Stappenbeck, T.S., Hooper, L.V., and Gordon, J.I. (2002). Developmental regulation of intestinal angiogenesis by indigenous microbes via Paneth cells. Proc. Natl. Acad. Sci. USA 99, 15451–15455.
Vogel, K.J., and Moran, N.A. (2011). Sources of variation in dietary requirements in an obligate nutritional symbiosis. Proc. Biol. Sci. 278, 115–121.
Wang, Z., Klipfell, E., Bennett, B.J., Koeth, R., Levison, B.S., Dugar, B., Feldstein, A.E., Britt, E.B., Fu, X., Chung, Y.M., et al. (2011). Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63.
Warnecke, F., Luginbühl, P., Ivanova, N., Ghassemian, M., Richardson, T.H., Stege, J.T., Cayouette, M., McHardy, A.C., Djordjevic, G., Aboushadi, N., et al. (2007). Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450, 560–565.
Watanabe, H., and Tokuda, G. (2010). Cellulolytic systems in insects. Annu. Rev. Entomol. 55, 609–632.
Werren, J.H., Baldo, L., and Clark, M.E. (2008). Wolbachia: master manipulators of invertebrate biology. Nat. Rev. Microbiol. 6, 741–751.
Wieczorek, H., Beyenbach, K.W., Huss, M., and Vitavska, O. (2009). Vacuolartype proton pumps in insect epithelia. J. Exp. Biol. 212, 1611–1619.
Wilson, A.C., Dunbar, H.E., Davis, G.K., Hunter, W.B., Stern, D.L., and Moran, N.A. (2006). A dual-genome microarray for the pea aphid, Acyrthosiphon pisum, and its obligate bacterial symbiont, Buchnera aphidicola. BMC Genomics 7, 50.
Wong, C.N., Ng, P., and Douglas, A.E. (2011). Low-diversity bacterial community in the gut of the fruitfly Drosophila melanogaster. Environ. Microbiol. 13, 1889–1900.
Xie, J., Vilchez, I., and Mateos, M. (2010). Spiroplasma bacteria enhance survival of Drosophila hydei attacked by the parasitic wasp Leptopilina heterotoma. PLoS ONE 5, e12149.
A8
A NEW VISION OF IMMUNITY: HOMEOSTASIS OF THE SUPERORGANISM30
Abstract
The immune system is commonly perceived as an army of organs, tissues, cells, and molecules that protect from disease by eliminating pathogens. However, as in human society, a clear definition of good and evil might be sometimes difficult to achieve. Not only do we live in contact with a multitude of microbes, but we also live with billions of symbionts that span all
________________
30 Eberl, G. 2010. A new vision of immunity: Homeostasis of the super organism. Mucosal Immunology 3:450–460. Reprinted with kind permission from Nature Publishing Group.
31 Department of Immunology, Institut Pasteur, Lymphoid Tissue Development Unit, Paris, France
32 CNRS URA1961, Paris, France.
Correspondence: G Eberl (gerard.eberl@pasteur.fr).
the shades from mutualists to potential killers. Together, we compose a superorganism that is capable of optimal living. In that context, the immune system is not a killer, but rather a force that shapes homeostasis within the superorganism.
Introduction
The capacity to discriminate between good and evil is crucial for survival. Our senses inform us on the nature of the environment and generate negative feelings of fear and disgust, or positive feelings of comfort and love. Accordingly, decisions are taken to follow a secure path. Societies have developed vast cultures to define good and evil to guarantee the survival of the system, be it a religion, a nation, or a political party. An extreme version of such a culture is Manichaeism, created during the third century AD in Babylon by the prophet Mani, and successfully spread within decades to all corners of the known world. Manichaeism describes the cosmological struggle between the good spiritual world of light and the evil material world of darkness, and prescribes how human society should guide this struggle to its resolution by separating good from evil. As esoteric as Manichaeism theology might sound, its dualistic principle seems nevertheless to pervade our usual perception and description of the world, and thereby to shape microbiology and immunology alike.
At the heart of the dualistic view in microbiology and immunology is the “germ theory of disease” or “pathogenic theory of medicine,” stating that microbes are the cause of a range of diseases. The first mention of such a theory goes back to 36 BC when Marcus Terentius Varro, a Roman scholar, wrote that “certain minute creatures, which cannot be seen by the eyes, which float in the air and enter the body through the mouth and nose, and there cause serious diseases” (Cato and Varro, 1934). Following up in 1020, the Persian polymath Avicenna stated in The Canon of Medicine that bodily secretions are contaminated by “foul foreign earthly bodies” before a person becomes infected (Syed, 2002). The formal demonstration of this theory by Robert Koch and Louis Pasteur in the late 19th century marks the birth of microbiology, hygiene, and vaccine development (Baxter, 2001). However, in the early 20th century, Ilya Metchnikoff proposed that bacteria are also agents of good. He suggested that lactic acid-producing bacteria prolong life by inhibiting the growth of putrefactive (proteolytic) bacteria, such as common gut microbiota members of the Clostridia class, which produce toxic substances and might provoke intestinal “auto-intoxication” (Metchnikoff, 2004). The term “probiotic,” defined as “a live microbial feed supplement, which beneficially affects the host animal by improving its intestinal microbial balance,” was coined in 1953 by Werner Kollath to contrast with antibiotics (Vergin, 1954). The use of probiotics in a number of ailments, including inflammatory bowel disease (IBD) and cancer, has shown some level of protective effects by strains
of the Lactobacillus, Bifidobacteria, Bacteroides, Escherichia, and Faecalibacterium genera (Round and Mazmanian, 2009).
The discovery of innate immune receptors or pattern-recognition receptors (PRRs) in the late 1990s marks a fundamental turn in immunology (Lemaitre et al., 1996; Medzhitov and Janway, 2000). PRRs recognize microbe-associated molecular patterns (MAMPs), thereby allowing the immune system to detect microbes and act accordingly. They are believed to universally elicit rejection and lead to antimicrobial immunity. Reflecting this view, MAMPs are in general termed PAMPs for pathogen-associated molecular patterns (Vance et al., 2009). Thereby, immunology has assigned to the immune system a dualistic perception of the world, aiming it at the detection and destruction of microbes. The inevitable corollary of this view posits that activation of the immune system leads or at least aims at destruction of the activator, and thus defines the activator as a pathogen. However, immune responses come in different blends, best characterized by generation of proinflammatory interferon-γ-producing Th1 cells and interleukin-17 (IL-17)-producing Th17 cells; IgE- and allergy-promoting Th2 cells; and anti-inflammatory IL-10-producing or Foxp3+ regulatory T cells (Weaver et al., 2007). Therefore, the current view holds that pathogens elicit proinflammatory responses, whereas innocuous microbes, such as probiotics, elicit anti-inflammatory responses if detected at all (Sansonetti, 2004). Considerable efforts are made to understand the distinction between pathogens and innocuous microbes, which point to differences in the structure and expression of PAMPs, virulence factors, and invasive properties (Vance et al., 2009).
Here, I challenge this dualistic view of microbes and of the immune system. I propose instead that microbes navigate between shades of good and evil, a position that is determined during interaction with the host, and a position that can change with host, tissue, and time. Facing the microbes, the immune system does not react to combat evil, but merely shapes the microbial environment to allow the organism to live with the microbes. It is not a fight between good and evil, it is rather an equilibrium between microbes and host that generates a superorganism (Gill et al., 2006).
The Superorganism
The Role of the Symbiotic Microbiota
The human intestine hosts an astronomical 1014 bacteria, roughly 100 times the number of cells in our body, and close to 1,000 distinct species (Backhed et al., 2005), not taking in account archea, fungi, and viruses. This microbiota is usually termed commensal, even though there is a considerable degree of mutualism with the host (Box A8-1). The microbes benefit from a selective environment that is regularly flooded with nutrients, and the host benefits from microbial activity that complements its digestive pathways, degrades xenobiotics, regulates epithelial homeostasis (Buchon et al., 2009), and provides a barrier
against potential pathogens (Benson et al., 2009; Brandl et al., 2008; Duerkop et al., 2009; Turnbaugh et al., 2007). Microbial communities reside on all body surfaces, including the entire length of the digestive tract, the vagina, and the skin. Altogether, the partnership of the host with its microbiota can be described as a new functional entity termed a superorganism (Figure A8-1). This superorganism encodes ~2 × 104 host genes and an estimated ~106 microbial genes (Backhed et al., 2005; Gill et al., 2006), and in addition to the mammalian metabolic pathways, operates a plethora of microbial metabolic pathways collectively termed the metabolome (Turnbaugh and Gordon, 2008).
The effect of the microbiota extends beyond complementation of the host. In germfree mice, the intestinal immune system is underdeveloped (Round and Mazmanian, 2009). Lymphoid tissues such as Peyer’s patches (PPs), mesenteric lymph nodes (LNs), and the splenic white pulp remain small, and isolated lymphoid follicles (ILFs), present in hundreds in the lamina propria of normal mice, fail to develop (Bouskra et al., 2008). Furthermore, intestinal lymphocyte populations are markedly reduced, including intra-epithelial T cells, lamina propria T cells, and IgA-producing B cells, and pro-inflammatory Th17 cells (Gaboriau-Routhiau et al., 2009; Ivanov et al., 2009; Niess et al., 2008) and innate IL-22-producing natural killer (NK)-like cells fail to be recruited or generated (Sanos et al., 2009; Satoh-Takayama et al., 2008). Expression of antibacterial peptides by Paneth cells is also reduced (Cash et al., 2006). Thus, the intestinal microbiota, mostly studied at the bacterial level, has an important role in the development and maturation of the immune system. Evidence accumulates that the collection of viruses, or virome, that stably infects the host, also has a major role in shaping the immune system, as discussed recently by Virgin et al. (2009).
Evolution of the Superorganism
Organisms are selected by their fitness within a particular environment. Traditionally, an organism is defined by its genome, and inter-individual competition sorts out the fittest genome. However, selection might also operate at higher levels of organization, such as the group where the social abilities of individuals may provide a collective advantage to the group and hence to each individual of the group (Wynne-Edwards, 1962). At a lower level, the mutualistic partnership between a host and its microbiota provides digestive, protective, and immune advantages to the superorganism. Selection can operate both at the individual host and microbial levels to evolve the best partnership, and the best partnership can be selected to evolve the fittest superorganism. Reflecting this selection process, only few bacterial phyla successfully colonize the human intestine, and the majority of bacterial species are members of either the Firmicutes or the Bacteroidetes phyla (Dethlefsen et al., 2007). On the host’s side, a restricted number (~50 known) of PRRs define the universe of MAMPs that are sensed by the innate immune system (Brown, 2006; Fritz et al., 2006; Kawai and Akira, 2009), a limited reactivity that is nevertheless expected to provide a broad vision of the microbial world to
The recent literature is relatively confusing regarding its usage of the terms “symbiosis” and “commensals.” Symbiosis is generally understood as a relationship between two organisms from which both organisms benefit. In that definition, symbiosis is synonymous with mutualism (Wilkinson, 2001).Furthermore, qualified as commensals are microbes living within a host. However, a number of such commensals are actually mutualists, helping for example in digestion and others are opportunistic pathogens, such as Staphylococcus aureus on the skin. Thus, the usage of the term “commensal” in that context may not be adequate. In this review, I use the original definitions of these terms to categorize the different types of microbes in their relationship with the host.
Symbiosis
Originated from the Greek words syn and biosis, meaning “with” and “living.” The original meaning of symbiosis is the “living together of unlike organisms,” first coined in 1879 by the mycologist Heinrich Anton de Bary (Wilkinson, 2001). The common usage of the term “commensal” should therefore be replaced by “symbiont” to designate the microbiota living within a host. The symbiotic relationships can be formally categorized as mutualistic, commensal, or parasitic, even though parasites are rarely considered symbionts.
Mutualism
Originated from the Latin word mutuus, meaning lent, borrowed, or mutual. A relationship between two organisms where both organisms benefit. For example,
the host. Beyond innate immunity, a virtually unlimited number of receptors are generated by lymphocytes to fill any potential gaps, a diversity suggested to allow vertebrate organisms to host a more complex microbiota (McFall-Ngai, 2007). Furthermore, the immune system must have co-evolved with the microbiota to adjust its reactivity and maintain homeostasis of the superorganism. In conclusion, the dualistic view that separates the host from its microbiota is of course valid in terms of separation of the genomes, but appears to be overruled at the functional level in the superorganism (Box A8-2).
Microbes and Immunity
The Nature of Microbes
Given the proximity with so many microbes, it is expected that we are in constant danger of invasion, and that lowering our guard should be dangerous. As
bacteria that expand in the intestinal niche and provide metabolic pathways complementing the digestive functions of the host.
Commensalism
Originated from the Latin word cum mensa, meaning “sharing a table.” A relationship between two organisms where one organism benefits but the other is unaffected. Microbes that expand in the intestine obviously benefit from the intestinal niche. However, it is more difficult to ascertain that microbes have no effect on the host, so as to be described as commensals.
Parasitism
Originated from the Greek words para and sitos, meaning “beside” and “food,” or one who eats at another’s table. A relationship between two organisms where one organism benefits at the expense of the other. Defines the behavior of a pathogen.
Superorganism
Originated from the Latin word supra, meaning “above,” and the Greek word organon, meaning “organ, instrument, tool.” In biology, an organism is a living system capable of autonomous metabolism and reproduction. A superorganism is a living system of a superior degree of complexity, consisting of many organisms. It may be defined more generally as a “collection of agents that can act in concert to produce phenomena governed by the collective.” Examples of superorganisms include ants and termite societies.
a matter of fact, immunosuppressed individuals are prone to opportunistic infections by ubiquitous and otherwise symbiotic microbes (Slack et al., 2009; Virgin et al., 2009). It seems, therefore, safe to state that the immune system protects us from microbe invasion. However, it is an inaccurate leap forward to state that the immune system protects us from pathogens. How could it define a pathogen? Does a microbe express markers of pathogens before attack? This fundamental question is at the root of the current vision to develop immunotherapies and vaccines (Sansonetti and Medzhitov, 2009; Vance et al., 2009). The discovery of PRRs suggested indeed that expression of PAMPs defines a pathogen, but this view is problematic. Toll-like receptor-4 (TLR-4), which recognizes lipopolysaccharides (LPS), a component of the outer membrane of gram-negative bacteria, best illustrates this problem. Given its wide expression, LPS can hardly be defined as a PAMP. Nevertheless, it has been shown that LPS expressed by prominent symbionts, such as Bacteroides, is modified and poorly recognized by TLR-4 (Sansonetti, 2004; Weintraub et al., 1989). However, potential pathogens such as
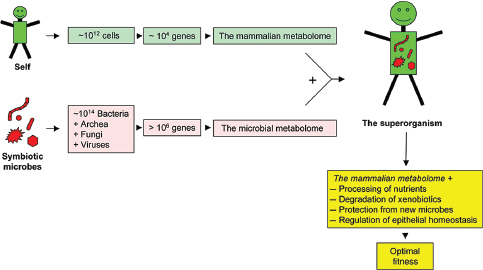
FIGURE A8-1 The superorganism. In the context discussed here, the superorganism is the composite of a host with its symbiotic microbiota. These two worlds have evolved to live together and establish an equilibrium that optimizes the fitness of the superorganism, and thereby the fitness of both the host and the members of the microbiota. The microbial metabolome (Turnbaugh and Gordon, 2008) complements the mammalian metabolome in a number of functions best described in the intestine (Turnbaugh et al., 2007), whereas the mammalian metabolome forms a niche that allows survival of selected microbes.
Helicobacter pylori (Tran et al., 2005) and Porpyromonas gingivalis (Darveau et al., 2004) also express variant forms of LPS that evade efficient recognition by TLR-4. Therefore, the term MAMPs seems to better describe the nature of bacterial moieties recognized by PRRs (Mackey and McFall, 2006; Vance et al., 2009), and cannot be the basis of pathogen recognition.
Recently, it was suggested that pathogenic microbes are recognized by “patterns of pathogenesis” (Vance et al., 2009), or POPs. Rather than by its structure, a pathogen would be defined by its characteristic behavior. A first POP is growth, as pathogens are able to grow in their host upon invasion. A second POP is cytosolic invasion, as many pathogens deliver active proteins, or virulence factors, into the cell host through syringe-like secretion systems or pores, which interfere with activation of the immune system (Sansonetti and Di Santo, 2007). A third proposed POP is disruption of the normal function of the host cell cytoskeleton, as bacterial pathogens and viruses have been shown to exploit the actin network to move within and between cells. During such pathogenic behaviors, MAMPs access forbidden compartments of host tissues and cells. In such a context, MAMPs would be recognized by PRRs as genuine PAMPs and elicit an appropriate inflammatory immune response. In accordance with this view, many PRRs are segregated to react only within a POP context (Barton and Kagan, 2009). In
The value of a concept or a model is measured by its predictive power. Therefore, what is the predictive power of the superorganism concept? What does the continuum model add to the classical dualistic models of immunity? Some leads are proposed here.
The Superorganism
The host and its symbiotic microbiota constitute a superorganism with superior efficiency in digestion, defense, and detoxification, to list a few items, as compared with the bare host (Figure A8-1). Many systems are actually affected in germfree mice (Smith et al., 2007), and therefore, the concept of superorganism is largely validated. Importantly the intestinal microbiota not only affects the intestine but also more distant organs, such as the pancreas (Wen et al., 2008), and the hypothalamic–pituitary–adrenal axis during stress response in mice (Sudo et al., 2004). The mechanism leading to such effects is still a matter of speculation, but a recent paper shows that microbiota-derived peptidoglycans are present in the serum and bone marrow where they enhance neutrophil function (Clarke et al., 2010). Thus, it is predicted, and testable, that homeostasis of the superorganism not only requires an extensive local crosstalk between symbionts and the immune system, but also a more general systemic effect of microbiota based on invaders or circulating MAMPs.
The Continuum Model
The continuum model questions the dualistic view of the immune system based on self/non-self discrimination or physiologic versus pathological inflammation (Figure A8-2). Instead, it proposes that triggers of immunity, microbes for that matter, and the immune responses they elicit are best described by a continuum of properties. The type of microbes and immune responses are defined by their position on this continuum, and the properties of individual microbes can vary between the two extremes depending on their interaction with the host. Thus it is predicted, and already documented (Barton et al., 2007; Brandl et al., 2008; Virgin, 2009), that microbes perceived as pathogens may become mutualists and that microbes perceived as mutualists may become pathogens, depending on the context. Another consequence of the continuum model is that proinflammatory immunity, which is viewed as elicited during a pathological situation, is also required to maintain steady state. Similarly, regulatory immunity is required to maintain steady state but is also involved during microbial infection to establish long-term resistance (Belkaid et al., 2002). It is predicted that blocking proinflammatory immunity during steady state may lead to loss of homeostasis and, conversely, that blocking regulatory immunity during pathology may lead to loss of homeostasis during steady state.
the intestine, TLR-5 is expressed mainly on the basolateral side of the intestinal epithelial cells (Gewirtz et al., 2001) and on dendritic cells in the intestinal lamina propria (Uematsu et al., 2006). Thus TLR-5 is not contacted by its flagellin ligand that is ubiquitous in the intestinal lumen, but by bacteria that cross the epithelial barrier and invade the forbidden compartment of the lamina propria. Some PRRs are only expressed within the cytosol, such as NOD-1 and NOD-2, as well as a number of components of the inflammasome and nucleic acid-recognizing molecules, and can only react to MAMPs that penetrate the cell (Vance et al., 2009). The definition of PAMPs in that context is nevertheless complicated by the existence of membrane transporters encoded by the host, as in the case of NOD-2 (Vavricka et al., 2004), or endosome portals for both NOD-1 and NOD-2 (Lee et al., 2009) that can translocate MAMPs into the cytosol.
The “patterns of pathogenesis” POP hypothesis was proposed with the aim to understand how the immune system recognizes pathogens, and thus was framed within a dualistic view of microbes and immunity (Vance et al., 2009). It can be taken a step further to suggest that a POP does not define a pathogen, but merely a pathogenic behavior that can be adopted by any microbe. The distinction may seem subtle, but allows a particular microbe to be considered a mutualist or a pathogen, depending on time and context. In that view, a pathogenic microbe is not a pathogen per se. Its pathogenicity is determined by its interaction with the host, and can change with host, location in the host, and the immune state of the host. The interaction between a microbe and its host determines the position of the microbe on the scale of mutualists to pathogens, and its position on the scale can change continuously. Many examples exist of bacteria and viruses (Virgin et al., 2009) that can change sides, such as the symbiotic intestinal bacteria that become a source of pathogens during colitis (Sartor, 2008), and pathogenic herpesviruses that cause an acute infection of the host before turning into a helpful ally against bacterial infection during latency (Barton et al., 2007). These and other examples will be discussed in detail below.
The Nature of Immunity
Microbes, be it bacteria, fungi, or viruses, grow and move. However, the niches offered by the host to the microbes in the gastrointestinal tract, the skin, and elsewhere considerably restrict the type of microbes able to colonize (Dethlefsen et al., 2007). For example, most bacteria colonizing the human intestine are members of only two phyla, the Firmicutes and the Bacteroidetes. In addition, containment of the microbiota is enforced by multiple epithelial cell layers, such as in the skin, or thin layers of epithelial cells in the digestive tract and lungs, which produce antibacterial peptides and are protected by mucus produced by Goblet cells. In addition, B cells in mucosal surfaces produce large amounts of IgA that are transported across the epithelial barrier and released into the lumen. There, IgA contribute to the containment of microbes beyond and
within the mucus, and to opsonization of microbes for sampling or destruction by phagocytes (Duerkop et al., 2009).
However, the quality of the containment can change, due for example to environmental variations, exposure to chemicals, and injury. Breaches might be generated and microbes might penetrate the forbidden compartments of host tissues and cells. When this happens, the immune system is triggered through PRRs, such as by TLR-5 on the basolateral side of epithelial cells and on phagocytes in the intestinal lamina propria, which activate cascades of proinflammatory pathways (Gewirtz et al., 2001; Uematsu et al., 2006). The consequence is recruitment and activation of polymorphonuclear cells, macrophages, and lymphocytes; elimination of the intrusive microbes; and eventual wound healing. Ideally, containment and homeostasis is re-established. The rules of engagement of proinflammatory immunity are best described by the “danger model,” which posits that modification of self, through invasive microbes and injury of tissues and cells, is the ultimate trigger (Matzinger, 1994, 2002). Thus, another dualistic view of immunity has emerged: one arm of immunity is involved in every day’s containment of well-behaving symbiotic microbes, whereas another arm of immunity is involved in the elimination of invasive bad-behaving pathogenic microbes that breach containment (Figure A8-2) (Duerkop et al., 2007; Sansonetti, 2004; Sansonetti et al., 2007).
It might be inferred from this view that homeostasis of the superorganism commands the immune system to fight breaches of containment at all cost. However, such a conclusion is a dubious leap forward on the role of proinflammatory immunity and the nature of homeostasis in the superorganism. Even though proinflammatory immunity may follow dualistic rules of engagement to fight the pathogenic behavior of microbes, it cannot avoid breaches of containment. Such breaches seem to occur constantly and are really part of homeostasis. Direct evidence of such breaches is difficult to obtain, but the elevated proinflammatory state of the intestinal immune system is well documented. In specific pathogen-free mice, the proportion of proinflammatory Th17 cells producing IL-17, which induces granulopoiesis and recruitment of neutrophils (Fossiez et al., 1996; Jones and Chan, 2002), is significantly higher in the intestine and skin than in any other tissues (Ivanov et al., 2006; Lochner et al., 2008). This state is induced by the intestinal microbiota (Gaboriau-Rothiau et al., 2009; Ivanov et al., 2009) and is regulated by IL-10 to avoid overdrive of intestinal proinflammatory immunity; IL-10-deficient mice suffer from severe colitis (Kuhn et al., 1993). The notion of “physiologic inflammation” has been suggested a hundred years ago by Metchnikoff in the context of homeostatic removal of dead cells (Metchnikoff, 1989), and was more recently discussed by Sansonetti and Di Santo (Sansonetti and Di Santo, 2007) in reference to immunity induced for containment of symbionts, distinct from proinflammatory immunity to pathogens. However, physiologic inflammation, which might be defined as immunity required for maintaining
homeostasis in the superorganism, appears also to include proinflammatory immunity (Box A8-2).
Homeostasis in the Superorganism
Activation of the Immune System During Homeostasis
Homeostasis in the superorganism is defined by the optimal cohabitation of the host with its microbiota. It is a dynamic equilibrium between host and microbes, where growth and movement of the microbes are constantly kept in check by mechanical, chemical, and immunological containment by the host. The dynamic nature of this equilibrium results from variations in the composition of the microbiota, caused by changes in environmental factors such as chemistry, food, and incoming microbes, and variations in the state of the host through mutations, injury, and other forms of stress. The host must have developed a flexible system that adapts to these variations and maintains homeostasis of the superorganism. The immune system perfectly matches this function: it contributes to containment and equilibrium through for example expression of IL-13 that induces production of mucus by Goblet cells (Wills-Karp et al., 1998), expression of IL-22
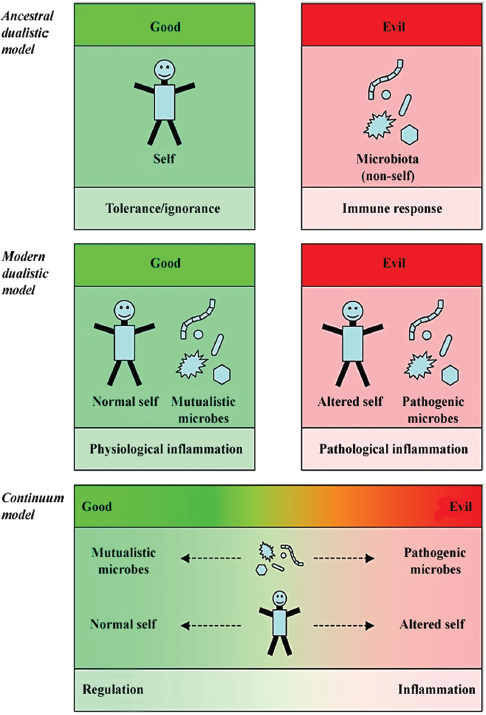
FIGURE A8-2 The dualistic and the continuum models of the microbial and immunological worlds. The ancestral view of immunology states that the immune system is educated not to react to self and to react to non-self. In that view, self is good and induces tolerance during lymphocyte maturation and selection, or anergy of self-reacting mature lymphocytes. Non-self, including microbes, as well as allotypic and xenotypic tissues, are evil and elicit an immune response aimed at destroying non-self. All can be boiled down to self/non-self discrimination (Moller, 1977; Nossal, 1991). The modern view of immunology states that the immune system reacts primarily to danger signals (Matzinger, 1994, 2002). The immune system is still educated to be tolerant to self, but the notion of self is modified. Good includes normal self and mutualistic microbes, whereas evil includes altered self such as dead cells releasing danger signals and pathogenic microbes that alter the antigenic landscape of normal self. In that context, the normal self induces a physiological level of inflammation that contributes to homeostasis through for example containment of the intestinal microbiota, whereas injury and pathogens induce pathological inflammation that leads to “full-blown” inflammation (Sansonetti and Di Santo, 2007). The continuum model states that the perceived duality of mutualistic and pathogenic microbes, normal and altered self, and regulatory or inflammatory immunity, represents extremes of a continuous reality. Microbes can express different levels of mutualistic or pathogenic properties and these levels can vary during interaction with the host. Similarly, the state of self and of immune responses can navigate between well-described extremes, and the most likely states are combination of these extremes.
that induces the production of antibacterial peptides by epithelial cells (Wolk et al., 2004), and generation of IgA-producing B cells (Duerkop et al., 2009). Furthermore, it can be activated to a higher state of inflammation by breaches in containment that provoke deviations from the equilibrium. Thus, the immune system emerges as a crucial force to maintain homeostasis (Figure A8-3).
In the superorganism, the immune system is never at rest. It is like a spring: the more microbes colonize the host niches or behave like pathogens, the stronger they pull the spring of immunity, and the stronger the spring of immunity pushes the microbes back. In germfree animals, the immune spring is close to rest, but in animals grown in a normal microbial world, the immune spring is always under tension, the tension required to maintain homeostasis. This tension also contributes to defining the niche for symbionts, and thus to select incoming microbes and preserve homeostasis. This principle is nicely illustrated by the following experiment: when mice, and humans, were treated with a mix of antibiotics to eradicate most of their intestinal microbiota, they became highly sensitive to infection by an antibiotic-resistant strain of Enterococcus (Brandl et al., 2008). Because of the lack of symbionts, intestinal epithelial cells expressed low levels of antibacterial peptides that normally provide selection against the Enterococcus strain. However, resistance was re-established by concomitant treatment of mice with LPS, which binds TLR-4 and re-induces the expression of antibacterial peptides. Thus, the symbiotic microbiota pulls the string of immunity and induces the production of antibacterial peptides that contribute to defining an intestinal niche permissive for symbionts, but mostly toxic for potential pathogens such as antibiotic-resistant Enterococcus.
Development of the Immune System for Homeostasis
In the absence of microbiota, the immune system does not fully develop. The set of lymphoid tissues is incomplete and immature, and lymphocytes populations in the intestine are reduced in numbers or fail to develop (Vergin, 1954). Particular members of the intestinal microbiota, such as segmented filamentous bacteria, are potent activators of intestinal T-helper cells, including Th17 cells (Gaboriau-Routhiau et al., 2009; Ivanov et al., 2009) and in germfree mice, Th17 cells and NKp46+ cells that produce IL-17 and/or IL-22 are markedly reduced (Gaboriau-Routhiau et al., 2009; Ivanov et al., 2009; Niess et al., 2008; Sanos et al., 2009; Satoh-Takayama et al., 2008). These cytokines have an important role in the containment of microbiota, as IL-17 induces the recruitment of neutrophils (Fossiez et al., 1996) and both cytokines induce the production of antibacterial peptides by epithelial and Paneth cells, such as β-defensins, RegIIIγ, and S100 family members (Ishigame et al., 2009; Liang et al., 2006; Wolk et al., 2004).
The microbiota-induced development of lymphoid tissues is particularly instructive. Secondary lymphoid tissues, such as LNs and PPs, are programmed to develop in the sterile environment of the fetus (Mebius et al., 2003). Lymphoid
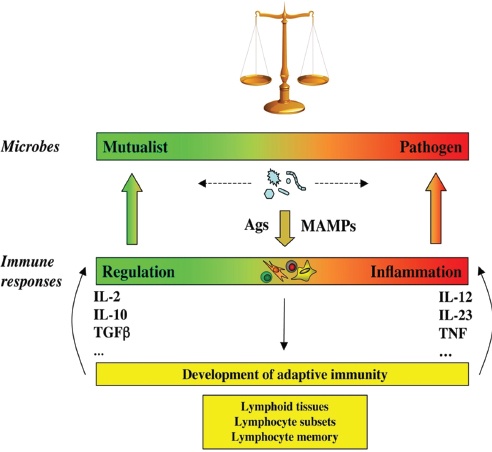
FIGURE A8-3 The continuum of microbial states and immune responses: a dynamic equilibrium. Microbes, including members of the symbiotic microbiota, are not inherently mutualistic or pathogen, but navigate between shades of mutualism and parasitism. Facing the microbes, the immune system is not designed to discriminate between mutualist or pathogens, but merely to react to signals, including MAMPs and antigens. The nature of the immune response is not purely regulatory or inflammatory, but more generally adjusts to the nature of the trigger it faces, like a spring that is pulled by the intensity of the microbial challenge. Furthermore, the immune system has the capacity to evolve when challenged, through generation of different types of lymphocyte subsets such as Th1, Th2, Th17, Treg, Th22 cells (Weaver et al., 2007), and follicular T helper cells (Nurieva et al., 2008), and generation of memory lymphocytes and lymphoid tissues (Round and Mazmanian, 2009). This adds another level of adaptability to the immune system and provides it with the necessary flexibility to maintain homeostasis of the superorganism. MAMP, microbe-associated molecular pattern.
tissue inducer cells are recruited to LN and PP anlagen where they activate specialized stromal cells to produce chemokines and cytokines for subsequent recruitment of B and T cells (Yoshida et al., 1999). After birth, bacterial colonization induces the enlargement of mesenteric LNs and PPs, but does not alter their basic organization, nor are microbes required to maintain this organization. By contrast, ILFs, present in the adult intestinal lamina propria as hundreds of small B-cell follicles, are not programmed during ontogeny, but induced by colonizing microbiota (Hamada et al., 2002). Peptidoglycans shed by dividing bacteria are recognized by NOD-1 in epithelial cells, which then upregulate the expression of β-defensin-3 and of the chemokine CCL20. Both are ligands of CCR6 expressed by lymphoid tissue inducer cells clustered in cryptopatches of the lamina propria. Lymphoid tissue inducer cells in cryptopatches then initiate local lymphoid tissue development and recruitment of CCR6+ B cells (Bouskra et al., 2008). Importantly, ILFs generate IgA-producing B cells that contribute to the containment of the intestinal microbiota, thus establishing a negative feedback loop on their own development (Tsuji et al., 2008). Therefore, ILFs seem to be an optimal system to maintain intestinal homeostasis, as the more bacteria are present in the intestine, the more ILFs are generated that produce IgA, but the more IgA is produced, the less bacteria will be capable of inducing the formation of ILFs. It remains to be established whether a “breach in containment” is required to activate NOD-1 and formation of ILFs, or whether a mechanism of active sampling of luminal content by epithelial cells and transfer to cytosolic NOD-1 is sufficient to activate this pathway (Lee et al., 2009).
Unexpected Mutualists for a Changing Homeostasis
The interaction between a microbe and its host determines the position of the microbe on the scale of mutualists to pathogens, and its position on the scale can change continuously. It may be added that not only the host but also the environment decides the position of the microbe on the scale. Environmental factors may include climate, chemicals, microbes (such as phages), or anything that conditions the fitness of the superorganism. In a changing context, new combinations of microbes and host may yield a competitive advantage to the superorganism, and old foes might become new mutualists. Conversely, old friends may turn bad.
A striking example has been reported in mice infected with a γ-herpesvirus (Barton t al., 2007). Early after infection, γHV68 undergoes lytic replication in a number of cell types, before establishing lifelong latency in memory B cells, macrophages, and dendritic cells. Such a latent infection confers mice resistance to infection with Listeria monocytogenes and Yersinia pestis through a state of increased and persistent immune activation. It was thus proposed that viral latency can define a mutualistic relationship between the host and the virus. In this case, the superorganism includes a former pathogen that eventually has been brought under control: its status within the superorganism changed from an invasive
pathogen to a contained mutualist. Homeostasis is modified as the proinflammatory state of the immune system is increased, providing an advantage to the superorganism in the face of Listeria infection.
Another example is immunological memory. Being exposed constantly to microbes that breach containment, the host develops a collection of memory lymphocytes. The host becomes thereby more reactive to such microbes and alters the niche available to the microbial community. As in the previous example, this evolution of the superorganism is expected to make homeostasis more resistant to pathogenic invasion and thus more robust. Two non-mutually exclusive mechanisms generate and maintain immunological memory: differentiation of long-lived memory lymphocytes during a primary immune response, and latent infection by the microbe or long-lasting antigen depots to uphold specific T- and B-cell activation (Antia et al., 2005; Sprent and Surh, 2002; Welsh et al., 2004; Zinkernagel, 2002). In another case of herpesvirus infection, cytomegalovirus establishes latency after systemic infection in both mouse and man (Snyder et al., 2008). In infected individuals, accumulation of cytomegalovirus-specific CD8+ T cells can reach the exceptional proportion of 10% of total CD8+ T cells, in a phenomenon known as memory inflation. It was shown that this high proportion of cytomegalovirus-specific CD8+ T cells was maintained by long-lived memory cells as well as by continuous generation of short-lived and functional T cells by the latent infection. Such latent infections are thus expected to protect the superorganism from secondary infections, and may operate upon childhood infections and vaccination with other microbes. In this case, homeostasis of the superorganism is modified as immunological memory supported in part by the latent cytomegalovirus infection increases resistance to new infections by itself or related viruses. The old foe remains a foe, but its “domestication” can make it a member of the superorganism to fight its wild relatives.
It has recently been discussed that the host is in a situation of metastable equilibrium with such old foes or pathogens, as a decrease in immunosurveillance, or progressive erosion of this equilibrium, may turn latent microbes into a life-threatening pathogen (Virgin et al., 2009). This view can be applied to the whole symbiotic microbiota, and is the basis of ruptures of homeostasis discussed in the next chapter.
Rupture of Homeostasis
A rupture in homeostasis can be caused by the environment, invading microbes, symbiotic microbes, or the host, and can have severe consequences if the superorganism cannot re-establish equilibrium between the microbiota and the host. Here, we will examine ruptures of homeostasis caused by alterations in the immune system, as well as two examples involving subtle alterations in the host metabolic and tissue regulation. Examples of progressive or abrupt rupture of homeostasis involving the host virome (Virgin et al., 2009), or strong pathogens
such as Shigella or Influenza A virus, are well described (Kobasa et al., 2007; Sansonetti, 2004) and will not be discussed here.
Weak Immunity
Too weak an immune system exposes the superorganism to a conversion of mutualists and commensals to pathogens. The immune forces that normally contain symbionts and establish equilibrium are at risk to be overwhelmed, and microbes that are not normally behaving as pathogens may invade forbidden areas and become pathogens. Immunocompromised people, such as patients suffering from AIDS, treated in intensive care units, receiving chemotherapy, or organ transplants, are exposed to life-threatening opportunistic infections by microbes that are otherwise harmless or ubiquitous symbionts. The latter include the yeast Candida albicans (d’Enfert, 2009), a member of the human microbiota of the skin, intestine, and vagina, and Pseudomonas aeruginosa and Staphylococcus aureus (Nordmann et al., 2007), both present in the skin microbiota (Gao et al., 2007). Recently, it has been reported that lipoteichoic acid derived from Staphylococcus strains inhibits inflammatory cytokine release from keratinocytes through a TLR-2-dependent mechanism (Lai et al., 2009), suggesting a possible mutualistic role of Staphylococci in skin homeostasis. Additional benefits these symbionts may provide to the host include exclusion of related microbes from the niche.
A number of examples have been reported on the loss of containment of the intestinal microbiota in mice deficient for components of the innate or the adaptive immunity. When specific pathogen-free mice are fed orally with the symbiont Enterobacter cloacae, dendritic cells collect bacteria in the PPs sub-epithelial dome before migrating to the draining mesenteric LN where live bacteria can be recovered (Macpherson and Uhr, 2004). In the absence of immunoglobulins in JH-deficient mice, containment of the symbionts is less efficient and significantly more bacteria are recovered from the mesenteric LNs. A similar loss of containment is found in mice deficient for the Toll-like receptor-signaling molecules Myd88 and Trif, or deficient in the phagocyte oxidative burst (Slack et al., 2009). When specific pathogen-free mutant mice are fed with the symbiont Escherichia coli K-12, live bacteria are recovered from the spleen, and serum IgG are produced against members of the symbiotic microbiota, indicating bacterial leakage into the bloodstream. Similar findings were reported earlier in mice deficient for Myd88 or TLR-4 (Fukata et al., 2005). The consequence of decreased containment of the intestinal microbiota is increased morbidity in Myd88- and JH- double-deficient mice (Slack et al., 2009), and increased susceptibility of TLR-4- or Myd88-deficient mice to IBD induced by sodium dextran sulfate (Fukata et al., 2005).
Strong Immunity
Too strong an immune system destabilizes the microbiota and causes dysbiosis, or alteration in the symbiotic microbial community. In the case of Drosophila flies, immune hyperactivity against intestinal symbionts can be fatal (Ryu et al., 2008). Wild-type flies harbor an intestinal community dominated by five bacterial species, including Gluconobacter strain EW707 and Acetobacter strain EW911. Mutant flies that show uncontrolled, bacteria-induced nuclear factor-κB activation, due to knockdown of the regulatory Caudal transcription factor, suffer excessive apoptosis of epithelial cells and progressed to death. It was shown that the mutant flies harbor an altered intestinal bacterial community, where EW911 becomes a minor population and EW707 expands. EW707 alone is pathogenic to germfree flies, but EW911 protect the flies from the damaging effect of EW707. Thus, even though EW707 is a “regular” member of the symbiotic microbiota, its population size has to be kept in check by the presence of other symbionts to maintain homeostasis in wild-type flies. A beneficial role of EW707 for the host during homeostasis remains to be established.
Furthermore, given the nearly unlimited source of antigens generated by the intestinal microbiota, an aggressive immune system can progress into overdrive, generate acute and chronic pathological inflammation, and cause severe damage to the tissue. Such as scenario unfolds during Crohn’s disease and mutations that map to the CARD15 locus significantly increase disease susceptibility in patients (Lesage et al., 2002). CARD15 codes for the innate receptor NOD-2, which recognizes muramyldipeptides shed mostly by Gram-positive bacteria (Giardin et al., 2003). Contradictory data have been reported on the effect of such mutations on the activity of NOD-2. It was found that mutations either increase NOD-2 activity and induce elevated nuclear factor-κB activity and IL-1β secretion (Maeda et al., 2005), or abolish NOD-2 activity and decrease containment through antibacterial peptides (Kobayashi et al., 2005). Decreased or lack of NOD-2 activity, reflecting the mutations found in Crohn’s patients, leads to increased stimulation in response to TLR-2 stimulation and consequent hyperactivity toward the microbiota (Strober et al., 2008).
Dysbiosis
Dysbiosis can be caused by dysregulated immunity and have life-threatening consequences as we just discussed. But dysbiosis can also be induced by subtle shifts in immunity or by non-immunological factors and elicit dramatic consequences.
Propionibacterium acnes is a major constituent of the microbiota on human skin and feeds on fatty acids produced by sebaceous glands (Gao et al., 2007). However, disruption of homeostasis can lead to skin disease caused by P. acnes such as acne. Acne unfolds as hair follicles undergo hyperkeratinization, which blocks sebum egress and causes its accumulation within the follicle
(Noble, 1984). Under such conditions, P. acnes expands and damages the follicle and local containment through increased production of bacterial enzymes. The immune system reacts to occasional breaches in containment and invasion of P. acnes and generates local inflammation and folliculitis. The situation worsens when other skin symbionts, such as Staphyloccocus aureus, penetrate the breaches and cause more severe infections.
Another, much publicized, example of dysbiosis is shown to lead to obesity. The intestinal microbiota of obese ob/ob mice and patients was sequenced and compared with that of lean mice and patients (Ley et al., 2006; Turnbaugh et al., 2006). The microbiome and its inferred metabolome, the collection of metabolic pathways encoded by the microbiota, show that the microbiota of obese individuals is more efficient than that of control individuals in harvesting energy from nutrients, in particular through enhanced catabolism of dietary polysaccharides (Turnbaugh et al., 2006). As a result, the consumable energy remaining in feces is lower in obese individuals. Remarkably, transfer of the cecal microbiota of ob/ob mice to wild-type controls induces a significant increase in body fat as compared with control transfers, showing the dominant effect of the dysbiosis present in ob/ob mice. These data also suggest that obesity can be inherited from the mother not only through genes and behavior, but also through symbiotic bacteria. It was further shown that ob/ob mice and obese patients have an increased proportion of Firmicutes versus Bacteroidetes symbionts, the two dominant phyla in the mouse and human intestine (Ley et al., 2005, 2006). However, the primary cause of dysbiosis in ob/ob mice and obese patients remains a fundamental and fascinating problem (Turnbaugh et al., 2008).
A striking effect of dysbiosis was also reported in the susceptibility of NOD mice to type-1 diabetes (Wen et al., 2008). A series of experiments was designed to assess whether progression to disease was dependent on proinflammatory signals delivered through innate receptors such as TLRs. Apparently confirming this hypothesis, Myd88-deficient NOD mice show marked resistance to development of the disease. T cells reactive to diabetes-associated peptides are found in the spleen and in the mesenteric LN of such mice, yet are markedly reduced in the pancreatic LNs. These data indicated that Myd88-deficient NOD mice do not suffer from systemic immunosuppression but rather from localized immunosuppression in the pancreatic LNs that drain both the pancreas and the intestine. Surprisingly, germfree Myd88-deficient NOD mice develop diabetes to the same extent as germfree or normally colonized Myd88-sufficient NOD mice, indicating a role for the intestinal microbiota in protection from disease progression in the absence of Myd88. A key experiment showed that Myd88-deficient mice develop an altered intestinal microbiota that somehow suppresses the generation of diabetogneic T cells in the pancreatic LN: when NOD newborn mice were raised with Myd88-deficient mothers, disease progression was mitigated. Myd88-deficient mice harbor a decreased ratio of Firmicutes versus Bacteroidetes symbionts as compared with Myd88-sufficient mice, as well as increased
proportions of Lactobacillaceae. A cause-to-consequence relationship has not been established yet in this model, but a probiotic mix containing four species of Lactobacillaceae has been previously shown to protect NOD mice from diabetes and induce a concomitant increase in IL-10 production (Calcinaro et al., 2005).
Restoration of Homeostasis for Prevention of Disease and Therapy
Prevention and therapy of a rupture in homeostasis, leading to disease, might be directed against the primary cause in the host or the microbiota, such as immune dysregulation or potential pathogens. It might also aim at restoring homeostasis through engineering of the symbiotic microbiota.
In cases of immunodeficiencies, the key is to maintain equilibrium between host and microbiota through restoration of normal immunity or a decrease in the microbial load through chemotherapy. In cases of pathological immune hyperactivity, unfolding for example during Crohn’s disease, the most widely used therapy is immunosuppression through administration of anti-tumor necrosis factor-α antibodies. Decreasing the bacterial load through antibiotic therapy would in principle also be a valid approach during acute inflammatory disease, but in the long run exposes the host to invasion by antibiotic-resistant microbes that take advantage of empty niches, and deprives the host of microbial metabolic pathways involved in protection, digestion, and catabolism of xenobiotics.
The study by Pamer and co-workers proposes a new strategy to combine antibiotic treatment with replacement of microbiota (Brandl et al., 2008). In their study, susceptibility of antibiotic-treated mice to an antibiotic-resistant Enterococcus could be mitigated through concomitant administration of LPS. The LPS induced TLR-4-mediated production of antibacterial peptides targeting the pathogenic bacteria, an effect normally induced by the symbiotic microbiota. Thus, in patients treated with antibody cocktails, some degree of homeostasis could be restored by complementing the dwarfed microbiota with selected microbiota-derived compounds. Therefore, deciphering the activity of microbial components on the immune system and beyond the immune system might lead to an extension of such replacement strategies. As microbial metabolic pathways would still be missing, such strategies would have to be complemented with specific diets.
IBD, including Crohn’s disease and ulcerative colitis, is associated with dysbiosis (Packey and Sartor, 2009; Tamboli et al., 2004). It has therefore been proposed that dysbiosis is a potential target for the treatment of IBD. Potential treatments are based on the use of antibiotics targeting specific types of bacteria associated with IBD, or on the use of pre- and probiotics that would favor the restoration of a normal microbiota. Nevertheless, a cause-to-consequence relationship between dysbiotic microbiota and IBD remains to be clearly established. An alternative strategy is based on the use of bacterial strains that exert an immunomodulatory effect on the immune system. The Firmicutes Faecalibacterium prausnitzii is significantly reduced in patients suffering from Crohn’s
disease (Sokol et al., 2008). In vitro, F. prausnitzii induces lower IL-12 and interferon-γ and higher IL-10 production by mononuclear cells, and in vivo, modulated TNBS (2,4,6-trinitro benzene sulfonic acid)-induced colitis. Similar anti-inflammatory effects and protection from IBD have been shown for different species of Lactobacilli, Bifidobacteria, Bacteroides, and E. coli (Round and Mazmanian, 2009). Thus, homeostasis might be restored through modifications of the microbial community through administration of probiotics or prebiotics that favor the emergence of such probiotics. This modified microbiota would exert an anti-inflammatory effect on the hyperactive immune system found in IBD patients and establish a new and potentially viable homeostasis. An important issue however is the stability of such modified homeostasis with time and its robustness in pathological settings.
We have discussed how stable dysbiosis, transmitted by the mother to her offspring or maintained by mutations in the immune system, promotes obesity (Turnbaugh et al., 2006) or prevents the progression of type-1 diabetes (Wen et al., 2008), respectively. These examples show how altered but stable associations between microbiota and host influence the metabolic and inflammatory state of the superorganism. It is therefore expected that a fine understanding of the crosstalk and equilibrium between microbiota and host, and eventually of the rules that govern the superorganism, should lead to preventive and therapeutic strategies in numbers of ailments that are characterized by disrupted homeostasis.
Conclusions
The superorganism is discussed here as the association of the mammalian host with its symbiotic microbiota. This concept offers a novel perspective of self and microbiota, where self is embedded in the microbial world and depends on it for full development and optimal survival. The superorganism requires forces, such as the immune system, to maintain homeostasis. During pathogenesis, the equilibrium between the microbiota and the immune system can become dangerously unstable and glide toward collapse and death of the superorganism. In that context, it appears important to decipher the cellular and molecular cross-talk between the symbiotic microbiota, the host, and its immune system. The molecular messengers of this crosstalk may lead to a new generation of preventive and therapeutic avenues: it will be possible to diagnose and understand ruptures of homeostasis, elaborate strategies to prevent progression to disease, and design therapies for return to homeostasis.
Conflict of Interest
The authors declared no conflict of interest.
Acknowledgments
I thank the following co-workers for discussions, suggestions, and critical reading of the paper: Marc Daëron, Ivo Gomperts-Boneca, Nadine Cerf-Bensussan, Matthew Albert, Pascale Cossart, and Jim Di Santo.
References
Antia, R., Ganusov, V.V. & Ahmed, R. The role of models in understanding CD8+ T-cell memory. Nat. Rev. Immunol. 5, 101–111 (2005).
Bäckhed, F., Ley, R.E., Sonnenburg, J.L., Peterson, D.A. & Gordon, J.I. Host–bacterial mutualism in the human intestine. Science 307, 1915–1920 (2005).
Barton, E.S. et al. Herpesvirus latency confers symbiotic protection from bacterial infection. Nature 447, 326–329 (2007).
Barton, G.M. & Kagan, J.C. A cell biological view of Toll-like receptor function: regulation through compartmentalization. Nat. Rev. Immunol. 9,535–542 (2009).
Baxter, A.G. Louis Pasteur’s beer of revenge. Nat. Rev. Immunol. 1, 229–232 (2001).
Belkaid, Y., Piccirillo, C.A., Mendez, S., Shevach, E.M. & Sacks, D.L. CD4+CD25+ regulatory T cells control Leishmania major persistence and immunity. Nature 420, 502–507 (2002).
Benson, A., Pifer, R., Behrendt, C.L., Hooper, L.V. & Yarovinsky, F. Gut commensal bacteria direct a protective immune response againstToxoplasma gondii. Cell Host Microbe. 6, 187–196 (2009).
Bouskra, D. et al. Lymphoid tissue genesis induced by commensals through NOD1 regulates intestinal homeostasis. Nature 456, 507–510 (2008).
Brandl, K. et al. Vancomycin-resistant enterococci exploit antibiotic-induced innate immune deficits. Nature 455, 804–807 (2008).
Brown, G.D. Dectin-1: a signalling non-TLR pattern-recognition receptor. Nat. Rev. Immunol. 6, 33– 43 (2006).
Buchon, N., Broderick, N.A., Chakrabarti, S. & Lemaitre, B. Invasive and indigenous microbiota impact intestinal stem cell activity through multiple pathways in Drosophila. Genes Dev. 23, 2333–2344 (2009).
Calcinaro, F. et al. Oral probiotic administration induces interleukin-10 production and prevents spontaneous autoimmune diabetes in the non-obese diabetic mouse. Diabetologia 48, 1565–1575 (2005).
Cash, H.L., Whitham, C.V., Behrendt, C.L. & Hooper, L.V. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 313,1126–1130 (2006).
Cato, M.P. &Varro M.T. Cato and Varro: On Agriculture (Loeb Classical Library, 1934).
Clarke, T.B. et al. Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat. Med. 16, 228–231.
Darveau, R.P. et al. Porphyromonas gingivalis lipopolysaccharide contains multiple lipid A species that functionally interact with both toll-like receptors 2 and 4. Infect. Immun. 72, 5041–5051 (2004).
d’Enfert, C. Hidden killers: persistence of opportunistic fungal pathogens in the human host. Curr. Opin. Microbiol. 12, 358–364 (2009).
Dethlefsen, L., McFall-Ngai, M. & Relman, D.A. An ecological and evolutionary perspective on human–microbe mutualism and disease. Nature 449, 811–818 (2007).
Duerkop, B.A., Vaishnava, S. & Hooper, L.V. Immune responses to the microbiota at the intestinal mucosal surface. Immunity 31, 368–376 (2009).
Fossiez, F. et al. T cell interleukin-17 induces stromal cells to produce proinflammatory and hematopoietic cytokines. J. Exp. Med. 183, 2593–2603 (1996).
Fritz, J.H., Ferrero, R.L., Philpott, D.J. & Girardin, S.E. Nod-like proteins in immunity, inflammation and disease. Nat. Immunol. 7, 1250–1257 (2006).
Fukata, M. et al. Toll-like receptor-4 is required for intestinal response to epithelial injury and limiting bacterial translocation in a murine model of acute colitis. Am. J. Physiol. Gastrointest. Liver Physiol. 288, 1055–1065 (2005).
Gaboriau-Routhiau, V. et al. The key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses.Immunity 31, 677–689 (2009).
Gao, Z., Tseng, C.H., Pei, Z. & Blaser, M.J. Molecular analysis of human forearm superficial skin bacterial biota. Proc. Natl. Acad. Sci. USA 104,2927–2932 (2007).
Gewirtz, A.T., Navas, T.A., Lyons, S., Godowski, P.J. & Madara, J.L. Cutting edge: bacterial flagellin activates basolaterally expressed TLR5 to induce epithelial proinflammatory gene expression. J. Immunol. 167,1882–1885 (2001).
Gill, S.R. et al. Metagenomic analysis of the human distal gut microbiome.Science 312, 1355–1359 (2006).
Girardin, S.E. et al. Nod2 is a general sensor of peptidoglycan through muramyl dipeptide (MDP) detection. J. Biol. Chem. 278, 8869–8872 (2003).
Hamada, H. et al. Identification of multiple isolated lymphoid follicles on the antimesenteric wall of the mouse small intestine. J. Immunol. 168,57–64 (2002).
Ishigame, H. et al. Differential roles of interleukin-17A and -17F in host defense against mucoepithelial bacterial infection and allergic responses. Immunity 30, 108–119 (2009).
Ivanov, I.I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell (2009).
Ivanov, I.I. et al. The orphan nuclear receptor RORγt directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell 126,1121–1133 (2006).
Jones, C.E. & Chan, K. Interleukin-17 stimulates the expression of interleukin-8, growth-related on-cogene-alpha, and granulocyte-colony-stimulating factor by human airway epithelial cells. Am. J. Respir. Cell Mol. Biol. 26, 748–753 (2002).
Kawai, T. & Akira, S. The roles of TLRs, RLRs and NLRs in pathogen recognition. Int. Immunol. 21, 317–337 (2009).
Kobasa, D. et al. Aberrant innate immune response in lethal infection of macaques with the 1918 influenza virus. Nature 445, 319–323 (2007).
Kobayashi, K.S. et al. Nod2-dependent regulation of innate and adaptive immunity in the intestinal tract. Science 307, 731–734 (2005).
Kuhn, R., Lohler, J., Rennick, D., Rajewsky, K. & Muller, W. Interleukin-10-deficient mice develop chronic enterocolitis. Cell 75, 263–274 (1993).
Lai, Y. et al. Commensal bacteria regulate Toll-like receptor 3-dependent inflammation after skin injury. Nat. Med. 15, 1377–1382 (2009).
Lee, J. et al. pH-dependent internalization of muramyl peptides from early endosomes enables Nod1 and Nod2 signaling. J. Biol. Chem. 284,23818–23829 (2009).
Lemaitre, B., Nicolas, E., Michaut, L., Reichhart, J.M. & Hoffmann, J.A. The dorsoventral regulatory gene cassette spatzle/Toll/cactus controls the potent antifungal response in Drosophila adults. Cell 86, 973–983 (1996).
Lesage, S. et al. CARD15/NOD2 mutational analysis and genotype-phenotype correlation in 612 patients with inflammatory bowel disease. Am. J. Hum. Genet. 70, 845–857 (2002).
Ley, R.E. et al. Obesity alters gut microbial ecology. Proc. Natl. Acad. Sci. USA 102, 11070–11075 (2005).
Ley, R.E., Turnbaugh, P.J., Klein, S. & Gordon, J.I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Liang, S.C. et al. Interleukin (IL)-22 and IL-17 are coexpressed by Th17 cells and cooperatively enhance expression of antimicrobial peptides. J. Exp. Med. 203, 2271–2279 (2006).
Lochner, M. et al. In vivo equilibrium of proinflammatory IL-17+ and regulatory IL-10+ Foxp3+ RORgamma t+ T cells. J. Exp. Med. 205, 1381–1393 (2008).
Mackey, D. & McFall, A.J. MAMPs and MIMPs: proposed classifications for inducers of innate immunity. Mol. Microbiol. 61, 1365–1371 (2006).
Macpherson, A.J. & Uhr, T. Induction of protective IgA by intestinal dendritic cells carrying commensal bacteria. Science 303, 1662–1665 (2004).
Maeda, S. et al. Nod2 mutation in Crohn’s disease potentiates NF-kappaB activity and IL-1beta processing. Science 307, 734–738 (2005).
Matzinger, P. The danger model: a renewed sense of self. Science 296,301–305 (2002).
Matzinger, P. Tolerance, danger, and the extended family. Annu. Rev. Immunol. 12, 991–1045 (1994).
McFall-Ngai, M. Adaptive immunity: care for the community. Nature 445,153 (2007).
Mebius, R.E. Organogenesis of lymphoid tissues. Nat. Rev. Immunol. 3,292–303 (2003).
Medzhitov, R. & Janeway, C. Jr Innate immune recognition: mechanisms and pathways. Immunol. Rev. 173, 89–97 (2000).
Metchnikoff, I.I. Lectures on the Comparative Pathology of Inflammation Delivered at Pasteur Institute in 1891 (Dover Press, New York, 1989).
Metchnikoff, I.I. The Prolongation of Life: Optimistic Studies (Springer Publishing Company, New York, 2004).
Moller, G. Mechanism of B-cell activation and self-non-self discrimination. Cold Spring Harb. Symp. Quant. Biol. 41 (Part 1), 217–226 (1977).
Niess, J.H., Leithauser, F., Adler, G. & Reimann, J. Commensal gut flora drives the expansion of proinflammatory CD4 T cells in the colonic lamina propria under normal and inflammatory conditions. J. Immunol. 180,559–568 (2008).
Noble, W.C. Skin microbiology: coming of age. J. Med. Microbiol. 17, 1–12 (1984).
Nordmann, P., Naas, T., Fortineau, N. & Poirel, L. Superbugs in the coming new decade; multidrug resistance and prospects for treatment of Staphylococcus aureus, Enterococcus spp. and Pseudomonas aeruginosa in 2010. Curr. Opin. Microbiol. 10, 436–440 (2007).
Nossal, G.J. Molecular and cellular aspects of immunologic tolerance. Eur. J. Biochem. 202, 729–737 (1991).
Nurieva, R.I. et al. Generation of T follicular helper cells is mediated by interleukin-21 but independent of T helper 1, 2, or 17 cell lineages.Immunity 29, 138–149 (2008).
Packey, C.D. & Sartor, R.B. Commensal bacteria, traditional and opportunistic pathogens, dysbiosis and bacterial killing in inflammatory bowel diseases. Curr. Opin. Infect. Dis. 22, 292–301 (2009).
Round, J.L. & Mazmanian, S.K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9,313–323 (2009).
Ryu, J.H. et al. Innate immune homeostasis by the homeobox gene caudal and commensal–gut mutualism in Drosophila. Science 319, 777–782 (2008).
Sanos, S.L. et al. RORgammat and commensal microflora are required for the differentiation of mucosal interleukin 22-producing NKp46+ cells. Nat. Immunol. 10, 83–91 (2009).
Sansonetti, P.J. & Di Santo, J.P. Debugging how bacteria manipulate the immune response. Immunity 26, 149–161 (2007).
Sansonetti, P.J. & Medzhitov, R. Learning tolerance while fighting ignorance. Cell 138, 416–420 (2009).
Sansonetti, P.J. War and peace at mucosal surfaces. Nat. Rev. Immunol.4, 953–964 (2004).
Sartor, R.B. Microbial influences in inflammatory bowel diseases.Gastroenterology 134, 577–594 (2008).
Satoh-Takayama, N. et al. Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense.Immunity 29, 958–970 (2008).
Slack, E. et al. Innate and adaptive immunity cooperate flexibly to maintain host–microbiota mutualism. Science 325, 617–620 (2009).
Smith, K., McCoy, K.D. & Macpherson, A.J. Use of axenic animals in studying the adaptation of mammals to their commensal intestinal microbiota. Semin. Immunol. 19, 59–69 (2007).
Snyder, C.M. et al. Memory inflation during chronic viral infection is maintained by continuous production of short-lived, functional T cells.Immunity 29, 650–659 (2008).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl. Acad. Sci. USA 105, 16731– 16736 (2008).
Sprent, J. & Surh, C.D. T cell memory. Annu. Rev. Immunol. 20, 551–579 (2002).
Strober, W., Kitani, A., Fuss, I., Asano, N. & Watanabe, T. The molecular basis of NOD2 susceptibility mutations in Crohn’s disease. Mucosal. Immunol. 1 (Suppl 1), S5–9 (2008).
Sudo, N. et al. Postnatal microbial colonization programs the hypothalamic–pituitary–adrenal system for stress response in mice. J. Physiol. 558, 263–275 (2004).
Syed, I.B. Islamic medicine: 1000 years ahead of its times. J. Islam. Med. Assoc. 2, 2–9 (2002).
Tamboli, C.P., Neut, C., Desreumaux, P. & Colombel, J.F. Dysbiosis in inflammatory bowel disease. Gut 53, 1–4 (2004).
Tran, A.X., Stead, C.M. & Trent, M.S. Remodeling of Helicobacter pylori lipopolysaccharide. J. Endotoxin. Res. 11, 161–166 (2005).
Tsuji, M. et al. Requirement for lymphoid tissue-inducer cells in isolated follicle formation and T cell-independent immunoglobulin a generation in the gut. Immunity 29, 261–271 (2008).
Turnbaugh, P.J. & Gordon, J.I. An invitation to the marriage of metagenomics and metabolomics. Cell 134, 708–713 (2008).
Turnbaugh, P.J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Turnbaugh, P.J. et al. The human microbiome project. Nature 449, 804–810 (2007).
Turnbaugh, P.J., Backhed, F., Fulton, L. & Gordon, J.I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe. 3, 213–223 (2008).
Uematsu, S. et al. Detection of pathogenic intestinal bacteria by Toll-like receptor 5 on intestinal CD11c+ lamina propria cells. Nat. Immunol. 7,868–874 (2006).
Vance, R.E., Isberg, R.R. & Portnoy, D.A. Patterns of pathogenesis: discrimination of pathogenic and nonpathogenic microbes by the innate immune system. Cell Host Microbe. 6, 10–21 (2009).
Vavricka, S.R. et al. hPepT1 transports muramyl dipeptide, activating NF-kappaB and stimulating IL-8 secretion in human colonic Caco2/bbe cells.Gastroenterology 127, 1401–1409 (2004).
Vergin, F. Antibiotics and probiotics. Hippokrates 25, 116–119 (1954).
Virgin, H.W., Wherry, E.J. & Ahmed, R. Redefining chronic viral infection.Cell 138, 30–50 (2009).
Weaver, C.T., Hatton, R.D., Mangan, P.R. & Harrington, L.E. IL-17 family cytokines and the expanding diversity of effector T cell lineages. Annu. Rev. Immunol. 25, 821–852 (2007).
Weintraub, A., Zahringer, U., Wollenweber, H.W., Seydel, U. & Rietschel, E.T. Structural characterization of the lipid A component of Bacteroides fragilis strain NCTC 9343 lipopolysaccharide. Eur. J. Biochem. 183, 425–431 (1989).
Welsh, R.M., Selin, L.K. & Szomolanyi-Tsuda, E. Immunological memory to viral infections. Annu. Rev. Immunol. 22, 711–743 (2004).
Wen, L. et al. Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 455, 1109–1113 (2008).
Wilkinson, D.M. At cross purposes. Nature 412, 485 (2001).
Wills-Karp, M. et al. Interleukin-13: central mediator of allergic asthma.Science 282, 2258–2261 (1998).
Wolk, K. et al. IL-22 increases the innate immunity of tissues. Immunity21, 241–254 (2004).
Wynne-Edwards, V.C. Animal Dispersion in Relation to Social Behavior ( Oliver & Boyd, London, 1962).
Yoshida, H. et al. IL-7 receptor α+ CD3- cells in the embryonic intestine induces the organizing center of Peyer’s patches. Int. Immunol. 11,643–655 (1999).
Zinkernagel, R.M. On differences between immunity and immunological memory. Curr. Opin. Immunol. 14, 523–536 (2002).