ATTACHMENT 2
COMPARISON OF MICROARRAY DATA FROM MULTIPLE LABS AND PLATFORMS
Rafael A. Irizarry
Department of Biostatistics
Johns Hopkins Bloomberg School of Public Health
Baltimore, Maryland
INTRODUCTION
Microarray technology is a powerful tool able to measure RNA expression for thousands of genes at once. This technology promises to be a useful tool to better understand the role of genetics in biologic responses to environmental contaminants. Various examples were presented in the 10th Meeting of the Committee on Emerging Issues and Data on Environmental Contaminants. In this extended abstract I give a short summary of a study organized to compare the three leading microarray platforms. I also give recommendations on experimental issues and statistical analyses. Many of the recommendations are applicable to general problems in technology assessment. More details are available in a Nature Methods paper (Irizarry et al. 2005) which includes various figures and tables referred to in this extended abstract.
MOTIVATION
As a statistician working in the Johns Hopkins Medical Institutions I give advice to various scientists working with microarrays. Various commercial vendors as well as custom made facilities provide many alternative platforms for researchers interested in gene-expression data. A frequently asked questions among those just getting started with this relatively new technology is “which platform performs best?” Various
studies have been published comparing competing platforms with mixed results: some find agreement (Kane et al. 2000; Hughes et al. 2001; Yuen et al. 2002; Barczak et al. 2003; Carter et al. 2003; Wang et al. 2003), others do not (Kuo et al. 2002; Kothapalli et al. 2002; Li et al. 2002; Tan et al. 2003). Why the disagreement? Two possibilities were that 1) different statistical assessments were used and 2) the lab effect was not explored. That different statistical assessments can lead to different conclusions is clear and we will give an example later. To see how the lab effect can result in different conclusions, consider that in all previous studies, platform variation was confounded with technician/lab variation. In the cases where the same lab created all the microarray data it was clear that they had more experience with one of the platforms being compared. The lab effect has been shown to be particularly strong. For example, in a 1972 paper, W.J. Youden (Youden 1972) pointed out how different Physics labs published speeds of light estimates with confidence intervals that made the differences between labs statistically significant (see Figure 2-1). Notice, that if we do not take the lab effect into account this would imply that the speed of light is different in the different labs! It is no surprise that similar effects are present in biology labs and that it does not only apply to microarray measurements.
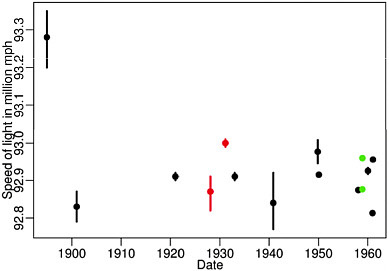
FIGURE 2-1 Speed of light estimates with confidence intervals (1900-1960). Source: Youden 1972. Reprinted with permission; copyright 1972, Journal Information for Technometrics.
Furthermore, in my opinion, previous studies had at least one of these problems: (1) precision/accuracy were not properly assessed (that is, the sensitivity/specificity trade-off was not considered and in general, assessments were based on validation of few genes). (2) There was not an a-priori expectation of truth. In general, RTPCR was considered the gold standard and measurements from that technology considered the “truth.” (3) The effect of preprocessing was not explored. (4) As mentioned, the lab effect not explored.
OUR STUDY
Together with various labs from District of Columbia/Baltimore area, that volunteered their time and materials, I conducted a study for comparing microarray technologies. To overcome the problems of previous studies, we followed methodology that can be summarized by the following steps: (1) We included platforms for which results from at least two labs were available. (2) To avoid a transportation effect, we considered only labs in DC/Baltimore area. Of those we asked, five Affymetrix labs, three two-color cDNA labs, and two two-color oligo labs agreed. (3) We send each lab technical replicates of two RNA samples. (4) In the samples sent to each lab, we included technical replicates of each of the two samples. This permitted us to assess precision. (5) We designed the two RNA samples to induce a-priori knowledge of differential expression of four genes. This permitted us to assess accuracy. (6) Finally, to provide more power to the assessment of accuracy we measured fold-changes for 16 strategically chosen genes. Details are available from the Nature Methods publication (Irizarry et al. 2005).
STATISTICAL RECOMMENDATIONS—WHAT MEASUREMENT?
In our study we evaluated what we consider to be the basic measurement obtained from microarrays: relative expression in the form of log ratios. Thus, for each lab we had two replicate measures of relative expression: M1 = log(B1/A1) and M2 = log(B2/A2), with A1, A2, B1, B2 representing the two pairs of technical replicates provided to each lab.
The first important recommendation is that when comparing and/or combining measurements from different platforms one should look at relative as opposed to absolute measures of expression. This is because
in microarray technology there exists a strong probe or spot effect. The Nature Methods publication (Irizarry et al. 2005) describes in some detail how this can lead to over-optimistic measures of precision when comparing within platform and over-pessimistic measures of agreement when comparing across platforms.
USING RELATIVE EXPRESSION INSTEAD OF ABSOLUTE EXPRESSION
Most experiments compare expression levels between different samples, thus in general this type of measure is readily available. In classification problems, it is typical to use distance measures which implicitely use relative measures. For example, the most popular methods subtract the across sample mean log expression for each gene, which produces relative (to the mean) expression measures.
PRECISION AND ACCURACY ASSESSMENTS
We believe it is important to assess precision in the context of accuracy. A platform that produces results that are perfectly reproducible (precise) is of little practical use if it fails to detect a signal (accuracy). This concept is particularly important with microarray platforms because different pre-processing methodology can lead to differences in precision and accuracy. Typically, better precision can be reached by sacrificing accuracy and vice-versa (Wu and Irizarry 2004).
PRECISION
In our study we used the following simple statistical assessments that have very intuitive interpretation in practice. To assess precision we looked at the standard deviation (SD) of M1 – M2 across all genes. This SD represents (roughly) the typical value of log-fold change when it should be 0. Correlation, which is a commonly used measure, does not have the same simple interpretation. Furthermore, in a comparison where most genes are not differentially expressed the correlation will be correctly attenuated towards 0 which might lead to the incorrect interpretation that the measurements are not reproducible. Note that measurement error should not correlate, but it should have small variance.
ACCURACY
To assess accuracy we computed the slope of M’s regressed against nominal log-fold change. This represents (roughly) the expected log-fold change when it should be 1. This measure combined with the precision assessment gives us a good overall idea of the ability of the technology to detect changes among a large number of instances of no change.
CAT PLOTS
In practice, we typically screen a small subset of genes that appear to be differentially expressed. Therefore, it is more important to assess agreement for genes that are likely to pass this screen. To account for this, we introduce a new descriptive plot: the correspondence at the top (CAT) plot. This plot is useful for comparing two procedures for detecting differentially expressed genes. To create a CAT plot we form a list of n candidate genes for each of the two procedures and then form curves that show the proportion of genes in common plotted against the list size n. These figures can be seen in the Nature Methods publication (Irizarry et al. 2005). As an assessment measure, we can give the proportion of genes in common in lists of specific sizes, for example 25, 50, and 100.
RESULTS
Our results demonstrated that precision is comparable across platforms. With the exception of one lab, all the labs performed similarly, and it is clear that the lab effect is stronger than the platform effect. All the labs appear to give attenuated log2-fold change estimates, which is consistent with previous observations. In general, the Affymetrix labs appear to have better accuracy than the two-color platforms, although the best signal measure was attained by a two-color oligo lab. While two labs (a two-color cDNA lab and a two-color oligo lab) clearly under performed, the differences among the other eight labs do not reach statistical significance. Among the best performing labs we found a relatively good agreement of about 40% among list sizes of 100 (that is, among these labs, there was about 40% agreement on the top 100 genes with the greatest fold change in expression). It is not obvious that for variables that are highly correlated it is hard to get agreement much higher than
40%. Figure 2-2 plots the expected agreement of two bivariate normal random variables against their correlation.
DISCUSSION
Overall, the Affymetrix platform performed best. However, it is important to keep in mind that this platform is typically more expensive than the alternatives. We also demonstrated that relatively good agreement is achieved between the Affymetrix labs and the best performing two-color labs. These results contradict some previously published papers that find disagreement across platforms (Kuo et al. 2002; Kothapalli et al. 2002; Li et al. 2002; Tan et al. 2003). The conclusions reached by these studies are likely due to inappropriate statistical assessments as well as the confounding of lab effects. The existence of the sizable lab effect was ignored in all previously published comparison studies. This permits the possibility that studies using, for example, experienced technicians may find agreement and studies using less experienced technicians may find disagreement. Figure 2-3 plots the precision assessment against experience for the Affymetrix labs.
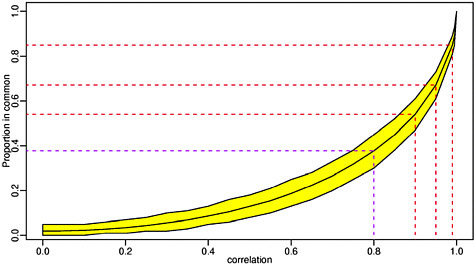
FIGURE 2-2 Expected agreement of two bivariate normal random variables against their correlation.
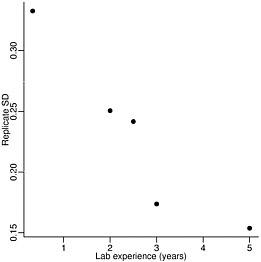
FIGURE 2-3 Precision versus years of experience for labs using Affymetrix microarrays. Source: Irizarry et al. 2005. Reprinted with permission; copyright 2005, Nature Methods.
Although we found relatively good across platform agreement it is quite far from being perfect. In all across-platform comparison, there was a small group of genes that had relatively large fold changes for one platform but not for the other. We conjecture that some genes are not correctly measured, not because the technologies are not performing adequately, but because transcript information and annotation can still be improved.
Our findings illustrate that improved quality assessment standards are needed. Assessments of precision based on comparisons of technical replicates appear to be standard operating procedure among, at least, academic labs. Precision and accuracy assessments are not informative unless performed simultaneously. Figure 2-4 shows observed versus nominal log-fold-changes for two labs with similar precision. Labs that perform well in terms of accuracy will show points near the diagonal line. Notice these two labs had similar precision, but Lab 3 had much better accuracy. We hope that our study serves as motivation for the creation of such standards. This will be essential for the success of microarray technology as a general measurement tool.
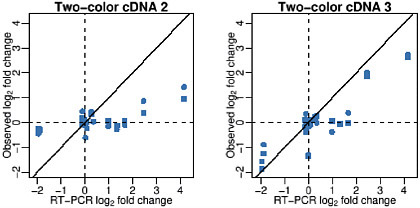
FIGURE 2-4 Observed versus nominal log-fold change for two labs (two-color cDNA 2 and two-color cDNA 3) with similar precision. Source: Irizarry et al. 2005. Reprinted with permission; copyright 2005, Nature Methods.
REFERENCES
Barczak, A., M.W. Rodriguez, K. Hanspers, L.L. Koth, T.C. Tai, B.M. Bolstad, T.P. Speed, and D.J. Erle. 2003. Spotted long oligonucleotide arrays for human gene expression analysis. Genome Res. 13(7):1775-1785.
Carter, M.G., T. Hamatani, A.A. Sharov, C.E. Carmack, Y. Qian, K. Aiba, N.T. Ko, D.B. Dudekula, P.M. Brzoska, S.S. Hwang, and M.S. Ko. 2003. In situ-synthesized novel microarray optimized for mouse stem cell and early developmental expression profiling. Genome Res. 13(5):1011-1021.
Hughes, T.R., M. Mao, A.R. Jones, J. Burchard, M.J. Marton, K.W. Shannon, S.M. Lefkowitz, M. Ziman, J.M. Schelter, M.R. Meyer, S. Kobayashi, C. Davis, H. Dai, Y.D. He, S.B. Stephaniants, G. Cavet, W.L. Walker, A. West, E. Coffey, D.D. Shoemaker, R. Stoughton, A.P. Blanchard, S.H. Friend, and P.S. Linsley. 2001. Expression profiling using microarrays fabricated by an ink-jet oligonucleotide synthesizer. Nat. Biotechnol. 19(4):342-347.
Irizarry, R.A., D. Warren, F. Spencer, I.F. Kim, S. Biswal, B.C. Frank, E. Gabrielson, J.G. Garcia, J. Geoghegan, G. Germino, C. Griffin, S.C. Hilmer, E. Hoffman, A.E. Jedlicka, E. Kawasaki, F. Martinez-Murillo, L. Morsberger, H. Lee, D. Petersen, J. Quackenbush, A. Scott, M. Wilson, Y. Yang, S.Q. Ye, and W. Yu. 2005. Multiple-laboratory comparison of microarray platforms. Nat. Methods 2(5):329-330.
Kane, M.D., T.A. Jatkoe, C.R. Stumpf, J. Lu, J.D. Thomas, and S.J. Madore.
2000. Assessment of the sensitivity and specificity of oligonucleotide (50mer) microarrays. Nucleic Acids Res. 28(22):4552-4557.
Kothapalli, R., S.J. Yoder, S. Mane, and T.P. Loughran, Jr. 2002. Microarray results: How accurate are they? BMC Bioinformatics 3:22
Kuo, W.P., T.K. Jenssen, A.J. Butte, L. Ohno-Machado, and I.S. Kohane. 2002. Analysis of matched mRNA measurements from two different microarray technologies. Bioinformatics 18(3):405-412.
Li, J., M. Pankratz, and J. Johnson. 2002. Differential gene expression patterns revealed by oligo-nucleotide versus long cDNA arrays. Toxicol. Sci. 69(2):383-390.
Tan, P.K., T.J. Downey, E.L. Spitznagel, Jr., P. Xu, D. Fu, D.S. Dimitrov, R.A. Lempicki, B.M. Raaka, and M.C. Cam. 2003. Evaluation of gene expression measurements from commercial microarray platforms. Nucleic Acids Res. 31(19):5676-5684.
Wang, H., R.L. Malek, A.E. Kwitek, A.S. Greene, T.V. Luu, B. Behbahani, B. Frank, J. Quackenbush, and N.H. Lee. 2003. Assessing unmodified 70-mer oligonucleotide performance on glass-slide microarrays. Genome Biol. 4(1):R5.
Wu, Z., and R.A. Irizarry. 2004. Preprocessing of oligonucleotide array data. Nat. Biotechnol. 22(6):656-658.
Youden, W. 1972. Enduring values. Technometrics 14:1-11.
Yuen, T., E. Wurmbach, R.L. Pfeffer, B.J. Ebersole, and S.C. Sealfon. 2002. Accuracy and calibration of commercial oligonucleotide and custom cDNA microarrays. Nucleic Acids Res. 30(10):e48.









