Informatics-Supported Cancer Research Endeavors
DISCUSSION POINTS HIGHLIGHTED BY INDIVIDUAL PRESENTERS
• Cloud-based informatics platforms aim to simplify interaction and information sharing between scientists and oncologists so that targeted treatments can begin faster; can manage billions of data points generated per patient; and can reduce time for data mapping and analysis from months to days to create a real-time, growing body of knowledge.
• The National Comprehensive Cancer Network (NCCN) outcomes database is one example of a collaborative data collection system that is relevant to both clinical practice and robust clinical outcomes research.
• Web-based EHR systems can facilitate real-time alerts, decision support capability, and multiple user applications.
• Patient-centered outcomes empower providers to make tailored recommendations to the individual patient based on data from other patients most like them.
• Secondary use of data that were collected for a different primary purpose generally involves some sort of adaptation or compromise. The challenge is to identify how this secondary use can best complement and extend the primary use of the data, for example, for generation of new hypotheses.
• Trust is the core underlying issue for concerns regarding consent, data privacy and security, accountability, and data ownership. Consumers and patients want their data to be protected, and they want medicine and health care to be advanced.
In this session, four cancer use cases were presented as examples of successful informatics-supported approaches to managing large, complex datasets. Panelists discussed data collection, storage, and retrieval; data analysis and reporting; and data sharing.
Spyro Mousses, vice president in the Office of Innovation and director of the Center for BioIntelligence at the Translational Genomics Research Institute (TGen), described a clinical trial using molecularly guided individualized therapy in pediatric cancer as a case example of successful alignment of biomedical science and informatics. To begin, Mousses described what he called “the evolution from evidence-based medicine to information-enabled medicine to intelligence-based medicine” and TGen’s “N = 1” approach to drug development (Figure 4-1).
An N = 1 Approach to Clinical Research
When seeking to develop a drug for a deadly disease such as cancer, investigators generally start with a broad target population from which they select a representative study cohort for a clinical trial comparing one therapeutic option against another. Evidence-based decisions on treatment are informed by statistical outcomes of the trial. For example, if the data indicate a 30 percent response rate to therapeutic option 1 and a 20 percent response rate to option 2, then therapeutic option 1 would be the drug of choice to be developed for all patients in the original target population (Figure 4-1A). Such a statistical approach is not ideal when dealing with a disease or condition that is clinically heterogeneous and molecularly complex, Mousses said.
Moving beyond basic evidence-based medicine, information-enabled
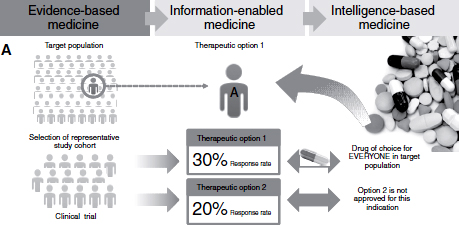
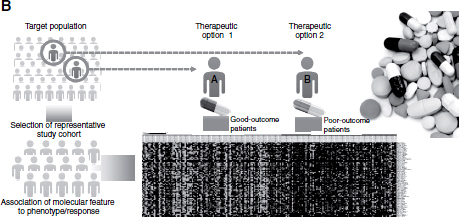
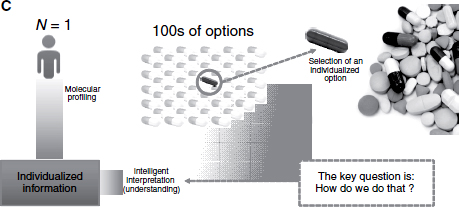
FIGURE 4-1 The evolution from evidence-based medicine (A) to information-enabled medicine (B) to intelligence-based medicine (C).
SOURCE: Mousses presentation (February 27, 2012).
medicine takes into account more detailed characteristics of the disease, such as expression of tumor markers or measurement of biomarkers. This approach also starts with a target population and the selection of a representative cohort for a clinical trial, but the cohort is then further stratified based on the association of a molecular feature with a phenotype or response. This stratification can allow one to determine that patient A would benefit more from therapeutic option 1, while patient B would be better treated with therapeutic option 2 (Figure 4-1B). Still, there are affected individuals who do not respond to either drug A or drug B.
To help address these concerns, TGen has been experimenting with an N of 1 approach where the focus is not on finding a representative population in which to test an investigative compound, but rather on finding a drug that meets the needs of a single individual based on his or her molecular profile (Figure 4-1C). This approach requires intelligent interpretation of individualized information and a mechanistic understanding of the disease to allow for predictions about which therapy is most appropriate. About a decade ago, TGen started to take the fundamental steps in the direction of molecular-based cancer treatments and has been using technologies ranging from simple gene expression and chemical assays to whole-genome sequencing to identify targets, Mousses said.
Molecularly Guided Individualized Cancer Therapy
As an example of the N of 1 approach to drug development, Mousses described an ongoing pediatric neuroblastoma clinical trial. To make the most of its molecular technologies, TGen needed innovative IT infrastructure. Through a collaboration with Dell, it created a cloud-based computing system that would both support the workflow of the trial and aid in repurposing the data. It is both a cloud to support personalized medicine, Mousses explained, and a repository to support translational research. As a result, the trial will be expanding to other pediatric and young adult cancers.
The process starts with clinical trial design, working within the framework of the regulatory agencies and in accordance with the IRBs. Following consent and enrollment of the patients, biopsies are collected and molecular profiling is done using next-generation sequencing technologies. Mousses noted that the collaboration with Dell has allowed faster and deeper profiling for each child and faster and improved data and clinical communication. A personalized treatment plan is then devised based on the profiling. This model for personalized medicine clinical trials, using molecularly
guided treatment and the Dell IT infrastructure and TGen cloud, is also being used for a new melanoma trial.
These types of trials are necessary experiments, Mousses said, to help define the IT requirements for intelligence-based personalized medicine using cloud-based informatics. The molecular profiling presents the first IT challenge, requiring high-performance computing and storage for 200 billion data points generated per patient. Another IT challenge is to match the patient profile to known information about therapeutic concepts, pathway concepts, and cellular processes, for example. This requires very large knowledge databases (e.g., pharmacogenomic databases) and presents numerous challenges. How should these complex, heterogeneous data be presented and processed? How can they be shared and exchanged across sites? There are also issues of security and privacy in the cloud. One of the models TGen is exploring is a hybrid cloud, where each clinical center would be able to warehouse its own data, keeping the patient data secure within the confines of the health care enterprise.
The TGen model for personalized medicine clinical trials facilitates intelligent use of the data to help each individual patient, Mousses said, but it does not yet allow for repurposing the data for secondary studies. TGen’s future vision is to provide a system that effectively links the pediatric oncology community, including software, hardware, and protocols that support data exchanges so that they are secure. Infrastructure for communication and collaboration is critical, Mousses said, and the system would include scientific, clinical, and community web portals. The goal is for the wider oncology community to have access to the network and be able to manage, analyze, and link patient data to pharmacological knowledge to identify potential treatment options. Information about the prediction and the outcome should feed into the model to create a real-time growing body of knowledge, not just about what is working, but about what is not working as well.
August Calhoun, vice president at Dell Healthcare and Life Sciences Services, added that the cloud increases computation and collaboration capacity by 1,200 percent and reduces the time needed for data mapping and analysis from months to days. The cloud is a shared resource that will be accessed over the Internet to do the complex analyses required to make
better-informed decisions about care and to share information in a consented and secure manner, he said.
CASE EXAMPLE: NATIONAL COMPREHENSIVE CANCER NETWORK OUTCOMES DATABASE
NCCN is an affiliation of 21 leading cancer centers throughout the United States. The overarching mission of the network, explained Kimary Kulig, vice president of clinical and translational outcomes research at NCCN, is to improve the quality, effectiveness, and efficiency of oncology practice so that patients can live better lives. NCCN seeks to enhance care through information resources, outcomes research, and clinical trials, and to develop information resources that are valuable to patients and other stakeholders within the health care delivery system.
To aid in fulfilling its mission, NCCN issues comprehensive oncology clinical practice guidelines that include clinical algorithms and supporting documentation for various tumor types. There are currently 56 individual guidelines covering 97 percent of all malignant diseases. Guidelines are developed and continually updated in collaboration with the 21 NCCN member institutions and efforts that involve more than 18,000 volunteer-hours per year from 900 clinicians who participate on guideline panels. Guidelines are categorized according to both level of evidence and degree of consensus among panel members.
NCCN Oncology Outcomes Database
To understand the extent to which NCCN member institutions adhered to the guidelines they helped to develop, NCCN launched an outcomes database. In addition to the primary goals of monitoring and benchmarking concordance with the guidelines, the database was also intended to be used to describe patterns and outcomes of care under the guidelines and to create a feedback loop to the physicians, institutions, and guideline panel members. Kulig noted that the outcomes database has also become a major data repository for research, aided by the fact that it is based on a common data dictionary and thus serves as a platform for multi-institutional health services research.
The first database, launched in 1997, was focused on breast cancer and currently includes data on close to 54,000 patients who are followed actively throughout their entire course of treatment until death. There are 17 institutions actively participating in this database. With the architecture in place, databases for other tumor types were subsequently launched, including non-Hodgkin’s lymphoma, colorectal cancer, non-small-cell lung cancer, and ovarian cancer.
Structure and Operations
The NCCN outcomes database is governed by the Scientific Office, a virtual office that includes investigators who are chairs of each of the tumor-specific databases. The data coordinating center is housed at the City of Hope Cancer Center in Los Angeles, California, and clinical research associates at each participating center abstract and de-identify the data for submission to a centralized, web-based database. Kulig noted that this is done under an IRB-approved protocol in every site and that most of the IRBs now have a waiver of consent unless the patient’s data are linked to specimen collection protocols. The patients included in the database have received all or most of their care at an NCCN institution, and each patient is followed longitudinally throughout the course of care (Kulig added that outside medical records are unfortunately not accessed for abstraction in this database). The NCCN main office in Fort Washington, Pennsylvania, employs project managers, quality assurance managers, statisticians, and analysts who work with the aggregate data from all of the databases and all of the sites.
There are some unique characteristics of the NCCN outcomes database, Kulig said, that differentiate it from other large datasets (e.g., the Surveillance, Epidemiology, and End Results Program [SEER]) or other institutional databases. The NCCN database has more than 300 different data elements that track the continuum of care for each patient. There is complete data on patient demographics, medical histories, family histories, and comorbidities. Detailed information about sites of metastases and biomarkers is also collected. All clinical events and interventions are collected, including diagnostics, hospital admissions, very detailed sequencing of therapies, and reasons for discontinuation of chemotherapy. Progression-free and overall survival data are also captured.
Kulig contrasted this to tumor registry data, which generally contain no information on comorbidity or diagnostic procedures and only limited data
on treatment and outcomes (limited to recurrence and survival data). She also drew a contrast to claims or billing data, which are not research-quality data to begin with; which have no information on staging, pathology, or histology; and from which treatment data can be very difficult to interpret. There is generally no biomarker information, and the outcome end points that can be derived from billing or claims data are usually in the form of resource utilization to which costs can be affixed.
NCCN has rigorous data quality assurance processes in place, starting with extensive data manager training. The system also includes online edit checking during web-based entry, programmed logic checks (e.g., flagging an entry if a prostate cancer patient is entered as female), quarterly quality assurance reports, and on-site audits.
Five NCCN sites are currently using electronic data transfer for the breast cancer database, Kulig said. This is particularly important at high-volume sites to maximize efficiencies (e.g., eliminates the need to enter the same data multiple times into various databases, such as the tumor registry, the NCCN database, or any internal databases). Electronic data transfer does need dedicated resources, including programming support to conform to the NCCN database requirements and changes. Lack of dedicated resources can lead to failed audits or poor-quality data, Kulig said. The fact that some sites are using electronic data transfer while others are not does add complexity to the system, and every proposed change by NCCN must be carefully considered for how it will affect electronic data transfer sites versus manual data entry sites.
Current Use
The primary use of the NCCN outcomes database is the annual analysis of institutional concordance to the NCCN guidelines. This is done systematically for each tumor database, comparing the care that was actually received in practice to the guideline that was in effect at the time the care was delivered. (Kulig noted that any treatment received as part of a clinical trial is considered concordant.) Each institution’s care is then benchmarked against the NCCN aggregate, and individual reports are provided to each institution for quality-improvement purposes. Individual patient summary reports can be generated from the database and are useful for viewing concordance status, treatment information, or visit history, for example.
The database is also very conducive to comparative effectiveness research, Kulig said, because it contains detailed information about the patient’s clinical
characteristics, comorbidities, and treatment. Institution-specific data are available to NCCN sites, and aggregate data are available upon request. The data are also available to non-NCCN entities for specific research queries, and Kulig noted that NCCN has provided data to pharmaceutical, biotechnology, information technology, and medical device companies.
Value of Collaborative Databases for Diverse Stakeholders
When we think of the continuum of data and how it is generated, Kulig said, controlled clinical trial data provide the strongest evidence for safety, efficacy, and even patient-reported outcomes. Such high-quality data can also provide information about the predictive and prognostic value of diagnostic testing and biomarkers.
Beyond the clinical trial, Kulig suggested, a lot of observational data generated every day in clinical practice is often underutilized or even ignored, including real-world safety and effectiveness data; patient-reported preferences and outcomes; adherence to, duration of, and reasons for discontinuation of therapy; biomarkers and diagnostic testing; and resource use and costs of care. A variety of stakeholders make use of real-world data for a variety of purposes (Box 4-1).
Personalized medicine has a particular need for real-world, real-time data. Kulig noted that lags in existing datasets, the NCCN dataset included, do not necessarily accommodate real-time analysis or cutting-edge biomarker-linked outcomes research. In addition, there are large bio-specimen repositories in numerous institutions that are not necessarily annotated or linked to clinical data.
In conclusion, Kulig said, the NCCN outcomes database is one example of a collaborative data collection system that is relevant to both clinical practice and robust clinical outcomes research. Observational, real-world data hold value for key stakeholders. The promise of personalized medicine in particular underscores the need for these types of data aggregation and exchange systems.
CASE EXAMPLE: IT INNOVATIONS FOR COMMUNITY CANCER PRACTICES
Cancer costs are rising more rapidly than inflation and other health care costs, and community cancer centers are facing more challenges than ever, said Asif Ahmad, senior vice president for information and technology
BOX 4-1
What Real-World Data Are Used by Whom?
Clinicians
• Treatment effectiveness
• Adverse events and safety
Food and Drug Administration
• Epidemiology
• Adverse events and safety
Payers
• Adverse events and safety
• Comparative effectiveness
• Resource use and costs
Patients and Caregivers
• Treatment effectiveness; real-world survival
• Adverse events and safety, symptoms and side effects, quality of life
• Costs of care
Manufacturers
• Epidemiology, current practice patterns, unmet medical need
• Treatment effectiveness versus comparators; real-world survival
• Side effects and adverse events, hospitalizations, resource use and costs
SOURCE: Kulig presentation (February 27, 2012).
services at McKesson Specialty Health. McKesson Specialty Health partners with community cancer care practices to help them manage increased competition from hospitals and clinics, declines in reimbursement, health care reform uncertainty, and rising health care costs. McKesson Specialty Health is the second-largest business unit of McKesson, which is one of the largest distributors of specialty pharmaceuticals and biologics. The company has long-term partnerships with about 3,000 oncologists, about 1,000 of whom are in the U.S. Oncology Network, and 2,200 multi-specialty practices.
Ahmad said that the foundation of McKesson Specialty Health is IT,
upon which is built the robust customer-facing technology from which analytic applications stem. The customers could be patients, pharmaceutical manufacturers, payers, or others. The focus is on developing an efficient, integrated technology suite built upon the core architecture.
The cornerstone of the McKesson system is the iKnowMed Electronic Health Record, one of the first totally hosted, web-based EHR systems, Ahmad said. iKnowMed includes charge capture functionality, safety alerts to help decrease errors, a cancer diagnosis and regimen library to aid point-of-care decision making, and clinical trial support to help increase clinical trial accruals. It integrates with other tools such as oral e-prescribing and drug inventory management. One benefit for practices is the ability to drive the workflow of the clinic from a single place. Another unique feature of the system is the support for real-time decision making. Ahmad added that all of the data feed into a common data management system that currently warehouses data for about 1.2 million patients, with 17,000 newly diagnosed cancer patients added to the McKesson database every month.
Ahmad reiterated that the customer-facing technology is all built upon the same structural data framework. For example, data in the common core framework could be used for real-time clinical alerts that could inform the clinician about a newly diagnosed patient for the purposes of clinical trial recruitment or alert the nurse to a scheduled appointment for a patient coming for the first dose of treatment. The same technology drives alerts for pharmaceutical manufacturers that may include aggregate, de-identified data regarding use of their products. The CARE, or Comprehensive Accrual REsource, tool for clinical research is also built on the same technology and facilitates identification of patients for personalized medicine trials. The same data framework is used to pull up the weekly and monthly financial reports for the practices. Rather than duplicating efforts, everything comes from one source. That data store is fed by the EHR, practice management tools, and patient-reported data from the patient portal. Ahmad noted that the technology architecture is able to use data from whatever practice management tools the practice has chosen to use.
Supporting Clinical Outcomes and Research
Current trends in health care reform, consumerism, pay for performance, and care delivery models are changing the landscape of medical practice. The old model of “patient plus prescription results in payment” is being replaced with one where demonstrated outcomes impact payment. The McKesson system allows researchers to harness clinical data to show cost-effectiveness of care alongside evidence for best possible outcomes. Another feature is the ability to provide a market intelligence report to manufacturers, which can show, for example, the penetration of a particular drug in the market compared to a competitor. The same data are integrated with the clinical trial management system, Ahmad said; this has facilitated the enrollment of more than 50,000 patients in various clinical trials over the past 8 years.
Because no defined standards are fully implemented with regard to EHRs, the data remain somewhat “dirty,” Ahmad said. To help address this, McKesson established a multidisciplinary data governance committee tasked with increasing stakeholder awareness of the importance of data quality and working collaboratively with all stakeholders to ensure the accuracy, completeness, and consistency of the clinical, administrative, and financial data that are entered into the database. The committee will consider data governance from a very comprehensive point of view, including, for example, common definitions, structure and standardization, data validation, data access and compliance with regulations, communication, prioritization, and benchmarks.
CASE EXAMPLE: SECONDARY USES OF DATA FOR COMPARATIVE EFFECTIVENESS RESEARCH
Paul Wallace, senior vice president and director of the Center for Comparative Effectiveness Research at The Lewin Group, a health care and health policy consultancy in Washington, DC, began with a brief overview of the evolution of comparative effectiveness research.
Comparative effectiveness research (CER) was defined in the Medicare Modernization Act of 2003 as “the conduct and synthesis of research comparing the benefits and harms of different interventions and strategies
to prevent, diagnose, treat and monitor health conditions in ‘real world’ settings.” Subsequently, the Agency for Healthcare Research and Quality (AHRQ) created programs to foster CER, and in 2009, the American Recovery and Reinvestment Act (ARRA) provided $1.1 billion to fund CER. The Patient Protection and Affordable Care Act of 2010 included further provisions to foster CER, including the creation of the Patient--Centered Outcomes Research Institute (PCORI). Chartered as an independent, nonprofit research organization with a sustainable funding stream, PCORI’s charge is to fund research that offers patients and caregivers the information they need to make important health care decisions.
Patient-centered outcomes research is a relatively new term, Wallace explained, and there has been some tension as to what it is and what it is not, as well as how it contrasts with personalized medicine. According to the PCORI working definition, patient-centered outcomes research
helps people and their caregivers communicate and make informed health care decisions, allowing their voices to be heard in assessing the value of health care options. This research answers patient-centered questions such as: Given my personal characteristics, conditions, and preferences, what should I expect will happen to me? What are my options and what are the potential benefits and harms of those options? What can I do to improve the outcomes that are most important to me? How can clinicians and the care delivery systems they work in help me make the best decisions about my health and health care? (PCORI, 2012).
Wallace added that from a practice perspective, patient-centered outcomes research will empower the oncologist to have more robust conversations with an individual patient about what is known about similar patients, how his or her situation compares and relates to others, and how the patient and practitioner can move forward on the basis of that information.
The draft research agenda recently released by PCORI indicates that 20 percent of funding will be allocated to accelerating patient-centered outcomes research and to methodological research, particularly for conducting observational research, Wallace said. There has been an evolution in how evidence is perceived (Table 4-1). Expert opinion based on case reports and case series is still important and credible, but it is based on chart review and clinician experience (essentially an N of 1). Evidence-based medicine is more from the perspective of population efficacy (an N of many). Data from clinical trials and systematic reviews are expressed as a mean, and subgroups of people are thereby excluded. While population efficacy is translated into a variety of practices, from performance management to
TABLE 4-1 The Evolving Evidence Perspective
| Study Type | Methods | Data Source/Organization | Perspective |
| Expert Opinion |
• Case Reports • Case Series |
• Charts • Experience |
Effects on Patients (N of 1) |
| Evidence-Based Medicine | • RCTs • Systematic Reviews • (Observation) |
• Trial Data & Databases • Meta-analysis • Reports & Series |
Population Efficacy (N of Many) |
| Comparative Effectiveness Research | • RCTs • Systematic Reviews • Observation |
• Trial Data & Databases • Meta-analysis • Large Population Databases • Reports & Series |
Population Effectiveness (N of Many) |
| Patient-Centered Outcomes Research | • RCTs • Systematic Reviews • Systematic Observation |
• Trial Data & Databases • Meta-analysis • Large Population Databases • Reports & Series • Patient-Generated Data |
Patient Effectiveness (Many N of 1s) |
NOTE: RCT = randomized clinical trial.
SOURCE: Wallace presentation (February 27, 2012).
guidelines, Wallace noted that there are pitfalls, not the least of which is that the data may not be particularly applicable to the individual patient in the room. Comparative effectiveness research uses large population databases to consider population effectiveness (also an N of many). The key difference between evidence-based medicine and CER, Wallace suggested, is that CER attempts to include those groups of people who were ineligible for and excluded from clinical trials, using observational data to complement and extend what was learned from the trial. Patient-centered outcomes research builds on all of the previously discussed dimensions, focusing on patient effectiveness (many N of 1s). In other words, what is the provider’s recommendation to the individual patient, based on as many people like that patient as can be found?
When considering “secondary use” of data, Wallace said, it is important to remember that this means the intended primary use was most likely for
a different purpose and the data were collected under different rules. As a result, secondary use of data generally involves some sort of adaptation or compromise. The challenge is to identify how this secondary use can best complement and extend the primary use of the data. Wallace outlined several approaches to secondary use.
Most secondary uses have been reactive and opportunistic, making use of any data that are already there. Wallace mentioned the Optum Natural History of Disease (NHD) Model as an example of how data can be used for secondary analysis. Using claims data, this application can answer questions such as, How many people within a population are taking a particular lipid drug and how do they differ from matched people with the same clinical situation who are not taking the same drug? While it would take about a year and a half to answer that question by going through charts, Wallace said, the NHD applications could answer within minutes, making use of big datasets that are optimally structured. Reactive secondary analysis can be useful for forming new hypotheses or for iterative testing to refine a hypothesis.
A different approach is to plan for secondary use. This may involve structured data capture, expanded common datasets outside clinical trials, or common intervention protocols. Planning for secondary analysis can help to answer questions that cannot be answered by classical experimental approaches. For example, What are the drivers from the patient side for choosing to undertake a third line of therapy? Can differences in costs and response rates for various lines of therapy be demonstrated? How do survival and costs for advanced cancer patients who opt not to have therapy compare to those who are treated?
Wallace cited a variety of ongoing efforts in secondary use of data that support CER and patient-centered outcomes research, including the FDA Sentinel system that uses data to track the safety of products on the market and the National Institute of Mental Health (NIMH) Registry of Individuals with Autism Spectrum Disorders to test hypotheses about etiology and health services use by patients and families.
The sustainability of these projects is an ongoing challenge, Wallace concluded. There is a necessary balance and overlap between funding and governance, and between data sources and data users. Surrounding this are issues regarding the use of distributed versus aggregated data, cost structure
for participation, preservation of privacy and confidentiality, and proprietary and ownership issues.
Following the presentation of case examples, a reaction panel moderated by Adam Clark, patient advocacy consultant at MedTran Health Strategies, discussed further some of the cross-cutting issues for informatics-supported cancer research and care. Panelists included Gwen Darien, director of the Pathways Project and cancer survivor, Deven McGraw, director of the Health Privacy Project at the Center for Democracy and Technology, James Cimino, chief of the NIH Laboratory for Informatics Development, and Steven Piantadosi, director of the Samuel Oschin Comprehensive Cancer Institute at Cedars-Sinai Medical Center.
Gwen Darien offered the patient’s perspective based on both her personal experience as a cancer survivor and input from her colleagues in the advocacy community. The advocacy community is not as engaged in health IT and informatics as it needs to be, she said. To create buy-in and engagement of the survivor advocacy community, that community should be engaged from the beginning, during the conceptualizing of the process and parameters of information exchange (rather than simply participating afterward). Are patients people that something is done to, she asked, or are they active partners in the formulation and creation of value? Data are an asset. What is the value of patients to health IT, and what do the patients get out of sharing their data in return? Who owns the information and what information do patients have access to?
Darien noted that a barrier to moving forward in informatics is the broad assumptions made about “the patient.” Some patients are extremely engaged, some disengage once their treatment is over, others do not want to know anything.
Building Trust: Privacy, Consent, and Ownership
Deven McGraw said that the end goal of privacy is not privacy itself, but trust. The goal is to build a trusted, accountable ecosystem for using data in ways that help individuals, communities, and populations. Privacy
rules are structured largely around tools such as patient consent and data minimization or de-identification. These tools are critically important, McGraw said, but they are not the end goal. They are tools to be used to build trust, along with other tools. It is also important to remember that consumers and patients want their data to be protected, and they want medicine and health care to be advanced. These competing interests need to be considered and balanced when developing privacy policies.
McGraw also suggested that too much time is spent focusing just on the issue of consent in lieu of addressing other important privacy protections. Consent is not the same as privacy. Consent ends up shifting the burden for protecting privacy to the patient. That said, when surveyed, people often say that they want to be asked before their data are used for research purposes. There are efforts now to obtain general consent for future research because it is not possible to define all of the potential research uses of the data being collected today, but this does not lead to a meaningful and informed consent for the patient, she said. Building trust in research requires research institutions to be mindful of the sensitivity of the data, to treat them with respect, and to make good decisions about how the data are to be used. McGraw suggested that one of the ways to rely less on consent and build trust is to improve transparency, both to the public at large and to cancer patients, about how patient data are used, the typical tools that institutions use to protect data, and oversight and accountability for those protections.
Consent as a policy issue is also important for researchers because their access to clinical data is based on informed consent. James Cimino pointed out that if the data are going to be reused in de-identified form, the original investigators are notified of reuse. This raises the issue of the ownership of the data and goes beyond the patient or the institution to the intellectual property of the investigators who are providing these data for sharing.
Workshop participant Alison Smith from C-Change asked what role additional penalties for inappropriate and irresponsible use of data played in the trust equation. McGraw expressed her concern that imposing penalties and creating the threat of legal liability can cause people to be unwilling to share data, because that is the path of least risk. Regulators need to do more than impose penalties, she said; they need to provide more guidance about how to comply with HIPAA Privacy Rule (IOM, 2009).
James Cimino highlighted several technical issues of data granularity in databases. When repositories collect data from different sources, one of the concerns is determining when the data collected are synonymous and when are they not. The level of data granularity in clinical care is not necessarily the same as the level of granularity in research data (e.g., noting the patient had pain versus reporting the severity of pain on a scale of 1 to 10 and where exactly the pain was). Capturing the context of the data is also important (e.g., was a complete blood count done because the patient had a fever, or was it part of the protocol data collection schedule?). Another technical aspect is the inclusion of genomic data. What is important to index? How can the system cope with the changing methods of how these data are being collected?
Steven Piantadosi added his perspective as a clinical trialist. Expanding on the comments by Wallace, he stressed that data that are intended and designed to address a particular therapeutic question are distinct from data that were produced for some other purpose. He urged caution when making inferences based on data that were not collected for that purpose. One example is when a safety signal emerges in a clinical trial designed to consider a therapeutic question. Even in a highly structured clinical protocol, it can be very difficult to determine whether or not the safety signal is real and actionable when the study was not designed specifically to study it. These are not new issues, and for historical perspective, Piantadosi referred participants to a 1984 article discussing the use of observational data from registries to make treatment comparisons (Green and Byar, 1984).
Kulig added that observational data are largely underutilized because of concerns from both a statistical and a clinical perspective. Wallace said that just because we do not have statistically significant data does not mean we do not make decisions. While the foundation of decision making is ideally experimental empirical data, there are many questions for which empirical data will not exist. Frameworks are needed that can help clinicians make progressively better care decisions for each individual patient, even in the absence of gold standard data. Piantadosi suggested that it is not a question of having the perfect data source, but rather a problem of bias. These databases are not unbiased sources of information on which to base definitive statements about therapeutic decisions. Mousses concurred that we should
not be making definitive statements about something that the data were not generated to address in the first place. The data can be mined, aggregated, and used to identify trends and design studies to test new hypotheses, he said. There may be bias in the data, but there may be a signal nonetheless that could guide further study.
Engaging Private Practice and Extramural Researchers
Piantadosi noted that the majority of faculty at Cedars-Sinai are in private practice (i.e., not employed directly by the hospital) and it is difficult to incentivize them appropriately with regard to informatics needs. If, as was noted earlier, 80 percent of patients are treated outside of academic centers, we have to find ways to incentivize the private practice community to use the tools and provide the data that are needed, he said.
Clark asked what would be needed for the governance or implementation of an interconnected system that would engage community practices in data exchange. One solution, Piantadosi said, is to buy the practice. Independent practices tend to see things in terms of time spent and cost, and they do not respond well to edicts from the parent institution. Practitioners may also be less computer savvy, either because of their background or the nature of the practice.
Cimino said that NIH intramural investigators are being encouraged to collaborate extramurally, and the clinical center is opening its doors to extramural investigators to bring in their patients for studies and to make use of some of the unique resources that NIH has. With regard to governance, there is currently no coordinated trans-institute effort to share data within or outside NIH, other than what individual researchers choose to do. The IT working group of an advisory council to the NIH director is considering how to better coordinate intramural and extramural work, and a report is expected in 2012.
Green, S. B., and D. P. Byar. 1984. Using observational data from registries to compare treatments: The fallacy of omnimetrics. Statistics in Medicine 3(4):361-373.
IOM (Institute of Medicine). 2009. Beyond the HIPAA privacy rule: Enhancing privacy, improving health through research. Washington, DC: The National Academies Press.
PCORI (Patient-Centered Outcomes Research Institute). 2012. Patient-centered outcomes research. http://www.pcori.org/patient-centered-outcomes-research/ (accessed April 26, 2012).




















