This paper was presented at a colloquium entitled “Genetic Engineering of Viruses and of Virus Vectors,” organized by Bernard Roizman and Peter Palese (Co-chairs), held June 9–11, 1996, at the National Academy of Sciences in Irvine, CA.
Site-specific integration by adeno-associated virus
(parvovirus/gene therapy/targeted integration/DNA replication/recombination)
R. MICHAEL LINDEN*, PETER WARD*, CATHERINE GIRAUD*, ERNEST WINOCOUR†, AND KENNETH I. BERNS*‡
*Department of Microbiology, Hearst Microbiology Research Center, Cornell University Medical College, 1300 York Avenue, New York, NY 10021; and †Department of Molecular Genetics, Weizmann Institute of Science, Rehovot 76100, Israel
ABSTRACT Adeno-associated virus (AAV) has attracted considerable interest as a potential vector for gene delivery. Wild-type virus is notable for the lack of association with any human disease and the ability to stably integrate its genome in a site-specific manner in a locus on human chromosome 19 (AAVS1). Use of a functional model system for AAV DNA integration into AAVS1 has allowed us to conclude that the recombination event is directed by cellular DNA sequences. Recombinant junctions isolated from our integration assay were analyzed and showed characteristics similar to those found in latently infected cell lines. The minimal DNA signals within AAVS1 required for targeted integration were identified and shown to contain functional motifs of the viral origin of replication. A replication mediated model of AAV DNA integration is proposed.
The human parvovirus, adeno-associated virus (AAV), has aroused considerable interest as a potential vector for human gene therapy. Among favorable properties of the virus are its lack of association with any human disease (1), the wide range of cell lines derived from different tissues which can be infected (2), and the ability of the virus to integrate into the genome of the infected cell to establish a latent infection (3). The latter property appears to be unique among mammalian viruses for two reasons. The first is that integration can occur in nondividing cells (4, 37), albeit at a lesser frequency than in dividing cells. Second, AAV integration occurs at a specific site in the human genome, on the q arm of chromosome 19 between q13.3 and qter (5–9). For several years our laboratory has studied the mechanism underlying the site specificity of AAV DNA integration.
To date, a number of experiments have been initiated to address the feasibility of gene transfer with AAV. In preliminary experiments nondividing cells [hematopoetic progenitor cells (10), neurons (11), photoreceptor cells (12), etc.] were shown to be stably transduced by these recombinant AAV vectors. The majority of vectors used have contained the inverted terminal repeats (ITR) as the only genetic information from wild-type AAV. These vectors do not integrate in a site-specific manner. Knowledge of the mechanisms leading to site-specific integration may lead to a superior class of AAV-based vectors.
In cell culture AAV does not undergo productive infection unless there is a coinfection with a helper adeno- (13, 14) or herpesvirus (15, 16). Rather, the virus penetrates to the nucleus where the viral genome is uncoated (K.I.B. and S. Adler, unpublished data). Little viral gene expression occurs, and that which does serves to repress further viral gene expression and to inhibit most viral DNA synthesis. In place of productive viral infection, the viral genome is integrated to establish a latent infection (2, 3, 17). The virus is maintained in the latent state indefinitely, thus perpetuating the viral genetic information. However, superinfection of the latently infected cell with adeno- or herpesvirus activates the viral genome, leading to viral gene expression and to rescue and replication of the viral genome with subsequent production of viral progeny (18). Cells in culture can also be made permissive for AAV-productive infection in the absence of helper adenoor herpesviruses by exposure to genotoxic chemicals or radiation (19–21). Because of its usual dependence on a helper virus for productive infection, AAV was originally considered to be defective. Our current model of the replication of the virus is that AAV has evolved to perpetuate its genetic information by the establishment of latency. When the cell is stressed, the AAV genome is activated to produce new progeny to leave the cell to seek a new host.
Our original studies mapped the specific site of integration (AAVS1) to a position on the q arm of chromsome 19 (8). An 8-kb fragment which contained the integration site was cloned and the 5′ approximately 4 kb were sequenced (7). The sequence contained several interesting features: (i) an open reading frame (ORF) that was expressed in several tissues at low levels detectable using reverse transcription-PCR (no match was found with ORFs representing known proteins); (ii) a higher than expected frequency of direct repeats of dodecanucleotides both upstream and downstream of the ORF; (iii) an overall GC content of 65% which rose to 82% upstream of the ORF in the first 1 kb; and (iv) a 35-mer which was repeated in tandem 10 times. This minisatellite sequence is found at about 60 sites in the human genome, all of which occur on the q arm of chromosome 19 (22). However, none of these features of AAVS1 was sufficiently distinctive to indicate uniqueness of the preintegration site within the the human genome. The possibility existed that the unique aspect of the specific site of integration was not in the sequence but in some higher order structure of the chromatin structure of 19 q.
To resolve the question of whether the specificity lay in the sequence, we have moved AAVS1 to another site in the cell. Our assumption was that if the determining factor was the sequence, the viral genome would integrate regardless of the location of AAVS1. To move the sequence we made use of an Epstein-Barr virus-based shuttle vector (p220.2; ref. 23) which could either be replicated extrachromosomally in a cell cycle-dependent manner in mammalian cells or as a plasmid in Escherichia coli. For persistence in a latent state in mammalian cells, p220.2 is dependent on the presence of the Epstein-Barr origin of DNA replication (oriP) that functions in the presence of EBNA1, the only viral gene product required to initiate replication at oriP. AAVS1 was inserted into the shuttle vector,
The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. §1734 solely to indicate this fact.
Abbreviations: AAV, adeno-associated virus; ITR, inverted terminal repeats; RBS, Rep binding site; TRS, terminal resolution site.
which was then transfected into human C17 cells (293 cells which constitutively express EBNA1). Pools of hygromycin-resistant C17 clones containing the shuttle vector (hygromycin serves as a selective marker for p220.2) were isolated and grown up. On average the clones contained 50–100 copies of the shuttle vector per cell. The cloned cells were infected with AAV and after 48 hr plasmid DNA was isolated and transfected into E. coli. The only colonies to form were those containing the selectable marker of the shuttle vector (ampR which is carried by p220.2). The fraction of such colonies which hybridized to an AAV probe was considered a reflection of the frequency with which AAV had integrated into the shuttle vector. Data are summarized in Fig. 1 (24). AAV did integrate into the shuttle vector which contained the entire 8.2 kb of AAVS1. By sequential deletion analysis it was possible to map the sequences required to direct site-specific integration to the first 510 nt of AAVS1. Thus, it was possible to conclude that AAV site-specific integration was determined by the DNA sequence on 19 q and that the critical sequence was contained within the first 510 bases of the AAVS1 sequence.
To delineate the critical signal sequences within the first 510 bases of AAVS1, a brief review of the molecular genetics and biology of AAV replication is required. Within the 4.7-kb genome there are two ORFs; the one in the right half of the genome encodes the three structural proteins; the ORF in the left half of the genome encodes four regulatory proteins, Reps 78, 68, 52, and 40, with overlapping amino acid sequences. (The Rep designation is used because a frame shift mutation anywhere within the ORF blocks DNA replication) (25). There are two promoters (at map positions 5 and 19) in the left half of the genome and both spliced and unspliced forms of the two transcripts are translated to synthesize the four Rep proteins. Reps 78 and 68 have essentially identical phenotypes and are involved in all phases of the AAV life cycle. These phenotypes depend on the physiological state of the cell. In the absence of helper virus (i.e., the nonpermissive state) Rep 68/78 represses AAV gene expression and inhibits viral DNA synthesis. It is required for site-specific integration and affects expression of a number of cellular genes, most by down regulation (26). In the presence of helper virus (i.e., the permissive state) Rep 68/78 is required for AAV gene expression and transactivates expression of the structural proteins; it is also required for viral DNA replication and rescue of the integrated viral genome. Interestingly, it inhibits expression of the helper adenovirus early genes (M.A.Labow and K.I.B., unpublished data).
The AAV genome contains an ITR of 145 nt (Fig. 2). The first 125 nt constitute an overall palindrome interrupted by two smaller internal palindromes of 21 nt, one immediately on either side of the overall axis of symmetry. When folded on itself to optimize potential base pairing, the palindromic sequence forms a T-shaped structure. The long stem of the T-shaped structure contains an RBS. When Rep binds to the ITR it interacts with at least one of the cross arms of the T (or small internal palindrome) and can make a site-specific nick between nt 124 and 125 (27). After nicking, Rep is covalently bound to the 5′ side of the nick and can function as a helicase (28). The ITR is the cis-active signal in the nonpermissive state for the negative regulation of gene expression and DNA replication. In the permissive state the ITR enhances gene expression, serves as the ori for DNA replication, and is required for rescue of the viral genome from the integrated state.
Thus, it was of interest to note that the 510 nt of AAVS1 sufficient to direct site-specific integration contained both an RBS and TRS in the appropriate orientation with a comparable spacing between them. A third signal of potential interest was a hexanucleotide homologous to an enhancer of meiotic gene conversion (M26) in fission yeast (29). This sequence was also present at approximately the same position (relative to RBS and TRS) at one end of the AAV genome. Biochemical experiments had shown that Rep 68 could bind at the RBS in AAVS1 and that oligomeric complexes of Rep could link the ITR of AAV to the corresponding sequences in AAVS1 (30). It was also shown in vitro that bound Rep could nick the sequence
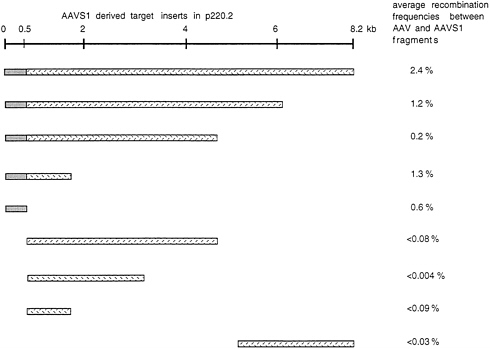
FIG. 1. AAVS1-derived target sequences cloned into p220.2 are graphically displayed. Boxes represent different fragments from AAVS1. Gray boxes highlight the 510-bp fragment sufficient as a target for integration. Average recombination frequencies are indicated for each subfragment. They were calculated as the fraction of E. coli colonies which hybridized to an AAV-specific probe. This fraction was considered to represent the frequency with which AAV had integrated into the shuttle vector. The data were taken from ref. 24.
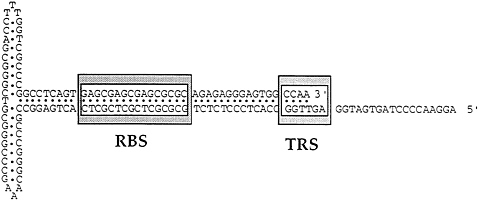
FIG. 2. An AAVITR is shown. The figure represents the T-shaped structure resulting from the palindromic sequence folded on itself to optimize potential base pairing. The stem contains a Rep binding site (RBS) and a terminal resolution site (TRS).
at AAVS1 TRS and initiate unidirectional DNA synthesis (31). To directly demonstrate that the putative signal sequences in AAVS1 were actually required for site-specific integration, a number of genetic analyses were performed (32). These are summarized in Fig. 3. Insertion of DNA fragments representing those portions of the 510 nt containing the TRS and RBS into the shuttle vector assay were required for site-specific integration; M26 was not. In further experiments, creation of mutations involving 2 or 3 nt in RBS or TRS were shown to block site-specific integration. Thus, we have directly demonstrated that defined signal sequences are required to direct site-specific integration. In additional experiments we have found that a 33-nt oligonucleotide containing these two signal sequences is sufficient to direct the integration process. It is of interest to note that these signals in the context of AAV constitute the minimal origin of DNA replication
Although the M26 sequence does not seem to play a direct role in site-specific integration, there are data which suggest that M26 may play a general role in destabilizing AAVS1 and thus may contribute indirectly to the integration reaction (32). Insertion of the 510-nt fragment into the shuttle vector led to rearrangement of the vector sequences in the C17 cells which was independent of the presence of AAV infection. Deletion of the sequences containing M26 has led to stabilization of the shuttle vector in C17 cells. Whether M26 has a comparable effect at the chromosomal level is unknown; there is at least one fragile site on the q-arm of 19 and it has been extremely difficult to map this region, because both cosmids and yeast artificial chromosomes have been very unstable.
Structures of a number of the recombinants produced using the shuttle vector assay have been determined (Table 1) (33). About 20% of the recombinants contained an intact AAV
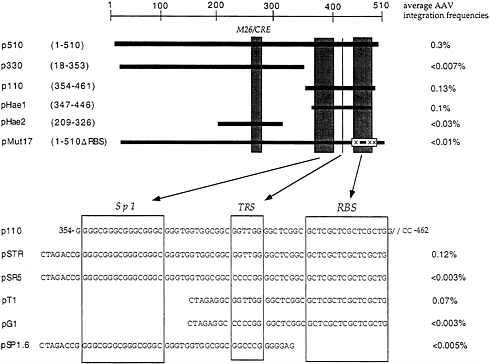
FIG. 3. Construct used for the genetic analysis to determine the recombination signals sufficient and necessary are indicated. The top panel represents subclones generated from the 510-bp fragment sufficient for AAV integration indicated in Fig. 1. Boxes [gray (Upper) and empty (Lower)] indicate known DNA signals identified within the 510-bp sequence. Nucleotide numbers indicated are given with respect to AAVS1. M26, enhancer of meiotic gene conversion in fission yeast; CRE, cyclic AMP response element; Sp1, transcription factor Sp1 consensus sequence; TRS, terminal resolution site; RBS, Rep binding site. (Lower) Synthetic oligonucleotides cloned into the shuttle vector p220.2. Recombination frequencies indicated are calculated as described in the text as well as in Fig. 1. Data were taken from ref. 32.
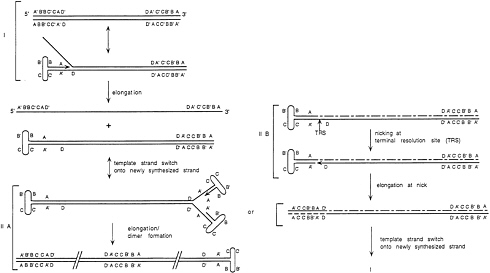
FIG. 4. A model for AAV DNA replication is shown. (I) Replication proceeds by single-strand displacement. (II) After reaching the end of the template strand AAV dimers can be produced (IIA) by folding of the two free ends to form hairpin structures. To be able to prime a second round of replication, the replication apparatus has to switch template strands onto the folded ITR containing the free 3′ OH group. Alternatively, the TRS can be nicked by Rep (IIB), allowing elongation from the nick creating a structure similar to step I.
genome in which the junctions with vector DNA occurred within the ITR. In every case, a portion of the ITR had been deleted. In 2 instances (out of 20) the integrant was more than unit length AAV, stretching from one partial ITR through the ITR at the other end of the genome which had been elongated to form a head-to-tail junction with a portion of a second viral genome that extended to map position 5. These two recombinants were quite interesting, because when the recombinant plasmid was transfected into 293 cells infected with adenovirus, the AAV integrant was rescued and replicated to make progeny virions. Most of the remainder of the recombinants contained less than a full-length viral genome. In most cases the AAV sequences missing were from the part of the AAV genome encoding the REP genes. The integrated DNA sequence extended from a point within the internal AAV sequence through the ITR at the 3′ end of the genome, which again had been elongated to form a head-to-tail junction and then extended in another sequence to map position 5 (as in the two full-length inserts described above). The large number of integrants with head-to-tail junctions suggests a circular intermediate, which could be the result of a limited form of rolling circle replication. Of particular note was that one junction between viral and vector sequences always involved AAVS1 sequences, frequently near RBS; but the other junction between viral and vector DNAs seemed to never occur within AAVS1. Probing the recombinants with oligonucleotides from the upstream AAVS1 sequences demonstrated that in a majority of cases the AAVS1 sequence immediately upstream of the integration event had been rearranged. Rearrangement of flanking cellular sequences in chromosomal integration has been reported, as has been rearrangement of viral sequences.
Table 1. Structural analysis of a subset of recombinants isolated from the shuttle vector assay
|
|
|
AAV junctions with |
AAV sequences involved in junctions |
|
||||
|
|
AAV nucleotide numbers |
AAVS1 |
p220.2 |
Unknown sequences |
ITR |
p5 |
Rep/capsid genes |
Head-to-tail ITR |
|
1 |
2323–4680, 126–268 |
● |
● |
○ |
○ |
● |
● |
● |
|
2 |
3872–4680, 126–251 |
● |
● |
○ |
○ |
● |
● |
● |
|
3 |
1352–4680, 126–287 |
● |
○ |
● |
○ |
● |
● |
● |
|
4 |
Full length, 1–4680 |
○ |
● |
● |
● |
○ |
○ |
○ |
|
5 |
Full length, 1–4680 |
● |
○ |
● |
● |
○ |
○ |
○ |
|
6 |
3106–4680, 126–287 |
● |
● |
○ |
○ |
● |
● |
● |
|
7 |
Greater than full length, 105–4680 and 105–287 |
● |
○ |
● |
● |
● |
○ |
● |
|
8 |
3172–4598 |
● |
● |
○ |
● |
○ |
● |
○ |
|
9 |
4303–4635 |
● |
● |
○ |
● |
○ |
● |
○ |
|
10 |
3819–4547 |
● |
● |
○ |
● |
○ |
● |
○ |
|
11 |
80–719 |
● |
● |
○ |
● |
○ |
● |
○ |
|
Structural characteristics of a number of recombinants produced using the shuttle vector assay are shown. Numbers 1–11 represent independent recombinants analyzed (●, was found; ○, was not present). Data are from ref. 33. |
||||||||
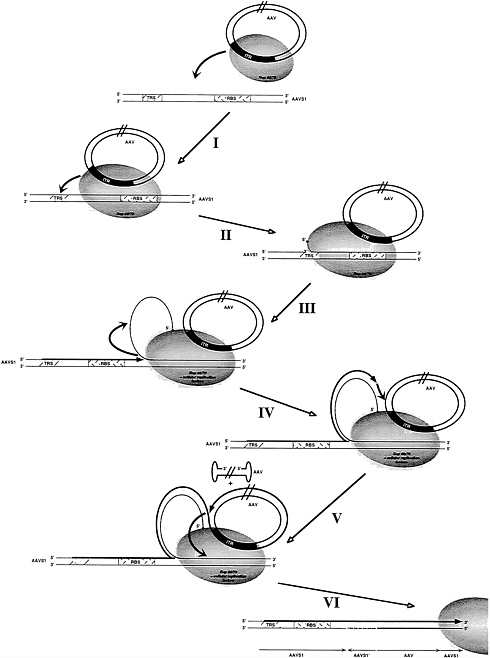
FIG. 5. Model for AAV site-specific DNA integration. Parallel lines in the AAV molecules indicate that the sizes of DNA structures in the figure are not drawn in their actual proportions. A thick grey line indicates the newly synthesized strand; the dashed line indicates the displaced strand of AAVS1. (I) Complex formation between AAV and AAVS1 is mediated by Rep 68/78. (II) Introduction of a strand-specific nick at the TRS in AAVS1 by rep 68/78 and assembly of cellular replication factors. (III) DNA synthesis by single-strand displacement originating at the TRS is followed by template strand switch onto the displaced strand. (IV) A second strand switch occurs onto AAV creating a link between AAVS1 and AAV sequences. (V) After synthesis of AAV DNA sequences a third template strand switch back onto AAVS1 results in a second link between viral and host DNA sequences. (VI) Repair of DNA structures containing noncomplementary strands by cellular enzymes results in integrated copies of AAV DNA within AAVS1. This figure is taken from ref. 32.
A model for the integration process has been developed (32). Because of the involvement of Rep proteins, RBS and TRS, it appears likely that viral DNA replication and localized DNA replication within AAVS1 are involved in integration. AAV DNA replication involves a single strand displacement mechanism (Fig. 4) (for a review, see ref. 34). A major feature of AAV replication is the ability of the elongating strand to switch templates (35). Replication initiates from the ITR in the folded state which serves as both the ori and the primer. When the template strand is fully copied, the 3′ end of the newly synthesized strand can fold on itself and begin synthesis of a second new strand, this time using the first daughter strand as the new template (i.e., by switching templates). It is suggested that the intrinsic tendency of a Rep-mediated replication complex to switch template strands is also the underlying mechanism in the generation of aberrant AAV DNA particles and defective interfering particles.
If the first hairpin structure created by the initial priming event has not been resolved, continuing synthesis will lead to a double stranded, dimeric form of AAV DNA (in Fig. 4IIA). Resolution of the hairpin structures is achieved by Rep cleavage at TRS (Fig. 4IIB). This leads to transfer of the original hairpin sequence from parental to daughter strand and creates a 3′ OH to serve as a primer for repair of the 5′ end of the parental strand.
In vivo, AAV DNA replication requires a co-infection with a helper virus, usually adenovirus. In vitro, it is possible to observe replication of full-length AAV DNA using an extract from uninfected HeLa cells (i.e., from nonpermissive cells) which has been supplemented with purified Rep 68 (36). A significant question was whether the only important adenovirus helper effect on AAV DNA replication was to allow synthesis of sufficient amounts of Rep. The fact that in vitro AAV DNA replication is greatly enhanced by the substitution of extracts from adenovirus-infected cells for those from uninfected cells proves this hypothesis incorrect. The major consequence of using the extract from adenovirus-infected cells is to enhance the ability of the elongating DNA strand to remain on the original template. Use of uninfected cell extract leads to premature strand switching and the consequent interruption of the normal replication process with the synthesis of defective DNA molecules (35). We believe that the enhanced probability of strand switching during DNA synthesis in the absence of a helper virus coinfection plays a major role in the integration process.
A model for the integration process must take into account the following properties. (i) Involvement of RBS and TRS. (ii) Rearrangement of the AAVS1 sequences at one junction. (iii) Presence of head-to-tail AAV junctions, (iv) Despite the requirements for very distinct integration signals (RBS and TRS) a model must account for the observation that integration junctions observed are scattered within ca. 1 kb of AAVS1 downstream of RBS and TRS. A simplified model which can account for these features is shown in Fig. 5. An oligomeric complex of Rep binds to the RBS on AAVS1 and to the RBS in the AAV ITR, thus linking a circularized duplex AAV molecule to AAVS1. This represents a protein (Rep) mediated alignment of the recombination partners AAV and AAVS1, initiating the nonhomologous recombination event observed. Rep then introduces a nick into the AAVS1 TRS. DNA synthesis initiates, displacing a single strand of AAVS1. The extension of replication determines the location of a junction with AAV subsequently formed (see requirement iv mentioned above). It should be noted that the displaced single strand is circular because Rep is covalently bound to the 5′ end and presumed to be still bound to the RBS. After limited extension the elongating strand switches to the displaced single strand as the template; note that copying of the displaced circular AAVS1 sequence leads to inversion of the sequence. When it reaches the end of the displaced strand (close to the RBS as observed in the shuttle vector model system), the elongating strand again switches templates, now onto the circular AAV DNA. After synthesis proceeds on the AAV template, the elongating strand reaches RBS where Rep is bound and the strand again switches to a new template on AAVS1 (alternatively onto p220.2 in the shuttle vector system). Eventually the single-strand gap involving the inserted AAV sequence and the inverted AAVS1 sequence is repaired. Undoubtedly, this model is simplified, but we believe that it is consistent with many of the features observed in AAV integration.
The proper conjunction of RBS and TRS required for integration is present only once in the data concerning the human genome in GenBank, likely explaining the apparent presence of only a single site for specific AAV integration. However, RBS has been noted at multiple sites in the human genome; in 14/15 cases analyzed, it appears in the 5′-untranslated regions of characterized genes; therefore, indicating a cellular counterpart of the Rep protein with possible regulatory functions, which also recognizes RBS. In addition it seems likely that AAV has evolved to take advantage of one copy of this recognition signal sequence.
Our current knowledge of the requirements for site-specific AAV DNA integration, together with our proposed model for an integration mechanism, may help in the design of improved AAV-based vectors for gene therapy. At this point it can be concluded that the Rep protein is an absolute requirement for the site specificity of AAV DNA integration. Finally, we propose that the full AAV ITR may not be necessary for targeted integration. Rather, it is possible that integrity of the ITR is only required for efficient rescue of integrated proviruses, a function not necessary, or even desirable, for stable, long term gene delivery.
This work was supported by a grant from the National Institute of Allergy and Infectious Diseases (AI22251) and by a grant from the National Institute of General Medical Sciences (GM50032). R.M.L. was supported in part by a fellowship from the Norman and Rosita Winston Foundation.
1. Blacklow, N.R., Hoggan, M.D., Kapikian, A.Z., Austin, J.B. & Rowe, W.P. (1968) Am. J. Epidemiol. 88, 368–378.
2. Berns, K.I., Pinkerton, T.C., Thomas, G.F. & Hoggan, M.D. (1975) Virology 68, 556–560.
3. Cheung, A.K., Hoggan, M.D., Hauswirth, W.W. & Berns, K.I. (1980) J. Virol. 33, 739–748.
4. Podsakoff, G., Wong, K.K., Jr., & Chatterjee, S. (1994) J. Virol. 68, 5656–5666.
5. Kotin, R.M. & Berns, K.I. (1989) Virology 170, 460–467.
6. Kotin, R.M., Siniscalco, M., Samulski, R.J., Zhu, X.D., Hunter, L., Laughlin, C.A., McLaughlin, S., Muzyczka, N., Rocchi, M. & Berns, K.I. (1990) Proc. Natl. Acad. Sci. USA 87, 2211–2215.
7. Kotin, R.M., Linden, R.M. & Berns, K.I. (1992) EMBO J. 11, 5071–5078.
8. Kotin, R.M., Menninger, J.C., Ward, D.C. & Berns, K.I. (1991) Genomics 10, 831–834.
9. Samulski, R.J., Zhu, X., Xiao, X., Brook, J.D., Housman, D.E., Epstein, N. & Hunter, L.A. (1991) EMBO J. 10, 3941–3950.
10. Zhou, S.Z., Cooper, S., Kang, L.Y., Ruggieri, L., Heimfeld, S., Srivastava, A. & Broxmeyer, H.E. (1994) J. Exp. Med. 179, 1867–1875.
11. Kaplitt, M.G., Leone, P., Samulski, R.J., Xiao, X., Pfaff, D.W., O’Malley, K.L. & During, M.J. (1994) Nat. Genet. 8, 148–154.
12. Ali, R.R., Reichel, M.B., Thrasher, A.J., Levinski, R.J., Kinnon, C., Kanuga, N., Hunt, D.M. & Bhattacharya, S.S. (1996) Hum. Mol. Genet. 5, 591–594.
13. Atchison, R.W., Casto, B.C. & Hammon, W. (1965) Science 149, 754–756.
14. Hoggan, M.D., Blacklow, N.R. & Rowe, W.P. (1966) Proc. Natl. Acad. Sci. USA 55, 1467–1471.
15. Buller, R.M., Janik, J.E., Sebring, E.D. & Rose, J.A. (1981) J. Virol. 40, 241–247.
16. Weindler, F.W. & Heilbronn, R. (1991) J. Virol. 65, 2476–2483.
17. Handa, H., Shiroki, K. & Shimojo, H. (1975) J. Gen. Virol. 29, 239–242.
18. Hoggan, M.D., Thomas, G.F. & Johnson, F.B. (1972) Continuous “Carriage” of Adenovirus Associated Virus Genomes in Cell Cultures in the Absence of Helper Adenovirus (North-Holland, Amsterdam).
19. Yakobson, B., Koch, T. & Winocour, E. (1987) J. Virol. 61, 972–981.
20. Yakinoglu, A.O., Heilbronn, R., Burkle, A., Schlehofer, J.R. & zur Hausen, H. (1988) Cancer Res. 48, 3123–3129.
21. Yakobson, B., Hrynko, T.A., Peak, M.J. & Winocour, E. (1989) J. Virol. 63, 1023–1030.
22. Das, H.K., Jackson, C.L., Miller, D.A., Leff, T. & Breslow, J.L. (1987) J. Biol. Chem. 262, 4787–4793.
23. Yates, J.L., Warren, N. & Sugden, W. (1985) Nature (London) 313, 812–815.
24. Giraud, C., Winocour, E. & Berns, K.I. (1994) Proc. Natl. Acad. Sci. USA 91, 10039–10043.
25. Hernonat, P.L., Labow, M.A., Wright, R., Berns, K.I. & Muzyczka, N. (1984) J. Virol. 51, 329–339.
26. Labow, M.A., Graf, L.H., Jr., & Berns, K.I. (1987) Mol. Cell. Biol. 7, 1320–1325.
27. Im, D.S. & Muzyczka, N. (1990) Cell 61, 447–457.
28. Im, D.S. & Muzyczka, N. (1992) J. Virol. 66, 1119–1128.
29. Schuchert, P., Langsford, M., Kaslin, E. & Kohli, J. (1991) EMBO J. 10, 2157–2163.
30. Weitzman, M.D., Kyostio, S.R., Kotin, R.M. & Owens, R.A. (1994) Proc. Natl. Acad. Sci. USA 91, 5808–5812.
31. Urcelay, E., Ward, P., Wiener, S.M., Safer, B. & Kotin, R.M. (1995) J. Virol. 69, 2038–2046.
32. Linden, R.M., Winocour, E. & Berns, K.I. (1996) Proc. Natl. Acad. Sci. USA 93, 7966–7972.
33. Giraud, C., Winocour, E. & Berns, K.I. (1995) J. Virol. 69, 6917–6924.
34. Berns, K.I. (1996) in Parvoviridae: The Viruses and Their Replication, eds. Fields, B.N., Knipe, D.M. & Howley, P.M. (Lippincott-Raven, Philadelphia), Vol. 2, pp. 2173–1220.
35. Ward, P.J. & Berns, K.I. (1996) J. Virol. 70, 4495–4501.
36. Ward, P., Urcelay, E., Kotin, R., Safer, B. & Berns, K.I. (1994) J .Virol. 68, 6029–6037.
37. Russell, D.W., Miller A.D. & Alexander, I.E. (1994) Proc. Natl. Acad. Sci. USA 91, 8915–8919.







