard: The Chilling True Story of the Largest Covert Biological Weapons Program in the World—Told from the Inside by the Man Who Ran It. New York: Random House. The Soviet military had tested smallpox. Although Moscow has denied that it ever conducted open-air testing of smallpox, a detailed report prepared by the Monterey Institute of International Studies Center for Nonproliferation Studies asserts that the former Soviet Union did conduct such tests on Vozrozhdeniye Island. For more on this program, see Bozheyeva, G., Y. Kunakbayev, and D. |
|
|
Yeleukenov. 1999. Former Soviet Biological Facilities in Kazakhstan: Past, Present and Future. Center for Nonproliferation Studies, Monterey Institute of International Studies: 6. |
|
66 |
Alibek, K. and S. Handelsman. 1999. Biohazard: The Chilling True Story of the Largest Covert Biological Weapons Program in the World—Told from the Inside by the Man Who Ran It. New York: Random House. |
|
67 |
Gould, D. and P. Folb. 2002. Project Coast: Apartheid’s Chemical and Biological Warfare Programme. Geneva: UNIDR. See also Burgess S. and H. Purkitt. 2001. The Rollback of South Africa’s Chemical and Biological Warfare Program. USAF Counter Proliferation Center. Maxwell Air Force Base, AL: Air War College. |
|
68 |
Institute of Medicine/National Research Council. 2005. An International Perspective on Advancing Technologies and Strategies for Managing Dual-Use Risks. Washington, DC: The National Academies Press: 42-43. |
|
69 |
Ibid. |
|
70 |
Institute of Medicine. 2003. Microbial Threats to Health: Emergence, Detection, and Response. Washington, DC: The National Academies Press. |
|
71 |
Institute of Medicine. 2003. Microbial Threats to Health: Emergence, Detection, and Response. Washington, DC: The National Academies Press. |
|
72 |
Specter, M. 2005. Nature’s bioterrorist. The New Yorker (February 28):50-61. |
|
73 |
For detailed discussions of antigenic drift and shift in influenza A virus, see Krug, R.M. 2003. The potential use of influenza virus as an agent for bioterrorism. Antiviral Research 57(1-2):147-150; and Wright, P.F. and R.G. Webster. 2001. Orthomyxoviruses. In: D.M. Knipe and P.M. Holwey eds, Field’s Virology 4th Ed. Philadelphia: Lippincott Williams & Wilkins: 1533-1579. |
|
74 |
Influenza viruses are defined by two protein components on the virus surface: haemagglutinin (H) and neuraminidase (N). |
|
75 |
Institute of Medicine. 2005. The Threat of Pandemic Influenza: Are We Ready? Washington, DC: The National Academies Press. |
|
76 |
Ibid. |
|
77 |
See www.who.int/csr/disease/avian_influenza/country/cases_table_2006_04_21/en/index.html, [accessed April 25, 2006]. |
|
78 |
Chen, H. et al. 2004. The evolution of H5N1 influenza viruses in ducks in southern China. Proceedings of the National Academy of Sciences 101(28):10452-10457; Institute of Medicine. 2005. The Threat of Pandemic Influenza: Are We Ready? Washington, DC: The National Academies Press; Keawcharoen, J. et al. 2004. Avian influenza H5N1 in tigers and leopards. Emerging Infectious Diseases 10(12):2189-2191. Available online at www.cdc.gov/ncidod/EID/vol10no12/04-0759.htm [accessed March 17, 2005]. |
|
79 |
Finlay, B.B. and S. Falkow. 1997. Common themes in microbial pathogenicity revisited. Microbiology and Molecular Biology Reviews 61(2):136–169. |
|
80 |
Casadevall, A. and L-A. Pirofski. 1999. Host-pathogen interactions: redefining the basic concepts virulence and pathogenicity. Infection and Immunity 67(8):3703-3713. |
|
81 |
Casadevall, A. and L-A. Pirofski. 1999. Host-pathogen interactions: redefining the basic concepts virulence and pathogenicity. Infection and Immunity. 67(8): 3703-3713; Falkow, S. 1997. What is a pathogen? ASM News 63:359–365; Finlay, |
|
90 |
Blaser, M. 1997. Ecology of Helicobacter pylori in the human stomach. Journal of Clinical Investigation 100(4):759–762. |
|
91 |
Staskawicz, B. et al. 2001. Common and contrasting themes of plant and animal diseases. Science 292(22):2285-2289. |
|
92 |
Plotnikova, J.M. et al. 2000. Pathogenesis of the human opportunistic pathogen Pseudomonas aeruginosa PA14 in arabidopsis. Plant Physiology 124(4):1776-1774; Woolhouse, M.E. et al. 2001. Population biology of multihost pathogens. Science 292(5519):1109-1112. |
|
93 |
Reeve, J.N. 1999. Archaebacteria then…Archaes now (are there really no archaeal pathogens?). Journal of Bacteriology 181(12):3613-3617; Eckburg, P.B. et al. 2003. Archaea and their potential role in human disease. Infection and Immunity 71(2):591–596. |
|
94 |
Lepp, P.W. et al. 2004. Methanogenic Archaea and human periodontal disease. Proceedings of the National Academy of Sciences 101(16):6176-6181. |
|
95 |
Worobey, M. 2000. Extensive homologous recombination among widely divergent TT viruses. Journal of Virology 74(16):7666-7670. Available online at www.cdc.gov/ncidod/EID/vol10no12/04-0759.htm [accessed January 4, 2006]. |
|
96 |
Shimono, N. et al. 2003. Hypervirulent mutant of Mycobacterium tuberculosis resulting from disruption of the mce1 operon. Proceedings of the National Academy of Sciences 100(26):15918; Foreman-Wykert, A. and Miller, J.F. 2003. Hypervirulence and pathogen fitness. Trends in Microbiology 11(3):105-108. |
|
97 |
Mouslim, C. et al. 2002. Conflicting needs for a Salmonella hypervirulence gene in host and non-host environments. Molecular Microbiology 45(4):1019-1027. |
|
98 |
Lorange, E.A., et al. 2005. Poor vector competence of fleas and the evolution of hypervirulence in Yersinia pestis. Journal of Infectious Diseases 191(11):1907-1912. |
|
99 |
The Committee recognizes that virulence can evolve to increase or decrease in a pathogen, in response to specific circumstances, such as how the pathogen is transmitted from person to person. |
|
100 |
Kagan, E. 2001. Bioregulators as instruments of terror. Clinics in Laboratory Medicine 21(3):607-618. See also, Wheelis, M. 2004. Will the new biology lead to new weapons? Arms Control Today 34(6):6-13. |
|
101 |
Casadevall, A. and L-A. Pirofski, 1999. Host-pathogen interactions: redefining the basic concepts of virulence and pathogenicity. Infection and Immunity 67(8): 3703-3713; Ingham, H.R. and P.R. Sisson. 1984. Pathogenic synergism. Microbiol. Sci. 1(8):206-208; Janeway, C.A., C.C. Goodnow and R. Medzhitov. 1996. Immunological tolerance: danger—pathogen on the premises! Current Biology 6:519-522. For an easy to read guide on Polly Matzinger’s work on molecular “danger signals” see en.wikipedia.org/wiki/Polly_Matzinger. |
|
102 |
Kagnoff, M.F. and Eckmann L. 1997. Epithelial cells as sensors for microbial infection. Journal of Clinical Investigation 100(1):6-10. |
|
103 |
Bäckhed, F. et al 2005. Host-bacterial mutualism in the human intestine. Science 307(5717):1915-1920; Rakoff-Nahoum, S. et al. 2004. Recognition of commensal microflora by Toll-like receptors is required for intestinal homeostasis. Cell 118(2):229-241. |
|
104 |
Kobayashi, K.S. et al. 2005. Nod2-dependent regulation of innate and adaptive immunity in the intestinal tract. Science 301(5710):731-734; Maeda, S., et al. 2005. Nod2 mutation in Crohn’s disease potentiates NF-kappaB activity and IL- |
|
|
1beta processing. Science 307(5710):734-738; Girardin, S.E. et al. 2003. Lessons from Nod2 studies: towards a link between Crohn’s disease and bacterial sensing. Trends in Immunology 24(12):652-658; Girardin, S.E. et al. 2003. Nod1 detects a unique muroopeptide from gram-negative bacterial peptidoglycan. Science 300(5625):1584-1587; Girardin, S.E. et al. 2003. Nod2 is a general sensor of peptidoglycan through muramyl dipeptide (MDP) detection. Journal of Biological Chemistry 278(11): 8869-8872; Fiocchi, C. 1998. Inflammatory Bowel Disease: Etiology and Pathogenesis. Gastroenterology 115(1):182-205. |
|
105 |
Nixdorff, K. and W. Bender. 2002. Ethics of university research, biotechnology and potential military spin-off. Minerva 40(1):15-35. |
|
106 |
Proliferation: Threat and Response. Available online at www.defenselink.mil/pubs/prolif97/annex.html#technical [accessed February 24, 2005]. |
|
107 |
Block, S. 1999. Living nightmares: Biological threats enabled by molecular biology. In Drell, S.D., A.D. Sofaer, and G.D. Wilson. 1999. The New Terror: Facing the Threat of Biological and Chemical Weapons. Stanford, CA: Hoover Institution Press: 39-75. |
|
108 |
Krug, R.M. 2003. The potential use of influenza virus as an agent for bioterrorism. Antiviral Research 57(1-2):147-150. |
|
109 |
Kobasa, D. et al. 2004. Enhanced virulence of influenza A viruses with the haemagglutinin of the 1918 pandemic virus. Nature 431:703-707. |
|
110 |
Tumpey, T.M. et al. 2005. Characterization of the reconstructed 1918 Spanish influenza pandemic virus. Science 310(5745):77-80; Taubenberger, J.K. et al. 2005. Characterization of the 1918 influenza virus polymerase genes. Nature 437(7060):889-893. Available online at www.nature.com/nature/journal/v437/n7060/full/nature04230.html [accessed January 4, 2006]. |
|
111 |
Ibid. |
|
112 |
Institute of Medicine/National Research Council. 2005. An International Perspective on Advancing Technologies and Strategies for Managing Dual-Use Risks. Washington, DC: The National Academies Press; also Bokan, S. et al. 2002. An evaluation of bioregulators as terrorism and warfare agents. ASA Newsletter 02-3(90):1. Available online at www.asanltr.com/newsletter/02-3/articles/023c.htm [accessed January 4, 2006]; Kagan, E. 2001. Bioregulators as instruments of terror. Clinics in Laboratory Medicine 21(3):607-618. |
|
113 |
In determining whether to list a biological agent, the Secretary of HHS, in consultation with scientific experts representing appropriate professional groups, is required to consider the agent’s effect on human health, its degree of contagiousness and methods by which the agent is transferred to humans, and the availability of immunizations and treatments for illnesses that may result from infection by the agent. The list was initiated in 1997, when the Antiterrorism and Effective Death Penalty Act of 1996 required the Secretary of HHS to establish and enforce safety procedures for the transfer of listed biological agents (select agents), including measures to ensure proper training and appropriate skills to handle such agents, and proper laboratory facilities to contain and dispose of such agents. An expanded list of pathogens and toxins went into effect on February 11, 2003. Agricultural plant and animal pathogens are now also included; other changes reflect taxonomic changes and a few reassessments of what constitutes the most dangerous biothreat agents. |
|
114 |
See Table 2-2 in National Research Council. 2004. Biotechnology Research in an Age of Terrorism. Washington, DC: The National Academies Press; 54-57. |
|
115 |
Our current biosafety system and select agents lists are mostly concerned with full systems or whole organisms. But as we start to construct new things via the combination of many functions in novel ways, the current scheme will not scale. Although beyond the scope of this study, governments and regulatory bodies may need to consider whether or not a biosafety system that is based at the “parts” level might be more useful. |
|
116 |
Meselson, M. 2000. Averting the hostile exploitation of biotechnology. The CBW Conventions Bulletin.48(June): 16-19. |
|
117 |
Information about current biological weapons capabilities summarized in Squassoni, S. 2004. Nuclear, biological, and chemical weapons and missiles: Status and trends. CRS Report for Congress, July 2 (RL30699); National Research Council. 2004. Biotechnology Research in an Age of Terrorism. Washington, DC: The National Academies Press. |
|
118 |
See www.opbw.org/ [accessed October 28, 2004]. |
|
119 |
Theodore Kaczynski, the Unabomber, was charged in 1998 with making and delivering four bombs that killed two men and maimed two scientists. In all, Mr. Kaczynski was alleged to have killed three people and injured 29, in 16 attacks between 1978 and 1995. See www.cnn.com/US/9805/04/kaczynski.sentencing/index.html [accessed January 4, 2006]. |
|
120 |
Hacking the genome. 2003/2004; 2600 The Hacker Quarterly 20(4), Winter 2003/2004. |
|
121 |
“National security experts and even … biologists themselves are concerned that rogue scientists could create new biological weapons—like deadly viruses that lack natural foes. They also worry about innocent mistakes: organisms that could potentially create havoc if allowed to reproduce outside the lab.”… [W]e live in an age that many tools and technologies can be turned into weaponry,” said Laurie Zoloth, a bioethicist at Northwestern University. “You always have the problem of dual use in every new technology.” See Elias, P. 2005. Light-sensitive bacteria used to create pictures:UCSF Scientists Make Living Film. Associated Press, November 24. Available online at www.montereyherald.com/mld/montereyherald/business/technology/13251114.htm?template=contentModules/printstory.jsp, [accessed January 4, 2006]. |
|
122 |
Much of this discussion draws from Chyba, C.F. 2002. Toward biological security. Foreign Affairs 81(3):122-136; and Chyba, C.F. and A.L. Greninger. 2004. Biotechnology and bioterrorism: An unprecedented world. Survival 46(2):143-162. |
|
123 |
Weapons of Mass Effect (i.e., truck bombs or hijacked airliners) are used, as Time magazine says, “to cause great loss of life and spread chaos and despair” among the populace. See www.worldnetdaily.com/news/article.asp?ARTICLE_ID=24804 [accessed June 14, 2005]. |
|
124 |
Weiss, L. 2003. Atoms for peace. Bulletin of the Atomic Scientists November/December:34-44. |
|
125 |
And because these nuclear materials advertise their presence by emitting various distinctive signatures as radioactive emissions from the source. |
|
126 |
Many states attached reservations to their instruments of ratification that |
|
|
had the effect of making this protocol an agreement to only ban first use, not retaliation. |
|
127 |
Chyba, C.F. and A.L. Greninger. 2004. Biotechnology and bioterrorism: An unprecedented world. Survival 46(2):143-162. For a more in-depth discussion of this point see, National Research Council. 2004. Biotechnology Research in an Age of Terrorism. Washington, DC: The National Academies Press. |
|
128 |
Meselson proposal to make use of biological weapon a crime against humanity; A Draft Convention to Prohibit Biological and Chemical Weapons Under International Criminal Law. The Draft Convention and a discussion about the need for such a convention may be found at www.sussex.ac.uk/Units/spru/hsp/CRIMpreambleFeb04.htm [accessed January 4, 2006]. |
|
129 |
It should be noted that a biological “arms race” is between protective measures and malevolent applications of potentially benevolent technologies, rather than between protective measures and offensive weapons programs. The protective technologies that are developed in such a competition are very unlikely to be classified (for all the reasons described) and hence may enable malicious applications of that same technology. This means that it is difficult for defensive applications to win, and bears on the question (which should be discussed to a greater extent) of whether defense can win an offense-defense competition. For a discussion of “can defenses run faster than offenses,” see the section with that name, 17-19 of Epstein, G.L. 2005. Global Evolution of Dual-Use Biotechnology: A Report of the Project on Technology Futures and Global Power, Wealth, and Conflict. Center for Strategic and International Studies. |
|
130 |
Some of the implications of creating a regime of “sensitive” information are discussed in Epstein, G.L. 2001. Controlling biological warfare threats: Resolving potential tensions among the research community, industry, and the national security community. Critical Reviews in Microbiology, 27(4):321-354, especially pp. 347-348. This analysis was extended in a presentation given to the Committee on June 23, 2004 titled “Sensitive Information in the Life Sciences.” A presentation very similar to that one, and available online, was delivered at the International Forum on Biosecurity in Lake Como on March 21, 2005 and can be found online at www7.nationalacademies.org/biso/Biosecurity_Epstein_2.0.ppt [accessed January 4, 2006]. |
|
131 |
Institute of Medicine/National Research Council. 2005. An International Perspective on Advancing Technologies and Strategies for Managing Dual-Use Risk. Washington, DC: The National Academies Press. |
|
132 |
Daar, A.S. et al. 2002. Top 10 biotechnologies for improving health in developing countries. Nature Genetics 32:229-232. |
|
133 |
We are mindful, however, that crops are indeed monocultures and thus exquisitely sensitive to epidemics of the next “new” fungus or virus; they usually require a lot of water (increasingly scarce in our world) and fertilizer (increasingly expensive and polluting); they are often disruptive of local social structures. As an exercise in practical GM crops, consider the lessons of Lansing, J. S. 1991. Priests and Programmers: Technologies of Power in the Engineered Landscape of Bali. Princeton, NJ: Princeton Univ Press. |
|
134 |
Hoyt, K. and S.G. Brooks. 2003/2004. A double-edged sword. International Security 28(Winter):123-148. |
|
135 |
Jackson, R.J. et al. 2001. Expression of mouse interleukin-4 by a recombinant ectromelia virus suppresses cytolytic lymphocyte responses and overcomes genetic resistance to mousepox. Journal of Virology 75(3):1205-1210. |
|
136 |
Cello, J. et al. 2002. Chemical synthesis of poliovirus cDNA: generation of infectious virus in the absence of natural template. Science 297(5583):1016-1018. |
|
137 |
Rosengard, A.M. et al. 2002. Variola virus immune evasion design: Expression of a highly efficient inhibitor of human complement. Proceedings of the National Academy of Sciences 99(13):8808-8813. |
|
138 |
Fraser, C.M. and D.R. Dando. 2001. Genomics and future biological weapons: The need for preventive action by the biomedical community. Nature Genetics 29(3):253-256. See, also, National Research Council. 2004. Seeking Security: Pathogens, Open Access, and Genome Databases. Washington, DC: The National Academies Press. |
|
139 |
Arora, N. and S.H. Leppa. 1994. Fusions of anthrax toxin lethal factor with shiga toxin and diptheria toxin enzymatic domains are toxic to mammalian cells. Infection and Immunity 62(11):4955-4961. |
|
140 |
Broad, W.J. 1998. Gene-engineered anthrax: Is it a weapon? New York Times (February 14): A4; Wade, N. 1998. Tests with anthrax raise fears that American vaccine can be defeated. New York Times (March 26). |
|
141 |
Aldous, P. 2001. Biologists urged to address risk of data aiding bioweapon design. Nature 414(6861): 237-238 as cited in Zilinskas, R.A. and J.B. Tucker. 2002. Limiting the contribution of the open scientific literature to the biological weapons threat. Journal of Homeland Security. Available online at www.homelandsecurity.org/journal/Articles/tucker.html [accessed December 30, 2005]. |
|
142 |
Hatta, M., et al. 2001. Molecular basis for high virulence of Hong Kong H5N1 influenza A virus. Science 293(5536):1840-1842. |
|
143 |
Gibbs et al. 2001. Recombination in the Hemagglutinin Gene of the 1918 ‘Spanish Flu.’ Science 293(5536):1842-1845. |
|
144 |
Tumpey, T.M. et al. 2005. Characterization of the Reconstructed 1918 Spanish Influenza Pandemic Virus. Science 310(5745):77-80; Taubenberger, J.K. et al. 2005. Characterization of the 1918 influenza virus polymerase genes. Nature 437:889-893. |
|
145 |
Boyce, N. 2002. Flu’s Worst Season. US News and World Report 133(6):50. |
|
146 |
Wang, X.G. et al. 2001. Purified Cholera Toxin B Subunit from Transgenic Tobacco Plants Possesses Authentic Antigenicity. Biotechnology and Bioengineering 72(4):490-494. |
|
147 |
Boyce, N. 2002. Should Scientists Publish Work that Could be Misused? US News and World Report 132(22): 60. |
|
148 |
Wein, L.M. and Y. Liu. 2005. Analyzing a bioterror attack on the food supply: The case of botulinum toxin in milk. Proceedings of the National Academy of Sciences 102(28):9984-9989. |
|
149 |
Berg, P. et al. 1975. Summary statement of the Asilomar Conference on recombinant DNA molecules.Proceedings of the National Academy of Sciences 72(6): 1981-1984. |
































































