3
Scientific Session III: Systems Biology
Moderated by Robert Cousins, Boston Family Chair in Nutrition, Director of the Center for Nutritional Sciences, and Affiliate Professor of Biochemistry, University of Florida
Throughout the 20th century, much of biology embraced reductionist goals. That is, researchers broke systems down into their component parts to understand them. This was perhaps most obvious in molecular biology, as scientists explored genes, the products of their expression, and the molecular pathways through which they act. The expectation has always been that an understanding of the individual pieces would eventually allow greater understanding of biological systems on a broader scale.
In the 21st century, biologists will find themselves dealing increasingly with integrated approaches, said Robert Cousins. Systems biology, which is the study of the interactions among the components of a biological system, exemplifies this integrated approach.
The ultimate goal of systems biology is to create a computational model that describes the system of interest. Quoting Leroy Hood, one of the pioneers of systems biology, Cousins remarked that systems biology generally follows a pattern of first defining the elements of a system and then defining a mathematical model for the system, performing simulations with that model, comparing the simulations with what is known about the real system, refining the model based on those comparisons, and repeating those steps again and again to zero in on a model that accurately captures the behavior of the system itself. Cousins recalled Hood’s comments from a conference at the National Institutes of Health in 2005: “Dr. Hood was asked: what are the two areas of science that are most amenable to the systems biology approach and his answer was plant molecular biology and nutritional sciences.”
GENETIC NETWORKS AND APPLIED SYSTEMS BIOLOGY
Presented by C. Ronald Kahn, President and Director, Joslin Diabetes Center, Harvard University
Diabetes and the Insulin-Signaling Network
Understanding a biological system demands understanding the details of its components, and their interactions. As an example, Ronald Kahn described his research into the pathways that control insulin action.
Kahn began by noting that the current epidemic of diabetes and obesity is having a tremendously deleterious effect on human health in the United States. He added, however, that “diabetes and obesity are really a small part of a much larger problem that we call the metabolic syndrome.” The metabolic syndrome includes a constellation of diseases that are linked to insulin resistance and, in many cases, to the effects of overnutrition and underactivity. These diseases include glucose intolerance and type II diabetes; obesity; hypertension; lipid abnormalities; accelerated atherosclerosis; fatty liver; reproductive dysfunction, particularly polycystic ovarian disease in women; and even Alzheimer’s disease and neurodegenerative diseases, which are more common in individuals with insulin-resistant states (see Box 3-1).
|
BOX 3-1 Glucose Metabolism and Insulin Resistance A stable level of glucose in the blood is necessary to provide energy to the brain, muscles, and organs; and excess energy is stored in fat tissue. When glucose levels in the blood decrease, pancreatic beta cells produce glucagon, which stimulates the liver to mobilize glycogen stores to release glucose into the bloodstream. When glucose levels rise, pancreatic alpha cells produce insulin, which inhibits glucose output from the liver and stimulates muscle and fat tissue to absorb glucose from the blood. In insulin resistance the liver, muscle, and fat do not respond to the presence of insulin, which in turn leads to elevated blood glucose levels. This increases signaling to tissue receptors, which respond, although at a minimal level, allowing glucose levels to remain elevated. In type II diabetes, however, tissue receptors are desensitized to glucose stimuli and blood sugar levels rise, leading to symptoms of diabetes and, if left uncontrolled, to diabetic complications. |
To understand the genesis of the metabolic syndrome and what it will take to address it, Kahn posed three questions:
-
What are the key tissues and pathways of insulin action that are involved in the development of insulin resistance and the metabolic syndrome?
-
What important features of the insulin-signaling network play a role in insulin-resistant states and the metabolic syndrome?
-
How do genes and the environment, especially diet, contribute to gene expression changes that might interact with the development of these syndromes?
To address these questions, Kahn primarily relied on research from the Diabetes Genome Anatomy Project, whose goals are “to use mainly genomics and to some extent proteomics to define normal action of insulin on gene and protein expression in cells, mice, and in humans; to define the abnormalities of gene expression in insulin-resistant states; and to determine the role of genetic variation of insulin-signaling proteins in this.” In short, the goals are to understand insulin action on a fundamental level and to use that understanding to pinpoint what goes wrong (and why) when insulin resistance develops.
The insulin-signaling network has two major pathways. The Ras-MAP kinase pathway is involved in regulating gene expression and cell growth and differentiation. The PI 3-kinase pathway modulates the metabolic effects of insulin; the ability to stimulate glucose transport into the cell; and initiates glycogen, lipid, and protein synthesis. To study these pathways and their roles in insulin resistance, the Kahn laboratory has produced mouse models with knockouts of single genes in the insulin-signaling pathway; compound knockouts, either homozygous or heterozygous, of the genes in the pathway; various types of tissue-specific knockouts; and the use of RNAi and shRNA to do gene knockdowns in various tissues (Figure 3-1). These mouse models and the cell lines created from them have allowed investigators to examine signaling proteins and their respective pathways and the physiologic effects of their gene-knockout strategies using a systems biology approach.
An important outcome from this approach is the discovery that the insulin-signaling pathways are not redundant. They are complementary pathways of insulin action, which increases their complexity. For example, unlike knockout mice, in humans there is no single flaw that leads to insulin resistance. According to Kahn, “In human Type II diabetes and in all of the precursor states related to the metabolic syndrome, we think we are looking at polygenic diseases created by partial defects in either
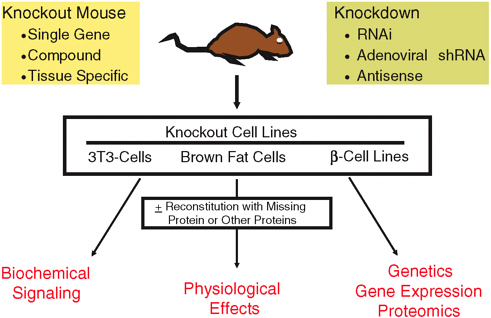
FIGURE 3-1 Knockout and knockdown technology used to study insulin-signaling pathways.
SOURCE: Presentation by C. Ronald Kahn, used with permission, January 9, 2007.
gene expression or gene function and that these lead to more subtle phenotypes.”
As a result, the Kahn laboratory created mice that had heterozygous knockouts for both the insulin receptor (IR) gene and the IR substrate 1 (IRS-1) gene, so that mice expressed both genes at only half the normal level. A heterozygous knockout of either gene failed to create a diabetic phenotype in the mice, leading to the question of what would happen if both genes had 50 percent defects.
The answer was somewhat surprising, in that the effect varied greatly depending on the genetic background of the mouse. In strain C57 Bl6 mice, 90 to 100 percent of the animals would develop diabetes within the first 6 months of life, but in strain 129 mice less than 2 percent would develop diabetes even if they were monitored to 2 years of age. “It turns out that this phenotypic difference is not due to differences in insulin secretion,” Kahn said, “but is due to additional defects in insulin action.” A genomewide scan and gene expression experiments to identify variations that contributed to the phenotypic difference revealed four loci on three chromosomes that play a role. Furthermore, not only did the loci contribute to the difference in how the IR and IRS-1 heterozygous knockouts affect the mice, but they also contributed to differences in how the
two strains developed insulin resistance in response to a high-fat diet. Interestingly, the C57 Black 6 mouse is fatter than the 129 mouse when they are on the same diet, and the C57 Black 6 mouse actually eats less and moves around more; the C57 Bl6 mouse was just more efficient at storing calories.
In looking for an explanation of the phenotypic differences between the mice, investigators identified one locus on chromosome 14 that had a strong correlation with the differences between the mice in their development of insulin resistance; however, there were large differences in the levels of gene expression between the strains of mice. Investigators identified about 250 differences in gene expression overall that could contribute to the phenotype, and 12 of them were located within a specific region on chromosome 14. Many of the genes showed a twofold, fivefold, or eightfold difference in expression between the two strains of mice. The difference in gene expression between these two normal strains of mice was greater in some cases than the difference between heterozygous knockouts or the RNAi and shRNA knockdowns. These experiments demonstrated that gene expression can be modified by disease status; in this case, the diabetic mouse had a gene expression pattern different from that of a healthy mouse of the same strain. “We not only have to consider the background genetics of the animal, but we have to consider all of the extrinsic factors which can be regulating gene expression,” Kahn said.
It is important to understand exactly what is causing the change in gene expression. In the case of diabetes, for example, the changes could be the result of a decrease in insulin levels, but they could also result from the metabolic state created by the diabetes. To distinguish between the two, the Kahn laboratory looked at gene expression in mice that were made diabetic by the administration of streptozotocin and the MIRKO (muscle insulin receptor-knockout) mouse, whose muscles cannot respond to insulin, and MIRKO mice made diabetic with streptozotocin. Each type of mouse has a large number of genes whose expression is different from those in the normal mouse, but there is a small overlap between these two sets of genes. The experiments revealed additional changes in gene expression, but they only partially overlapped with the diabetic state in the normal mouse. Therefore, there are genes that can be viewed as being directly regulated by the insulin signal, for example, those that are changed in the MIRKO mouse, those that are changed in the diabetic animal but not in the MIRKO mouse, and those that are discordantly regulated between diabetes and insulin signaling (that is, the absence of insulin signaling causes an increase in gene expression, but the hyperglycemic state represses expression; in effect, the steady-state levels are not changed).
Researchers face a tremendous challenge in understanding the details
of biological systems. Nutrigenomics will be a complex process. Evaluation of the most well-characterized of metabolic disorders like obesity, type II diabetes, and the metabolic syndrome will require rethinking the entire nature of this system because not only can insulin directly regulate metabolism and gene expression in its target tissues, but the action of insulin is also affected tremendously by nutrients, the genetic background of the animal, and the nutritional state at the time of study. Only by dissecting all of these individual components will real understanding of the true control of gene expression in response to hormones and nutrients be achieved.
GENOME-SCALE RECONSTRUCTION OF THE HUMAN METABOLIC NETWORK
Presented by Bernhard Palsson, Professor of Bioengineering and Adjunct Professor Medicine, University of California, San Diego
One of the core goals in systems biology is to construct models of networks, particularly computational models that can be tested and interrogated mathematically. It is only by creating a well-tested, mostly complete model that a network can be truly understood. This process is done in steps, explained Bernhard Palsson. The first step is the acquisition of data, the second is reconstruction of the network and the creation of a mathematical model that captures the reconstruction (Box 3-2), and the third is computation with the model to make predictions about the
|
BOX 3-2 Network Reconstruction Bernhard Palsson described network reconstruction as being analogous to a map of Los Angeles, California, highways. There is the physical network, the set of roads itself, but also the functional state, that is, the arrangement of cars and other vehicles on the roads. That functional state at 5 p.m. will be very different from that at 5 a.m. Similarly, a gene network or a metabolic network will have different states, depending on the situation. It is generally not possible to compute the particular functional state of a network because that would require more information than is practically available. It is possible, however, to compute the allowable functional states on the basis of various constraints. What all the cars on the road are doing may not be known, but it is certain that there is not a line of traffic traveling through downtown at 80 mph. For a reconstruction, according to Palsson, one can determine a “cone” of allowable solutions so that everything inside the cone represents a possible state of the system and everything outside is a state that is not possible. |
network’s behavior. “The promise of systems biology that these computational models can be used for prospective experimentation is just beginning to be realized.”
Network reconstruction (Step 2) can be approached in two different ways: from the top down or from the bottom up. The top-down approach performs various statistical calculations with data from a high-throughput data set and looks for correlations. In a gene network, for example, one would look for pairs of genes that were always expressed at the same time or for genes that were negatively correlated, with one of them on whenever the other was off, and vice versa. The approach is a “very coarse-grained resolution” of how gene networks function; nevertheless, it does have some utility.
The bottom-up approach, on the other hand, requires more human effort: reading the literature, manually evaluating the process step by step, and evaluating the data component by component. Each connection between components stems from experimental evidence and is described mathematically in terms of input and output, addition and subtraction, and starting point and end point. The bottom-up approach aims to be very accurate and has well-defined chemical interactions among cellular components. The results are self-consistent, in which no information is included in the database until it has been checked and rechecked against experimental results and against other items in the database. The contents of the databases are biochemically, genetically, and genomically accurate and represent knowledge in a structured format. By comparison, the top-down approach aims at being comprehensive; but in trying to measure everything simultaneously, the results are often inconsistent and the final conclusions soft or suggestive rather than coherent and solid. These are often called “inference methods,” based on observation and correlation.
As a “context for content,” such models can bring the top-down and bottom-up approaches together. This is because the most useful models are those that can integrate data derived by the top-down and bottom-up approaches. To date, bottom-up reconstructions have been done for the metabolic networks of a number of microbes, including Escherichia coli, Haemophilus influenzae, and Helicobacter pylori, and have proven to be quite useful. “These models are actually quite good at computing the consequences of gene knockouts. They can also compute the optimal growth rates of cells. Surprisingly, they can predict the outcome of adaptive evolution, which is quite a complicated biological process.” Microbial reconstructions have also been used to study horizontal gene transfer and the evolution of a complex bacterial genome to a simple bacterial genome. Importantly, reconstructions have proven to be a valuable tool in filling the gaps in literature knowledge. Once a map is put together, knowledge gaps remain that emphasize areas in research that require further explora-
tion. However, it is now possible to develop automated methods that fill these gaps and generate hypotheses that lead to biochemical experiments and the discovery of unknown functions of organisms.
The Palsson laboratory has completed a genome-scale reconstruction of the human metabolic map. “It took 18 months for six people to do this, and the map is now complete and has been published,” he said. “This will be of interest to everybody studying nutrigenomics.” To create the reconstruction, the group began with data from various databases, such as the human genome sequence and lists of reactions and metabolites. After the assembly of all those data, the group members reviewed the entire assembly, component by component, fitting the components together, followed by comprehensive evaluations of literature to fill in everything that was not found in databases.
According to Palsson, it is relatively easy to go through an annotated genome, identify the metabolic genes, determine the biochemical reactions that certain gene products catalyze and elementally balance them, determine their cellular location, and then review the relevant literature. Finally, functional testing of the network is performed once it has been put into a mathematical framework wherein the network is checked for gaps. The first version of the reconstruction, Recon 1, accounts for 1,496 open reading frames, 2,004 proteins, 2,766 metabolites, and 3,311 metabolic and transport reactions. However, this is not considered a complete map. For example, metabolomic profiling of human blood and tissues and the construction of lipid maps has documented thousands of potential metabolites and lipids. Not all of these identified compounds may be real; some of them will likely be “false peaks,” while others could be real. “We now will be able to reconcile metabolic data against this map, and hopefully we will find metabolites that are missing and need to be included in the network,” Palsson said. “We still have some things to discover.” After the reconstruction was completed, the Palsson group tested it by using their model to carry out 288 functional tests to check for metabolic capabilities known to exist in human physiology.
Another application important to nutrigenomics is computational interrogation of the model. That is, once the model has been validated, it can be used to study aspects of metabolism that are not easily assessed through laboratory experimentation. It can be used, for instance, to study disease states of metabolism and determine how they differ from normal states or to perform computational experiments on the effects of various nutrients on the metabolic network.
Perhaps most importantly, the reconstruction should serve as the basis for what Palsson hopes will become a rapidly expanding resource. “We now have a platform,” he said, “which different groups can expand to different cell types based on expression profiling of particular organ-
elles as well as other types of high-throughput data that may be of interest. This will be a community resource that I would like to build on the web and have people contribute to and iteratively build. I may not be the best person to do that, so maybe we can get a group of bioinformatics labs to continue this process.”
EMERGING TECHNOLOGIES: NANOTECHNOLOGY
Presented by Martin Philbert, Professor of Toxicology and Senior Associate Dean for Research, University of Michigan School of Public Health
Nutrigenomics researchers will need a variety of tools for studying the cell and its contents, including DNA, proteins, and nutrients. As an example of some of the innovative tools that are becoming available, Martin Philbert described how he, Raoul Kopelman, and their coworkers have used nanotechnology to peer into living cells and see their activity in real time and in three dimensions.
“Nano” refers to the scale of a structure, in which at least one relevant dimension is 100 nanometers or less, and “nanotechnology” is the use of nanoscale structures in research and development. At the nanoscale, materials tend to have properties that are very different from those of the same materials at larger scales. Philbert explained that much of the power of nanotechnology comes from a bottom-up approach to constructing nanostructure devices, which makes it possible to create molecular assemblies with unique properties that can be manipulated with an unprecedented degree of control.
An example of a practical nanostructure application is the use of zinc oxide nanoparticles in sunscreen. When these particles are less than 100 nanometers in diameter, they are able to capture high-energy photons and convert them into low-energy photons, which are less harmful to the skin, thus allowing for a sun protection factor (SPF) of 50 or 60 instead of the usual SPF 15 or 20.
A molecular application developed in the Philbert laboratory is the creation of nanostructures that can be used as intracellular sensors. These nanosensors are able to enter a cell and measure metabolic activity with minimal perturbation of cell function. These nanosensors are termed “PEBBLEs” (probes encapsulated by biologically localized embedding). “We can make these within the range of 20 to 600 nanometers in diameter with very fine control of the mean diameter of the sensors,” Philbert said. “At 20 nanometers one PEBBLE occupies one part per billion of the neuronal somata of the average anterior horn motor neuron cell body, so
a large number of them can be inserted without interfering significantly with cellular functions.
A key feature of PEBBLE structures is that they can be engineered to fluoresce in the presence of various small molecules or ions, such as oxygen, nitric oxide, calcium, potassium, and zinc, among many others. By flooding a cell with the PEBBLE nanosensors and then monitoring the cell under a microscope, investigators can observe the distribution of oxygen, nitric oxide, or other target molecules throughout the cell and monitor its evolution in time. With a little cleverness combined with a thorough knowledge of physics, chemistry, and cellular biology, it is possible to measure a great number of cellular metabolic activities, including changes in temperature, viscosity, and local magnetic and electric fields. “The beauty of this,” Philbert said, “is that we can now begin to add richness to the data sets in local areas in the cell where there may be intracellular electrophoresis contributing to the association of proteins, many of which have very large electrical dipoles.”
Another area of research pursued by the Philbert laboratory is the incorporation of Photofrin (porfimer sodium), a light-activated cancer drug, into nanostructures. Normally, Photofrin cannot cross the blood-brain barrier and thus cannot be used to treat brain cancer by conventional means. Because of the size of nanostructures, however, when the drug is incorporated into them it can cross the blood-brain barrier and target the tumor cells. A laser device is then used to activate the Photofrin and destroy the tumor cells. In experiments with rodent models, the treatment has dramatically increased survival rates. Without treatment, the brain tumors kill 100 percent of mice within 10 to 12 days. With treatment, about 60 percent of the mice are still alive after 39 days and about 40 percent are alive after 3 months.
Nanostructures are not expensive to make, costing about a dollar a kilogram for the materials incorporated into the most common ones. Synthesis demands a great deal of expertise, however, and the high cost is acquired through the requisite toxicology screening and federal approval processes. “Still,” stated Philbert, “their potential to do so many things that cannot be done in any other way makes them well worth the investment.”










