Infectious diseases have always been with us, and always will. As Nobel Laureate Dr. Joshua Lederberg observed (Lederberg 2000), it is a competition between their genes and our brains. In addition to advances in medical products (such as drugs and vaccines) to treat or prevent natural infectious agents, multiple voices have argued that current advances in biological research and biotechnology would enable the development of bioengineered pathogens (Lindler et al. 2005; Petro et al. 2003), called by the U.S. Department of Defense (DoD) the “unknown-unknowns”1 because of their unknown profile and their unknown potential threat to warfighters and the public at large.
The United States and other governments have identified both the need to prevent the development of such designed pathogens and the need for a strategy to develop medical interventions, that is, medical countermeasures, against this unfamiliar group of potential infectious agents.
Preventing the development of biothreats would rely on predictive reasoning or covert discovery of the effort to develop such agents. However, while access to the methodologies, materials, and knowledge base of molecular biology particularly and bioscience more generally increases, the size and scale of such intelligence- and data-gathering capability decrease, making the reliable detection of such efforts possibly more difficult.
A responding strategy for addressing the threat of these novel unknown-unknowns is to thoroughly study the foundations and patterns of host-parasite or host-microbe evolutionary dynamics and patterns of interaction. All human infectious diseases had an origin in some preceding host-parasite system either of ancient or more recently recognized origin. The evolution of variola virus (causative agent of smallpox) is traced to an East African rodent host species 16,000-68,000 years ago, and Yersinia pestis is thought to have diverged from a Y. pseudotuberculosis lineage over the past 1,500-20,000 years, possibly as the bacterium adapted to life in the flea host (Achtman et al. 1999; Eppinger et al. 2010). An example of a modern emerging human disease is HIV whose origin is in nonhuman
_________________________
1 The term “unknown-unknown(s)” refers to pathogen(s) that may not be known or knowable because they currently may not exist. Due to the current or future possibility that they may exist, they are considered potential threats (e.g., a novel, genetically engineered, or created pathogen).
primates (Rambaut et al. 2004). An infinite array of patterns that yield favorable opportunities for a pathogen probably does not exist, and conversely, defense mechanisms that the host is capable of generating are probably few.
If the host is a collection of different environments and opportunities for exploitation by a pathogen, then a successful pathogen must bring the proper tools to exploit that opportunity (tailored adaptation strategy), and those tools probably fit into (recognizable) major patterns (mechanisms of pathogenesis). Any pattern or patterns of specific adaptations by these pathogens may be targeted for medical countermeasures, and mechanisms of pathogenesis that are similar or shared by different host species (human and nonhuman) may be used to demonstrate comparable efficacy of countermeasures when such assessment cannot be performed in humans.
ADDRESSING THE UNKNOWN THREAT THROUGH THE TRANSFORMATIONAL MEDICAL TECHNOLOGIES INITIATIVE2
The Transformational Medical Technologies (TMT; see Box 1-1) reflects a key transition point in the DoD’s philosophy about biological threats and the approach to developing medical countermeasures (MCMs). The overarching strategy for the TMT, conceived for the Quadrennial Defense Review (QDR) in 2006 by the DoD in its Chemical and Biological Defense Program Medical Research and Development, Testing & Evaluation (RDT&E) Plan, is as follows:
- The key to defending against unpredictable or unknown threats (e.g., bioengineered pathogens) lies not in expending resources to uncover the types of advanced agents that humans could face but rather in exploring and comparing the underlying pathophysiological patterns in the interaction of pathogen and host by using advanced scientific approaches, such as systems biology.
- In addition to the traditional method of looking for vulnerable pathogen targets, the strategy assumed the possibility of targeting broadly used host pathways for intervention. The TMT could ostensibly achieve broad protection against a variety of threats by looking at both host- and pathogen-based targets.
- The TMT’s strategy hypothesized that the key to defending against unknowns could be found in understanding potentially commonly evolved pathways and developing medical countermeasures focused on pathogenesis patterns, rather than on specific pathogens and the traditional “one-bug, one-drug” approach.
- The strategy suggested that pathogens that occupy similar “pathogenesis niches”, e.g., viruses that produce hemorrhagic responses in hosts, or bacteria that survive by exploiting an intracellular niche, acquired evolutionarily similar mechanisms or biochemical tools to achieve these niche-specific outcomes.
_________________________
2 The Transformational Medical Technologies Initiative (TMTI) became Transformational Medical Technologies (TMT) and is referred to as such throughout the report. In 2011 the Department of Defense moved the TMT to a Program Manager under the auspices of the Joint Program Executive Office for Chemical and Biological Defense, as the efforts have matured to advanced development. The Committee has addressed its report to the TMT.
BOX 1-1
Transformational Medical Technologies Initiative
The Department of Defense’s Transformational Medical Technologies Initiative (TMTI; now known as Transformational Medical Technologies; TMT) was organized in 2006 to boost countermeasure development with a “transformational” approach focusing on countermeasures with a broad enough therapeutic or preventive profile to defend against unanticipated and novel threats. The methods used to develop these countermeasures should be quickly adaptable to new threats once the genomic sequence of the pathogens is determined. The TMTI was to be transformational in shepherding the development of medical countermeasures and diagnostic products from early research through development phases - an “end-to-end” approach.
The development of medical countermeasures and diagnostic products by the TMTI was notable. The countermeasure work that is most relevant to this study includes investment in a wide range of projects including so-called platforms - technologies that can be used to quickly produce countermeasures against different targets. One example of the platform approach is the synthesis of antisense oligonucleotides that target the sequence of a biothreat agent by attacking the agent RNA with a complementary, or “antisense”, synthesized oligonucleotide strand that binds the agent’s RNA and leads to its destruction by the host cell (see Warren et al. 2010). Other funded research included efforts to create potentiators of the immune system.
In 2010, the TMTI became a program, instead of an initiative, known simply as Transformational Medical Technologies, or TMT. In 2011 the Department of Defense moved the TMT to a Program Manager under the auspices of the Joint Program Executive Office for Chemical and Biological Defense, as the efforts have matured to advanced development.
The QDR directed the DoD “to develop broad-spectrum MCMs against the threat of genetically engineered bio-terror agents” (DoD 2006, p 5), recognizing that emerging disease and potentially genetically engineered pathogens, could be leveraged as effective agents to wage “asymmetric warfare.” As a result, the TMT was created to provide new solutions for the warfighter that could be broadly generalized, even to unknown threats. The TMT was designed to incorporate systems biology approaches to understand patterns of pathogenesis for the purposes of developing and targeting medical countermeasures at various single or combined targets to achieve a broad level of protection or treatment (see Figure 1-1 for notional approach).
The program’s principal thrust has been to build a capability to respond to an event by using platform technologies to identify and counter unknown biological threat agents. Technologies have been developed that accelerate the process of definitive pathogen characterization, as well as the design and deployment of MCMs. The response capability that was tested in a real-world situation against the 2009 H1N1 influenza outbreak resulted in an effective medical countermeasure against the H1N1 virus. The countermeasure was produced using an antisense oligonucleotide therapeutic platform.
Another core area of the TMT has been the development, through Federal Drug Administration licensure, of “broad-spectrum” therapeutics. A defining feature of this MCM effort has been the use of intervention strategies that target multiple classes of pathogens, as opposed to the conventional “one-bug, one-drug” paradigm. Such approaches may defeat pathogens directly (through antibiotics or antivirals), exploit host targets attacked by multiple threat agents, or enhance host defenses by modulating the host’s immune response. An example of this approach is the targeting of the human protein TSG101, the product of the tumor susceptibility gene 101 that participates in the intracellular movement of proteins. This protein is “hijacked” by viral components to bring intracellular proteins to the surface of the cell for eventual virus assembly and budding (e.g., HIV and Ebola; Martin-Serrano et al. 2001). Once TSG101 is exposed on the cell surface it can be directly targeted using monoclonal antibodies to then help eliminate infected cells. Developed with TMT funding, one monoclonal antibody has demonstrated in vitro efficacy against many virus types, including HIV-1 and drug-
resistant HIV (Chen et al. 2010), and all forms of influenza (both seasonal and pandemic; Bonavia et al. 2010). Recent work in murine models has shown protection against a number of filoviruses through the development of broad-spectrum antivirals (FGI-106, FGI-103) whose cellular target remains undefined (Aman et al. 2009; Warren et al. 2010).
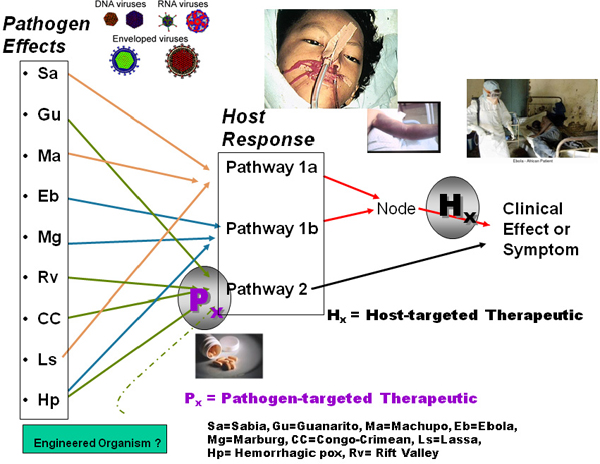
FIGURE 1-1 TMT concepts for broad capability against general categories or clusters of pathogens. These concepts exploit a particular life-history strategy, thereby achieving protection against unknown pathogens with similar pathogenesis patterns. In this graphic representation, major hemorrhagic fever viruses from different taxa (on the left side represented by two letters) are hypothesized to use one of three pathogenesis pathways to successfully infect a host, while the host has several major specific response pathways. Broad ability to defend against this entire class of pathogens through treatment would depend on developing a combination of therapeutics that act at several different but complementary points within the overall pattern (designated by Px and Hx), so that any unknown or engineered organism attempting to exploit this potential pathogenesis model would effectively be prevented from fulfilling its goal.
According to the DoD, the TMT is unique among U.S. government medical countermeasure efforts because it supports the full spectrum of drug development by funding basic research through advanced product development. Investigational New Drug (IND) filings for two hemorrhagic fever viruses (Marburg and Ebola) have resulted from this program, and additional IND filings are anticipated in the near future. The TMT’s integrated product development program has also sought out
and created public-private partnerships with pharmaceutical companies whose longer-term goals are compatible with the infectious-disease interests of the program. The program has become an integral component of the larger national effort to combat biological threats, whether they result from acts of terrorism or emerging infectious disease.
The study presented here was identified as an important topic by the National Research Council’s (NRC’s) Standing Committee on Biodefense for the Department of Defense. The Standing Committee was organized by NRC in 2007 at the request of the Office of the Secretary of Defense (Special Assistant for Chemical and Biological Defense and Chemical Demilitarization Programs). The Committee’s work focused on the DoD’s TMT and the challenges faced by the broader community in the development of medical countermeasures against biothreat agents. The TMT’s goals and approach were to be transformational (see Box 1-1). In focusing on bottlenecks and obstacles to the development of medical countermeasures the Standing Committee became aware of the Food and Drug Administration’s (FDA’s) Animal Rule (see Appendix A; 21 CFR Parts 314 and 601 [2002]). The Animal Rule is largely considered a step forward in addressing the fact that the efficacy of countermeasures against most biothreat agents cannot be tested in humans because it is unethical for humans to be given the disease against which the countermeasures are intended to work. However, while the FDA’s action to create a pathway for testing the effectiveness of countermeasures without human clinical trials was well received, experience since its promulgation demonstrated that it is not a facile pathway for assessing the efficacy of a countermeasure in humans based on the product’s efficacy in animals. This past decade has shown that the Animal Rule presents its own set of challenges, including developing appropriate animal models of pathogenesis and extrapolating results from animals to humans. Recognizing the need for focused attention on the issues, the Standing Committee and the DoD asked the NRC to organize a separate ad hoc committee to produce a report addressing issues related to animal models for testing countermeasure efficacy (see the complete Statement of Task in Appendix E).
Although this report was funded with the DoD and TMT’s needs in mind, the charge was to address a challenge that is widely accepted to be a major obstacle for the entire scientific and research community working on the development of medical countermeasures. The Committee on Animal Models for Assessing Countermeasures to Bioterrorism Agents was asked to:
- Evaluate how well the existing TMT-employed or candidate animal models reflect the pathophysiology, clinical picture, and treatment of human disease as related to the agents of interest.
- Address the process and/or feasibility of developing new animal models for critical biodefense research, placing emphasis on the need for a robust and expeditious validation process in terms of FDA’s Animal Rule.
- Evaluate alternatives to the use of animal models based on the premise of the Three Rs (refinement, reduction, and replacement of animal use; such venues would include but not be limited to in vitro work, computational modeling, new biotechnological tools, surrogate diseases, etc.) vis-à-vis the Animal Rule and FDA licensure. The evaluation will also consider the development of more humane models for infectious diseases research that do not incorporate death as an endpoint (i.e., humane endpoints).
In the chapters to follow, the Committee on Animal Models for Assessing Countermeasures to Bioterrorism Agents lays out its findings and offers potential solutions for the DoD to address the challenge of exclusively using animal models to demonstrate the effectiveness of a medical countermeasure in lieu of human efficacy data. The Committee did not consider animal models to evaluate the safety of products developed under the Animal Rule, as under the rule’s provisions “safety evaluation of products is not addressed in this rule” (FDA 2002, p 37989). Further, the Committee did not evaluate the Animal Rule or the FDA’s approach to assess product efficacy under the rule.
Chapter 2 looks at the adequacy of current animal model systems including an assessment of the data provided by these models versus available human data for filovirus-induced hemorrhagic fevers, anthrax, and tularemia. Chapter 3 discusses the history of the Animal Rule and relevant ethical issues. Chapter 4 explores the need for additional animal models to augment current capabilities and introduces the issue of qualification of models to be used for both hypothesis testing and regulatory purposes (i.e., toxicology studies). It suggests the compartmentalization of an animal model to match specific aspects of efficacy demonstration to individual components of the model rather than to results from the whole organism. Finally, chapter 5 considers what approaches and refinements should be applied now to current animal models for the TMT and recommends the exploration of advanced technologies and new types of genetically modified animals. It further discusses the potential value of supplementing the veterinary and clinical care of an experimental animal subjected to these pathogens to align with the clinical treatment received by the human patient.
Achtman M, Zurth K, Morelli G, Torrea G, Guiyoule A, Carniel E. 1999. Yersinia pestis, the cause of plague, is a recently emerged clone of Yersinia pseudotuberculosis. Proc Natl Acad Sci USA 96(24):14043-14048.
Aman MJ, Kinch MS, Warfield K, Warren T, Yunus A, Enterlein S, Stavale E, Wang P, Chang S, Tang Q, Porter K, Goldblatt M, Bavari S. 2009. Development of a broad-spectrum antiviral with activity against Ebola virus. Antiviral Res 83(3):245-251.
Bovania A, Diaz LS, Santos D, Cassella J, Fesseha Z, Bamba D, Sui B, Li W-B, Duan R, Chen L-M, Donis RO, Goldblatt M, Kinch MS. 2010. Antibody targeting of TSG101 on influenza-infected cells. Virus Adaptation and Treatment 2:147-157.
Chen H, Liu X, Li Z, Zhan P, De Clercq P. 2010. TSG101: A novel anti-HIV-1 drug target. Curr Med Chem 17(8):750-758.
DoD [U.S. Department of Defense]. 2006. Quadrennial Defense Review Report. Available online (http://www.defense.gov/qdr/report/Report20060203.pdf), accessed May 2011.
Eppinger M, Worsham PL, Nikolich MP, Riley DR, Sebastian Y, Mou S, Achtman M, Lindler LE, Ravel J. 2010. Genome sequence of the deep-rooted Yersinia pestis strain Angola reveals new insights into the evolution and pangenome of the plague bacterium. J Bacteriol 192(6):1685-1699.
FDA [Food and Drug Administration]. 2002. New Drug and Biological Drug Products; Evidence Needed to Demonstrate Effectiveness of New Drugs When Human Efficacy Studies Are Not Ethical or Feasible. Fed Regist 67(105):37988-37998.
Lederberg J. 2000. Infectious history. Science 288:287-293.
Lindler L, Choffnes E, Korch G. 2005. Definition and overview of emerging threats. In: Lindler L, Lebeda F, Korch G, eds. Biological Weapons Defense: Infectious Disease and Couterbioterrorism. New Jersey: Humana Press. p 351-360.
Martin-Serrano J, Zang T, Bieniasz PD. 2001. HIV-1 and Ebola virus encode small peptide motifs that recruit Tsg101 to sites of particle assembly to facilitate egress. Nat Med 7(12):1313-1319.
Petro JB, Plasse TR, McNulty JA. 2003. Biotechnology: Impact on biological warfare and biodefense. Biosecur Bioterror 1(3):161-168.
Rambaut A, Posada D, Crandall KA, Holmes EC. 2004. The causes and consequences of HIV evolution. Nat Rev Genet 5(1):52-61.
Warren TK, Warfield KL, Wells J, Enterlein S, Smith M, Ruthel G, Yunus AS, Kinch MS, Goldblatt M, Aman MJ, Bavari S. 2010. Antiviral activity of a small-molecule inhibitor of filovirus infection. Antimicrob Agents Chemother. 54(5):2152-2159.
Warren TK, Warfield KL, Wells J, Swenson DL, Donner KS, Van Tongeren SA, Garza NL, Dong L, Mourich DV, Crumley S, Nichols DK, Iversen PL, Bavari S. 2010. Advanced antisense therapies for postexposure protection against lethal filovirus infections. Nat Med 16(9):991-994.








