ATTACHMENT 3
STATISTICAL ANALYSIS OF TOXICOGENOMIC MICROARRAY DATA
Wherly Hoffman, Ph.D. and Hui-Rong Qian, Ph.D.
Statistical and Information Sciences
Eli Lilly and Company
Indianapolis, Indiana
Microarray is a relatively new technology allowing scientists to measure the expression level, or mRNA abundance of thousands of genes in a single experiment. Combined with other “omics” technologies, e.g., proteomics and metabonomics, this advance provides a completely new systems biology approach to investigate biologic problems, and makes it possible to study the huge dynamic gene interaction matrix in cells. The application of microarray in toxicity studies leads to the emergence of a new research area, toxicogenomics. Genomic studies help scientists understand the mechanism of gene function and pathway, and how a drug may alter a biologic process to treat a disease. Toxicogenomic experiments are conducted to do this with the goal of identifying the no-observed-effect level (NOEL) or no-observed-adverse-effect level (NOAEL). Since it may take a long time to develop an adverse effect while the gene expression change is relatively immediate, toxicogenomics studies have the potential for an early and accurate detection of toxic effects. Different technologies have emerged in this research area. In particular, Affymetrix’s GeneChip has been widely adopted in the drug discovery process in the pharmaceutical industry. Moving from making a scientific conjecture of a compound’s toxicity to analyzing gene expression profiles to concluding the association between the toxicity and gene alteration is a complex process. Statistical considerations are vital in each step of the process.
This presentation is focused on the statistical analysis of Affymetrix GeneChip. microarray data. The Affymetrix GeneChip technology, sta-
tistical hypotheses, and statistical analysis methods will be discussed in order. Visualization of the data and analysis results will be included. Conventional hypothesis testing on such a massive amount of data leads to severe multiplicity issues. Proper multiplicity adjustments for P values will be discussed to help distill massive amounts of information into useful information for each compound.
AFFYMETRIX GENECHIP TECHNOLOGY
On an Affymetrix GeneChip, each gene (probe set) is represented by a set (11-20 pairs) of paired short oligonucleotides of 25-base long, called perfect match (PM) and mismatch (MM) oligos. PM oligos match the gene sequence exactly so after hybridization with labeled sample RNA, they reflect the expression signal, MMs have the same sequences as PMs except that the middle base is changed to its complementary nucleotide. MMs are designed to capture the non-specific hybridized signals, or background signals. There are dozens of algorithms to extract a robust signal intensity from these 11-20 pairs of PMs and MMs for each probeset (Cope et al. 2004). The three most commonly used are MAS 5 from Affymetrix (Affymetrix 2002), the robust multi-array average (RMA) by Irizarry et al. (2003), and the model-based expression index (MBEI) by Li and Wong (2001). It is still not settled as to which method is the best. The final choice is often up to the researcher’s personal preference. In this presentation, signals extracted using MAS 5 from Affymetrix are statistically analyzed.
STATISTICAL HYPOTHESES
Every experiment is designed to answer certain scientific questions. It is important that before conducting an experiment, the researchers define scientific questions and statisticians translate the scientific questions into statistical hypotheses and determine appropriate statistical analyses. It is also important to make sure that at the end of the experiment, appropriate and right amounts of data have been collected for statistical analyses and for answering the scientific questions. Considering the expensive price tag of microarray chips and the large amount of time and other resources needed to carry out these experiments, it would be unwise to have problematic designs that could not provide answers to the scientific questions. In most microarray experiments, the primary goal is to iden-
tify differentially expressed genes under different treatment conditions. We will examine important sources of variation and discuss some exploratory and inferential analyses. We will provide some examples to show how different scientific questions lead to different experimental designs and statistical hypotheses.
Microarray measures the expression abundance of essentially all the genes in a genome. With thousands of measurements and relatively few subjects (tens, rarely over a hundred), any difference in the conditions of the subjects to be tested would cause a large number of genes to show expression changes. Therefore, it is very important to understand and control different sources of variability in microarray experiments. Typically there are two sources of random variability: biologic variation and technical variation. The biologic variation exists in the tested subjects. Sometimes it is possible to reduce the biologic variation, e.g., by using more homogenous individuals. Technical variation lies in the sample preparation and the microarray technology itself, including tissue collecting, RNA isolation, labeling, chip hybridization, etc. As the microarray technology matures, the chip-to-chip variation decreases. Still, large variation is observed during the sample preparation. For example, different labeling kits could lead to a big difference in the expression signals. Even with the same labeling kit, samples processed on different days may yield very different signals.
A related issue is the pooling of RNA samples from animals. During the early discovery phase when resources are more limited, pooling of the samples from animals in the same group may be necessary. An example of this is for establishing a surrogate assay to screen peroxisomal proliferation activated receptors (PPAR). While pooling samples allows researchers to use fewer chips, it loses the ability to measure individual expression and provide the estimation of biologic variation for proper statistical tests. It may be advantageous if samples are very cheap and easy to obtain (like in cell cultures), and a partial pooling is conducted in which samples from the same treatment are pooled to form multiple independent pools; thus the biologic variation can still be properly estimated. In general, for follow-up evaluation of observed toxicity, RNA samples from animals are not pooled. For example, upon observing heart weight changes or skeletal muscle necrosis in rats, microarray technology is applied to help find biomarkers and one genechip is used for each animal.
Another important issue is to avoid potential bias by randomization and/or proper blocking. For example, the processing batch effect may be significant. Two samples can appear very different if processed on two
different days, especially if it involves a change of reagents. It has been consistently shown that samples, even without any treatment, are different if the samples are collected at different times. This difference can be well detected by microarray technology.
EXPLORATORY DATA ANALYSIS BY VISUALIZATION
After a microarray experiment is done and data are generated, it is important to explore the data using some basic analytical and graphical tools to detect obvious patterns, potential outliers, errors during the experiments, etc. The following analysis and visualization approaches serve these purposes well:
-
Side-by-side box plot of log signals of each chip. It is helpful to check the chip signal distribution and to see if normalization has been applied.
-
Pairwise chip signal correlation. A very low correlation coefficient (CC) of one chip with other chips is an indication of a quality problem of this chip. High average CC within a treatment group and low average CC between treatment groups indicate a large treatment effect in an experiment. However, the variances of different genes are different. In the Affymetrix platform, a large signal intensity corresponds to a large variance on a raw scale, and thus has a high influence on the CC calculation. On a log scale, a large signal intensity corresponds to a small variance. The cube root transformation could be applied to stabilize the variance.
-
MVA plot of a pair of chips or average signals of treatment groups. This is a scatter plot of average log differences of a pair of chips vs. the average mean of their log signals. This would show clearly the dependence of the variance on signal intensity. It is also helpful to see if the data scatter around 0, a good approach to check if extra normalization is needed.
-
Principal components analysis (PCA). PCA helps reduce the data dimension and reveal the general pattern in data. A clear departure of a chip from other chips with the same treatment on a PCA plot indicates the existence of outliers. PCA plots may also reveal expression patterns that are due to unrealized blocking effects, arising from the sample preparation or array processing, the bias to eliminate during analysis.
Statistical Analysis and Visualization of Results
Although in the beginning of microarray technology, people assumed that a single model for expression could be applied to all genes on an array, it is understood now that the expression of each gene is different and should be modeled on a gene-by-gene basis. The analysis of microarray data depends on the questions we want to answer and the experimental design. To identify differentially expressed genes, in a two-group comparison, a simple t-test is usually applied. Considering the high variability when expression signal is small, it has been proposed to add an inflation factor to stabilize the variance (Tusher et al. 2001), or use a regularized t-test to minimize the dependence of variance on signal intensity (Baldi and Long, 2001). When the experiment has more than two treatment groups, an ANOVA t-test may be applied to better estimate the variability and lead to more powerful tests. In a more complicated situation, e.g., with technical replicates, or with repeated measurements, a mixed effects model may be used (Chu et al. 2002).
Sometimes in an experiment, we have multiple time points or multiple doses, and the interest is to investigate the temporal expression pattern or dose response. We may fit the data to a linear regression model to measure the general expression trend. We will present some examples to show different analysis approaches for different data. In general, when the test is applied to each gene, the statistical approach is not different from what has been used in classic biologic research. However, after the initial statistical test, since it is “fishing” significant gene expression changes out of thousands of genes, adjustments should be made to the resulting P values to control the false positive rate.
In a typical microarray experiment, usually there are hundreds or even thousands of genes being identified as significant. Some visualization tools to show the general patterns of analysis results are listed here:
-
MVA plot with highlighted significant genes. As described above, this plot is helpful to show the signal intensities of significant genes, and to check if the data are properly normalized.
-
Volcano plot. This is the plot of P values or adjusted P values with fold changes, showing if the significant genes are equally distributed over up- or down-regulation.
-
Clustering of genes. There are different ways of clustering over different distance matrices. The general purpose is to group genes with similar expression patterns together. Physiological
-
data if available can be included in the clustering to identify genes closely correlated with the phenotypes.
-
Heat map (see Figure 3-1). When there are a large number of genes that are significant, after clustering, we may rescale the gene expression data within each gene and use a heat map to show the expression patterns.
-
For individual genes of interest, the expression signals of each sample may be plotted, overlaid with any physiological data of interest if available. This helps researchers decide for a particular gene if outliers (or non-responders) exist, if modeling approaches are appropriate, etc.
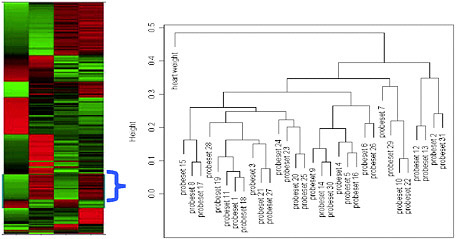
FIGURE 3-1 Heat map and gene cluster with physiological data. The cluster shows only the highlighted portion of the heat map.
MULTIPLICITY ADJUSTMENT
The power of microarray to monitor the expression of thousands of genes in one experiment enables the researchers to investigate the gene function systematically at the genome level. However, testing thousands of genes in a single experiment introduces a high rate of false positives when no efforts are taken to control the false positive rate. For example, the new human chip U133plus2 contains 54,000 probesets. If there is no treatment effect for a comparison so that all the P values are from a uni-
form distribution (0, 1), then there would be about 2,700 significant probesets (P < 0.05).
Bonferroni is one approach to control the false positive rate by controlling the family-wise error rate (FWER). Due to its conservativeness, when applied in microarray data analysis, it leads to too few rejections, i.e., the test power is too low to detect real useful information about gene expression changes. Some of the most recent progress in addressing multiple testing problems is the introduction of false discovery rate (FDR) by Benjamini and Hochberg (1995). FDR is defined as the expected rate of erroneous rejection of hypotheses among total rejected hypotheses. Let M be the total hypotheses being tested, R be the number of rejected null hypotheses, and V the number of falsely rejected true null hypotheses, then
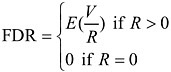
With this setting, the conventional per comparison error rate (PCER) and
Instead of controlling the FWER, procedures controlling FDR control the error rate of false rejections in the rejected hypotheses. Under complete null hypotheses (i.e., all the null hypotheses are true), FDR and FWER are the same. When some of the null hypotheses are not true, FDR is smaller than FWER, thus FDR-controlling procedures are more powerful than FWER-controlling procedures (but note they control two error rates). In microarray experiments, the researchers try to identify genes showing differential expressions across treatment groups. They are more interested in how many genes identified in a microarray experiment will fail to be confirmed in the follow-up studies, i.e., the false-positive rate among the “discoveries.” This provides a perfect setting to apply FDR-controlling methods.
When addressing the multiplicity issue and deciding what gene expression changes are significant using FDR or other approaches, the researchers assume that all gene expressions are independent and equally
important. These assumptions are typically not true. A list of genes of interest may be available before the microarray experiment is conducted. What FDR cutoff values to use to pick follow-up genes and how to apply the FDR adjustment will be discussed.
REFERENCES
Affymetrix, Inc. 2002. Statistical Algorithms Description Document [online]. Available: http://www.affymetrix.com/support/technical/whitepapers/sadd_whitepaper.pdf [accessed April 10, 2006].
Baldi, P., and A.D. Long. 2001. A Bayesian framework for the analysis of microarray expression data: Regularized t-test and statistical inferences of gene changes. Bioinformatics 17(6):509-519.
Benjamini, Y., and Y. Hochberg. 1995. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B Met. 57(1):289-300.
Chu, T.M., B. Weir, and R.D. Wolfinger. 2002. A systematic statistical linear modeling approach to oligonucleotide array experiments. Math. Biosci. 176(1):35-51.
Cope, L.M., R.A. Irizarry, H.A. Jaffee, Z. Wu, and T.S. Speed. 2004. A benchmark for Affymetrix GeneChip expression measures. Bioinformatics 20(2):323-331.
Irizarry, R.A., B. Hobbs, F. Collin, Y.D. Beazer-Barclay, K.J. Antonellis, U. Scherf, and T.P. Speed. 2003. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4(2):249-264.
Li, C., and W.H. Wong. 2001. Model-based analysis of oligonucleotide arrays: Expression index computation and outlier detection. Proc. Natl. Acad. Sci. USA 98(1):31-36.
Tusher, V.G., R. Tibshirani, and G. Chu. 2001. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98(18):5116–5121.








