2
Beyond the Gut: Insights from Other Host-Microbe Systems
OVERVIEW
Equally compelling but less studied than the host-microbe environment of the human gut, microbial communities in plants, insects, and the soil are also likely to yield important scientific and medical insights. The two papers in this chapter offer intriguing perspectives on nonhuman host-microbe systems, beginning with a description of the microbial communities that exist on and inside plants and how these microbial communities contribute to protecting their hosts from infectious disease. The first paper is also coauthored by workshop presenter Jo Handelsman.
Contending that the advancement of infectious disease research lies in understanding the nature of cooperation within microbial communities and its contribution to host health, the authors lead a guided tour of insights, derived from studies of plant pathology, that raise the possibility of harnessing the natural microbial communities of humans for disease prevention. She notes that many microbiological milestones—among them the germ theory of disease and the discovery and characterization of viruses—were first achieved by plant biologists, but were not recognized at the time by their peers in other disciplines. Thus, while it is not surprising that studies of plants and their associated microbial communities have added considerably to knowledge of host-microbe relationships, these findings have not been widely appreciated, nor have they been well integrated with current understanding of the human gut microbiota.
Handelsman and coworkers have demonstrated that intermicrobe communications that lead to disease could be disrupted, and that beneficial lines of communication could be protected against pathogenic saboteurs. For example, they
observed that plant diseases can be suppressed by treatments that modify the microbial community of the root to make it more like the community in the soil, a conclusion which they have dubbed the “camouflage hypothesis.” By analogy, the human intestinal microflora may influence the success of pathogens by either presenting a barrier to invasion that is predicated on the composition of the entire community, or conversely by potentiating activity, facilitating infection, or aggravating disease symptoms.
The second contribution to this chapter, by plant biologist Brian Staskewicz, describes similarities among the strategies used by plant and animal pathogens and compares the defenses mounted against them by their disparate hosts. The complexity of disease resistance in plants is illustrated through Staskewicz’s description of his laboratory’s efforts to describe the function and regulation of a key disease resistance protein in the plant Arabidopsis thaliana, in response to the pathogenic bacterium, Pseudomonas syringae.
Conserved cellular defense responses in plants resemble certain innate immune responses to pathogens in vertebrates and insects, suggesting that these defense pathways may be inherited from a common ancestor. The preponderance of conserved motifs and, presumably, mechanisms among plant and animal proteins involved with innate immunity has encouraged communication and even collaboration among the scientists who study these systems in widely different species—an unfortunately rare occurrence that may yield significant insights on the structure, function, and evolution of innate immunity.
IT TAKES A VILLAGE: ROLE OF INDIGENOUS MICROBIAL COMMUNITIES IN INFECTIOUS DISEASE
Christina Matta and Jo Handelsman1
Summary
The microbial communities that reside on and inside plants and animals are a key to host health. In addition to contributing to digestion and nutrition, they present a formidable barrier to pathogens, which may invade the community or arise from it. The future of infectious disease research needs to focus on ways to enlist the natural community in prevention of disease rather than solely on the traditional warfare model, which requires direct killing of the pathogen with a chemical agent. The future of this field will depend on insights into the nature of cooperation within communities and the features that make them resistant to invasion. In these areas, the field of plant pathology, or infectious disease of plants, has much to offer. Plant pathologists have long recognized the role of the micro-
bial community in host health. But biomedical research in the late 20th and early 21st century has tended to ignore recent developments in plant pathology that may enrich medical understanding of human disease. Examination of 19th and early 20th century microbiology illustrates the power of communication between scientists who study plant and human diseases. In fact, the development of the germ theory and experimental evidence for it were predicated on an intimate discussion among scientists who studied plants and humans. For both the plant sciences and biomedicine to benefit from each other’s expertise, we need to rebuild the ties and foster collaboration between plant pathology and human infectious disease.
Crossing the Disciplinary Divide: Microbiology in Historical Perspective
The role plant sciences have played in shaping the content and methods of modern bacteriology has been overshadowed by the popular appeal of medical breakthroughs. Indeed, to the lay public, the very terms microbiology and bacteriology, are almost synonymous with contagious disease of humans. Likewise, medical microbiologists have tended to overlook their discipline’s debts to botany, plant pathology, soil science, and the agricultural sciences. Historically, however, medical and nonmedical microbiology were enmeshed in an intricate, subtle relationship based upon shared methodologies that were equally applicable in both medical and nonmedical contexts. In many instances, breakthroughs heralded as triumphs of medical research echoed knowledge or practices that had been standard or assumed in the plant sciences for years or decades. The relevance of plant research to microbiology more broadly—something that biologists of all specialties actively recognized and drew upon even into the mid-20th century—has largely faded from view.
Cohn, Koch, and the Germ Theory
Plant pathology has historically offered novel approaches for investigating disease causality and pathogenesis that have proven crucial to the understanding of human and animal infectious disease. From Mathieu du Tillet’s extensive epidemiological studies of wheat smut in 1755, to de Bary’s thorough demonstration in 1863 that Phytophthora infestans was without a doubt the cause of late blight of potato, to USDA plant pathologist Erwin Frink Smith’s insistence that crown gall of tomatoes could provide a model for understanding tumor growth in human cancers, plant scientists have repeatedly anticipated medical methods and discoveries (Campbell et al., 1999).
Even Robert Koch, who is generally credited as the architect of the germ theory (Brock, 1988), worked within an intricate web of knowledge derived by botanists, bacteriologists, and mycologists. In the 19th century, changes in academic botany in Europe led to the inclusion of bacteria with other microscopic
organisms such as algae and fungi—organisms traditionally considered plants. It is no surprise, then, that the most prominent of the early bacteriologists was a botanist: Ferdinand Cohn, professor of botany at the University of Breslau (now Wroclaw, Poland). (Figure 2-1) As early as 1872, Cohn had presented strong evidence that bacteria cause disease when he published his observations of bacteria in diseased organisms and the loss of disease-causing ability when an infectious fluid was filtered. Moreover, Cohn’s taxonomic and morphological study of bacteria provided the observations necessary to quell continuing debates about pleomorphism, a doctrine that posited that all bacteria were a single species and that the shape of a bacterium was either determined by the organism’s stage in its life cycle or by environmental conditions (or some combination of both). If the doctrine of pleomorphism were correct, specific causality of disease would not be possible; if distinct bacterial species did not exist, then it would be impossible to identify a particular set of disease symptoms with one particular organism. Cohn amassed enough morphological, developmental, and physiological data on various species of bacteria to create a taxonomic system with four “tribes” of bacteria, which contained genera that were differentiated based upon both physiological and morphological criteria (Cohn, 1872).
Without Cohn’s careful observations and focused arguments against pleomorphism, the germ theory of the 1870s and 1880s could not have become the basis for our current understanding—that contagious diseases are each caused by a unique microorganism. It was, in fact, Cohn to whom a then-unknown physician named Robert Koch turned for an expert opinion on his demonstration of the life cycle of Bacillus anthracis.
Some microbiologists have argued that Cohn’s claims about monomorphism
FIGURE 2-1 Left to right: Ferdinand Julius Cohn (1828–1898), Heinrich Anton de Bary (1832–1888), and Robert Koch (1843–1910).
SOURCE: Handelsman (2005).
could not have been confirmed until after Koch published his pure culture techniques (Brock, 1961), but this assessment overlooks a very important detail: that pure culture techniques had been standard practice in mycology at least since the late 1860s when mycologist Oskar Brefeld proposed a set of techniques for cultivating fungi on sterile media and in pure culture. It was these methods that botanist and developmental mycologist Anton de Bary, for example, used in his studies of plant pathogens such as Phytophthora infestans and Puccinia graminis. Furthermore, botanist Joseph Schroeter (one of Cohn’s students) had begun experimenting with obtaining pure cultures of pigmented bacteria as early as 1870. By the time Koch published his methods for studying pathogenic organisms in culture in 1881, then, methods for isolating individual organisms and reintroducing them to host tissue were widespread in mycology and already under development in bacteriology.
A Mosaic from Tobacco to Polio
By far the most notable example of research in plant science that profoundly influenced the development of medical science is tobacco mosaic virus (TMV). In 1892, Russian botanist Dmitri Ivanowski published his observations that filtering sap from tobacco plants infected with tobacco mosaic disease did not prevent the sap from infecting other plants, even when he used the filtration techniques developed by Charles Chamberlain that were so effective in removing pathogenic bacteria from media (Ivanowski, 1892). Six years later, the Dutch soil microbiologist and plant pathologist Martinus W. Beijerinck attributed the symptoms to an unfilterable, noncellular, infectious particle that he termed a virus (Beijerinck, 1898). In 1901, U.S. Army physician Walter Reed echoed this conclusion when he pronounced yellow fever as nonbacterial, arguing instead that it was caused by a filterable agent.
The exact particle involved in TMV remained unidentified until 1935, when biochemist Wendell Stanley purified the virus and isolated its crystalline form. Stanley’s work provided the first physical evidence of a material infectious agent, and as a result, TMV became an experimental model for early research in virology. Both plant scientists and medical microbiologists recognized that the results of research on TMV and the methods used therein could be applied to other viral diseases. Stanley himself used techniques he had developed for working with TMV in his research on influenza. More important, however, Stanley regarded TMV as a means of unlocking the secrets of poliovirus. In the late 1940s and early 1950s, his laboratory used TMV as a standard against which to compare the viral structures, mutations and corollary amino acid changes, and host-viral interactions of the three types of poliovirus then recognized (Creager, 2002). The methods of crystallization and purification that Stanley had pioneered using a plant virus were crucial to understanding the polio epidemic of the mid-20th century.
Communication Breakdown Between Independent Universes
Since the 1950s, communication between the plant sciences (and related fields such as soil microbiology) and biomedicine has disintegrated, even when the questions that engage practitioners in both areas are the same. The breakdown of this interdisciplinary exchange has led biomedical scientists to overlook recent breakthroughs in the plant sciences that might advance their own scientific progress.
Trapping Promoters on Two Continents
In 1988, for example, Michael Daniels’ group at the John Innes Institute published their elegant method for isolating promoters from the plant pathogen Xanthomonas campestris, which involved cloning fragments of the X. campestris genome upstream of a promoterless tetracycline resistance determinant (Osbourn et al., 1987). He inoculated the clone library into plants that were grown in a solution containing tetracycline, ensuring that only those clones that carried a promoter that was induced inside the plant would survive. Daniels’ paper pre-dated the introduction of IVET, or in vivo expression technology, into the microbiological literature by six years, but the methods were essentially the same, except that the host was a plant rather than an animal (Figure 2-2). In the first IVET paper, the Mekalanos group at Harvard cloned fragments of the Salmonella typhimurium genome upstream of a promoterless purA gene in a purine auxotroph and injected the resulting library into mice, which provide a sufficiently purine-limited environment that selects for prototrophic strains (Mahan et al., 1993); therefore, the bacterium would only survive if the DNA upstream of the purA gene carried a promoter that was active inside the animal. The surviving clones could then be retrieved from the spleens of infected mice, and the promoters and genes they regulate could be identified in S. typhimurium, from which they arose. This powerful strategy has been applied to many host-microbe relationships and has provided sweeping insights into mutualistic and pathogenic relationships, yet despite Daniels’ priority of publication, his paper is only rarely cited in papers about IVET or promoter trapping (Brown and Allen, 2004).
Protection, Probiotics, and Biocontrol
Probiotics, or consumable preparations of live bacterial cultures, have become fashionable in health food stores, in food animal production facilities, and research laboratories as a means of preventing or curing numerous ailments in animals and humans. Explanations for the benefits of such preparations have, at their core, two proposed mechanisms: first, that the probiotic strains outcompete detrimental bacteria at the site of pathogen infection, thus preventing the onset of disease symptoms. The second mechanism claims that probiotics induce a systemic response in the host that enhances resistance to infection. Yet neither of
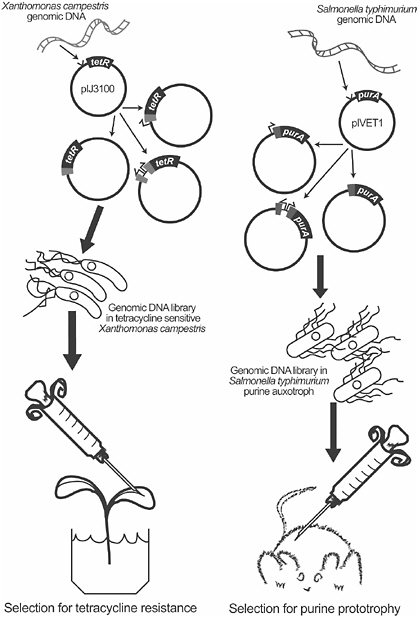
FIGURE 2-2 Schematic depiction of the first strategies to select in vivo for promoters that are induced by plant (left panel) or animal (right panel) hosts (Mahan et al., 1993; Osbourn et al., 1987). The strategies both use a chemical condition in the host (tetracycline or low availability of purines) to select for clones that carry host-inducible promoters. The scheme on the right is a simplified version of that published by Mahan et al. (1993); it does not show, for example, the integration of the vector into the chromosome, which was an aspect of the original strategy that differentiated it from the Osbourn et al. method (1987).
SOURCE: Handelsman (2005). Figure 2-2 was created with the help of Jo Handelsman’s graduate student, Zakee Sabree.
these mechanisms is new to probiotics; they are both established mechanisms that govern the successful biological control (or biocontrol) of plant disease. Biocontrol agents, in the form of microbial inoculants applied to seed, soil, or leaves, have been used to enhance plant health and agricultural yield since the early 20th century. These agents typically work by suppressing a disease—some biocontrol strains colonize plant surfaces and exclude pathogens by competitive or antagonistic strategies such as occupying desirable niches or reducing pathogen populations through secretion of antibiotics. Others incite systemic acquired resistance, an induced response that makes the entire plant less susceptible to infection. Although both past and present work on biological control provides rich data that could inform the much younger field of probiotics, the potential for integration of these fields has not been recognized in current literature on probiotics.
Because of the breakdown in communication, therefore, knowledge developed within the plant sciences community has not been adopted by the biomedical community as successfully as in the 19th and early 20th centuries. Given the frequency with which conceptual change in the plant and agricultural sciences has enriched—and, often, preceded—similar developments in biomedicine, overlooking recent breakthroughs in the former can only be detrimental to the future growth of biomedicine. Of special importance at present is the seminal work in plant pathology and microbial ecology that considers the role of microbial communities in infectious disease.
Role of Communities in Infectious Disease
Pathogens live in a microbiological stew. When between hosts, they must survive in the environment among competitors who garner their scarce resources. When they contact a host, pathogens often must breach a protective microbiological barrier to reach the host tissue and induce disease. Pathogens frequently constitute part of this protective barrier, making a switch from commensal to pathogen whose triggers and governors are poorly understood.
The study of pathogenesis has traditionally ignored the microbial context of disease. It is time to examine the role of the microbial communities that live on and in host plants and animals to identify factors that reduce the effectiveness of host-associated communities as barriers and that incite commensals in these communities to turn on their host and cause disease. Plant disease, more often than human disease, has been studied in a microbial context, so there is much to learn from plant pathology that may affect our understanding of pathogenesis in humans.
Suppressive Soils, Biocontrol, and Latent Pathogens
The study of plant pathogens has typically included attention to microbial communities, in part because some attempts to assign responsibility for an effect
to a single species have been dismal failures, indicating that mechanisms of disease suppression include coordination of responses in a community. Suppressive soils are an excellent case in point. It has long been observed that after growing a crop at the same site for many consecutive years, the frequency of certain diseases decreases. The soil becomes “suppressive” to that disease. Although the effect is demonstrably of biotic origin, disappearing upon biocidal treatment and spreading to other soils when a small soil inoculum is transferred to a new site, attempts to isolate a single organism responsible for the suppressiveness have been unsuccessful (Baker and Cook, 1974). Analysis of microbial communities in suppressive soils indicates that they differ from nonsuppressive soils in composition and structure, and certain disease-suppressive microorganisms have been isolated, but the exact complement of organisms required for suppressiveness has not been identified in most cases.
We found that introduction of Bacillus cereus, a biocontrol agent that suppresses root diseases of alfalfa and soybean plants caused by oomycete pathogens, altered the structure of the community associated with soybean roots, which is known as the rhizosphere community (Gilbert et al., 1993). Whether this community change is required for disease suppression, or just associated with it, remains unknown, but some features of the community change are noteworthy. First, B. cereus induces a dramatic increase in the proportion of the community that is represented by Cytophaga johnsoniae, a member of the Cytophaga/Flavobacterium group of the Bacteroidetes phylum (Gilbert et al., 1993). When B. cereus is isolated from field-grown roots, it frequently carries along with it “hitchhikers,” what are difficult to separate from it initially. Interestingly, these coisolates are most often members of the Cytophaga/Flavobacterium group, and B. cereus provides a growth benefit for them under some conditions (Dunn et al., 2003).
In addition to the increase in representation of the Cytophaga/Flavobacterium group, B. cereus induces a global change in composition of the rhizosphere community that causes it to resemble the community in raw soil more than a typical rhizosphere. This finding led us to suggest the “camouflage hypothesis,” which posits that microbial communities can prevent detection of a root by pathogens by altering the chemical composition of root exudates, making the root resemble soil, thereby “camouflaging” the root to resemble its surroundings (Gilbert et al., 1994). Many results reported in the first half of the 20th century are consistent with the camouflage hypothesis. For instance, biocontrol agents, plant disease resistance genes, and certain cultural practices have all been shown to be associated with reduction of disease as well as a change in the rhizosphere community that makes it resemble the soil more than the typical rhizosphere (Gilbert et al., 1993, 1996). As the study of plant disease became more reductionist in the latter half of the 20th century, attention to microbial communities decreased and a causal relationship between the composition of the rhizosphere community and disease suppression was not established. The camouflage hypothesis is attractive
because a broad-based mechanism of disease suppression that is founded upon the contributions of many organisms that comprise a community might be more robust to the evolution of pathogens than are disease resistance genes and synthetic pesticides.
The camouflage hypothesis might have parallels in human infectious disease. The gut community, for example, may govern pathogen success by presenting a barrier to invasion that is predicated on the composition of the entire community, not one species. Alternatively, members of the natural community may potentiate the activity of the pathogen, facilitating infection or aggravating disease symptoms.
Future Focus
Understanding both plant and human infectious disease will be enhanced by understanding the communities in which the pathogens must function. The following key questions about community-pathogen interactions are especially crucial to constructing a comprehensive understanding of disease.
Community Robustness
To induce disease, many pathogens must colonize their hosts. If the tissue they infect carries a normal microflora, then the pathogen must invade the community on the host to cause disease. Little is understood about the nature of community robustness (resistance to and recovery from change) or the basis for pathogen invasiveness of communities. Identifying the components of community structure (species richness, functional redundancy, abundance of certain members) that contribute to robustness and the genes that make pathogens invasive will advance the understanding of the role of communities in disease (Handelsman et al., 2005).
Commensal-Pathogen Switch
Many pathogens reside peacefully in the communities that colonize infection sites on healthy host tissue and are triggered by certain cues to incite disease. Perturbation of the community, changes in host resistance, or convergence of physical or chemical features of the environment (such as availability of certain dietary components) to stimulate pathogen population or alter gene expression in the pathogen may be responsible for the switch from commensal to pathogen. An understanding of community structure and function that accounts for the effects of such changes will provide the basis for building quantitative models that predict disease events.
Microbial Cooperation
Little is known about polymicrobial diseases, but there is no doubt that in both plants and animals, certain diseases require the presence of two or more species. For example, a particular sequence of invasion events is required for biofilm formation on teeth. Certain bacteria require the prior establishment of other species in order to colonize (Kolenbrander et al., 2002). Although Streptococcus mutans is directly responsible for tooth decay, other members of the community are required for its establishment. In plant disease, similar synergies are abundant. Verticillium wilt disease of potatoes, for example, is substantially intensified on potato plants when their roots are infected by the root lesion nematode Pratylenchus penetrans (MacGuidwin and Rouse, 1990).
Mathematical Models
Predictive models that integrate biological and physical characteristics of the community are needed to generate principles of microbial ecology. Models that predict invasion events in many different communities will reveal the variables that govern robustness, invasion, and microbial cooperation, thereby forming the basis for predicting outcomes in new experimental or biological landscapes. Modeling has traditionally provided the basis for establishing broad principles in areas of biology as diverse as enzyme kinetics and macroecology (Anderson and May, 1986; Berg, 1983; Levin et al., 1989; Michaelis and Menten, 1913). Similarly, it is such models that will furnish microbial ecology with the power to revolutionize approaches to avoiding, preventing, treating, and curing infectious disease of plants and animals.
Conclusion
Much is made these days of interdisciplinary science that spans many fields, often uniting the biological and physical or mathematical sciences. These efforts have yielded extraordinary collaborations and insights into the natural world. But collaborations among closely related fields also remain powerful. The history of communication and collaboration between plant pathologists and medical microbiologists is a testament to this power. Such interactions contributed to the development of pure culture techniques, establishment of the germ theory of disease, and understanding of viral structure and function among many other great discoveries that punctuate the early history of microbiology. The more recent division between the fields has led to embarrassing ignorance at its best and wasteful delays in progress at its worst.
As one final case in point, consider the career of Selman Waksman. Waksman was trained as a soil microbiologist and considered himself an agricultural scientist throughout his career. Yet, through his discovery of streptomycin, produced by actinomycetes from soil, he made one of the most significant advances in
medicine in the 20th century, for which he was awarded the 1952 Nobel Prize in Medicine. Waksman expressed great satisfaction in the fact that the National Academy of Sciences created divisions for both of his two great loves, agriculture and microbiology, during his lifetime (Waksman, 1958).
Waksman’s discovery came toward the end of the Second World War—the first war in which there were fewer deaths from infection than from battle itself—because of the availability of antibiotics. Similarly, in the 21st century, we now face medical challenges of daunting proportions. An impending flu pandemic, the spread of antibiotic resistance among bacterial pathogens, and the recognition of the role of bacteria in far more conditions than we ever anticipated all present us with an opportunity. Plant pathology has addressed the role of microbial communities in plant disease, host genes that mediate resistance to disease, and environmental factors in development of disease epidemics. And as the United States confronts the potential for human epidemics derived from natural events or bioterrorism, we need to be equally cognizant of the vulnerability of our food supply to pathogens and microbial toxins. Perhaps the urgency of preparing for dire events will encourage the cooperation between these two fields that have so much to share.
THE MOLECULAR BASIS OF BACTERIAL INNATE IMMUNITY IN ARABIDOPSIS THALIANA
Brian Staskawicz
University of California, Berkeley
As Jo Handelsman has described in her contribution to this workshop (see the previous paper), plant disease epidemics have changed the course of human history on many occasions. One such epidemic, a rust that wiped out coffee plantations in British colonies, turned the country into a nation of tea drinkers. Today, the interactions of plants and infectious diseases—the major interest of my laboratory—are studied in model systems such as Arabidopsis thaliana, a plant that researchers have developed as a sort of “fruit fly” for plant genetics. Figure 2-3 depicts Arabidopsis, as well as rice, the plant that feeds most of the world, and the various classes of pathogens known to infect these plants, and indeed, all other higher eukaryotes.
Arabidopsis is a member of the Brassica family, which also includes such familiar vegetables as broccoli and cauliflower. There are many advantages to using Arabidopsis as a model system for studying plant-pathogen interactions. The plant completes a life cycle, from seed to seed, within seven weeks, and its entire genome has been sequenced. It is easily transformed (to produce a transgene) with Agrobacterium tumefaciens, known as nature’s genetic engineer due to its facility for introducing DNA into plant (and yeast) cells. Arabidopsis transformation is performed by soaking the plant, while in flower, in a beaker contain-
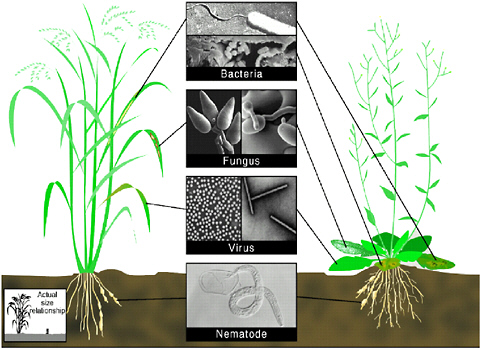
FIGURE 2-3 A rice plant (left) and Arabidopsis thaliana (right), a model plant for host-pathogen interactions. The establishment of numerous pathosystems in the genetically tractable plant species A. thaliana leads to rapid identification of components of host resistance and defense signaling pathways. Within each group, related bacterial, fungal, viral, and nematode pathogens cause diseases in both rice and Arabidopsis. Scanning electron micrographs (center panels) and disease reaction phenotypes of representative phytopathogens of Oryza and Arabidopsis are shown. The rice bacterial pathogen X. oryzae pv. oryzae causes chlorotic water-soaked stripes on rice leaves and lesions on Arabidopsis leaves. The bacterial pathogen P. syringae induces small water-soaked chlorotic lesions on Arabidopsis. The fungus Erysiphe cihoracearum causes powdery mildew disease on Arabidopsis. The most important fungal pathogen of rice is Magnoportha grisea, which produces gray necrotic lesions on all parts of the shoot. Tobacco mosaic virus infects and spreads throughout the Arabidopsis plant with few detectable symptoms. The spherical form of rice tungro virus causes yellow discoloration of the leaves. The plant parasitic nematode infects and causes disease in both rice and Arabidopsis.
SOURCE: Baker et al. (1997).
ing genetically engineered Agrobacterium in a liquid medium. Seeds from transformed plants are harvested and screened, with a typical transformant yield of 0.5 percent.
Thanks to the ease of conducting genetic research on Arabidopsis, a large international community of scientists works with this organism. An additional tool at our disposal is a comprehensive collection of transposon mutants in
Arabidopsis, created through random insertion of Agrobacterium transfer DNA (T-DNA) throughout the genome. The ends of the inserted transgenes have been sequenced and catalogued, so researchers who wish to examine a specific knockout mutant can simply order seed and grow it.
Breeding for Pathogen Resistance
One of the major techniques for controlling plant disease is through classical plant breeding. The science of plant breeding is considered to have begun in 1906 in England, when the first (intentional) genetic cross was performed in wheat. The basis of plant breeding for disease resistance is to identify pathogen-resistant germplasm at the geographical origin of the plant species, where the plant and its pathogen coevolved. Researchers have discovered many wild species of agronomically important plants that can be genetically crossed with their cultivated relatives. This repertoire of diversity is the source of all disease resistance genes in a particular plant species.
To introduce resistance genes into cultivated plants, breeders create interspecific hybrids by taking pollen from a wild species and crossing it into the agronomic species. This is followed by selection for resistance and many rounds of backcrossing to produce a stable, resistant cultivar—a process that can take as long as 10 years. Most resistance traits that have been incorporated through plant breeding are single dominant genes because plant breeders like to look at single traits, and they find them in the first generation; however, there are many instances of multiple and recessive genes controlling resistance. Because of the ease of manipulation of single-gene traits, a century of plant breeding for disease resistance has produced agronomic fields that are genetically uniform. The human equivalent of this situation would occur if every one of us had exactly the same genetic resistance to flu virus, and the virus mutated to avoid our defenses: it would kill us all. Plant breeding establishes a huge selection pressure for pathogens to mutate, and they do. Then the breeding cycle begins all over again, with continued natural genetic variation occurring in both host resistance (R) genes and genes for pathogen effector proteins.
A Model System of Plant Innate Immunity
Immunity to pathogens, while strategically similar among plants and animals, manifests itself in different ways in each kingdom. In keeping with their lack of mobility, plants have very sophisticated ways of dealing with both biotic and abiotic stress. The plant’s innate immune system employs hundreds of germline encoded pathogen receptors, most of which recognize type III effector proteins of bacterial pathogens. Plants have no system for adaptive immunity comparable to that in animals.
We study plant immunity through a discipline we call functional patho-
genomics, in which we examine both the host (Arabidopsis) and the pathogen (Pseudomonas syringae). Arabidopsis can be manipulated in much the same way as a microbe: everything you can do with yeast or with Pseudomonas, you can do with Arabidopsis, with the exception of gene replacement. Arabidopsis has 125 megabases of DNA and about 25,000 genes, but many of these genes are regulated as families and are therefore redundant, as is typical of plants. For example, a particular protein might be encoded by a family of genes, each of which functions only in the leaves, or in the roots, or in the stem. This redundancy complicates genetic forward screens and makes it difficult to reveal recessive mutations, but it also means that the Arabidopsis genome is far simpler than 25,000 genes might suggest.
The pathogen in our study system, P. syringae, is a gram-negative bacterium. It is a facultative parasite that lives on the leaf surface, and has been found to obtain populations of 105 colony-forming units per centimeter squared on the leaf surface without causing disease. Under appropriate environmental conditions, P. syringae enters the leaf through the stomates (openings in the epidermis where gas exchange occurs) or through wounds. It multiplies in the intercellular spaces and remains outside plant cells. Another interesting trait of P. syringae is its capacity to produce ice nucleation proteins; by ordering water molecules, these proteins can induce premature freezing, which kills the host plant. The advantage of this trait for the bacterium appears to be in promoting dissemination: bacteria on the leaf surface become incorporated into water droplets that return to the atmosphere and, eventually, rain down on other plants.
P. syringae has an inner and an outer membrane; the plant cell has a very complex plant cell wall, a plasma membrane, and various membrane-bound organelles within its cytoplasm. The bacterium delivers as many as 40 different effector proteins through the plant cell wall in order to suppress or modify the host plant’s immune response and thereby permit colonization, much as effector proteins from animal pathogens do. A type III secretion system, essentially a molecular syringe, injects the bacterial effector proteins into the plant cell. All 40 or so effector proteins have different (and some have multiple) targets within the plant cell that are either destroyed or altered in some fashion.
In response to such invasions, plants have evolved resistance proteins to deal with specific bacterial effector proteins. Plant resistance proteins have long been hypothesized to recognize—directly or indirectly—specific bacterial effector proteins. This notion, known as the gene-for-gene hypothesis, was first proposed in the 1940s by the plant pathologist H. H. Flor (Loegering and Ellingboe, 1987; Staskawicz, 2001). This concept is the essence of innate immunity in plants. However, until a decade or so ago, all that was known about plant R genes is that they typically segregated as a single trait. It was expected that R genes would encode receptors that served as ligands for the bacterial effector proteins, and that effector protein binding would cause the resistance protein to trigger a signal cascade leading to the disease resistance phenotype.
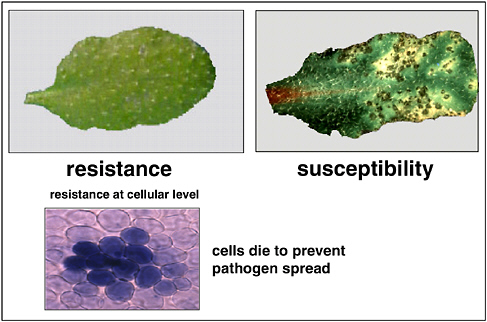
FIGURE 2-4 Phenotypes of bacterial disease resistance. Leaves from two isogenic plants that differ only as follows: the plant on the left has a single dominant disease resistance gene (RPS2); the plant on the right has a recessive (inactive) allele (rps2) of the same gene. Following inoculation of both plants with P. syringae, the plant on the left expresses resistance to the pathogen. At the microscopic level, it can be seen that infected cells in the resistant plant are undergoing programmed cell death, in a localized hypersensitive reaction.
SOURCE: Staskawicz (2005).
In 1994, our lab and several others cloned some disease resistance genes, and we have since found that in only a couple of cases, the ligand-receptor model does not apply. However, a key finding from these studies is that resistance genes confer upon plant cells the ability to undergo programmed cell death upon recognizing a pathogen. By committing suicide, infected cells prevent the pathogen from spreading through the plant. This phenomenon, known as the hypersensitive reaction, is depicted in Figure 2-4.
Structural Comparisons of Disease Resistance Proteins
There are two main classes of Arabidopsis disease resistance proteins: the cytosolic “nucleotide-binding site plus leucine-rich repeat” (NBS-LRR) type, of which there are several subtypes, and the membrane-bound “leucine-rich repeat transmembrane kinase” (LRR-TM kinase) type. Figure 2-5 shows the distribution of the 150 disease resistance gene loci among the five Arabidopsis chromo-
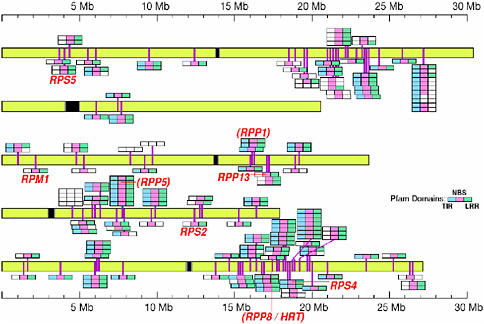
FIGURE 2-5 Arabidopsis disease resistance genes.
SOURCE: Michelmore (2001).
somes. The clustering of the R genes is thought to promote allelic diversity, and perhaps recombination as well (through unequal crossing over or possibly through gene conversion). After all, the 150 R genes in Arabidopsis not only encode the capacity for resistance to bacteria, but also resistance to viruses, fungi, nematodes, and insects. There must be some mechanism(s) that enable the plant to deal with the diversity of pathogen effector genes that it encounters.
Figure 2-6 shows similar motifs that occur in animal innate immune receptors and plant disease resistance proteins. For example, the Arabidopsis flagellin sensitivity 2 (FLS2) protein and the mammalian toll-like receptor 5 (TLR5) proteins both contain leucine-rich repeats and transmembrane domains; however, in the analogous cytoplasmic location to the FLS2 protein kinase domain, TLR5 has the toll interleukin 1 receptor. Both FLS2 and TLR5 interact with the bacterial protein, flagellin. The pervasive similarities in plant and animal innate immune receptor structure have become a focus for interaction among plant and animal molecular biologists studying these systems, as the entire community attempts to learn how these molecules actually function.
Disease Resistance at the Molecular Level
To illustrate the intricacy of disease resistance protein function and the methods we have used to elucidate how plant and pathogen components of disease
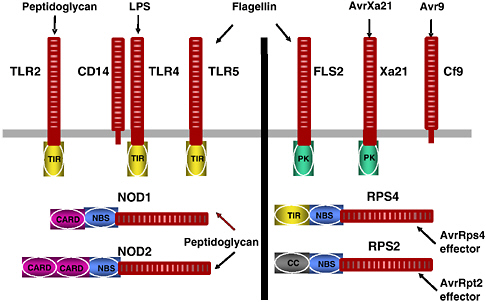
FIGURE 2-6 Representative protein motifs found in immune receptors of plants and animals. Schematic diagram of several proteins present in plants (left) and animals (right) that initiate signal transduction cascades after pathogen infection.
SOURCE: Staskawicz (2005).
resistance work together, I will describe research in my laboratory on a particular Arabidopsis disease resistance protein, RPS2 (Resistance to P. syringae 2). This protein recognizes a P. syringae effector protein called AvrRpt2. As shown in Figure 2-6, RPS2 is a membrane-associated NBS-LRR-type protein, with a coiled-coil “leucine zipper” nucleotide binding site at its N-terminus. As shown in Figure 2-7, RPS2 forms a complex with another Arabidopsis protein called RIN4 (for RPM1-interacting protein 4; RPM1 is also a disease resistance protein, Resistance to P. syringae pv maculicola 1).
It turns out that this system is under negative regulation, which appears to be a common theme of signal transduction networks in plants. In this case, the negative regulator RIN4 renders RPS2 inactive. This was shown to be the case by overexpressing RPS2 relative to RIN4, either transgenically or transiently, which produces a constitutive defense response: cell death. On the other hand, if RIN4 is overexpressed relative to RPS2, disease resistance is inhibited, so these plants are more susceptible to P. syringae. Homozygous rin4 mutants were found to be lethal in the presence of the RPS2 gene; this makes sense, because a constitutively activated defense response would result in whole-plant suicide by the hypersensitive response.
Recent work in my laboratory shows how these proteins function in the presence of AvrRpt2, the P. syringae effector protein (Axtell and Staskawicz, 2003;
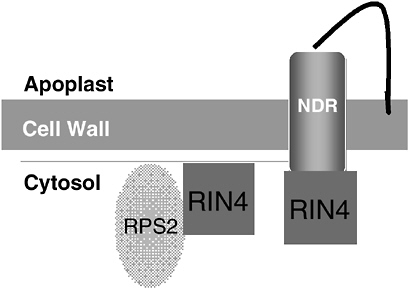
FIGURE 2-7 Model for RPS2-mediated innate immunity. RIN4 complexes with disease resistance proteins. RPS2 forms a complex with RIN4, a negative regulator of RPS2 activity (see Figure 2-8). RIN4 also forms a complex with and negatively regulates NDR1, a glycosylphatidylinositol membrane-anchored protein that is presumed to be involved in disease resistance downstream of the RPS2-RIN4 interaction.
SOURCE: Staskawicz (2005).
Axtell et al., 2003; Belkhadir et al., 2004). The bacterium delivers an inactive form of AvrRpt to the plant cell via a type III secretion system. The effector protein becomes active following processing by a plant protein factor (cyclophilin), after which 71 N-terminal amino acids are cleaved to produce the mature, 184-residue effector protein. After cloning and sequencing AvrRpt2, we analyzed its predicted secondary structure and found it had functional resemblance to the cysteine protease, staphopain.
We also isolated the plant protein factor that enables the processing of AvrRpt2 into its active form, and learned that it was common to all eukaryotes. This was significant because it meant that we could develop an assay for AvrRpt2 processing in yeast, which is even easier to manipulate than Arabidopsis. We took a biochemical approach, using the pure recombinant full-length AvrRpt2 protein and assaying chromatographic (hydrophobic interaction ion exchange gel filtration) fractions from yeast in order to isolate the eukaryotic proteins that processed the bacterial effector protein. Analysis of the active fraction by tandem mass spectrometry identified the eukaryotic processing factor as cyclophilin, a peptidyl-prolyl isomerase (PPIase) that is thought to act as a molecular chaperone
or folding catalyst. Cyclophilin has been widely studied in association with protein folding, signal transduction, protein trafficking and assembly, and cell cycle variation; it is required for assembly of the HIV-1 virion. Clearly, it is an important constituent of eukaryotic cells.
Next we used cyclophilin cDNA from yeast to find the orthologous gene in Arabidopsis. Pure recombinant protein from the Arabidopsis gene was then successfully tested in place of the yeast fraction in the processing assay. It worked as predicted, and we also found that cyclosporin A, which is known to bind specifically to cyclophilin, inhibited effector processing, while rapamycin, which inhibits another class of PPIases, did not.
The actual cleavage of AvrRpt2 occurs through autocatalysis, following processing (presumably refolding) by cyclophilin. Two regions encoding a 7-amino acid sequence of the autocatalytic cleavage site in AvrRpt2 occur in the negative regulator of disease resistance, RIN4. This suggests that RIN4 is inactivated when cleaved byAvrRpt2, which in turn permits activation of RPS2-mediated disease resistance. These findings are summarized in Figure 2-8. Researchers in my lab and others are now trying to determine the molecular basis of RPS2 activation.
A bioinformatics prediction has found putative AvrRpt2 cleavage sequences in approximately 20 proteins in the Arabidopsis genome, all of which are thus potential targets of that effector protein. By extension, it seems likely that among the 40 effector proteins produced by P. syringae, there are others that have multiple targets within the plant cell.
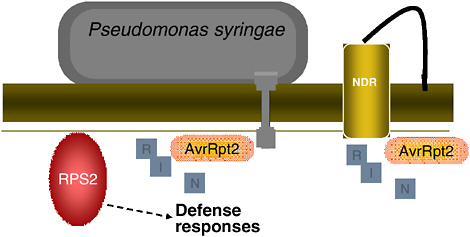
FIGURE 2-8 Model of RPS2-mediated immunity in Arabidopsis. In the presence of P. syringae effector protein AvrRpt2, RIN4 is eliminated and RPS2 disease resistance is activated.
SOURCE: Staskawicz (2005).
REFERENCES
Anderson RM, May RM. 1986. The Dynamics of Human Host-Parasite Systems. Princeton, NJ: Princeton University Press.
Axtell MJ, Staskawicz BJ. 2003. Initiation of RPS2-specified disease resistance in Arabidopsis is coupled to the AvrRpt2-directed elimination of RIN4. Cell 112(3):369–377.
Axtell MJ, Chisholm ST, Dahlbeck D, Staskawicz BJ. 2003. Genetic and molecular evidence that the Pseudomonas syringae type III effector protein AvrRpt2 is a cysteine protease. Molecular Microbiology 49(6):1537–1546.
Baker KF, Cook RJ. 1974. Biological Control of Plant Pathogens. San Francisco, CA: W.H. Freeman.
Baker B, Zambryski P, Staskawicz B, Dinesh-Kumar SP. 1997. Signaling in plant-microbe interactions. Science 276(5313):726–733.
Beijerinck MW. 1898. Ueber ein contagium vivum fluidum als ursache der fleckenkrankheit der tabaksblätter. Verhandelingen der Koninklyke Akademie van Wettenschappen te Amsterdam 65:3–21.
Belkhadir Y, Nimchuk Z, Hubert DA, Mackey D, Dangl JL. 2004. Plant Cell 16(10):2822–2835.
Berg HC. 1983. Random Walks in Biology. Princeton, NJ: Princeton University Press.
Brock T. 1961. Milestones in Microbiology. Englewood Cliffs, NJ: Prentice-Hall.
Brock T. 1988. Robert Koch: A Life in Medicine and Bacteriology. Berlin, Germany: Springer-Verlag.
Brown DG, Allen C. 2004. Ralstonia solanacearum genes induced during growth in tomato: An inside view of bacterial wilt. Molecular Microbiology 53:1641–1660.
Campbell CL, Peterson PD, Griffith CS. 1999. The Formative Years of Plant Pathology in the United States. St. Paul, MN: APS Press.
Cohn F. 1872. Untersuchungen über Bacterien. Beiträge zur Biologie der Pflanzen. 1:127–224.
Creager ANH. 2002. The Life of a Virus: Tobacco Mosaic Virus as an Experimental Model, 1930–1965. Chicago, IL: University of Chicago Press.
Dunn AK, Klimowicz AK, Handelsman J. 2003. Use of a promoter trap to identify Bacillus cereus genes regulated by tomato seed exudate and a rhizosphere resident, Pseudomonas aureofaciens. Applied Environmental Microbiology 69(2):1197–1205.
Gilbert GS, Parke JL, Clayton MK, Handelsman J. 1993. Effects of an introduced bacterium on bacterial communities on roots. Ecology 74:840–854.
Gilbert GS, Handelsman J, Parke JL. 1994. Root camouflage and disease control. Phytopathology 84:222–225.
Gilbert GS, Clayton MK, Handelsman J, Parke JL. 1996. Use of cluster and discriminant analyses to compare rhizosphere bacterial communities following biological perturbation. Microbial Ecology 32:123–147.
Handelsman J. 2005 (March 17). Session IV: Novel Approaches for Mitigating the Development of Resistance. Presentation at the Forum on Microbial Threats Workshop Ending the War Metaphor: The Changing Agenda for Unraveling the Host-Microbe Relationship, Washington, D.C., Institute of Medicine, Forum on Microbial Threats.
Handelsman J, Robinson CJ, Raffa KF. 2005. Microbial communities in lepidopteran guts: From models to metagenomics. In: McFall-Ngai M, Henderson B, Ruby EG, eds. The Influence of Cooperative Bacteria on Animal Host Biology. New York: Cambridge University Press. Pp. 143–168.
Ivanowski D. 1892. Ueber die mosaikkrankheit der tabakspflanze. St. Petersburg Academy of Imperial Sciences Bulletin Series 4 35:67–70.
Kolenbrander PE, Andersen RN, Blehert DS, Egland PG, Foster JS, Palmer RJ Jr. 2002. Communication among oral bacteria. Microbiolgy and Molecular Biology Reviews 66:486–505.
Levin SA, Hallam TG, Gross LJ. 1989. Applied Mathematical Ecology. New York: Springer-Verlag New York, Incorporated.
Loegering WQ, Ellingboe AH. 1987. H.H. Flor: Pioneer in Phytopathology. Annual Review of Phytopathology 25:59–66.
MacGuidwin AE, Rouse DI. 1990. Role of Pratylenchus penetrans in the potato early dying disease of Russet Burbank Potato. Phytopathology 80:1077–1082.
Mahan MJ, Slauch JM, Mekalanos JJ. 1993. Selection of bacterial virulence genes that are specifically induced in host tissues. Science 259(5095):686–688.
Michaelis L, Menten M. 1913. Die kinetik der invertinwirkung. Biochemistry Zeitung 49:333–369.
Michelmore R. 2001. Physical location of Arabidopsis sequences related to NBS-encoding plant R-genes. [Online]. Available: http://niblrrs.ucdavis.edu/At_RGenes/RGenes_Phylogeny/At_RGenes_on_Chromosomes.html [accessed April 28, 2006].
Osbourn AE, Barber CE, Daniels MJ. 1987. Identification of plant-induced genes of the bacterial pathogen Xanthomonas campestris pathovar campestris using a promoter-probe plasmid. European Molecular Biology Organization Journal 6(1):23–28.
Staskawicz B. 2001. Genetics of plant-pathogen interactions specifying plant disease resistance. Plant Physiology 125(1):73–76.
Staskawicz BJ. 2005 (March 16). Session I: Host-Pathogen Interactions: Defining the Concepts of Pathogenicity, Virulence, Colonization, Commensalism, and Symbiosis. Presentation at the Forum on Microbial Threats Workshop Ending the War Metaphor: The Changing Agenda for Unraveling the Host-Microbe Relationship, Washington, D.C., Institute of Medicine, Forum on Microbial Threats.
Waksman SA. 1958. My Life with the Microbes. London, UK: Scientific Book Club.






















