Overview of the Cancer Informatics Landscape
DISCUSSION POINTS HIGHLIGHTED BY INDIVIDUAL PRESENTERS
• Cancer researchers and care providers are facing an overwhelming volume of data from a multitude of sources and are hampered by the inability to merge those data or to communicate effectively across disciplines and stakeholders because of divergent standards, lack of interoperability, and other barriers.
• The most successful informatics tools will be those that integrate research and clinical data in an organized and efficient manner.
• A research information exchange system integrates data from multiple sources (extracting, transforming, harmonizing, and profiling for quality and accuracy) and then makes them available to diverse stakeholders per their queries. The information is provided based on the same data elements, but the presentation depends on who is asking for it (researchers, patients, clinicians, or administrators).
• Guiding principles for an integrated data warehouse include relevant standards for data entry, deep annotation, a good query interface, and sharing (via entry back into the database) of any new data derived from the analysis of data, specimens, or images stored in the data warehouse.
• Research data sources span the spectrum from electronic health records (EHRs) to disease registries to clinical research protocol repositories, with varying degrees of completeness, quality, and research utility. In silico research depends on having complete and valid information.
• caBIG is undergoing renovations and new informatics project review criteria are being implemented; NCI is open and receptive to communications from interested parties.
In the first session, an overview of the current status of cancer informatics was provided from the perspectives of cancer centers, cancer cooperative groups, and clinical translational researchers. Panelists also discussed the lessons that could be learned from the ongoing evolution of NCI’s caBIG.
STRUCTURED, INTEROPERABLE RESEARCH AND CLINICAL INFORMATION SYSTEMS—OR THE LACK THEREOF
Rapid advances in technology have led to a dramatic increase in the output of genomic and molecular data related to cancer biology, said Lawrence Shulman, chief medical officer and chief of the Division of General Oncology at the Dana-Farber Cancer Institute. These emerging data can inform our understanding of basic cancer biology, epidemiology, and behavior, as well as response to therapies, toxicity of therapies, and optimal care for an individual patient or cohort. However, the sheer volume of information presents significant data management and analysis challenges and is becoming overwhelming from a clinical decision-making standpoint.
To be optimally useful, data should be structured in a database, and we are still in the learning stages of how best to structure genomic and molecular data, Shulman noted. For clinical data to be useful, they should contain certain critical elements. From an oncology perspective, examples of key data elements include patient demographics; tumor type and anatomic and non-anatomic staging; treatment plan, treatment intent (e.g., curative or palliative), and actual treatment; tumor response; toxicity; patient-reported outcomes; and disease-free and overall survival. The nation is moving, albeit
slowly, toward the adoption of electronic health records (EHRs) to facilitate efficient clinical practice and decision making. However, many of the data included in EHRs are not in a structured format (i.e., are entered as free text). One must often read through the notes of clinicians, and it can be challenging to discern exactly what has happened to the patient, Shulman said.
Databases That Foster Learning
Shulman stressed that the most successful informatics tools will be those that interconnect research, clinical activities, and data in an organized and efficient manner, with as broad a database as possible. Citing a 2010 IOM workshop on evidence-driven practice in cancer care, he explained that the patient is the center of the system around which there is a cycle of aggregating information (including routinely collected real-time clinical data), analyzing that information, making new discoveries, and applying those discoveries to improve the care of individual patients (Abernethy et al., 2010; IOM, 2010) (Figure 2-1). The American Society of Clinical Oncology has recently launched CancerLinQ,1 a rapid-learning system based on this model that is being pilot-tested first for breast cancer. Shulman described an example in place at Dana-Farber, called the Synergistic Patient and Research Knowledge Systems (SPARKS) (Box 2-1; Figure 2-2). This system links clinical data (e.g., EHR, surgical, and pathology data), tissue sample information, cancer registry information, patient-reported outcomes, translational research data (e.g., gene expression), and operations data (e.g., billing, scheduling, and other visit information).
Shulman offered the “life cycle” of a gene mutation as a practical example of the value of an integrated data system. In this scenario, a basic researcher discovers a gene mutation in the laboratory, but its clinical significance is unclear. An association with a clinical syndrome is then determined. Translational research ties that gene mutation to a clinical outcome (e.g., prognosis, response to therapy). The clinical significance of the mutation is validated, and testing for the mutation may have therapeutic implications.
![]()
1 See http://www.asco.org/ASCOv2/Practice+%26+Guidelines/Quality+Care/CancerLinQ+-+Building+a+Transformation+in+Cancer+Care.
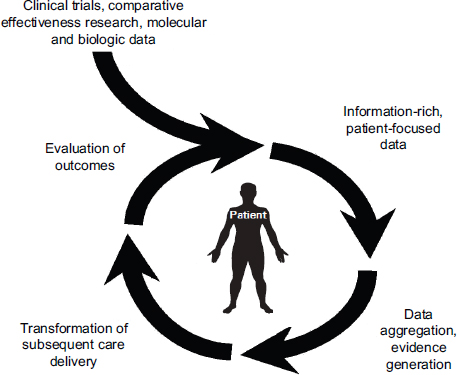
FIGURE 2-1 Rapid-learning health care system for cancer care.
SOURCE: Abernethy et al., 2010; IOM, 2010.
At Dana-Farber, researchers can query both a clinical data repository and a consented research database. A “transient data mart” houses data involved in a current query, which is then purged when the query is completed. Data are de-identified, and queries are covered under an umbrella protocol so that additional institutional review board (IRB) approval is not required. Investigators could be seeking an actionable mutation in multiple tumor types and could query, for example, the aggregate number of patients who have human epidermal growth factor receptor 2 (HER2) amplification in breast, gastric, and salivary gland tumors and have responded to trastuzumab. Queries can be very specific. For example, an investigator interested in hormone resistance might query the frequency of a particular mutation in women who have metastatic breast cancer that is estrogen receptor (ER)-positive and HER2-positive, who are between the ages of 50 and 65, and who had progressive disease while on tamoxifen. It is also possible to access identified data with IRB approval, and investigators recruiting for a
BOX 2-1
Dana-Farber Synergistic Patient and Research Knowledge Systems (SPARKS)
Vision To provide a cutting-edge, collaborative institutional informatics framework to accelerate scientific discoveries and their translation into clinical practice to enable early diagnosis, personalized treatment, cure, and prevention of cancer and related diseases.
Objective Implement policies, standards, systems, and tools that facilitate collection, integration, mining, analysis, and interpretation of biomedical data to accelerate scientific discoveries and their translation into personalized medicine and clinical practice.
Long-term goal Establish an integrated, patient-centric clinical genomic data model and systems for enabling translational research and personalized medicine.
NOTE: See also Figure 2-2.
SOURCE: Shulman presentation (February 27, 2012).
particular investigational protocol could query for actual patients who meet specific eligibility criteria.
Robust EHR Systems and Research Databases
In closing, Shulman stressed the need for robust EHR systems and robust research databases. All clinical data in EHRs should be codified or structured, he said. EHRs should include detailed data on patient demographics, tumor characteristics and staging, and treatment histories, as well as codified treatment responses and treatment resistance development and codified genomic and molecular data. Ideally, EHR systems would be interoperable and have standards for data entry. Similarly, he said there is need for robust, interoperable research databases containing structured genomic and molecular data, entered according to defined standards, and these research databases should link with relevant clinical databases. Shulman said that we are not even close to these aspirations.
Such databases often exist at the laboratory level and, in many cases,
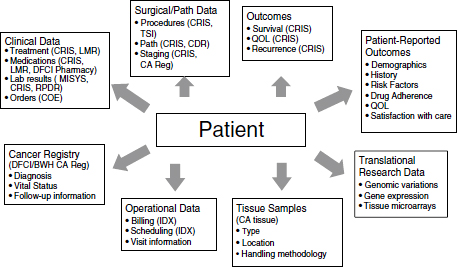
FIGURE 2-2 Dana-Farber Synergistic Patient and Research Knowledge Systems (SPARKS).
Note: BWH = Brigham and Women’s Cancer Center; CA = cancer; CDR = Clinical Data Repository; COE = computer order entry for chemotherapy and all medications; CRIS = Clinical Research Information System; DFCI = Dana-Farber Cancer Institute; IDX = IDX operating system; LMR = longitudinal electronic medical record; Path = pathological; QOL = quality of life; Reg = registry; RPDR = Research Patient Data Registry.
SOURCE: Shulman presentation (February 27, 2012).
at the institutional (e.g., cancer center) level. To be maximally effective, however, Shulman said, there is a need for database efforts at a national or even international level, incorporating data from academic centers as well as community practices (where about 80 percent of patients in the United States receive their cancer care).
CANCER CENTER INFORMATICS: CONNECTING WITH PATIENTS
To integrate new technologies into the standard of care there must be demonstrated value, explained William Dalton, president, CEO, and center director of the H. Lee Moffitt Cancer Center & Research Institute.2 A given treatment or technology, however, may not provide the same value
![]()
2 In July 2012, Dr. Dalton assumend the position of CEO for the newly formed M2Gen Personalized Medicine Institute at the Moffitt Cancer Center.
to all patients. Evidence must be generated regarding what works for which individuals or cohorts. Dalton outlined four elements of a personalized medicine approach that can lead to overall improved health care:
1. Addresses health care as a public issue and seeks to improve access, affordability, and quality of care by developing an information system to assist in making clinical decisions based on outcomes and comparative effectiveness;
2. Integrates new technologies into the standard of care in an evidence-based fashion to identify populations at risk, personalize treatment, and improve individual outcomes;
3. Provides an approach to identify the best treatment for individual patients based on clinical and biological characteristics of patients and their disease; and
4. Creates a network of health care providers, patients, and researchers who contribute and share information from individual patients to ultimately improve the care of all patients by learning from the experience of others (Dalton et al., 2010).
Improved medical care begins with data that provide information, from which we derive knowledge and develop wisdom, and we are still in the data phase of this journey, Dalton said. In implementing a research information exchange system that will serve the key stakeholders, cancers centers face technical, cultural (e.g., academic versus industry), regulatory, and financial challenges. From a technical and regulatory standpoint, data sharing raises many issues that must be addressed, for example, technical architecture, intellectual property concerns, privacy and security, and human subjects’ protections.
Another, perhaps underappreciated, technical aspect is data governance, Dalton said. How are the validity and quality of the data entered into the system ensured? Semantic interoperability or harmonization is needed; the system must serve users who are looking for the same data but in a different context or with different semantics and syntax, depending on who is asking. Multiple data sources can actually help ensure data quality, Dalton suggested.
The challenge is how to develop an integrated network information system that can manage the ever-increasing amount of information being
generated in basic, translational, and clinical research that is needed to support a personalized medicine approach. Dalton pointed out that a single information system can serve multiple stakeholders, including researchers, patients, administrators, and clinicians. Information is provided based on the same data or “truth,” but the presentation will depend on who is asking for it. As noted by Shulman, a database can be used by researchers to identify a patient cohort for data analysis or for clinical trial recruitment. Researchers might also seek data on comparative effectiveness or to do molecular profiling. The same information system, Dalton said, should allow patients to have their own personalized health record, which they can interact with and contribute to. The same platform would also be able to support evidence-based decision making by clinicians and administrators.
Dalton offered an illustration showing how an information exchange system could be designed (Figure 2-3). The goal was to develop a system
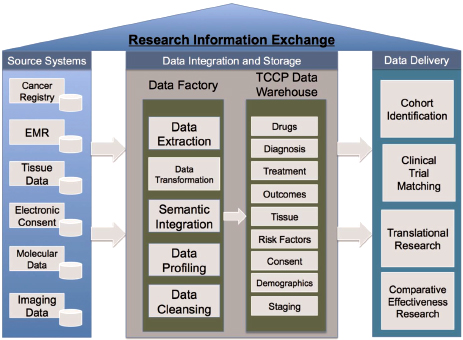
FIGURE 2-3 Example of a research information exchange system at the Moffitt Cancer Center, integrating data from multiple sources and providing them to diverse stakeholders.
NOTE: EMR = electronic medical record; TCCP = Total Cancer Care Protocol.
SOURCE: Fenstermacher et al., 2011. Reprinted with permission from The Cancer Journal.
that could identify the same patient in multiple source data systems that may use different identification numbers or ways of describing the patient. The data are loaded into a “data factory” where they are extracted, transformed, harmonized, and profiled for quality and accuracy. Data are then stored in the warehouse to be delivered to stakeholders per their queries. In response to a question about the broad integrity of the informatics enterprise, Dalton stressed the emphasis that the data factory places on ensuring the quality of data before they enter the data warehouse. In addition, multiple sources of data on the same individual allow for algorithm checks for quality.
It was noted by a participant that a challenge in obtaining patient consent for inclusion in such systems is ensuring that patients understand what they are consenting to, specifically that their data may be used for studies that are not yet defined to answer questions that have not yet been thought of. A multiphased consent process, where patients can opt in or opt out of different parts of the program, may be most appropriate, but it is a complicated and lengthy process. Dalton shared the example of a large, ongoing observational study in which patients consented to be studied throughout their lifetime, including collecting tumor samples and being recontacted should the investigators find something that might benefit them. An analysis of patient comprehension found that over time, many had forgotten that they had consented to this ongoing interaction. To help address this, patients receive a card within 2 weeks of consenting that thanks them for enrolling in the study, including a phone number to call if they have questions or do not recall consenting. A patient portal has also been developed, Dalton said, which again thanks patients for consenting and explains that they can obtain information about how their data are being used and how they can opt out at any time.
CANCER COOPERATIVE GROUP INFORMATICS: CONNECTING RESEARCHERS
Robert Comis, president and chair of the Coalition of Cancer Cooperative Groups and group chair of the Eastern Cooperative Oncology Group (ECOG), provided an overview of the NCI Clinical Trials Cooperative Group Program. As a result of an ongoing reconfiguration of the cooperative group structure, the program now comprises five major groups:
1. Alliance for Clinical Trials in Oncology—merging the Cancer and Leukemia Group B (CALGB), the North Central Cancer Treatment
Group (NCCTG), and the American College of Surgeons Oncology Group (ACOSOG);
2. Children’s Oncology Group;
3. ECOG-ACRIN Cancer Research Group—merging the American College of Radiology Imaging Network (ACRIN) and ECOG;
4. NRG Oncology Group—merging the National Surgical Adjuvant Breast and Bowel Project (NSABP), the Radiation Therapy Oncology Group (RTOG), and the Gynecologic Oncology Group; and
5. SWOG (formerly the Southwest Oncology Group).
Comis explained that the cooperative group environment includes academic centers, large community practices, smaller community practices, and biomedical researchers spanning the physical sciences through the clinical sciences. It is geographically dispersed and now includes international sites.
Historically, cooperative group clinical trial results have been crucial to setting the standards of cancer care. The primary mode of data collection has been case report forms, which are stored in individual group databases. There are some ad hoc interfaces for reporting and local information technology systems for managing tissue samples.
In 1999, NCI established the Cancer Trials Support Unit (CTSU), which, Comis said, now serves around 1,700 sites and thousands of investigators, supporting patient enrollment and randomization, data collection using common data elements, and regulatory and administrative activities.
Informatics Tools Used by the NCI Cooperative Group Program
Comis described the development and implementation of the Medidata Rave Clinical Data Management System software now used by the Cooperative Group Program. In 2005, the cooperative groups recognized that there was a need for a unified data system across the program. Specifications were developed; a request for proposals was sent to several vendors; and in 2008, a contract was awarded to Medidata. The unsuccessful vendors then filed formal protests, stalling the start of the contracted work. Following resolution of the disputes, the program was officially initiated in April 2011, resulting in Rave, a web-based system for capturing, managing, and reporting clinical research data that enables the user to record patient information using forms customized per study (visit, lab, and adverse event data). The program is now managed by the NCI Cancer Therapy and Evaluation Program (CTEP), with the assistance of the CTSU and Medidata, and
involves all of the cooperative groups. Hundreds of group representatives are currently engaged in classroom, webinar, and e-learning training on how to use the Rave system.
Another effort Comis described focuses on coordinating the tissue banking activities across cooperative groups. The Group Banking Committee, sponsored by NCI, is developing processes and standards for a national groupwide tissue bank virtual repository. Comis noted that each tissue bank has developed its own biospecimen information management system. Some systems integrate multiple banks, and some systems are integrated with institutional information technology (IT) systems. In addition, some groups have specimen tracking systems, which connect data entry at clinical sites with trial operations systems and bank inventory systems. The goal is to connect the various group IT structures to a single database that can be queried and is available to all researchers throughout the country.
Opportunities for an Innovative Informatics Structure
Mitchell Schnall, group chair of ACRIN, described some of the opportunities for cooperative groups in building an innovative informatics structure. Cooperative groups are a rich source of diverse biosamples and images that are associated with structured clinical information, often with long-term follow-up. In addition, there is a large array of other information, ranging from gross medical and histological images to molecular data profiling genes and proteins, that needs to be integrated with the clinical information.
The ECOG-ACRIN vision, Schnall explained, is to generate an integrated data warehouse incorporating the individual case report form, such as that from the Medidata Rave system, with imaging data, laboratory data, tissue and specimen repository inventory, digitized pathology, and -omics data (e.g., genomics, metabolomics, proteomics) as well as patient-reported outcomes and claims data.
Schnall offered several guiding principles for moving forward with such a system. First, it will be necessary for the cooperative groups to embrace relevant standards. Deep annotation should be encouraged (e.g., spatial annotation), a good query interface is important, and any new data that are derived from analysis of ECOG-ACRIN specimens or images should be entered into the data warehouse to further the value of the warehouse as a community resource. With regard to deep annotation, Schnall said that cancer control moves along a pathway from prevention to detection and characterization
and then through treatment, response assessment, adaptation of therapy, and surveillance. We need to understand the data in terms of where they were derived along this disease pathway. In addition, data can be structured relative to patient level, disease level, lesion level, and even sublesion level.
One of the tools ACRIN could contribute to an integrated data warehouse, for example, is TRIAD, a standards-based server and database that was developed to facilitate the exchange of imaging data. Currently in use at more than 200 sites, TRIAD is compliant with Good Clinical Practice (GCP) and houses image data from more than 100,000 cases, which are integrated with clinical data from the Medidata Rave system.
One concern with current image repositories, Schnall noted, is that information defining a specific location on the image is not retained after the analysis is finished. There is variability within a single tumor or among multiple lesions in a single patient, and such specificity is needed to be able to track lesions over time and over modalities and to tie pathology data directly to a specific lesion. A standard that ACRIN is embracing is called Annotation and Image Markup (AIM), which allows reviewers to set up a region of interest in an image for which they can add metadata describing that region of interest. This is very valuable for indexing specific lesions, Schnall said. Another goal is to be able to link a specific pathological or histological section with the location in the anatomic image that it came from. This goes beyond simply saying that a particular bit of data came from a patient that had breast cancer, to knowing what part of the histological specimen the molecular data came from and what specific part of the tumor the pathology sample came from.
The ECOG-ACRIN goal for the future is to have a central operations “cloud” linked to the administration and operations functions and the data sources. Interested stakeholder communities (e.g., clinical sites, the scientific community, scientific programs, patients, NCI) would then be able to interface with that cloud.
CLINICAL TRANSLATIONAL RESEARCH INFORMATICS: CONNECTING THE STEPS OF THE RESEARCH PROCESS
Bradley Pollock is chair of the Department of Epidemiology and Biostatistics at the University of Texas Health Science Center at San Antonio. He is also chair-elect of the Biostatistics, Epidemiology, and Research Design Committee of a national consortium funded by NIH through a Clinical & Translational Science Award (CTSA). He outlined the steps in
the research process for clinical translational studies as follows: hypothesis formation, study design and planning, data acquisition, statistical modeling and analysis, drawing valid inferences, and translation. While data are the critical basic building blocks, the development of evidence-based practice guidelines is driven by the entire research process.
Hypothesis Driven Versus Hypothesis Generating
Traditionally, the first step of the research process is to develop the hypothesis for which one will then design an experiment and collect data. Now, research is experiencing a paradigm shift as a result of the ever-increasing generation and availability of observational data Pollock said. We now have data, but remain in search of hypotheses. While hypothesis-generating work is important, he noted that the most novel oncology discoveries have been made using the traditional hypothesis-driven research framework.
In clinical research, attention to study design is extremely important. Good design leads to efficient use of data, and study design can have profound implications for validity. As a testament to the importance of study design for obtaining meaningful results, Pollock noted that the New England Journal of Medicine now requests full protocols for all clinical trials.
The randomized, controlled trial (RCT) is the gold standard for clinical studies; however, an RCT is not applicable or feasible in all situations. In designing observational studies, Pollock suggested that there may be more methodological hurdles to overcome than for RCTs. He cited a recent report of how the inclusion of incident versus prevalent cases in an observational study of postmenopausal hormone replacement therapy affected the results (Danaei et al., 2012). There have been concerns about the discordance between randomized versus observational studies of the effects of hormone replacement therapy on cardiovascular disease in women. In their meta-analysis, the researchers found that exclusion of prevalent users of hormone replacement therapy decreased the discrepancies between observational and randomized studies. Pollock questioned whether nuances such as prevalent versus incident use could reliably be discerned from EHRs when conducting observational studies.
In designing a research study, one generally begins by combing the literature.
Unfortunately, the value of the literature for informing study design is negatively affected by publication bias and the lag time between study findings and their final publication. The trial registry, ClinicalTrials.gov, is a useful resource but is limited to clinical trials, and entries do not provide a level of detail, rigor, or standardization necessary for scientific analysis. There is a real need, Pollock said, for accessible meta-study data for all types of study designs (not just clinical trials).
In this regard, Pollock drew attention to the Human Studies Database Project, a database of past and ongoing human studies, both interventional and observational. The primary project participants are CTSA institutions. Pollock said that the goal is to enable computational reuse of human studies data for activities such as systematic reviews, planning future studies, scientific portfolio analysis, and research networking. A subcomponent of the project is the Ontology of Clinical Research (OCRe), focused on developing an ontology to deal with issues of study design, interventions and exposures, participants, outcomes, and statistical analysis.
Informatics Challenges for Translational Research
Studies Using Existing, Non-Research Data
Research data sources span the spectrum from EHRs to disease registries to clinical research protocol repositories, with varying degrees of completeness, quality, and research utility. Pollock concurred with Shulman regarding the need for structured data elements. In EHRs, for example, useful information about drug exposure would include dose, schedule, intensity, area under the curve, dose modifications, reasons for stopping a drug, and patient pharmacokinetics, pharmacodynamics, and pharmacogenomic characteristics. These data are not generally included in the EHR, however. The major challenge when using existing information, Pollock said, is the inability to go back and fill in the missing data needed for a particular research investigation. Another concern with mining existing data is the potential for systematic biases in large clinical data repositories. More data is not necessarily better, Pollock noted, and biases can be amplified. In silico research depends on having complete and valid information.
Lack of Harmonization
From the perspective of a clinical translational researcher, Pollock concurred with the challenges highlighted by the cancer centers and cooperative
groups. Lack of harmonization is a key concern, and he noted that at his own institution, as in many academic health centers, there is more than one EHR system in place. Clinical data systems and research data systems do not routinely interoperate. Another issue for clinical research is the choice of clinical trial management system (CTMS) and whether to use a commercially available system or an open source system. There are sustainability and cost issues to consider. Open source is technically free, but not without cost because a lot more development time goes into implementing an open source platform; however, it may be more sustainable when moving forward.
Regulatory Barriers
There are also regulatory barriers for public use datasets. As chair of the advisory committee for the Texas Cancer Registry, Pollock expressed concerns about the heterogeneity of across-state permissions to combine data from multiple state cancer registries (in some cases, four levels of approval are required before researchers can utilize the data). There are also within-state restrictions; for example, until very recently, linking hospital discharge data and Texas Cancer Registry data was not permitted.
Tools, Technology, and Big Datasets
Imaging data are highly dimensional, and technical advances, such as the ability to collect real-time functional imaging data, further increase data dimensionality. There also has been an explosion in -omics data and analysis tools. Decision support tools are available, but they require large-scale validation and constant updating. Limitations to data storage and networking are also a major issue. In one genome sequencing laboratory Pollock had visited, for example, it was possible to keep active datasets on the server for only about 1 week, after which the data had to be pulled off to create space for new data.
Despite the challenges, big datasets can facilitate hypothesis generation and study planning. Data can be used to assess the feasibility of a study. Big datasets could also lower the cost of conducting clinical translational research, Pollock said, by offering more precollected data, more automation, and more interconnectivity.
Moving Clinical Translational Informatics Forward
Pollock reiterated that research is a process, and more data do not necessarily lead to more discovery. He offered several suggestions to help evolve the clinical translational research process:
• Bring clinical data up to the same standards as high-quality research data.
• Devise statistical methodologies and study designs for use with clinical data.
• Develop better data mining and filtering approaches to sort through massive datasets.
• Connect genomic and molecular data with clinical data.
• Structure clinical data appropriately to support research.
• Ensure that these processes are guided in a way that is compatible with a research framework.
caBIG—THE VISION AND THE REALITY
Daniel Masys, affiliate professor of biomedical and health informatics at the University of Washington, Seattle, and chair of the new caBIG oversight subcommittee of the NCI Board of Scientific Advisors, shared the history and vision of caBIG and some of the lessons learned since its launch in 2004. The Cancer Biomedical Informatics Grid was launched by NCI to help address the growing problem of the overwhelming volume of data from a multitude of sources and the inability to merge those data or to communicate effectively across disciplines and stakeholders because of divergent standards and terms and other barriers.
To define the priority areas for caBIG, a very extensive market determination exercise was undertaken to define the unmet needs of the NCI-supported cancer centers (Box 2-2). As a result, caBIG was envisioned as a common, widely distributed infrastructure that would permit the research community to focus on innovation (rather than on the details of managing information), with the intent that raw and published cancer research data would be available for data mining and integration into reanalyses and meta-analyses. It would be built on shared vocabulary, data elements, and data models that would facilitate information exchange and would have a collection of interoperable applications and tools developed with common standards.
• Clinical data management tools
• Distributed data sharing/analysis tools
• Translational research tools
• Access to data
• Tissue and pathology tools
• Cancer center integration and program management
• Common data elements and standards
• Meta-data analysis
• Shared vocabulary and ontology tools and databases
• Statistical data analysis tools
• Visualization and imaging tools
• Proteomics
• Microarray and gene expression tools
• Licensing and intellectual property issues
• Staff resources
• High-performance computing
• Integration and interoperability
SOURCE: Masys presentation (February 27, 2012).
In 2010, NCI director Harold Varmus called for a high-level review of caBIG. In its report released in March 2011, the working group concluded that the need for caBIG is greater now then when it was conceived (NCI, 2011). In addition, there was very strong community support from cancer centers for the original caBIG vision and goals of interoperability and standards-based exchange of data. The working group also found, however, that the many successes of the program have been offset by several problems and that the overall impact of caBIG in transforming cancer research had not been commensurate with the level of the investment (about $300 million).
The report highlighted findings in three main areas:
1. creation and management of standards for data exchange, and support for community-based software already in place;
2. impact and track record of caBIG initiatives and tools with regard to life science or integrative cancer research, clinical data management, infrastructure, and community engagement; and
3. program administration, contracts management, and budget.
The working group concluded that the greatest impact of caBIG thus far had been in the first area. The review found that the program had been very effective in catalyzing progress in the development of community-driven standards for data exchange and interoperability; development, maintenance, enhancement, and dissemination of tools developed by academic researchers; and community dialogue on the interoperability of clinical and research software tools.
The group found that the main problems with the caBIG approach that limited its uptake and impact included a “cart-before-the-horse grand vision”; a technology-centric approach to data sharing; unfocused expansion; a one-size-fits-all architectural approach; an unsustainable business model for both NCI and users; and a lack of independent scientific oversight.
As a result, the working group issued five immediate tactical recommendations. They are, as summarized by Masys:
1. Institute an immediate moratorium on all ongoing internal and commercial contractor-based software development projects while initiating a mitigation plan to lessen the impact of this moratorium on the cancer research community.
2. Institute a 1-year moratorium on new projects, contracts, and subcontracts by caBIG.
3. Provide a 1-year extension on current caBIG-supported academic efforts for development, dissemination, and maintenance of new and existing community-developed software tools.
4. Establish an independent oversight committee, representing academic, industrial, and government (NCI, National Institutes of Health [NIH]) perspectives to review planned initiatives for scientific merit and to recommend effective transition options for current users of caBIG tools.
5. Conduct a thorough audit of all aspects of the caBIG budget and expenditures.
The independent oversight committee that was called for in recommendation 4 (chaired by Masys) met for first time in July 2011. To begin to address the criticisms in the March report, the committee developed review
criteria for informatics projects (similar to the evaluation criteria used by study sections), outlining how it would assess each of the ongoing caBIG activities, Masys noted (Box 2-3).
Looking Forward: A Three-Step Approach to Success in Informatics Innovation
In response to a request from the NCI director, Masys consulted with other experts to try to define the “recipe for success,” that is, are there comparable programs that have been successful and what can we learn from them? Masys found that two applications had “gone viral”: the Research Electronic Data Capture (REDCap) developed through a CTSA, and the Informatics for Integrating Biology and the Bedside (i2b2), developed with funding from NIH’s National Center for Biomedical Computing. In considering how these applications were different from caBIG, Masys and colleagues drafted the following general steps to success in informatics innovation.
1. Do not repeat the mistakes of the past.
• Do not try to solve all clinical and translational research information technology problems in one framework.
• Do not worship standards over functionality.
• Do not try to have enterprise software adopted by fiat from above.
• Do not try to buy adoption software products, because for the sponsor, those costs grow ever larger.
• Recognize that organizations that cannot afford ongoing staffing and help desk functions for software should not be expected to adopt software even if it is free or provides some income to the adopter. The acquisition cost is dwarfed by the support, maintenance, and integration costs.
2. Understand the basic truth about IT complexity.
Increased functionality that is built at the expense of increased complexity is always at risk of
• delays in development,
• inability of local implementers and users to understand what has been built and how to use it, and
• being overtaken by other approaches that have a better price-to-performance ratio (e.g., grid computing versus web services).
BOX 2-3
NCI Informatics Project Review Criteria
1. Does the activity, application, or resource meet a well--articulated and attainable need of basic, translational, or clinical research, or cancer care (i.e., is there a “driving biological or clinical project” and are the intended users members of the project team)?
2. How will success or failure be evaluated? Analogous to stopping rules for clinical protocols, what will be the stopping rules for ending the project if it either fails to meet its technical objectives or fails to be adopted even if technically successful?
3. Will the activity, resource, or application, if successful, make some objectively measurable incremental progress toward an overall vision of interoperability of data and systems? Will it enable data sharing and make use of and/or enhance open international standards for research?
4. Is the activity, resource, or application designed to anticipate change in a rapidly expanding knowledge base of science and practice? Flexibility and generalizability are important characteristics for longevity in an era of agile science.
5. Is the intended deliverable of the project achievable in the time frame and budget proposed?
6. Will the output of the project be broadly implementable by organizations of varying size and sophistication? Will it be used broadly by organizations and institutions outside of NCI cancer centers (e.g., other NIH centers or academic research organizations)?
7. Is there a documented plan for long-term maintenance, enhancement, and fiscal sustainability of the activity, application, or resource and its user base?
8. What is the user base and has there been a stakeholder assessment to ensure that the activity, application, or resource will indeed meet a currently unmet need or a reasonably anticipated future need?
9. Is the project generalizable and likely to create value or address broad needs across the community of cancer centers and investigators? Alternatively, would this activity, resource, or application be perceived as a “pet project” of an “in” group?
10. Does the activity, resource, or application have enough market value to gain adoption without incentives? Or if financial or policy incentives are required, are they justified?
SOURCE: Masys presentation (February 27, 2012).
3. Observe the informatics research and development “do’s.”
• Solve one significant challenge at a time.
• Use small, nimble development teams led by domain experts.
• Keep development-to-implementation intervals short.
• Deploy software that can solve at least one problem that users or adopters care about within 12 months of adoption.
• Demonstrate success first with a smaller group of the most advanced sites and then let others follow.
• Create software that makes adoption of standards easier (not harder) than nonstandardized alternatives.
• Let the market prioritize and vet the standards.
• Invest in simple interfaces between applications, not architectures.
• Make interested health care organizations demonstrate willingness to invest their own assets and time for enterprise software.
• Allow intraorganizational and interorganizational needs and technologies to diverge as needed, to maximize productivity.
With regard to increasing the probability of successful adoption of informatics innovation in cancer research specifically, Masys recommended focusing data sharing efforts (both the standards for sharing and the applications to do it) on those data for which there is a preexisting motivation to share. Areas in which researchers really need one another’s data and scientific problems that simply cannot be solved within one laboratory or institution are prime areas on which to focus sharing efforts (e.g., those studying rare alleles who are trying to assemble a cohort of interest for a study). Cooperative groups are a good example of this, he noted. The current challenges are less technological and more policy oriented (e.g., IRB issues, privacy concerns).
Increasing the uptake of innovative informatics in cancer care is a more difficult task, given the low penetrance of EHRs in U.S. health care and current economic pressures. Therefore, Masys advised aligning NCI efforts with the EHR adoption incentives of the Office of the National Coordinator for Health Information Technology (discussed further by Mostashari in Chapter 5). A plausible transition goal for NCI, he suggested, would be to make it easier for every oncology practice in America to care for a patient on a clinical trial protocol. Building “public libraries” of decision support tools to guide providers and patients through clinical protocols would be an important contribution in this regard.
Community Participation in Moving Informatics Forward
During the discussion, there was much interest in involving patients as well as the research community in efforts to advance cancer informatics. Questions remain regarding how best to interact with NCI on caBIG activities, the potential role of publicly developed apps, open source software, the development of institutional systems, and the importance of patient-reported outcomes data.
NCI is very open and receptive to communications from interested parties, Masys said. George Komatsoulis, interim director for the Center for Biomedical Informatics and Information Technology at NCI, added that caBIG is continuing to move forward, and there are “workspaces” in integrative basic biology, clinical trials, data standards, imaging, biospecimens, and other areas where individuals can bring their ideas to the attention of the scientific advisory group and the caBIG program staff.
Lynn Etheredge of George Washington University asked if a national apps strategy for cancer informatics would be appropriate, that is, whether individuals could begin to develop the needed functionalities. Masys noted that apps, as they are commonly understood, tend to be fairly small-scale computer programs. They have value as lightweight applications that handle very common tasks on a personal level, often on a personal device such as a smartphone or tablet. However, their level of complexity is usually around two orders of magnitude less than enterprise-level software (e.g., that used by researchers for regular data inputs required for supporting sponsors), with lower levels of data manipulation, storage, and security. Enterprise-level software requires a more in-depth approach to implementation and, in health care and clinical research, may also involve workflow modification.
Steven Piantadosi asked about open source software relative to caBIG. Masys responded that open source software is part of the caBIG infrastructure: i2b2 is open source, while REDCap is an academic consortium model where a limited number of people have the ability to contribute to the code base. Masys cautioned that in a full open source software model (where anybody in the community can contribute to source), modules can be introduced that have unintended transitive consequences (i.e., interfere with other functions). As such, some open source models use quality control mechanisms such as curators or stewards.
Mendelsohn noted that many organizations are trying to develop their own internal standardized record of clinical trials and asked whether these organizations should continue to spend scarce resources, time, and effort
developing these independent information systems in light of the national effort.
Masys responded that any major cancer center is going to have two overlapping spheres of development, one that is outwardly focused and one that is local and specific to the organization. There will then be transition applications that facilitate sharing sets of data by formatting them and ensuring that they meet the standard for external collaborations. The cooperative groups, for example, are well served by tools such as Medidata Rave, which, while not perfect, is implemented on a large scale and provides a conceptual basis for sharing information, he said.
Mark Gorman, cancer survivor and patient advocate, expressed support for the patient-centered elements of the information systems discussed, for example, those designed to collect patient-reported outcome information. There is also great potential for informatics to support patients in their decision making and in managing their own care. Dalton noted that patient portals are very popular, with more than 70 percent of new cancer patients at Moffitt creating their portal before their first visit. Shulman added that the power of the systems discussed is the multiple sources of complementary data obtained by different methodologies, and this includes the patient portals being implemented by many cancer centers.
Abernethy, A. P., L. M. Etheredge, P. A. Ganz, P. Wallace, R. R. German, C. Neti, P. B. Bach, and S. B. Murphy. 2010. Rapid-learning system for cancer care. Journal of Clinical Oncology 28(27):4268-4274.
Dalton, W. S., D. M. Sullivan, T. J. Yeatman, and D. A. Fenstermacher. 2010. The 2010 Health Care Reform Act: A potential opportunity to advance cancer research by taking cancer personally. Clinical Cancer Research 16(24):5987-5996.
Danaei, G., M. Tavakkoli, and M. A. Hernán. 2012. Bias in observational studies of prevalent users: Lessons for comparative effectiveness research from a meta-analysis of statins. American Journal of Epidemiology 175(4):250-262.
Fenstermacher, D. A., R. M. Wenham, D. E. Rollison, and W. S. Dalton. 2011. Implementing personalized medicine in a cancer center. Cancer Journal 17(6):528-536.
IOM (Institute of Medicine). 2010. A foundation for evidence-driven practice: A rapid learning system for cancer care: Workshop summary. Washington, DC: The National Academies Press.
NCI (National Cancer Institute). 2011. An assessment of the impact of the NCI Cancer Biomedical Informatics Grid (caBIG®). http://deainfo.nci.nih.gov/advisory/bsa/bsa0311/caBIGfinalReport.pdf (accessed July 11, 2012).
























