Out Far and in Deep: Shifting Perspectives in Ocean Ecology
Peter A. Jumars
Darling Marine Center, University of Maine
ABSTRACT
The pace of scientific advance in ocean ecology since the Ocean Ecology: Understanding and Vision for Research (OEUVRE) workshop is impressive. New food web models reveal the stabilizing influence of weakly interacting species. Hence a reason for population instability becomes the absence or disappearance of these stabilizers. Time-series analysis of the planktonic community and nutrients in the Central North Pacific gyre similarly have led to a much clearer focus on controls of patterns and rates of change in ecosystem structure. This focus on change lends itself also to the extraction of anthropogenic effects through potentially powerful statistical methods such as intervention analysis—if timing of onset of an anthropogenic perturbation is known. Realizing this power for past events, however, requires extending time series backward through paleontological evidence. Advances in understanding of form and function in the organisms that produce microfossils would greatly accelerate such progress and can be expected to follow from impressive recent gains in understanding of how body sizes and shapes interact with bacterial chemotactic capabilities and how copepods distinguish prey from predators hydromechanically. Ocean ecology approaches are accelerating toward Gordon Riley's goal for biological oceanography of having parsimonious equations effectively describing in predictive fashion the interactions of populations of organisms with their abiotic environments as well as with each other.
The people along the sand
All turn and look one way . . .
They cannot look out far.
They cannot look in deep . . .
—Robert Frost
"Neither out Far nor in Deep"
INTRODUCTION
Oceanographers pride themselves in taking a perspective different from the one embodied in Robert Frost's classic poem about human proclivity to stare out to sea but not to see. Scientists of all types pride themselves in being able to adopt multiple, tentative perspectives simultaneously in order to design measurements and experiments that can help to determine which perspectives reflect more accurately the rules that nature follows. It is difficult therefore to summarize perspectives, even within a subdiscipline of oceanography. Approximately 40 scientists met in February 1998, however, to make just such a summary in the form of past successes and future directions of biological oceanography, as supported by the National Science Foundation. The group chose the acronym "OEUVRE" for "Ocean Ecology: Understanding and Vision for Research" to describe its effort. Its report is available at http://www.joss.ucar.edu/joss-psg/project/oce-workshop/, and its separate publication is planned, so I do not repeat it here.
Several months later, I continue to be surprised by the outcome of the meeting. The first surprise was the pleasant one of seeing how much progress has been made in 30 years. Barber and Hilting in a companion paper in this volume have covered some of the major "Achievements in Biological Oceanography," so I will deal primarily with "the vision
thing." I cannot claim to have originated the ideas because I was present both for discussions of future research by all 40 of the participants and for the task of writing them up. The vision necessarily is clouded, however, by the second and far less pleasant surprise of the meeting, that anthropogenic effects on marine ecosystems are ubiquitous and probably have been since the advent of commercial whaling. The third surprise on which I focus here is how quickly perspectives in all areas of ocean ecology have been changing since the meeting—and in the direction of making long-difficult problems suddenly more tractable.
For OEUVRE we chose to summarize research opportunities in subject matter categories that are cross-cutting and untraditional. I use the same headings here to facilitate the reader' s testing of my assertion of rapid progress against the OEUVRE report. Immediately after the headings, I reproduce the questions that OEUVRE identified under these headings as being both pressing and well poised for progress in the next two decades, and I spend the first few paragraphs in each section explaining the topic heading. I devote most space, however, to developing one or two examples of striking progress since the February 1998 meeting. I make no pretense of balanced coverage of the topic area or questions; scientific progress rarely is even across all fronts. Further, each participant or other ocean ecologist would be likely to choose somewhat different examples. My examples take highly variable space and referencing to develop, depending on the background provided in the OEUVRE report.
The exercise reveals several symptoms of substantial shift in perspectives. Unsuccessful search for a superstable marine ecosystem has ended. Along with this failure comes a new focus on how and why marine ecosystems change over time—and on which changes may contain an anthropogenic component. Coherent, succinct models are emerging of sensory systems and behaviors at spatial scales and Reynolds numbers for which humans have no native intuition at the same time that high-technology sensor systems are being deployed that allow unprecedented spatial and temporal resolution in human exploration of the sea. For many reasons, some of the most revealing exploration now is in time rather than space. Indeed this essay focuses more on how perspectives are changing than on summarizing old or new perspectives and is clearly derivative in that sense as well.
FUNCTIONAL SIGNIFICANCE OF BIOLOGICAL DIVERSITY IN BIOGEOCHEMICAL DYNAMICS
-
How do environmental and biotic factors determine the distributions and activities of key species or functional groups important to biogeochemical cycles in space and time?
-
What are the important interactions among marine biota, global climate, and biogeochemistry?
Species diversity has received broad attention for many good reasons. The intent of the topic heading was to focus on function and the mapping of biological diversity onto functional diversity in biogeochemical transformations. Analytic and predictive models of ecosystem function for the foreseeable future require some aggregation of organisms into functional groups (e.g., bacterivores or sulfate reducers). The level of aggregation that is useful depends on the question and discriminatory ability at hand, but it is clear that much effort remains to be spent on assigning organisms to biogeochemically functional groups, with due attention to taxonomy and physiology. Excitement is palpable about the maturation of DNA methodologies for both species identification and identification of potential to catalyze specific reactions (e.g., presence of genes that code for nitrogenase) and the maturation of RNA technologies that can assess whether that potential is being realized.
Anticlimax
The central gyres of the ocean present many interesting questions. Collectively they constitute the largest habitat type on Earth and one of the oldest. Among them, the Central North Pacific Gyre (CNP) individually is the largest ecosystem on Earth, and my focus is primarily on this example. A small mistake in understanding of geochemical processes in such habitats can integrate into a large problem with global budgets. One geochemical problem of long standing is an inability to balance the fixed nitrogen budget for the global ocean. In most summaries, the loss terms exceed production substantially. Central gyres present a problem both individually and collectively, in that they seem to use more nitrogen in new production than can be accounted for (e.g., McGillicuddy and Robinson, 1997).
A parallel, long-running argument between geochemists and biologists has been about whether the oceans ultimately are limited in primary production by nitrogen or by phosphorus. The argument is partly a semantic one about what is meant by "ultimately" (i.e., time scale) and by "limited" (i.e., abundance or production of any or most phytoplankton), but it revolves around the issue that phosphorus has no substantial atmospheric source, whereas nitrogen is the largest component of the atmosphere. At its most simplistic, the assertion sometimes is made that because nitrogen-fixing organisms exist, the oceans cannot ultimately be nitrogen limited. Nitrogen fixation is notoriously expensive in energy and phosphorus, however (partly because it must be done anaerobically), and more sophisticated arguments revolve around whether rates of nitrogen fixation are slow enough to make phosphorus the effectively limiting nutrient because its availability restricts nitrogen fixation. There has long been evidence of phosphate as well as nitrate limitation in gyres, even of less demanding taxa than nitrogen fixers (e.g., Perry, 1972, 1976). Moreover, nitrogen fixers like Trichodesmium may well be limited or co-limited by iron
(Falkowski, 1997) and their vertical migration may be critical in obtaining nutrients.
The classic view of the gyres is as superstable, very species-rich but biomass-poor ecosystems, both in the pelagic realm and on the seabed (Hessler and Jumars, 1974). Early in the exploration of the CNP, climax communities were a popular ecological concept, and the gyres looked like end-member examples. For this reason, early exploratory cruises and station locations were named ''Climax." Species diversity is high, and zooplankton samples taken years apart were as similar in species composition as samples from the same cruise (McGowan and Walker, 1985).
Flaws in the idea of oligotrophic gyres as nutrient impoverished appeared in the form of evidence that some phytoplankton were growing at high rates (e.g., Laws et al., 1984). Flaws appeared in the idea of constancy or static stability after the number of visits grew (Venrick et al., 1987), but disintegration of the idea came from time series funded under JGOFS (Joint Global Ocean Flux Study). The CNP showed two seasons of enhanced new production, one in winter based on enhanced physical mixing and one in summer based on enhanced nitrogen fixation, and these pulses showed substantial interannual variability (Karl et al., 1996). Further analysis of the time series and integration with all prior data suggest a doubling of primary production and a shift from dominance by eukaryotes to dominance by prokaryotes in the mid-1970s (Karl, in review). Some evidence links decadal-scale change in the Central North Pacific to the same large-scale ocean-atmosphere interactions that drive El Niño-Southern Oscillation (Karl et al., 1995). Not even this diverse community can resist basin-scale changes imposed by physics and chemistry.
Wind and eddy activity that can be important in bringing nitrate closer to the surface intuitively is unsteady and difficult to integrate over scales in time and space appropriate to the balance of nutrient budgets, and it is not hard to imagine that this component has interannual variability. The perspective shift underway, however, is that unusual lack of physical mixing also leads to enhanced new production, which is based instead on nitrogen fixation (Karl, in review) and on nitrate transport through vertical migration by mats of the diatom Rhizosolenia (Villareal et al., 1999). That is, the CNP's new production is minimized at some intermediate and probably "typical" input of physical energy, and lack of energy input leads to important biological "events." The theme of physical control of functional groups that effect drawdown of nutrients (including CO2) extends to the Southern Ocean (Arrigo et al., 1999). Margalef (1978) must be pleased.
Time series clearly have power in exploring patterns of temporal variation and cross-correlation and have been key in shifting perspective away from stable, steady climax. They have made central gyres obvious places to improve the global ocean nitrogen budget, making the notion of oligotrophic seas as deserts even less tenable. To what extent nitrogen fixation is limited by phosphorus and trace metals (Falkowski, 1997), and to what extent it occurs in heterotrophic bacteria as well as cyanobacteria, is unclear (Karl, in press). Grazer influences on rates of nitrogen fixation and on the food web fates of microbially fixed nitrogen beg for exploration. The extent to which and reasons why higher trophic levels are more (or less) stable in composition than primary producers are unquantified. The success of Ironex II and newspaper reports of parallel successes in the Southern Ocean make large-scale manipulation of phosphorus and trace-metal concentrations a tantalizing prospect for oligotrophic gyres, and the existing time series can suggest the season and duration that would be effective. Making explicit, mechanistic, a priori predictions of consequences and their time scales is certain to enhance greatly the knowledge gained from any discrepancies observed.
FUNCTIONAL ECOLOGY (OF INDIVIDUALS, WITH RAMIFICATIONS FOR POPULATIONS, COMMUNITIES, AND ECOSYSTEMS)
-
How are mass and momentum transfer and other environmental forces integrated with information to influence behavior?
-
How does performance change with size and form?
Functional ecology is not a well-established term in the popular ecological lexicon, despite the fact that a journal of the British Ecological Society bears this name. Loosely, by contrast to numerical ecology, it refers to the performance of individuals in the context of environmental features, including other organisms. Perhaps the best-known functional responses of individuals are "filtering" and ingestion rates as functions of food concentration: rectilinear, hyperbolic, and sigmoidal responses have been described. As a consequence of rapid advances in understanding of mechanics of food encounter and handling, it can now be argued that it is better to use arrival rate of food items in the sensory field of the forager as the independent variable (instead of food concentration) in predicting ingestion rates. Ambient fluid motion, for example, can alter rate of encounter without any change in food concentration.
Tactile Senses of Copepods: Discriminating Food from Foe
Among the most difficult phenomena about which to gain intuition are ones outside human sensory experience. Mechano- and chemosensing at low to intermediate Reynolds numbers may be among the most alien; hence they require accurate description before they can be appreciated and succinct, logical description before they can be intuited. Advances on the front of understanding mechanosensory ecology recently have been stunning; data with broad scatter suddenly have collapsed onto simple curves defined by sys
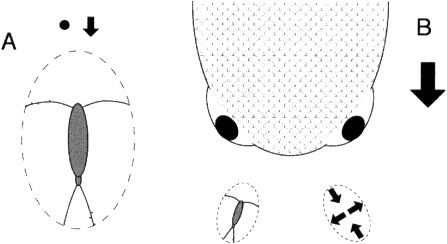
FIGURE 1. The two perspectives of copepod mechanosensing as predator and prey, respectively. A few mechanosensory hairs are sketched in on one antenna and the opposite caudal furca. The parcel of water within which the copepod is embedded is indicated by the dashed ellipse. A. Detection of small prey is via local perturbation of the velocity field. B. Detection of a predator by deformation of the water parcel in which the copepod is embedded. Qualitative features of the deformation are shown in the ellipse containing arrows. Based on analyses in Kiørboe and Visser (1999).
tematic decomposition of fluid dynamic phenomena into their constituent motions.
Kiørboe and Visser (1999) have idealized the motion of both prey and predator as flow around a sphere and have provided clear intuition for the mechanical perspective of a copepod in detecting and distinguishing moving prey and predators (Figure 1). This intuition comes not only from the calculated decomposition, but also from clever experiments that expose copepods to simplified and easily quantified fluid dynamic components of this decomposition. Arrays of mechanoreceptors on the antennae detect prey as local velocity variations. Independently performed experiments and numerical simulations (Bundy et al., 1998) show that the flow field generated by a swimming copepod also can contribute to detection of nonmoving, particulate prey.
Predators of copepods, on the other hand, are large in comparison with the copepod and are detected as larger-scale flow-field deformations that influence the whole space monitored by the copepod; the copepod knows that a predator is near when the stimulus affects the full array of sensors but deflects them with spatially varying velocities. Successful fish predators detect the copepod at a distance visually and decelerate as they approach, dropping the deformation produced by their bow wave below the threshold intensity that elicits the copepod's "jump" escape behavior. This threshold sits orders of magnitude above the neurophysiological detection limits of the mechanoreceptors and just above the level of deformation caused routinely by ambient turbulence. Kiørboe and Visser's (1999) analysis brings intuitive understanding of the process.
The qualitative gain in intuition for mechanoreception by copepods itself is compelling, but the gain is even more impressive when stimuli are quantified. Clearance rates by the ubiquitous omnivore Oithona similis are predictable over three orders of magnitude and on radically differing food particles (swimming protists and settling fecal pellets). Moreover, detection of settling particles by coprophagous copepods is found in calculations and experiments to be highly nonlinearly but predictably dependent upon particle size and settling velocity, to the point where detection of the most rapidly settling pellets at natural copepod abundances is virtually certain. Since McCave's (1975) seminal analysis, large particles or aggregates have been thought to account disproportionately for the flux of material to the sea-floor. Kiørboe and Visser's (1999) analysis suggests instead that particles of intermediate settling velocity that provide less mechanical stimulus for detection may be more successful in running the suspension-feeder gantlet. Particle-type-and settling-velocity-dependent degradation rates are among the most poorly constrained parameters in global carbon budgets, and this new analysis of mechanosensory abilities provides substantial help in the form of new perspectives and predictions from an unexpected direction. Suddenly complicating this range of issues in vertical transport of carbon still further is the documentation of spontaneous assembly of gels (Chin et al., 1998).
Diversity in Bacterial Tactics
Another important part of the carbon cycle is uptake of dissolved organic carbon by heterotrophic bacteria. Escherichia coli, a resident of the human large intestine and colon, has provided the universally used model of chemotaxis. Digestion in humans can be expected to yield large-scale
gradients relative to the body size of a bacterium (Dade et al., 1990), and the tumble-and-run approach makes good sense in the episodically stirred gut environment. E. coli runs in a straight line, then tumbles and goes off at a random heading from the original. Run duration increases with nutrient concentration, biasing the random walk and moving the bacterium up-gradient (Berg, 1993).
Most suggested sources of dissolved organic matter for bacteria are small in length and hence subject to rapid diffusion. In the case of leaking or exuding phytoplankton, the source may be quasi-steady, but in the cases of sloppy feeding, fecal pellet ejection, or autolysis, the source will be not only small but also short lived. An important question is whether marine bacteria show the same chemotactic behaviors as E. coli. Small bacteria apparently cannot find their way up gradient, and taxis is not practiced by bacteria smaller than about 0.6 gin in diameter (Dusenbery, 1998a). Below this size, the course of the bacterium would be changed too rapidly by Brownian rotation to allow directed movement. Conversely, the elongate shapes of chemotactic bacteria generally observed are particularly resistant to Brownian rotation and thereby increase the time over which the organism practically can integrate stimulus strength, and hence increase greatly its sensitivity to chemical gradients (Dusenbery, 1998b).
Recently, swimming paths of large marine bacteria have been recorded optically by taking advantage of light scatter under dark-field illumination (Blackburn et al., 1998). Unlike E. coli, they do not turn in a random direction at the end of a run but instead double back at close to 180° from the original heading, with some deviation due to Brownian rotation. This visualization was done in still water, but simulations in shear fields expected from decaying turbulence suggest that doubling back is far more effective than the tumble-and-run behavior in staying near a small, spherical source (Luchsinger et al., in review). There clearly is diversity in chemotactic strategies and patterns among marine bacteria (e.g., Barbara and Mitchell, 1996); E. coli is no longer an appropriate universal model. Models also suggest yet undocumented tactics. Quite contrary to intuition, bacteria can in principle use spatial sensing (difference in concentration on two parts of the cell) rather than temporal sensing (difference in concentration over time) to detect stimulus gradients (Dusenbery, 1998c).
Letting released enzymes do the searching for particulate material appears to be another useful bacterial strategy in either large aggregates of particles or in sediments (Vetter et al., 1998; Vetter and Deming, 1999) and explains the paradox of oversolubilization of aggregates by bacteria (more soluble products made than used; Smith et al., 1992). Further challenges to biological-physical modeling, to measurement of both bacterial tactics and carbon dynamics and even to the discrimination of dissolved from particulate carbon are the rapid self assembly and state changes of biogenic polymer gels (Chin et al., 1998).
The conceptual simplifications provided by Kiørboe and Visser (1999), steadily increasing abilities to, visualize flows and organisms both optically and acoustically, and increasing computational capabilities poise the study of fluid dynamic and chemical interactions with and among organisms for rapid advance. These advances promise in particular to help understand the vast and beautiful morphological diversity of protists and phytoplankton (though some modem classifications include the phytoplankton and even macroalgae with protists) living at low Reynolds numbers. Easier numerical modeling than at high Reynolds numbers is partial compensation in these regimes for lack of intuition about life in a fluid environment filled with dynamical chemical and physical signals.
STRUCTURING DYNAMICS OF BIOLOGICAL ASSEMBLAGES
-
Over what spatial scales are marine populations connected via dispersal of early life stages?
-
What are the dynamics of marine food webs, and how will they respond to environmental perturbations?
-
As models and synoptic data now are used to forecast the weather, can one forecast changes in physical-chemical-biological interactions in the sea that affect fisheries yields, food web dynamics, and ecosystem services that the sea provides?
This section covers population and community ecology but in the specific context of the ocean. Important facets of this context are the unquantified connectedness of subpopulations and the pervasiveness of fluid transport of propagules. This unquantified connectedness remains the greatest obstacle to rational establishment of marine preserves and management policy in general.
The OEUVRE workshop, to no one's surprise, cited identification of strong indirect effects through experimental manipulation as one of the great successes of marine ecology over the last 30 years and prediction of which interactions would be strong ones among the greate it challenges for the future. For few communities are the majority of interaction strengths known, but it is clear that in communities of even modest diversity the majority are weak (Paine, 1992). and identifying the important ones by manipulating all the species individually is a daunting empirical task. I left the workshop seeing no clear route to progress through predictive theory, either.
Theory suddenly has jumped to the rescue and shifted perspective by 180° by putting focus not on the strong interactors but on the weak ones. McCann et al. (1998) departed from classic food web models in two ways. They modeled functional responses as saturating rather than linear in prey concentration, and they allowed population abundances to be away from equilibrium values. With these additions to realism, food web models better reflect the added
stability observed as food web complexity increases. A deep insight from these models is that weakly interacting species in general damp oscillations. Although the paper was not specifically about marine communities, it is worth noting that communities with food webs rather than simple food chains, with omnivores rather than food specialists, with intraguild predation (predation on a competitor, particularly its young), and with allochthonous food supply are particularly stabilized. The majority of these characteristics apply to the majority of marine food webs. Berlow (1999) amplified this advance by demonstrating the efficacy of weak interactions in generating spatial and interannual variation in community structure with a simple mussel-barnacle-whelk system.
This shift in perspective is remarkable. When the goal was identification of strong interactors, they were hard to guess. When the goal instead is to find weak interactors with potentially large damping or amplifying effects, suspects jump to mind. It is easy to predict a cottage industry among ocean ecologists in the manipulation of omnivores, for example. Observations of strong interactions, particularly when the same species interact strongly in one place and not in another (e.g., the starfish-barnacle-mussel triad in Washington State and southeast Alaska; cf. Paine, 1980), translate into questions about what stabilizing species were missing in the former. Anchovies and sardines in this view exhibit such dramatic oscillations (Bakun, 1997) because they live in simplified food chains.
I cannot help but comment on how pleased I think that Gordon Riley would be to see ocean ecologists employ these new equations and a deeper understanding of the physical and chemical processes of the sea to write quantitative descriptions and predictions of marine ecosystem processes. He sought to combine understanding of the chemistry and physics of the oceans with the Lotka-Volterra equations systematically to dissect the workings of marine ecosystems. He supported use of the term "biological oceanography" to get away from the "grab-bag of semi-defined concepts [that he perceived to dominate the ecology of his day] to clear, step-wise analytical approaches to variation in nature" (Mills, 1995, p. 39). OEUVRE coined the term "ocean ecology" in this spirit but also to encompass the pressing need to obtain at least recent paleo-information about the workings of marine ecosystems.
HUMAN IMPACTS AND HABITAT LINKAGES
• How then can one understand the multiple-scale and pervasive human impacts on the sea in the face of the confounding effects of weather and climate change? Resolving and understanding anthropogenic and natural sources of variability and change on coastal to basin scales is arguably the greatest challenge to oceanographic science for the foreseeable future.
Anthropogenic effects from injection of nutrients and pollutants and from removal of predators certainly are pervasive. The challenge, given that a fully "natural" community free of anthropogenic effects appears to be a purely theoretical construct in the pejorative sense, is to avoid the trap of scurrying to understand the magnitudes of anthropogenic impacts without making the effort to understand their mechanisms. In the period since the OEUVRE workshop, I have not discovered any comparably perspective-shifting contributions to the ones mentioned under the other headings. This situation makes me continue to support the position taken by the OEUVRE group, that mankind is doing many manipulations without understanding their consequences and that a greater effort needs to be devoted to taking advantage of these manipulations to uncover the consequences and their mechanisms as the consequences arise rather than afterward. The surest method is to predict these consequences and then learn from the errors in the predictions as the perturbation proceeds.
Time-Series and Intervention Analysis
It is possible to look back at the other sections of this paper, however, and to note that time-series analysis has played a central role in gaining understanding that the CNP ecosystem is dynamically stable, and not statically stable, and that time series similarly have played a large role in dissecting food web interactions. It is not too soon to think about the consequences of increasing atmospheric inputs to the CNP from the industrialization of Asia and to ask how to resolve them from more natural variation. Time series again seem to be prominent in the answers.
One kind of time-series analysis, called "intervention analysis" (Box et al., 1994), appears to hold particular promise for characterizing some anthropogenic effects because it was developed to do so in the context of atmospheric pollutants. The procedure is to collect a long time series that includes the period before a change in policy or other anthropogenic perturbation whose timing is known. For example, one can look for changes in the record of lead deposition after the switch to lead-free fuels. The procedure is to fit an explicit time-series model to the pre-change data set. The model is then used to forecast the post-change data set, and the residuals from this forecast contain the treatment effect. Statistical power of this method depends on the length and simplicity of the pre-change time series (i.e., the ability to fit an explicit time-series model before the perturbation). Many ocean ecologists will be reluctant to trade traditional replication in space for replication in time, but if the whole system has been altered, then replication in space is elusive.
Ocean ecologists also must become as creative in extending time series backward as they are in extending them forward. Geochemists have made great contributions in this regard with chemical proxies for temperature and nutrients
and with biomarkers for some taxa. Archaeology through analysis of Indian middens has contributed to dissection of the sea otter-urchin-kelp interaction in the Aleutian Island chain (Estes et al., 1998). Where could additional effort by ocean ecologists produce the greatest extension and resolution back in time? The successes noted under functional ecology give reason to expect dramatic progress soon in understanding form and function in marine microfossils through understanding of form and function in today's fossilizable organisms. As the costs and benefits of simple shapes yield to analysis (e.g., Dusenbery 1998b), costs and benefits of more complex morphologies seem less daunting to study. Body form, spination, and mechanical properties of phytoplankton and protist individuals and chains certainly contain environmental information to be read. Continued development of "biomarker" compounds also certainly will be repaid. More conjecturally, establishing the extent to which buried bacterial communities reflect the conditions above and on the seafloor at some previous time (i.e., while the surface mixed layer of sediments was in contact with the overlying water) versus their environmental conditions at present may allow extraction of other paleoenvironmental information. At issue is the length of time that bacteria can survive in inactive state and be interrogated by molecular means in this biochemically messy medium.
ACKNOWLEDGEMENTS
I thank John Cullen and Dick Barber for constructive criticism of earlier drafts of this paper and Thomas Kiørboe and David Karl for sharing their unpublished manuscripts.
REFERENCES
Arrigo. K.A., D.H. Robinson, D.L Worthen, R.B. Dunbar, G.R. DiTullio, M. VanWoert, and M.P. Lizotte. 1999. Phytoplankton community structure and drawdown of nutrients and CO2 in the Southern Ocean. Science 283:365-367.
Bakun, A. 1997. Radical interdecadal stock variability and the triad concept: A window of opportunity for fishery management science. Pp. 1-18 in T.J. Pitcher, P.J.B. Hart, and D. Pauly (eds.), Reinventing Fisheries Management. Chapman and Hall, London.
Berg, H.C. 1993. Random Walks in Biology. Princeton University Press, Princeton, New Jersey.
Berlow, E.L. 1999. Strong effects of weak interactions in ecological communities. Nature 398:330-334.
Blackburn, N., T. Fenchel, and J. Mitchell. 1998. Microscale nutrient patches in planktonic habitats shown by chemotactic bacteria. Science 282:2254-2256.
Box, G.E.P., G.M. Jenkins. and G.C. Reinsel. 1994. Time Series Analysis: Forecasting and Control. Prentice-Hall, Englewood Cliffs, New Jersey.
Bundy, M.H., T.F. Gross, H.A. Vanderploeg, and J.R. Strickler. 1998. Perception of inert particles by calanoid copepods: Behavioral observations and a numerical model. J. Plankton Res. 20:2129-2152.
Chin, W.C., M.V. Orellana, and P. Verdugo. 1998. Spontaneous assembly of marine dissolved organic matter into polymer gels. Nature 391:568-572.
Dade, W.B., P.A. Jumars, and D.L. Penry. 1990. Supply-side optimization: Maximizing absorptive rates. Pp. 531-556 in R.N. Hughes (ed.), Behavioural Mechanisms of Food Selection. Springer-Verlag, Berlin.
Dusenbery, D.B. 1998a. Minimum size limit for useful locomotion by free-swimming microbes. Proc. Natl. Acad. Sci. USA 94:10949-10954.
Dusenbery, D.B. 1998b. Fitness landscapes for effects of shape on chemotaxis and other behaviors of bacteria. J. Bacteriol. 180:5978-5983.
Dusenbery, D.B. 1998c. Spatial sensing of stimulus gradients can be superior to temporal sensing for free-swimming bacteria. Biophysical J. 74:2272-2277.
Estes, J.A., M.T. Tinker, T.M. Williams, and D.F. Doak. 1998. Killer whale predation on sea otters linking oceanic and nearshore ecosystems. Science 282:473-476.
Falkowski, P.G. 1997. Evolution of the nitrogen cycle and its influence on the biological sequestration of CO2 in the ocean. Nature 387:272-275.
Hessler, R.R., and P.A. Jumars. 1974. Abyssal community analysis from replicate box cores in the central North Pacific. Deep-Sea Res . 21:185-209.
Karl, D.M. In press. A sea of change: Biogeochemical variability in the North Pacific Subtropical Gyre. Ecosystems.
Karl, D.M., J.R. Christian, J.E. Dore, D.V. Hebel, R.M. Letelier, L.M. Tupas, and C.D. Winn. 1996. Seasonal and interannual variability in primary production and particle flux at Station ALOHA. Deep-Sea Res. 43:539-568.
Karl, D.M., R.M. Letelier, D.V. Hebel, L. Tupas, J.E. Dore, J.R. Christian, and C.D. Winn. 1995. Ecosystem changes in the North Pacific subtropical gyre attributed to the 1991-92 El Niño. Nature 373:230-234.
Kiørboe, T., and A.W. Visser. 1999. Predator and prey perception in cope-pods due to hydromechanical signals. Mar. Ecol. Prog. Ser. 179:81-95.
Laws, E.A., L.W. Haas, P.K. Bienfang, R.W. Eppley, W.G. Harrison, D.M. Karl, and J. Marra. 1984. High phytoplankton growth and production rates in oligotrophic Hawaiian coastal waters. Limnol. Oceanogr. 29:1161-1169.
Luchsinger, R.H., B. Bergersen, and J.G. Mitchell. In review. Bacterial swimming strategies and turbulence. Biophysical Journal.
Margalef, R. 1978. Life-forms of phytoplankton as survival alternatives in an unstable environment. Oceanologica Acta 1:4923-509.
McCann, K, A. Hastings, and G.R. Huxel. 1998. Weak trophic interactions and the balance of nature. Nature 395:794-798.
McCave, I.N. 1975. Vertical flux of particles in the ocean. Deep-Sea Res. 22:491-502.
McGillicuddy, D.J., and A.R. Robinson. 1997. Eddy-induced nutrient supply and new production in the Sargasso Sea. Deep-Sea Res. 44:1427-1450.
McGowan, J.A., and P.W. Walker. 1985. Dominance and diversity maintenance in an oceanic ecosystem. Ecol. Monogr. 53:103-118.
Mills, E.L. 1995. From marine ecology to biological oceanography. Helgoländer Meeresunters. 49:29-44.
Paine, R.T. 1980. Food webs, linkage, interaction strength and community infrastructure. J. Anim. Ecol. 49:667-685.
Paine, R.T. 1992. Food-web analysis through field measurement of per capita interaction strength. Nature 355:73-75.
Perry, M.J. 1972. Alkaline phosphatase activity in subtropical central North Pacific waters using a sensitive fluorometric method Mar. Biol . 15:113-119.
Perry, M.J. 1976. Phosphate utilization by an oceanic diatom in phosphorus-limited chemostat culture and in the oligotrophic waters of the central North Pacific. Limnol. Oceanogr. 21:88-107.
Smith, D.C., M. Simon, A.L. Alldredge, and F. Azam. 1992. Intense hy-drolytic activity on marine aggregates and implications for rapid particle dissolution. Nature 359:139-142.
Venrick, E.L., J.A. McGowan, D.R. Cayan, and T.L. Hayward. 1987. Climate and chlorophyll a: Long-term trends in the central North Pacific Ocean. Science 238:70-72.
Vetter, Y.A., and J.W. Deming. 1999. Growth rates of marine bacterial isolates on particulate organic substrates solubilized by freely released extracellular enzymes. Microbial Ecol. 37:86-94.
Vetter, Y.A., J.W. Deming, P.A. Jumars, and B.B. Krieger-Brockett. 1998. A predictive model of bacterial foraging by means of freely released extracellular enzymes. Microbial Ecol. 36:75-76.
Villareal, T.A., C. Pilskaln, M. Brzezinski, F. Lipshultz, M. Dennett, and G.B. Gardner. 1999. Upward transport of oceanic nitrate by migrating diatom mats. Nature 397:423-425.








