The completion of the human genome sequence in 2001 and the technologies that have emerged from the Human Genome Project have ushered in a new era in biomedical science. Using technologies in genomics, proteomics, and metabolomics, together with advanced analytical methods in biostatistics, bioinformatics, and computational biology, scientists are developing a new understanding of the molecular and genetic basis of disease. By measuring, in each patient sample, thousands of genetic variations, mutations, or changes in gene and protein expression and activity, scientists are identifying previously unknown, molecularly defined disease states and searching for complex biomarkers that predict responses to therapy and disease outcome.
This new understanding is beginning to shape both the ways in which diseases are managed and how new drugs and tests are being developed and used. For example, Oncotype DX (Paik et al., 2004) is a multiparameter gene expression test that helps determine which patients with early stage breast cancer are at higher risk of recurrence and thus may be more likely to benefit from chemotherapy, while allowing women at lower risk to safely forgo chemotherapy. These patients avoid the toxicities, cost, and quality-of-life issues associated with treatment. Increasingly, drugs are being developed to target specific disease subtypes or mutations, and companion diagnostic tests are being developed to identify the subsets of patients most likely to respond or least likely to suffer serious side effects.
Despite great promise, progress in translating such “omics-based” tests into direct clinical applications has been slower than anticipated. This has been attributed to the time-consuming, expensive, and uncertain development
pathway from disease biomarker discovery to clinical test; the underdeveloped and inconsistent standards of evidence to assess biomarker validity; the heterogeneity of patients with a given diagnosis; and the lack of appropriate study designs and analytical methods for these analyses (IOM, 2007). Some also have questioned the excitement afforded omics-based discoveries, suggesting that advancements will have primarily modest effects in patient care (Burke and Psaty, 2007).
Nevertheless, patients themselves recognize the promise of molecularly driven medicine and are looking to the scientific community to provide validated, reliable clinical tests that accurately measure and predict response to treatment and provide more effective ways of screening for disease. Among scientists and clinicians, omics-based tests are seen as presenting opportunities for important new clinical trial design strategies and hopefully reducing the time and cost of developing new treatments (Macconaill and Garraway, 2010).
As is true in all areas of scientific research, rigorous standards must be applied to assess the validity of any study results, particularly if the study involves patients. Recently, the scientific community raised serious concerns about several omics-based tests developed to predict sensitivity to chemotherapeutic agents, developed by investigators at Duke University. The initial papers describing these omics-based tests garnered extensive attention because results suggested a potential major advance in the discovery and use of omics-based tests to direct choice of therapy for individual cancer patients. Almost from the time of initial publication, however, concerns were raised about the validity of these gene expression–based tests; Keith Baggerly and Kevin Coombes of MD Anderson Cancer Center first approached the Duke University principal investigators, Anil Potti and Joseph Nevins, with questions on November 8, 2006 (Baggerly, 2011), soon after the October 22 electronic publication of the article (PubMed, 2006). Clinical investigators at their institution were interested in using the methods, but the statisticians could not reproduce the results with the publicly available data and information. These concerns were heightened upon the publication of an article by Baggerly and Coombes (2009), detailing several errors in the development of the tests, inconsistencies between primary data and data used in the articles, and the inability to reproduce results reported by the investigators. In addition, in July 2010, a letter to the director of the National Cancer Institute (NCI) signed by a group of more than 30 respected statisticians and bioinformatics scientists brought additional scrutiny to these concerns, especially because these omics-based tests were being used in clinical trials to direct patient care (Baron et al., 2010).
Between October 2007 and April 2008, three cancer clinical trials were launched at Duke University, in which patients with lung cancer or breast cancer were assigned to a chemotherapy regimen on the basis of the test results (see Appendix B for additional details).
Dr. Varmus asked the IOM to conduct an independent analysis of the omics-based tests developed at Duke and define evaluation criteria for ensuring high standards of evidence for the development of omics-based tests prior to their use in clinical trials. In an interview for the Cancer Letter, Dr. Varmus summarized the committee’s task:
The Duke episode, from my perspective, was simply another way of illus trating the dangers of not doing it right, not having the right kinds of safeguards. And with my various colleagues, including colleagues at Duke, I asked the Institute of Medicine to do a study. The intention was not to investigate wrongdoing, because that was going to be taken care of in other ways, but to think about what needs to be in place to ensure that correct evaluation of new approaches to cancer care had been undertaken, that we met competing standards, and that the evidence base for changing diagnosis itself or evaluation of responses or, more importantly, choice of therapies—was based on good evidence. I asked the IOM … to think carefully about what kinds of hoops people need to jump through before new information about cancer is actually used in the clinical setting. The risks are high here. (Goldberg, 2011, p. 4)
NCI biostatistician Lisa McShane provided further motivation for the committee’s work:
I have witnessed the birth of many omics technologies and remain excited about their potential for providing important biological insights and their potential to lead to clinical tests that might improve care for cancer patients. It is important, however, that we understand the challenges and potential pitfalls that can be encountered with use of these technologies. Some unfortunate events at Duke University involving the use of genomic predictors in cancer clinical trials were a major impetus for the formation of this committee. We need to take a step back to evaluate the process by which tests based on omics technologies are developed and determined to be fit for use as a basis for clinical trial designs in which they may be used to determine patient therapy. (McShane, 2010, p. 1-2)
The scientific community needs to address these gaps if we are to realize the full potential of omics research in patient care. Omics technologies not only hold great promise, but also pose substantial risks if not properly developed and validated for clinical use.
COMMITTEE APPOINTMENT AND CHARGE
With support from NCI, the Food and Drug Administration (FDA), the Centers for Disease Control and Prevention, the U.S. Department of Veterans Affairs, the American Society for Clinical Pathology, and the College of American Pathologists, an IOM committee was charged to identify appropriate evaluation criteria for developing clinically applicable omics-based tests and to recommend an evaluation process for determining when predictive tests using omics-based technologies are fit for use in clinical trials, especially those in which the assay is used to direct patient care (Box 1-1). The IOM appointed a 20-member committee with a broad range of expertise and experience, including experts in discovery and development of omics-based technologies, clinical oncology, biostatistics and bioinformatics, clinical pathology, ethics, patient advocacy, development and regulation of diagnostic tests, university administration, and scientific publication.
An ad hoc committee will review the published literature to identify appropriate evaluation criteria for tests based on “omics” technologies (e.g., genomics, epigenomics, proteomics, and metabolomics) that are used as predictors of clinical outcomes. The committee will recommend an evaluation process for determining when predictive tests based on omics technologies are fit for use as a basis for clinical trial design, including stratification of patients and predicting response to therapy in clinical trials. The committee will identify criteria important for the analytical validation, qualification, and utilization components of test evaluation.
The committee will apply these evaluation criteria to predictive tests used in three cancer clinical trials conducted by Duke University investigators (NCT00509366, NCT00545948, NCT00636441). For example, the committee may assess the analytical methods used to generate and validate the predictive models, examine how the source data that were used to develop and test the predictive models were generated or acquired, assess the quality of the source data, and evaluate the appropriateness of the use of the predictive models in clinical trials.
The committee will issue a report with recommendations regarding criteria for using models that predict clinical outcomes from genomic expression profiles and other omics profiles in future clinical trials, as well as recommendations on appropriate actions to ensure adoption and adherence to the recommended evaluation process. The report will also include the committee’s findings regarding the three trials in question.
Before the IOM committee convened for its first meeting, investigators at Duke concluded that the omics-based tests used in the three clinical trials were invalid. They terminated the clinical trials, and began the process of retracting the papers describing the development of the tests. As a result, the committee did not undertake a detailed analysis of the data and computer code used in the development of those tests. Rather, the committee focused on how errors in the development process resulted in those tests being used in clinical trials before they were fully validated, and on developing best practices that would prevent invalid tests from progressing to the clinical testing stage in the future.
A rigorous process was undertaken in the development of the committee’s recommendations that included a review of the field of omics-based research, the processes necessary for verification and validation of omics-based tests, examination of what transpired in the development of the omics-based tests listed in the statement of task as well as other case studies of omics-based test development selected by the committee, and identification of the parties responsible for funding, oversight, and publication of results. Recommendations developed by the committee should be considered a roadmap critical to omics-based test development. The recommendations address the roles and responsibilities of all partners involved in the process, including individual scientists, their institutions, funding agencies that support the work, journals that publish the results of these studies, and FDA, which ultimately helps to define how these tests will make their way to clinical application.
Outside the Scope
The processes and criteria for adoption and use of omics-based tests in standard clinical practice are outside the scope of this report. The process of taking an omics-based test into clinical trials to evaluate a test for clinical utility and use is described, but no recommendation is made on how, finally, to take a test from the clinical trial setting into clinical practice. However, discussion of this step is critical for understanding the recommendations of the committee because this step may involve using an omics-based test to direct patient management in clinical trials, which is within the charge of the committee. Regardless, if an omics-based test is to be considered for use in clinical practice, one of three pathways needs to be followed to determine clinical utility, and all of these require a fully specified and validated omics-based test. When considering the parties responsible in the development of omics-based tests, the committee considered international funders to be outside the scope of the recommendations. Issues specific to tests that fall outside the committee’s definition of omics-based tests, such as single gene tests and whole genome sequencing, are also not addressed.
It is important to note that the IOM’s study is in no way linked to the concurrent scientific misconduct investigation at Duke University, and that inquiries about misconduct were not within this committee’s purview.
Definitions
Precise definitions and use of correct terminology are important for ensuring understanding, especially given the complexity of the rapidly expanding field of omics. The committee defined terminology that was central to its deliberations and recommendations (Box 1-2). Where possible, the committee used widely accepted definitions, such as those from the Biomarkers Definition Working Group. The terms “analytical validation,” “clinical validation,” and “clinical utility” have been adapted from the widely used definitions of the Evaluation of Genomic Applications in Practice and Prevention initiative, established by the Centers for Disease Control and Prevention (Teutsch et al., 2009). The committee has adapted this terminology by incorporating statistics and bioinformatics validation through use of the term “clinical/biological validation.”
Analytical Validation: Traditionally, “assessing [an] assay and its measurement performance characteristics, determining the range of conditions under which the assay will give reproducible and accurate data.a With respect to omics, assessing a test’s “ability to accurately and reliably measure the … analyte[s] … of interest in the clinical laboratory, and in specimens representative of the population of interest.”b
Biomarker: “A characteristic that is objectively measured and evaluated as an indicator of normal biological processes, pathogenic processes, or pharmacologic responses to a[n] … intervention.”c
Clinical Utility: “Evidence of improved measurable clinical outcomes, and [a test’s] usefulness and added value to patient management decision-making compared with current management without [omics] testing.”b
Clinical/Biological Validation: Assessing a test’s “ability to accurately and reliably predict the clinically defined disorder or phenotype of interest.”b
Cross-validation: A statistical method for preliminary confirmation of a computational model’s performance using a single dataset, by dividing the data into multiple segments, and iteratively fitting the model to all but one segment and then evaluating its performance on the remaining segment.
Effect Modifier: A measure that identifies patients most likely to be sensitive or resistant to a specific treatment regimen or agent. An effect modifier is particularly useful when that measure can be used to identify the subgroup of patients for whom treatment will have a clinically meaningfully favorable benefit-to-risk profile.
High-Dimensional Data: Large datasets characterized by the presence of many more predictor variables than observations, such as datasets that result from measurements of hundreds to thousands of molecules in a relatively small number of biological samples. The analysis of such datasets requires appropriate computing power and statistical methods.
Omics: Scientific disciplines comprising study of related sets of biological molecules. Examples of omics disciplines include genomics, transcriptomics, proteomics, metabolomics, and epigenomics.
Omics-Based Test: An assay composed of or derived from many molecular measurements and interpreted by a fully specified computational model to produce a clinically actionable result.
Overfitting: Occurs when the model-fitting process unintentionally exploits characteristics of the data that are due to noise, experimental artifacts, or other chance effects that are not shared between datasets, rather than to the underlying biology that is shared between datasets. Overfitting leads to a statistical or computational model that exhibits very good performance on the particular dataset on which it is fit, but poor performance on other datasets. Although not unique to omics research, the chance of overfitting increases when the model has a large number of measurements relative to the number of samples.
Preanalytical Variables: Aspects of sample collection and handling that need to be standardized and documented prior to test development and use.
Predictive Factor: An effect modifier of treatment.
Prognostic Factor: A measure correlated with a clinical outcome in the setting of natural history or a standard of care regimen; It is a variable used to estimate the risk of or time to clinical outcomes.
Statistics and Bioinformatics Validation: Verifying that the omics-based test can perform its intended task. Ideally, this involves assuring that the test can accurately predict the clinical outcome of interest in an independent set of samples that were not used in developing the test. Such validation is particularly important as omics tests typically involve computational models whose parameters can be over-fit in any single dataset, leading to an overly optimistic sense of the test’s accuracy.
SOURCES: a Wagner, 2002; b Teutsch et al., 2009; c Biomarkers Definitions Working Group, 2001.
The committee provides additional scientific and technical definitions in Chapters 2, 3, 4, the Glossary, and Appendix C.
Introduction to Biomarkers
The set of biological information measured and analyzed in a validated omics-based test is an example of a biomarker. This section introduces the concept and history of biomarkers. The scientific literature provides definitions of the term “biomarker” as well as some of the principal uses of biomarkers. A widely used definition of a biomarker is “a characteristic that is objectively measured and evaluated as an indicator of normal biological processes, pathogenic processes, or pharmacologic responses to a[n] … intervention” (Biomarkers Definitions Working Group, 2001). A recent IOM report on Evaluation of Biomarkers and Surrogate Endpoints in Chronic Disease provided the following description of biomarkers:
Biomarkers are measurements of biological processes. Biomarkers include physiological measurements, blood tests and other chemical analyses of tissue or bodily fluids, genetic or metabolic data, and measurements from images. Cholesterol and blood sugar levels are biomarkers, as are blood pressure, enzyme levels, measurements of tumor size from MRI or CT, and the biochemical and genetic variations observed in age-related macular degeneration. Emerging technologies have also enabled the use of simultaneously measured “signatures,” or patterns of co-occurring sets, of genetic sequences, peptides, proteins, or metabolites as biomarkers. These signatures can also be combinations of several of these types of measurements; ideally, each component of a signature is identified. (IOM, 2010, p. 2-3)
Biomarkers can be measurements of macromolecules (DNA, RNA, proteins, lipids), cells, or processes that describe a normal or abnormal biological state in an organism. Biomarkers may be detected and analyzed in tissue, in circulation (blood, lymph), and in body fluids (urine, stool, saliva, sputum, breast nipple aspiration, etc.). Biomarkers have many important potential roles in settings such as discovery research, clinical practice, and public health practice; these and other biomarker uses are described in Table 1-1 (IOM, 2010).
Uses intended for clinical practice include risk assessment, screening, diagnosis, prognosis, prediction of response to therapy (effect modifiers), prediction of clinical outcome (surrogate endpoints), and patient monitoring during and after treatment (Table 1-2).
It is important to understand a key distinction between two types of biomarkers: prognostic factors and effect modifiers. Prognostic factors are correlated with a clinical outcome in the setting of a specified clinical regimen. They are used to estimate the risk of or the time to clinical outcomes. However, a pure prognostic factor does not predict whether future, additional patient management strategies or therapies will be effective. Conversely, an effect modifier identifies patients most likely to be sensitive
TABLE 1-1 Categories of Biomarker Use
|
|
|
| Use | Description |
|
|
|
| Discovery | Identification of biochemical, image, or other biomarkers associated with a disease, condition, or behavior of interest; biomarkers identified may be screened for many potential uses, including as a target for intervention to prevent, treat, or mitigate a disease or condition |
| Early product development | Biomarkers used for target validation, compound screening, pharmacodynamic assays, safety assessments, and subject selection for clinical trials, and as endpoints in early clinical screening (i.e., Phase I and II trials) |
| Surrogate endpoints for claim and product approvals | Biomarkers used for Phase III clinical testing or to substantiate claims for product marketing when the effect of treatment on the biomarker reliably predicts the effect of treatment on a direct measure of how a patient feels, functions, or survives |
| Clinical practice | Biomarkers used by clinicians for uses such as risk stratification, disease prevention, screening, diagnosis, prognosis, therapeutic monitoring, and posttreatment monitoring |
| Clinical practice guidelines | Biomarkers used to make generalized recommendations for healthcare practitioners in the areas of risk stratification, disease prevention, treatment, behavior/lifestyle modifications, and more |
| Comparative efficacy and safety | Biomarkers used in clinical studies looking at the relative efficacy, safety, and cost effectiveness of any or all interventions used for a particular disease or condition, including changes in behavior, nutrition, or lifestyle; these studies are a component of comparative effectiveness research |
| Public health practice | Biomarkers used to track public health status and make recommendations for prevention, mitigation, and treatment of diseases and conditions at the population level |
|
|
|
| SOURCE: Adapted from IOM, 2010. | |
or resistant to a specific treatment regimen or agent. Effect modifiers are particularly useful when they can be used to identify the subgroup of patients for whom treatment will have a clinically meaningful favorable benefit-to-risk profile. In oncology, effect modifiers are also referred to as predictive factors, treatment-guiding biomarkers, or treatment selection biomarkers (Henry and Hayes, 2006; McGuire et al., 1990). While many people frequently use the term “predictive factor” rather than “effect modifier,” the use of this term is problematic because most dictionaries indicate
TABLE 1-2 Use of Biomarkers in Clinical Practice
|
|
|
| Clinical Biomarker Use | Clinical Objective |
|
|
|
| Disease risk stratification | Assess the likelihood that disease will develop (or recur) |
| Screening | Detect and treat early-stage disease in the asymptomatic population |
| Diagnosis/differential diagnosis | Definitively establish the presence and precise description of disease |
| Classificationa | Classify patients by disease subset |
| Prognosis | Estimate the risk of or the time to clinical outcomes |
| Prediction/treatment stratificationa | Predict response to particular therapies and choose the drug that is mostly likely to yield a favorable response in a given patient |
| Therapy-related risk management | Identify patients with a high probability of adverse effects of a treatment |
| Therapy monitoringb | Determine whether a therapy is having the intended effect on a disease and whether adverse effects arise |
| Posttreatment monitoring | Early detection and treatment of advancing disease or complications |
|
|
|
| a Companion diagnostic biomarkers include features from several of these categories. These tests identify whether an individual’s molecular profile associated with a disease pathophysi-ology is likely to respond favorably to a particular therapeutic. Examples include KRAS– cetuximab, HER2–herceptin, and estrogen receptor–tamoxifen. b Dose optimization is a subset of this category. SOURCE: Adapted from IOM, 2007, 2010. |
|
that the adjectives predictive and prognostic have very similar meanings. This report uses the term “effect modifier.” It should be noted that a bio-marker can be both prognostic and an effect modifier.
More detailed descriptions of biomarker types and examples, as well as the types of clinical studies and trials in which biomarkers are developed, are given in Appendix C.
Evaluation of Biomarkers and Surrogate Endpoints
As outlined in the statement of task, this committee was charged with identifying appropriate evaluation criteria for tests based on omics technologies, including criteria for the analytical validation, qualification, and
utilization components of test evaluation. The terminology of analytical validation, qualification, and utilization stems from the IOM consensus report Evaluation of Biomarkers and Surrogate Endpoints in Chronic Disease (IOM, 2010). The 2010 committee recommended a three-step biomarker evaluation framework consisting of analytical validation, qualification, and utilization, and intended the framework to be applicable to a diverse range of biomarker uses, including panels of biomarkers. The qualification step of biomarker evaluation is parallel to this report’s clinical/ biological validation step.
The 2010 IOM report emphasized the importance of a test’s intended use when making determinations in the utilization stage of biomarker evaluation. If a test’s validation did not reach the level needed for its intended use, the test would be sent back for further development. The interdependence of the steps in the evaluation process is highlighted in Figure 1-1.
This report’s process for discovery and development of omics-based tests can be viewed as an example of how the process above can be applied in a more specific case. The 2010 report covered all types of biomarkers and surrogate endpoints, including single and multiple analyte, molecular or imaging, and quantitative or qualitative biomarkers. Omics-based biomarkers are generally quantitative, involve measurement of multiple analytes, and involve use of computational models.
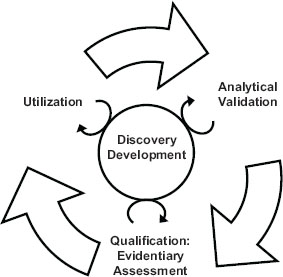
FIGURE 1-1 The steps of the biomarker evaluation are interdependent.
SOURCE: IOM, 2010.
Omics-Based Biomarkers and Omics-Based Tests
Omics-based tests can be considered a complex form of a biomarker test, using a defined set of measurements combined with a precisely defined computational model as a clinical test, for any of the purposes defined above for biomarkers. Several features distinguish omics-based biomarkers and omics-based tests from other biomarkers and biomarker-based tests. Most importantly, an omics-based test is derived from complex high-dimensional data; these data are often generated through measurement of many more variables per sample than the total number of biological samples used to generate the dataset. These data are used to produce a computational model1 that can be used to analyze samples from individual patients. High-dimensional data are particularly prone to overfitting, which can result in a computational model that functions well on the samples used for test development, but is inaccurate on any other sample. With careful analysis and a series of studies leading to a valid test, an omics-based test can be used to help a clinician make a decision about a patient’s care.
Several other characteristics distinguish omics-based tests from other medical technologies, including regulation and oversight of the development process and the difficulty in defining the biological rationale behind the test.
Omics-based tests and other clinical laboratory tests are subject to a different regulatory framework than drugs; for example, there are more pathways for regulation of devices—the regulatory category under which omics-based tests fall—than there are for drugs. Also, test development is more likely to occur in an academic setting than for drugs. Because the regulatory and oversight requirements for clinical laboratory tests are both different and less clear than for drugs, a greater burden is placed on the institutions to oversee biomarker-based test research and development. While pharmaceutical companies follow well-established drug development pathways and have many process controls in place for strong oversight of drug development and manufacturing, academic institutions are not as accustomed to overseeing the development of medical products. More work may be needed in academic institutions to reach an appropriate level of oversight.
The frequent lack of a clear biological rationale further distinguishes omics-based tests from other biomarker-based tests. It is usually possible
1 Includes all data processing steps, normalization techniques, weights, parameters, and other aspects of the model, as well as the mathematical formula or formulas used to convert the data into a prediction of the phenotype of interest.
to explain the biological rationale behind a single-biomarker test: The test is useful because the biomarker plays a role in disease pathology or other biological process under investigation. Examples of single-biomarker tests include HER2 tests or blood levels of low-density lipoprotein (LDL) cholesterol. For omics-based tests, however, the opposite is often true: It is generally not possible to explain the biological reasons why the test works. This difference puts an additional burden on the statisticians and bio informatics experts involved in test validation. Because of the risk of overfitting involved with omics-based test development, the need for rigor, validation, and accountability is even higher than for other biomarker-based tests.
ENGAGEMENT OF STAKEHOLDERS AND IMPLEMENTATION OF THE RECOMMENDATIONS
The future of omics-based tests and the realization of the promise they hold may well depend on the adoption of the recommendations put forth in this report. This report will be relevant to multiple audiences, including the responsible parties to whom the recommendations are directed and the various scientific disciplines and professions involved in the discovery and translation of omics-based biomarkers and tests. These responsible parties include the biomedical and clinical research community, investigators, institutions—public and private, commercial and nonprofit—funders of omics research, and journals that publish the results of omics research and clinical trials. Finally, the general public, as potential clinical trial participants and the beneficiaries of the products developed through this research, also may be interested. However, the technical aspects of this report are intended for a scientific audience. The omics-based test development process is complex and thus calls for a complex and rigorous methodology.
Many different scientific disciplines and professions are involved in the discovery, development, and validation of omics-based tests. Investigators that conduct the studies and the institutions that oversee their research are key to adoption and implementation of these recommendations. Laboratory scientists in many fields engage in omics research. Quantitative scientists, including those trained in biostatistics and bioinformatics, are an essential part of the scientific team in omics research because of the need to manage and use large datasets and to generate and validate the predictive models. Clinical researchers are responsible for the design and implementation of clinical trials that assess the clinical utility of new tests and can play a major role in the adoption of these recommendations. Organizations that fund this research and scientific journals that publish the results are also integral to advancing this research. FDA, as a regulatory agency, has a substantial role in oversight of this science as well.
Chapters 2 through 4 describe the recommended omics-based test evaluation process that the committee was charged with developing.
Chapter 2 provides an overview of the science and technology underpinning omics research and the recommended discovery and confirmation process prior to development and validation of a clinical omics-based test.
Chapter 3 describes the processes for defining the clinical test method and for assessing analytical and clinical/biological validation prior to use of an omics-based test in a clinical trial to evaluate the clinical utility and use of the test.
Chapter 4 describes the process for assessing the clinical utility and use of a new omics-based test.
Chapter 5 examines the roles of responsible parties in the development of omics-based tests: the roles of investigators and institutions in ensuring high-quality discovery and test development, the roles of journals and funders in the publication and financing of research to develop omics-based tests, and the role of FDA.
Chapter 6 presents an overview of lessons learned from the case studies, which are described in detail in Appendix A and B.
Recommendations are presented in Chapters 2-5 and in the Summary.
Baggerly, K. A. 2011. Forensics Bioinformatics. Presented at the Workshop of the IOM Committee on the Review of Omics-Based Tests for Predicting Patient Outcomes in Clinical Trials, Washington, DC, March 30-31.
Baggerly, K. A., and K. R. Coombes. 2009. Deriving chemosensitivity from cell lines: Forensic bioinformatics and reproducible research in high-throughput biology. Annals of Applied Statistics 3(4):1309-1334.
Baron, A. E., K. Bandeen-Roche, D. A. Berry, J. Bryan, V. J. Carey, K. Chaloner, M. Delorenzi, B. Efron, R. C. Elston, D. Ghosh, J. D. Goldberg, S. Goodman, F. E. Harrell, S. Galloway Hilsenbeck, W. Huber, R. A. Irizarry, C. Kendziorski, M. R. Kosorok, T. A. Louis, J. S. Marron, M. Newton, M. Ochs, J. Quackenbush, G. L. Rosner, I. Ruczinski, S. Skates, T. P. Speed, J. D. Storey, Z. Szallasi, R. Tibshirani, and S. Zeger. 2010. Letter to Harold Varmus: Concerns about Prediction Models Used in Duke Clinical Trials. Bethesda, MD, July 19. http://www.cancerletter.com/categories/documents (accessed January 18, 2012).
Biomarkers Definitions Working Group. 2001. Biomarkers and surrogate endpoints: Preferred definitions and conceptual framework. Clinical Pharmacology and Therapeutics 69(3):89-95.
Burke, W., and B. M. Psaty. 2007. Personalized medicine in the era of genomics. Journal of the American Medical Association 298(14):1682-1684.
Goldberg, P. 2011. A year at NCI: Harold Varmus reflects on provocative questions, Duke scandal, financial disaster and grant review. Cancer Letter 37(29):1-7.
Henry, N. L., and D. F. Hayes. 2006. Uses and abuses of tumor markers in the diagnosis, monitoring, and treatment of primary and metastatic breast cancer. Oncologist 11(6):541-552.
IOM (Institute of Medicine). 2007. Cancer Biomarkers: The Promises and Challenges of Improving Detection and Treatment. Washington, DC: The National Academies Press.
IOM. 2010. Evaluation of Biomarkers and Surrogate Endpoints in Chronic Disease. Washington, DC: The National Academies Press.
Macconaill, L. E., and L. A. Garraway. 2010. Clinical implications of the cancer genome. Journal of Clinical Oncology 28(35):5219-5228.
McGuire, W. L., A. K. Tandon, D. C. Allred, G. C. Chamness, and G. M. Clark. 1990. How to use prognostic factors in axillary node-negative breast cancer patients. Journal of the National Cancer Institute 82(12):1006-1015.
McShane, L. 2010. NCI Address to the Institute of Medicine Committee on the Review of Omics-Based Tests for Predicting Patient Outcomes in Clinical Trials. Meeting 1. Washington, DC. December 20.
Paik, S., S. Shak, G. Tang, C. Kim, J. Baker, M. Cronin, F. L. Baehner, M. G. Walker, D. Watson, T. Park, W. Hiller, E. R. Fisher, D. L. Wickerham, J. Bryant, and N. Wolmark. 2004. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. New England Journal of Medicine 351(27):2817-2826.
PubMed. 2006. PubMed entry for Genomic Signatures to Guide the Use of Chemotherapeutics by Potti et al., Nature Medicine, 2006. http://www.ncbi.nlm.nih.gov/pubmed/17057710 (accessed October 18, 2011).
Teutsch, S. M., L. A. Bradley, G. E. Palomaki, J. E. Haddow, M. Piper, N. Calonge, D. Dotson, M. P. Douglas, and A. O. Berg. 2009. The Evaluation of Genomic Applications in Practice and Prevention (EGAPP) Initiative: Methods of the EGAPP Working Group. Genetics in Medicine 11(1):3-14.
Wagner, J. A. 2002. Overview of biomarkers and surrogate endpoints in drug development. Disease Markers 18(2):41-46.
















