3
Subglacial Environments: Biological Features
INTRODUCTION
To develop a research and management plan for subglacial aquatic environments, one of the principal issues that needs to be resolved is whether these systems contain viable organisms. In the case of microbes, viable means only that they are capable of growth, and it is possible that subglacial environments contain viable but not actively growing microbes. If there are actively growing microbes in subglacial aquatic environments, this will dictate that a specific course of action be taken during the exploration and studies of such environments to ensure that these ecosystems are not profoundly changed or irreversibly altered. While no subglacial aquatic environments have been sampled directly to date, a general understanding of some of the key physiochemical characteristics of these environments has developed (Siegert et al. 2003; Christner et al. 2006). The key questions that need to be addressed are the following: (1) Can these environments support cell maintenance and growth (Box 3.1) and thereby be considered active systems? (2) What are the likely source populations? (3) Have the populations evolved to be significantly different from their source.
REQUIREMENTS FOR LIFE
The minimum requirements for proliferation of microorganisms are an inoculum of viable cells; available water; electron donors and acceptors for biological energy supply (Box 3.1); a source of carbon, nitrogen, phosphorus, sulfur, iron, and other elemental constituents of biomolecules; sufficient time for reproduction; and the absence of severe physical, chemical, or biological conditions that would destroy cells or preclude growth. Severe conditions may include extremes in temperature, pH, pressure, water activity, salinity, and the presence of ionizing radiation. It should be noted, however, that to date, in terms of the extent of microbial life, no geographic limitations are known. Evidence of microbial life has been found in the most extreme environments. These environments include submarine hydrothermal vents (Baumgartner 2002; Kashefi and Lovley 2003), terrestrial hot springs (Brock 1997;
|
BOX 3.1 Biological Energy Supply All life uses adenosine triphosphate (ATP) for storing energy and driving cellular processes that require energy (such as flagellar rotation or protein synthesis). Therefore sufficient free energy must be biologically available (e.g., in the form of redox couples) and exceed a finite minimum in order to drive the formation of ATP from adenosine diphosphate (ADP). The presence of free energy in an ecosystem that equals or exceeds this finite minimum is necessary to sustain life and conserve ecosystem functions (Hoehler 2004; von Stockar et al. 2005). From a thermodynamic standpoint it is not possible to couple with complete efficiency all energy released during metabolism to ATP synthesis. The thermodynamic efficiency of the ecosystem must therefore account for the proportion of free energy of metabolism that can be captured and stored along with that proportion lost as heat (Hoehler 2004; von Stockar et al. 2005). Thus, the minimum free energy that must be available for biological processes in any given environment is the combination of the ATP synthesis energy, stoichiometry of ion release during ATP synthesis, and thermodynamic efficiency (Schink and Stams 2002). This quantity can be referred to as the biological energy quantum (BEQ) and provides a predictive measure of the favorability of biological activity analogous to the use of Gibbs free energy change to determine the likelihood of a chemical reaction proceeding (Hoehler 2004). Coupled to the presence of an energy quantum that must be generated in a chemical reaction inside a biological system is the concept of maintenance energy (ME), which suggests that organisms require a certain minimum rate of energy intake to maintain molecular and cellular order and function (Hoehler 2004). Three levels of increasing ME requirement can be described (Morita 1997): (1) cell survival, (2) cell maintenance, and (3) active cell growth. Both BEQ and ME can be affected by a variety of environmental factors such as temperature (BEQ and ME requirements should decrease with temperature). BEQ and ME requirements thus define the minimum substrate concentration and substrate generation rate that must be sustained by a given environment in order to support life (Hoehler 2004). For some time the figure of −20 kJ mol−1 was described in the literature as the minimum quantum of free energy that could be converted by a biological system (Schink 1997). More recently, a free energy value as low as −4.5 kJ mol−1 was reported (Jackson and McInerney 2002) to support growth of a co-culture of butyrate-degrading organisms with methanogens in which one species lives off the products of another. Although the ability to measure thermodynamic minima such as BEQ and ME is still very much a work in progress, this result implies that microbial communities could be supported even in extreme subglacial aquatic environments despite ultra-oligotrophic substrate concentrations and the absence of sunlight. |
Rothschild and Mancinelli 2001), brine channels in sea ice (Thomas and Dieckmann 2002), glaciers and ice shelves (Christner et al. 2003; Mueller et al. 2005), the subterranean deep biosphere (Pedersen 1993), cold deep-sea environments (Yayanos 1995; Bartlett 2002), deserts (Drees et al. 2006), and pH and salinity extremes (Vreeland et al. 2000; Fütterer 2004).
Subglacial aquatic systems are extreme environments for microbial life with low temperatures, elevated pressures, and no direct contact with the atmosphere or sunlight. Indirect evidence suggests that they may also have a high gas content (including oxygen) and low inorganic and organic nutrient concentrations. Most temperate latitude species are unlikely to proliferate and are also unlikely to compete effectively with species
that have been selected for over long periods of time by this unusual combination of properties. However, Lake Vostok and other subglacial aquatic systems are liquid-water environments with a number of features that are conducive to microbial growth, and the growth of a small subset of contaminating cells cannot be precluded. Growth may be especially favored for those microbes that are introduced from viable populations in glacial ice higher in the borehole (Chapter 3) and within drilling fluids that contain both organic substrates and living cells (Chapter 4).
Persistent liquid water, stable conditions, and extended time for reproduction are features of subglacial aquatic environments that make them suitable habitats for microbial growth. In this respect they contrast with many polar environments that are subject to intense freeze-thaw cycles (e.g., shallow lakes and ponds, ice shelf ecosystems), physical disruption (e.g., sea ice), and extreme aridity (e.g., polar desert soils, rock surfaces).
POTENTIAL IMPEDIMENTS TO LIFE IN SUBGLACIAL AQUATIC ENVIRONMENTS
Adverse conditions in the subglacial environment that could inhibit or preclude growth include low temperature, elevated pressure, and any reactive oxygen species favored by high oxygen tensions. Despite their extremely cold environments, many microbial species in the polar environment are psychrotolerant (able to grow at low temperatures) rather than psychrophilic (optimal growth at low temperatures). All of these microbes grow slowly under extremely cold conditions, yet they can sometimes achieve large standing stocks because losses are low (Vincent 2000). These includes well-developed benthic microbial mats in polar lakes with ultra-oligotrophic water columns (Bonilla et al. 2005) and microbial communities in the anoxic sediments of Antarctic saline lakes (Mancuso et al. 1990). It is also well known that microbial communities grow in polar oceans at temperatures down to −1.8°C, and can be sustained at much lower temperatures (−20°C) in cold saline environments such as the brine channels in sea ice (Junge et al. 2004).
The deep ocean habitat is similar to that of subglacial aquatic environments, being both high pressure and cold. Despite these two adverse conditions, microbes can live as deep as 10,000 m in the ocean (Parkes et al. 1994; Kormas et al. 2003), well above the pressures found in subglacial aquatic environments where they are less than or equal to 400 atmospheres, equivalent to 4000 m in the ocean. Supersaturated oxygen habitats are also known to harbor active microbial communities, including the perennially ice-covered lakes of the McMurdo Dry Valleys (Wharton et al. 1986). The upper waters of Lake Vostok, however, are predicted to contain extreme oxygen levels more than an order of magnitude higher. McKay et al. (2003) calculated that in situ gas concentrations in Lake Vostok would be up to 2.5 L per liter of lake water (at standard temperature and pressure), equivalent to 750 mg O2 L−1. Even higher values (up to 1300 mg O2 L−1) have been estimated by Lipenkov and Istomin (2001).
These oxygen tensions could be a severe constraint on growth if accompanied by the formation of reactive oxygen species (see below), particularly for microbial contaminants that would largely comprise taxa that are adapted to less severe conditions. Resistant cells and endospores attached to particles might find more favorable oxygen conditions for proliferation as they sink deeper in the water column.
Information about the chemistry and microbiology of Lake Vostok is limited to exhaustive analysis of the accreted ice. As described earlier (Chapter 2), the accreted ice makes up the bottom 84 m of the ice core retrieved above Lake Vostok. Jouzel et al. (1999) observed that this ice extended from 3539 m below the surface of the Antarctic ice sheet to its bottom at about 3750 m and differed from the overlying glacial ice in isotopic properties, total gas content, crystal size, and electrical conductivity. They concluded that this section of the Vostok core consists of ice refrozen from Lake Vostok water.
Although the chemical composition of subglacial aquatic environments remains highly speculative, the analysis of glacial and accretion ice from Vostok implies that the elemental requirements for microbial growth could be satisfied in the lake and that there are many possible electron donors and acceptors for biological production by chemotrophic and heterotrophic organisms (Table 3.1).
However, the various measurements of these concentrations in Lake Vostok ice differ by orders of magnitude (De Angelis et al. 2004). This may reflect differences in protocols between laboratories or real differences associated with sampling a heterogeneous ice core (Price 2007). One suggestion that accounts for some of the heterogeneity (Bell et al. 2002) involves the movement of the ice across the lake. In this scenario, one level of the accretion ice in the ice core formed when that site in the ice sheet was above a shallow embayment; another level of accretion ice formed later when the site was above a deep pelagic part of the lake.
Additional substrates could enter the lake environment by glacial scour of the surrounding bedrock and the transfer of dissolved and particulate minerals by advection
TABLE 3.1 Possible Electron Donors and Acceptors in Subglacial Aquatic Environments in the Presence (Aerobic) or Absence (Anaerobic) of Oxygen
|
Conditions |
Electron Donor |
Electron Acceptor |
Metabolic Process |
|
Aerobic |
H2 |
O2 |
H oxidation |
|
|
HS-, S0, S2O32-, S4O62- |
O2, NO3- |
S oxidation |
|
|
Fe2+ |
O2 |
Fe oxidation (low pH) |
|
|
Mn2+ |
O2 |
Mn oxidation |
|
|
NH4+, NO2- |
O2 |
Nitrification |
|
|
CH4 and other C-1s |
O2 |
(C-1) oxidation |
|
|
CH4 |
O2 |
Methane oxidation |
|
|
Organic compounds |
O2 |
Heterotrophic metabolism |
|
Anaerobic |
H2 |
NO3- |
H oxidation |
|
|
H2 |
S0, SO42- |
S0 and sulfate reduction |
|
|
H2 |
CO2 |
Acetogenesis |
|
|
H2 |
CO2 |
Methanogenesis |
|
|
S0, SO42- |
NO3- |
Denitrification (inorganic) |
|
|
Organic compounds |
NO3- |
Denitrification (organic) |
|
|
Organic compounds |
S0, SO42- |
S0 and sulfate reduction |
|
|
Organic compounds |
|
Fermentation |
|
SOURCE: Modified from Siegert et al. (2003). |
|||
directly or after melting of the bottom ice. Such inputs could include mineral particles of the type seen in the upper accretion ice of Lake Vostok that are thought to be derived from the shore of the lake. Microbial growth on substrates in glacial rock flour, such as metal sulfides, has been documented in suboxic and anoxic glacial bed environments (Christner et al. 2006); for example, microbial production has been suggested for an ecosystem contained within the basal ice of the Greenland Ice Sheet. Here, the lower 13 m of the GISP2 (Greenland Ice Sheet Project) ice core (at a depth of 3053 m) contains high concentrations of bacteria (>108 cells cm−3) attached to abundant clay particles; Tung et al. (2006) suggest that these bacteria thrive by reduction of Fe(III) within the clay, in the process oxidizing organic molecules such as phenol and humic acids (both detected in the ice) to CO2. The chemical properties of this interface, however, reflect the freeze-over of an anaerobic wetland and are likely to differ greatly from subglacial environments in Antarctica.
In addition, some subglacial aquatic environments (e.g., Lake Vostok) may have formed as aerial lakes at a time prior to the cooling of Antarctica. Formation of lakes in this manner would provide early sources of sediments that could harbor carbon and nutrients that subsequently could continue to be recycled within the resulting subaerial system. Other characteristics of subglacial aquatic environments that may contribute potential sources of carbon and energy to support microbial life include tectonic activity, the presence of hydrothermal vents, gas hydrate deposits, and cold seeps containing inorganic solutes.
Microbial communities growing in the interface between ice and rock could also provide a source of biota and organic carbon into the lake. The detection of thermophiles (Table 3.2) in a clone library analysis of samples of Vostok accretion ice (Bulat et al. 2004) has led to speculation about the possibility of hydrothermal vents and an ecosystem based on chemolithotrophy by heat-loving microbes. Bulat et al. (2006) have recently suggested that fault vents and microbial ecosystems are likely to be in a shallow side basin or outside the lake, with horizontal transfer of microbiota to the 5G borehole site by the overlake ice flow. It is likely that such bacteria may not survive in the frigid waters of Vostok, but perhaps could provide an allochthonous (external) source of organic carbon to the lake ecosystem.
Glacial meltwaters at latitude 80 degrees south contain relatively high concentrations of dissolved organic nitrogen and nitrate (Vincent and Howard-Williams 1994), suggesting long-range and upper atmosphere input mechanisms that could deliver reduced and oxidized nitrogen compounds into subglacial aquatic environments via their overlying ice. Nitrification has been identified as one potential mechanism for energy supply in subglacial systems (Table 3.1), with potential ammonium levels of 3 μg L−1 in meltwaters from the glacial ice (Siegert et al. 2003). The nitrification intermediate N2O has been detected in the Vostok ice column, and experiments to date with the psychrophilic nitrifier Nitrosomonas cryotolerans show that nitrification could occur down to at least minus 12°C (Sowers et al. 2006).
Measured dissolved organic carbon (DOC) concentrations in the accretion ice of Lake Vostok vary greatly, from undetectable to around 80 μmol L−1. In part this may reflect vertical heterogeneity in the ice column but it may also result from major differences in DOC analytical methodologies among laboratories. Further uncertainty surrounds the appropriate partition coefficient for the distribution of solutes between ice and lake water during freeze-up (Siegert et al. 2003). It is also known that this partitioning acts differentially, with higher-molecular-weight organic compounds excluded
TABLE 3.2 Taxonomic Identification of Clone LV3607bR-40A Recovered from Accreted Ice Sample at a Depth of 3607 m
|
Closest Neighbors and Their Features |
||||||
|
GenBanka (similarity, %) |
Acc. No. |
Representative |
Sampling Size |
Temperature Profile (°C) |
Source |
References |
|
100 |
AB009828 |
Hydrogenophilus thermoluteolus TH-1 |
Hot springs, Izu district, Japan |
52b |
Crustal fluid, adjacent soil |
Gogo et al. 1977 Hayashi et al. 1999 |
|
99.5 |
AJ131694 |
Hydrogenophilus hirschii yel5a |
Angel Terrace Spring, Yellowstone National Park, USA |
50-67 |
Crustal fluid |
Stohr et al. 2001 |
|
99.1 |
AB009829 |
Hydrogenophilus thermoluteolus TH-4 |
Hot springs, Izu district, Japan |
50b |
Crustal fluid, adjacent soil |
Gogo et al. 1977 Hayashi et al. 1999 |
|
96.3 |
AF026979 |
Unidentified beta-proteobacterium OPB30 (rDNA study only) |
Obsidian Pool, Yellowstone National Park, USA |
75-95 |
Sediments |
Hugenholtz et al. 1998 |
|
aOnly long, published 16S rDNA sequences are included in the analysis. bTemperature profile for culturing. SOURCE: Copied from Bulat et al. 2004. |
||||||
to a greater extent than smaller organic molecules (Belzile et al. 2002). On the basis of ice-water partitioning measured in Antarctic Lake Bonney, Christner et al. (2006) calculated that Lake Vostok may contain 17-250 μmol L−1 organic carbon, which may be sufficient to sustain microbial populations if continually replenished.
Oxygen Effects
The surface waters of Lake Vostok are inferred to be supersaturated in oxygen given that ice melting into the lake contains air clathrate hydrates while the accretion ice formed during freeze-up of the lake water is gas-free. McKay et al. (2003) estimated that for a water residence time of 13,300 years, this pumping mechanism would result in oxygen tensions about 50 times higher than those found in surface lake waters in equilibrium with the atmosphere. These hyperbaric concentrations would likely have toxic and potentially lethal effects on many species of microbes if they were associated
with the formation of reactive oxygen species (ROS), but such effects have yet to be evaluated.
Molecular oxygen itself is relatively unreactive, and biomolecules such as proteins and nucleic acids are little affected by high oxygen tensions. However in the presence of ionizing radiation or after reaction with certain compounds, molecular oxygen is converted to ROS. In the cellular environment, oxygen reacts with reduced flavins (a constituent of redox enzymes) to produce superoxide and hydrogen peroxide. The latter can react with free iron to generate hydroxyl radicals. All of these products are stronger oxidants than oxygen itself and can have wide-ranging damaging effects on nucleic acids, proteins, and membrane constituents (Imlay 2002). These oxidative stress effects have been widely studied, including in laboratory or environmental systems during complete darkness (e.g., Bechtold et al. 2004).
Microorganisms have several lines of defense that can protect them from the damaging effects of ROS production (Sies 1997). These include the production of non-enzymatic antioxidants, such as tocopherol and carotenoids, and powerful enzymatic systems that react with and detoxify oxidants, in particular superoxide dismutases, catalases, and glutathione peroxidases. The effects of high oxygen tensions and ROS production on microbial metabolism and growth are therefore highly variable among species. For example, a 90-minute exposure of hyperbaric hyperoxia (100 percent O2 at 2 atmospheres) inhibited growth of Staphylococcus aureus by 3-25 percent, but had no effect on the growth of Escherichia coli (Tsuneyoshi et al. 2001). It has also long been known from medical experience that even obligate anaerobes vary greatly in their oxygen sensitivity (Tally et al. 1975).
It should also be noted that subglacial aquatic environments may include those in which anoxic conditions or low oxygen tensions prevail, and where even O2-sensitive microbes may survive and grow. Aquatic environments that are flushed by hydraulic through-flow may not accumulate high oxygen concentrations, depending on the source waters. Aquatic environments may be stratified as in the perennially ice-covered lakes of the McMurdo Dry Valleys. These have supersaturated oxygen concentrations in their upper water columns, but also contain anoxic bottom waters and sediments. Particles in suspension in the water column may also provide interior microhabitats in which oxygen concentrations are much lower than in the surrounding water.
SOURCE POPULATIONS
Bacterial Transport
Biogeography is the study of the distribution of biodiversity over space and time and the mechanisms that generate and sustain this diversity, which include speciation, extinction, dispersion, and species interactions (Martiny et al. 2006). As described by Fenchel (2003), many macroscopic animal and plant species exhibit biogeographic patterns resulting from their limited distribution ranges on Earth due to requirements for specific habitats and climates as well as the influence of historical events (e.g., continent separations). These occurrences have combined to produce taxonomic groups that have evolved over the course of geologic time within continents or geographically isolated regions such as lakes, mountain ranges, or oceanic islands. These barriers to migration have resulted in the presence of groups restricted to specific locations (Fenchel 2003).
The existence of biogeographic patterns in microbes on the other hand is possible but not easy to identify (reviewed in Bell et al. 2005; Martiny et al. 2006). This uncertainty is due to a lack of information in many critical areas. For example, universal concepts that define a bacterial species are lacking (Box 3.2) and only now are methods being developed that are sensitive enough to resolve differences between microbes. However, some concepts of biogeography from studies in macroecology may be applicable to microbial ecological investigations, although at least some differences in biogeographical concepts are likely to exist between the two. For instance, Fenchel (2003) notes that microbial population sizes are enormous by comparison with macroorganisms and argues that microbial distribution can best be understood in terms of habitat characteristics. Furthermore, the probability of microbial dispersal is higher and the probability of local extinction is much lower in microbes than in macroorganisms. The effect and relative influence of historical contingencies on the existence of microbial biogeographic patterns also continue to be discussed (Martiny et al. 2006).
Many potential mechanisms exist for bacterial dispersion. For example, it has been calculated that about 1018 viable microbes annually are transported through the atmosphere between continents (Griffin et al. 2002). Other evidence concerning particle transport by air currents comes from the record of volcanic ash described by Basile et al. (2001). These authors report, “There is a record of 15 ash layers in the past 500,000 years in the Vostok ice core. Nine layers originate from activity of the South Sandwich volcanic arc, three from southern South America, one from the Antarctic Peninsula (Bransfield Strait), and one from West Antarctica (Marie Byrd Land province). In spite of the low frequency of occurrence of visible tephra layers in the Vostok core (one event every 20 kyr [thousand years]), the overall atmospheric pathway of these ash events appears consistent with the almost continuous advection of continental dust from South America.”
Movement by global ocean currents is another mechanism by which microbes can be transported long distances. Additional mechanisms include transport by birds and by the increasing presence and activity of humans in Antarctica. Of particular importance to the study of subglacial aquatic environments is their potential connectivity, which may allow the movement of microbes beneath the ice sheet. The surface of the Antarctic ice sheet acts as vast collector for microbiota deposited from the atmosphere (Vincent 1988), and a subset of this microflora may retain viability and even metabolize within the snow and glacial ice (Price and Sowers 2004; Price 2007). These communities of viable cells and spores may ultimately reach the subglacial aquatic environments to provide a continuous inoculum at the melting glacier ice-lake water interface.
Microscopic and culture analyses of Vostok glacial ice from the 1500-2750 m stratum (>240,000 years old) have shown the presence of a wide variety of microorganisms: bacteria (including actinomycetes), yeasts, fungi, and microalgae (Abyzov 1993; Abyzov et al. 1998), although questions remain concerning the presence of contaminants introduced during sampling. However, Abyzov (1993) points out that in the deepest ice he examined (2405 m) only spore-forming bacteria remained. As shown in Figure 3.1, the numbers of small cells containing DNA is higher in the glacial ice than in the accreted ice. Abyzov (1993, p. 284) had the following to say about the microbes he isolated from the Vostok ice core down to a depth of 651 m: “Psychrotolerant and psychrophilic microorganisms could be brought into the central parts of the Antarctic continent with the air flows from sub-Antarctic islands, littoral zones of the continent, or its ice-free area, but it cannot be excluded that mesophilic strains isolated from the
|
BOX 3.2 Bacterial Speciation Processes The definition of a bacterial species remains an elusive concept in microbiology in large part due to the fact that bacteria are clonal and reproduce asexually. As a result, the Biological Species Concept first espoused by Ernst Mayr (1942), which broadly states that groups of actual or potentially interbreeding natural populations can reproduce only among themselves to the exclusion of all others, is not applicable. At present, somewhat arbitrary operational definitions based on phenotypic characterizations (Bochner 1989; Mauchline and Keevil 1991) and molecular biological tools such as the association of genomic DNA in standardized DNA/ DNA hybridizations (typically >70 percent) and/or gene sequence identity of ≥97 percent for the16S ribosomal RNA (rRNA), are most commonly used to define bacterial species (Wayne et al. 1987; Stackebrandt and Goebel 1994; Konstantinidis and Tiedje 2005). The ever-increasing amount of information derived from comparative genomic analyses is providing fundamentally new insights into the question of what constitutes a bacterial species while further revealing that bacterial genomes are not static entities; many mechanisms or combinations thereof can contribute to both maintenance and change of genome structure and content over time (Ward and Fraser 2005). Among the most important forces contributing to the evolution of bacterial species are the interplay between the rates of mutation and homologous recombination (where homologous in this case denotes a substitution of some portion of a genomic sequence with a sequence of high similarity ) and the influence of biogeography in the divergence of bacterial lineages through adaptations and separation into ecological niches (Nesbø et al. 2006). Genetic variations are the source upon which evolutionary forces such as genetic drift and natural selection act. Some mutations in DNA are spontaneous, random events that can result from several possibilities. For instance, mutations can arise by the exposure of DNA to ionizing radiation, ultraviolet radiation, and some chemicals. Normally DNA sequences are copied precisely during replication; however, errors in DNA replication can result in changes in gene sequence. In addition, a variety of recombinatory processes can occur in nature that contribute to genetic variability by joining DNA from different biological sources (Graur and Li 2000). Basic concepts of population genetics suggest that most mutations are deleterious to the fitness of an individual (ability of the individual to survive and reproduce) and are selected against causing their removal from the population, a process referred to as negative or purifying selection. Mutations can also result in alleles (alternative forms of the gene) of similar fitness that are considered neutral mutations and are subject to loss from or fixation within a population as a result of stochastic forces. Occasionally, a mutation may produce an allele that increases the fitness of an individual. These advantageous mutations will be subjected to positive selection and will tend to be fixed more rapidly in a population than to neutral mutations (Graur and Li 2000). |
ice sheet originated from the islands and continents of the temperate latitudes of the Southern Hemisphere.”
EVOLUTION OF LIFE IN SUBGLACIAL AQUATIC ENVIRONMENTS
Antarctic subglacial aquatic environments represent potentially rich and largely unexplored storehouses of genetic information. As such, a variety of biogeochemical processes could potentially be at work, resulting in unique metabolically active microbial assemblages in terms of structure and function. The age of the oldest ice in the ice sheet is 1 million years (EPICA 2004; Peplow 2006), and this has been taken to indicate the length of time of microbial isolation of Lake Vostok. Although 1 million years is not a considerable amount in terms of prokaryotic evolution, some changes
|
Homologous recombination is recognized as a vital process to maintain chromosome integrity by facilitating the repair of damaged DNA that is mediated largely through pathways controlled by the enzyme RecA (Ivanic-Bacce et al. 2006). However, homologous recombination is also increasingly being recognized as central to the creation of genetic variability and as an important force in genome change within and between prokaryotic lineages (Fraser et al. 2007). Within prokaryotic lineages, sources of change can result from rearrangement, duplication, or loss of chromosomal segments during DNA replication. Further recombination facilitates the exchange of DNA between lineages from a donor to a recipient, a process referred to as horizontal or lateral gene transfer (Ochman et al. 2000). Among the best-identified mechanisms of lateral gene transfer are those of transformation, conjugation, and transduction. Conjugation and transduction require an intermediary to genetic exchange, a plasmid in the case of conjugation and a bacteriophage or bacterial virus in the case of transduction. Alternatively, transformation, the third mechanism, involves the uptake of free DNA from the environment (Graur and Li 2000). Recombination is not restricted to the transfer of homologous DNA from a donor to a recipient in bacteria. One possible result of recombinatorial events is the integration of heterologous DNA from the donor to the recipient along with flanking homologous DNA. Therefore, under some circumstances DNA from distantly related lineages can also be transferred. Additional mechanisms that can facilitate this type of integration include transposable elements, such as insertion sequences, transposons, and retroelements (Bennet 2004). Site-specific recombination systems such as integrons (reviewed in Mazel 2006) have also been recognized as important mobile elements in acquiring exogenous genes and shaping genome change. In mammalian reproduction, recombination accompanies each reproductive event. The rate of recombination in bacteria does not approach this frequency (Levin and Bergstrom 2000). However, it is possible that bacterial recombination can exceed mutation as a source of genetic variability, raising the possibility that at least a modified form of the Biological Species Concept may be applicable to some bacteria under certain circumstances. For example, related bacterial lineages may share a core set of genes via recombination in a common gene pool (Doolittle and Papke 2006). At present there are a number of competing ideas concerning how mechanisms that foster genetic variability and the forces of evolution interact to both maintain sufficient genetic similarity (effectively preserving species) and to create genetic divergence between species. Currently an intense debate reigns as to the relative importance of barriers to genetic exchange by homologous recombination, which is broadly captured by two models: “the bacterial species model” (Doolittle and Papke 2006; Nesbø et. al 2006) versus natural selection of adaptive mutants that use similar resources and inhabit similar physical locations (“the ecotype model”) (Cohan 2002). As the search proceeds, to unlock the underlying principles that define a bacterial species will require the continued integration of results from diverse areas of biology. The exploration of subglacial aquatic environments will provide a unique lens through which to view and study these questions. |
may have occurred to help these organisms adapt to the cold, dark, oligotrophic environment (Tiedje 1989).
The waters of Lake Vostok have been isolated from direct atmospheric contact for at least 15 million years (Christner et al. 2006), giving a much longer time for evolutionary processes to operate (Box 3.3), and the founding populations in the lake may even be older if the original microbiota were derived from Antarctic bedrock or sediments. Then they could have been isolated from the surface microbial populations prior to the formation of Lake Vostok, as much as 35 million to 40 million years ago (Tiedje 1999).
As discussed by Tiedje (1999), studies on the rate of genetic change of microbes in nature indicate that speciation could take 10 million to 100 million years. An often cited estimate of speciation is the divergence of Escherichia coli and Salmonella enterica
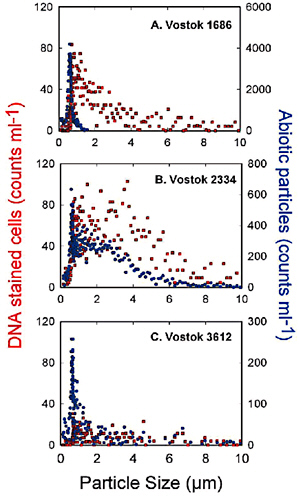
FIGURE 3.1 Flow cytometer data showing the relative abundance and size distribution of cellular and abiotic particles in Lake Vostok glacial ice (A = 1686 m; B = 2334 m) and accretion ice (C = 3612 m). Cellular particles were characterized by their fluorescence induced with the DNA stain CYTO 60. The presence of attached bacteria is also supported by the flow cytometer data in this figure. Note the rather scattered and large size of DNA containing particles. Using epifluorescence microscopy, these “large” cells are actually multiple cells aggregated on abiotic particles. SOURCE: From Priscu et al. (in press).
lineages from one another, which is estimated to have occurred approximately 100 million years ago (Lawrence and Ochman 1998). A division rate of 10 generations per year would give rise to a change of only one base pair per gene in 1 million years for a gene with 1000 base pairs (Tiedje 1999). If the microbes have been isolated from the surface microbes in bedrock or sediments for 35 million to 40 million years, significant genetic changes may have taken place. Tiedje (1999) points out that isolation of microbes in bedrock or sediments for millions of years also occurs elsewhere on Earth.
While speciation is not a well understood process (Box 3.2), lower rates of metabolic activity and growth may lengthen the time between bacterial generations, which will further increase the amount of time needed for significant genetic changes in genomes to take place. Further, connectivity of subglacial aquatic environments (Chapter 2) will serve to lessen genetic isolation and barriers to gene flow, although it may serve to increase the effective population for recombination (Fraser et al. 2007). Given
|
BOX 3.3 Natural Selection and Genetic Drift Evolution represents the process of change in the genetic content of populations. As such, the common denominator of evolution is the change in gene frequencies with time. Changes in allele (alternative forms of a gene) frequency can occur as a result of natural selection or genetic drift. Natural selection is the process by which organisms possessing the most favorable genetic adaptations outcompete other organisms in a population, resulting in their tendency to displace the less-adapted ones. Natural selection is dependent on the existence of variations of the same trait that confer different survival advantages or disadvantages (Graur and Li 2000). In contrast, genetic drift is a stochastic process by which changes in gene frequencies result from the accumulation of small, random, neutral mutations over time (Kimura 1983, 1989). The forces of genetic drift and natural selection rarely act in isolation, and their relative strengths depend on several factors including effective population size. In large populations, genetic drift will occur slowly. Therefore even weak selection of an allele can have significant effects on its frequency; beneficial alleles will increase (positive selection), while harmful ones will be eliminated (negative or purifying selection) (Graur and Li 2000). However, in species with a small effective population size, genetic drift will predominate. In small effective populations, deleterious alleles can increase in frequency leading to fixation in a population purely as a result of chance since the relative importance of genetic drift is greater. Species with a small effective population size are also subject to a greater probability of extinction because they are more vulnerable to genetic drift, leading to stochastic variation in their gene pool, demography, and environment. Any allele, regardless of its effect on fitness (deleterious, beneficial, or neutral), is more likely to be lost from a small population or gene pool than from a large one, resulting in a reduction in the number of variants of a given allele (Graur and Li 2000). |
at most, millions of years during which evolutionary forces will have been at work, the metabolically active and growing microorganisms present in these communities should represent those better adapted to the relevant conditions but will still be related to microorganisms found in other environments today.
Is it possible to detect the end product of evolutionary change in subglacial environments? Many microbes may be present as viable cells that remain dormant until they encounter conditions that allow their metabolism and growth, as described by the “rare biosphere” concept (Pedrós-Alió 2007). These rare components would be undetectable with standard clone library analysis. However, emerging new technologies (Church 2006; Sogin et al. 2006) may increase the number of rare genotypes by several orders of magnitude or more.
CURRENT EVIDENCE FOR LIFE IN SUBGLACIAL AQUATIC ENVIRONMENTS
Biogeochemical Evidence for an Active Bacterial Flora in Lake Vostok
The total dissolved solids (TDS) in the lake water can be estimated by applying ice-water partition coefficients to the major ion concentrations in the accretion ice (Christner et al. 2006). These ice-water partition coefficients are assumed to be the same
as those calculated from the permanent ice cover and water column of Lake Bonney in the McMurdo Dry Valleys. In this way, Christner et al. (2006) estimated the TDS concentrations of water in the embayment and the main lake to be 34 and 1.5 mmol L−1, respectively. The former value is well above typical concentrations in temperate lakes. In the same way, the concentrations of non-purgeable organic carbon (NPOC), amino acids, and major ions can be estimated. In the 5G core the concentration of NPOC in the glacial ice ranged from 2 to 80 μmol L−1. In Lake Vostok, Christner et al. (2006) predicted that the NPOC concentrations would range from 17 to 250 μmol L−1. Total amino acids (dissolved free and combined amino acids plus protein) in glacial ice constituted 0.6-2 percent of the NPOC, while they made up 0.01 to 2 percent of NPOC in the accretion ice. Although there was a significant correlation between the NPOC and amino acid concentration in the accretion ice (r = 0.83), no significant relationship existed between these parameters in the glacial ice (r = 0.015).
From these data plus information on the acid-extractable amino acid concentrations, Christner et al. (2006) concluded the following: relative to the main body of the lake, the waters near the shoreline represent a source of amino acids. This, together with the cell density data, implies that this region has elevated rates of heterotrophic biological activity and associated amino acid transformations.
Biological Evidence for an Active Bacterial Flora in Lake Vostok
Analysis of samples from the accretion ice of Lake Vostok by epifluorescence microscopy (Figure 3.2) and scanning electron microscopy has shown the presence of intact, DNA-containing cells; however there are conflicting reports on their concentrations. Christner et al. (2006) reported that the number of bacterial cells counted in accretion ice depths between 3539 and 3572 m ranged from 98 ± 18 to 430 ± 23 cells mL−1. These data overlap with the range 200 to 3000 cells mL−1 reported for samples between 3541 and 3611 m (Karl et al. 1999; Priscu et al. 1999; Abyzov et al. 2001). Price (2007) notes that other reports indicate concentrations less than 10 cells mL−1, but suggests that these disparities may simply reflect sampling error if the microbes are located in veins between the large ice crystals. The bacterial abundance is two- to seven-fold higher in accretion ice than in the overlying glacial ice, implying that Lake Vostok is a source of bacterial carbon beneath the ice sheet. Scanning electron microscopy (Priscu et al. 1999) has shown that microbial cells are often associated with organic and inorganic particles.
When a sample of the accreted ice was melted, Karl et al. (1999) were able to measure 14CO2 produced from added 14C-glucose over incubations of 2, 11, and 18 days. They were careful to state, however, that the laboratory results were of potential activity only and in situ rates might be much lower.
Biological Evidence for Viability of the Bacterial Flora in Lake Vostok
Christner et al. (2006) report that differential cell staining revealed that the majority (75 to 99 percent) of cells in the accretion ice are potentially viable, which is comparable to the ~80 percent value reported by Miteva et al. (2004) from a single depth of “silty” basal ice in the Greenland GISP2 core. Positive rates of glucose respiration in four of the six samples tested by Christner et al. (2006) corroborated the presence of viable heterotrophic cells. When the respiration data were normal-
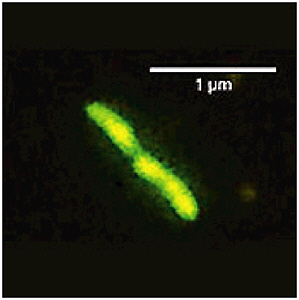
FIGURE 3.2 Bacterial sample from the Lake Vostok accretion ice analyzed by epifluorescence microscopy. SOURCE: Karl et al. (1999).
ized to the lake temperature of −3°C, their measurements of glucose mineralization (0.005 ± 0.003 to 0.2 ± 0.005 nmol C L−1 day−1) were on the low end of the range reported by Karl et al. (1999) for an accretion ice core from 3603 m (0.3 ± 0.1 to 0.5 ± 0.4 nmol C L−1 day−1 at −3°C). However, Christner et al. (2006) found that in samples where respiration rates could be measured there were no significant levels of incorporation of substrate into macromolecules. On these same samples, Christner et al. (2006 p. 25) reported: “Our enrichment culturing studies on these samples have required up to 6 months of incubation in the dark at 4°C before growth is initiated (Arnold et al. 2005; Christner and Priscu unpublished). After this apparent period of resuscitation, the microbial (bacteria and yeast) species recovered and grew to visible densities on both liquid and agar-solidified media in <10 d, and many of the isolates were capable of growth to the stationary phase in 40 h at 22°C. When non-growing and sub-lethally injured cells are placed in a growth situation (e.g., Dodd et al. 1997; Aldsworth et al. 1999), metabolism must first be initiated to repair incurred cellular damage before growth and reproduction can occur. Thus, if our incubations had been conducted over a longer time frame, we would expect such carbon-fed cells to eventually initiate the growth phase.”
Evidence for an Autochthonous or Lake-Dwelling Microbial Flora
Microbiological and molecular-based studies of the accretion ice by six laboratories indicate that this ice contains low but detectable amounts of prokaryotic cells and DNA (Karl et al. 1999; Priscu et al. 1999; Abyzov et al. 2001; Christner et al. 2001; Bulat et al. 2004). Christner et al. (2006) state that molecular identification of microbes in Vostok accretion ice by culturing and small subunit rRNA gene amplification, show close agreement with present-day microbiota. These were identified to lie within the bacterial phyla Proteobacteria (α, β, and γ), Firmicutes, Actinobacteria, and Bacteroidetes (Table 3.3). Further, they state (Christner et al. 2006; p. 2496): “Con-
TABLE 3.3 Microbial Diversity, Number of Cells, and Number of Inclusions Observed in Different Horizons of the Vostok 5G Deep Core Accretion Ice
|
Ice Depth (m below surface) |
Region of the Accretion Ice Sampled |
Inclusion Number (m−1) (mean) |
Investigator |
Type of Microorganisms Detected |
Total Numbers of Microorganisms Detected (cells /ml) |
|
3541 |
Shallow embayment / grounding line (3540-3584 m) |
0-5 (8.6) |
Abyzova |
Micrococci, short rods, pollen, many diatoms |
< 200 |
|
3555 |
|
15 |
Abyzova |
Cytophaga spp. and many Coccolithophoridae |
600 |
|
3565 |
|
5-10 |
Abyzova |
Caulobacter spp. |
250 |
|
3579 |
|
5-10 |
Abyzova |
Many micrococci, short rods, and Coccolithophoridae |
100 |
|
3585 |
|
0-5 |
Abyzova |
Cytophaga spp. |
250 |
|
3587 |
|
0-5 |
Sambrottob |
Fungi, rod-shaped bacteria, diatoms, pollen grains |
Not quantified |
|
3590 |
|
0-5 |
Priscuc |
Acidovorax sp., Comamonas sp., Afipia sp., Actinomyces sp. |
2800-36,000 |
|
3592 |
|
0-5 |
Abyzova |
Many different kinds |
900 |
|
3593 |
Shallow open lake grounding line (3593-3608 m) |
0-5 (4) |
Christnerd |
Brachybacterium sp., Paenibacillus sp., Methylobacterium sp., Sphingomonas sp. |
Not quantified |
|
3598 |
|
5-10 |
Abyzova |
Many Coccolithophoridae |
500 |
|
3603 |
|
5-10 |
Karle |
Gram-negative (?) rods, cocci, and vibrios |
200-300 |
|
3606 |
|
5-10 |
Abyzova |
Many different kinds |
700 |
|
3611 |
Open lake (3608-3623 m) |
0 (0) |
Abyzova |
Mostly diatoms and cyanobacteria |
100 |
|
3619 |
|
0 |
Rogers and Bulatf |
Serratia liquefaciens, Comamonas acidovorans, Rhodoturla creatinovora, Kocuria sp., Friedmanniella Antarctica, Hydrogenophlus theroluteolus |
Not quantified |
|
aData from Abyzov et al. (2005). Techniques used included light microscopy and scanning electron microscopy. Coccolithophoriade, cyanobacteia, actinobacteria, rod and coccus-shaped bacteria, and fungal conidia and hyphae were dectected in all ice horizons examined. bData from Sambrotto and Burckle (2005). Techniques included culturing and microscopy. cData from Priscu et. al. (1999). Techniques included microscopy and polymerase chain reaction (PCR) amplification. dData from Christner et al. (2005). Techniques included microscopy and PCR amplification. eData from Karl et al. (1999). Techniques included microscopy, flow cytometry, and 14C uptake. fData from S.O. Rogers and S. Bulat, unpublished. Techniques included culturing and PCR amplification. SOURCE: From Bell et al. (2005). |
|||||
sidering that metabolic inferences for the γ and δ-proteobacterial clones are based on distant phylogenetic relationships, confident predictions about physiology are unachievable. Although equivocal, the clustering of the identified small subunit rRNA gene sequences among aerobic and anaerobic species of bacteria with metabolisms dedicated to iron and sulfur respiration or oxidation implies that these metals play a role in the bioenergetics of microorganisms that occur in Lake Vostok.”
Likelihood of Growth
Rates of proliferation of introduced as well as native microbes are likely to be greatly reduced in subglacial aquatic environments by the low ambient temperatures and high pressure conditions. Growth rates may be suppressed further by the inhibitory effects of high oxygen concentrations and by low concentrations of inorganic nutrients and organic carbon substrates.
Christner et al. (2006) applied Price and Sower’s (2004) model to address the potential for survival, maintenance, and growth of microheterotrophs in Lake Vostok and concluded that carbon supply rates would be adequate for maintenance but not growth. It should be noted, however, that their upper estimate of in situ NPOC concentrations of 250 μmol L−1 (3 mg C L−1) greatly exceeds the total organic carbon in offshore marine environments of around 75 μmol L−1 (0.9 mg L−1).
Furthermore, organic carbon may vary among aquatic environments and spatially within aquatic environments and could achieve higher values than the estimates derived from Vostok accretion ice. For example, the organic carbon supply from melting ice and subglacial waters entering at the northern end of Lake Vostok could be higher than at the accretion ice end, more than 200 km away, particularly if there are organic inputs from active microbial processes at the ice sheet-rock interface. Many of the aquatic environments may be stratified and organic carbon levels could be higher in the bottom waters of each lake associated with microbial decomposition and release from the sediments.
Lake sediments typically contain orders of magnitude higher bacteria, nutrient, and DOC concentrations. The upward plumes associated with normal geothermal heating could potentially transport these constituents from the sediments to the upper water column in some aquatic environments. Subglacial lake sediments may also provide a microbial refuge from high oxygen concentrations in the overlying lake water.
Another of the interesting features of subglacial aquatic environments that distinguishes them from surface aquatic environments but is similar to deep marine environments is the absence of ultraviolet (UV) photochemistry. The exposure of natural surface waters to sunlight has multiple effects on DOC availability for microbial processes, including the photodegradation of higher-molecular-weight materials into more biologically reactive constituents, and the polymerization of certain autotroph-derived organic compounds into forms that are less available for microbial activity (Tranvik and Bertilsson 2001). This absence of UV effects will thus have both positive and negative implications for microbial growth in subglacial aquatic environments.
In summary, growth rates are unknown for bacteria in subglacial ecosystems, but they are likely to be slow and possibly negligible in some situations. Analogous systems include the deep subseafloor biosphere where bacterial turnover times have been estimated as up to 22 years (Schippers et al. 2005). These slow rates of metabolism and growth also have implications for the extent of genetic adaptation to the subglacial
environment given that rates of evolution tend to be slow under conditions of low temperature (Gillooly et al. 2005) and low productivity (Horner-Devine et al. 2003).
BIOLOGY—CONCLUSIONS
Summary
The subglacial aquatic environments of the Antarctic lie beneath kilometers of ice. Microbes of a number of types are found in low numbers throughout all depths of the ice sheet, carried to the Antarctic by atmospheric winds and deposited at the surface. Some of these survive their extremely slow downward travel through the ice and are still capable of respiration and growth when sampled in cores from thousands of meters below the surface. It is, therefore, highly likely that the subglacial aquatic environments contain microbes capable of growth and are constantly being inoculated with microbes from the ice sheet. The important question then centers on the probability that these environments support growing and evolving microbial populations. After considering the requirements for microbial growth and the chemical and physical conditions predicted for subglacial environments, the committee concluded that the hypothesis of microbial life and growth cannot be rejected. Therefore, future exploration of these environments must assume an aquatic ecosystem containing growing microbial populations. This assumption affects issues of preservation of habitats and minimization of contamination during exploration. The uncertainty can be resolved only by direct sampling of the lakes and flowing waters beneath the ice sheet.
Requirements for Life
The proliferation of microorganisms requires, at a minimum an inoculum of cells; water; electron donors and acceptors (e.g., reduced iron, sulfides, organic matter, oxygen) for biological energy supply; a source of nutrients (e.g., C, N, P, S, Fe, and other elemental constituents of biomolecules); sufficient time for reproduction; and the physical, chemical, or biological conditions that prevent cell destruction and promote growth. Subglacial aquatic environments have not been studied directly; therefore no unequivocal evidence exists to confirm the presence or absence of life in these ecosystems.
Subglacial Aquatic Environments and Potential Microbial Communities
Two types of evidence have been used to predict what will be found in waters beneath the ice sheet. First, microbes exist in all extreme environments on Earth where there is water, from a depth of 10,000 m in the ocean to surface saline lakes and from subzero temperatures in the brine channels of ocean ice to hot springs. Subsurface microbial communities of land and ocean are driven by abiotically produced materials from the Earth’s interior such as sulfides, hydrogen, and carbon dioxide. Based on this, it appears likely that actively growing microbes will be present in subglacial waters. Second, although Lake Vostok has not been studied directly, there are samples of the chemistry and microbiology of the ice that has been derived from the lake (accreted ice); these indicate that there are no environmental conditions that would rule out microbial life within the lake.
Another complication for predicting the present conditions in the subglacial waters is that rock particles from glacial scour and sediments from preglacial lakes might be present. These materials could be sources for biota as well as for electron donors and acceptors for chemotrophic and heterotrophic organisms.
No matter what their origin, the substrates required for microbial growth appear to be present in the water column of subglacial aquatic systems at low concentrations, although a richer microbial habitat may be provided in the underlying sediments. The supply rate of these substrates is one of the key unknowns; however, the concentrations of growth substrates are considered to be low, suggesting that these environments are oligotrophic. As a consequence, microbes adapted to these environments by necessity will most likely have highly proficient metabolic processes capable of taking advantage of extremely oligotrophic conditions including efficient recycling strategies. Depending on the level of carbon and nutrients available to the microbial community, metabolic activity and growth could be very low in some aquatic environments and some fraction of the community may represent a viable but metabolically inactive state. It is likely that microbial communities, if present and active, will only be able to grow extremely slowly.
The growth of accretion ice in Lake Vostok may have created an environment where oxygen concentrations are 50 times higher than in normal lake waters. This situation is likely to occur because enclathrated hydrates of atmospheric air contained in glacial ice are released into the lake when this ice melts. Accretion ice excludes gases as it grows; the result is supersaturation of the upper levels of the lake water with gases. Unless the gases are removed by mixing into the deep waters or by transport out of the lake, high concentrations of oxygen may result in the production of superoxide radicals and other reactive oxygen species that could be detrimental to life. Alternatively, if the subglacial lakes are stratified there may be low oxygenic levels and anoxic bottom waters and sediments where microbes could exist.
Microbes in Glacial Ice and Subglacial Waters
The microbes in glacial and accretion ice have been examined by several methods. Microscopic and culture analysis of the Vostok ice core revealed bacteria, yeasts, fungi, and microalgae in glacial ice greater than 240,000 years old. In the oldest ice, only bacterial spores were found. Microbiological and molecular-based studies of the accretion ice found low but detectable amounts of bacterial cells and DNA. Molecular identification of microbes show that they are closely related to microbiota from the surface of the Earth: Proteobacteria, Firmicutes, Actinobacteria, and Bacteroidetes. Microscopic examination revealed bacteria, pollen, diatoms, and coccolithophorids. Of these, only the bacteria would be able to grow in the lake water.
Bacteria and yeasts from the accretion ice, thus presumably from Lake Vostok, respired glucose and grew on liquid and agar media. The total numbers of microorganisms detected in the accretion ice ranged from 100 to 900 cells mL−1.
Actual growth rates are unknown for bacteria in subglacial ecosystems, but are expected to be slow and possibly negligible in some situations. Analogous systems include the deep subseafloor biosphere, where one report estimated that the bacterial turnover time is up to 22 years (Schippers et al. 2005). These slow rates of metabolism and growth also have implications for the extent of genetic adaptation to the subglacial
environment given that rates of evolution tend to be slow under conditions of low temperature (Gillooly et al. 2005) and low productivity (Horner-Devine et al. 2003).
The Possibility of Microbial Evolution
If the source population of microbial communities in subglacial aquatic environments is derived from the overlying glacial ice, these populations have been isolated for at least 1 million years, which is the age of the oldest Antarctic ice. Against the backdrop of 3.7 billion years during which microbes have been present on Earth, 1 million years is not a long time in terms of evolution. In the case of Lake Vostok, rates of evolution are likely to slow under the conditions of low temperature and low microbial productivity. The result is likely to be that the metabolically active and growing microorganisms present in these communities should represent those better adapted to the ambient conditions but will still be related to microorganisms found in other environments today.
A much longer time for evolution may have occurred if, however, the original inoculum to Lake Vostok dates back to the time of the isolation of the lake from the overlying atmosphere more than 15 million years before the present. This time of isolation could be even longer if the founding populations were derived from rocks or sediments. In this latter case, the microbial populations could have been isolated from the surface approximately 35 million to 40 million years ago, prior to the formation of the lake.
Samples of microbes from water and sediment are the only way to answer these questions of the uniqueness of microbial life in subglacial aquatic systems. Although the effect of the extreme conditions is unknown, there is a chance that the microbes have been isolated long enough that significant genetic divergence of microbial lineages has occurred.



















