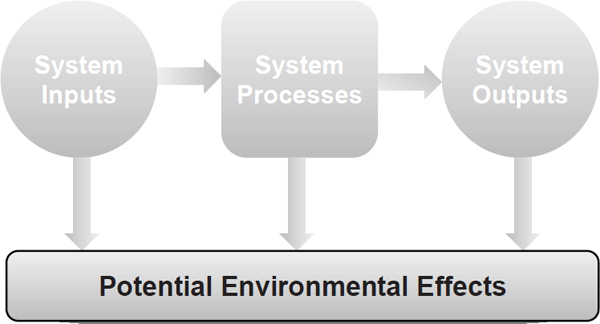
As with production and use of any fuels, aspects of biofuel production and use have benefits and adverse effects. This chapter discusses potential environmental effects from the production and use of algal biofuels, the potential influence of perceived or actual impacts on societal acceptance, and some of the health impacts potentially emanating from the specific environmental effects. Potential environmental effects discussed in this chapter include those resulting from land-use changes, water quality, net greenhousegas (GHG) emissions, air quality, biodiversity, waste generation, and effects from genetically engineered algae (with an emphasis on new or enhanced traits).
Where possible, this chapter discusses the potential for algal biofuels to improve aspects of sustainability compared to petroleum-based fuels and other biofuels and the potential for mitigating negative effects along the life cycle of algal biofuel. Environmental indicators of sustainability and data to be collected to assess sustainability are suggested. In some environments and biofuel management systems, metrics for assessing environmental performance are easy to measure and adequate baseline data are available, but that is not the case in all systems.
A number of potential environmental concerns are evident, and if the concerns are not addressed they could become significant risks under large-scale deployment. As in any other industrial or agricultural enterprise, once they are recognized, such risks can be managed by standards or regulations so that industry is required to reduce effects to acceptable levels. For the sake of comprehensiveness, a number of potential environmental risks are mentioned in this chapter, but some are less likely to occur than others. Some of the environmental risks might require exploratory assessment and subsequent monitoring to ensure that they do not become sustainability concerns if algal biofuel production is scaled up.
Producing algal biofuels could improve or harm water quality depending on the resource input and management used in algae cultivation, weather events, integrity of infrastructure, and processing of spent water. Water-quality concerns associated with commercial-scale production of algal biofuels, if sufficient culture waters are released to natural environments, include eutrophication of waters, contamination of groundwater, and salinization of water sources. Potential water-quality benefits are reduced runoff of herbicides and insecticides compared to corn-grain ethanol or soybean-based biodiesel because of their reduced use, and reduced eutrophication if there are no releases of culture water or if algae are used as a means to remove nutrients from municipal waste, confined animal feeding operations, and other liquid wastes. Water-quality effects will depend on the nutrient content of the algal culture medium; whether feedstock production systems are sealed, artificially lined, or clay lined; and the likelihood of extreme precipitation events. Leakage of culture fluid to groundwater or surface water could occur if the integrity of the pond or trough system is compromised, if flooding occurs, or if spills occur during transfers of fluid during process stages or waste removal, but most of these events could be avoided with proper management.
5.1.1 Releases of Culture and Process Water
As discussed in Chapter 4, the water for algae cultivation is likely to be reclaimed and reused to reduce the water requirement and consumptive water use. The liquid effluent also can be recycled from anaerobic digestion of lipid-extracted algae to produce biogas (Davis et al., 2011). If harvest water is to be released instead of recycled, it or effluent from anaerobic digestion would contain nitrogen (N) and phosphorus (P), the concentrations of which depend on the nitrogen and phosphorus taken up by the harvested algal biomass (Sturm and Lamer, 2011). Released waters could be more saline than receiving waters, particularly if water from saline aquifers is used for algae cultivation. Such point-source discharge will be regulated by the Clean Water Act, and a National Pollutant Discharge Elimination System permit would have to be obtained to operate the algae cultivation facilities (EPA, 2011a). However, permit violation has been observed in some biofuel refineries
that use terrestrial crops as feedstock (Beeman, 2007; Smith, 2008; EPA, 2009b; Buntjer, 2010; Meersman, 2010; O’Sullivan, 2010). Regulation and compliance assurance would address concerns about release of harvest water.
The potential for accidental release of cultivation water exists; for example, clay or plastic liners could be breached through normal weathering or from extreme weather events, some of which are predictable. High precipitation or winds could lead to overtopping of ponds or above-grade raceways. In those cases, the entire contents of algal cultures could be lost to surface runoff and leaching to surface water or groundwater. Siting in areas prone to tornadoes, hurricanes, or earthquakes would increase the likelihood of accidental releases. However, producers are likely to take preventive measures when extreme weather events are forecasted, and they would put effort into preventing accidental releases of cultivation water because such events could adversely affect their profit margin.
5.1.2 Eutrophication
5.1.2.1 Potential Environmental Effects
Large-scale algae cultivation requires the provision of large quantities of nutrients, especially nitrogen and phosphorus, to ensure high yield (see section Nutrients in Chapter 4). Even where nitrogen and phosphorus are not in oversupply, the total nutrient concentrations in algal biomass will be high. Although accidental release of cultivation water into surface water and soil is unlikely, such an event could lead to eutrophication of downstream freshwater and marine ecosystems, depending on the proximity of algal ponds to surface and groundwater sources.
Eutrophication occurs when a body of water receives high concentrations of inorganic nutrients, particularly nitrogen and phosphorus, stimulating algal growth and resulting in excessive algal biomass. As the algae die off and decompose, high levels of organic matter and the decomposition processes deplete oxygen in the water and result in anoxic conditions (Smith, 2003; Breitburg et al., 2009; Rabalais et al., 2009; Smith and Schindler, 2009). In some cases, eutrophication-induced changes could be difficult or impossible to reverse if alternative stable states can occur in the affected ecosystem (Scheffer et al., 2001; Carpenter, 2005).
Eutrophication effects have been well studied, and they depend on the nutrient loadings to the receiving waters and the volume and residence time of water of these systems (Smith et al., 1999; Smith, 2003). High nutrient loading could lead to anoxia in the deep cool portion of lakes or in hypoxia in the receiving water bodies. Potential biotic effects of eutrophication include changes in algal density and in the structure and biomass of the broader ecological community (Scheffer et al., 1997; Reynolds et al., 2002; Smayda and Reynolds, 2003). Fish yield is affected by phytoplankton1 biomass and by the nutrient ratios in the edibility of phytoplankton (Oglesby, 1977; Bachmann et al., 1996).
Nutrient levels play a key role in determining the productivity and structure of the primary producing community in estuaries and coastal marine waters (Deegan et al., 2002; Smith, 2006) and by extension, the productivity and structure of higher trophic levels. Nutrient-enriched shallow marine systems tend to have a reduced seagrass community (Burkholder et al., 1992; Hauxwell et al., 2003) because elevated nitrogen concentrations and loadings adversely affect seagrass (Efroymson et al., 2007 and references cited therein).
_______________
1 A collection of microscopic photosynthetic organisms that float or drift in fresh water or sea water.
In high-nitrate environments, seagrasses can be shaded by epiphytic algae and macroalgae (Drake et al., 2003) or sometimes by phytoplankton blooms (Nixon et al., 2001). Seagrasses affect the entire estuarine food web because they stabilize sediments; serve as habitats and temporary nurseries for fish and shellfish; are sources of food for fish, waterfowl, benthic invertebrates, or manatees; and provide refuges from predation. Eutrophication and other nutrient-related effects could be a concern for cultivation of microalgae or macroalgae in large suspended offshore enclosures (for example, Honkanen and Helminen, 2000).
Eutrophication also has implications for social acceptability (Codd, 2000), for example, because of eutrophication-related aesthetic concerns (Grant, 2010), and aesthetics can affect the recreational value of water bodies. It is unknown whether rare releases of culture water or the physical appearance of open ponds for algae cultivation could have negative effects on the social acceptability of algal biofuels.
5.1.2.2 Opportunities for Mitigation
Quantifying water losses from raceways, ponds, or photobioreactors would indicate whether repairs of small leaks are necessary. These culture systems can be designed and tested to withstand natural disasters that are possible during the lifetime of the infrastructure. In coastal locations, for example, facility and infrastructure designs would need to consider the probabilities that hurricane winds and water surges could reach the algae cultivation site (Guikema, 2009). Mitigation plans for accidental releases would be desirable. Open-pond algae cultivation also can be sited in locations that are not prone to hurricanes or away from lakes and streams. With respect to harvest water, engineering solutions can maximize recycling.
5.1.3 Waterborne Toxicants
5.1.3.1 Potential Environmental Effects
Some compounds present in algal ponds or photobioreactors could be toxic to humans or other organisms depending on exposure levels. Herbicides often are added to open systems to prevent growth of macrophytes and for selective control of algae (NALMS, 2004), but their application likely would be regulated as in the case of agriculture. If wastewater or oil well-produced water (Shpiner et al., 2009) is used as a water source for algae cultivation, heavy metals could be present. Wastewater could include industrial effluent (Chinnasamy et al., 2010) and municipal wastewater that has undergone various levels of treatment (Wang et al., 2010). The composition and amount of toxicants vary by the type of wastewater. Produced water (water contained in oil and gas reservoirs that is produced in conjunction with the fossil fuel) may contain high levels of organic compounds, oil and grease, boron, and ammonia (NH3) (Drewes et al., 2009). Many algal species including cyanobacteria, diatoms, and chlorophytes can bioconcentrate heavy metals (Watras and Bloom, 1992; Vymazal, 1995; Mathews and Fisher, 2008). Mercury could be introduced into feedstock production waters if unscrubbed flue gas from coal-fired power plants is used as a carbon dioxide (CO2) source (O’Dowd et al., 2006). Therefore, potential risks from using each type of produced water need to be identified so that adequate containment and mitigation measures can be implemented in cultivation and processing.
Waterborne toxicants (toxic substances made or introduced into the environment anthropogenically, not including algal toxins) potentially pose risk to humans or other
animals if exposures occur. Occupational exposures could be significant, especially during the harvesting phase. Thus, monitoring of toxic compounds in the culture media is important. Potential toxicity exposure to animals through drinking is discussed in the section on terrestrial biodiversity. The release of culture waters to natural environments could pose other risks to animal consumers. Toxic concentrations and doses for various chemicals are available in the Environmental Protection Agency (EPA) Integrated Risk Information System database for humans (EPA, 2012), in Suter and Tsao (1996) for aquatic biota, in Sample et al. (1996) for terrestrial wildlife, and in other government and independent compilations. Cultivation of algae in wastewater may require special handling and means of containment. Monitoring for the presence of toxicants or pathogens might be necessary to ensure the quality of the culture water.
5.1.3.2 Opportunities for Mitigation
Monitoring of metals and other compounds in water sources, nutrient sources, and culture media in demonstration facilities would provide information about whether waterborne toxicants pose a significant concern. If so, technical solutions for removing waterborne toxicants would be needed to prevent occupational and ecological exposures. Mercury is removed from flue gas in some configurations of coal-fired electric-generating units (EPA, 2010). However, mercury removal is ineffective for certain types of coal and plant configurations (NETL, 2011). Contaminants in flue gas could place another constraint on the type of coal-fired electricity facilities that would be suitable for providing CO2 for algae cultivation (see sections Estimated Land Requirements and Estimated Nutrient Requirements in Chapter 4).
5.1.4 Groundwater Pollution
5.1.4.1 Potential Environmental Effects
Open ponds may not be suitable for many soil types without using lining, and a thorough review of potential effects on surface water and groundwater quality would have to be conducted if clay-lined ponds are to be used. If outdoor ponds are poorly lined or the lining fails as a result of wear, then seepage of the pond water into the local groundwater system could occur. Clays that are compacted and graded have structural integrity that can be comparable to synthetic liners (Boyd, 1995). However, integrity can be compromised by poor construction. Nitrate leaching has been observed below structured clay soils (White et al., 1983), but the qualitative applicability of these results to clay-lined algal ponds is unknown. Local terrestrial vegetation might take up some of the culture media released through seepage. In some areas, if open ponds contain high concentrations of dissolved inorganic nitrate, seepage may contribute to concerns related to nitrate poisoning if the groundwater is used for drinking by livestock or humans.
Withdrawal of freshwater adjacent to briny aquifers or injection of saline wastewater into the ground could result in salinization of groundwater if fresh water and briny aquifers are not well separated. Salinization of groundwater is a potential problem for some agricultural lands where irrigation is prevalent (Schoups et al., 2005). However, one of the key advantages of algal biofuel is that the feedstock could be produced on nonarable land (Ryan, 2009; Assmann et al., 2011), so salinization of agricultural lands as a result of freshwater withdrawal for algae cultivation is not likely.
5.1.4.2 Opportunities for Mitigation
Using sealed algal cultivation systems would practically eliminate the potential for leakage, barring catastrophic breaches. Where open systems are used, technologies (such as the development of impermeable, long-lived liner systems) and regional solutions for minimizing nutrient leakage could be deployed, and regulations to minimize leakage could be developed. For example, Phyco BioSciences uses a trough system that has a lightweight, fabricated liner. The liner is expected to eliminate leakage or minimize percolation to less than 0.01 percent (Cloud, 2011). Potential preventive measures might include specifications for soil type, combined with defined values for the minimum depth from the pond bottom to groundwater. Moreover, local regulations likely require lined ponds, which would reduce the probability of leakage of waters but contribute to capital costs and lead to temporary system closures when the liners are replaced because of wear or failure. Measures to prevent inadvertent discharge of water (for example, overflow corridors or basins) during extreme weather events would be helpful in preventing water pollution.
5.1.5 Wastewater Treatment
Wastewaters derived from municipal, agricultural, and industrial activities potentially could be used for cultivating algal feedstocks either in open ponds or in photobioreactors for algal biofuels and could provide an environmental benefit. Microalgae have been used in wastewater treatment for a long time (Oswald et al., 1957), where they provide photosynthetically produced oxygen for the bacterial breakdown of organic compounds present in the waste (Benemann, 2008). Microalgae have been shown to be effective for wastewater treatment in diverse systems including oxidation (stabilization) ponds and shallow raceway systems and using both phytoplankton and periphyton (Green et al., 1995; Hoffmann, 1998; Pittman et al., 2011; Sandefur et al., 2011). High rate algal ponds (HRAPs), which are shallow, open raceway ponds used for treating municipal, industrial, and agricultural wastewater, combine heterotrophic bacterial and photosynthetic algal processes (Park et al., 2011). The ponds allow the growth of high-standing crops of algae, which remove nitrogen and phosphorus from the wastewater (Sturm et al., 2012). The concept of adapting HRAPs for the purpose of biofuel production was proposed more than five decades ago (Oswald and Golueke, 1960). Park et al. (2011) reviewed the potential benefits and opportunities of using HRAPs for wastewater treatment and harvesting the algae for energy or fuel production. The feasibility and scale of such systems will be determined by the amount of wastewater, the availability of land near the facilities generating the wastewater and produced water, and the climatic conditions of the region. (See also Chapter 4.) If wastewater is used, the wastewater treatment rate and the harvesting schedule would determine the maximum volume of ponds or photobioreactors.
A major goal of wastewater treatment is removal of nitrogen and phosphorus (Pittman et al., 2011). In conventional treatment systems, phosphorus is especially difficult to remove (Pittman et al., 2011). In advanced wastewater treatment, phosphorus typically is either chemically precipitated using aluminum- or iron-based coagulants to form an insoluble solid, or it is stripped from the water by microbial activity (EPA, 2007). The recovered phosphorus is then buried in a landfill or treated to create sludge fertilizer (Pittman et al., 2011). Given that readily available supplies of phosphorus may begin running out by the end of the 21st century (Vaccari, 2009), conservation and stewardship of U.S. phosphorus supplies are essential. Recycling nutrients from wastewater and using them for further algae production could be an attractive option for using otherwise discarded nutrients (Exhibit 9.7 and associated text in DOE, 2010b; see also section Nutrients in Chapter 4).
Algae-based treatments have been found to be as efficient as chemical treatment in removing phosphorus from wastewater (Hoffmann, 1998). Moreover, because harvested algal biomass contains the nutrients that were absorbed during cellular growth, wastewater-integrated systems can perform an important nutrient removal service. In laboratory-scale experiments, more than 90 percent of nitrogen and 80 percent of phosphorus were removed from primary treated sewage by the green alga Chlorella vulgaris (Lau et al., 1995). Similarly, laboratory cultures of Chlorella and Scenedesmus removed 80 to 100 percent of NH3, nitrate, and total phosphorus from wastewater that already had undergone secondary treatment (Martinez et al., 2000; Zhang et al., 2008; Ruiz-Marin et al., 2010). Sturm et al. (2012) performed a six-month, pilot-scale algal production experiment using large (10 cubic meters) outdoor bioreactors fed by effluent from the secondary clarifier of the wastewater treatment facility in Lawrence, KS. They reported only a 19 percent removal of dissolved nitrogen and a 43 percent removal of dissolved phosphorus from this treated effluent. These differences in nutrient removal observed may be related, in part, to the different scales of the studies. The ultimate level of nutrient removal benefit may depend on the level of wastewater treatment that occurs prior to nutrient uptake in the algal cultivation systems and on the chemical and ecological conditions that exist in the wastewater-fed production system.
Algae have the potential to remove nutrients from agricultural or industrial wastewater. Some studies have found high efficiencies of removal of nitrogen and phosphorus from wastewater containing manure (Gonzalez et al., 1997; Wilkie and Mulbry, 2002; An et al., 2003), and this wastewater also could be used as input to algal biofuel systems. Algal biofuel systems have the potential to increase water quality and to promote municipal or agricultural wastewater treatment systems with improved sustainability. However, the maintenance of lipid-rich strains and the manipulation of growth conditions to promote high lipid production have yet to be demonstrated consistently for outdoor pond systems, including wastewater treatment ponds (DOE, 2010b). Industrial wastewaters have lower nutrient concentrations and higher toxicant concentrations, and thus are less likely to be used to generate the algal biomass necessary for commercial-scale production of biofuels (Pittman et al., 2011).
Integrated algal biofuel production systems can remove many other pollutants, such as metals and organic contaminants, including endocrine disruptors (Mallick, 2002; Munoz and Guieysse, 2006; Ahluwalia and Goyal, 2007; DOE, 2010b). Whether pollutant uptake by algae is desirable depends on whether coproducts are to be produced with algal biofuels or whether the lipid-extracted algae are to be used for nutrient recycling. Pollutant removal by these systems would improve water quality, but it also could pose a potential risk if organisms such as migrating waterfowl directly or incidentally consumed high metal content algae during the cultivation process, or if humans or wildlife were exposed chronically to the dried algae during biomass processing. Uptake of pollutants by algae is not desirable if residual biomass is to be used for human cosmetic products or animal feed.
5.1.6 Comparison of Pathways
The pathways described in Chapter 3 affect the types, probabilities, and magnitudes of water-quality effects (Table 5-1). For example, slow releases of nutrients to natural environments (and increased potential for eutrophication and groundwater pollution) are common for open systems but not for closed systems. Herbicides likely would be used only in open systems. The water quality benefit for wastewater treatment is achieved only if wastewaters are used as nutrient sources, but the scenarios in Chapter 3 do not specify this.
TABLE 5-1 An Illustration of Potential Benefits and Adverse Effects to Water Quality from Different Pathways for Algal Biofuel Production
|
|
||||||
|
Potential Effect |
Pathway | |||||
|
|
||||||
|
Open-pond, salt water, producing biodiesel, recycling nutrients and water |
Open-pond, salt water, producing biodiesel + coproducts |
Open-pond, salt water, producing FAMEa, recycling nutrients and water |
Photobioreactor, salt water, direct synthesis, recycling water |
Open-pond, salt water, producing biomass, pyrolysis, recycling some nutrients and water |
||
|
|
||||||
|
Releases of Culture Water |
Slow releases from seepage, overtopping likely, |
Slow releases from seepage, overtopping likely, |
Slow releases from seepage, overtopping likely, |
No slow releases, catastrophic breaches rare |
Slow releases from seepage, overtopping likely, catastrophic breaches rare |
|
|
|
||||||
|
Eutrophication and Related Effects |
Rare, only when large volume releases occur |
Rare, only when large volume releases occur |
Rare, only when large volume releases occur |
Very rare, only when large volume releases occur |
Rare, only when large volume releases occur |
|
|
Waterborne Toxicants |
Herbicides, heavy metals may be present and pose occupational or ecological exposures and risks |
Herbicides, heavy metals may be present and pose occupational or ecological exposures and risks |
Herbicides, heavy metals may be present and pose occupational or ecological exposures and risks |
Heavy metals may be present and pose occupational exposures and risks |
Herbicides, heavy metals may be present and pose occupational or ecological exposures and risks |
|
|
Groundwater Pollution |
Possible, depending on soil type, distance to groundwater, and frequency of release |
Possible, depending on soil type, distance to groundwater, and frequency of release |
Possible, depending on soil type, distance to groundwater, and frequency of release |
Rare, only when catastrophic breaches occur |
Possible, depending on soil type, distance to groundwater, and frequency of release |
|
|
Wastewater Treatment |
Algae may treat wastewater if wastewater is used as nutrient source |
Algae may treat wastewater if wastewater is used as nutrient source |
Algae may treat wastewater if wastewater is used as nutrient source |
Algae may treat wastewater if wastewater is used as nutrient source |
Algae may treat wastewater if wastewater is used as nutrient source |
|
|
|
||||||
aFatty-acid methyl esters.
5.1.7 Sustainability Indicators
Proposed sustainability indicators for water quality include aqueous concentrations and loadings of nutrients, herbicides, metals, and salinity of groundwater (GBEP, 2012). These indicators are standard measures for quality of water and wastewater (Eaton et al., 2005). Concentrations of nutrients are included because they relate to benefits or potentially adverse effects on water quality (for example, eutrophication). These usually are measured quantities, and baseline levels and natural variability also can be measured. Loadings are field measures or simulation results representing the contribution of released algal biofuel culture media to receiving waters. These may be compared to other loadings to those waters.
• Nitrate concentration in streams and groundwater.
• Total nitrogen concentration in streams, lakes, reservoirs, and estuaries.
• Total phosphorus concentration in streams, lakes, reservoirs, and estuaries.
• Nitrate loading to streams and groundwater.
• Total phosphorus loading to streams.
• Herbicide concentrations in streams.
• Herbicide loading to streams.
• Metal concentrations in streams.
• Metal concentrations in cultures.
• Salinity of groundwater.
5.1.8 Information and Data Gaps
Good design and engineering would minimize the potential for releases of water and nutrients from open-pond systems to surface water and to ground water. Toxicant concentrations (for example, metals) need to be characterized, particularly if wastewater or produced water is used as culture medium. Information on the nutrient removal efficiencies of commercial-scale facilities would be needed if algal biofuel production is to be combined with wastewater treatment.
5.2.1 Potential Environmental Effects
Land-use change is a change in anthropogenic activities on land, which often is characterized in part by a change in land cover, including the dominant vegetation. Land-use changes play a role in the sustainability of algal biofuel development because of associated environmental effects, such as net GHG emissions, changes in biodiversity, and changes in ecosystem services such as food production. Moreover, there is growing societal concern about the spatial and temporal scales of some types of conversions, such as deforestation and urbanization. The impacts of algal biofuel development will depend in part on the type of land conversion, the extent (area) of land use that has changed, the intensity of land disturbance and management, and the duration of the change (for example, whether it is reversible).
Commercial-scale production of algal biofuels will require substantial land area for each facility (see Chapter 4), and the large-scale deployment of algal biofuels will lead to conversion of lands from other existing uses. Land conversion for ponds, processing facilities, and refineries for most products will be localized, and potential land conversion for related infrastructure, such as roads and power lines to the facilities, will be more diffuse and will involve linear features. This section focuses on land-use change (LUC) associated with algae cultivation, because change associated with feedstock processing or refining facilities is not different in kind from that of other liquid fuel sources.
High-value lands used by agriculture, by other commodity industries, and for residential purposes are unlikely to be used for algae cultivation because algae cultivation does not require fertile soils and because capital and operating costs would have to be kept low for algal biofuel companies to operate close to the profit margin (Table 5-2). Similarly, the conversion of forestland is unlikely because of the high costs of clearing and site preparation and the high value for residential or recreational use. Land-use change for algal biofuels is
TABLE 5-2 A Summary of the Committee’s Judgment on the Likelihood of Land (or Water Surface) Conversion to Algae Cultivation Ponds and Facilities, Based on Value for Other Land (or Surface Water) Uses
|
|
||
|
Land Type |
Possible or Likely |
Unlikely |
|
|
||
|
Productive agricultural land |
X |
|
|
Marginally or unproductive agricultural land |
X |
|
|
Desert |
X |
|
|
Brownfields |
X |
|
|
High-value coastal land |
X |
|
|
Low-value coastal land |
X |
|
|
Forest land |
X |
|
|
Rangeland, low-density grazing land |
X |
|
|
Parks and conservation land |
X |
|
|
Wetlands |
X |
|
|
Residential land |
X |
|
|
Industrial parks |
X |
|
|
Urban land other than brownfields |
X |
|
|
Former catfish pond lands |
X |
|
|
Offshore |
X |
|
|
|
||
NOTE: Low-value land is assumed to be used to cultivate algae for biofuels.
more likely to involve brownfields2, rangelands, deserts, scrubland, abandoned farmland, or unproductive farmland, some of which may be on coasts or in near-shore marine waters. On coasts, dredge spoil islands might be additional options for use. For example, Phycal, an algal biofuel company, is using fallow land in Hawaii that was previously a pineapple plantation but is no longer economically viable for that use. Sapphire, another company operating in the Southwest, plans to develop nonagricultural land for algae cultivation. (Siting requirements are described in Chapter 4.) Competing land demands could change over time and may influence the landscape of algal biofuels. For example, some of the same lands that are attractive for algal biofuel development are also attractive for large-scale solar power development (BLM and DOE, 2010).
Direct land-use change generally is defined as a direct cause-and-effect link between biofuel development and land conversion in the absence of strong external mediating factors. Direct land-use change occurs within the biofuel production pathway when land for one use is dedicated for biofuel production. However, in practice, direct land-use change from biofuel production generally is assumed to include lands used for feedstock production, processing, storage, and refining areas. Indirect land-use change occurs when biofuel production causes new land-use changes elsewhere domestically or in another country through market-mediated effects (NRC, 2011).
Direct and indirect land-use changes could affect the net GHG emissions of biofuels (NRC, 2011). Direct land-use change can result in carbon sequestration or net GHG emissions, depending on the type of land conversion and prior land use. For example, converting from annual-crop production to perennial-crop production can enhance carbon storage on that piece of land (Fargione et al., 2008). Conversely, clearing native ecosystems to produce row-crops would result in a one-time release of a large quantity of GHGs into the atmosphere (Fargione et al., 2008; Gibbs et al., 2008; Ravindranath et al., 2009). In the
_______________
2 Brownfields are “industrial or commercial propert[ies] that [remain] abandoned or underutilized because of environmental contamination or the fear of such contamination” (Environmental Law Institute, 2012).
context of algae cultivation, converting pastureland to algal ponds is likely to contribute to GHG emissions. Perennial pasture is effective in sequestering carbon in soil (Franzluebbers, 2010; Gurian-Sherman, 2011). Removal of such vegetation would result in a one-time loss of carbon and the elimination of any potential for further carbon sequestration if the land is to be left as a pasture. In contrast, if the algae cultivation ponds are installed on degraded land that is not storing much carbon, immediate emissions from the conversion will be minimal.
Indirect land-use change could occur if the use of land to cultivate biofuel feedstocks replaces and ultimately reduces the production levels of crops destined for a commodity market. The lowered production of those commodities could drive up market prices, which in turn could trigger agricultural growers to clear land elsewhere to grow the displaced crops in response to market signals (Babcock, 2009; Zilberman et al., 2010). However, as stated above, because algal feedstock cultivation does not require fertile cropland, arable land likely will not be used for algal biofuels (Sheehan et al., 1998; Gong and Jiang, 2011), and displacement of commodity crops by algae is unlikely. In addition, protein from lipid-extracted algae potentially can replace soybean or other terrestrial crops as feedstuff (Wijffels and Barbosa, 2010) and reduce the demand for land by terrestrial crops. The nutritional compatibility of algal feedstuff and the animal diet would have to be examined.
Pasture and rangeland could be converted to algae cultivation, and displacement of these land uses by algae also may or may not result in other indirect effects. If the pasture or rangeland is surplus and not in use, then repurposing the land will not incur indirect land-use change (ILUC). In contrast, if algae cultivation displaces grass-fed cattle production, producers might decide to change to corn-fed cattle production. Changing from grass-fed to corn-fed cattle production also would exert pressure on the corn-grain market. Alternatively, if existing pasture and rangeland is limiting beef production, such that removing some of this land would decrease production, then grass-fed cattle production might be replaced elsewhere. The indirect land-use changes not only affect ecosystem services, but result in changes in GHG emissions that have to be considered in life-cycle GHG assessments for algal biofuels.
If the indirect effects of algal biofuel production are to be quantified, then the potential biodiversity, water quality, and water balance impacts would include those associated with indirect land conversions. Previous quantification of indirect effects of biofuels generally has been limited to GHG effects and food security effects.
As in the case of terrestrial-crop biofuels, market-mediated indirect land-use changes are difficult to ascertain, and estimates of associated GHG emissions are highly uncertain (NRC, 2011). Although complex models have been used to project the extent of indirect land-use changes as a result of terrestrial-crop biofuels, the committee is not aware of similar projections for algal biofuels. Algae cultivation is less likely to incur indirect land-use changes because it does not require prime agricultural land. Converting crop lands to new crops (algal biofuels) also will require new ownership or a willingness on the part of farmers to grow a new commodity. Growing algal biofuels will require differing work schedules than row crop farming. Even if cropland is not to be converted to algal ponds, the above discussion of potential pasture conversion illustrates a potential for indirect land-use change.
5.2.2 Comparison of Pathways
With respect to land-use change, the primary relevant difference among the pathways in Chapter 3 is the difference between the land required for open-pond and photobioreactor systems (see Chapter 4). The spatial and temporal scales of land-use change would be commensurate with those of land use.
5.2.3 Potential Opportunities for Mitigation
In general, algal biofuel development will avoid forestland and land with agricultural value. Avoiding pastureland and areas of high biodiversity or recreational value also would eliminate some of the sustainability concerns associated with commercial development of algal biofuels.
5.2.4 Sustainability Indicators
Land-use change is not consistently proposed as a criterion for sustainability, even though it often is considered a factor influencing aspects of the sustainability of biofuel (for example, GHG emissions, biodiversity, water quality, and soil quality). Therefore, some compilations of sustainability indicators do not include indicators of sustainable land use (for example, McBride et al., 2011). However, there are aspects of land use, such as infrastructure, impervious surfaces, and some disturbances, that may be long lasting or irreversible and may not be adequately considered using indicators of other categories of sustainability. Potential indicators of sustainable land use include percent impervious surface (Sutton et al., 2009; Uphoff et al., 2011; Weiland et al., 2011) and land disturbance area. Changes in impervious surface area affect the water cycle and watershed dynamics, as well as terrestrial and aquatic habitats. The area of land disturbed can be considered a measure of sustainability. Land disturbance areas can be normalized based on a land-condition factor (Eq. 5-1) that captures the degree to which aspects of development, processing, infrastructure, potential accidents, and use of energy change the land from its natural state (Lenzen and Murray, 2001, 2003) and its ability to provide ecosystem services.
![]()
Table 5-3 shows examples of land condition factors that can be multiplied by disturbed area to give a currency of disturbance. This metric relates to ecological footprint methods that sometimes are applied to energy comparisons (Stoglehner, 2003), but it does not attempt to encompass effects on water, GHG emissions, and other ecological impacts that can be more controversially subsumed in ecological footprints (Fiala, 2008; Özdemir et al., 2011).
TABLE 5-3 Illustrative Land Condition Factors for Land-Cover Changes Relevant to Algal Biofuel Production
|
|
|
|
Land Use or Land Cover Type |
Land Conditionc |
|
|
|
|
Builta (refineries, offices) or paved |
1.0 |
|
Ground denuded of vegetation but no pond |
0.8 |
|
Earthen pond or raceway containing algal culture |
0.5 |
|
Lined pond or raceway containing algal culture |
0.6 |
|
Partially disturbed grazing landb |
0.2 |
|
|
|
aValue taken from Lenzen and Murray (2003). Reprinted with permission from Elsevier.
bValue taken from Lenzen and Murray (2003) and can represent areas between algal biofuel facilities, where grazing may occur. Reprinted with permission from Elsevier.
cLand condition factors capture the proportion of disturbed land or relative intensity of disturbance, with land in a natural state having a land condition factor of 0 and paved land having a land condition factor of 1. Land condition factors are multiplied by disturbance area to allow comparison of disturbed areas of different intensities and scales.
Trends in land-use change related to algal biofuel production are important to quantify. However, until there is a history of commercial development of algal biofuel production facilities, probable land-use changes and trends will need to be projected based on economic and social drivers and environmental contributing factors.
Where important or rare ecosystem services are provided by the baseline land use, a measure of those services could serve as a sustainability indicator for algal biofuels. The services of pastures, rangelands, and coastal waters that might be displaced by feedstock production facilities would be important to quantify. Relevant metrics would be:
• National or regional area of grassland and shrubland devoted to livestock grazing; however, data are lacking on the acreage used for livestock grazing (The H. John Heinz III Center for Science and the Environment, 2008).
• Number of livestock fed on grasslands and shrublands (West, 2003; The H. John Heinz III Center for Science and the Environment, 2008).
• Pasture yield calculated on a per-area or per-forage biomass basis (methods described in Burns, 2008).
A less direct indicator of livestock numbers or biomass would be area covered by grassland and shrubland (West, 2003; The H. John Heinz III Center for Science and the Environment, 2008). Additional sustainability indicators have been suggested for brownfield redevelopment efforts. Some of these are summarized in Wedding and Crawford-Brown (2007) and would be appropriate where algal biofuel production is sited on brownfields.
The potential to mitigate GHG emissions is one of the motivations to develop biofuels. The basis of mitigation is that carbon emissions from combusting a biofuel are cancelled by the corresponding capture in photosynthesis. This said, the net GHG emissions of producing biofuels and coproducts are not zero because of carbon and other GHGs emitted in processing. In this section, the results of life-cycle assessment (LCA) studies of GHG emissions are reviewed critically.
5.3.1 Life-Cycle GHG Emissions of Algal Biofuels
Primary GHG emissions from algal biofuels are expected to be connected to the use of energy in the processing chain (see section Energy in Chapter 4). The translation of energy use to GHG emissions is complicated by variability in the carbon overhead of different forms of energy, in particular electricity. The average direct GHG emissions of electricity production in the United States is 606 grams of CO2 equivalent per kilowatt hour (EIA, 2002). Depending on the mix of fossil fuels, hydropower, nuclear, wind, and other sources providing power to the grid, emissions vary by state from 13 to 1,017 grams CO2 equivalent per kilowatt hour (EIA, 2002). The approach taken by many analysts is to use a national average emission factor (Liu et al., 2011).
LCA results for net GHG emissions for algae biofuel production vary from a net negative value (that is, a carbon sink) to positive values substantially higher than petroleum gasoline (Table 5-4). As with the case for energy use (see Chapter 4), drivers of variability in CO2 emissions are nutrient source, productivity and process performance, and the credit associated with coproducts. For example, Sander and Murthy (2010) assumed that residual algal biomass substitutes for corn in ethanol plants. Corn is energy intensive to produce; the
TABLE 5-4 Results from Sample Studies of Life-Cycle CO2 Emissions of Algal Biodiesel Production in Common Normalization
|
|
|
|
Source |
Life-Cycle CO2 Emissions (kg CO2 equivalent per liter of biodiesel) |
|
|
|
|
Clarens et al. (2010) |
8.7 |
|
Lardon et al. (2009) |
4.0 |
|
Stephenson et al. (2010) |
0.6 |
|
Sander and Murthy (2010) |
-0.8 |
|
Campbell et al. (2011) |
-1.1 |
TABLE 5-5 Meta-Analysis Results for Ranges in Carbon Emissions Over the Life Cycle of Algal Biofuel Estimated by Various Studies
|
|
|
|
Scenario |
GHG Emissions (kilograms CO2 eq per liter biodiesel) |
|
|
|
|
Virgin CO2, no coproduct |
5.9-8.2 |
|
Virgin CO2, w/ coproduct |
2.0-4.2 |
|
Recycled CO2, no coproduct |
2.1-2.9 |
|
Recycled CO2, w/ coproduct |
(-1.2)-(-2.9) |
|
|
|
NOTE: The direct carbon emissions of driving an average gasoline automobile is about 0.15 kg CO2 eq per kilometer.
SOURCE: Liu et al. (2012). Reprinted with permission from Elsevier.
GHG credit from replacing corn with oil-extracted algae as a feedstock for ethanol results in a negative carbon balance. For reference, the direct carbon emission of combusting gasoline is about 2.7 kg CO2 equivalent per liter of fuel (Farrell et al., 2006).
The vast differences in results in Table 5-4, ranging from a net carbon credit to emissions far larger than those from petroleum-based diesel, present a challenge for interpretation. Liu et al. (2012) performed a meta-analysis of these studies to analyze variability in processing energy by replacing differences in data and assumptions for nutrients and coproducts with common data (Lardon et al., 2009; Clarens et al., 2010; Jorquera et al., 2010; Sander and Murthy, 2010; Stephenson et al., 2010; Campbell et al., 2011). Differences in nutrient sourcing and coproducts are treated via four scenarios: virgin versus recycled CO2 and no coproducts versus coproducts. The common coproduct system used is generation of bioelectricity from gas generated by anaerobic digestion with the electricity generated substituting for carbon emissions from the U.S. grid. Table 5-5 shows the ranges in results from the six treated studies, after normalization, for the four scenarios.
These meta-analysis results suggest that the CO2 source and coproducts are critical factors in the GHG balance. It is, however, premature to conclude that algal biofuels based on recycling CO2 and producing biogas has net negative GHG emissions. The variability in Table 5-5 is based on differences in energy data and assumptions in the six existing studies. It is not yet clear if current LCA analyses of algal biofuel production systems will accurately reflect the energy use of a real-world, scaled-up system.
None of the studies above addresses the potential issue of indirect land-use change from biofuels. As stated earlier, it is possible that conversion of pastureland to algae cultivation facilities would necessitate conversions to pastureland elsewhere. However, uncertainties are too great to quantify this probability or to calculate net GHG emissions under these assumptions. (See section Land-Use Change in this chapter.)
While many agricultural processes emit non-carbon GHGs such as nitrous oxide (N2O) and methane (Weber and Matthews, 2008), these emissions have not been established empirically as significant for algae cultivation. N2O could be emitted from cultivation systems, and these emissions would need to be quantified in the future for cultivation conditions that might promote N2O or methane emission. One study of a single species quantified N2O emissions from algal culture under laboratory conditions (Fagerstone et al., 2011). In this study of Nannochloropsis salina with nitrate as a nitrogen source, elevated N2O emissions were observed under a nitrogen headspace (photobioreactor simulation) during dark periods, but N2O emissions were low during light periods. In contrast, when the headspace consisted of air (open-pond simulation), N2O emissions were negligible. Denitrifying bacteria were present.
Denitrification is the microbial reduction of nitrate and nitrite with generation of N2O and, ultimately, gaseous nitrogen. Anaerobic environments are required for the transformation, but high rates of denitrification occur where oxygen is available alternately, then unavailable (Kleiner, 1974). In rivers, ponds, lakes, and estuaries, the production of N2O is correlated with nitrate concentrations in the water (Stadmark and Leonardson, 2005). The denitrification rate depends on the underlying soil and the liner’s permeability.
Whether anaerobic denitrification is the only potential pathway for N2O generation in algal cultivation systems is unclear. Weathers (1984) has shown that certain Chlorophyceae in axenic culture evolve N2O when using nitrite as a nitrogen source. Florez-Leiva et al. (2010) found that coastal open-pond systems containing Nannochloris emitted large quantities of N2O during senescence. They speculated that oxidation of ammonium (NH4) by bacteria was the likeliest N2O-generation pathway under the observed aerobic conditions. Proper management of the algal cultivation systems, which would prevent senescence of algae and maintain aerobic conditions in ponds, likely would keep N2O emissions to low levels.
Methanogenesis can occur in freshwater and marine sediments, waterlogged soils, marshes, and swamps where oxygen is low. These conditions might prevail in some ponds with substantial biomass or other organic matter in the sediment. Methane is released when organic acids, alcohols, celluloses, hemicelluloses, and proteins are degraded. Methane production is related to water temperature (Stadmark and Leonardson, 2005) and is maximized at neutral pH (Alexander, 1977). Methanogenesis is suppressed by nitrogen compounds that bacteria can use as electron acceptors, including nitrate and nitrite (Bollag and Czlonkowski, 1973), but these compounds may be reduced easily in oxygen-depleted environments. Methanogenesis and denitrification might be enhanced if the culture fails. During catastrophic failure of the culture, the dense algal cultures in algal biofuel ponds can become anaerobic and emit a variety of volatile nitrous or sulfur compounds as well as methane. However, culture failures would be expected to be short-term and rare occurrences if algal biofuel companies are to maintain a profit margin.
5.3.2 Opportunities for Mitigation
The opportunities for mitigating energy use discussed in the section Energy in Chapter 4 apply to reduction of GHG emissions. There is additional potential to mitigate GHGs by using low-carbon energy sources for processing and by substituting for carbon-intensive coproducts. For example, the carbon benefit of generating bioelectricity is larger in areas where the grid relies on fossil fuels. The yields for producing and properties of different coproduct options are poorly understood. The potential for N2O and methane emissions could be reduced through thorough mixing and proper management of algae cultivation (Fagerstone et al., 2011).
5.3.3. Data and Method Gaps
The data gaps for estimating energy use and the method gaps in reducing energy use discussed in the section Energy (Chapter 4) apply to reduction of GHG emissions.
5.3.4 Sustainability Indicators
An appropriate sustainability indicator for GHG emissions is the amount of CO2 equivalent emitted per unit energy produced, which has been selected as an indicator for GHG emissions of biodiesel and commonly has been used in discussing energy-related GHG emissions (GBEP, 2011; Mata et al., 2010).
5.4.1. Potential Environmental Effects
The introduction of large bodies of water in arid or semi-arid environments could alter the local climate of the area by increasing humidity and reducing temperature extremes. Similarly, the introduction of large-scale, open-pond algal cultivation systems in arid or semi-arid environments, where much of algae production in the United States is projected to take place (see Chapter 4), could affect local climate and ecosystems. The use of photobioreactors would not likely alter local climate.
Studies of reservoirs provide some useful ecological information. Reservoirs created by the damming of rivers could affect evaporation rates of the surrounding landscape, leading to changes in vegetation cover and terrestrial species diversity (Huntley et al., 1998). Large dams can affect surrounding climate and precipitation, particularly in Mediterranean and semi-arid climates (Degu et al., 2011).
5.4.2 Sustainability Indicators
The sustainability indicators for potential changes in local climate are trends in relative humidity and trends in temperature distribution statistics.
5.4.3 Information and Data Gaps
While parallels can be drawn from the introduction of large reservoirs in arid regions, the variability in size, geography, and production methods that will emerge as the algae industry grows will necessitate additional research to fully understand and address the impacts associated with local climate alteration.
5.5.1 Potential Environmental Effects
The air quality impacts of algal biofuel production will depend on system design. Different air quality issues arise in conjunction with the different steps of the algal biofuel supply chain. Thus, this section is organized by the steps along the production pathways. The wide range of potential organisms for producing algal biofuels and the wide range of final fuel products result in a broad range of possible air emissions.
This section focuses on the air quality emissions unique to algal biofuel production and does not consider emissions of fossil fuels used to power processing equipment or emissions of fossil fuels that may be used in manufacturing fertilizer or pesticides. The purpose of the chapter is to consider emissions unique to algal biofuel production so that appropriate indicators are identified. However, emissions from fossil fuels used along the production pathway of algal biofuel would need to be considered in any LCA of the airquality impacts of different algal biofuel designs. Further, how algal biofuels will be scaled up and how air quality might change with increasing scale is uncertain.
5.5.1.1 Open-Pond Cultivation
The committee is not aware of any measured emissions of atmospheric pollutants from algal biofuel feedstock ponds published in the literature. Under normal running conditions in open ponds, the cultures are aerobic, and low emissions of volatile organic compounds (VOCs) are expected (Rasmussen, 1974; A. Ben-Amotz, Israel Oceanographic and Limnological Research Ltd., personal communication on November 1, 2011; Zuo et al., 2012). However, macroalgae and microalgae growing in natural marine environments are known to be important sources of VOCs, including isoprene and monoterpenes (Giese et al., 1999; Shaw et al., 2010). Researchers from Texas A&M University currently are screening and quantifying VOCs from a wide variety of marine and freshwater algal cultures and algal paste. Three of the species tested are being grown for biofuels in open raceways, open ponds, and closed photobioreactors, with test samples derived from cultures being grown in treated wastewater with CO2 enrichment. In preliminary findings, 45 VOCs have been identified (P. Zimba, Texas A&M University, unpublished data).
Other emissions expected are aerosols that may be emitted directly or created in the atmosphere through reactions of gaseous emissions of precursor gases of sulfur dioxide (SO2), nitrogen oxides (NOx), NH3, and VOCs. Aerosols could include algae and nutrients, as well as a wide range of compounds that are produced by microalgae, including toxins. (See section Pathogens and Toxins later in this chapter.) Microalgae in the natural marine environment are known sources of sulfate aerosols (for example, Liss et al., 1997).
A large number of algae produce odorous secondary metabolites (reviewed in Smith et al., 2008), but those algae are not likely to be selected for large-scale production. The odors are produced during aerobic growth as secondary metabolites. Other odorous compounds are associated with the decay of algae under anaerobic conditions where bacteria break down the organic material and produce hydrogen sulfide and NH3, both of which have a strong odor. In open ponds intended for algae cultivation, anaerobic conditions are minimized. Emissions from photobioreactors would be lower than those from open ponds if undesirable gaseous products and odorous chemicals are scrubbed before gas exchange with the outside environment is permitted.
5.5.1.2 Drying
Drying processes may produce coarse and fine particulates, including algae and lysed algae. The concentrations of particulates in air will depend on the technologies used; for example, belt dryers and convective systems will lead to greater local emissions than passive solar drying. Whether emissions move beyond the facility will depend on the level of containment. Particulates could be an occupational hazard even in closed facilities. In confined areas, dust could be an explosion hazard. Poor drying methods also can give rise to decomposition of biomass and release of VOCs, amines, methane, and other compounds.
5.5.1.3 Extraction
Most proposed algal biofuel processing methods involve extraction of lipids or other compounds from cells using organic solvents. Extraction with organic chemicals, by necessity, results in some solvent emissions, and the quantities emitted depend on the technology applied. The most common solvent that is openly discussed by manufacturers is hexane (Demirbas, 2009; Lardon et al., 2009; Gong and Jiang, 2011). In an environmental assessment, Sapphire Energy, Inc., an algal biofuel company, noted that “less than 50 ppm of hexane will remain in the algal solids after the hexane recovery process. This residual hexane will be emitted fugitively from the algal solids to the atmosphere during conveyance to the IABR [Integrated Algal Biorefinery] oil purification process” (USDA-RD, 2009). Desirable properties of these solvents are low cost, recoverability, low toxicity, nonpolar structure, and poor extractor of non-lipid cell components (Rawat et al., 2011). Hexane is used as an extractant of vegetable oils in biodiesel production with fugitive hexane emissions (Hess et al., 2009). Compliance with regulatory standards likely would minimize release of solvents.
5.5.1.4 Pyrolysis
Technologies to convert total biomass to drop-in liquid fuels are being tested. These processes may have additional feed inputs and will have different air emissions from those from production of esterified or green diesels. Pyrolysis of biomass yields three energy products—solids (char), liquids (bio-oils), and gases—in various proportions depending on the temperature, pressure, residence time, and other factors. The gases are recycled to provide energy for the system and thus do not contribute directly to air emissions except for any fugitive emissions that might escape the system. The heating of the pyrolysis units might contribute a small amount of NOx and carbon monoxide (CO). Additional energy, likely supplied by natural gas may be required to sufficiently dry the algal biomass prior to pyrolysis. Particulate emissions, acid gases, and hydrocarbon emissions from pyrolysis are not characterized in the literature. The bio-oil produced from whole-cell pyrolysis will require additional upgrading to produce transportation fuels. The upgrading can be done with a separate hydrotreating step or a process similar to the Integrated Hydropyrolysis and Hydroconversion process. In either case, input of hydrogen is required. The production of hydrogen produces low levels of NOx (Spath and Mann, 2001) and makes a CO2 stream that could be used to supply the algae cultivation.
5.5.1.5 Anaerobic Digestion
Anaerobic digestion for processing wastewater from algal biofuel production facilities is described in Chapter 2. NH3 has been observed to be present in biogas from anaerobic digestion at concentrations up to 450 ppm (Schomaker, 2000). The concentration of NH3 in biogas would depend on the nitrogen content of the particular feed material. Early work by Golueke et al. (1957) found that anaerobic digestion of algae yielded N2 and NH3 concentrations of the order of 4 to 5 percent (volumetric average) of the total gas production. NH3 would not be released to air around the facility because of the desire to recycle nutrients required for cultivation.
5.5.1.6 Transportation Emissions
The primary categories of environmental effects associated with the end use of biofuels in vehicles are evaporative emissions and tailpipe emissions from fuel combustion.
TABLE 5-6 Comparison of Typical Tailpipe Emissions from Biofuels to Conventional Gasoline or Diesel
|
|
|||
|
BIOETHANOL (E10) |
BIOETHANOL (E85) |
BIODIESEL (B20 and B100) |
FISCHER-TROPSCH |
|
|
|||
|
|
|
|
|
|
|||
NOTES: E10 is a fuel blend of up to 10% denatured ethanol. E85 is a fuel blend of up to 85% denatured ethanol. B20 is a blend of 20% biodiesel and 80% petroleum diesel and is the most common biodiesel blend in the United States. B100 is 100% biodiesel. Fischer-Tropsch (F-T)synthesis converts a mixture of CO and hydrogen (which may be derived from biomass) into liquid hydrocarbons.
SOURCES: Dufey (2006), EPA (2002a,b,c), and Graham et al. (2008).
Generally, the type and quantities of emissions vary depending on fuel characteristics (for example, chemical properties and blends), age of the vehicle or other equipment, power output of engine, operating condition of engine, how the vehicle or other equipment is operated, and ambient temperature (Graham et al., 2008; Yanowitz and McCormick, 2009; Ginnebaugh et al., 2010). Using biofuels in place of petroleum-based fuels decreases emissions of some air pollutants while increasing others (Table 5-6; NRC, 2011). EPA established emission standards for tailpipe emissions of CO, hydrocarbons, NOx, and particulate matter to which vehicle manufacturers and refiners have to comply (EPA, 2009a).
5.5.1.7 Life-Cycle Assessment
Emissions of air pollutants need to be assessed over the life cycle of algal biofuels and compared to petroleum-based fuels and other alternatives. The Hill et al. study (2009) and data therein (NRC, 2011) illustrate the importance of such assessment. They found that although the uses of gasoline and terrestrial-plant biofuels (corn-grain ethanol and cellulosic ethanol) release similar amounts of VOC, PM, NOx, SOx, and NH3, emissions from the production stages are significantly different between petroleum-based fuels and biofuels. Biofuels emit higher quantities of VOCs, NOx, NH3, and PM2.5 than petroleum-based fuels (Hill et al., 2009). The committee is not aware of any LCA of such air pollutants for algal biofuels. Such analysis is critical in assessing whether biofuel production and use result in air quality improvement compared to fossil fuel and it provides information on stages in the supply chain that are key contributors to air pollutants.
TABLE 5-7 An Illustration of Potential Contributions from Different Stages of the Algal Biofuel Supply Chain to Air Pollutants
|
|
||||||
| Potential Effect | Pathway | |||||
|
|
||||||
|
Open-pond, salt water, producing biodiesel, recycling nutrients and water |
Open-pond, salt water, producing biodiesel + coproducts |
Open-pond, salt water, producing FAME, recycling nutrients and water |
Photobioreactor, salt water, direct synthesis, recycling water |
Open-pond, salt water, producing biomass, pyrolysis, recycling some nutrients and water |
||
|
|
||||||
|
Emissions from Culture |
VOCs, nitrous oxides, methane, aerosols possible |
VOCs, nitrous oxides, methane, aerosols possible |
VOCs, nitrous oxides, methane, aerosols possible |
None |
VOCs, nitrous oxides, methane, |
|
|
Odors from Culture |
Some odors |
Some odors |
Some odors |
No odors |
Some odors |
|
|
Drying |
Particulates generated, amount depending on drying process |
Particulates generated, amount depending on drying process |
Particulates generated, amount depending on drying process |
No drying |
Particulates generated, amount depending on drying process |
|
|
Extraction |
Some solvent vaporization |
Some solvent vaporization |
Some solvent vaporization |
No extraction so no solvent vaporization |
No extraction so no solvent vaporization |
|
|
Pyrolysis |
Not applicable |
Not applicable |
Not applicable |
Not applicable |
Particulate emissions, hydrocarbon slip, and acid gases all possible from combustion of off-gas |
|
|
Anaerobic Digestion |
Possibility of low or negligible NH3 release |
Not applicable |
Possibility of low or negligible NH3 release |
Not applicable |
Not used |
|
|
|
||||||
5.5.2 Comparison of Pathways
With respect to air quality, the differences in expected effects among the pathways in Chapter 3 depend on the type of culture system (open versus closed), the drying process, and whether or not extraction and pyrolysis steps are present in the pathway (Table 5-7).
5.5.3 Potential Social Acceptability Effects
Algae produce a number of aerosols and secondary metabolites, some of which may be noxious (for example, malodorous) or harmful to humans. Similarly, some supply-chain processes, such as extraction and drying, may emit solvents or particulates that could affect local air quality if not contained. If an algal biofuel facility is located near human populations, measures likely will be taken to contain or limit the release of any products that negatively affect local air quality or are perceived to be a risk to public health. The health costs of some types of air emissions were discussed in Hill et al. (2009). Depending on the quantity of these outputs, and the proximity of population centers to a production facility, the reduction in air quality and perceived health and quality-of-life risks may impact the
siting and permitting processes, making it more difficult for developers to secure land and obtain permits. If the public is not made aware of these potential effects prior to the siting and permitting of a facility, there is a risk that the production of undesirable compounds will be viewed as unacceptable after the construction of the facility has been completed. If this is the case, litigation or protests may slow or shut down operations, resulting in financial losses for the developer and negative attention for the industry at large.
5.5.4 Opportunities for Mitigation
The more contained a process is, whether it is the biomass cultivation process, drying, solvent extraction, pyrolysis, or digestion, the lower the emissions to air will be. Therefore, photobioreactors could have reduced air-quality impacts compared to open-pond systems. However, full LCA of the air pollutant emissions associated with the production of the bioreactor materials and system operation also would be needed to assess whether photobioreactors represent a small or negligible impact on air quality. Although passive processes (for example, solar drying) reduce air quality impacts compared to active processes that generate dust or increase volatilization rates, they are not practical solutions at large scale. Siting facilities at a distance from human population centers and ecological species of concern would mitigate potential adverse effects of air pollution on humans.
5.5.5 Sustainability Indicators
Appropriate sustainability metrics for air quality would depend on the processes used in algal biofuel production. Concentrations would have to be measured or modeled at scales appropriate to bound regulatory levels or potential human health or annoyance effects. These may include:
• For open pond systems, concentrations of VOCs and odorous secondary metabolites.
• For active drying processes, concentrations of particulates in air.
• For extraction processes, air concentrations of the solvent used.
• For pyrolysis, particulates, hydrocarbons, and acid gases.
5.5.6 Information and Data Gaps
Measuring air emissions from large open ponds can provide information for occupational and other environmental exposure estimates that can be compared to thresholds for human health or environmental effects. Information and data gaps include the relationship between particular drying technologies and the types and concentrations of particulates released, releases of solvents during extraction, likely concentrations of NH3 in air during anaerobic digestion, and chemicals potentially released during pyrolysis. That information would be submitted when the biorefineries seek air-quality permits.
5.6 SPECIES INVASIVENESS AND AQUATIC BIODIVERSITY
Species invasiveness is a concern unique to biofuels produced from algae and vascular plants. In addition, changing land use or altering landscapes to produce algal biofuel feedstocks can affect biodiversity. Effects of many biofuel feedstocks on biodiversity and mechanisms leading to those effects are beginning to be understood. However, existing studies (Fargione et al., 2009; Fletcher et al., 2011; Wiens et al., 2011) focus primarily on
terrestrial ecosystems and terrestrial plant biofuel feedstocks, rather than on aquatic systems and algal feedstocks.
5.6.1 Distributions of Algae in Natural Aquatic Environments
Many cyanobacteria and eukaryotic microalgae are cosmopolitan in their spatial (biogeographical) distributions and therefore could not be invasive if released in regions included in their broad habitat range. However, they are not necessarily found in every location where their habitat requirements (for example, pH, salinity, temperature, moisture, and climate) are met, so their distribution is often mosaic-like (Hoffmann, 1996). Other algae may be endemic to particular regions, for example, some cyanobacteria in Swedish lakes (Rott and Hernandez-Marine, 1994) and particular marine species (Hoffman, 1994). Endemic species could become invasive if transported elsewhere, but these species could also exist in low numbers in other locations even though they have not been recorded there. Algae may have broader distributions than what has been recorded because of the lack of sampling on some continents (especially of benthic habitats) and because of the lack of detection of organisms at low densities (Hoffmann, 1996). Coastal marine macroalgae tend to be less cosmopolitan in their spatial distribution than phytoplanktonic cyanobacteria and microalgae. Macroalgae have narrower temperature, light, substratum, and nutrient preferences.
The wide range of processes that could transport microalgae away from open water also could contribute to their dispersal and consequentially to a broad distribution. Many algal species can be transported by air (Grönblad, 1933). Vectors of algae include aquatic insects (Stewart et al., 1970), dragonflies, wasps (Maguire, 1963), fish (digestive tracts, Velasquez, 1940), beetles (references in Kristiansen 1996), water-living mammals such as raccoons (Maguire, 1963), minks (Irenee-Marie, 1938), and muskrats (Roscher, 1967). The most important vectors of algae are birds (Atkinson, 1972; Kristiansen, 1996). In one study of 16 species of waterfowl, 86 species of algae were found on the feet, 25 species on the feathers, and 25 species on the bills. Most algae survived out of surface waters for four hours, but most did not survive for more than eight hours (Schlichting, 1960).
Some species of algae may appear to be rare. Whitford (1983) explains that species of freshwater algae may appear to be rare for several reasons (for example, infrequent historical collections, species with long-lived spores that do not easily germinate, and species that are highly specific in their habitat requirements), but that very few freshwater species are actually rare. This suggests that few rare species of algae could be displaced by invasive algae used to produce biofuel feedstocks.
5.6.2 Releases of Algae to Natural Environments
Releases of improved nongenetically engineered or genetically engineered strains of algae from biofuel production cultures to natural environments can be expected to be common, especially from open ponds. Releases may occur during the feedstock production stage or possibly during the harvesting or drying stages. Releases probably will occur most often through aerosolization, although leakages from ponds or weather-related spillage (for example, high tides and heavy storms) also are possible.
The probability of release from an open pond would be related to pond area and freeboard space (that is, the distance between normal water level and the top of the cultivation pond), the direction and speed of prevailing winds, the frequency and quantity of
precipitation (for example, rain splash), distance to water bodies, the probability and intensity of visitation by potential vectors such as aquatic birds and mammals, and the absolute abundance (in cells per mL) of the species that might be released into the environment. Humidity affects the survival of unicellular algae (Ehresmann and Hatch, 1975). Survival rates differ among algal groups. In one study climatic characteristics such as temperature, relative humidity, rainfall, wind velocity, and hours of sunshine affected the release and vertical transport of algae (Sharma and Singh, 2010).
Atmospheric density of algae is affected by aerosolization rate (Sharma and Singh, 2010), wind speed, and rainfall, as well as survival rate. The abundance of algae in the atmosphere also depends on taxonomy of the algae. In one study, cyanobacteria had the highest density, whereas chlorophytes and diatoms were much less common (Sharma and Singh, 2010; Wilkinson et al., 2011).
Dissemination to distant sites can occur through the air, through water, and by boats (Alexander, 1971) or animal vectors. The wide range of vectors that could remove algae from open ponds include aquatic insects (Stewart et al., 1970), dragonflies (Maguire, 1963), birds (Atkinson, 1972), and raccoons (Maguire, 1963), among others. Closed photobioreactor systems would have a much lower risk of release and transport of algae. Harvesting operations from open or closed systems could be a major potential route for loss of microalgae to the surrounding environment.
If algae require culture media with characteristics substantially different from the surrounding natural environment (especially if the algae have narrow tolerance limits to nutrients concentrations, pH, or salinity), then releases to the local landscape likely would result in low survival rates. Survival rate would be further reduced if the cultured species is not tolerant of desiccation (Hoffmann, 1996).
5.6.3 Potential Environmental Effects
Environmental concerns associated with releasing algae from biofuel facilities into natural waters include the potential for species invasiveness, alteration of nutrient recycling and trophic relationships, and the displacement of rare algal species. Although some researchers and producers are considering the use of regionally native species that are adapted to the local climate (Odlare et al., 2011), other algal production facilities may use nonnative species or species that have been selected and bred or genetically modified for desirable characteristics for algal biofuel production. Some of the nonnative or improved species may be invasive in some environments. Invasive algae can compete with native species for light, space, or nutrients, and have different tolerances for stressors, compared to native species (White and Shurin, 2011). Thus, invasive species can affect community composition and ecosystem processes (Strayer et al., 2006). Successful invasions are characterized by the invasive potential of the invader and the invasibility of the native community (Lonsdale, 1999). Species that are not invasive in one environment may be invasive when introduced to a different habitat (Raghu et al., 2006). For example, an algal species that thrives in saline waters may not survive or may invade freshwater ecosystems, even if released in a large quantity. Whether the ecological niches of invaders and the invaded community overlap is a predictor of success as well (Mehnert et al., 2010). Whether a particular cultured algal species poses a threat as an invasive species to the surrounding aquatic environments needs to be considered. Some of the same characteristics that can make a species desirable as a biofuel feedstock, for example, rapid growth, vegetative propagation, pest resistance, and robustness in culture, also are those associated with invasiveness.
TABLE 5-8 An Illustration of Potential Environmental Effects Resulting from Species Invasion by Algae Cultivated for Fuels
|
|
|||||
|
Potential Effect |
Pathway | ||||
|
|
|||||
|
Open-pond, salt water, producing biodiesel, recycling nutrients and water |
Open-pond, salt water, producing biodiesel + coproducts |
Open-pond, salt water, producing FAME, recycling nutrients and water |
Photobioreactor, salt water, direct synthesis, recycling water |
Open-pond, salt water, producing biomass, pyrolysis, recycling some nutrients and water |
|
|
|
|||||
|
Transport of Invasive Algae into New Environments |
Possible but unlikely with appropriate controls |
Possible but unlikely with appropriate controls |
Possible but unlikely with appropriate controls |
Impossible unless accidental breach of photobioreactor |
Possible but unlikely with appropriate controls |
|
|
|||||
Releases of some exotic algal species, particularly from open-pond cultures, could threaten the integrity of local and regional ecosystems (Ryan, 2009). Blooms of exotic species could displace native species, with adverse impacts on organisms that feed on those species propagating through aquatic food webs. An example is the diatom Didymosphenia geminata (also known as Didymo or Rock Snot) that can cause dense algal blooms. The blooms block sunlight and cause a local decline in native plant and animal life. Historically, D. geminata occurred mostly in northern latitudes in nutrient-poor waters, but it now has been observed in nutrient-rich water at lower latitudes—possibly a genetic variant that has broader tolerances than the original genotype (see Global Invasive Species Database; ISSG, 2012).
5.6.4 Comparisons of Pathways
The primary variable that is different among the pathways in Chapter 3 and would influence the likelihood of species invasions and changes in biodiversity is whether the pond system is open or closed (Table 5-8).
5.6.5 Opportunities for Mitigation
Algal species known to be noninvasive or unlikely to cause harmful blooms could be selected for large-scale cultivation for fuels. Invasiveness varies in different natural environments, and site-specific assessments might be necessary to reduce risks of invasion. Moreover, species that are intolerant of conditions in natural waters (for example, salinity) in the vicinity of the biofuel facility may be selected to minimize the risk of invasion if released.
Landscape design also may be considered to limit any potential impacts of releases of algae from pond systems. Placing systems well away from waterways and wetlands where pond algae may thrive could reduce or minimize the likelihood of blooms of released species. When considering the factors that affect the probability of release and the abundance of released organisms above, then mitigation measures might include shields from wind and mechanisms to discourage vectors.
5.6.6 Sustainability Indicators
Indicators of sustainable ecological communities include metrics of aquatic diversity and invasiveness of algae. One category of such metrics would be diagnostic traits for invasiveness. Qualitative metrics that are related to invasiveness, but not necessarily diagnostic, include:
• Fast growth in natural environments.
• Wide habitat tolerances, for example, tolerances for temperature, light, and nutrients.
• Pest and herbivore resistance.
• Aggressive competition for resources, for example, light, nutrients, or space.
More direct metrics of aquatic biodiversity that relate to the sustainability of biofuels are recommended by McBride et al. (2011) and are pertinent here:
• Presence of taxa of special concern. These may include rare fish, aquatic invertebrates, or macrophytes.
• Habitat area of taxa of special concern, which for aquatic organisms might translate to stream reach length for taxa of concern.
Additional sustainability indicators for aquatic biodiversity might include the types of metrics found in recovery plans for species protected under the Endangered Species Act (Table 5-9).
TABLE 5-9 Recovery Goals for Endangered and Threatened Species in U.S. Fish and Wildlife Service Recovery Plans
|
|
|
|
Type of Recovery Goal |
Metric for Recovery |
|
|
|
|
Population |
Total population size |
|
Demography |
Age structure of population |
|
Habitat |
Total range (presence/absence) |
|
|
|
SOURCE: Adapted from Efroymson et al. (2009), whose sources were Campbell et al. (2002) and Gerber and Hatch (2002). Reprinted with permission from Elsevier.
5.7.1 Landscape Pattern of Development
5.7.1.1 Potential Environmental Effects
The pattern of landscape conversion for any new infrastructure could affect terrestrial species and community diversity through at least three distinct mechanisms that also apply to algal biofuel production or other energy production (McCabe, 1994; DOE, 2009, 2010a; Garvin et al., 2011):
• Displacement of terrestrial vegetation and wildlife habitat from the facility area and replacement with a pond or photobioreactor containing a monoculture or a few species of algae.
• Reduction in local wildlife habitat area below the threshold needed for the species.
• Fragmentation of wildlife habitat such that mates are more difficult to find or mi-gration corridors are disrupted. The magnitude of land requirements (discussed in Chapter 4) and the types of conversions (discussed in section Land-Use Change in this chapter) influence the magnitude of potential effects on ecological populations and communities.
Displacement of native vegetation and individual vertebrates usually is limited to the area of the facility, but some species are sensitive to human infrastructure and tend to be displaced to distances beyond the boundaries of the facility, for example, female sage grouse avoiding nesting within 950 meters of infrastructure associated with natural gas fields (Holloran et al., 2010).
Extensive infrastructure, especially from multiple facilities, could fragment habitat for some wide-ranging vertebrates. Fragmentation of habitat is determined less by the area of a facility than by the dimensions compared to significant habitat types or corridors. One measure of fragmentation is the ratio of the perimeter (patch edge length) to the area of a habitat patch (Dale and Pearson, 1997). Thus, a linear facility would tend to be fragmenting in more environments than one that is closer to square. However, the latter configuration is more practical for system maintenance, so extensive linear facilities are not considered. Other potential measures of fragmentation include the percent of the landscape occupied by a given habitat, the number or density of habitat patches within a given area (more patches means greater fragmentation), and the degree of connectedness or isolation among habitats (McGarigal et al., 2005).
Even where habitat is not fragmented, human infrastructure and associated disturbance could reduce the habitat area beyond minimum levels required by certain species. Carlsen et al. (2004) review critical patch sizes (contiguous habitat area necessary to conserve a population) required by many species, such as the minimum patch size that can sustain a viable population. They found that few studies examined behavioral or population dynamics associated with large areas of contiguous habitat, which also contained smaller patches of unsuitable or disturbed lands (as in algal biofuel development or oil and gas development). An exception is a theoretical study of American badger at an oil production site that investigated the effects of increasing areas of patches of disturbance on an otherwise highly suitable matrix of tallgrass prairie in Oklahoma (Jager et al., 2006). Critical disturbance areas would depend on the species of concern, the habitat type, habitat suitability, and type of infrastructure.
Impacts on terrestrial vegetation and wildlife could vary widely, depending on the specific sites chosen and the land-use baseline and dynamics prevailing in the absence of algae cultivation and algal biofuel refineries. According to Wigmosta et al. (2011), within the land area potentially suitable for biofuels, land cover types consisted of 42 percent shrub or scrub, 19 percent herbaceous, 14 percent evergreen forest, 10 percent pastureland, 8 percent deciduous forest, and 7 percent other lands including mixed forest, barren, and low-intensity developed. As discussed in Chapter 4, the most favorable conditions in terms of land and water requirements were in the Gulf Coast region. Shrub-scrub habitat in the United States is widely distributed but is threatened by changes in land-use patterns; numerous bird species dependent on this habitat type are in decline (NRCS and WHC, 2007). Development of large areas of shrub-scrub for ponds, up to 181,000 square kilometers (using figures from Wigmosta et al., 2011), could accelerate this decline.
5.7.1.2. Opportunities for Mitigation
The presence and abundance of wildlife need to be assessed prior to construction, as is done for facilities that are subject to environmental assessment (DOE, 2010a). Landscape design could minimize potential effects on biodiversity. Dale et al. (2011) suggest that incorporating design considerations recommended for bioenergy could prevent or minimize adverse effects on terrestrial biodiversity, for example by maintaining corridors for movement of terrestrial wildlife. In planning the size of individual ponds, their density on the landscape, and associated production facilities, managers would have to consider potential environmental impacts on biodiversity.
5.7.2 Wildlife Drinking
5.7.2.1 Potential Environmental Effects
Open algal ponds may be sources of water to wildlife that may prove beneficial in arid conditions or harmful if toxic to certain species. The risks of animals being exposed to salinity or chemicals in water from algae cultivation ponds and having adverse effects from drinking or dermal exposures are unknown.
Toxicity from salt exposure is possible. This occurs when salt or chloride are accumulated in blood at toxic levels and, in the case of birds, at rates too high to be excreted by salt glands. For example, mortality from sodium toxicity has been observed at hypersaline playa lakes of southeast New Mexico (Meteyer et al., 1997). However, the water for algae cultivation is not likely to be hypersaline. Coastal bird species have specialized organs to accommodate high salt levels (Hughes, 2003). Lethal and sublethal salinity concentrations for some species are summarized in a U.S. Department of the Interior report (1998), with toxicity threshold values for ducks ranging from 9 to 20 parts per thousand (compared to the salinity of most seawater at 35 parts per thousand).
Many chemical and behavioral factors could influence exposure of wildlife to salt and other chemicals in open-pond systems. For example, artificial water developments in desert environments are sometimes an important water source for local bird populations (Lynn et al., 2006), but can be less important for some migratory species (Lynn et al., 2006) or animals that may have a strong fidelity for specific water sources (Dickens et al., 2009). If ponds are sited near wastewater treatment facilities and CO2 sources (that is, near population centers), then water is unlikely to be rare in the landscape and wildlife will have many options for water sources. Ponds with dense algae might not be as attractive to wildlife as more pristine
water, but this hypothesis is untested. Similarly, the effect of dense algae on the attractiveness of ponds for wildlife drinking is unknown. For oil-field wastewater evaporation ponds, bird exposures appear to be episodic, coinciding with migration behavior (Ramirez, 2010). To consider potential exposures of wildlife to toxicants in culture water from algal biofuel facilities and their potential effects, analogies may be made to agricultural evaporation ponds and oil-field wastewater evaporation ponds.
In the western part of the San Joaquin Valley, California, agricultural evaporation ponds have been developed where other options for disposal of drainage water are limited. Birds use evaporation ponds for resting, foraging, and nesting (Evaporation Ponds Technical Committee, 1999). One study of shorebird use in California’s Central Valley found that agricultural evaporation ponds are very attractive to these birds (Shuford et al., 1998). In another study, northern pintails (Anas acuta) wintering in Tulare Basin, CA, were found not to use or select agricultural drain-water evaporation ponds or sewage treatment ponds (which might appear similar to some algal biofuel ponds) and to prefer flooded fields and marshes (Fleskes et al., 2003).
For agricultural evaporation ponds, the primary wildlife concern has been the concentration of selenium (Evaporation Ponds Technical Committee, 1999); its environmental transformations and accumulation have been studied (Gao et al., 2007). Whether selenium might represent a significant exposure in algal ponds depends on the availability of selenium in source water and in underlying soils if pond water seeps out. Some investigators suggest that waterfowl exposed to waters from agricultural evaporation ponds might be at risk from uranium toxicity (Duff et al., 1997). Uranium accumulation in pond sediments was attributed in part to decaying algae. Arsenic dynamics also have been studied as a potential concern (Ryu et al., 2010).
In another potentially analogous example, birds (Ramirez, 2010), as well as bats, amphibians, reptiles, small mammals, game species, and insects (Ramirez, 2005), have been observed to be attracted to large (0.4 to 2 hectares) evaporation ponds from oilfield wastewater disposal facilities in the western United States. Bird fatalities from those ponds generally are attributed to oil, but sodium toxicity and surfactants have been implicated in some cases (Ramirez, 2010).
Attraction to algal ponds could be a major problem if they contain toxic chemicals or pathogens at harmful concentrations. Fish injuries (![]() ada, 1998) and bird fatalities (Osborn et al., 2000) have affected the social acceptability (and therefore the sustainable development) of hydropower and wind energy, respectively. Adaptive management can play a role in mitigating any adverse effects on wildlife through exposure via drinking.
ada, 1998) and bird fatalities (Osborn et al., 2000) have affected the social acceptability (and therefore the sustainable development) of hydropower and wind energy, respectively. Adaptive management can play a role in mitigating any adverse effects on wildlife through exposure via drinking.
5.7.2.2 Opportunities for Mitigation
The committee is not aware of any reports of wildlife drinking being a concern in existing open-pond algae cultivation facilities. As the number and size of facilities increase, concentrations of potential toxicants in water and wildlife drinking exposure needs to be monitored to ensure that the latter is not a concern. The fail-safe mitigation for wildlife exposure to salinity or any toxicants in culture waters is to use closed photobioreactor systems. Moreover, salinity concerns would be eliminated through the use of fresh water, though Chapter 4 discusses resource constraints for fresh water at commercial scales of development. Mitigations for open-pond systems might include netting to prevent exposure (as in the oilfield wastewater evaporation ponds), but this would be expensive and only necessary if wildlife exposure proves to be a problem. Rapid stirring could make ponds less suitable as wildlife drinking habitat than still water. Other wildlife deterrents may be
TABLE 5-10 An Illustration of Potential Environmental Effects on Wildlife from Different Pathways of Algal Biofuel Production
|
|
|||||
|
Potential Effect |
Pathway | ||||
|
|
|||||
|
Open-pond, salt water, producing biodiesel, recycling nutrients and water |
Open-pond, salt water, producing biodiesel + coproducts |
Open-pond, salt water, producing FAME, recycling nutrients and water |
Photobioreactor, salt water, direct synthesis, recycling water |
Open-pond, salt water, producing biomass, pyrolysis, recycling some nutrients and water |
|
|
|
|||||
|
Alteration of Terrestrial Habitat |
Displacement of vegetation and vertebrates (habitat loss), possible |
Displacement of vegetation and vertebrates (habitat loss), possible |
Displacement of vegetation and vertebrates (habitat loss), possible |
Displacement of vegetation and vertebrates (habitat loss), possible fragmentation of habitat, probably at a smaller spatial scale than for other pathways |
Displacement of vegetation and vertebrates (habitat loss), possible |
|
|
|||||
|
Wildlife Exposed to Saline Water or Toxicants |
Exposure and adverse effects are possible |
Exposure and adverse effects are possible |
Exposure and adverse effects are possible |
Exposure to water in a closed system is not possible |
Exposure and adverse effects are possible |
|
|
|||||
possible. Some of the mitigations used for oilfield wastewater evaporation ponds, such as covering the surface with plastic balls to make the ponds less attractive to birds (Ramirez, 2010), are not options for photosynthetic fuel sources. Similarly, mitigation strategies used in agricultural evaporation ponds, such as steepening pond slopes or maintaining deep water levels that reduce suitability of bird feeding habitat, are not practical for algae cultivation that requires shallow ponds (Evaporation Ponds Technical Committee, 1999). As open ponds are monitored for chemical contaminants, toxicity thresholds for these chemicals will help determine when culture waters need to be disposed and renewed.
5.7.3 Comparisons of Pathways
As with land-use change, regarding the landscape pattern of development, the primary relevant difference among the pathways in Chapter 3 is the difference between the land required for open-pond and photobioreactor systems (see Chapter 4). For wildlife drinking, the primary variable of interest is closed versus open systems (Table 5-10).
5.7.4 Sustainability Indicators
Metrics of terrestrial biodiversity for the sustainability of biofuels that are recommended by McBride et al. (2011) and pertinent to issues related to the landscape pattern of development include:
• Presence of taxa of special concern (presence).
• Habitat area of taxa of special concern (hectare).
Habitat area can be a proxy for population size (Turlure et al., 2010). As with aquatic diversity metrics, additional sustainability indicators for terrestrial biodiversity might be obtained from recovery plans for species listed under the Endangered Species Act (Table 5-9).
For wildlife exposures to salinity and contaminants in drinking water, sustainability indicators would include:
• Dosage received by wildlife (direct measure).
• Number of vertebrate fatalities from drinking from algal ponds per year (direct measure).
• Concentrations of toxicants, toxins, or salinity in culture medium (less direct measure).
• Abundance of vertebrates drinking from open ponds per year (less direct measure).
5.7.5 Information and Data Gaps
Patterns of development of algal biofuel facilities in relation to wildlife corridors have not been studied because locations for future development are uncertain. The spatial scale and landscape pattern of these developments needs to be understood to simulate the effects on wildlife populations. As algae cultivation expands in number and scale, the potential for wildlife drinking needs to be assessed at sites. If wildlife drinking is observed, then concentrations of toxicants in source waters and culture waters need to be measured to ensure that there is no threat to wildlife health. Alternatively, measures to deter wildlife drinking can be implemented.
5.8 ENVIRONMENTAL EFFECTS OF GENETICALLY ENGINEERED ORGANISMS
5.8.1 Potential Environmental Effects
The environmental sustainability of genetically engineered feedstocks for bioenergy (Wolt, 2009; Moon et al., 2010) and the potential implications of regulations on sustainable development of the industry (Moon et al., 2010; Strauss et al., 2010) have been considered previously, but the emphasis has been on engineered terrestrial crops (Moon et al., 2010) rather than algae. Some algal biofuel companies, such as Algenol and Synthetic Genomics, are conducting research on genetically engineered organisms for algal biofuel production (Gressel, 2008). In a hypothetical, worst-case scenario, genetically engineered algae that have been introduced to natural environments might persist and become so abundant that they create harmful algal blooms (Snow and Smith, 2012). Clearly, any adverse effects of released genetically engineered algae, if observed, would affect the sustainable development of algal biofuel technologies. The evaluation of potential effects of genetically engineered algae will be a complex undertaking, given the diversity of organisms, range of engineered functions, and range of environments potentially receiving the engineered organisms (Tiedje et al., 1989). This section of the report addresses the novel traits and genetic structure of genetically engineered cyanobacteria and microalgae for biofuels and whether they have unique or more uncertain risks. (Potential genetic manipulation methods are discussed in Chapter 2.)
Past broad assessments of the risks of genetically engineered organisms have concluded that the product (novel traits) is more important than the process (genetic engineering techniques) for evaluating risk (NRC, 1987; Tiedje et al., 1989; Snow et al., 2005). However, novel traits may be more common when the process for creating new algae involves direct genetic manipulation than when horizontal gene transfer occurs in evolutionary time.
Several traits of algae for biofuels may be modified through genetic engineering methods. Most are intended to increase biomass or oil productivity, though some could be designed to minimize survival or reproduction following release. Increasing productivity could involve objectives such as enhancing lipid content as a precursor to biodiesel (which could involve growing cells in nitrogen-deficient or silicon-deficient media), introducing biological pathways that permit direct production of fuels that need minimal processing prior to distribution and use, modifying cells to secrete feedstock or fuel directly into the culture medium, modifying carbohydrate metabolism in cells (increasing glucan storage, decreasing starch degradation), increasing tolerance to stressors (such as salt, light, pH, temperature, glyphosate) (Radakovits et al., 2010), and improving resistance to disinfectants. Some of these engineered traits and intended or unintended accompanying traits could affect either the suitability of algae for biofuel production purposes or their survival and physiology when released into natural systems. Phenotypic changes that could lead to potential major ecological effects of released organisms include those that result in increases in physiological tolerance or altered substrate use or that change the species’ geographic range (Tiedje et al., 1989).
Predictors of potential adverse effects of genetically engineered algae include probability of release, abundance of organisms released (predictor of establishment), survival rate and fitness, reproduction rate, probability of dissemination to distant sites, interactions with other organisms, probability of genetic exchange, and probability of an adverse effect (Alexander, 1985). New traits potentially can influence these factors, but few of these relationships are understood. Cell density in the culture medium could be affected by engineered traits. The scale and frequency of releases might determine whether the release leads to a self-sustaining (established) population (Tiedje et al., 1989). The survival rate of a genetically engineered microalga or cyanobacterium will be determined by a combination of the species identity, the genetic modification(s), and the environment to which it is released. Algae with high lipid content probably will be more attractive to predators. Some researchers suggest that most genetically engineered organisms will have lower fitness in receiving environments than unmodified organisms (Tiedje et al., 1989). Algae could be cross-bred or engineered to have high growth rates under specific culture conditions, and some of these might have high growth rates under specific natural conditions. New traits conferred on algae by genetic modifications would determine whether and how community interactions might be altered. Radakovits et al. (2010) pointed out that it is uncertain how genetically engineered strains will perform in scaled-up production systems with varying conditions and with wild-type competitive strains.
Genetic exchange might lead to unexpected effects. Snow et al. (2005) asserted that genetic exchange between recombinant microbes and indigenous microbes is probable. The three types of horizontal transfer are transformation of free deoxyribonucleic acid (DNA), conjugation, and transduction. The transfer of genes between microorganisms is common in some species (Snow et al., 2005). About 1 to 20 percent of the genomes of bacteria consist of DNA acquired recently (in an evolutionary context), predominantly from other prokaryotes but also from eukaryotes, for example, metazoa (Ochman et al., 2000; Koonin et al., 2002; Snow et al., 2005). Genetic exchange between prokaryotes and eukaryotes is not well studied (Rogers et al., 2007) beyond specific pairwise interactions such as T-DNA transfer from Agrobacterium species to plant cells (Gelvin, 2010). Some ancient, evolutionary scale transfers have been recorded. For example, a gene for plastidtargeted fructose bisphosphate aldolase was transferred from red algae to some Prochlorococcus and Synechococcus species (Rogers et al., 2007). Another study, for example, showed evidence for the horizontal transfer of a self-splicing, homing intron from a cyanobacterium
(Calothrix) to the chloroplast genome of Euglena myxocylindracea (Sheveleva and Hallick, 2004). Gene transfer between dissimilar organisms is possible, though rare. There is evidence that nuclear genes encoding chloroplast proteins have been transferred from an alga to an ascoglossan sea slug that consumes the algae (Pierce et al., 2003). Horizontally transferred genes can code for selectable traits, such as antibiotic resistance, pathogenicity, and metabolic enzymes (Snow et al., 2005). Horizontal gene transfer depends on the density of organisms with which exchange is possible.
The principal adverse effects of any algae, whether genetically engineered or not, could include health and ecological effects from toxin production, ecological effects from blooms, and species replacement. These potential effects are discussed elsewhere in this report. The potential propagation of antibiotic resistance markers also would be a concern, but species containing these markers would unlikely be used for commercial-scale production of biofuels. Toxin production by genetically engineered algae is unlikely because toxin-producing strains would be avoided, or strains probably would be engineered to remove toxin genes. A genetically engineered strain might have a lower risk of adverse impact than a natural strain that has not had such modifications. Categories of potential ecological risks from genetically engineered organisms that were highlighted by Tiedje et al. (1989) and Snow et al. (2005) would need to be considered in assessments of released algae. These include creating new or more effective pathogens, affecting nontarget species, disrupting biotic communities and ecosystems, reducing biodiversity or species-genetic diversity, or degrading valuable biological resources, many of which are discussed in the section on invasive species. Little evidence is available to evaluate the potential for any of these effects. Species that are genetically engineered to become more tolerant of environmental stressors, such as salt or temperature, could bloom in habitat conditions where blooms previously have not occurred. Species replacement is a potentially delayed effect (Tiedje et al., 1989). Whether exposure to genetically engineered organisms or genetic exchange with these organisms poses any potential hazards depends on the particular traits of the organism (Snow et al., 2005).
Most approaches to risk assessment suggest that familiarity with genetically engineered organisms is an important predictor of risk (Efroymson, 1999). That is, genetically engineered organisms that have a history of safe use in applications similar to proposed uses (for example, at similar densities in similar ecosystems) would not be likely to threaten environmental sustainability of algal biofuels. Similarly, microorganisms that are not developed from dissimilar source organisms but rather are created from closely related organisms (see EPA, 1990) are less likely to have new traits and to cause adverse effects.
5.8.2 Social Acceptability of Genetically Engineered Algae
If algal biofuel companies are moving toward the use of genetically engineered algae, popular and political resistance could be anticipated. Some concerns over genetically engineered algae depend on the capability of these algae to survive and invade natural environments (as in the case of invasive algae) outside a production environment where temperature, nutrient loads, salinity, and pH all can be optimized. People have expressed concerns regarding the release of genetically engineered microorganisms, ranging from impacts of large-scale releases (and failure of control mechanisms) on biodiversity to ecosystem and evolutionary processes (Hagedorn and Allender-Hagedorn, 1997). Other concerns regarding genetic technologies relate to the unnaturalness of organisms (Tenbult et al., 2005; Connor and Siegrist, 2011); these cannot be abated through technical mitigation. It is the public
perception of risk, and not necessarily the scientific basis for risk, that will be preeminent in determining acceptability of genetically engineered algae to communities. Concerns over genetically engineered algae and perceptions of risks associated with introducing nonnative species into new geographies will need to be addressed.
While concerns surrounding the use of genetically engineered algae for energy production are likely to be raised as the industry continues to develop, the United States remains one of the most accepting countries in the developed world in terms of the adoption rates of genetically engineered crops (USDA-ERS, 2011). In 2008, U.S. farmers planted more than 32 million acres of Monsanto’s “triple-stack” genetically engineered corn, and it is estimated that this number will increase to approximately 56 million acres by 2015 (Kaskey, 2009).
Nonetheless, social acceptability of gene technology depends on the type of application. In one study, medical applications were perceived to be more beneficial, less hazardous, and more ethical than food applications (Frewer et al., 1995). In a Swiss study, lay people distinguished between acceptability of medical and non-medical applications of gene technology but not among agricultural, nutritional, and industrial applications (Connor and Siegrist, 2011). Further, prevailing concerns over the use of genetically engineered crops in the United States are related to human health and food safety rather than potential ecological risks (Kamaldeen and Powell, 2000). However, concerns about genetically engineered microorganisms in surveys and in the popular press have related more to environmental effects than to health or ethical issues (Hagedorn and Allender-Hagedorn, 1997), and these concerns might be expected to dominate for microalgae. Coproduct markets such as health supplements, food additives, and cosmetics could attract additional scrutiny from consumers. It is unknown whether the U.S. public may be more tolerant of the use of biofuels from genetically engineered algae as an energy source than if the crops were grown for food.
Social acceptability of a new technology also depends on how a decision is framed. For example, Wolfe and Bjornstad (2003) suggested that options regarding the use of genetically engineered organisms for hazardous waste remediation likely would be presented in the context of multiple technology options. It is less likely that stakeholders evaluating the use of genetically engineered algae in their regions would be explicitly weighing the relative benefits and risks of different liquid fuels produced elsewhere.
Social acceptability of gene technology depends on trust (Siegrist, 2000). Whether the public is more willing to accept the use of naturally occurring algal strains than those that have been genetically altered for maximum fuel production might depend on the engagement of managers of the facility, other stakeholders, and the public.
5.8.3 Opportunities for Mitigation
Containment of genetically engineered algae might be desirable as a precaution against unknown effects and societal concerns. Physical containment of released algae will be difficult or impossible. Physical containment solutions, such as those proposed for vascular plants (for example, fences and border plants; Moon et al., 2010), are ineffective against released algae. Containment options might include using species that require saline water in freshwater environments or those that have a nutrient requirement that is not met outside of the photobioreactor or pond. The use of environment-dependent “molecular switches” has been proposed to increase the likelihood of community acceptance of genetically engineered crops (Chapotin and Wolt, 2007). Similarly, some modified traits could reduce fitness in natural environments. For example, reduced light harvesting antennae not only would increase growth in ponds but also would reduce the ability to compete with native algal
species for light in natural waters (Sayre, 2011). Moreover, removing bicarbonate pumps, which increases fitness in the cultivation environment with high CO2, could reduce the competitive ability to take up inorganic carbon in natural waters (Sayre, 2011). If algicides are used, they would be effective against non-target organisms. Terminator genes that cause released cells to die could be developed so that those genes would be suppressed in a photobioreactor but derepressed in natural environments (Sayre, 2011). If algal strains that cannot produce toxins are used, potential risk is minimized. If genetically engineered organisms are released, monitoring should be undertaken so that effects of particular organism-environment combinations can be better understood (Snow et al., 2005).
5.8.4 Sustainability Indicators
Sustainability indicators for genetically engineered algae generally would be the same as indicators for native algae. That is, if effects of concern include biodiversity or water quality, appropriate metrics are described in those chapters. However, the sustainability goals for genetically engineered algae likely would include two other issues:
• Minimizing dissemination of genetically engineered algae.
• Establishing methods to determine whether an observed effect was caused by a genetically engineered alga.
Abundance of genetically engineered algae released to water could be measured through species-specific tests if the species were not native. Moreover, some modified traits, such as altered antennae, might be detectable microscopically and thus quantifiable in water. Particular DNA sequences also might be detectable. Moreover, markers could be added to algae to allow easy measurement in specific media.
5.8.5 Information and Data Gaps
The ecological risks of a release of genetically engineered microorganisms have to be carefully assessed before they are used in commercial-scale algal biofuel production. Whether there are plausible scenarios under which genetically engineered algae, or organisms that acquire genes from the genetically engineered algae, could proliferate to levels that might harm humans or the environment in some way needs to be examined. More information is needed on potential relationships between traits that are targets for modification and behavior of cyanobacteria or microalgae that could alter rates of release, survival, growth, transport, genetic exchange, and ecological or human health effects. Little research to date has been conducted in the United States on behavior of genetically engineered algae in open ponds, in part because EPA notification guidelines can lead to delays for researchers. Information is needed on the social acceptability of the use of genetically engineered algae for biofuels, particularly in open systems.
5.9.1 Potential Environmental Effects
Sustainability of a production process is enhanced by recycling raw materials and minimizing of waste. If the oil-extracted biomass is recycled or made into coproducts, a
source of waste would be reduced or eliminated. Anaerobic digestion is another method of waste disposal and can generate electricity as a coproduct (Chapter 2). For the disposal of waste biomass, blow down of solids from production and recycling ponds, and saltwater, companies are considering landfilling waste, underground injection, and diverting processed water to sewage systems.
Solid waste from algal biofuel manufacturing processes is most likely to be generated as sludge from an anaerobic digester from which the volatile organic acids have been converted to methane and CO2; the methane is useful as a fuel supplement for the process. Anaerobic digestion in many cases is followed by aerobic digestion to convert dissolved solids to sludge, concentrated by settling in the large aerobic settlers. Such systems have been operated commercially for decades and most likely will be incorporated into algal biofuel production + concentrations of digested processes. Golueke et al. (1957) reported average NH4+ sludge in the range of 1600 to 1850 milligrams per liter for anaerobic digestion of algae, which is comparable to some of the high values reported for piggery waste (Sukias and Tanner, 2005; Sukias and Craggs, 2011). Another source of solid waste is the spent synthetic plastic liner from open ponds or closed bioreactors that will need to be disposed periodically.
According to Jim Sears (J. Sears, A2BE Carbon Capture, personal communication on September 22, 2011), who chaired the “Committee on Technical Standards” for the Algal Biomass Organization, “there are as many proposed processes for producing algal biofuels as there are companies.” Thus, whether generation of waste products would be a concern cannot be known until operations at commercial scale are in place and compositions can be ascertained. Maximizing recycling would reduce the need for waste product disposal.
5.9.2 Opportunities for Mitigation
Recycling of nutrients is the obvious mitigation for waste generation. Algenol, an algal biofuel company, plans to recycle seawater waste for cultivation (less than 6 liters of seawater waste per liter of ethanol is produced if photobioreactors last greater than 6 years).
If digested sludge is produced, municipal waste treatment plants usually spread the nutrient-rich waste on designated land, with the benefit of conditioning and nourishing the soil. Algal fuel process sludge from wastewater treatment is not expected to be significantly different. The National Pollutant Discharge Elimination System permitting process governs discharge of sludge; most states have a permitting process under this federal program. The composition of the sludge is monitored to ensure compliance with the permit.
5.9.3 Sustainability Indicators
Many sustainability indicators relevant to waste are described elsewhere, for example, quantifying recycling of nutrients and salinity of ground water. If saline wastewater is injected to groundwater, then sustainability indicators also could include annual volume injected per volume of reservoir per year.
5.9.4 Information and Data Gaps
Information is needed about the types and rate of waste generation for most algal biofuel production processes. When and if processes move toward commercialization, state and local regulations will govern the acceptable disposal of waste, which will necessarily be well characterized by then.
5.10.1 Potential Environmental Effects
5.10.1.1 Algal Toxins
Known toxin-producing strains are not likely to be used in algal biofuel production systems. Indeed, many species have food grade status or are being used as feed in aquaculture. However, some species regarded as benign may in fact produce toxins previously unknown. Examples of these include newly discovered euglenoid toxins (Zimba et al., 2010) and free radical toxins (Moeller et al., 2007). In addition, contaminating toxin-producing algae and cyanobacteria could potentially colonize production systems, especially open ponds.
Human toxins that are produced by cyanobacteria have been found in freshwater, marine, and estuarine organisms and include hepatotoxins, cytotoxins, dermatotoxins, and neurotoxins among others (Smith et al., 2008). Ecotoxicity from algal toxins is observed in fish (Zimba et al., 2001b), shellfish (Lance et al., 2011), or invertebrate herbivores such as Daphnia and shrimp (Zimba et al., 2006; Sarnelle, 2010). Toxins can affect viability, growth, and fecundity of many organisms (Plumley, 1997).
The chemical structures of freshwater toxins are probably more diverse than those of marine toxins, including alkaloids, phosphate esters, macrolides, chlorinated diaryllactones, and penta- and hepta-peptides (Rouhiainen et al., 1995; Smith et al., 2008). Species of toxin-producing algae in the divisions Euglenophyceae, Bacillariophyceae, Dinophyceae, Haptophyceae, and Raphidophyceae have been documented. Some cyanobacteria also are producers of toxins and probably are responsible for the production of most freshwater algal toxins from harmful algal blooms (Plumley, 1997). Toxicity of some compounds can exceed that of curare (Zimba et al., 2001a). Irritants and allergens also are produced by certain algae. Toxin production in some cyanobacteria is influenced by environmental variables and competition (Moeller et al., 2007; Briand et al., 2008), though the physiological and ecological causes of toxin production are largely unknown (Paerl and Millie, 1996; Carrick, 2011). Harmful blooms of toxin-producing algae are not the sole source of algal toxins, nor are algal toxins always associated with blooms (Plumley, 1997). Moreover, blooms cannot be predicted with accuracy in natural environments (Carrick, 2011), and this likely applies to open biofuel cultivation systems as well.
Both freshwater and marine forms of toxin-producing algae could colonize production systems. The current state of knowledge about phytoplankton community composition is not sufficient to predict whether toxin-producing strains could invade and bloom in algal biofuel production systems, even if these systems are seeded either initially or continuously with non-toxigenic algal strains.
Compounds presently not known to be harmful because of their presence in low concentrations in small-scale, low-intensity algal biomass production may have harmful impacts when concentrated 100,000 times during the harvesting and drying phases. Concentrated cultivation methods may lead to the identification of previously unknown toxic material (Moeller et al., 2007; Zimba et al., 2010). If the lipid-extracted algae are to be used in value-added coproducts, the quality of those products would have to be monitored.
The outdoor open-pond production systems likely will develop diverse algal populations (Smith et al., 2010). Monitoring algal composition is critical to maintaining desired
characteristics for processing biomass to fuels, ensuring that coproducts from lipid-extracted algae are safe for use, and minimizing downstream effects of water-soluble toxins.
5.10.1.2 Human and Animal Pathogens in Algal Cultivation Systems
Cultivated Spirulina, Chlorella, and Haematococcus have been used to produce foodgrade products, and the presence of pathogens has not been a concern. However, algal production systems are diverse communities that may contain pathogens particularly if municipal wastewater, wastewater from concentrated animal feeding operations (CAFOs), biosolids (sewage sludge), or manures are used as water or nutrient supplies. Although the algal cultivation systems using wastewater are similar to the thousands of algal wastewater ponds in the United States, different occupational exposures might arise because the algal biomass being handled in algal biofuel production is larger in quantity (that is, higher sludge mass in algae cultivation for fuels than in algal wastewater pond) and higher in concentration (from harvesting and drying before processing to fuels). The density and probability of particular pathogens in wastewater is related to the level of treatment, with greater pathogen numbers and diversity in primary treated sewage than secondary treatment. Primary treatment is the sedimentation of solids, secondary treatment is the removal of suspended and dissolved organic materials, and tertiary treatment is the removal of inorganic constituents such as nitrogen and phosphorus. Biosolids (sewage sludge), for example, may include bacterial, viral, protozoan, or helminth pathogens (EPA, 2011b). In a study of mesophilic anaerobic digested biosolids from 18 locations in the United States, Clostridium perfringens, Shigella, Campylobacter, Salmonella, enteric viruses, and adenoviruses were detected, but Ascaris and Escherichia coli 0157:H7 were not. The original wastewater would be expected to contain at least these species. In another study of treated wastewater and biosolids in Michigan, adenovirus, enterovirus, and norovirus were detected in 100, 70, and 10 percent of samples, respectively (Simmons and Xagoraraki, 2011). The taxonomic identities and abundances of pathogens in biosolids (and by extension, wastewater) are determined by the incidence of infection within the wastewater-generating community and the particular wastewater treatment process used (Straub, 1993; EPA, 2011b). Survival of some pathogens from biosolids in soil has been studied (Zerzghi et al., 2009), but the survival of human and animal pathogens in algal biofuel cultures is only beginning to be investigated.
Where pathogens are present in algal cultures, there could be occupational health effects or environmental effects (if release occurs). The presence of fecal coliforms or other pathogens would limit the options for coproducts.
5.10.2 Comparison of Pathways
The key difference in pathways is that photobioreactors are less likely to be colonized by toxin-producing strains than open ponds (Table 5-11).
5.10.3 Opportunities for Mitigation
Selecting strains known not to produce toxins will mitigate toxin concerns in closed systems and aid in mitigating toxin concerns for open systems, though toxins could be
TABLE 5-11 An Illustration of Potential Effects from Toxins and Pathogens from Different Pathways of Algal Biofuel Production
|
|
|||||
|
Potential Effect |
Pathway | ||||
|
|
|||||
|
Open-pond, salt water, producing biodiesel, recycling nutrients and water |
Open-pond, salt water, producing biodiesel + coproducts |
Open-pond, salt water, producing FAME, recycling nutrients and water |
Photobioreactor, salt water, direct synthesis, recycling water |
Open-pond, salt water, producing biomass, pyrolysis, recycling some nutrients and water |
|
|
|
|||||
|
Toxins in Algal Culture |
Possible if toxin-producing strains colonize production systems |
Possible if toxin-producing strains colonize production systems |
Possible if toxin-producing strains colonize production systems |
Very unlikely. Strain is selected that does not produce toxins, and contamination with toxin-producing strain very unlikely. |
Possible if toxin-producing strains colonize production systems |
|
Pathogens in Algal Culture |
Possible if wastewater is used as nutrient source and possible colonization by environmental pathogens |
Possible if wastewater is used as nutrient source and possible colonization by environmental pathogens; possible movement of spores into coproducts |
Possible if wastewater is used as nutrient source and possible colonization by environmental pathogens |
Possible if wastewater is used as nutrient source |
Possible if wastewater is used as nutrient source, very unlikely colonization by environmental pathogens because of low residence time |
|
|
|||||
produced by non-target algae in the open-pond community. Genomic approaches could be used to screen for genes required for toxin synthesis in candidate algal strains for biofuel production (La Claire, 2006; Ianora et al., 2011). Periodic monitoring could ensure that wellknown toxins are not produced.
Minimizing sources of human and animal pathogens in algal culture could include:
• A high level of treatment or sterilization of wastewater. For example, Phycal passes wastewater through a 0.2 micrometer filter prior to use and uses ultraviolet sterilization for initial treatment as well as treatment of recycled water. Reuse of wastewater for algal biofuel production could follow established wastewater reuse regulations.
• Using agricultural grade fertilizers (for example, Sapphire Energy, Inc.).
• Use of high-pH brines to reduce survival of pathogen competitors of cyanobacteria (for example, Phyco BioSciences).
5.10.4 Sustainability Indicators for Algal Toxins and Pathogens
Indicators of sustainable development of algal biofuels include metrics of algal toxins and pathogens in water, which consist of concentrations of toxins in water, measures of toxic effects (for example, in animal models) that are diagnostic of particular toxins, or
genetic markers of toxin production. Methods to distinguish some toxin-producing strains from other strains are available. For example, an oligonucleotide probe can distinguish hepatotoxic from neurotoxic Anabaena and these strains from Nostoc spp. (Rouhiainen et al., 1995). Information supporting genetic markers of toxin production is increasingly available, for example, a PCR-based test to assess the potential for microcystin occurrence (Nonneman and Zimba, 2002). However, these tests cannot identify unknown toxins, and they can give false-positive results where toxins are not expressed, for example, where multigene families are needed for arrangement (Zimba et al., 2010).
Directly measuring all pathogens in algal culture media is not generally practical. Indicator species are often microorganisms that are nonpathogenic, abundant, and associated with the presence of a suite of pathogens (EPA, 2011b). For example, densities of fecal coliform and Salmonella can be used as indicators for assessing the efficiency of wastewater treatment (40 CFR 136). These can be measured via culture methods or through quantitative polymerase chain reaction tests for fecal indicator bacteria that provide sameday information (Dorevitch et al., 2011). The abundance of fecal indicator bacteria can be related to disparate pathogens, such as protozoa (Dorevitch et al., 2011). However, typical indicator organisms are not useful in all media or for all pathogens. For example, Pepper et al. (2010) found that indicator organisms in Class B biosolids were not correlated with the numbers of pathogenic organisms. Criteria for selecting non-pathogen indicator organisms of pathogens in waters have been summarized by EPA (2011b), based on information in Gerba (2009) and NRC (2004). These include attributes of organisms and testing methods. Analytical methods for detecting low densities of pathogens have not been sufficiently developed and tested to be recommended (EPA, 2011b). Any potential indicators would have to be tested in algal ponds or photobioreactors for potential relationships with pathogen levels. Attributes of non-pathogen indicator organisms of pathogens in waters and attributes of methods for detecting pathogen indicator organisms are described in an EPA report Problem Formulation for Human Health Risk Assessment of Pathogens in Land-applied Biosolids (EPA, 2011b).
5.11.1 Potential Environmental, Health, and Social Acceptability Effects
Health effects from and social acceptability of algal biofuels could be affected if open ponds are poorly managed and provide habitats for mosquito larvae. Photobioreactors and raceways would not represent mosquito habitat unless there is substantial leakage of culture fluid, and puddles are formed. Mosquitoes lay their eggs opportunistically in standing water, which can vary from large lakes to small puddles or buckets. The full area of algae cultivation ponds would not be optimal habitat because of the required stirring and agitation for adequate mixing of nutrients and light exposure. Females of most mosquito species only infrequently lay eggs in flowing or agitated water (Lothrop and Mulla, 1996; Mogi and Motomura, 1996). Moreover, waters that are in motion can interfere with the surface tension required for mosquitoes’ respiratory siphons to function (Schober, 1966). However, any relatively still edges of open ponds (analogous to stream banks and floodplains along moving streams) and outlying puddles or open-water storage vessels would be suitable for the growth of mosquito larvae. Because algae constitute food for mosquito larvae, the high nutrient and carbon content of algal cultivation systems (when and where the water is relatively still) can be prime habitat (Rydzanicz and Lone, 2003). The turbidity
of some cultivation systems would provide refuges from visual predators (Jacob et al., 2008; Jackson et al., 2009). Fewer mosquito species can tolerate saline conditions than fresh water (Patrick and Bradley, 2000), but some species can tolerate salinities of 100 percent sea water (Grueber and Bradley, 1994).
Providing habitat for mosquitoes could be a concern for human health and the acceptability of algal biofuel production for several reasons including:
• Mosquitoes are considered a pest and a nuisance that may not be tolerated by people living near a cultivation facility. Communities near proposed constructed wetland sites sometimes object to siting based on the anticipation of a mosquito problem (Anderson et al., 2007).
• Mosquitoes are vectors for numerous human infectious diseases in the United States, such as Eastern equine encephalitis, La Crosse encephalitis, St. Louis encephalitis, West Nile virus, Western equine encephalitis, and Dengue fever (recently reported in Florida). West Nile virus is also hypothesized to be a factor in the decline of sage grouse (Naugle et al., 2004).
• If the ponds for algae cultivation become breeding grounds for mosquitoes, there is a risk that the larvae will become a pest, reducing algal population densities below economically productive levels through predation.
5.11.2 Opportunities for Mitigation
Measures could be taken to control mosquito and other pest populations in and around algae cultivation ponds. The extensive use of agitators, aerators, and fountains decrease the suitability of open ponds for mosquito habitat (Jackson et al., 2009) and distribute nutrients and algae in the system. If standing water at cultivation facilities is minimized, mosquitoes and associated health effects should not be a problem.
Other mitigation options include site-specific surveys that can inform mosquito management agencies regarding the timing, species, and abundance of mosquitoes to develop disease-reduction plans (Anderson et al., 2007). Control options include chemical treatments like insecticides and biological methods such as the introduction of natural predators such as mosquitofish that consume mosquito larvae. If some of these measures are used without prior consultation and acceptance by the public, or if it is perceived that a population control method poses a threat to the human health or well-being, local communities might not accept algae production as a viable source of energy.
5.11.3 Sustainability Indicators
The sustainability indicators for mosquito-borne diseases are density of mosquito larvae in ponds and changes in incidence of mosquito-borne diseases attributable to cultivation ponds.
Reducing GHG emissions from the transportation sector has been one of the primary motivations for using alternative liquid transportation fuels. Therefore, the life-cycle GHG emissions are key factors in considering the sustainable development of algal biofuels. Published estimates of GHG emissions span a wide range, with some studies suggesting that algal biofuel production has high GHG emissions. The utility of these LCAs is that they point out key drivers of CO2 emissions in the algal biofuel supply chains and indicate
the aspects or processes that could benefit from research and development for improving GHG emissions.
Some concerns of medium importance to consider include:
• The presence of waterborne toxicants or pathogens in algal cultivation systems if waste streams (flue gas or wastewater) are to be used as sources of nutrients or water. Their presence would affect occupational safety and the safety of coproducts if the residual algal biomass is used to produce certain coproducts to maximize recycling and to improve process economics.
• Effects from land-use changes if pasture and rangeland are to be converted to algae cultivation. Displacing pasture and rangeland could incur direct and indirect landuse changes that would affect the net GHG emissions of algal biofuels.
• Air quality emissions over the life cycle of algal biofuels. Emissions from the pro-cessing facilities and tailpipe emissions will be regulated, but emissions from other parts of the supply chain also need to be considered. The committee is not aware of any published studies that include measured emissions of air pollutants from open-pond cultivation.
• Potential effects on local climate. The introduction of large-scale algal cultivation systems in arid or semi-arid environments could alter the local climate of the area by increasing humidity and altering temperature extremes.
• Releases of cultivated algae to natural environments and potential alteration of species composition in receiving waters.
• Effects on terrestrial biodiversity from changing landscape pattern as a result of infrastructure development for algal biofuels.
• Potential adverse effects and unintended consequences of introduction of genetically engineered algae for biofuel production.
• Waste products from processing algae to fuels.
This chapter discussed the potential environmental effects of algal biofuel production. Some of those effects require assessment and monitoring to ensure that they do not pose serious sustainability concerns (for example, potential land conversion, air emissions, effects on biodiversity, waste products from algal biofuel production systems, and potential presence of pathogens and unknown or unidentified toxins). Other environmental effects discussed could be avoided with proper management and good engineering designs (for example, release of culture water leading to eutrophication, seepage of culture water into local ground water, and habitats for mosquito larvae).
SUMMARY FINDINGS FROM THIS AND EARLIER CHAPTERS
Algal biofuels have the potential to contribute to improving the sustainability of the transportation sector, but the potential is not yet realized. Additional innovations that require research and development are needed to realize the full potential of algal biofuels. (See Chapters 2 and 3 for biological and engineering innovations needed and Chapter 4 for resource recycling.)
Engineering solutions to enhance algae cultivation, to facilitate biomass or product collection, and to improve processing of algae-derived fuels can increase the EROI and reduce the GHG emissions of algal biofuel production. (See Chapters 2 and 3 for engineering solutions and Chapter 4 for a discussion on EROI.)
40 CFR 136 (Code of Federal Regulations). TITLE 40—Protection of Environment; PART 136—Guidelines Establishing Test Procedures for the Analysis of Pollutants, U.S. Government. No. 40. Available online at http://ecfr.gpoaccess.gov/cgi/t/text/text-idx?c=ecfr&tpl=/ecfrbrowse/Title40/40cfr136_main_02.tpl. Accessed July 20, 2012.
Ahluwalia, S.S., and D. Goyal. 2007. Microbial and plant derived biomass for removal of heavy metals from wastewater. Bioresource Technology 98(12):2243-2257.
Alexander, M. 1971. Biochemical ecology of microorganisms. Annual Review of Microbiology 25:361-392.
____________. 1977. Soil Microbiology. 2nd Edition. New York: John Wiley and Sons.
____________. 1985. Ecological consequences—Reducing the uncertainties. Issues in Science and Technology 1(3):57-68.
An, J.Y., S.J. Sim, J.S. Lee, and B.W. Kim. 2003. Hydrocarbon production from secondarily treated piggery waste-water by the green alga Botryococcus braunii. Journal of Applied Phycology 15(2-3):185-191.
Anderson, A.L., K. O’Brien, and M. Hartwell. 2007. Comparisons of mosquito populations before and after construction of a wetland for water quality improvement in Pitt County, North Carolina, and data-reliant vectorborne disease management. Journal of Environmental Health 69(8):26-33.
Assmann, A., A. Braun, S. John, A. Lei, and S. Southard. 2011. The Potential for Micro-Algae and Other “Micro-Crops” to Produce Sustainable Biofuels. Master of Science, University of Michigan, Ann Arbor.
Atkinson, K.M. 1972. Birds as transporters of algae. British Phycological Journal 7:319-321.
Babcock, B.A. 2009. Measuing unmeasurable land-use changes from biofuels. Iowa Agriculture Review Summer:4-6.
Bachmann, R.W., B.L. Jones, D.D. Fox, M. Hoyer, L.A. Bull, and D.E. Canfield. 1996. Relations between trophic state indicators and fish in Florida (USA) lakes. Canadian Journal of Fisheries and Aquatic Sciences 53(4):842-855.
Beeman, P. 2007. Biofuel plants generate new air, water, soil problems for Iowa. Available online at http://www.desmoinesregister.com/article/20070603/BUSINESS01/706030325/Biofuel-plants-generate-new-air-water-soil-problems-for-Iowa. Accessed January 21, 2012.
Benemann, J.R. 2008. Opportunities and challenges in algae biofuel production. Available online at http://www.fao.org/uploads/media/algae_positionpaper.pdf. Accessed October 21, 2011.
BLM (Bureau of Land Management) and DOE (U.S. Department of Energy). 2010. Solar Energy Development Draft Programmatic Environmental Impact Statement (Draft Solar PEIS). Washington, DC: Bureau of Land Management and U.S. Department of Energy.
Bollag, J.M., and S.T. Czlonkowski. 1973. Inhibition of methane formation in soil by various nitrogen-containing compounds. Soil Biology and Biochemistry 5(5):673-678.
Boyd, C.E. 1995. Bottom soils, sediment, and pond aquaculture. New York: Chapman and Hall.
Breitburg, D.L., D.W. Hondorp, L.A. Davias, and R.J. Diaz. 2009. Hypoxia, nitrogen, and fisheries: Integrating effects across local and global landscapes. Pp. 329-349 in Annual Review of Marine Science.
Briand, E., C. Yepremian, J.F. Humbert, and C. Quiblier. 2008. Competition between microcystin- and non-microcystin-producing Planktothrix agardhii (cyanobacteria) strains under different environmental conditions. Environmental Microbiology 10(12):3337-3348.
Buntjer, J. 2010. Heron Lake BioEnergy fined. Daily Globe, December 27. State and Regional News section.
Burkholder, J.M., K.M. Mason, and H.B. Glasgow. 1992. Water-column nitrate enrichment promotes decline of eelgrass Zostera-Marina—Evidence from seasonal mesocosm experiments. Marine Ecology-Progress Series 81(2):163-178.
Burns, J.C. 2008. ASAS centennial paper: Utilization of pasture and forages by ruminants: A historical perspective. Journal of Animal Science 86(12):3647-3663.
![]() ada, G.F. 1998. Efforts to reduce the impacts of hydroelectric power production on reservoir fisheries in the United States. International Review of Hydrobiology 83(Special Issue):43-50.
ada, G.F. 1998. Efforts to reduce the impacts of hydroelectric power production on reservoir fisheries in the United States. International Review of Hydrobiology 83(Special Issue):43-50.
Campbell, P.K., T. Beer, and D. Batten. 2011. Life cycle assessment of biodiesel production from microalgae in ponds. Bioresource Technology 102(1):50-56.
Campbell, S.P., J.A. Clark, L.H. Crampton, A.D. Guerry, L.T. Hatch, P.R. Hosseini, J.J. Lawler, and R.J. O’Connor. 2002. An assessment of monitoring efforts in endangered species recovery plans. Ecological Applications 12(3):674-681.
CARB (California Air Resource Board) 2010. Low Carbon Fuel Standard—Indirect Effects. Sacramento: California Environmental Protection Agency.
Carlsen, T.M., J.D. Coty, and J.R. Kercher. 2004. The spatial extent of contaminants and the landscape scale: An analysis of the wildlife, conservation biology, and population modeling literature. Environmental Toxicology and Chemistry 23(3):798-811.
Carpenter, S.R. 2005. Eutrophication of aquatic ecosystems: Bistability and soil phosphorus. Proceedings of the National Academy of Sciences of the United States of America 102(29):10002-10005.
Carrick, H.J. 2011. Niche modeling and predictions of algal blooms in aquatic ecosystems. Journal of Phycology 47(4):709-713.
Chapotin, S.M., and J.D. Wolt. 2007. Genetically modified crops for the bioeconomy: Meeting public and regulatory expectations. Transgenic Research 16(6):675-688.
Chinnasamy, S., A. Bhatnagar, R.W. Hunt, and K.C. Das. 2010. Microalgae cultivation in a wastewater dominated by carpet mill effluents for biofuel applications. Bioresource Technology 101(9):3097-3105.
Clarens, A.F., E.P. Resurreccion, M.A. White, and L.M. Colosi. 2010. Environmental life cycle comparison of algae to other bioenergy feedstocks. Environmental Science and Technology 44(5):1813-1819.
Cloud, B. 2011. Questionnaire reply from Phyco BioSciences Inc. Received by the NRC Committee on Sustainable Development of Algal Biofuels on July 25.
Codd, G.A. 2000. Cyanobacterial toxins, the perception of water quality, and the prioritisation of eutrophication control. Ecological Engineering 16(1):51-60.
Connor, M., and M. Siegrist. 2011. The power of association: Its impact on willingness to buy GM food. Human and Ecological Risk Assessment 17(5):1142-1155.
Dale, V.H., K.L. Kline, L.L. Wright, R.D. Perlack, M. Downing, and R.L. Graham. 2011. Interactions among bioenergy feedstock choices, landscape dynamics, and land use. Ecological Applications 21(4):1039-1054.
Dale, V.H., and S. Pearson. 1997. Quantifying habitat fragmentation due to land use change in Amazonia. Pp. 400-410 in Tropical Forest Remnants: Ecology, Management, and Conservation of Fragmented Communities, W.F. Laurance and R.O. Bierregaard, eds. Chicago, IL: University of Chicago Press.
Davis, R., A. Aden, and P.T. Pienkos. 2011. Techno-economic analysis of autotrophic microalgae for fuel production. Applied Energy 88(10):3524-3531.
Deegan, L.A., A. Wright, S.G. Ayvazian, J.T. Finn, H. Golden, R.R. Merson, and J. Harrison. 2002. Nitrogen loading alters seagrass ecosystem structure and support of higher trophic levels. Aquatic Conservation-Marine and Freshwater Ecosystems 12(2):193-212.
Degu, A.M., F. Hossain, D. Niyogi, R. Pielke, J.M. Shepherd, N. Voisin, and T. Chronis. 2011. The influence of large dams on surrounding climate and precipitation patterns. Geophysical Research Letters 38. DOI: 10.10.29/2010GLO46482.
Demirbas, A. 2009. Production of biodiesel from algae oils. Energy Sources Part A—Recovery Utilization and Environmental Effects 31(2):163-168.
Dickens, M.J., D.J. Delehanty, J.M. Reed, and L.M. Romero. 2009. What happens to translocated game birds that “disappear”? Animal Conservation 12(5):418-425.
DOE (U.S. Department of Energy). 2009. Construction and Operation of a Proposed Cellulosic Ethanol Plant, Range Fuels Soperton Plant, LLC (formerly Range Fuels Inc.) Treutlen County, Georgia. Washington, DC: U.S. Department of Energy.
____________. 2010a. Algenol Integrated Biorefinery for Producing Ethanol from Hybrid Algae. Washington, DC: U.S. Department of Energy.
____________. 2010b. National Algal Biofuels Technology Roadmap. Washington, DC: U.S. Department of Energy, Energy Efficiency and Renewable Energy.
Dorevitch, S., M. Doi, F.C. Hsu, K.T. Lin, J.D. Roberts, L.C. Liu, R. Gladding, E. Vannoy, H. Li, M. Javor, and P.A. Scheff. 2011. A comparison of rapid and conventional measures of indicator bacteria as predictors of waterborne protozoan pathogen presence and density. Journal of Environmental Monitoring 13(9):2427-2435.
Drake, L.A., F.C. Dobbs, and R.C. Zimmerman. 2003. Effects of epiphyte load on optical properties and photosynthetic potential of the seagrasses Thalassia testudinum Banks ex Konig and Zostera marina L. Limnology and Oceanography 48(1):456-463.
Drewes, J., P. Xue, D. Heil, and G. Wang. 2009. Multibeneficial Use of Produced Water Through High-Pressure Membrane Treatment and Capacitive Deionization Technology. Denver, CO: U.S. Department of the Interior Bureau of Reclamation.
Dufey, A. 2006. Biofuels production, trade and sustainable development: Emerging issues. London: International Institute for Environment and Development (IIED).
Duff, M.C., C. Amrhein, and G. Bradford. 1997. Nature of uranium contamination in the agricultural drainage water evaporation ponds of the San Joaquin Valley, California, USA. Canadian Journal of Soil Science 77(3):459-467.
Eaton, A.D., L.S. Clesceri, E.W. Rice, A.E. Greenberg, and M.A.H. Franson, eds. 2005. Standard Methods for the Examination of Water and Wastewater, 21st ed. Washington, DC: American Public Health Association, American Water Works Association, and Water Environment Federation.
Efroymson, R., H. Jager, V. Dale, and J. Westervelt. 2009. A framework for developing management goals for species at risk with examples from military installations in the United States. Environmental Management 44(6):1163-1179.
Efroymson, R.A. 1999. Regulating risk: Oversight of microbial products of biotechnology under the Toxic Substances Control Act. Journal of Environmental Assessment Policy and Management 1:329-347.
Efroymson, R.A., D.S. Jones, and A.J. Gold. 2007. An ecological risk assessment framework for effects of onsite wastewater treatment systems and other localized sources of nutrients on aquatic ecosystems. Human and Ecological Risk Assessment 13(3):574-614.
Ehresmann, D.W., and M.T. Hatch. 1975. Effect of relative humidity on survival of airborn unicellular algae. Applied Microbiology 29(3):352-357.
EIA (U.S. Energy Information Adminstration). 2002. Updated State-Level Greenhouse Gas Emission Coefficients for Electricity Generation 1998-2000. Washington, DC: U.S. Energy Information Adminstration.
Environmnetal Law Institute. 2012. Glossary of brownfields terms. Available online at http://www.brownfieldscenter.org/big/glossary.shtml. Accessed August 17, 2012.
EPA (U.S. Environmental Protection Agency). 1990. Points to consider in the preparation and submission of TSCA Premanufacture Notices (PMNs) for microorganisms. Washington, DC: U.S. Environmental Protection Agency.
____________. 2002a. Clean alternative fuels: Biodiesel. Available online at http://www.afdc.energy.gov/afdc/pdfs/epa_biodiesel.pdf. Accessed February 1, 2012.
____________. 2002b. Clean alternative fuels: Ethanol. Available online at http://www.afdc.energy.gov/afdc/pdfs/epa_ethanol.pdf. Accessed February 1, 2012.
____________. 2002c. Clean alternative fuels: Fischer Tropsch. Available online at http://www.afdc.energy.gov/afdc/pdfs/epa_fischer.pdf. Accessed February 1, 2012.
____________. 2007. Advanced wastewater treatment to achieve low concentration of phosphorus. Seattle, WA: U.S. Environmental Protection Agency, Region 10, Office of Water and Watersheds.
____________. 2009a. Emsisions standardard reference guide. Available online at http://www.epa.gov/otaq/standards/basicinfo.htm#3. Accessed January 13, 2012.
____________. 2009b. Ethanol Plant Clean Air Act Enforcement Initiative. Available online at http://www.epa.gov/oecaerth/resources/cases/civil/caa/ethanol/index.html. Accessed February 14, 2011.
____________. 2010. Available and Emerging Technologies for Reducing Greenhouse Gas Emissions from Coal-Fired Electric Generating Units. Washington, DC: U.S. Environmental Protection Agency.
____________. 2011a. Clean Water Act. Available online at http://cfpub.epa.gov/npdes/cwa.cfm?program_id=45. Accessed April 18, 2012.
____________. 2011b. Problem Formulation for Human Health Risk Assessments of Pathogens in Land-applied Biosolids. Cincinnati, OH: National Center for Environmental Assessment.
____________. 2012. Integrated Risk Information System. Available online at http://www.epa.gov/IRIS/. Accessed February 17, 2012.
Evaporation Ponds Technical Committee. 1999. Evaporation Ponds. Available online at http://www.water.ca.gov/pubs/groundwater/evaporation_ponds_final_report__san_joaquin_valley_drainage_implementation_program/06-evapponds.pdf. Accessed October 10, 2011.
Fagerstone, K.D., J.C. Quinn, T.H. Bradley, S.K. De Long, and A.J. Marchese. 2011. Quantitative measurement of direct nitrous oxide emissions from microalgae cultivation. Environmental Science and Technology 45(21):9449-9456.
Fargione, J., J. Hill, D. Tilman, S. Polasky, and P. Hawthorne. 2008. Land clearing and the biofuel carbon debt. Science 319(5867):1235-1238.
Fargione, J.E., T.R. Cooper, D.J. Flaspohler, J. Hill, C. Lehman, T. McCoy, S. McLeod, E.J. Nelson, K.S. Oberhauser, and D. Tilman. 2009. Bioenergy and wildlife: Threats and opportunities for grassland conservation. BioScience 59(9):767-777.
Farrell, A.E. 2006. Ethanol can contribute to energy and environmental goals. Science 312(5781):1748-1748.
Fiala, N. 2008. Measuring sustainability: Why the ecological footprint is bad economics and bad environmental science. Ecological Economics 67(4):519-525.
Fleskes, J.P., R.L. Jarvis, and D.S. Gilmer. 2003. Selection of flooded agricultural fields and other landscapes by female northern pintails wintering in Tulare Basin, California. Wildlife Society Bulletin 31(3):793-803.
Fletcher, R.J., B.A. Robertson, J. Evans, P.J. Doran, J.R.R. Alavalapati, and D.W. Schemske. 2011. Biodiversity conservation in the era of biofuels: Risks and opportunities. Frontiers in Ecology and the Environment 9(3):161-168.
Florez-Leiva, L., E. Tarifeño, M. Cornejo, R. Kiene, and L. Farías. 2010. High production of nitrous oxide (N2O), methane (CH4) and dimethylsulphoniopropionate (DMSP) in a massive marine phytoplankton culture. Biogeosciences Discussions 7:6705-6723.
Franzluebbers, A.J. 2010. Achieving soil organic carbon sequestration with conservation agricultural systems in the southeastern United States. Soil Science Society of America Journal 74(2):347-357.
Frewer, L.J., C. Howard, and R. Shepherd. 1995. Genetic engineering and food: What determines consumer acceptance? British Food Journal 97(8):31-36.
Gao, S., K.K. Tanji, R.A. Dahlgren, J. Ryu, M.J. Herbel, and R.M. Higashi. 2007. Chemical status of selenium in evaporation basins for disposal of agricultural drainage. Chemosphere 69(4):585-594.
Garvin, J.C., C.S. Jennelle, D. Drake, and S.M. Grodsky. 2011. Response of raptors to a windfarm. Journal of Applied Ecology 48(1):199-209.
Gelvin, S.B. 2010. Plant proteins involved in Agrobacterium-mediated genetic transformation. Annual Review of Phytopathology 48:45-68.
Gerba, C.P. 2009. Environmental indicators. in Environmental Microbiology, 2nd edition, R.M. Maier, I.L. Pepper and C.P. Gerba, eds. NY: Academic Press. Pp. 485-498.
Gerber, L.R., and L.T. Hatch. 2002. Are we recovering? An evaluation of recovery criteria under the U.S. Endangered Species Act. Ecological Applications 12(3):668-673.
Gibbs, H.K., M. Johnston, J.A. Foley, T. Holloway, C. Monfreda, N. Ramankutty, and D. Zaks. 2008. Carbon payback times for crop-based biofuel expansion in the tropics: The effects of changing yield and technology. Environmental Research Letters 3(3):034001.
Giese, B., F. Laturnus, F.C. Adams, and C. Wiencke. 1999. Release of volatile iodinated C1-C4 hydrocarbons by marine macroalgae from various climate zones. Environmental Science and Technology 33(14):2432-2439.
Ginnebaugh, D.L., J. Liang, and M.Z. Jacobson. 2010. Examining the temperature dependence of ethanol (E85) versus gasoline emissions on air pollution with a largely-explicit chemical mechanism. Atmospheric Environment 44(9):1192-1199.
Golueke, C.G., W.J. Oswald, and H.B. Gotass. 1957. Anarobic digestion of algae. Applied Microbiology and Biotechnology 5:47-55.
Gong, Y.M., and M.L. Jiang. 2011. Biodiesel production with microalgae as feedstock: From strains to biodiesel. Biotechnology Letters 33(7):1269-1284.
Gonzalez, L.E., R.O. Canizares, and S. Baena. 1997. Efficiency of ammonia and phosphorus removal from a Colombian agroindustrial wastewater by the microalgae Chlorella vulgaris and Scenedesmus dimorphus. Bioresource Technology 60(3):259-262.
Graham, L.A., S.L. Belisle, and C.L. Baas. 2008. Emissions from light duty gasoline vehicles operating on low blend ethanol gasoline and E85. Atmospheric Environment 42(19):4498-4516.
Grant, J. 2010. Coastal communities, participatory research, and far-field effects of aquaculture. Aquaculture Environment Interactions 1(2):85-93.
Green, F.B., T.J. Lundquist, and W.J. Oswald. 1995. Energetics of advanced integrated waste-water pond systems. Water Science and Technology 31(12):9-20.
Gressel, J. 2008. Transgenics are imperative for biofuel crops. Plant Science 174(3):246-263.
Grönblad, R. 1933. Contribution to the knowledge of sub-aërial Desmids. Societas Scientiarum Fennica. Commentationes Biologicae 4:1-10.
Grueber, W.B., and T.J. Bradley. 1994. The evolution of increased salinity tolerance in larvae of aedes mosquitos—A phylogenetic analysis. Physiological Zoology 67(3):566-579.
Guikema, S.D. 2009. Infrastructure design issues in disaster-prone regions. Science 323(5919):1302-1303.
Gurian-Sherman, D. 2011. Global Warming and Pasture-Raised Beef Production in the United States. Cambridge, MA: Union of Concerned Scientists.
Hagedorn, C., and S. Allender-Hagedorn. 1997. Issues in agricultural and environmental biotechnology: Identifying and comparing biotechnology issues from public opinion surveys, the popular press and technical/regulatory sources. Public Understanding of Science 6(3):233-245.
Hauxwell, J., J. Cebrian, and I. Valiela. 2003. Eelgrass Zostera marina loss in temperate estuaries: Relationship to land-derived nitrogen loads and effect of light limitation imposed by algae. Marine Ecology-Progress Series 247:59-73.
The H. John Heinz III Center for Science and the Environment. 2008. The State of the Nation’s Ecosystems 2008: Measuring the Lands, Waters, and Living Resources of the United States. Washington, DC: The Heinz Center.
Hess, P., M. Johnston, B. Brown-Steiner, T. Holloway, J. de Andrade, and P. Artaxo. 2009. Air quality issues associated with biofuel production and use. Pp. 169-194 in Biofuels: Environmental Consequences and Interactions with Changing Land Use., R.W. Howarth and S. Bringezu, eds. Ithaca: Cornell University.
Hill, J., S. Polasky, E. Nelson, D. Tilman, H. Huo, L. Ludwig, J. Neumann, H.C. Zheng, and D. Bonta. 2009. Climate change and health costs of air emissions from biofuels and gasoline. Proceedings of the National Academy of Sciences of the United States of America 106(6):2077-2082.
Hoffman, L. 1994. Biogeography of marine blue-green algae. Archiv für Hydrobiologie Supplement 75:137-148.
Hoffmann, J.P. 1998. Wastewater treatment with suspended and nonsuspended algae. Journal of Phycology 34(5):757-763.
Hoffmann, L. 1996. Geographic distribution of freshwater blue-green algae. Hydrobiologia 336(1-3):33-40.
Holloran, M.J., R.C. Kaiser, and W.A. Hubert. 2010. Yearling greater sage-grouse response to energy development in Wyoming. Journal of Wildlife Management 74(1):65-72.
Honkanen, T., and H. Helminen. 2000. Impacts of fish farming on eutrophication: Comparisons among different characteristics of ecosystem. International Review of Hydrobiology 85(5-6):673-686.
Hughes, M.R. 2003. Regulation of salt gland, gut and kidney interactions. Comparative Biochemistry and Physiology—A Molecular and Integrative Physiology 136(3):507-524.
Huntley, B., R. Baxter, K.J. Lewthwaite, S.G. Willis, and J.K. Adamson. 1998. Vegetation responses to local climatic changes induced by a water-storage reservoir. Global Ecology and Biogeography Letters 7(4):241-257.
Ianora, A., M.G. Bentley, G.S. Caldwell, R. Casotti, A.D. Cembella, J. Engstrom-Ost, C. Halsband, E. Sonnenschein, C. Legrand, C.A. Llewellyn, A. Paldaviciene, R. Pilkaityte, G. Pohnert, A. Razinkovas, G. Romano, U. Tillmann, and D. Vaiciute. 2011. The relevance of marine chemical ecology to plankton and ecosystem function: An emerging field. Marine Drugs 9(9):1625-1648.
Irenee-Marie, F. 1938. Flora Desmidiale de la Region de Montreal. LaPrairie, Canada.
ISSG (Invasive Species Specialist Group). 2012. Global Invasive Species Database. Available online at http://www.issg.org/database/species/ecology.asp?si=775.
Jackson, M.J., J.L. Gow, M.J. Evelyn, N.E. Meikleham, T.J.S. McMahon, E. Koga, T.J. Howay, L. Wang, and E. Yan. 2009. Culex mosquitoes, West Nile virus, and the application of innovative management in the design and management of stormwater retention ponds in Canada. Water Quality Research Journal of Canada 44(1):103-110.
Jacob, B.G., E.J. Muturi, E.X. Caamano, J.T. Gunter, E. Mpanga, R. Ayine, J. Okelloonen, J.P.M. Nyeko, J.I. Shililu, J.I. Githure, J.L. Regens, R.J. Novak, and I. Kakoma. 2008. Hydrological modeling of geophysical parameters of arboviral and protozoan disease vectors in Internally Displaced People camps in Gulu, Uganda. International Journal of Health Geographics 7.
Jager, H.I., E.A. Carr, and R.A. Efroymson. 2006. Simulated effects of habitat loss and fragmentation on a solitary mustelid predator. Ecological Modelling 191(3-4):416-430.
Jorquera, O., A. Kiperstok, E.A. Sales, M. Embiruçu, and M.L. Ghirardi. 2010. Comparative energy life-cycle analyses of microalgal biomass production in open ponds and photobioreactors. Bioresource Technology 101(4):1406-1413.
Kamaldeen, S., and D.A. Powell. 2000. Public Perceptions of Biotechnology. Guelph, Ontario: University of Guelph.
Kaskey, J. 2009. Monsanto, Dow Chemical win approval for modified corn (update3). Available online at http://www.bloomberg.com/apps/news?pid=newsarchive&sid=a57J5HHLMOg4. Accessed June 19, 2012.
Kleiner, D. 1974. Quantitative relations for the repression of nitrogenase synthesis in Azotobacter vinelandii by ammonia. Archives of Microbiology 101:153-159.
Koonin, E.V., Y.I. Wolf, and G.P. Karev. 2002. The structure of the protein universe and genome evolution. Nature 420(6912):218-223.
Kristiansen, J. 1996. Dispersal of freshwater algae—A review. Hydrobiologia 336(1-3):151-157.
La Claire, J.W. 2006. Analysis of expressed sequence tags from the harmful alga, Prymnesium parvum (Prymnesiophyceae, Haptophyta). Marine Biotechnology 8(5):534-546.
Lance, E., F. Alonzo, M. Tanguy, C. Gerard, and M. Bormans. 2011. Impact of microcystin-producing cyanobacteria on reproductive success of Lymnaea stagnalis (Gastropoda, Pulmonata) and predicted consequences at the population level. Ecotoxicology 20(4):719-730.
Lardon, L., A. Helias, B. Sialve, J.P. Stayer, and O. Bernard. 2009. Life-cycle assessment of biodiesel production from microalgae. Environmental Science and Technology 43(17):6475-6481.
Lau, P.S., N.F.Y. Tam, and Y.S. Wong. 1995. Effect of algal density on nutrient removal from primary settled waste-water. Environmental Pollution 89:58-66.
Lenzen, M., and S.A. Murray. 2001. A modified ecological footprint method and its application to Australia. Ecological Economics 37(2):229-255.
Lenzen, M., and S.A. Murray. 2003. The Ecological Footprint—Issues and Trends. Sydney: The University of Sydney.
Liss, P.S., A.D. Hatton, G. Malin, P.D. Nightingale, and S.M. Turner. 1997. Marine sulphur emissions. Philosophical Transactions of the Royal Society B-Biological Sciences 352(1350):159-168.
Liu, X., A.F. Clarens, and L.M. Colosi. 2011. Algae biodiesel has potential despite inconclusive results to date. Bioresource Technology 104:83-86.
Lonsdale, W.M. 1999. Global patterns of plant invasions and the concept of invasibility. Ecology 80(5):1522-1536.
Lothrop, B.B., and M.S. Mulla. 1996. Diel patterns of oviposition and influence of agitated water surface in Chironomus anonymus (Diptera: Chironomidae). Journal of the American Mosquito Control Association 12(2 Part 1):215-219.
Lynn, J.C., C.L. Chambers, and S.S. Rosenstock. 2006. Use of wildlife water developments by birds in Southwest Arizona during migration. Wildlife Society Bulletin 34(3):592-601.
Maguire, B. 1963. Passive dispersal of small aquatic organisms and their colonization of isolated bodies of water. Ecological Monographs 33(2):161-185.
Mallick, N. 2002. Biotechnological potential of immobilized algae for wastewater N, P and metal removal: A review. Biometals 15(4):377-390.
Martinez, M.E., S. Sanchez, J.M. Jimenez, F. El Yousfi, and L. Munoz. 2000. Nitrogen and phosphorus removal from urban wastewater by the microalga Scenedesmus obliquus. Bioresource Technology 73(3):263-272.
Mata, T.M., A.A. Martins, and N.S. Caetano. 2010. Microalgae for biodiesel production and other applications: A review. Renewable and Sustainable Energy Reviews 14(1):217-232.
Mathews, T., and N.S. Fisher. 2008. Evaluating the trophic transfer of cadmium, polonium, and methylmercury in an estuarine food chain. Environmental Toxicology and Chemistry 27(5):1093-1101.
McBride, A.C., V.H. Dale, L.M. Baskaran, M.E. Downing, L.M. Eaton, R.A. Efroymson, C.T. Garten Jr, K.L. Kline, H.I. Jager, P.J. Mulholland, E.S. Parish, P.E. Schweizer, and J.M. Storey. 2011. Indicators to support environmental sustainability of bioenergy systems. Ecological Indicators 11(5):1277-1289.
McCabe, T.R. 1994. Assessing values of Arctic wildlife and habitat subject to potential petroleum development. Landscape and Urban Planning 28(1):33-45.
McGarigal, K., S. Cushman, and C. Regan. 2005. Quantifying terrestrial habitat loss and fragmentation: A protocol. Available online at http://www.umass.edu/landeco/teaching/landscape_ecology/labs/fragprotocol.pdf. Accessed July 25, 2012.
Meersman, T. 2010. Minnesota ethanol plants’ price is pollution. Star Tribune, October 11. Available online at http://www.startribune.com/local/104746614.html?elr=KArks:DCiU1PciUiD3aPc:_Yyc:aUoaEYY_1Pc_bDaEP7U. Accessed January 21, 2011.
Mehnert, G., F. Leunert, S. Cirés, K.D. Jöhnk, J. Rücker, B. Nixdorf, and C. Wiedner. 2010. Competitiveness of invasive and native cyanobacteria from temperate freshwaters under various light and temperature conditions. Journal of Plankton Research 32(7):1009-1021.
Meteyer, C.U., R.R. Dubielzig, F.J. Dein, L.A. Baeten, M.K. Moore, J.R. Jehl, and K. Wesenberg. 1997. Sodium toxicity and pathology associated with exposure of waterfowl to hypersaline playa lakes of southeast New Mexico. Journal of Veterinary Diagnostic Investigation 9(3):269-280.
Moeller, P.D.R., K.R. Beauchesne, K.M. Huncik, W.C. Davis, S.J. Christopher, P. Riggs-Gelasco, and A.K. Gelasco. 2007. Metal complexes and free radical toxins produced by Pfiesteria piscicida. Environmental Science and Technology 41(4):1166-1172.
Mogi, M., and M. Motomura. 1996. Possibile Culex pipiens pallens control by improvement of flow rates in water channels of Saga City, southwest Japan. Journal of the American Mosquito Control Association 12(4):647-650.
Moon, H.S., J.M. Abercrombie, A.P. Kausch, and C.N. Stewart. 2010. Sustainable use of biotechnology for bioenergy feedstocks. Environmental Management 46(4):531-538.
Munoz, R., and B. Guieysse. 2006. Algal-bacterial processes for the treatment of hazardous contaminants: A review. Water Research 40(15):2799-2815.
NALMS, (North American Lake Management Society). 2004. Position statement 4. Use of herbicides in lakes. Available online at http://www.nalms.org/media.acux/adbd6227-81e9-411a-be86-cc2691380d0f. Accessed January 31, 2012.
Naugle, D.E., C.L. Aldridge, B.L. Walker, T.E. Cornish, B.J. Moynahan, M.J. Holloran, K. Brown, G.D. Johnson, E.T. Schmidtmann, R.T. Mayer, C.Y. Kato, M.R. Matchett, T.J. Christiansen, W.E. Cook, T. Creekmore, R.D. Falise, E.T. Rinkes, and M.S. Boyce. 2004. West Nile virus: Pending crisis for greater sage-grouse. Ecology Letters 7(8):704-713.
NETL (National Energy Technology Laboratory). 2011. Secure and reliable energy supplies—Coal becomes a “future fuel.” Available online at http://www.netl.doe.gov/KeyIssues/future_fuel.html. Accessed January 12, 2011.
Nixon, S., B. Buckley, S. Granger, and J. Bintz. 2001. Responses of very shallow marine ecosystems to nutrient enrichment. Human and Ecological Risk Assessment 7(5):1457-1481.
Nonneman, D., and P.V. Zimba. 2002. A PCR-based test to assess the potential for microcystin occurrence in channel catfish production ponds. Journal of Phycology 38(1):230-233.
NRC (National Research Council). 2004. Indicators for Waterborne Pathogens. Washington, DC: National Academies Press.
____________. 2011. Renewable Fuel Standard. Potential Economic and Environmental Effects of U.S. Biofuel Policy. Washington, DC: The National Academies Press.
____________. 1987. Field testing genetically modified organisms: framework for decisions. Washington, DC: National Research Council.
NRCS (Natural Resources Conservation Service) and WHC (Wildlife Habitat Council). 2007. Biology - 62 – Fish and Wildlife Habitat Management Leaflet No. 42, Scrub-Shrub Bird. Available online at http://policy.nrcs.usda.gov/viewerFS.aspx?hid=21225. Accessed April 4, 2012.
O’Dowd, W.J., H.W. Pennline, M.C. Freeman, E.J. Granite, R.A. Hargis, C.J. Lacher, and A. Karash. 2006. A technique to control mercury from flue gas: The thief process. Fuel Processing Technology 87(12):1071-1084.
O’Sullivan, J. 2010. SD ethanol plants fined $225k for water violations. Available online at http://thepostsd.com/2010/12/12/sd-ethanol-plants-fined-225k-for-water-violations/. Accessed February 14, 2011.
Ochman, H., J.G. Lawrence, and E.A. Grolsman. 2000. Lateral gene transfer and the nature of bacterial innovation. Nature 405(6784):299-304.
Odlare, M., E. Nehrenheim, V. Ribé, E. Thorin, M. Gavare, and M. Grube. 2011. Cultivation of algae with indigenous species—Potentials for regional biofuel production. Applied Energy 88(10):3280-3285.
Oglesby, R.T. 1977. Relationships of fish yield to lake phytoplankton standing crop, production, and morphoedaphic factors. Journal of the Fisheries Research Board of Canada 34(12):2271-2279.
Osborn, R.G., K.F. Higgins, R.E. Usgaard, C.D. Dieter, and R.D. Neiger. 2000. Bird mortality associated with wind turbines at the Buffalo Ridge wind resource area, Minnesota. American Midland Naturalist 143(1):41-52.
Oswald, W.J., and C.G. Golueke. 1960. Biological transformation of solar energy. Advances in Applied Microbiology 2:223-262.
Oswald, W.J., H.B. Gotaas, C.G. Golueke, and W.R. Kellen. 1957. Algae in waste treatment. Sewage and Industrial Wastes 29:437-455.
Özdemir, E.D., M. Härdtlein, T. Jenssen, D. Zech, and L. Eltrop. 2011. A confusion of tongues or the art of aggregating indicators—Reflections on four projective methodologies on sustainability measurement. Renewable and Sustainable Energy Reviews 15:2385-2396.
Paerl, H.W., and D.F. Millie. 1996. Physiological ecology of toxic aquatic cyanobacteria. Phycologia 35(Supplement):160-167.
Park, J.B.K., R.J. Craggs, and A.N. Shilton. 2011. Wastewater treatment high rate algal ponds for biofuel production. Bioresource Technology 102(1):35-42.
Patrick, M.L., and T.J. Bradley. 2000. The physiology of salinity tolerance in larvae of two species of Culex mosquitoes: The role of compatible solutes. Journal of Experimental Biology 203(4):821-830.
Pepper, I.L., J.P. Brooks, R.G. Sinclair, P.L. Gurian, and C.P. Gerba. 2010. Pathogens and indicators in United States class B biosolids: National and historic distributions. Journal of Environmental Quality 39(6):2185-2190.
Pierce, S.K., S.E. Massey, J.J. Hanten, and N.E. Curtis. 2003. Horizontal transfer of functional nuclear genes between multicellular organisms. Biological Bulletin 204(3):237-240.
Pittman, J.K., A.P. Dean, and O. Osundeko. 2011. The potential of sustainable algal biofuel production using wastewater resources. Bioresource Technology 102(1):17-25.
Plumley, F.G. 1997. Marine algal toxins: Biochemistry, genetics, and molecular biology. Limnology and Oceanography 42(5):1252-1264.
Rabalais, N.N., R.E. Turner, R.J. Díaz, and D. Justi. 2009. Global change and eutrophication of coastal waters. ICES Journal of Marine Science 66(7):1528-1537.
Radakovits, R., R.E. Jinkerson, A. Darzins, and M.C. Posewitz. 2010. Genetic engineering of algae for enhanced biofuel production. Eukaryotic Cell 9(4):486-501.
Raghu, S., R.C. Anderson, C.C. Daehler, A.S. Davis, R.N. Wiedenmann, D. Simberloff, and R.N. Mack. 2006. Adding biofuels to the invasive species fire? Science 313(5794):1742.
Ramirez, P. 2010. Bird mortality in oil field wastewater disposal facilities. Environmental Management 46(5):820-826.
Ramirez, P.J. 2005. Oilfield-produced wastewater discharges into wetlands: Benefits and risks to wildlife. Environmental Geoscience 12:65-72.
Ravindranath, N.H., R. Mauvie, J. Fargione, J.G. Canadell, G. Berndes, J. Woods, H. Watson, and J. Sathaye. 2009. Greenhouse gas implications of land use change and land conversion to biofuel crops. Pp. 111-125 in Biofuels: Environmental Consequences and Interactions with Changing Land Use, R.W. Howarth and S. Bringezu, eds. Ithaca, NY: Cornell University.
Rasmussen, R.A. 1974. Emissions of biogenic hydrogen sulfide. Tellus XXVI (1-2):254-260.
Rawat, I., R. Ranjith Kumar, T. Mutanda, and F. Bux. 2011. Dual role of microalgae: Phycoremediation of domestic wastewater and biomass production for sustainable biofuels production. Applied Energy 88(10):3411-3424.
Reynolds, C.S., V. Huszar, C. Kruk, L. Naselli-Flores, and S. Melo. 2002. Towards a functional classification of the freshwater phytoplankton. Journal of Plankton Research 24(5):417-428.
Rogers, M.B., N.J. Patron, and P.J. Keeling. 2007. Horizontal transfer of a eukaryotic plastid-targeted protein gene to cyanobacteria. BMC Biology 5:26.
Roscher, J.P. 1967. Algal dispersal by muskrat intestinal contents. Transactions of the American Microscopial Society 86:497-498.
Rott, E., and M.C. Hernandez-Marine. 1994. Pulvinularia suecica, a rare stigonematalean cyanophyte. Algol Studies 75:313-322.
Rouhiainen, L., K. Sivonen, W.J. Buikema, and R. Haselkorn. 1995. Characterization of toxin-producing cyanobacteria by using an oligonucleotide probe containing a tandemly repeated heptamer. Journal of Bacteriology 177(20):6021-6026.
Ruiz-Marin, A., L.G. Mendoza-Espinosa, and T. Stephenson. 2010. Growth and nutrient removal in free and immobilized green algae in batch and semi-continuous cultures treating real wastewater. Bioresource Technology 101(1):58-64.
Ryan, C. 2009. Cultivating Clean Energy: The Promise of Algae Biofuels. New York: Natural Resources Defense Council.
Rydzanicz, K., and E. Lone. 2003. Species composition and seasonal dynamics of mosquito larvae in the Wrocław, Poland area. Journal of Vector Ecology 28(2):255-266.
Ryu, J.H., S.D. Gao, and K.K. Tanji. 2010. Speciation and behavior of arsenic in evaporation basins, California, USA. Environmental Earth Sciences 61(8):1599-1612.
Sample, B.E., D.M. Opresko, and G.W. Suter, II. 1996. Toxicological Benchmarks for Wildlife: 1996 Revision. Oak Ridge, TN: Oak Ridge National Laboratory.
Sandefur, H.N., M.D. Matlock, and T.A. Costello. 2011. Seasonal productivity of a periphytic algal community for biofuel feedstock generation and nutrient treatment. Ecological Engineering 37(10):1476-1480.
Sander, K., and G.S. Murthy. 2010. Life cycle analysis of algae biodiesel. International Journal of Life Cycle Assessment 15(7):704-714.
Sarnelle, O. 2010. Effects of cyanobacteria on fitness components of the herbivore Daphnia. Journal of Plankton Research 32:471-477.
Sayre, R. 2011. Sustainable Algal Biofuels. Presentation to the NRC Committee on Sustainable Development of Algal Biofuels on June 13.
Scheffer, M., S. Carpenter, J.A. Foley, C. Folke, and B. Walker. 2001. Catastrophic shifts in ecosystems. Nature 413(6856):591-596.
Scheffer, M., S. Rinaldi, A. Gragnani, L.R. Mur, and E.H. vanNes. 1997. On the dominance of filamentous cyanobacteria in shallow, turbid lakes. Ecology 78(1):272-282.
Schlichting, H.E., Jr. 1960. The role of waterfowl in the dispersal of algae. Transactions of the American Microscopial Society 74:160-166.
Schober, H. 1966. Agitation of water surfaces by sprinkling to prevent mosquito breeding. Mosquito News 26:144-149.
Schomaker, A. 2000. Anaerobic digestion of agro-industrial wastes: Information networks technical summary on gas treatment. Available online at http://agrienvarchive.ca/bioenergy/download/AD_techsum_bio-gas_AD-NETT.pdf. Accessed February 1, 2012.
Schoups, G., J.W. Hopmans, C.A. Young, J.A. Vrugt, W.W. Wallender, K.K. Tanji, and S. Panday. 2005. Sustainability of irrigated agriculture in the San Joaquin Valley, California. Proceedings of the National Academy of Sciences of the United States of America 102(43):15352-15356.
Sharma, N.K., and S. Singh. 2010. Differential aerosolization of algal and cyanobacterial particles in the atmosphere. Indian Journal of Microbiology 50(4):468-473.
Shaw, S.L., B. Gantt, and N. Meskhidze. 2010. Production and emissions of marine isoprene and monoterpenes: A review. Advances in Meteorology 2010(Article ID 408696).
Sheehan, J., T. Dunahay, J. Benemann, and P. Roessler. 1998. A Look Back at the U.S. Department of Energy’s Aquatic Species Program: Biodiesel from Algae. Golden, CO: National Renewable Energy Laboratory.
Sheveleva, E.V., and R.B. Hallick. 2004. Recent horizontal intron transfer to a chloroplast genome. Nucleic Acids Research 32(2):803-810.
Shpiner, R., S. Vathi, and D.C. Stuckey. 2009. Treatment of oil well “produced water” by waste stabilization ponds: Removal of heavy metals. Water Research 43(17):4258-4268.
Shuford, W.D., G.W. Page, and J.E. Kjelmyr. 1998. Patterns and dynamics of shorebird use of California’s Central Valley. Condor 100(2):227-244.
Siegrist, M. 2000. The influence of trust and perceptions of risks and benefits on the acceptance of gene technology. Risk Analysis 20(2):195-203.
Simmons, F.J., and I. Xagoraraki. 2011. Release of infectious human enteric viruses by full-scale wastewater utilities. Water Research 45(12):3590-3598.
Smayda, T.J., and C.S. Reynolds. 2003. Strategies of marine dinoflagellate survival and some rules of assembly. Journal of Sea Research 49(2):95-106.
Smith, J.L., G.L. Boyer, and P.V. Zimba. 2008. A review of cyanobacterial odorous and bioactive metabolites: Impacts and management alternatives in aquaculture. Aquaculture 280(1-4):5-20.
Smith, S. 2008. Pollution violations may test public support for biodiesel. Available online at http://www.biodieselmagazine.com/articles/2383/pollution-violations-may-test-public-support-for-biodiesel. Accessed January 21, 2011.
Smith, V.H. 2003. Eutrophication of freshwater and coastal marine ecosystems—A global problem. Environmental Science and Pollution Research 10(2):126-139.
____________. 2006. Responses of estuarine and coastal marine phytoplankton to nitrogen and phosphorus enrichment. Limnology and Oceanography 51(1):377-384.
Smith, V.H., and D.W. Schindler. 2009. Eutrophication science: Where do we go from here? Trends in Ecology and Evolution 24(4):201-207.
Smith, V.H., B.S.M. Sturm, F.J. deNoyelles, and S.A. Billings. 2010. The ecology of algal biodiesel production. Trends in Ecology and Evolution 25(5):301-309.
Smith, V.H., G.D. Tilman, and J.C. Nekola. 1999. Eutrophication: Impacts of excess nutrient inputs on freshwater, marine, and terrestrial ecosystems. Environmental Pollution 100(1-3):179-196.
Snow, A.A., D.A. Andow, P. Gepts, E.M. Hallerman, A. Power, J.M. Tiedje, and L.L. Wolfenbarger. 2005. Genetically engineered organisms and the environment: Current status and recommendations. Ecological Applications 15(2):377-404.
Snow, A.A., and V. Smith. 2012. Genetically engineered algae for biofuels: A key role for ecologists. Bioscience 62(8):765-768.
Spath, P.L., and M.K. Mann. 2001. Technical Report: Life Cycle Assessment of Hydrogen Production via Natural Gas Steam Reforming. Golden, CO: National Renewable Energy Laboratory.
Stadmark, J., and L. Leonardson. 2005. Emissions of greenhouse gases from ponds constructed for nitrogen removal. Ecological Engineering 25(5):542-551.
Stephenson, A.L., E. Kazamia, J.S. Dennis, C.J. Howe, S.A. Scott, and A.G. Smith. 2010. Life-cycle assessment of potential algal biodiesel production in the United Kingdom: A comparison of raceways and air-lift tubular bioreactors. Energy and Fuels 24:4062-4077.
Stewart, K.W., L.E. Milliger, and B.M. Solon. 1970. Dispersal of algae, protozoans, and fungi by aquatic hemiptera, trichoptera, and other aquatic insects. Annals of the Entomological Society of America 63(1):139-147.
Stoglehner, G. 2003. Ecological footprint—A tool for assessing sustainable energy supplies. Journal of Cleaner Production 11(3):267-277.
Straub, T.M. 1993. Hazards from pathogenic microorganisms in land-disposed sewage sludge. Review of Environmental Contamination and Toxicology 132:55-91.
Strauss, S.H., D.L. Kershen, J.H. Bouton, T.P. Redick, H. Tan, and R.A. Sedjo. 2010. Far-reaching deleterious impacts of regulations on research and environmental studies of recombinant DNA-modified perennial biofuel crops in the United States. Bioscience 60(9):729-741.
Strayer, D.L., V.T. Eviner, J.M. Jeschke, and M.L. Pace. 2006. Understanding the long-term effects of species invasions. Trends in Ecology and Evolution 21:645-651.
Sturm, B.S., E. Peltier, V. Smith, and F. De Noyelles. 2012. Controls of microalgal biomass and lipid production in municipal wastewater-fed bioreactors. Environmental Progress and Sustainable Energy 31:10-16.
Sturm, B.S.M., and S.L. Lamer. 2011. An energy evaluation of coupling nutrient removal from wastewater with algal biomass production. Applied Energy 88(10):3499-3506.
Sukias, J.P.S., and R.J. Craggs. 2011. Digestion of wastewater pond microalgae and potential inhibition by alum and ammoniacal-N. Water Science and Technology 63(5):835-840.
Sukias, J.P.S., and C.C. Tanner. 2005. Ponds for livestock wastes. Pp. 408-432 in Pond Treatment Technology, A. Shilton, ed. London: IWA Publishing.
Suter, G.W., II, and C.L. Tsao. 1996. Toxicological Benchmarks for Screening of Potential Contaminants of Concern for Effects on Aquatic Biota on Oak Ridge Reservation: 1996 Revision. Oak Ridge, TN: Oak Ridge National Laboratory.
Sutton, P.C., S.J. Anderson, C.D. Elvidge, B.T. Tuttle, and T. Ghosh. 2009. Paving the planet: impervious surface as proxy measure of the human ecological footprint. Progress in Physical Geography 33:510-527.
Tenbult, P., N.K. de Vries, E. Dreezens, and C. Martijn. 2005. Perceived naturalness and acceptance of genetically modified food. Appetite 45(1):47-50.
Tiedje, J.M., R.K. Colwell, Y.L. Grossman, R.E. Hodson, R.E. Lenski, R.N. Mack, and P.J. Regal. 1989. The planned introduction of genetically engineered organisms—Ecological considerations and recommendations. Ecology 70(2):298-315.
Turlure, C., J. Choutt, H. Van Dyck, M. Baguette, and N. Schtickzelle. 2010. Functional habitat area as a reliable proxy for population size: Case study using two butterfly species of conservation concern. Journal of Insect Conservation 14(4):379-388.
Uphoff, J.H., M. McGinty, R. Lukacovic, J. Mowrer, and B. Pyle. 2011. Impervious surface, summer dissolved oxygen, and fish distribution in Chesapeake Bay subestuaries: Linking watershed development, habitat conditions, and fisheries management. North American Journal of Fisheries Management 31:554-566.
USDA-ERS (U.S. Department of Agriculture Economic Research Service). 2011. Adoption of Genetically Engineered Crops in the United States. Available online at http://www.ers.usda.gov/data/biotechcrops/. Accessed June 18, 2012.
USDA-RD (U.S. Department of Agriculture Rural Development). 2009. Environmental Assessment for Sapphire Energy Inc.’s Integrated Algal Biorefinery (IBR) Facility in Columbus, New Mexico. Washington, DC: U.S. Department of Agriculture.
USDI (U.S. Department of the Interior). 1998. National Irrigation Water Quality Program Information Report No. 3: Guidelines for Interpretation of the Biological Effects of Selected Constituents in Biota, Water, and Sediment. Denver, CO.
Vaccari, D. 2009. Phosphorus: A looming crisis. Scientific American 300(6):54-49.
Velasquez, G.T. 1940. On the viability of algae obtained from the digestive tract of the Gizzad Shad, Dorosotnu cepediunum. American Midland Naturalist 22:376-412.
Vymazal, J. 1995. Algae and Element Cycling in Wetlands. Boca Raton, FL: Lewis Publishing.
Wang, L., M. Min, Y. Li, P. Chen, Y. Chen, Y. Liu, Y. Wang, and R. Ruan. 2010. Cultivation of green algae Chlorella sp in different wastewaters from municipal wastewater treatment plant. Applied Biochemistry and Biotechnology 162(4):1174-1186.
Watras, C.J., and N.S. Bloom. 1992. Mercury and methylmercury in individual zooplankton—implications for bioaccumulation. Limnology and Oceanography 37(6):1313-1318.
Weathers, P.J. 1984. N2O evolution by green algae. Applied and Environmental Microbiology 48(6):1251-1253.
Weber, C.L., and H.S. Matthews. 2008. Food-miles and the relative climate impacts of food choices in the United States. Environmental Science and Technology 42(10):3508-3513.
Wedding, G.C., and D. Crawford-Brown. 2007. Measuring site-level success in brownfield redevelopments: A focus on sustainability and green building. Journal of Environmental Management 85(2):483-495.
Weiland, U., A. Kindler, E. Banzhaf, A. Ebert, and S. Reyes-Paecke. 2011. Indicators for sustainable land use management in Santiago de Chile. Ecological Indicators 11(5):1074-1083.
West, N.E. 2003. History of rangeland monitoring in the U.S.A. Arid Land Research and Management 17(4):495-545.
White, L.F., and J.B. Shurin. 2011. Density dependent effects of an exotic marine macroalga on native community diversity. Journal of Experimental Marine Biology and Ecology 405(1-2):111-119.
White, R.E., S.R. Wellings, and J.P. Bell. 1983. Seasonal variations in nitrate leaching in structured clay soils under mixed land-use. Agricultural Water Management 7(4):391-410.
Whitford, L.A. 1983. On rare freshwater algae. Transactions of the American Microscopial Society 102(4):401-403.
Wiens, J., J. Fargione, and J. Hill. 2011. Biofuels and biodiversity. Ecological Applications 21(4):1085-1095.
Wigmosta, M.S., A.M. Coleman, R.J. Skaggs, M.H. Huesemann, and L.J. Lane. 2011. National microalgae biofuel production potential and resource demand. Water Resources Research 47: WH00H04.
Wijffels, R.H., and M.J. Barbosa. 2010. An outlook on microalgal biofuels. Science 329(5993):796-799.
Wilkie, A.C., and W.W. Mulbry. 2002. Recovery of dairy manure nutrients by benthic freshwater algae. Bioresource Technology 84(1):81-91.
Wilkinson, D.M., S. Koumoutsaris, E.A.D. Mitchell, and I. Bey. 2011. Modelling the effect of size on the aerial dispersal of microorganisms. Journal of Biogeography 39:89-97.
Wolfe, A.K., and D.J. Bjornstad. 2003. Making decisions about hazardous waste remediation when even considering a remediation technology is controversial. Environmental Science and Technology 37(8):1485-1492.
Wolt, J.D. 2009. Advancing environmental risk assessment for transgenic biofeedstock crops. Biotechnology for Biofuels 2(1):27.
Yanowitz, J., and R.L. McCormick. 2009. Effect of E85 on tailpipe emissions from light-duty vehicles. Journal of the Air and Waste Management Association 59(2):172-182.
Zerzghi, H., C.P. Gerba, J.P. Brooks, I.L. Pepper. 2009. Long-term effects of land application of Class B biosolids on the soil microbial populations, pathogens, and activity. Journal of Residual Science Technology 7:51-61.
Zhang, E.D., B. Wang, Q.H. Wang, S.B. Zhang, and B.D. Zhao. 2008. Ammonia-nitrogen and orthophosphate removal by immobilized Scenedesmus sp isolated from municipal wastewater for potential use in tertiary treatment. Bioresource Technology 99(9):3787-3793.
Zilberman, D., G. Hochman, and D. Rajagopal. 2010. On the inclusion of indirect land use in biofuel regulations. University of Illinois Law Review 2011:413-434.
Zimba, P.V., A. Camus, E.H. Allen, and J.M. Burkholder. 2006. Co-occurrence of white shrimp, Litopenaeus vannamei, mortalities and microcystin toxin in a southeastern USA shrimp facility. Aquaculture 261(3):1048-1055.
Zimba, P.V., C.P. Dionigi, and S.S. Brashear. 2001a. Selective toxicity of exogenous L-lysine to cyanobacteria, relative to a chlorophyte and a diatom. Phycologia 40(5):483-486.
Zimba, P.V., L. Khoo, P.S. Gaunt, S. Brittain, and W.W. Carmichael. 2001b. Confirmation of catfish, Ictalurus punctatus (Rafinesque), mortality from Microcystis toxins. Journal of Fish Diseases 24(1):41-47.
Zimba, P.V., P.D. Moeller, K. Beauchesne, H.E. Lane, and R.E. Triemer. 2010. Identification of euglenophycin—A toxin found in certain euglenoids. Toxicon 55(1):100-104.
Zuo, Z.J., Y.R. Zhu, Y.L. Bai, and Y. Wang. 2012. Acetic acid-induced programmed cell death and release of volatile organic compounds in Chlamydomonas reinhardtii. Plant Physiology and Biochemistry 51:175-184.




















































