Developing Biomarker-Based Tools for Cancer Screening, Diagnosis, and Treatment
INTRODUCTION
Research has long sought to identify biomarkers that could detect cancer at an early stage, or predict the optimal cancer therapy for specific patients. Fueling interest in this research are recent technological advances in genomics, proteomics, and metabolomics that can enable researchers to capture the molecular fingerprints of specific cancers and fine-tune their classification according to the molecular defects they harbor. The discovery and development of new markers of cancer could potentially improve cancer screening, diagnosis, and treatment. Given the potential impact cancer biomarkers could have on the cost effectiveness of cancer detection and treatment, they could profoundly alter the economic burden of cancer as well.
Despite the promise of cancer biomarkers, few biomarker-based cancer tests have entered the market, and the translation of research findings on cancer biomarkers into clinically useful tests seems to be lagging. This is perhaps not surprising given the technical, financial, regulatory, and social challenges linked to the discovery, development, validation, and incorporation of biomarker tests into clinical practice. To explore those challenges and ways to overcome them, the National Cancer Policy Forum held the conference “Developing Biomarker-Based Tools for Cancer Screening, Diagnosis and Treatment: The State of the Science, Evaluation, Implementation, and Economics” in Washington, D.C., from March 20 to 22, 2006.
At this conference, experts gave presentations in one of six sessions:
-
Brief overview of technologies, including genomics, proteomics, metabolomics, and functional imaging
-
Overcoming the technical obstacles, with presentations on informatics and data standards, and biomarker validation and qualification
-
Coordinating the development of biomarkers and targeted therapies, with a clinical investigator and representatives from industry and the National Cancer Institute offering their perspectives
-
Biomarker development and regulatory oversight, including current regulations governing biomarker tests as well as new clinical trial designs needed to incorporate biomarker tests that predict patient responders
-
Adoption of biomarker-based technologies, with discussion on what motivates private insurers and Medicare to cover biomarker-based tests and what various organizations consider when recommending such tests be adopted into clinical practice
-
Economic impact of biomarker technologies, with an exploration of cost-effectiveness analyses of biomarker tests and a payor perspective on the evaluation of such tests
In addition, seven small group discussions explored the policy implications surrounding biomarker development and adoption into clinical practice:
-
Clinical development strategies for biomarker utilization
-
Strategies for implementing standardized biorepositories
-
Strategies for determining analytic validity and clinical utility of biomarkers
-
Strategies to develop biomarkers for early detection
-
Mechanisms for developing an evidence base
-
Evaluation of evidence in decision making
-
Incorporating biomarker evidence into clinical practice
This document is a summary of the conference proceedings, which will be used by an Institute of Medicine (IOM) committee to develop consensus-based recommendations for moving the field of cancer biomarkers forward. The views expressed in this summary are those of the speakers and discussants, as attributed to them, and are not the consensus
views of the participants of the workshop or of the members of National Cancer Policy Forum.
OVERVIEW OF TECHNOLOGIES USED TO DISCOVER CANCER BIOMARKERS
Technology is constantly evolving and recent technological advances have made it easier to discover many potential cancer biomarkers through high-throughput screens. Advances in imaging technology also are furthering the discovery and use of biomarkers. The goal of the first session of the conference was to provide a brief overview of the technologies currently being used to identify and develop cancer biomarkers (Figure 1).
Genomics, Proteomics, and Metabolomics
Todd Golub, MD, of the Dana-Farber Cancer Institute, began by discussing several of the genomics-based techniques commonly used to discover biomarkers for cancer detection or for patient stratification for therapy. Some of these techniques detect changes at the DNA level (are DNA-based), whereas others detect changes at the RNA level and are considered RNA-based.
Dr. Golub explored which type of genomics test—DNA based or RNA based—would be likely to serve as a better biomarker if cost were not an issue. DNA-based tests are advantageous because DNA is more stable than RNA, and because most changes related to cancer occur at the DNA level, he said. But he noted that perhaps one could make a stronger argument for RNA-based tests because not only can they detect oncogenic RNA missteps, but molecular signatures at the RNA level also help reveal upstream DNA-level abnormalities that could contribute to a cancer. These abnormalities include base substitutions, and amplifications or deletions that alter the copy number or heterozygosity of specific genetic sequences. Dr. Golub noted that studying epigenetic changes in DNA, such as methylation, and genome rearrangements, such as chromosome translocations, can also lead to discovery of important cancer biomarkers, although he did not have time to address these topics in his presentation.
Although early genetic analyses of cancers focused on detecting changes in the copy number of genes, Dr. Golub stressed that it is also important to screen for loss of heterozygosity (LOH). LOH can occur without a change
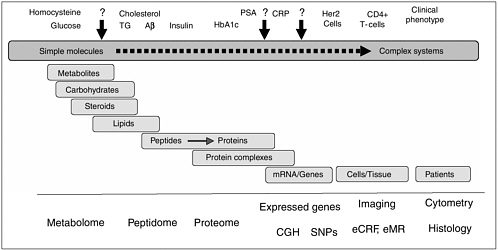
FIGURE 1 The spectrum of potential biomarkers suggests that no single technology can cover the enteir biomarker space. TG = triglycerides; Aβ = β-amyloid; HbA1c; PSA = prostate-specific antigen; CRP = C-reactive protien; CEG = comparative genomic hybridization; Snp = single nucleotide polymorphism; eCRF = electronic case report from; eMR = electronic medical record.
SOURCE: Adapted from Schulman presentation (March 20, 2006).
in gene copy number, he noted, if both alleles for a specific gene have been mutated or epigenetically altered. This copy-neutral LOH may account for as much as half of all LOH in the genome.
Two main types of arrays are used to detect changes in copy number or LOH linked to cancer. Single nucleotide polymorphism (SNP) arrays have between 50k and 500k SNPs across the genome and can detect both copy number changes and other forms of LOH. Comparative genomic hybridization arrays can detect changes in copy number of DNA content, but are unable to detect LOH in which the copy number remains the same. For this reason, Dr. Golub prefers SNP arrays for detecting cancer biomarkers. Higher density SNP arrays can give sharper resolution by reducing the signal-to-noise ratio than lower density SNP arrays, he pointed out. But the optimal amount of density that is the most cost-efficient means for detecting cancer biomarkers remains to be determined.
Standard DNA sequence analysis of tumor samples as a means of detecting cancer biomarkers has numerous drawbacks, which Dr. Golub pointed out. Not only is it difficult and costly to do, but it is frequently inaccurate, causing false negatives because of normal tissue contamination of the tumor samples used. Most tumor samples contain a mixture of normal cells, such as inflammatory cells, as well as tumor cells. Because the Sanger sequencing results are an average of both the normal and tumor cells in a sample, normal genome contamination can obscure mutations in tumor cells that might serve as cancer biomarkers.
However, newer techniques, such as single-molecule sequencing, may substantially lower the cost of sequencing, and should avoid the problems of normal cell contamination that plague standard sequencing efforts. “I think this is exactly the type of technology, even if cost neutral, that would dramatically accelerate our ability to detect important mutations in cancer,” Dr. Golub said.
To exemplify this, Dr. Golub reported on results from his colleagues at Dana-Farber who used single-molecule sequencing to detect a mutation that was linked to resistance of the drug Iressa in a lung cancer patient. The lung fluid sample the researchers analyzed only had 3 percent tumor cells, and a standard Sanger sequencing analysis missed the mutation.
Once a genetic signature with likely clinical relevance has been discovered, custom-made arrays that only have the gene sequences of interest need to be made for preclinical or clinical testing. Dr. Golub described a few genetic signature amplification and detection platforms useful for such testing, including a Luminex bead-based method. For this method, the genetic
material is amplified using polymerase chain reaction. The genetic signature is then read not on microarrays, but on miniscule color-coded beads that are detected by lasers in a flow cytometer. This is an inexpensive way to detect genetic signatures, costing about 50 cents for every 100 transcript signatures. One can also use the standard mRNA expression profiling platforms that are commercially available. These are all sufficiently accurate and precise to be used in a clinical setting to detect genetic signatures, according to Dr. Golub. Cost and throughput will be significant drivers of this technology, he added.
The next presentation was on proteomics and metabolomics technologies, given by Howard Schulman, PhD, of PPD Biomarker Discovery Sciences. One of those techniques, which Dr. Schulman described as the traditional proteomic workhorse, uses two-dimensional polyacrylamide gels for the separation stage. This is a slow process that is less amenable to high-throughput. Surface-enhanced laser desorption/ionization is a high-throughput technology that can more quickly separate the proteins in a sample, but identifying the protein peaks is a challenge. That identification process can be bypassed by using software to differentially identify patterns of protein peaks to find a molecular fingerprint that can distinguish cancerous from noncancerous tissue. This fingerprint is based on the amounts of all the various proteins detected, without knowledge of what those individual proteins are, Dr. Schulman noted. However, it can be problematic to translate mass spectroscopy fingerprints into a clinical diagnostic test without identifying or further characterizing those proteins.
One- and multidimensional liquid chromatography are also used to separate peptides in a sample (after protein digestion) that a mass spectrometer can differentially quantify and then identify (Figure 2). But the amplitude for each of the peptides can vary depending on the composition of the mixture, which makes it hard to compare one person’s sample with another’s, and one batch run versus another. This has proven problematic for researchers trying to develop cancer biomarkers based on differential quantification, otherwise known as molecular fingerprinting.
To improve such differential accuracy, researchers developed a method called isotope-coded affinity tags several years ago. This technique labels a portion of a sample with a mass tag and runs both labeled and unlabeled samples through the mass spectrometer at the same time. The labeled sample serves as a sort of baseline control for the unlabeled sample. This helps normalize or eliminate a lot of the peak amplitude variability due to differences in mixture composition. But this is a more costly method because
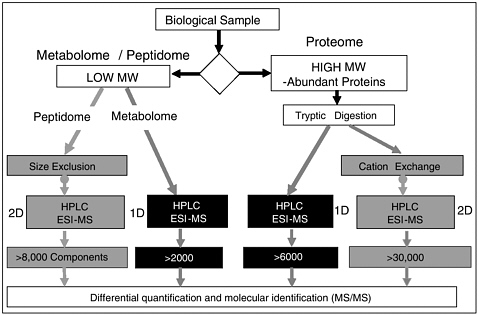
FIGURE 2 One-dimensional and multidimensional liquid chromatography LC-LC/MS. LC = liquid chromatography; MS = mass spectroscopy; MW = molecular weight; HPLC = high-performance liquid chromatography; ESI = electrospray ionization.
SOURCE: Schulman presentation (March 20, 2006).
of the need for the reagents, and it has some bias introduced by the type of tag used, according to Dr. Schulman. The field is rapidly adopting a label-free approach in which chromatographic separation techniques and mass spectrometry are coupled with software-based solutions for normalizing the variation in amplitude signal due to differences in mixture composition to yield accurate differential expression data.
Dr. Schulman concluded his talk by noting that the current state of proteomics is comparable to the early days of microarrays, which could detect about one-sixth the number of genetic sequences that can now be detected. But proteomics is still highly effective even without the ability to profile every protein, he said. He noted that one can profile more than a thousand proteins by using multidimensional chromatography. But the tradeoffs with more fractionation are lower throughput (due to slower processing) and higher costs. The advantage of proteomic and metabolomic profiling is that you can sample readily accessible tissues, such as plasma and urine, that are ideal for monitoring biomarkers in clinical trials and testing diagnostics.
He also noted that the lowest abundance proteins, such as cytokines or other signaling molecules, will likely require antibody-based protein chips to complement liquid chromatography separation techniques. Sensitivity to such proteins could also be increased by using samples likely to have higher concentrations of biomarkers of interest. For example, cerebral spinal fluid could be tapped to find biomarkers for lymphoma metastases in the central nervous system, or prostatic fluid could be used to detect biomarkers for prostate cancer. Affinity capture of protein subcategories, such as phosphorylated proteins, could also selectively profile lower abundance proteins of interest.
Drs. Schulman and Golub stressed the need to experimentally validate the biological basis and importance of detected genetic or proteomic differences in a disease process. For example, researchers in Dr. Golub’s laboratory used high-density expression arrays to detect an RNA signature in bone marrow samples that correlated with response to a drug for myelodysplastic syndrome. They found a group of genes that were only highly expressed in patients who responded to the drug. Many of these genes previously had been identified as markers for late red blood cell differentiation, leading to the hypothesis that such differentiation may be predictive of drug response.
To test this idea, they induced normal immature blood cells to differentiate into red blood cells. They found that all of the genes, whose boosted expression was linked to drug response in their biomarker discovery study, also had heightened expression during the red blood cell differentiation that occurred in their experiments. This validated their hypothesis and put the concept of genetic signature for drug response on firmer footing. “The most valuable and robust biomarkers will be those that have some component of experimental validation accompanying them,” Dr. Golub said. He added that “the challenge looking forward is going to be to move from simply cataloging mutations or genome abnormalities to coalescing those abnormalities into more of a molecular taxonomy that brings biological understanding to this catalog. The more we can integrate these anonymous molecular signatures with biological knowledge, the more they’re likely to stick.”
Dr. Golub also pointed out the need to develop biomarker diagnostics that can easily be used on the paraffin-embedded or formalin-fixed tissues that are routinely collected in the clinic. “We need to make the technology work for those routinely collected samples rather than retrain the medical community to collect samples in a different way,” Dr. Golub said.
Drs. Golub and Schulman noted that a lack of good-quality samples can be a stumbling block for biomarker discovery. Rarely are enough samples collected in a clinical trial, and those samples that are collected are usually fixed in formalin, which can affect their ability to be analyzed in a mass spectrometer. Dr. Schulman suggested that pharmaceutical and biotechnology companies have experimental medicine groups that are best positioned to collect the samples required to discover biomarkers.
But the biggest impediment for biomarker development, which Drs. Golub and Schulman both cited, was a lack of a critical mass of research in the discovery phase. “The bottleneck is not so much on the regulatory side or the validation side, but that not enough of the discovery effort has been made,” said Dr. Schulman.
As to whether such efforts at biomarker discovery should take a hypothesis-driven or open-ended approach, Drs. Golub and Schulman agreed that both approaches were necessary. Open-ended discovery aims at uncovering a molecular understanding of a particular type of cancer that may eventually lead to useful biomarkers. A hypothesis-driven approach, in contrast, is more streamlined at finding molecular changes likely to predict a response to therapy or some other useful clinical endpoint. There is a role for both these approaches, Dr. Schulman said. But he added that pharmaceutical companies are unfortunantely more likely to conduct a hypothesis-driven search for biomarkers that predict drug response than to support a more open-ended search. Dr. Golub noted that the danger of conducting only hypothesis-driven research on biomarkers is that it does not address the challenge of “how do we get beyond discovering what we already know, in terms of biological knowledge?”
Molecular Imaging
Next, Michael Phelps, PhD, of the University of California, Los Angeles, discussed molecular imaging biomarkers for drug discovery, development, and patient care. He described how positron emission tomography (PET) can be used as a molecular camera to image in vivo processes at the molecular level. But PET is more than an imaging device, as it also can be used analytically to perform a variety of quantitative biochemical and biological assays.
There are currently about 600 PET probes for metabolism, receptors, enzymes, DNA replication, gene expression, antibodies, hormones, drugs, and other compounds in nanomole amounts. Typical antibody probes get
broken down too quickly to be effective for PET imaging, but there are modified antibody probes that are small molecule versions of the original antibodies and retain the active end. Most PET probes were developed from probes used in in vitro assays so as to translate that assay into an in vivo measurement. Ninety percent of PET probes were developed from drugs, Dr. Phelps reported.
Over the past few years, PET scanners have merged with computed tomography (CT) scanners to combine the anatomical definition of the CT with the biological assay capability of the PET scan. Researchers have also created microPET/CT machines to image biological processes and drug responses in mice.
Because PET probes are administered in nanomole amounts, measures can be performed on biological processes without disturbing the processes or causing pharmacologic mass effects, Dr. Phelps noted. Not only can PET scans be safely done, but studies show they are more accurate than magnetic resonance imaging or CT scans for the diagnosis and staging of cancer, for assessing therapeutic response, and for detecting cancer recurrence.
To detect cancer, technicians usually use a PET probe that images the heightened glucose metabolism of cancer cells. To predict or determine response to therapy, a number of different types of probes are used, depending on the type of cancer and type of treatment. The PET assay can enable stratification of patients according to whether they have the therapeutic target. For example, a probe that detects DNA replication may be used to predict whether a cancer will respond to a chemotherapy that blocks such replication. A probe for an estrogen receptor may be used to determine if breast cancer metastases are likely to respond to hormonal therapy. PET is especially useful for revealing whether a tumor is responding to therapy. It can detect within a day, for example, whether patients’ tumors are not responding to Gleevec, thereby quickly determining if patients should receive a different drug, Dr. Phelps pointed out.
PET imaging also has advantages over standard techniques for assessing the pharmacokinetics and pharmacodynamics of drugs, he added. For example, standard pharmacokinetic assessments are based on measurements of how quickly a drug is cleared from the blood. In contrast, by using labeled drugs as probes, PET can precisely measure the concentration of the drug, not just in the blood, but in all tissues over time, he noted.
Dr. Phelps described a recent innovation in PET technology that uses “click chemistry” to create PET probes. This technique involves combining two small molecules with low to moderate affinity to the target, but high
affinity to each other. They collectively latch onto the target as they bind to each other. The end result is that they bind to the target with an extremely high affinity that is the product of the affinities of the two molecules. These probes dramatically increase the sensitivity and physical resolution of PET imaging. Because the probes are such small molecules, they can access surface receptors, cells, and even the cell nucleus.
Dr. Phelps concluded his talk by noting there are “PET pharmacies” scattered all over the world that use automated chemistry to make and ship labeled molecular PET probes. There are also “labs on a chip” that enable researchers to custom build their own PET probes using click chemistry and other techniques.
In response to comments by Drs. Golub and Schulman regarding where the bottleneck is in biomarker development, Dr. Phelps noted that as one gets closer to introducing a biomarker into a clinical setting, Food and Drug Admininstration (FDA) premarket regulation can become very limiting. He suggested that regulatory bodies work with researchers to change the criteria by which drugs and molecular diagnostics are evaluated.
MEETING THE TECHNICAL CHALLENGES OF BIOMARKER VALIDATION AND QUALIFICATION
Appropriate analysis and interpretation of biomarker data presents enormous challenges, especially with the advent of genomic and proteomic technologies that can generate a tremendous amount of data on individual samples. Three speakers at the conference addressed the technical challenges involved with validating the accuracy and clinical relevance of cancer biomarkers. John Quackenbush, PhD, of Harvard University spoke about experimental design considerations and data reporting standards to aid the validation of biomarkers. David Ransohoff, MD, of the University of North Carolina also discussed experimental design, and the shortcomings of recent cancer biomarker studies that should be avoided in future studies. John Wagner, MD, PhD, of Merck and Co., Inc., gave a pharmaceutical company’s perspective on what is required to validate a cancer biomarker and establish its relevance to useful clinical endpoints.
Dr. Quackenbush began this session by noting that with microarray technologies, researchers tend to do more hypothesis-generating experiments than hypothesis-driven experiments. But despite a lack of an experimental hypothesis, one still needs to think critically about experimental design and how data are collected, managed, and analyzed, he said. All of these steps
play crucial roles in determining whether the results derived from biomarker studies are clinically meaningful and valid in broader populations than in the original test population.
Drs. Quackenbush and Ransohoff stressed that the same issues that apply to standard hypothesis-driven clinical studies are also applicable to studies in genomics, proteomics, and metabolomics, which they collectively called “omics.” “This is an exciting era because we have very powerful tools to measure the biology [of cancer], but the rules of evidence about validity have not changed,” said Dr. Ransohoff. “New reductionist methods mean lots more data, but not necessarily more knowledge, and the rules of evidence about how you go from data to knowledge haven’t changed.”
Dr. Quackenbush cited a need for the development of more cutting-edge bioinformatics tools to help with data analysis, and called for collaborations between bench researchers and bioinformatics specialists to develop those tools. Dr. Ransohoff acknowledged that bioinformatics is important, but pointed out that many of the problems in data analysis and interpretation of the omics field are not new problems stemming from the nature of the technology. Instead, they are age-old problems well known to clinical epidemiologists: overfitting of data, bias, and sample sizes that are disproportionately small compared to the number of variables measured. Researchers in the omics field do not pay enough attention to these experimental design flaws that can distort the accuracy and reproducibility of their results, Dr. Ransohoff said.
Overfitting of data is a problem in a number of omics studies, Drs. Quackenbush and Ransohoff asserted. Overfitting can occur when a large number of predictive variables are fit to a small number of subjects. A model can fit perfectly by chance in these situations, even if there is no real relationship, Dr. Ransohoff pointed out. He cited a study by Richard Simon1 in which Dr. Simon simulated a genomics study by making up patients, assigning them genes with various degrees of expression, and randomly assigning whether or not they had cancer. Dr. Simon then did a multivariable analysis to see if he could find a genetic signature model that discriminated between patients who had cancer versus those who did not. He found that, depending on how he did his analysis, he could make a discrimination model fit the data almost perfectly (98 percent of the time). He was able to achieve high-accuracy assessments of predictive genetic
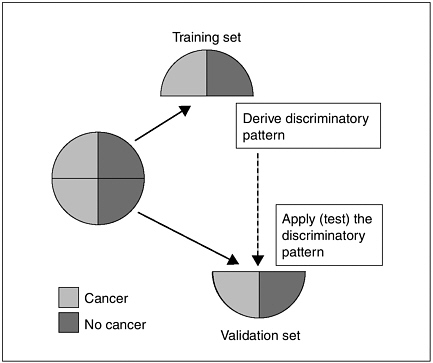
FIGURE 3 Method of dividing original sample to assess reproducibility and overfitting.
SOURCE: Ransohoff presentation (March 20, 2006).
signatures, even though they did not truly determine which patients had cancer, as this was randomly assigned.
The way to check for overfitting is to assess the reproducibility of the results in a new group of subjects who are totally independent from the original group (Figure 3). But such assessments often are not done in omics studies, according to Dr. Ransohoff. Instead, results from a new group of subjects are often combined with those from the original group to further assess the accuracy of a predictive genetic signature or proteomic pattern.
Having a large enough sample population can help avoid the problem of overfitting, Dr. Quackenbush noted. “If we find a biomarker, or a set of patterns that we use as a biomarker, in 20 to 30 samples when we’re looking at tens of thousands of genes, there’s a high likelihood that when we go to a larger population, many of those genes that we see in the small sample set won’t hold up as robust markers,” he said.
According to Dr. Ransohoff, overfitting helps to explain why a number of studies of cancer biomarkers, including a Dutch study that recently reported a gene expression signature as a predictor of breast cancer survival,2 showed initial highly promising results that did not hold up quite as strongly when researchers tried to duplicate them in different study populations. One reanalysis of the original data from seven RNA expression and cancer prognosis studies3 found that in five of them, results were no better than chance. Dr. Ransohoff pointed out that many of these studies were conducted at well-respected institutions and published in major journals, such as Lancet and the New England Journal of Medicine. “If our best institutions don’t know when the data are strong enough to support claims like this, then there’s something genuinely difficult about the entire field,” he said.
Dr. Ransohoff said much of the faulty study designs of omics research, and their readily accepted findings by major journals, stems from a culture clash between bench scientists and clinical researchers. “A culture clash hinders exploration when you get people from these fields in the same room and they really can’t communicate with one another because the molecular biologists don’t understand enough about clinical or observational epidemiology and biostatistics, and the epidemiologists and biostatisticians may be intimidated and don’t know enough about molecular biology and biochemistry,” he said.
Bias is another common problem in experimental research that is not addressed adequately by many in the omics field, according to Dr. Ransohoff. Bias is the systematic difference between compared groups that alters the accuracy of the conclusions stemming from the comparison. Bias is such a common and serious problem in research that “results of a study must be regarded as being guilty of bias until proven innocent,” he said. Just one bias can be a fatal flaw in a study.
As Dr. Quackenbush noted, “I’ve looked at people’s datasets, even published datasets, where they claim differences between two groups, and when I look at it, I see the primary difference being the difference between two hospitals or two collection protocols.” As an example of bias, Dr. Ransohoff reported on the reanalysis of the data from studies of the highly acclaimed
proteomics test for ovarian cancer, which supposedly could detect ovarian cancer in blood serum with near 100 percent accuracy.4 When statistician Keith Baggerly scrutinized the methods used to assess the accuracy of the study results, he discovered significant nonbiologic experimental bias between the cancer and control groups. He found that the researchers ran their proteomic analyses of ovarian cancer samples on different days than when they ran the same analyses on noncancer samples. Because of mass spectrometer drift over time, this created a bias because a “signal,” from the machine, was introduced into one group but not the other, making the proteomics test result invalid.5
In clinical research, the bias of baseline inequality is usually avoided easily and effectively by using randomization, but researchers still go to great lengths simply to report that there are no statistical differences in the baseline conditions of the study populations they are comparing. In contrast, the features needed to assess “baseline inequality” are seldom reported in the same detail in much “omics” research. According to Dr. Ransohoff, “the process to deal with bias is routinely ignored by authors, reviewers and editors in omics research.” A number of factors could cause bias in omics research, including differences in how samples are collected, handled, and stored, or in how the assay is run. But such details are rarely reported when this research is published, he said. “When I want to find out what’s happened in an article, I’ve got to go to a Gordon conference and take the researcher out for a walk in the woods and interview [him or her] for an hour. But, of course, that’s what our method sections are supposed to do,” Dr. Ransohoff said. “Our methods sections are failing the scientific community in much ‘omics’ research.”
Dr. Quackenbush also stressed the need for data and methods to be openly reported in a readily accessible fashion so that other researchers can review them and/or compare the reported data to their own research results. Such reviews and comparisons are key to validating particular biomarkers. But to do such reviews and comparisons, researchers need to know the biological characteristics of the study samples, including relevant clinical information, how the samples were collected and analyzed, and what the
results were. This information is often missing in published journal articles or data published online, Dr. Quackenbush noted.
To counter that lack of information, he and others at the Microarray Gene Expression Data Society created a guide for authors, editors, and reviewers of microarray gene expression papers. The Minimum Information About a Microarray Experiment (MIAME) guide6 requires researchers to report effectively on their entire process of collecting, managing, and analyzing data so that the data can be reused and interpreted by others. The MIAME guide was published in 2001, and has been readily adopted by several scientific journals as a requirement for publication. The guide has led to the development of standards in other fields, including metabolomics and proteomics, according to Dr. Quackenbush.
Numerous challenges in the reporting of data still need to be addressed, however, Dr. Quackenbush pointed out. One challenge is to develop a standard format for consistently describing and entering clinical data, such as the estrogen receptor status of a tumor sample, into a database so that the information can be accessed easily by others. “A rose by any other name is a rose, you just can’t find it in the database,” he said. He suggested “carrots and sticks” from research funders and journals to encourage more standardized reporting of data.
Standard data formats are especially needed so researchers can compare genomic, proteomic, and metabolomic datasets to each other. Such cross-domain comparisons will enable researchers to move more rapidly from the discovery of biomarkers to their applications in the clinic, Dr. Quackenbush said. A centralized repository of omics data would be helpful to make such comparisons, he added, but such a repository does not currently exist. Most researchers are not keen on creating an omics database, because such work is considered “blue-collar science,” he said. “It’s not very sexy—nobody is going to win a Nobel Prize for creating a database, yet bringing such data together and integrating it is absolutely essential if we want to look beyond these demonstration studies that have been done and really do the largescale clinical studies we’d like to be able to do.”
There also is a need to develop tools that can visualize and interpret omics data in a way that is easy for clinicians to access and understand. Otherwise, omics tests will not be readily adopted in a clinical setting. “You don’t want to have to send your data off to a statistician in order to tell a
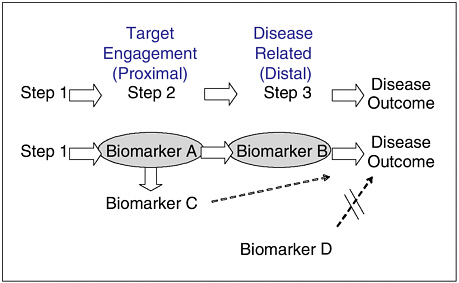
FIGURE 4 Target-engagement markers (Biomarker A) versus disease-related markers (Biomarker B). Pathophysiology is typically a multistep process. A putative biomarker may be (i) involved in one of the steps of the pathophysiology of a disease outcome (Biomarker A), (ii) related to, but not directly involved in, the pathophysiology of a disease outcome (Biomarker C), or (iii) not involved in the pathophysiology of a disease outcome (Biomarker D).
SOURCE: Wagner presentation (March 20, 2006). Adapted from Wagner (2002).
patient whether or not he or she is going to be resistant to chemotherapy,” Dr. Quackenbush noted.
Dr. Wagner explored a new angle of biomarker validity in his talk by showing how pharmaceutical companies classify biomarkers and tailor their degree of validity assessments according to the type of biomarker and how it will be used. He began his talk by pointing out how many biomarkers fall at various intervals on the pathophysiology path from the initial trigger or cause of a disease to final disease outcome (Figure 4). Biomarkers that occur close to the actions of the target are termed target-engagement biomarkers. Those that are closer to the disease outcome are called disease-related biomarkers. Target-engagement biomarkers help one understand how well a drug is acting on a target, whereas disease-related biomarkers are used to assess the effect of a particular drug on a disease.
Some biomarkers are not directly related to pathophysiology, yet are still useful. One example is hemoglobin A1c, Dr. Wagner noted. This
is a measure of hemoglobin with glucose molecules attached (glycated hemoglobin). When there are higher than normal levels of blood glucose, as occurs with diabetes, more hemoglobin becomes glycated. Blood levels of hemoglobin A1c serve as an excellent surrogate endpoint in diabetes drug trials, yet this biomarker has nothing to do with the diabetes disease process—that is, the glycation of hemoglobin has no impact itself on the health of the patient.
Dr. Quackenbush also pointed out that “there are many examples of biomarkers that exist outside of the realm of omic technologies that are clinically useful even though they don’t have a clear mechanistic interpretation.” Both prostate-specific antigen (PSA) and carcinogenic embryonic antigen are biomarkers used clinically to manage patients, he noted, but they do not explain tumor behavior. So although finding a mechanistic interpretation can help validate a biomarker, one shouldn’t rule out the usefulness of a biomarker if its mechanism of action cannot be directly related to a disease process, Drs. Quackenbush and Wagner pointed out. “If we focus too strongly on just looking at mechanistic understanding in order to develop biomarkers, we may be throwing out the baby with the bathwater,” Dr. Quackenbush said.
Another way pharmaceutical companies classify biomarkers is according to the purpose for which they will be used (Box 1). Exploratory biomarkers are usually used to generate hypotheses and are mainly seen as research and development tools. Demonstration biomarkers are considered one step up from that and termed probable or emerging biomarkers, according to FDA parlance.
|
BOX 1 Biomarker Types Characterization—known or established biomarker that often aids drug development decision making. Demonstration—a probable or emerging biomarker. Disease-related—used to assess the effect of a particular drug on a specific disease process. Dose stratifier—an indicator of the optimal dose of a specific drug for a specific patient. |
|
Early compound screening—biomarker used early in drug development to detect likely effective drug candidates, that is, those that affect a specific drug target. Early response indicator—biomarker that objectively indicates early in treatment whether the patient is responding to the treatment; for example, PET imaging of tumor size. Exploratory—used to generate hypotheses; a research and development tool. Partial surrogate endpoint—indicator of the effectiveness of treatment in early (Phase I/II) clinical trials. Improvement of a partial surrogate endpoint is necessary for, but not sufficient to, ensure improvement of the primary clinical endpoint of interest. Partial surrogate endpoints serve as indicators of whether to continue the clinical testing of new drugs and progress to Phase III trials. Patient classifier—marker that classifies patients by disease subset. Pharmacodynamic—marker that indicates drug activity and informs dose and schedule selection of a drug. Relapse risk stratifier—indicator of the degree of risk for relapse after initial therapy. Response predictor—a measurement made before treatment to predict whether a particular treatment is likely to be beneficial. Risk management—marker for patients or subgroups with high probability of experiencing adverse effects from their treatment, such as a marker for a drug metabolism subset. Risk stratifier—indicator of the probability of an event (e.g., metastasis) or time to the event. Surrogate endpoint—an outcome measure that is thought to correlate with the primary clinical endpoint (outcome) of interest, and is used in place of the primary endpoint to determine whether the treatment is working. Target-engagement—indicator of how well a drug is acting on a target. Tumor progression indicator—a measurement that provides early detection of tumor progression following treatment; for example, an increase in PSA levels can indicate progression of prostate cancer. SOURCE: This box is based on information presented by Drs. Janet Woodcock, John Wagner, and Richard Simon at the workshop. |
TABLE 1 Research and Regulatory Use of Qualified Disease-Related Biomarkers
|
Exploration |
Hypothesis generation |
|
Demonstration |
Decision making, supporting evidence with primary clinical evidence |
|
Characterization |
Decision making, dose finding, secondary/tertiary claims |
|
Surrogacy |
Registration |
|
SOURCE: Wagner presentation (March 20, 2006). Adapted from PhRMA Biomarker Working Group, FDA Advisory Committee Meeting (2004). |
|
Characterization biomarkers are known or established biomarkers that often aid drug development decision making, and surrogacy biomarkers can substitute for clinical endpoints in drug efficacy studies.
All biomarkers undergo some degree of validation and qualification. Dr. Wagner defined qualification as the evidentiary process of linking a biomarker with biology and clinical endpoints, generating data that are scientifically and clinically meaningful within the context of its intended use. This contrasts with validation of the biomarker assay, which is obtaining reliable biomarker data that meet the experiment or study objective. The degree of validation and qualification of biomarkers should fit their purpose, and depend upon whether they are target-engagement biomarkers or disease-related biomarkers (Table 1).
Exploratory biomarkers require a minimum set of assay validation experiments, but demonstration or characterization biomarkers require more advanced assay validation. This is especially true if they will be used as a basis for drug development decisions, such as whether a drug is effective, or at what dose the drug should be used. A target-engagement biomarker that is used in drug development decision making would need some advanced validation, but would not be subject to qualification assessments, whereas a disease-related biomarker that would be used for such decision making would undergo qualification assessments, said Dr. Wagner.
COORDINATING THE DEVELOPMENT OF BIOMARKERS AND TARGETED THERAPIES
Only a fraction of cancer patients will respond to a given cancer therapy, with responders being as low as 1 percent for drugs that target
specific genetic and molecular changes in cancer cells. Such targeted treatments often require biomarkers that can reliably predict patients likely to respond in order to show efficacy in clinical trials, let alone in the clinical setting at large. But development of biomarker-based tests to predict drug responders has lagged and is often undertaken outside of the company developing the drug. Progress in this field potentially could be accelerated by coordinating the development of biomarkers and new drugs. The goal of the third session of the conference was to discuss current incentives and disincentives for the development of biomarkers for targeted cancer therapies, and ways to encourage cooperation and resource sharing.
Therapeutics Industry Perspective
Paul Waring, PhD, of Genentech opened this session by summarizing the state of the art for developing clinically useful biomarker tests to predict patients likely to respond to targeted cancer therapies. He discussed the first successful attempt in this regard, which was the codevelopment of the breast cancer drug Herceptin with a diagnostic test that predicted whether breast cancer patients would be likely to respond to it. Herceptin targets the gene human epidermal growth factor receptor 2 (HER2), which is overexpressed in about 25 percent of breast cancer cases due to amplification of the gene. Genentech, which developed the drug, also developed an assay to select patients likely to respond for its clinical trial of Herceptin.
Due to the diagnostic test’s ability to enrich the study population with drug responders, a clinical trial was able to show that Herceptin lengthened the survival time of about 25 percent of women with metastatic breast cancer who overexpress the HER2 gene. If the study population had not been enriched with responders, a mathematical model revealed the clinical efficacy of the drug would have been difficult, if not impossible, to demonstrate with the number of patients typically recruited for a clinical trial. “This is clearly a huge success that raised the paradigm for personalized medicine and predictive tests in targeted therapies,” Dr. Waring said.
The diagnostic assay used to select patients for the clinical trial proved to be unsuitable for commercialization, however, so Genentech partnered with DAKO to codevelop an immunohistochemistry (IHC) diagnostic test, known as the HercepTest®, which is now widely used in clinical practice. This test was validated during the Phase III clinical trial by showing equivalence to the clinical trial assay. Both the drug and the test were approved jointly by the FDA in September 1998.
Studies have shown that there are high false-positive (Table 2) and false-negative (Table 3) rates in the general community for HER2 immunohisto-chemistry testing as well as the more accurate fluorescent in situ hybridization (FISH) test. Although large, more experienced laboratories generally perform both these tests well with low false-positive and -negative rates, small-volume laboratories, particularly those that use home-brew immuno-histochemistry tests, were shown in these studies to have unacceptably high false-positive and false-negative rates, Dr. Waring reported. “The problem isn’t so much with the tests themselves, but where the tests are performed,” he said. Genentech’s estimation of the situation is that each year about 5,000 U.S. patients receive Herceptin without any clinical benefit, and about 7,000 patients who could derive benefit are not being treated because of a false-negative test result. “This keeps me awake at night and is a very serious problem,” Dr. Waring said.
To rectify this situation, Dr. Waring recommended implementation of standardized testing and mandatory participation in HER2 quality assurance testing programs akin to what is in place in the United Kingdom. Such best practice programs allow laboratories to compare their performance against reference materials and other laboratories and hence identify whether they have a testing problem (Ellis et al., 2004). The accompanying educational material and instructional assistance allows most laboratories to identify and rectify their problems. In the UK HER2 QAP program, which publishes its collective results (Rhodes et al., 2004), retesting of over 100 European laboratories on 6 successive occasions resulted, over a 2 year period, in a significant improvement in the number of laboratories achieving acceptable HER2 test results. Dr. Waring added that “generally, the pathology community isn’t ready yet in many ways to adopt these predictive tests for therapeutic decision making. I think for more sophisticated tests, they’re going to have to be performed in central reference laboratories that have very rigorous accreditation processes.”
Dr. Waring described more recent and less successful attempts to develop diagnostic tests that predict responsiveness to targeted cancer therapies. He discussed the DAKO test for expression of epidermal growth factor receptor (EGFR), which was used to detect colorectal cancer patients likely to respond to cetuximab. Colorectal cancer patients were not entered into the clinical trials of cetuximab unless they had a positive result in the EGFR test (had 1 percent or greater tumor cells showing positivity). These trials revealed that between 10 and 20 percent of patients responded, and led to the approval of the drug by the FDA in 2004.
TABLE 2 False-Positive HER2 Test Results
|
|
Local vs. Central FISH |
Local vs. Central HercepT |
Local HercepTest vs. Central |
Local Homebrew vs. Central est FISH HercepTest |
|
NCCTG N9831 (n=970) |
15% |
20% |
— |
31% |
|
B-31 (n=104) |
— |
14% |
21% |
— |
|
Small volume (n=79) |
— |
19% |
23% |
48% |
|
Large volume (n=24) |
— |
4% |
4% |
0% |
|
B-31 amendment (n=204) |
2% overall |
2% overall |
2% overall |
2% overall |
|
HER-First (n=1,434) |
— |
23% (any IHC) |
26% (any IHC) |
— |
|
SOURCES: Adapted from Waring (2006). Adapted from Perez et al. (2004), Paik et al. (2002), Reddy et al. (2006). |
||||
TABLE 3 False-Negative HER2 Test Results
|
|
Local vs. Central FISH |
Local vs. Central HercepTest |
Local HercepTest vs. Central |
Local Homebrew vs. Central HercepTest |
|
NCCTG N9831 |
15% |
20% |
— |
31% |
|
N9831 (n=970) |
— |
— |
— |
— |
|
B-31 (n=104) |
— |
— |
— |
— |
|
Small volume (n=79) |
— |
— |
— |
— |
|
Large volume (n=24) |
— |
— |
— |
— |
|
B-31 amendment (n=204) |
— |
— |
— |
— |
|
HER-First (n=1,434) |
— |
11% (any IHC) |
14% (any IHC) |
— |
|
SOURCES: Adapted from Waring (2006). Adapted from Reddy et al. (2006). |
||||
But an analysis of the trials and other studies has revealed that there is no correlation between clinical benefit and EGFR positivity, either by the number of positive cells or by staining intensity, Dr. Waring pointed out. This is probably because the staining pattern for EGFR is quite heterogeneous, he said. Some tumors may only show focal areas that are positive, so a positive result may depend on which piece of the tumor is examined. “The EGFR test was able to accelerate or increase the probability that cetuximab would be approved and in that regard it was a success. But I don’t think it has been a success in terms of testing in community practice,” said Dr. Waring.
He also pointed out that although initial studies indicated that more than 70 percent of the responders to Tarceva had mutations in EGFR, testing positive did not correlate with a survival advantage in small cell lung cancer patients because of the complex biology of the disease. Studies have shown that although patients who have EGFR mutations initially respond to these drugs, surviving tumor cells may acquire secondary resistance mutations and then progress, resulting in no survival benefit. But the drug also slows the growth of tumors in patients who do not have EGFR mutations, which can result in improved survival time.
Dr. Waring concluded his talk by suggesting ways to enhance harnessing the power of cancer biomarkers. He recommended designing and powering clinical trials to answer diagnostic questions as well as therapeutic questions. Although large numbers of patients are accrued to clinical trials of cancer drugs, many of their samples are not available or are of inadequate quantity or quality to enable the testing needed to find a molecular signature that correlates with clinical outcome. He also recommended that the clinical utility of predictive diagnostic tests be demonstrated. The test has to significantly impact therapeutic decision making, he said. He also noted the importance of making distinctions between clinical assays used to enroll patients in clinical trials of unproven therapies versus those used to test patients in clinical practice prior to making therapeutic decisions.
Diagnostics Industry Perspective
The next talk was given by Robert Lipshutz, PhD, of Affymetrix. Dr. Lipshutz gave the diagnostics industry perspective on incentives and disincentives to develop biomarker-based cancer diagnostics whose utility is linked to targeted therapies. He noted that companies such as Affymetrix, which develop the platforms used in many microarray diagnostic tests, often
partner with drug companies, diagnostic companies, academic institutions, and/or Clinical Laboratory Improvement Amendments (CLIA)-certified laboratories to move a diagnostic test from its initial discovery to its commercial application in a clinical setting. To encourage academic medical centers to undertake biomarker discovery endeavors using their gene testing platform, Affymetrix will offer them special pricing so as to share the risk of pursuing such discovery efforts, as well as the opportunity to develop any biomarkers discovered by working with its partners—companies that will validate and bring the biomarker-based tests to market.
Although the cost of developing a diagnostic is relatively small compared to the cost of developing a drug, so too is the overall return, Dr. Lipshutz noted. “You don’t have a lot of blockbuster diagnostics on the market,” he said. A number of costs and risks are linked to every step of developing a biomarker-based test, he pointed out. If the diagnostic is only going to be useful if the targeted therapy gains FDA approval, the risk of the new drug failing clinical trials must be added to the risk of developing a new diagnostic. This is a major disincentive for diagnostic companies, he noted.
But on the plus side, if the drug does make it to market with its label requiring the diagnostic test, then there is reduced cost and risk linked to marketing the diagnostic because both are shared with the company that developed the new drug. However, its reimbursement rate may be too low for the diagnostic company to earn enough revenue on the test, even if there is a huge demand for it. Also, Dr. Lipshutz noted that the actual market for diagnostics linked to specific cancer therapeutics is smaller than the average diagnostic test, and thus is likely to generate less revenue. This, too, can act as a disincentive to diagnostic companies.
For simple diagnostics, such as the IHC tests already on the market, the costs and risks were low so it was relatively easy to have diagnostic companies develop these tests, Dr. Lipshutz said. But they may be less inclined to develop more complicated diagnostics that might have to undergo an extensive in vitro diagnostics (IVD) approval process with the FDA to reach the market, he added. He explained the IVD process has more extensive testing requirements than the home-brew development process often used for diagnostic tests, which only requires CLIA certification of the laboratory performing the test (Figure 5).
Dr. Lipshutz suggested an alternative regulatory model to reduce the risks and costs of developing biomarker diagnostic tests linked to new targeted cancer treatments. In this model, researchers should use a biomarker
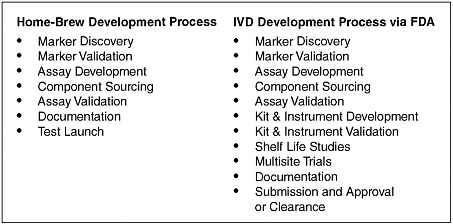

FIGURE 5 IVD developmental process for FDA approval or clearance vs. home-brew test development.
SOURCE: Adapted from Lipshutz presentation (March 20, 2006).
assay that meets CLIA requirements during clinical trials of the new therapeutic for which its use will be linked. If the drug is then approved by the FDA, the diagnostic test would also enter the market via CLIA-certified labs. But linked to the diagnostic approval would be the stipulation that further testing be done for the diagnostic test so it is later evaluated by the FDA as an IVD.
Dr. Lipshutz concluded his talk by pointing out the need for improved standards for sample preparation and controls for expression reagents, SNPs, and copy number. He also reiterated the need for statistical standards to evaluate the patterns seen in the omics field.
NCI Perspective
The next speaker was James Doroshow, MD, of the National Cancer Institute (NCI), who discussed the agency’s goals and funding initiatives in regard to cancer biomarkers. He pointed out a number of new initiatives the agency has undertaken that should further the cancer biomarker field.
One of these is a $100 million investment in a program to develop and test new animal models molecularly engineered to mimic human cancers. These animal models can be used to predict the pharmacodynamics for new cancer drugs, and can ease the development of assays that can predict effectiveness or safety of new drugs in clinical trials. In a later presentation,
Dr. Charles Sawyers, MD, of the University of California, Los Angeles, stressed the usefulness of these animal models in the discovery of genetic signatures that not only indicate promising genetic targets for drugs, but that can be used to test patients for the presence of such targets. He also noted that the predictive power of preclinical models could be transformed by parallel experiments in genetically engineered mice.
The NCI also increased its support of efforts to develop and validate pharmacodynamic in vitro assays well in advance of early phase clinical trials. In addition, it recently opened a new laboratory in its Frederick, Maryland facility to develop its molecular toxicology profiling capabilities so as to speed the development of new agents. The agency also has an extensive collection of synthetic and natural products, as well as biologics and tumor and animal cell lines or models that are provided free-of-charge to cancer researchers.
The Institute supports several programs designed to supplement the limited resources in academia to support the transition from molecular targets to drugs. These include the National Cooperative Drug Discovery Group, which is a consortium of about seven or eight academic groups and pharmaceutical companies. Over the past 20 years, this consortium received about $200 million in NCI funds, and in return has generated efforts that led to the approval of five new cancer drugs, including cetuximab.
NCI’s Rapid Access to NCI Discovery Resources program develops assays for investigators who submit promising model targets that survive the competitive external review process. The NCI’s Rapid Access to Intervention Development program provides the expertise of its staff and additional in-house resources to academic or nonprofit investigators in the extramural community. These individuals compete to have NCI develop their lead compounds into those suitable for submission into clinical trials. Such development may include pharmacology or toxicology studies, efficacy studies in animals, or the formulation of bulk drug. During its nearly 8-year existence, the program has fostered 24 investigational new drug applications at the FDA, Dr. Doroshow reported.
He acknowledged the need for more NCI resources earmarked to supporting the development of biomarker assays, including validation efforts. “It’s almost impossible now to get a peer-reviewed grant to develop an assay. That’s something we either have to correct in terms of the peer review process, or by doing the assays for our investigators that we work with closely,” he said. NCI is currently developing a new program to address this shortcoming, he added.
Clinical Investigator Perspective
Dr. Sawyers gave the clinical investigator perspective on the discovery and development of cancer biomarkers useful in predicting response to targeted therapies. He began his presentation by showing how developments in his lab led to the discovery of a genetic test for predicting resistance to Gleevec or other drugs that target the BCR-ABL translocation in chronic myelogenous leukemia patients. Because this test was simple to develop, it was not difficult to convince a diagnostic company to undertake this project, and the test was launched commercially this past year.
In contrast, when researchers at the University of California, San Francisco, and the University of California, Los Angeles, discovered two biomarkers that predicted response to EGFR inhibitors in glioblastoma patients, the discovery was not readily adopted and developed by a diagnostics company. These biomarkers were more challenging to develop into an assay, according to Dr. Sawyers, because they consisted of two noncommercial antibodies that would probably be quickly outdated by DNA-based diagnostics. Reluctance to develop the assay also stemmed from the likelihood that it would only be applied to the relatively small number of glioblastoma patients, rather than a larger market. Recent movement away from the standard of single-drug treatment for glioblastoma to multiple-drug therapy also made it difficult to confirm the effectiveness of the assay, he added. Because no diagnostics company has developed the assay, only the original discoverers of the biomarkers use them to test glioblastoma patients. Their labs are not really set up to do such extensive testing, Dr. Sawyers noted.
An important deterrent to academic researchers discovering and developing cancer biomarkers is the high cost associated with such efforts, he pointed out. Genomic tests can add more than $1 million to the cost of running a clinical trial, he estimated. “I personally feel it is worth making these investments to do the experiment, but as many of us know, it’s not easy to come up with those kinds of funds, even if a trial is actually quite compelling,” he said.
Drs. Sawyers and Lipshutz also addressed intellectual property issues. Dr. Sawyers noted that the kinds of information generated from genetic signature analyses are going to be broadly useful because “there will be a limited number of cancer pathways and lots of drugs will be going at these same pathways from different companies and different angles. So there will be a need for a broad base of pathway markers and I see them as sort of
precompetitive knowledge.” Discovery costs for those pathways and biomarkers should be shared among academia and pharmaceutical, platform, and diagnostics companies, he said. Incentives for commercialization of molecular diagnostic assays must be retained without compromising the need for open access to data, he added. Such open access is critical for metaanalysis of datasets from different trials.
Barbara Weber, MD, a representative from GlaxoSmithKline, noted that her company concurs with Dr. Sawyer’s view that biomarker efforts should be precompetitive. Her drug company has released publicly and immediately all its biomarker data in the hopes of encouraging other large pharmaceutical companies to do the same. “The competitive advantage comes from having good molecules that get properly developed, and we can only benefit by making those data publicly available,” she said.
In his talk, Dr. Lipshutz discussed how intellectual property uncertainties can act as a disincentive for diagnostic companies to develop tests that may require the licensing of multiple sources of genetic information. For example, one company that uses Affymetrix’s microarray platform plans to use a few hundred genes for their diagnostic tests, but they estimate they would have to examine 20,000 pieces of intellectual property patents before pursuing such tests. Dr. Lipshutz deplored the patenting of natural products and natural laws, which he called patenting obvious information. The Supreme Court is currently evaluating one such patent case7 and its decision will impact the diagnostics arena, he said.
In the meantime, it has been proposed that patent pools be established so there can be “one-stop shopping” to gain access to all the genetic or other such information needed for a diagnostic test. He also suggested the academic community develop more rational economic models and best practice guidelines for the licensing of intellectual property patents.
BIOMARKER DEVELOPMENT AND REGULATORY OVERSIGHT
Biomarker assays are often widely marketed as laboratory services, without FDA clearance or approval. Such assays usually have undergone analytical validation, which indicates the laboratory accuracy of the tests for detecting what they are supposed to detect. But often there are scanty clinical data on predictive value, such as how accurately the tests determine
a clinical parameter such as disease diagnosis. However, biomarker tests used to screen for or to diagnose cancer, or to develop a treatment plan have considerable potential for harm as well as benefit. As sophisticated biomarker tests that take advantage of the latest developments in molecular biology begin to enter the market, questions have been raised regarding the level of oversight that is warranted for them. The fourth session of the conference explored recent FDA initiatives regarding biomarkers, ways to design new drug clinical trials that use biomarkers, and how biomarkers should be regulated.
FDA Critical Path Initiative
Janet Woodcock, MD, of the FDA opened the session by noting the recent explosion of new scientific knowledge, particularly within molecular biology, and the doubling of funding that biomedical research has received in the past decade. Yet paradoxically, 2004 marked a 20-year low in the introduction of new molecular-entity drugs on the international market, and there has been a decade-long downward trend for new drugs and biologics submitted to be evaluated by the FDA.
To address this mismatch between innovations in biomedical research and lack of a corresponding surge in novel drugs, the FDA issued a white paper in March 2004 called “Innovation or Stagnation: Challenges and Opportunities on the Critical Path to New Medical Products.” The paper noted that this mismatch was caused by using 20th-century tools to evaluate 21st-century advances, and that there is a need to apply new science to the tools used to evaluate new medical products. This is especially true regarding biomarkers, the paper pointed out.
Dr. Woodcock noted that despite the hundreds of candidate biomarkers that are published each year, few ever reach a high enough level of clinical correlation to enable decisions in product development or patient management. “Getting that clinical correlation information that we need is very difficult and costly and it just isn’t done,” she said. “The process for developing biomarkers for various uses is really broken.”
She pointed out that a biomarker is not the same as the assay that is developed to analyze the biomarker, and that this assay requires analytical validation. But it is not yet known how to best prove the performance characteristics of a biomarker-based test that employs newer technologies, especially because many lack a gold standard for comparison. She also stressed the wide range of biomarker uses from pharmacodynamic assays to
disease diagnosis, and reiterated the need to tailor the qualification package to the biomarker’s intended use. For example, an assay used to screen for disease has a much higher bar than a pharmacodynamic assay used in a drug development program. Dr. Woodcock stated that the agency plans to clarify its regulatory acceptance of biomarkers for various uses in future draft guidances.
New trial designs and methods are needed that incorporate biomarkers, especially if there is codevelopment of a diagnostic and a therapeutic, the FDA white paper also pointed out. These trials should use biomarkers that predict patient responders to make the trial more efficient and informative. “The clinical trial process has been highly observational in its conduct, primarily because we don’t have the tools to look at the basis for individual response so we look at population responses. But these trials are extremely expensive and it really is important that we get maximum information when we subject human subjects to experiments.”
The FDA white paper also called for more development of bioinformatics, which would encourage the sharing of data and databases so that “we can learn generalizable knowledge about biomarkers, rather than knowledge that simply stays in a particular trial or drug development program,” Dr. Woodcock said. There should be standardization of terminology to allow pooling of data and construction of computer-based, quantitative disease models in which biomarker performance data can be incorporated for trial modeling and simulation, she added.
Dr. Woodcock ended her talk by describing the public-private consortia the FDA has fostered to support biomarker development. These include the Critical Path Institute. This nonprofit institute is a consortium consisting of pharmaceutical industry partners with the goal of qualifying new animal safety biomarkers for predicting human toxicities. The companies that participate in this consortium share and cross-validate existing proprietary markers and data that are accrued on them. Another consortium that is under way is an outgrowth of the Oncology Biomarker Qualification Initiative created to join the FDA, NCI, and Centers for Medicare & Medicaid Services (CMS) efforts to foster biomarker development. This led to the development of a nonprofit public-private partnership to qualify fluoro-2-deoxy-D-glucose (FDG)-PET scanning as a marker for drug response in non-Hodgkin’s lymphoma.
These consortia are vital, Dr. Woodcock asserted, because “the availability of biomarkers is a common good. It is good for patients and clinicians as well as for researchers and medical-product developers. One
company, research or funding source is unlikely to have adequate resources to complete the needed work.”
Oversight of Diagnostic Tests
Mr. Heller, a partner at Wilmer Cutler Pickering Hale and Dorr, LLP, gave the next talk. Mr. Heller discussed the FDA’s role in regulating biomarker tests and explored some recent precedent-setting initiatives the FDA has taken in regard to regulating innovative biomarker-based assays. Mr. Heller began his talk by stating that the FDA has regulatory jurisdiction over all in vitro instruments and reagents that are “intended for use in the diagnosis of disease or other conditions, or in the cure, mitigation, treatment, or prevention of disease” in the human population because they are considered devices.
The FDA defines “intended use” as the objective intent of persons legally responsible for labeling a device. In order to determine intended use, the FDA closely considers a device marketer’s advertising, labeling claims, product distribution, product websites, and other objective information, said Mr. Heller. The FDA does not regulate in vitro devices that are intended for research purposes only. Instead, the sellers of such devices must comply with a labeling requirement that states the product is for research only and not for clinical purposes. But, “the amount of grayness that attaches to research-only status is profound, and it is something the agency has been wrestling with for years,” said Mr. Heller. If someone markets a product for research use and is aware that it is used diagnostically, the agency can assert jurisdiction, and regulate the assay as a device. When the agency asserts jurisdiction, this typically results in premarket submissions to the FDA under its premarket approval (PMA)8 or premarket notification (510[k])9 requirements.
Mr. Heller discussed how “home-brew” tests, those that are developed by a laboratory in-house for in-house use, present regulatory challenges to the FDA. The FDA, through an exercise of its enforcement discretion,
has withheld its authority to regulate home-brew diagnostic tests, thus not requiring premarket submissions before their commercial use. Because the data needed for premarket applications are costly and time consuming to procure and assemble, that regulatory treatment appeals to laboratories who devise tests that are essentially in competition with commercially available assays.
Home-brew tests are subject to the regulations of the Clinical Laboratory Improvement Amendments (CLIA), which mandate that each lab create its own performance specification and provide evidence of accuracy, reproducibility, and analytic specificity for the target patient population of a home-brew test. But Mr. Heller emphasized that although the FDA does not regulate laboratories, it asserts that it has jurisdiction to do so, and the CLIA does not displace the Federal Food, Drug, and Cosmetic Act. “The agency’s choice not to enter laboratories, I think, represents a resource judgment and a sensitive approach to prioritizing resources,” Mr. Heller said.
But this self-imposed limitation of the FDA raises some potential problems, according to Mr. Heller. For example, he noted that laboratories can license intellectual property for home-brew tests to other laboratories. He suggested that “from a public health point of view, there is very little difference in whether the test moves through commerce itself or the IP is licensed and then the test is performed pursuant to a specific recipe, with royalties paid for each test performed.” “As things go forward, I think this will present a challenge to [the] FDA and maybe suggests [the need for] a modern means of regulation, including possibly statutory adjustment,” he added.
In order to maximize its efficiency in regulating and ensuring the safety and effectiveness of home-brew tests, the FDA regulates commercial analyte-specific reagents (ASRs), which are used to develop home-brew tests.10,11 ASRs are defined as “antibodies, both polyclonal and monoclonal,
specific receptor proteins, ligands, nucleic acid sequences, and similar reagents which, through specific binding or chemical reaction with substances in a specimen, are intended for use in a diagnostic application for identification and quantification of an individual chemical substance or ligand in biological specimens.”12 Laboratories that produce ASRs must register with the FDA and satisfy the agency’s Quality System Regulation (good manufacturing practices). However, most ASRs are not subject to premarket review. Mr. Heller noted that sellers of reagents assert that “many products [on] the market are either research-use only or analyte-specific reagents, whether they necessarily meet those clear definitions or not.”
Mr. Heller briefly described an instance in which the FDA made a decision to regulate a microarray product as a device based on its intended clinical use despite the manufacturer’s characterization of the product as an ASR, which does not require premarket review. Specifically, Roche Molecular Diagnostics planned to introduce a microarray genetic test for drug metabolism (AmpliChip CYP 450) into marketplace in 2003. After reviewing the product and requesting information from the company, the FDA decided that the product was “of substantial importance in preventing impairment of human health” and its technological characteristics “would cause it to differ from existing or reasonably foreseeable ASRs.” This determination resulted in denying a 510(k) exempt status accorded to Class I ASRs and resulted in the requirement to submit a premarket notification before marketing.13 The FDA suggested that if the device were found to be not substantially equivalent, the company could seek de novo classification. De novo classification became part of the Federal Food, Drug, and Cosmetic Act in 1997 to provide the FDA with a cost-effective means of avoiding an automatic classification of novel devices into a Class III, PMA status. If a novel device has a lower risk profile that permits the device to be regulated in Class I or II, then the agency has 60 days after receiving a request for de novo to classify the device.14 In this case, both Roche’s microarray and Affymetrix’s scanner used with the microarray were found not substantially equivalent to a predicate device and both were placed into Class II under
|
12 |
21 C.F.R. § 864.4020(a). |
|
13 |
Letter from OIVD to Roche Molecular Diagnostics Re: AmpliChip, http://www.fda.gov/cdrh/oivd/amplichip.html. |
|
14 |
In order to be eligible for de novo classification, a 510(k) submitter must submit a request to the agency within 30 days of receiving a not substantially equivalent order proposing and justifying a Class I or II classification. |
the de novo classification procedure. As a result, each was marketed without a PMA.
Mr. Heller noted that the FDA is very interested in molecular diagnostics and is still trying to determine to what extent it will implement its jurisdiction over new diagnostic devices, as indicated by a number of recent FDA activities. He gave several examples where the FDA asserted regulatory authority over products that manufacturers thought would be outside of the FDA’s jurisdiction. Mr. Heller described a meeting and letters in 2004 between the FDA and the developer of a new serum protein test that used mass spectrometry for ovarian cancer screening (OvaCheck). After reviewing the product information and corresponding with the developer, the FDA allowed tests to be run in labs under CLIA without premarket review, but it considered the software used to analyze the results to be a device subject to its regulation and requiring premarket approval.
An April 2003 FDA draft guidance (which is non-binding) on multiplex genetic tests states that tests that interrogate several analytes are not ASRs and require premarket submissions. The focus of the document is on nucleic-acid-based analyses, but the guidance also indicates that it is applicable to protein and tissue arrays. Based on this guidance document, the FDA sent a warning letter to the Nanogen Corporation on August 11, 2005, in which it wrote that the Nanochip Molecular Biology Workstation, Nanochip Electronic Microarray, and several ASRs were not approved as a system or as components. The agency was concerned that the NanoChip array, and the system as a whole, would be used in clinical diagnostics, and was therefore not a research-use only product, as the company had alleged. Similarly, that same month, Access Genetics received a warning letter regarding marketing of test packages for several genetic tests. In addition to notifying a company of concerns with its practices, warning letters can be used by the FDA to clarify how it defines boundaries for its regulatory jurisdiction, according to Mr. Heller.
Mr. Heller noted that biomarker tests used to identify likely responders to drugs will be regulated as devices in parallel with their corresponding drug candidates, and those for higher risk conditions will require PMAs. He added that the FDA Guidance on Pharmacogenomic Data Submissions (March 2005) recommends submitting pharmacogenomic data when the data will be used to make approval-related decisions and when the data are relied upon to define, for example, trial inclusion or exclusion criteria, the assessment for prognosis, dosing, or labeling, or used to support the safety and efficacy of a drug. If a test shows promise for enhancing dosing,
safety or effectiveness, or will be specifically referenced on the label, the FDA recommends co-development of the device and drug.
In its April 2005 concept paper on co-development, the FDA addressed the use of a single test with a single drug. Co-development applies when use of an in vitro diagnostic is mandatory for drug selection for patients, or when optional use during drug development may assist in understanding disease mechanisms and in selecting clinical trial populations. Co-development applies to a device/drug combination product, as well as to in vitro devices and drugs sold separately. The concept paper on co-development explicitly states that drug selection biomarkers, particularly for high-risk conditions, are expected to be subject to PMAs.
Mr. Heller concluded his talk by noting that because more than one center at the FDA will often be involved in approval or clearance decisions, the agency should focus on ensuring coordination among its centers to facilitate the clearance or approval of molecular diagnostics. He suggested that the agency should also focus on clarifying which in vitro tests are considered research only, the FDA’s role in regulating or not regulating labs, the agency’s reliance on the CLIA, and what does and does not constitute an ASR. “Frankly, without these understandings, many folks have products out there, some in perfectly good faith, not knowing that, from an agency perspective, they may be in violation of the law,” he said. He added that except for the highest risk in vitro diagnostic devices, the FDA should seriously consider de novo classification as the standard means of clearing novel molecular diagnostics to ensure safety and effectiveness, so that important diagnostics/prognostics reach health care professionals and patients as soon as possible.
Designing Clinical Studies of Biomarkers
The next speaker was Richard Simon, DSc, of the NCI. Dr. Simon focused on ways that biomarkers are transforming the design of clinical trials, and how they should be appropriately regulated. The conventional wisdom is that there should be broad eligibility of patients in clinical trials. But this notion is outdated now that there is increasing evidence that many kinds of cancers are heterogeneous in pathogenesis and sensitivity to treatment. This results in the effectiveness of many drugs being missed in traditional clinical trials because the proportion of patients who would benefit from the drug was too small to make its presence felt among the majority. “I think it is almost the rule, rather than the exception in cancer therapy,
that we treat the majority for the benefit of the minority,” Dr. Simon observed.
Instead, he noted that enriching trial populations with likely responders not only will reduce the cost of a clinical trial, but will make it more likely that participants will benefit from the drug being tested. “Cancer clinical trials of molecularly targeted agents may benefit a relatively small proportion of patients, but the benefit for the sensitive subset can be very substantial,” he pointed out. New cancer drug development, consequently, increasingly relies on a biomarker classifier that selects a target patient population for treatment. However, the focus of a clinical trial that uses a classifier is to evaluate the effectiveness of a new drug, not to validate the classifier, he said.
Dr. Simon gave several examples of how clinical trials could be designed to incorporate a classifier. In one trial design, a classifier that predicts responsiveness is used to restrict the eligibility of patients to a prospectively planned evaluation of a new drug such that only those who “pass” the responsiveness test are entered into the study and randomized into treatment or control groups (Figure 6).
In another trial design, the responsiveness diagnostic is not used to restrict eligibility, but to structure a prospective analysis plan. The purpose of this trial is to evaluate treatment versus control overall, as well as for a predefined subset of likely responders (Figure 7). The purpose of the trial is neither to reevaluate the components of the classifier, nor to modify or refine it, Dr. Simon stressed.
In the second study design, effectiveness of the new drug in patients is compared to results in controls in the overall study population. If statistical significance (p less than .04) is found, one can claim effectiveness of the drug for the eligible population as a whole. Otherwise, one would perform a single subset analysis that evaluates the drug in the classifier-positive patients, and would claim effectiveness for these patients if statistical significance (p less than .01) is found. The overall study type 1 error of .05 is split between the overall test and the subset test.
The second study design is commonly used when there is not complete confidence that the biomarker used as a classifier will predict response, Dr. Simon noted. The key features of this trial design are that it has a prespecified analysis plan with a single predefined subset. “Saying that the study should be stratified is not enough. You really need a completely well-defined analysis plan as to how you are going to use that subset,” he said.
One can size the trial based on what is needed for the overall analysis. If the results in this analysis are not statistically significant, one could continue accruing for the predetermined subset until a large enough population is reached for a subset analysis. Alternatively, if an interim analysis reveals that there is a large treatment effect for the subset, then one could continue to accrue the classifier-negative patients until there is a large enough population to assess whether the new drug benefits them as well.
A guiding principle for all these study designs is that the data used to develop the classifier must be distinct from the data used to test the hypothesis about treatment effect in subsets determined by the classifier, Dr. Simon pointed out. He added that archived samples from a conventional nontargeted clinical trial could be used to develop the classifier of a subset of likely responders. That subset hypothesis would then be tested in a separate trial. But he noted that it is not possible to use many genetic analysis techniques on archived samples because of the way the samples are preserved.
Dr. Simon concluded his talk by asserting that extensive FDA regulation of biomarkers used in clinical trials is not appropriate. There should be no requirement for demonstrating that the classifier or any of its components are “validated biomarkers of disease status” nor should one have to repeat the classifier development process on independent data, he said. He also does not believe the FDA should regulate how DNA microarrays are used for classifier development in early (Phase I and II) clinical trials.
“If we have developed a classifier in Phase I and Phase II studies, we need to know that we can reproducibly measure that with some assay, and then we need to know something about treatment effect on the subset determined by that classifier. But I don’t think it is appropriate to regulate all of the possible ways we could develop that classifier,” Dr. Simon said. “[The] FDA can slow effective utilization of this technology, either by over-regulating classifier development or by not providing sponsors with a clear and practical roadmap of what is required.” He added that some aspects of the FDA guidelines on biomarkers are inappropriate for treatment selection biomarkers.
ASSESSMENT AND ADOPTION OF BIOMARKER-BASED TECHNOLOGIES
Once cancer biomarker tests enter the market, they have to overcome additional hurdles before they are widely used clinically. How readily
biomarker tests are adopted in the clinic depends, in part, on how extensively they are reimbursed by health insurers, and how highly they are recommended by various organizations, particularly those that promulgate practice guidelines. Reimbursement policy, in turn, can impact industry marketing and development strategies. The goal of the fifth session of the conference was to examine current and developing strategies for medical decision making and insurance coverage of biomarker-based tests.
Federal Programs for Technology Assessment
Alfred Berg, MD, MPH, of the University of Washington began this session by recounting how the United States Preventive Services Task Force (USPSTF) generates its evidence-based practice recommendations. These recommendations, although not officially binding, generally become the standard of care for medical practice in the United States. A member of the current USPSTF, Dr. Berg explained that it is a rotating, interdisciplinary panel, which regularly publishes its guidelines and recommendations on the web.15 Its mission is to produce scientific evidence-based reviews of preventive interventions given to asymptomatic patients in primary-care clinical settings.
Prior to conducting their reviews, the Task Force selects a panel of expert generalists who have not already taken public stands on the preventive intervention the panel is reviewing. The analytical framework for the review is specified in advance. It includes assessing how an intervention affects morbidity and mortality, as well as what adverse effects are linked to the intervention, and how the benefits and risks compare to those of standard treatment. The panel does an explicit and prospective quality review of relevant journal articles that meet its stringent criteria. “I emphasize prospective,” said Dr. Berg. “We feel strongly that one should specify in advance exactly what you are looking for and not change your mind once you get into the literature.”
The reviews are then summarized in evidence tables and the literature is formally linked to recommendations and clinical discussion. Recommendations for interventions that have a net benefit are coded A, B, or C, with the most benefit seen in A recommendations, and the smallest seen in C designations. The C designation is essentially no recommendation because there is fair to good evidence that the benefits and harms are closely
balanced. Interventions with zero benefit or those that have negative net effects are coded D, and those for whom the evidence is poor are termed I. The quality of the evidence is also considered. To receive an A recommendation, for example, there must be good-quality evidence of a substantial benefit. A substantial benefit seen in a poorly controlled study will not suffice (Table 4).
Many recommendations end up in I territory, Dr. Berg noted. I stands for insufficient because the evidence is insufficient to recommend for or against the intervention. It can be insufficient due to poor quality of the studies done on the intervention, or a lack of studies. An I rating is also given if there are good-quality studies, but their results conflict with each other.
The Task Force recently reviewed the evidence regarding screening for prostate cancer with the PSA test. It gave the use of this test for this purpose a designation of I. Although it found good evidence that screening can detect early stage prostate cancer, there was mixed and inconclusive evidence that such early detection improves health outcomes. In addition, it found very strong evidence that screening and subsequent treatment are both linked to important harms, and concluded that the benefits of treating early prostate cancer are unknown. “The conclusion is not to not do PSA screening,” Dr. Berg noted. “The conclusion is that the evidence is insufficient to be able to give clear advice. Our advice to clinicians is that if you are going to do it, do it with care and make sure the patient knows what [he is] getting into.”
As Dr. Berg pointed out, the infrequent patient who “wins the lottery” and has a lethal prostate cancer detected at an early stage with PSA screening could receive enormous benefit from such detection. But most screened patients will not receive that benefit. Studies suggest that to prevent one death from prostate cancer in 8 years, one would have to screen about 1,000 men with the PSA. These men would be subject to such potential harms
TABLE 4 Recommendation Codes
|
Quality of Evidence |
Net Benefit |
||||
|
Substantial |
Moderate |
Small |
Zero/Negative |
||
|
Good |
A |
B |
|
C |
D |
|
Fair |
B |
B |
|
C |
D |
|
Poor |
|
I |
|
||
|
SOURCE: Adapted from Berg presentation (March 21, 2006). |
|||||
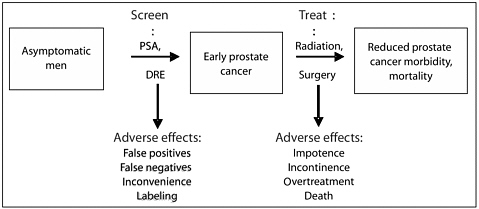
FIGURE 8 Analytic framework for prostate cancer.
SOURCE: Adapted from Berg presentation (March 21, 2006).
as false-positive tests, anxiety, and treatment-linked erectile dysfunction, incontinence, and bowel dysfunction (Figure 8).
“This is a classic dilemma for the patient and the clinician trying to decide whether prostate cancer screening is a good idea for one personally; trying to balance the potential for an enormous benefit against a somewhat more likely potential for harm,” Dr. Berg said. The conclusion of the Task Force was basically to let the patient decide whether he wants to receive PSA screening after age 50.
Dr. Berg summed up the findings of the Task Force by noting that their review of biomarker-based tests and other screening tests for skin, bladder, lung, ovarian, pancreatic, oral, and testicular cancer led to I or D recommendations. The only cancer screening tests they actually recommended with B or A ratings were for breast, colorectal, or cervical cancer, and none were biomarker-based tests.
Dr. Berg finished his talk by discussing a new panel sponsored by the Centers for Disease Control and Prevention called the Evaluation of Genomic Applications in Practice and Prevention (EGAPP). Like the Preventive Services Task Force, it is a nonregulatory panel that is expected to make evidence-based recommendations. The goal of EGAPP is to establish and evaluate a systematic and sustainable mechanism for premarket and postmarket assessments of genomic tests in the United States.
“Screening tests are often implemented before the science is fully in place,” Dr. Berg noted. “A concern shared by clinicians, patients, regula-
tors, and insurance carriers is that some genomic tests may be released and marketed prematurely. So one of the things that EGAPP hopes to do is to collect what information we do have and assist folks in making a more informed decision.”
EGAPP is in the second year of its 3-year existence. It has developed a number of brief summaries of genetic tests. (It chose to review those tests based on health burden of the applicable diseases, and availability, misuse, or impact of the tests.) It is currently working on developing an appropriate analytic framework, as well as a study search strategy and standard for assessing study quality that can be used to review genomic clinical tests. The panel has defined the relevant categories of outcomes for genomic tests. In addition to considering how the test will affect diagnostic determinations, therapeutic choice, and patient outcomes, EGAPP also considers the impact of the test on the families that are related to the person being tested, as well as the impact to society at large.
EGAPP has reviews under way for tests for the drug and toxin metabolizing enzyme, CYP450; the genetic biomarker for colon cancer, HNPCC; and ovarian cancer screening, for which it will be testing its methods. It also plans to do fast-track reviews for tests that have limited data. These include the test for EGFR, and a test for UGT1A1, a drug and toxin metabolizing enzyme that affects susceptibility to chemotherapy side effects. The final expected outcome of the panel is three to five major reviews, two to three fast-track reviews, and a document on methods and evaluation.
Dr. Berg noted that his work on the panel made him aware that there is a lack of information on many important areas related to genetic tests, such as the frequency of genetic variation in the general population, and gene penetrance (what percentage of people with a specific gene allele actually express the allele and show its corresponding phenotype). There are also few clinical trials that compare a genomic intervention with no intervention, and many studies do not assess all the relevant outcomes, he said. Often little attention is paid to documenting the harms of a genomic test, or to its cost and feasibility. Instead, most attention is focused on the potential benefits of a particular test.
Insurance Coverage Decisions and Practice Guidelines
The next speaker was William McGivney, PhD, of the National Comprehensive Cancer Network (NCCN). Dr. McGivney previously was Vice President for Clinical and Coverage Policy of Aetna Health Plans and
currently is part of the IOM Medicare Coverage Advisory Committee. He spent much of his talk noting the factors that payors weigh when considering coverage decisions for various diagnostic and treatment interventions, and how those decisions are influenced by societal pressures.
In the early 1990s, pressure from large companies, who wanted to reduce the costs of the health insurance they were providing for their employees, led to the development of strict evidence-based reimbursement decisions, according to Dr. McGivney. But the adverse publicity and lawsuits this approach triggered led to insurance companies seeking other ways of reducing costs, such as reducing how much they reimburse hospitals, physicians, and other health care providers. However, there is still a need to reduce costs and improve health care. Various options are being considered in this regard, including increasing patient copayments and patient participation in treatment decisions, and evaluating and improving the quality of care based on adherence to guidelines and quality measures, Dr. McGivney said.
Biomarker tests present another health care expense that could be a cost challenge for insurers. But their additional cost might be offset by the opportunity to better direct appropriate treatment and derive greater patient benefit for each health care dollar spent. Dr. McGivney noted, “that is the promise of biomarkers, so payors are looking at them as a way to manage and improve utilization and effectiveness by applying them as inclusive criteria even in preauthorization and medical necessity determinations.”
When making reimbursement decisions, some payors only consider whether a biomarker provides information that helps manage patients, whereas others also consider what patient outcomes the use of the biomarker improves and carefully examine the evidence in that regard, Dr. McGivney said. Cost effectiveness is not used as a criterion for coverage determinations, he said. But cost does affect the intensity at which a payor reviews the evidence for a reimbursement decision. He noted that “until test kits hit $3,000 and are going to be used in, say, 500,000 patients per year, they may not really care. But at some point, there will be a threshold in terms of dollars, where the payors begin to take a hard look at the impact of the test on their bottom lines.”
Dr. McGivney spent part of his talk explaining how many health insurers make their reimbursement decisions. To be reimbursed by an insurer, a technology usually must receive approval from the FDA or some other government regulatory agency. There also has to be sufficient scientific
evidence that the technology improves the net health outcome, and must be as beneficial as any established alternatives. The improvement in health benefit this technology provides must also occur outside of a research setting. He noted that the definition of health outcome in cancer is moving away from complete and partial responses to progression-free survival, as the disease becomes more of a chronic condition.
Other unspoken factors also influence reimbursement decisions, Dr. McGivney added. For example, there can be less certainty about the effectiveness of treatments for life-threatening diseases, especially when children are involved. “At Aetna, my unspoken principle was that we paid for everything for kids under 21,” he said.
Dr. McGivney concluded his talk by discussing how NCCN guidelines affect clinical care and reimbursement decisions. These guidelines are internationally recognized as the standard for clinical policy and coverage decision in oncology, and are used by CMS and other private payors, he said. They are developed by 1,000 clinicians and patient representatives, who serve on 48 panels focused on individual cancers or supportive care issues. The NCCN guidelines are current, specific, and continually updated, according to Dr. McGivney.
Like the recommendations given by the USPSTF, those given by NCCN specify the level of evidence and consensus. Biomarker tests are increasingly being included in NCCN guidelines, Dr. McGivney noted. “Biomarkers clearly address the direction of each treatment pathway for individual patient subpopulations,” he said. Some cancer biomarker tests, such as those for HER2 or the estrogen receptor, play important roles in NCCN guidelines for the treatment of breast cancer (Figure 9). Others, such as urinary urothelial tumor markers, are considered optional additions because the evidence for their effectiveness is not as strong.
Sometimes an NCCN recommendation may contradict what is specified in an FDA label. For example, NCCN recommends that no patient be included or excluded from cetuximab therapy for colorectal cancer on the basis of EGFR test results. In contrast the FDA label for this drug specifies that it be used for the treatment of EGFR-expressing colorectal carcinoma. The decision to link EGFR test results to cetuximab use on the drug label was based on the limited available evidence at the time, and may have also entailed political considerations, Dr. McGivney said.

FIGURE 9 National Comprehensive Cancer Network (NCCN) clinical practice guidelines in oncology, v.2.2006.
SOURCE: McGivney presentation (March 21, 2006).
CMS Coverage of Biomarkers
The final talk in this session was given by Jim Rollins, MD, PhD, of CMS. He explained that CMS bases its coverage of a new diagnostic test on its accuracy and whether the test will lead to a better health outcome. To assess the accuracy, sensitivity and specificity measures may not be adequate, he added, and instead the agency may focus on the test’s analytic validity, clinical validity, and clinical utility. Often CMS will not accept surrogate markers for survival, such as effect on tumor size, he said. Cost is not a consideration when CMS makes its reimbursement decisions, he added.
Certain factors germane to the older population CMS serves (85 percent are 65 and older) and its limited mandate affect CMS coverage of biomarker-based tests, Dr. Rollins noted. The Medicare statute covers diagnosis and treatment but does not specify a benefit category for screening tests or preventive care, so it is unlikely to reimburse for biomarker-based
tests that are used to screen for cancer and/or predict cancer susceptibility. That could potentially change in the future, however, as the Secretary of Health and Human Services’ Advisory Committee for Genetics Health in Society has recommended that Congress add a preventive health benefit to areas that are currently being covered by CMS, according to Dr. Rollins. CMS also is not given authority to conduct research. But CMS can give coverage for a medical intervention conditional on the agency’s concurrent collection of data on the intervention while reimbursing it. A guidance document on Coverage with Evidence Development is pending on this matter.
Cancer biomarkers used to monitor or manage the care of patients with cancer, including those that predict recurrence, are usually covered by Medicare. For example, the agency on a national level covers the use of CA-125 for peritoneal and ovarian cancer patients. Locally in California, CMS covers the use of the OncoTypeDX test, a genetic test that predicts breast cancer recurrence.
Ninety percent of CMS coverage decisions are made locally, but national decisions take precedence over local ones, Dr. Rollins explained. Vendors, physicians within CMS, or those in private practice can request local or national coverage decisions. If there is a great deal of inconsistency between regions over coverage of a particular intervention or test, the agency may evaluate it and generate a national coverage decision.
The CMS and the Agency for Healthcare Research and Quality recently reviewed the literature on biomarkers for cancer as to how they are used (whether for diagnosis or for the management or monitoring of patients) and how effective they are for certain forms of cancer. This review will be posted on the CMS website, and eventually will be expanded with accuracy assessments, according to Dr. Rollins.
CMS may require more evidence to cover a biomarker test than would be required by the FDA for the test’s approval, Dr. Rollins noted. That is because many of the studies submitted to FDA review do not include sufficient numbers of people 65 or older, so their results may not be applicable to the Medicare population. Also, the FDA may approve a particular technology based on the requirement that the vendor will do postmarketing analysis and surveillance. But often vendors do not provide this additional information, so CMS may not cover the use of that technology until there is sufficient evidence to fully evaluate it, Dr. Rollins said.
ECONOMIC IMPACT OF BIOMARKERS
The rapidly increasing cost of medical care is a major concern and has led to a greater interest in the cost effectiveness of medical interventions. The high cost of health care is often attributed, in part, to the adoption of expensive new technologies. These include new targeted therapies for cancer, which, like more traditional therapies, only benefit a fraction of the patients for whom they may be indicated.
However, appropriate patient selection via accurate diagnostic biomarker tests to predict responsiveness could substantially improve patient outcome and thus increase the cost effectiveness of treatment. Similarly, if biomarker-based screening tests could be developed to detect cancer at an earlier, more easily treated stage, these new biomarker technologies could have a substantial impact on the economic burden of cancer by reducing the cost of treatment, as well as the overall burden and consequence of disease. The goal of the last session of the conference was to examine how the cost effectiveness of biomarker tests and the value of the information they provide affects their acceptance by health care payors, such as insurance companies and CMS.
Cost-Effectiveness Analysis
The first speaker at this session was Andrew Stevens, MD, of the United Kingdom’s National Institute for Health and Clinical Excellence (NICE). This organization assesses the value of various medical interventions. Their assessments are used to set the nation’s health service priorities. His talk was followed by that of health economist and physician David Meltzer, MD, PhD, of the University of Chicago. Naomi Aronson, PhD, of the BlueCross BlueShield Technology Evaluation Center (TEC), was the final speaker. BlueCross BlueShield provides health insurance for one out of three privately insured Americans. The company uses its TEC’s scientific reviews of medical interventions when making reimbursement decisions.
Dr. Stevens began the session by noting the need for having cost effectiveness as the “fourth hurdle” in health care, after safety, efficacy, and quality. Such a hurdle is imperative given limited financial resources and the high costs of innovative treatments. For example, treatment with imatinib (Gleevec) can cost as much as $66,000 per patient, he pointed out. Cost-effectiveness analyses are often used by Great Britain and other nations with
socialized medicine to determine how best to ration the health care services it provides, according to Dr. Stevens.
NICE only approves treatments that are both clinically effective and cost effective, although it does give due consideration to notions of equity and innovation. Cost-effectiveness analyses assess the value of a medical treatment by noting its costs relative to its health benefits. That way, one can choose an intervention for which the cost relative to benefit is less than a threshold value. Health benefits are measured with an index called QALY, for quality-adjusted life-years. This index combines measures of quality of life with length of life. In Great Britain, treatments that cost less than $35,000 per QALY are generally approved, whereas those that cost more than $52,000 per QALY are rarely approved. In his talk, Dr. Meltzer noted that the cost-effectiveness threshold for medical interventions in the United States is between $50,000 and $100,000 per QALY.
But cost-effectiveness appraisals have many shortcomings in their methods that can affect their accuracy, all the speakers at this session pointed out. How valid they are depends on the validity of their measures of health outcomes. But that can be adversely affected by basing them on inadequately controlled studies, studies that do not consider the most useful comparators, or studies that are not long enough to truly assess the health outcome of interest. The use of surrogate markers that do not adequately reflect health outcomes can also be a problem. In addition, quality-of-life measures can vary according to subpopulation, Dr Stevens noted, and cost assessments may not be comprehensive enough.
“So there’s an awful lot of subjective analysis in these [cost-effectiveness] appraisals, however scientific the documents [used to make them] seem,” he said. In her presentation, Dr. Aronson concurred and added that although her center has done some cost-effective analyses for educational purposes, “there isn’t any clear science for the cost-effectiveness threshold. I think it is troubling because often we see cost-effectiveness analyses brought to our attention in a lobbying mode by the sponsors of a technology,” who claim the technology should be reimbursed because it is cost effective. But Dr. Meltzer pointed out in his talk that despite their limitations, cost-effectiveness analyses were well accepted and broadly used in the biomedical arena.
Dr. Stevens spent much of his discussion elaborating on the experience NICE has had in evaluating or employing various biomarker diagnostics in their assessments of medical interventions. The value of a biomarker
depends on what type it is and how it is used, he noted. For example, he considers a test for antibodies to hepatitis C an “exposure biomarker” test for liver cancer.16 NICE found this biomarker was not useful in determining who should initiate treatment with interferon and ribavirin (as opposed to watchful waiting) because such costly early treatment only increased the QALY from 21 to 22 years. Such exposure biomarkers are not valuable because of their large lead time and low predictive power, he said.
PSA is a useful biomarker for prostate cancer recurrence or prognosis, but NICE called for more clinical trial evidence when evaluating PSA as a screening test for prostate cancer. In addition to the standard measures of a screening test, such as false-positive and false-negative rates, NICE wanted measures of how the test affected patient health outcomes. Even those patients whose biopsies indicate that they are true positives for the PSA test may not develop an aggressive prostate cancer that requires treatment, Dr. Stevens pointed out. This can be problematic because the treatment for prostate cancer has many severe side effects, he added.
NICE accepted the absence of the Philadelphia chromosome in the bone marrow as a useful surrogate biomarker for improved health outcome for patients with chronic myelogenous leukemia who were treated with Gleevec (Figure 10). This chromosome has the translocation that causes the cancer-triggering mutation that Gleevec targets. The agency recommended offering Gleevec to such patients, despite its high cost, because it was much more effective and had significantly fewer side effects than the standard alternative treatment for this type of leukemia (Figure 11).
The final biomarker example Dr. Stevens presented was the use of O6-methylguanine-DNA methyltransferase (MGMT) methylation status17 in glioma patients to distinguish a treatable subgroup. Treatment with temozolomide in addition to radiotherapy surpasses NICE’s cost-effectiveness threshold. But such treatment only in the subgroup likely to respond, as indicated by MGMT methylation status, gives results that suggest it may be cost effective. MGMT methylation status and other response-predicting biomarkers “have the potential to refine disease and therapy and improve cost effectiveness,” Dr. Stevens said. But he added that their impact on cost effectiveness depends on whether they induce a cost
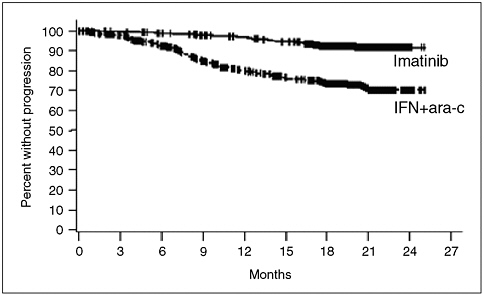
FIGURE 10 Developing new rational therapies—Philadelphia chromosome and imatinib. IFN = interferon alpha; ara-c = cytosine arabinoside.
SOURCE: Stevens presentation (March 21, 2006).
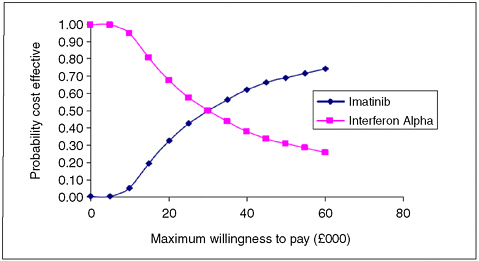
FIGURE 11 Cost-effectiveness acceptability curve for IFN- and imatinib. The willingness of the National Health Service to pay for a treatment depends on the probability of its cost effectiveness. As cost effectiveness increases, high cost is less of a deterrent to providing the treatment.
SOURCE: Stevens presentation (March 21, 2006).
backlash from drug manufacturers. These companies may increase the price of their drugs to make up for the loss in income due to treatment markets narrowed by biomarker tests for patient responsiveness, he noted.
Dr. Meltzer also pointed out how the value of a diagnostic test, including a biomarker test, depends on how it is used. The cost effectiveness of the Pap test substantially decreases, he showed, when it is used annually or every 2 years, as opposed to every 3 years, because the more frequent use only lengthens a patient’s life by an average of a day or two. “These simple analyses can be very revealing,” he said. They show that one cannot simply determine whether a test “is good or bad,” because such determinations depend, in part, on how the test is used.
He expanded on this concept by showing mathematically how self-selection of a medical treatment by patients occurs because they tend to opt out of a therapy when it is not effective. This self-selection can substantially improve the cost effectiveness of the treatment. But most cost-effective analyses only consider the costs and benefits of a diagnostic or treatment for the entire general population, he noted, and do not consider self-selection. “The results of standard cost effectiveness analyses can be very misleading because in modeling, self-selection is very important,” he said.
Requiring copayments for treatments increases self-selection, which in turn also increases the cost effectiveness of the treatment, he added. This suggests a framework for designing copayment strategies to enhance the cost effectiveness of therapies, he said. Nonetheless, he noted that reimbursement systems are not necessarily the right tool to increase value. Decision aids, for example, might be a better tool, he pointed out.
Biomarker diagnostics would be valuable if they encouraged the selective use of treatments. This would substantially increase the cost effectiveness of the treatments, Dr. Meltzer noted. “Our efforts need to go toward getting the right treatment to the right person,” he said. “Having a framework to account for heterogeneity in patient benefits is key to valuing diagnostic tests.” But he added that “biomarkers can also be used incorrectly in the wrong population. If we use biomarkers outside the context in which they have been developed and use them to find a disease, for example, for which we don’t know there is a benefit to treating, then the biomarker is not necessarily going to give us much benefit. So biomarkers are incredibly exciting if they are used right, and dangerous if we don’t control the way in which they are used.”
Another example of this was the use of COX-2 inhibitors, Dr. Meltzer said. Prior to the release of data showing their cardiovascular side effects,
COX-2 inhibitors were shown to be highly cost-effective drugs for patients at high risk of gastrointestinal bleeding. But the drugs were not cost effective in people at low risk of such bleeding. However, most COX-2 inhibitors were used in the United States by people at low risk of bleeding, so the actual cost effectiveness was poor because of how they were used, Dr. Meltzer pointed out. “We need to think about tests and interventions, not just as they would be used under ideal circumstances, but as they are used in practice,” he said.
The Value of Information and Research
In his talk, Dr. Meltzer also showed how mathematical models used to calculate the value of a diagnostic can also be used to calculate the value of information gained by research. These models calculate the difference between the expected outcome with the information garnered from a study and the expected outcome without that information. Such an analysis was used to show the value of research on Alzheimer’s disease treatments and wisdom teeth removal that led NICE to invest in such studies, Dr. Meltzer said.
The value of information analysis was also used to calculate the value of biomedical research supported by the National Institutes of Health (NIH). For this calculation, University of Chicago economists showed that biomedical research increases life expectancy in this country by about 3 months per year. By putting a dollar value on that increase in life expectancy for all U.S. citizens, they calculated that biomedical research was worth about $3 trillion a year. This calculation was used to successfully lobby Congress for an increase in the NIH budget, Dr. Meltzer noted. But he pointed out that the real value of research can be far less than expected, in part because it does not always generate the complete information needed to improve a health outcome. For example, he estimated that the expected value of perfect information about prostate cancer generated from research would be $21 billion, but the expected value of more limited information about certain aspects of the disease would be only $1 billion.
Technology Assessment in the Private Sector
Dr. Aronson of BlueCross BlueShield’s TEC gave the next presentation. TEC has a staff of physicians, epidemiologists, pharmacists, and
medical editors who review and write up scientific assessments of the clinical evidence for various medical interventions. These assessments are used by an independent Medical Advisory Panel, composed mainly of academic researchers, when deciding the insurance company’s medical policy, Dr. Aronson explained.
She stressed that the medical policy decisions on which procedures are clinically beneficial are made separately from coverage and reimbursement policy decisions that determine who should receive such clinical benefits and at what rate of reimbursement. In the development of their medical policy, costs and coverage are not considered, Dr. Aronson pointed out, although they are factored into determinations of premium rates, and into the contracts made with health care providers that specify reimbursement rates.
The TEC assessments, some of which are published online at www.bcbs.com/tec, consider whether a medical procedure or treatment improves health outcomes by increasing the length and/or quality of life, or by increasing the ability to function. But the organization encounters many challenges when conducting their assessments. According to Dr. Aronson, these challenges include inadequate quality of studies done on a topic, selective reporting and publication bias, and incomplete data from studies that are published; an example is that they do not consider important variables needed to determine medical policy decisions.
Often there is a lack of prospective, randomized, double-blinded, and placebo-controlled clinical studies, Dr. Aronson pointed out. Many clinical studies also lack clearly defined patient populations, relevant comparators, and intention-to-treat analyses of all participants by initial group assignment. The studies often do not have the long-term follow-up needed to adequately assess the health outcomes of a medical intervention. The end result is a lack of robust evidence on the effects of an intervention, and how those effects compare to other interventions, Dr. Aronson said.
The way adverse effects are reported in studies is also problematic, she added. These effects often are not systematically and consistently classified across studies, and are not presented in a way that can be easily synthesized. In addition, usually the frequency of adverse effects, rather than their severity, is reported. “This is tremendously frustrating to us,” she said. “We feel like when we do systematic reviews, we can only do half our job. Most of what is available to us is efficacy outcomes, but often, we are really lacking what we need to know about adverse effects.”
Another challenge is a lack of direct evidence for the value of diagnostics. Often performance characteristics of the test are used to fill in a model
of how the technology can detect a condition or change its management such that there is an improved health outcome. But such an approach can be overly simplistic, Dr. Aronson said, and “there are times when the model is so complicated that you will need direct evidence for a diagnostic; that is to have it tested in a randomized controlled trial, much as if it were an intervention.”
For example, there is a test for a variation in the gene that codes for the drug-metabolizing enzyme cytochrome P450 (CYP450). This variant hampers the enzyme’s ability to metabolize certain drugs, including warfarin. Therefore, a person who has the gene might benefit by having lower doses of those drugs. But other factors also can slow down drug metabolism. These factors include other enzymes, coexisting disease, age, diet, and interactions with other drugs. Given this complex scenario, it is not clear how useful a test for just one influence on drug metabolism will be for patients who take warfarin, Dr. Aronson said (Figure 12). “I don’t think you can jump from the observation about CYP 450 to the conclusion that this is a good control for personalizing warfarin dose. We think something like this is so complicated that it needs to be tested, and I expect many biomarkers for prediction of response may fall into that category,” she said.
A final challenge to making assessments of medical interventions or tests that Dr. Aronson discussed was the “file drawer problem” of researchers not publishing their nonsignificant results or results that do not favor the
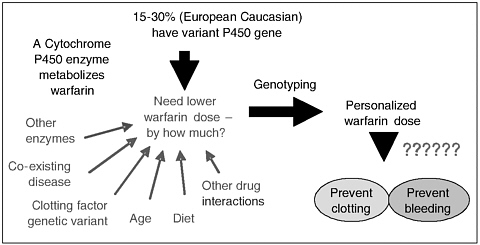
FIGURE 12 Direct evidence for diagnostics.
SOURCE: Adapted from Aronson presentation (March 21, 2006).
drug company that sponsors the research. “We certainly concur [with], and will integrate into our own process, the principles of the International Committee of Medical Journal Editors that call for prospective registration of clinical trials in a public database,” she said.
During the next part of her talk, Dr. Aronson addressed some concerns she has about biomarker diagnostics, including their ability to slip through regulatory cracks and not be sufficiently evaluated by adequately designed studies. She noted the regulatory gap for biomarker diagnostics, especially those that are considered home brews. “I see CLIA focusing on lab quality, the FDA focusing on analytic performance, manufacturing quality, clinical validity. Where does clinical utility come in?” she said. She added that she and her colleagues at TEC readily embrace the efforts of EGAPP and believe their model for analyzing genetic tests will serve TEC well.
She criticized the design of many studies to formally assess tumor markers as being inconsistent and inadequate, and questioned the frequent use of biomarkers as surrogates for outcomes in clinical trials. “A correlate does not a surrogate make,” she said, because the biomarker may not be in the causal pathway of the disease, there may be multiple causal pathways, or there may be unintended adverse effects of an intervention. It is difficult to know where to draw the line on when a biomarker can adequately serve as a surrogate. She asked, “what shall we trust to demonstrate a health outcome? Under what circumstances? And if we draw the bar there, what are the consequences? Once accepted into clinical practice, it is often difficult to obtain higher level evidence.”
Dr. Aronson concluded her talk by discussing cost issues linked to the use of biomarkers. “I think we are, on an ongoing basis, encountering something very troubling in terms of the new technologies—that they bring benefits, but they are small benefits at high costs,” she said. Even so-called cost-effective interventions may not be affordable, she added. “I am not sure we can afford everything that is a good buy, or at least at the good buy rate of $50,000 per QALY. Whatever the value of the intervention is, it, alone, cannot ultimately trump affordability,” she said.
The consequences of not capping the costs of medical interventions and procedures are high premium rates, which foster a decline in employers offering health benefits to employees, Dr. Aronson said. The end result is that more individuals will be uninsured. The number of uninsured in the United States now is around 45 million—a number that equals the number of Medicare beneficiaries, Dr. Aronson observed.
During the questioning that followed her talk, Dr. Aronson said that although BlueCross BlueShield does fund some health services research, “a research agenda is not really appropriate to our mission nor affordable.” The company has a financial responsibility to keep administrative costs, including research costs, to a minimum to maintain premium affordability, she said.
CLINICAL DEVELOPMENT STRATEGIES FOR BIOMARKER UTILIZATION DISCUSSION
On the final day of the conference, representatives from each of the seven small group discussions that met during the previous 2 days gave summaries of their groups’ discussions. Discussion moderator Stephen Friend, MD, PhD, of Merck and Co., Inc., started the summary of his group’s discussion of clinical development strategies for biomarker utilization by listing the main challenges that his group identified to the clinical development of biomarkers. These challenges were:
-
Gaining more access to patient materials and data;
-
Coordinating the development of diagnostics and treatments;
-
Providing incentives for diagnostic companies;
-
Developing “smarter” clinical trial designs; and
-
Better integrating basic science and clinical research efforts.
Patient biopsy tissue and other patient materials collected during clinical trials are invaluable for researchers trying to discover or develop biomarkers. But a number of issues contribute to making a lack of patient materials a limiting factor in biomarker clinical development, Dr. Friend said. There is a general lack of sample collection, which is especially true for patients having relapses of their cancer. In addition, the samples are often of poor quality, are misclassified by pathologists such that normal cells are mistaken for cancerous cells, or are preserved in a way that precludes their use in various biomarker studies. Furthermore, annotating and storing samples can be costly, so few investigators or institutions are willing to undertake these endeavors and then provide the samples and data to others.
Even if good-quality patient samples can be accessed, investigators may not be able to use many of them because of inadequate data about the patients from whom they were collected. Variability in the way the data are entered and categorized in a database can also make it difficult, if not
impossible, for investigators to retrieve the information they need to include the patients in their studies. Another major barrier that can impede research on stored patient samples is a lack of informed consent forms that are broad enough to encompass new uses of the tissue samples beyond the use for which they were initially collected. In addition, rules in various academic institutions may restrict the sharing of patient materials and corresponding data with other researchers because of competitive and financial pressures. These patient materials are often seen as having some inherent intellectual property value.
Group members had a number of ideas for how to overcome patient sample-related barriers to biomarker development. Suggestions included reexamining the existing informed consent process and making it more flexible as to the range of studies that can be conducted on the tissues collected, providing more funds for the annotation of collected patient materials, and reexamining current academic center rules on collected data, particularly as related to intellectual property. Some members of the discussion group suggested imposing penalties on investigators or academic institutions unwilling to share patient samples and data, whereas other members thought that offering rewards for such behavior was a better alternative.
Some of this group’s discussion focused on how to provide incentives for diagnostic companies to translate a biomarker discovery into a clinically marketed test. Dr. Friend noted that diagnostic companies have small profit margins that limit their willingness to undertake major financial risks or costly development endeavors. But a diagnostic company might be more willing to pursue a biomarker test if it could be coupled with a therapeutic that a drug company is developing such that the risks, revenues, and products are shared between the two companies.
Another way to make diagnostic biomarker development more appealing is to give efforts in this regard “type two patents.” Australian political philosopher Thomas Pogge coined this term for patents that reward research and development work that results in useful drugs or diagnostics that normally would have a low margin of return. The financial reward of the patent is based on how much of a health benefit the drug or test provides, and makes up for the more limited revenue gained from the sales of the product. Federal governments create funds to support such a patent reward system, which helps make the early development of diagnostic biomarkers less risky. In this manner, the financial worth of a diagnostic is based more on its value to society rather than on its sales revenue.
One member of the group, Richard Frank, MD, PhD, of GE Healthcare, suggested another way to provide an incentive for diagnostics companies to do more research and development on biomarkers. He proposed that these companies be allowed access to government-funded tissue and databanks to conduct biomarker research with the option of having exclusive rights to any biomarker tests that evolve from such research. But other members of the group questioned the need to grant such exclusive rights. Current NIH policy does not grant exclusivity in the licensing of tests that emerge from research on its tissue samples or data.
Another way to make diagnostic development less risky, the group suggested, would be to develop biomarkers for key steps along the biochemical pathways that cause various types of cancer. Researchers suspect there are a limited number of these pathways, which play a role in a wide range of cancers. “If you got those pathway biomarkers, then they are not dependent on a particular individual drug that is going through the pipeline, but instead could apply to any company drug,” Dr. Friend said.
He noted that there needs to be a certain level of rigor to a response-predicting diagnostic used in a clinical trial. But the development of such a rigorous biomarker diagnostic often lags behind that of a related drug, so the diagnostic is not ready to enter Phase III testing at the same time as the drug. To solve that timing issue, group participants suggested the development of common shared databases that can be used to develop biomarkers appropriate for predicting response to drugs. They also suggested that industry do more precompetitive investing in research on biomarkers that predict drug response, and make greater use of pathway biomarkers. The FDA might also consider linking its approvals of therapeutics to related response-predicting diagnostics such that one is contingent on the other.
Members of the group also recognized the dynamic nature of the field of biomarkers that predict drug response. Some of these biomarkers are developed after the drugs they predict response to are already on the market, while others are found to be predictive for more drugs than the ones whose labels specify their use. This can restrict the use of biomarker tests because off-label use is often not reimbursed. Consequently, the group suggested considering the need to have more dynamic ways of modifying drug labels based on emerging data.
Group members also suggested considering the consequences of increasingly tight restrictions on off-label use of diagnostics and therapeutics. Not only do such restrictions limit the use of already developed biomarkers, but they limit the amount of resources that drug companies
can devote to developing biomarkers by requiring them to sink large sums of money into conducting clinical trials for added indications, Dr. Friend said. However, some group members noted that a benefit to label restrictions could be that more patients would enter clinical trials in order to be reimbursed for a drug used for an off-label purpose.
Another major topic of this group’s discussion was the need for smarter clinical trial designs that invest in earlier use of biomarkers, especially to define responsive subpopulations prior to Phase III trials. One person in the group noted that the bulk of a company’s drug development costs are for developing unsuccessful drugs—those that do not “pass” clinical trials and enter the market. The use of biomarkers to enrich the number of responders in a clinical trial population should therefore lower development costs overall, he noted, if it makes it more likely that drugs would fare well in clinical trials.
Group participants suggested there should be better integration of basic science and clinical research efforts. The personalized medicine approach that the latest findings in molecular biology suggest does not fit into the traditional models for running clinical trials and developing therapeutics or diagnostics. A “third culture” is needed to bridge the gap between the basic and clinical world, as well as to connect academic and industrial realms, according to group member Dr. Phelps.
Dr. Friend noted that biomarker validation is a “no-person’s land” with respect to funding and effort. Academics are not likely to take on this endeavor because they cannot build their careers on such efforts. Pharmaceutical companies also may not be willing to undertake certain biomarker validations if the tests limit their current market for drugs.
Members of the group suggested that NCI support a program that brings together basic, clinical, and perhaps even industry researchers working on a common group of biomarkers—those that define particular oncogenic pathways, for example. This program could be modeled after the Specialized Programs of Research Excellence (SPORE), which supports a mix of basic and clinical researchers working on the same cancer type.
However, David Carbone, MD, of Vanderbilt University, cautioned against emphasizing pathway-specific as opposed to disease-specific research because “the requirements for biomarkers—sensitivity, specificity, precision and accuracy—are quite different in different diseases.” He gave the example of epidermal growth factor receptor- (EGFR-) targeted drugs for lung and breast cancer and noted that fundamentally different biomarkers are going to be needed for these two diseases. Dr. Friend agreed with the
importance of pathways being seen within the context of what tissues they operate in, but added “it does not take away from the need of taking raw data, aggregating it, and looking at pathways.”
Dr. Friend concluded his summary with the group’s idea that demonstration studies be funded for oncology drugs already on the market. These studies could demonstrate the feasibility and utility of developing robust response biomarkers.
STRATEGIES FOR IMPLEMENTING STANDARDIZED BIOREPOSITORIES DISCUSSION
This discussion group session was summarized briefly by Harold Moses, MD, of Vanderbilt University. His summary was supplemented with a more detailed written synopsis by Maria Hewitt, PhD, of the IOM.
Dr. Moses noted that the discussion session began with a report by Carolyn Compton, MD, Director of NCI’s Office of Biorepositories and Biospecimen Research. She pointed out that NCI’s initiatives to further personalized medicine all depend on human biospecimens. Through an extensive internal and external review process, NCI has identified major biorepository-related barriers to furthering these initiatives, as well as potential solutions. This effort led to the development of NCI Guidelines for Biorepositories,18 the second generation of which is currently being developed in collaboration with the College of American Pathologists and other relevant extramural groups.
The first-generation guidelines include recommendations for the following:
-
Common best practices for research biorepositories
-
Quality assurance and quality control programs
-
Informatics systems
-
Ways to address ethical, legal, and policy issues (e.g., informed consent, privacy, data security protections, Institutional Review Board oversight, ownership of and access to biospecimens and data)
-
Standardized reporting mechanisms
-
Administration and management structure
The NCI’s second-generation guidelines will propose evidence-based standard operating procedures (SOPs). There is widespread recognition that one size does not fit all, and that SOPs may vary depending on the analytic goal.
NCI recently established an Office of Biorepositories and Biospecimen Research and launched a pilot test of the proposed National Biospecimen Network (NBN). This pilot study will be conducted in 11 prostate SPOREs to evaluate the use of best practices. Dr. Compton reported that NBN’s key requirements for a new biorepository system include:
-
Representation of all cancer types, and all populations
-
Access through a timely, centralized peer-review process
-
Ethical and privacy compliance through a chain of trust
-
Resources provided without intellectual property restrictions
-
Pathology and clinical annotation (including longitudinal)
-
State-of-the-art information technology system to streamline the research process
-
Communication and outreach efforts
-
Best practice- and data driven-based SOPs to enable reproducible and comparable (additive) results
Brent Zanke, MD, PhD, of the Ontario Cancer Research Network then discussed the Ontario Tumour Bank, which collects, stores, and distributes tissues at six clinical centers that follow defined SOPs. The Canadian biorepository has centralized data collection, which protects patient privacy, and accrues 3,000 samples a year. A web-based system19 allows researchers to browse the central database for specimens that meet their study requirements. The Ontario Tumour Bank offers the following products:
-
Fresh frozen tumor
-
Frozen plasma
-
Frozen buffy coat
-
Paraffin-embedded tumor
-
Normal tissue adjacent to tumor samples
Future plans are to offer paraffin sections, stained sections, and tissue microarrays. Researchers can retrieve extensive data, including specimen
quality, from the web-accessible database on available specimens. Requests for specimen access are made through a controlled application process. A tissue ethics committee oversees the program. Samples are provided at a discount to participating centers and academic researchers in Ontario, and at reimbursement costs to others.
The Ontario biorepository was established with a $10 million (Canadian) investment from the Ontario Provincial Government Ministry of Research and Innovation. Similar biorepositories operate in Great Britain and other countries in Europe, according to Dr. Moses. The Ontario system is not directly exportable to the United States because the United States lacks a national health care system and centralized control of hospitals and provider networks. “Our country is way behind,” Dr. Moses said. “The reason the Ontario Tumour Bank can do it for $10 million is that their surgeons, pathologists, etc., are on government salary.”
These presentations led to a general discussion on how to fund biorepositories in the United States. Group participants noted that NCI alone cannot bear the costs of supporting national biorepositories, and suggested public-private consortia as a means for supporting biorepositories. Industry has much to gain and should find that sharing costs serves its interest along with academic centers and philanthropists, participants pointed out. The biorepositories would require a large initial investment, Dr. Moses noted, but could be self-sustaining through fees charged for providing high-quality material. Group members also suggested involving CMS and other health care payors in a discussion of supporting biorepository efforts, as they could reimburse pathologists’ fees for processing specimens.
In a discussion following Dr. Moses’s presentation, David Parkinson, MD, of Amgen (now at Biogen Idec) noted that the biorepository set up by the Multiple Myeloma Research Consortium could serve as a model. This organization is funded by philanthropy and aims to accelerate the development of novel, cutting-edge treatments for multiple myeloma by catalyzing, promoting, and facilitating collaborative research between industry and academia. The Consortium shares its well-annotated and extensive tissue collection with academic and industry researchers. These investigators normally would not focus their efforts on such a rare type of cancer, but do so because of the ease with which these materials are made available to them, Dr. Parkinson said.
Discussant Margaret Spitz, MD, MPH, of the MD Anderson Cancer Center at the University of Texas explored the unique needs of population-based registries and epidemiologic studies. Specimens from control subjects
are very important, as are prediagnostic specimens. In addition to clinical information about samples, epidemiologists need information on environmental exposure, family history, and risk factors. Deidentification of specimens can be problematic because long-term follow-up is often necessary. Some large cohort studies have lost funding for longer term follow-up and face the problem of what to do with their patient samples. A plan is needed prospectively to deal with this issue. NIH might support biorepositories as part of large cohort studies, the group suggested. “We need to look at mechanisms for funding the preservation of these biorepositories as a matter of course because they are just too valuable to let deteriorate because of current funding issues,” Dr. Moses said.
In a discussion that followed Dr. Moses’s presentation, Dr. Carbone described the long-term expenses involved in supporting a biorepository. “It is not readily appreciated how complex managing a tissue collection program really is if you want to do good science,” he said. “And it does not end with plunking the sample in liquid nitrogen. The most valuable thing you have in these tissue banks is detailed clinical information that evolves over time. We have to go back every 3 months and go over every sample in our tumor bank and update the status on patients, including what chemotherapies they got. It is very difficult and expensive for very focused questions in a typical SPORE grant with a very limited tissue collection. In one disease site we are talking about it costing between $250,000 and $500,000 a year. The cost of doing anything on a grand scale would be enormous.” Dr. Moses noted that NCI currently spends about $50 million a year supporting biorepositories.
A major topic of his discussion group was patient-informed consent. There was an appeal for improved and standardized consent forms that could be used nationally. These forms should resolve current disparities in government rules regarding informed consent, the group suggested. For example, certain government agencies require informed consent before using tissues from patients who have died, while others do not have such a requirement.
The privacy provisions of the Health Insurance Portability and Accountability Act (HIPAA) also raise some obstacles for biorepositories, especially in the area of needing to acquire new patient consent to gain access to tissues for research other than the study for which the samples were originally collected. How hospitals interpret HIPAA rules also varies widely.
The National Cancer Policy Forum was scheduled to examine the effect of HIPAA on biomedical research at its June meeting,20 Dr. Moses said.
Who owns patient specimens? This was another major issue tackled by the discussion group, which reported that an answer to this question is currently being decided in the courts. One case is testing whether the investigator or hospital owns patient samples. Some research consortia have clearly specified, in advance, issues related to access and ownership of samples.
The discussion group also touched on the need for common data elements being reported for specimens in biorepositories. In addition, some group members suggested biorepositories invest in electronic medical record systems to facilitate gathering of clinical and other data.
In a discussion following Dr. Moses’s presentation, Drs. Carbone, Ransohoff, and Friend questioned the logic of investing in large centralized biorepositories because most studies require specific specimens from specific cohorts of patients, and those specimens must be handled in certain ways. “You cannot make some sample adequate for a particular [research] question just by annotation. We have to be careful not to overinvest in large data repositories until we give some thought to exactly what questions we would be able to answer if we really collected all of the data,” said Dr. Ransohoff.
Dr. Carbone added, “A much more valuable way to spend resources would be to dramatically increase funding for biospecimens associated with particular clinical investigations or interventions such as cooperative group trials. If you could dump money in support of biospecimen collections into Phase III randomized trials, instead of getting 20 samples out of 1,000 you could get 600, which would give you a specimen collection that is much more valuable than catching things that are thrown in the trash in surgical pathology.”
Drs. Friend and Ransohoff suggested starting a new initiative to support centralized biorepositories by funding a collection of just one or two tumor types and focusing on specific hypotheses to “prove that it works and can be a shining example for others,” Dr. Friend said. Dr. Moses agreed and reiterated that NBN is funding such a pilot project in its prostate SPOREs.
STRATEGIES FOR DETERMINING ANALYTIC VALIDITY AND CLINICAL UTILITY OF BIOMARKERS DISCUSSION
Moderator Dr. Howard Schulman reported on his group’s discussion about strategies for determining analytic validity and clinical utility of biomarkers. He noted that the group’s comments reflected a widespread belief that discussions on biomarkers should differentiate between qualification (clinical validity and utility) and validation (assay validity). Similarly, distinctions should be made for biomarkers used only by pharmaceutical companies during the initial stages of drug development versus those used in clinical trials that affect clinical decisions. For example, biomarkers used only by pharmaceutical companies to determine if a drug they are developing is acting on its target would not have to undergo scrutiny by the FDA, but more oversight is needed for a biomarker used to stratify patients into responders and nonresponders.
Group members stated that the type of technology used for an assay will influence acceptance criteria for its analytical validation. They suggested that one should consider the context and risk/benefit equation when determining validation and qualification acceptance criteria. In other words, the test consequences determine the standard, and assays that affect clinical decisions should meet the highest standards. The discovery phase should be guided by good science without being encumbered by regulations, Dr. Schulman said. However, quality control samples, platform standards, or proficiency testing of laboratories may be needed when a biomarker test is used to predict patient response or determine dosing.
Discussion participants from all of the various interest groups expressed the view that increased access to and standardization of databases is necessary to advance biomarker science. Often investigators cannot access the clinical information they need to conduct biomarker studies. Even if it is collected, it may not be entered into a database in a way that makes it easy to retrieve and use.
The current FDA regulatory approach to biomarkers, which involves having heightened regulation of biomarkers used for clinical decisions and less regulation for those used early on in development, was generally agreed on by the group, Dr. Schulman said. But there was a lack of agreement on acceptance criteria for diagnostics paired with new therapeutics when the diagnostic is a clinical laboratory test that is subject only to CLIA oversight. Members of the group thought it was not necessary to have rigorous regulatory requirements for response-predicting biomarkers used in the initial
stages of drug development. It was pointed out that a biomarker used to stratify patients in a clinical trial is not equivalent to a diagnostic test used for the same purpose on the market, and therefore could be subject to a lower standard. Many group members believed that there is a need for a realistic algorithm for combining the development of therapeutics and diagnostics.
In addition, group members thought incentives for developing diagnostics are lacking because the diagnostics business is not a high-margin business. There was tremendous enthusiasm from different interest groups for a variety of consortia that can further precompetitive work on biomarkers. In the discussion after his presentation, Dr. Schulman noted that a consortium for biomarker validation that includes the FDA, CMS, and various pharmaceutical companies is already under way. He added that “there is a general feeling that there is an opportunity to do something right on a bigger scale where oftentimes the intellectual property issues are not problematic.”
Group participants noted that investigators who discover biomarkers often do not understand what is required for analytical or clinical validation. “Most people in discovery sites are not familiar with the whole process that one has to go through if you are a diagnostic company,” said Dr. Schulman. “It is quite rigorous and for some of these diagnostics you actually have to have data on 6,000 patients whereas oftentimes, people do a study on 20 patients and think they have discovered a diagnostic and are ready to put it out there.”
There is a critical need for standards for the new technologies used in biomarker-based assays, Dr. Schulman reported. Standards would be helpful for microarrays and other genomics technologies, proteomics, and metabolomics. These standards could help solve current problems with interpreting results, such as how to deal with uncertainties in protein identification when using mass spectrometer data and how to compare results garnered from different technologies. Another issue is how to establish consistent standards for the same technology, such as for mass spectrometry, whose resolution and other determinants depend on the exact instrumentation used in a laboratory. Even established genetic probes can generate uncertain results when applied to microarrays, Dr. Schulman pointed out.
Members of his discussion group thought there was a role for the National Institute of Standards and Technology and the FDA in resolving some of these standardization issues. Consortia could also be helpful in this regard, he added. In the discussion that followed his presentation, Helen
Francis Lang, PhD, from Affymetrix, Inc. pointed out that “the FDA and a large number of stakeholders are involved with platform companies in establishing quite strict standards in terms of the use of microarrays, controls, and interpretation of data.”
In regard to the discovery and development of biomarkers, group members noted that biomarkers are best vetted and promoted to greater degrees of qualification as more studies are conducted with them. “In a way, it is a communal process,” Dr. Schulman said. “This is one of the reasons for sharing both samples and information because the more people work on the same set of biomarkers, the more we learn about their flaws and good points and come up with a better test.” To further that communal process, the group suggested that investigators publish all raw data from their biomarker studies.
Group participants also noted that better access to clinical specimens would be a boost to diagnostic development, but a number of obstacles must be overcome. As other groups have pointed out, HIPAA can make it difficult to access clinical material and linked clinical information. To overcome those difficulties, members of Dr. Schulman’s discussion group suggested facilitating studies by multiple groups that use the same high-quality tissue repository, akin to what is already done successfully in Germany and Holland. These repositories should have extensive annotation of their samples so that the clinical characteristics of interest to investigators are documented. For these samples, much more is needed beyond healthy and diseased distinctions, Dr. Schulman noted.
For biorepositories to be useful, they should be tailored to the types of research investigations that will be done when using them, he added, and contain the kinds of samples and patient information needed to test specific research hypotheses. “For the millions of samples that are stored today in various banks in the United States, most are never touched,” Dr. Schulman noted. “That is one reason to link the hypothesis that needs to be tested to the samples collected. If you just collect a priori you may not have the right samples to address your questions.” The quality of tissue collections is also critical, so members of the discussion group suggested involving clinical pathologists in the collection process from the start, as well as training them so that sample preparations better meet the needs of researchers.
STRATEGIES TO DEVELOP BIOMARKERS FOR EARLY DETECTION DISCUSSION
Scott Ramsey, MD, PhD, of Fred Hutchison Cancer Research Center gave the synopsis for his group’s discussion on strategies to develop biomarkers for early detection. He noted that most members of his group were “end users” of biomarkers, including clinicians such as oncologists and general internists. They focused their discussion on the use of biomarkers to screen for cancer.
Participants in this group noted that often biomarkers are developed without extensive thought about how exactly they will fit into the clinical pathway, such as how they will affect clinical decisions on treatment or other interventions. “When developing biomarkers for early detection, we must keep end use and value in mind,” Dr. Ramsey said. “I have been asked to do clinical studies on biomarkers and when I ask the developer to draw out a decision tree or a pathway and tell me where the biomarker fits in that pathway, I often get four or five different approaches. It is pretty clear that they are looking for whatever sticks in terms of how the biomarker might be used in the clinical pathway.”
Biomarker tests are not useful, for example, if they detect a cancer somewhere in the body, but lack enough specificity to lead to a treatment plan. “There is nothing I can imagine worse than having a blood test that shows cancer and not having the foggiest idea what to do with that information as far as the patient is concerned,” said discussant Larry Norton, MD, of Memorial Sloan-Kettering Cancer Center.
Biomarkers may also detect heightened risk for a specific cancer, when there are no known prophylactic measures to reduce that risk. Biomarker tests that detect whether people are at higher risk of developing a specific cancer, such as those for breast cancer-related mutations in the BRCA genes, are problematic in that regard. As there are no known measures to substantially reduce the risk of breast cancer in women who test positive for these mutations short of prophylactic mastectomies, the clinical usefulness of the test is questionable, some group members asserted.
Another related area that many individuals in the group thought was often neglected by biomarker test developers was the potential for biomarker tests to harm patients. False results, or positive results for a disease that would never have manifested clinically, can cause patients to be overtreated. Group members thought this concern should be addressed early in the development process before biomarker tests are disseminated,
marketed, and adopted. Studies should clearly define the risk/benefit ratio of a biomarker test prior to its use in the clinic, Dr. Ramsey said.
Group participants noted that patient perspectives and preferences for screening tests are very influential. Patients may have expectations about a specific test that are overly optimistic, both in terms of what the test can reveal and in terms of whether it is likely to be negative in their case. Many patients do not have a good sense of the risks and benefits of moving down a pathway of testing using biomarkers. This is especially true for tests that predict heightened risk of developing a cancer. Patients may pressure their physicians to give them these tests, even when there is inadequate evidence regarding the risks and benefits of testing. Alternatively, patients may acquiesce to being tested, even though they have no intention of submitting to treatment if a test is positive. Group members noted the importance of recognizing patient preferences when developing biomarker tests.
Biomarker developers should also consider whether there is a clinical need for a biomarker-based test. Does it help doctors and their patients, or does it just complicate their care? For example, a biomarker-based test for colorectal cancer screening may be superfluous because several good tests that accurately detect this type of cancer are already in use, some discussion participants noted. Often development of a biomarker test is driven by advances in basic science and what new techniques are available, but not necessarily by the clinical need for the test, the group noted. Furthermore, a new understanding of cancer etiology due to basic science findings does not always translate into tests and interventions that are helpful to patients.
Group members recognized that the regulatory oversight for biomarker tests is evolving as a moving target. Some participants expressed concern about off-label use of biomarkers for indications other than those for which they were originally intended. This off-label use can be problematic, as it has been for the PSA test. This test was originally developed for prognostic and surveillance purposes for men diagnosed with prostate cancer. But it was rapidly put into practice as a screening test for prostate cancer, despite a lack of evidence on its risks and benefits in that regard. Tests used for early detection of cancer should be assessed for that purpose, the group stated, with some measure of benefit and risk.
In addition to the FDA’s regulatory role, group members noted that coverage decisions play a role in determing the use of biomarker tests in clinical settings. Coverage decisions may keep biomarkers that have not been assessed adequately off the market, or may prevent them from being used inappropriately, such as for screening, discussants noted. But they also
appreciated the fact that insurers are extremely responsive to pressures from advocacy groups and politicians, and will reimburse for a test when pressured to do so. Much of the screening technology that Medicare reimburses was mandated by legislation, noted Dr. Ramsey.
Dr. Ramsey finished his presentation by stating his group’s awareness that translating a promising discovery into a validated biomarker for early detection of cancer is enormously challenging. As members of other discussion groups noted, this translational process requires access to high-quality, highly annotated patient samples collected in a nonbiased way. They suggested biomarker developers use existing, prospectively collected samples, such as those collected as part of the Women’s Health Initiative study.
Smaller biorepositories that stem from private collections can also be useful in this regard, Dr. Norton pointed out, but locating those samples can be difficult. To help researchers find the clinical samples they need, he created a “virtual” repository, which catalogs what is available and uses a computer program to match investigators to appropriate specimen collections. This virtual repository was used to delineate the various types of invasive breast cancers based on tumor cells’ estrogen receptor, HER2, progesterone receptor, or EGFR status.
Translating biomarkers into clinically useful tests also requires prospective, randomized clinical trials to assess their risks and benefits. These trials are extremely large, lengthy, and costly. In some cases they may not be feasible because of difficulties in accruing enough patients, especially if current practice patterns make it unlikely that people will knowingly accept randomization into a control group. This is especially true if a biomarker test is put into clinical practice before its clinical usefulness is fully assessed. Such premature clinical adoption is often the case for off-label use of such tests, or for tests that enter the market as a home-brew clinical laboratory test. But premature and inappropriate adoption of biomarker tests could be even costlier for society, the group noted. According to Dr. Ramsey, “if a biomarker diffuses into clinical use and we really do not know what it is doing to folks, the cost of that could be enormous and exceed the cost of doing a clinical trial itself.”
Members of the discussion group stressed that clinical trials of biomarker tests should be designed so that the diagnostic test is tied to a therapeutic intervention. “There has to be a plan for what you do if the test is positive and that has to be built into the trial early on,” said Dr. Norton. But other group members asserted that conducting such trials early in biomarker development is not feasible because there are not enough resources
(both funds and patients) to conduct such large expensive trials for all the biomarkers currently in development.
Discussant Walter Koch, PhD, of Roche Molecular Systems suggested that such trials be reserved for screening biomarker tests for which there is a diagnostic test that can help determine whether the cancer it detects would need to be treated; for example it may be indolent. He also suggested additional criteria for selecting the most promising candidate tests on which to conduct clinical trials; criteria would include a clinical need for the tests and the availability of effective treatments for the cancers they detect. Another criterion suggested by group members was that the potential screening biomarker should first show promising results when tested using well-annotated archived patient specimens collected for other prospective clinical studies, such as the Women’s Health Initiative or the NCI’s Prostate, Lung, Colon, and Ovarian screening trial. Group members said a conceptual framework was needed for deciding which biomarker tests should proceed to clinical trials.
The group also discussed which type of biomarker tests—those based on multiple markers versus those based on a single marker—are more advantageous to pursue. Discussant Hongyue Dai, PhD, of Rosetta Inpharmatics said he thought multiple markers are more powerful in terms of measurement certainty, sensitivity, or specificity. “With multiple markers, you can do a pattern match, and not simply rely on a high or low judgment based on threshold,” he said.
Others were concerned about the use of a pattern biomarker test that stemmed from overfitting of data, but Dr. Dai said overfitting could be avoided by predefining the pattern prior to testing its predictive power. He added that a pattern biomarker test that considers multiple steps on the same cancer-causing pathway is more likely to be accurate than one that relies on just one step of the pathway. But it can be difficult and arbitrary to draw the threshold on pattern biomarker tests as well, according to BrianTaylor, PhD, of Expression Pathology. “Let’s say you are looking at 20 different biomarkers [in your pattern test]. If 12 of those fit, does that mean that your test has worked, or does it have to be 8 or 17? How do you draw those lines?”
The group recognized that NIH traditionally has not funded translational work for biomarker tests, so it is difficult to find funding to run clinical trials on them. There is also a lack of incentives for academics to undertake such trials because the academic career and reward structure does not encourage translational work. In addition, few incentives exist for
industry to undertake such costly long-term clinical trials, which will not necessarily reward companies with higher revenues, Dr. Ramsey said.
MECHANISMS FOR DEVELOPING AN EVIDENCE BASE DISCUSSION
Dr. Sawyers was the moderator who summarized his group’s discussion on mechanisms for developing an evidence base. He pointed out that many people who participated in the discussion also participated in the earlier discussion on clinical development strategies for biomarker utilization. Consequently, some of his group’s conclusions echo those of the earlier group, which was summarized by Dr. Friend.
Members of Dr. Sawyers’s group suggested creating public-private consortia to develop different types of biomarkers. Participants in each consortium would be those parties most likely to use and benefit from the type of biomarker the consortium develops. For example, surrogate endpoint markers are beneficial to all parties conducting clinical trials for the purpose of achieving FDA approval for a drug to enter the market. An example of such a consortium was one created to develop CD4 count and HIV viral load as surrogate endpoints for clinical trials used to gain FDA approval of various antiretroviral drugs for HIV infection. Participants in this consortium were the pharmaceutical companies that were developing drugs to treat patients with HIV.
Biomarkers that predict adverse reactions to drugs actually protect the public so their development should be a publicly funded goal, proposed one discussant. But Dr. Sawyers said that pharmaceutical firms should help pay for their development as they help these companies decide whether to subject drugs to further testing in clinical trials. Consortia to develop pathway biomarkers were also suggested by group members, who broke these biomarkers down into two subcategories: signaling biomarkers and cellular response markers.
Signaling markers detect aberrations in a specific biochemical signaling pathway in tumor cells. For example, these markers include disease-causing changes in the ras or the EGFR genes or proteins. Signaling markers are best suited for determining a prognosis and for choosing an appropriate treatment plan. Cellular response markers measure more general processes such as tumor cell proliferation, apoptosis, or angiogenesis. Ideally, these markers should be measured noninvasively, via serum tests or imaging, to
reveal whether tumors are progressing and how a treatment is affecting the targeted tumor.
The development of surrogate endpoint markers, adverse reaction biomarkers, and pathway biomarkers would be precompetitive activities that should not require exclusivity. Therefore, all interested parties would benefit by pooling their activities and sharing the development costs, Dr. Sawyer noted. This is in contrast to diagnostics that will be used only when paired to the use of specific drugs, such as the HercepTest, which is used to predict response to Herceptin. The group suggested that both the diagnostic company and the drug company for a paired diagnostic and treatment share the development costs for these types of biomarkers.
After group participants suggested that public-private partnerships could be established to facilitate development of candidate biomarkers, they explored further which groups should be involved in these partnerships. As previous discussions have noted, academia does some discovery work on biomarkers. But academia typically is not involved in the development of robust diagnostic assays because of a lack of expertise in the industrialization aspects and because of a lack of academic rewards and funding sources for this type of research. Start-up diagnostic companies also are not likely to develop biomarker assays because of the low profit margins of diagnostic tests, which make them unattractive to investors. “There was some discussion that if we wait and hope that this happens through free enterprise, we could be waiting awhile,” Dr. Sawyers noted. Consequently, group participants suggested a national effort to drive biomarker development, with NCI as the most likely agency to further this effort and support academic researchers doing this type of work.
A public-private partnership that furthers biomarker development could be modeled after the SNP Consortium. This nonprofit foundation was organized to provide public genomic data, and it was supported by pharmaceutical and technical companies and the Wellcome Trust medical research charity. One discussant indicated that a main impetus for forming the foundation was to prevent academic institutions and industry from claiming intellectual property rights on each SNP they discovered in the human genome. Avoiding intellectual property claims could be an impetus for starting a biomarker consortium as well. The group noted that such claims on each possible biomarker could be a huge impediment to having diagnostic companies develop assays for the biomarkers. Several people in the group felt strongly that biomarker information should be in the public domain, with some stating “the real value of the intellectual property comes
from developing the assays and not just linking an mRNA to a possible outcome,” Dr. Sawyers reported. This raised the problem of how to give diagnostic companies exclusive rights so that they are encouraged to fully develop and commercialize a biomarker.
The group came up with several incentives for biomarker development. Defining the FDA approval path for a biomarker diagnostic more clearly, and linking the approval path for paired diagnostics and therapeutics so both companies share the risks and development costs would provide incentives for biomarker development. It was also suggested that there be patent extensions of innovative biomarker diagnostics to reward the ground-breaking work that one or two companies do that is then used by competing companies to develop similar products. Precedents exist for this enhanced exclusivity in the development of pediatric interventions, and have been proposed for the development of anti-infectives, Dr. Frank noted.
Finally, group participants suggested working with payors to define the cost effectiveness of biomarker tests. “There was a sense that the cost effectiveness of a biomarker was not really appreciated,” he said. “If it were, then reimbursement paradigms could be built in that would incentivize companies to make them sooner.” Group discussants also suggested working with payors to establish alternatives to basing reimbursement decisions on evidence generated from large, long-term clinical trials. CMS and other insurers often require more evidence than does the FDA for a biomarker’s effectiveness prior to reimbursing its clinical use, Dr. Sawyers noted. Several group members suggested that evidence could be generated after the test enters the clinic via community-based postmarketing studies. These studies could be facilitated by using an electronic medical records infrastructure.
Dr. Sawyers concluded his summary by discussing his group’s suggestion that there be a demonstration project to develop biomarkers for drugs already on the market. This project could show the value of using biomarkers to identify the group of patients most likely to respond to the drug, or to identify and exclude those likely to have severe adverse reactions to the drug. Such a proof-of-concept experiment could lay out a path for developing biomarkers and could provide lessons about the appropriate business model to follow and regulatory issues to consider. The reason to use approved drugs for the demonstration project is because patients already taking the drugs can be easily accrued into a study, Dr. Sawyers said. One discussant suggested demonstrating the usefulness of biomarkers that indicate the safety of a number of drugs in a class. Another discussant suggested
using the demonstration project to show the value of biomarkers in predicting responsiveness for two or three drugs widely used in oncology.
If a demonstration project had high-impact findings, it could serve as a catalyst that would spur investment into diagnostic companies and lead more academic institutions and industry to pursue biomarker discovery and development, the group pointed out. Several discussants thought some “success stories” via such a demonstration project would overcome the inertia that is preventing extensive biomarker development. The science needed to do such work is already in place, they noted, and what is lacking is leadership and funding. As an example of a biomarker demonstration project, Dr. Sawyers mentioned the pilot project already under way that was previously described by Dr. Woodcock in her presentation. This is a nonprofit public-private partnership to qualify FDG-PET as a marker for drug response in non-Hodgkin’s lymphoma. Dr. Sawyers’ group also reiterated the need for annotated, quality-assured patient samples that are readily available to further efforts to develop biomarkers.
EVALUATION OF EVIDENCE IN DECISION MAKING DISCUSSION
Dr. Ramsey was the moderator who provided the summary of the discussion on evaluation of evidence in decision making. This discussion group noted that many biomarker-based tests in wide use today were never thoroughly evaluated for analytic validity, clinical validity, or clinical utility in relation to standards. Consequently, their value is often unknown. Group members suggested that this lack of standardized evaluation be eliminated for new tests because the developmental and clinical costs of these tests are quite expensive, and costs also can be incurred if tests are used inappropriately and/or cause undue harm to patients.
Some group participants agreed there is a need for more uniform standards for biomarker evaluation. Dr. Ramsey said there is no consistency regarding standards among organizations and regulatory programs such as the FDA, CLIA, the College of American Pathologists (CAP), and the American Society for Clinical Oncology (ASCO). Each organization has its own set of standards for biomarker tests that are based on different criteria. There is even variability within these organizations, the group noted. In a discussion following Dr. Ramsey’s summary, Dr. Dai pointed out that scientific journals also have their own set of standards for biomarkers. For example, if researchers want to publish gene expression biomarkers, journals
may ask them to compare the biomarkers to what is already available. They may even require that researchers use a specific statistical modeling technique when making such comparisons.
Group members thought the ASCO guidelines for tumor biomarkers for breast or colorectal cancer21 could serve as a potentially useful model in terms of how one might set standards for evaluating whether biomarkers are ready for clinical use. These guidelines established the appropriate levels of evidence needed for different types of clinical decisions made based on biomarker test results. For example, the highest level of evidence was required for a biomarker assay that would indicate the need to deny specific care, that is, one that indicates drug resistance.
However, there was no group consensus on what standards should be required or recommended for cancer biomarkers. This lack of consensus stemmed, in part, from recognizing that there is no gold standard for many of the new kinds of assays used to detect cancer biomarkers, and the evolving nature of those technologies. This made many in the group reluctant to specify standards. In addition, the group recognized that broad, generalized standards alone are not sufficient; guidelines may also need to be use specific and even target specific.
Because the technologies for genomics and proteomics assays are rapidly evolving, the group noted, standards have to be adaptable to the changes in technology that are continually occurring. There is also such a wide range of uses for biomarkers in the cancer arena that standards for one use, such as a surrogate endpoint in clinical trials, may not be applicable to another use, such as a predictor of patient responsiveness. In addition, the standards for a biomarker that predicts responsiveness to a drug may vary depending on the type of cancer on which it is tested, such as lung or breast. However, basic generalized criteria should be met for all clinical tests including biomarker-based tests, the group members recognized. They agreed with Dr. Ransohoff’s assertion in his presentation that the standards of clinical epidemiology still apply to biomarker-based tests.
Working against a common desire to fully evaluate biomarker tests and ensure they meet certain standards is the desire of companies to bring such tests to market as quickly as possible to generate revenues to compensate for development costs. In addition, companies that are developing biomarker tests to be used in combination with specific drugs are often under time pressure to put the drug and the diagnostic on the market at the same time.
Because diagnostic development often lags behind drug development for paired diagnostics and therapeutics, shortcuts may be taken in evaluations of the diagnostic, some discussants pointed out.
Because of such financial and time pressures, companies usually seek the fastest and easiest entry into the market, such as CLIA certification for a home-brew laboratory test, rather than a more rigorous evaluation process by the FDA that might require them to conduct clinical studies. Consequently, few biomarker-based tests are designated Class III devices, which require clinical evidence of their effectiveness and safety.
Competition with other companies also prods the makers of biomarker diagnostics to lower the standards bar in order for their products to go to market before those of their competitors. As Sharon Kim, MBA, of Precision Therapeutics observed, “The challenge has been not just to set your own quality standards for yourself, but you worry and wonder what your potential competitors might be held to because there is no standard, and so are you holding yourself to too stringent of a standard, knowing there may be someone else out there that may place a lower level-of-evidence bar or variability bar out there? While the FDA has the ability to come in and regulate, they have elected not to, and so it is more self-regulated. Even for CLIA-governed or CAP-governed labs, there is no specific cookbook or guidance you can go to.”
Industry representatives in the discussion group pointed out that companies often evaluate their biomarker diagnostics in phases, with a more complete evaluation of their broader applications not occurring until after the tests enter the market. For example, a cancer detection test may at first only be evaluated for its accuracy and predictive value in high-risk populations because this evaluation can be done relatively quickly compared to one done in the general public. But to create a greater market for their products, companies may evaluate them for broader uses once they are already on the market for a more restrictive indication. In that way, companies can quickly bring their products to market and begin gaining revenue on them to help cover the costs of further evaluations. But once a test is on the market, there are few ways to control, beyond coverage decisions, how the test is clinically used.
As was noted in Mr. Heller’s presentation at the conference, the high-variability and rapidly evolving approach to the FDA’s regulation of biomarker diagnostics has created uncertainty as to what evaluations industry needs to do of their tests and what standards to apply, Dr. Ramsey said. The group spent some time discussing whether health insurance payors should
set the standards for biomarker diagnostics. The group noted that if they did, it would add another layer of variability, uncertainty, and complexity that would be problematic for the developers of the tests, especially if there was no agreement among health insurers in this regard.
The group also considered whether the FDA, CMS, and perhaps other stakeholders should work together to develop more uniform standards for the evaluation of biomarkers. But consensus was not reached on this issue, in part because of the tradeoffs involved. Having these agencies set uniform standards would be beneficial in that companies would know what to expect and what would be required of them regarding the evaluation and performance of their biomarker diagnostics. “As long as they are not overly burdensome, they would help us defend our experimental design if we could refer to something else that had been published and widely accepted. That way when the data were reviewed our study design wouldn’t be questioned, which could help speed things through [an FDA approval process],” said Lynne McBride of BD BioSciences.
But Dr. Aronson, the session moderator, said, “there are decisions that come out of CMS and the FDA that are more political than rational and health plans do not follow them.” But she added that it would be valuable to gather together a community of stakeholders to help establish the evidence base needed for biomarkers used clinically.
In a discussion following Dr. Ramsey’s summary, Dr. Waring stressed the need to engage the pathology community when setting standards for biomarker tests. “When we are talking about predictive tests that determine treatment decisions for patients with serious life-threatening diseases, I think that the pathologists and the pathology community are often the afterthought in this process. We need to engage them very early and make sure they understand the consequences of the decisions and that they maintain quality testing,” he said. He noted that CAP and ASCO would be meeting in a few weeks to try to develop common guidelines for HER2 testing.
The group discussed further Dr. Waring’s presentation on the variability among laboratories on the accuracy of the IHC test for HER2. Part of that variability stemmed from the manual, visual, and subjective nature of the test, the group noted. But it is likely that such variability in accuracy will crop up again for other biomarker tests, Dr. Ramsey said. The group debated whether there should be additional measures of quality assurance in such tests. Suggested quality assurance measures included proficiency testing akin to what is now required for cytotechnologists who read Pap
smears, volume requirements akin to what is required for radiologists who read mammograms, and requirements for collecting, analyzing, and reporting data on test performance. There was no consensus on which measures, if any, should be pursued to improve the quality of biomarker testing.
INCORPORATING BIOMARKER EVIDENCE INTO CLINICAL PRACTICE DISCUSSION
Moderator Robert McDonough, MD, of Aetna U.S. Healthcare summarized his group’s discussion on incorporating biomarker evidence into clinical practice. He noted that there are many sources of information on biomarkers that reach clinicians, including journals, colleagues, product vendors, patients, popular media, practice guidelines, clinical trials abstracts, meetings, and continuing medical education. But when the group evaluated what prompts clinicians to adopt biomarker tests into their clinical practices, evidence-based information was not high on the list. “If you are looking at the screening for cancers, there is no correlation between the strength of the evidence and adoption,” said discussant Mark Fendrick, MD, of the University of Michigan.
For example, an impressive 75 percent of the target population undergoes regular screening for prostate cancer, despite the fact the USPSTF gave it an unimpressive I rating. This is in contrast to the 50 percent of the target population who undergo regular colon cancer screening, which the USPSTF gave its highest rating because of its proven effectiveness. Academic practitioners appear to be more influenced by evidence, however, and may delay adopting a new test until there is evidence showing its effectiveness, several discussants agreed. This is in contrast to community practitioners, who may more readily adopt a new test or drug, even when there is little to no evidence of its clinical value. As a consequence, once a product enters the market, it may be impossible to gather the evidence on a test’s clinical value because of difficulties accruing patients to serve as controls for the trials needed to gather that evidence.
Other factors beyond evidence appeared to be more important in influencing the incorporation of biomarker tests into clinical practice, the group noted. The most influential factor they identified was reimbursement for a test at a sufficient level. “If you look at the adoption of CT scans, PSA testing, or even COX-2 inhibitors, until they were paid for, they were not used,” said Dr. Fendrick. Because most diagnostics are relatively inexpensive, insurers are more likely to reimburse their costs without scrutinizing
the evidence base for the test, the group also noted. “If they didn’t pay for even low-cost biomarkers unless they were validated in a proper way, that would be an incentive to do those [validation] studies,” said discussant Dr. Carbone.
The promotion that health insurers and employers do for various tests also influences their use, some discussants pointed out. For example, insurers often promote preventive health tests, such as those used to screen for various cancers, via informational mailings and their websites. “Some employers give discounts on health insurance to employees who undergo a self-assessment that indicates what types of screening and other health maintenance measures they should undertake,” Dr. Carbone said. “I think it is widely adopted when you give people a buck to do it.”
Another highly influential factor was whether the test was adopted by what the group called “thought leaders.” A thought leader is someone who other members of a group look to as an authority. A thought leader may be misinformed, but he or she is still influential. In academic settings, thought leaders tend to be the lead investigators of clinical studies or the chairs of departments. In clinical practices the thought leader “is the clinician down the hall who seems to be knowledgeable about what is new in medical technology,” Dr. McDonough said. He said one discussant noted that physicians who practice in groups seem to adopt technology more rapidly than solo practitioners, possibly because of the presence of thought leaders in group practices.
Another potential driver for the uptake of new biomarker tests is patient requests for the tests, the group noted. Studies reveal that if a patient asks for a drug by name, there is an 80 percent chance that a physician will prescribe it, Dr. Fendrick observed. Presumably patients have the same influence over the tests they request, he suggested.
Through promotional efforts, product manufacturers also influence doctors and patients to use their biomarker tests, Dr. McDonough noted. “What I always thought was an important factor was the guy who knocks on your door—the vendor of the new device or new drug or new test,” he said during his group’s discussion. Dr. Waring also noted that for a test such as the FISH test for HER2, used to determine patient responsiveness to a specific treatment, the pharmaceutical company that provides that treatment may pay the costs of the test if it is not covered by an insurance provider. This is especially the case in Europe where national health plans may not offer the test as part of their services. “Roche until recently was paying for those tests to be performed in their own central laboratories,” he
said. “So these tests were becoming available not because of reimbursement issues—they were being made available by the pharmaceutical company for business reasons.”
Other influences on the clinical adoption of a biomarker test hinge on features of the test itself, the discussion group said. Ease of interpretation is one such feature. If the test is easy to interpret and has a simple positive versus negative result, it will be adopted more readily than a test whose results require “some kind of complex algorithm to understand,” said Dr. McDonough. Clinicians are also more inclined to adopt tests that are reliably accurate and have timely results. “If you need to make a decision today, and the test is going to take 2 weeks, regardless of how easy or reliable that test is, it may not be very clinically useful,” said Dr. McDonough. Clinicians are also more likely to adopt tests if there is little to no risk in using them, and there are no alternative tests or test-linked treatments. Insurers are also more likely to reimburse for both the test and treatment, for those that are linked, if there are no treatment alternatives and the disease the drug targets is life threatening, the group noted.
Inconvenience to the patient is another important test feature that influences its adoption in the clinic. Physicians are more likely to prescribe a simple blood test than an endoscopic procedure or a test that requires a stool sample, Dr. McDonough pointed out. Practitioners are also more likely to use a test that will influence their clinical decision making. “Is it a test that might give you some idea of the prognosis of lung cancer, but will not actually influence the type of therapy you might actually give to the patient? If the test does not seem to have any influence on the clinical management we would hope that would make it less likely that a clinician would use it,” Dr. McDonough said.
Like other discussion groups, Dr. McDonough’s group recognized that low profit margins on diagnostic tests act as a disincentive to the development of biomarker tests and their evaluation in clinical trials. This led to the suggestion by Dr. McGivney that payors help subsidize some of this clinical research. “A payor who is asking for evidence should actually support, in part, the development of some of that evidence,” he said. Dr. McDonough said that some insurers, such as Aetna, do pay for routine costs of their patients in clinical trials. But Dr. McGivney countered that there is an increasing trend for payors not to cover such costs.
Given that reimbursement levels highly influence the adoption of clinical tests, other discussants suggested that payors tailor their copay amounts for biomarker tests based on a test’s value or degree of evidence to support
any positive impact on patient outcomes. Zero copayment amounts could be allotted for those biomarker tests that are highly cost effective and likely to affect clinical management. High copayments could be required for tests whose cost effectiveness is questionable due to a lack of evidence on their benefits.
But the group recognized that “it would not be easy to structure a benefit program to that fine a degree of assigning copays based on someone’s assessment of cost effectiveness,” Dr. McDonough said. There would be legal issues that might be difficult to overcome, such as varying state regulations that affect copayment levels. In addition, both legislators and the insurance clientele might look askance at plans that specify high copayments for treatment-linked tests for life-threatening illnesses.
For payors to more adequately influence the adoption of biomarker tests, those tests need to have their own Current Procedural Terminology (CPT) codes, group members noted. These identifying codes are established by the American Medical Association and are used to report medical procedures and services to health insurers. Health insurers then specify reimbursement rates for each code. CPT codes are also used for developing guidelines for medical care review. “Many of these biomarkers do not have specific CPT codes,” said Dr. McDonough. “They are defined by process steps so that the insurer, even if they were willing to scrutinize biomarkers, often find it difficult to know what type of biomarkers are being used. What this means is that many of these biomarkers are being incorporated into clinical practice without much scrutiny.”
This is especially true for home-brew tests, which are always defined by process steps. These tests, therefore, bypass scrutiny by both regulators and reimbursers, the group noted. Even when a test has been approved by the FDA, some discussants said, there is no guarantee that laboratories will use that test. Instead, they may offer their own home-brew version of the test, which may not be as acurate. Home-brew versions of the HercepTest, Dr. Waring said, help explain the variability in accuracy among laboratories.
In a discussion following Dr. McDonough’s summary, Dr. Ramsey gave an overseas perspective of health care payors playing a role in gathering clinical data to evaluate new products. For example, the United Kingdom’s National Health Service pays for a new drug at an agreed upon price, with the requirement that data on the drug’s effectiveness be collected in a patient registry. If the drug does not show effectiveness at the expected level, the drug’s price is reduced so that the total reimbursement over time
reflects the actual quality of life gain observed. He thought such risk sharing in drug development was valuable, and noted that the group’s suggestion that payors cover the costs of clinical trials on biomarker tests would put all the burden of risk on insurance companies. He suspected they would balk at such a suggestion and reiterated that risk sharing has some value.