5
Portals to Knowledge: Information Technology, Research, and Training
Eric G. Jakobsson
University of Illinois at Urbana-Champaign
INFORMATION TECHNOLOGY WILL BLUR THE BOUNDARY BETWEEN RESEARCH AND EDUCATION
Let me tell you what I mean by “information technology,” or the appropriate information environment for any field of study. What I mean is that you ought to be able to fire up your Web browser and have, at your fingertips, all the important information sources, plus the tools to analyze that information, calculational tools to correlate and make sense out of the raw data, visualization tools when that is appropriate, simulation tools, and molecular dynamics and quantum chemistry tools. It all ought to be available, point and click, on your Web browser. That is the kind of information environment that I am suggesting will blur the boundary between research and education.
If you create that environment, you will not only make the access to this information and the use of it more efficient for the research scientist, but you will also make it accessible to the student. You are going to incorporate this environment into the classroom, and students will use it for undergraduate research and for exploring and learning in a hands-on way, at least in terms of the computational sense of things, and in a hands-on way about the nature of their subject.
INFORMATION TECHNOLOGY WILL BLUR THE BOUNDARIES BETWEEN DISCIPLINES
Information technology is already blurring the boundaries between disciplines. We used to have students climb the ladder of knowledge. There was a ladder for chemistry, a ladder for biology, a ladder for physics, and so forth, and they were next to each other. Now, students navigate the web of knowledge. On the Web, you can move sideways as well as vertically. You can move in any direction as easily as any other direction.
If I want to know something about nitrogen, for example, and I look on the Web, it is equally easy for me to access a description of industrial processes for synthesizing ammonia, or a description of the natural processes by which nitrogen is cycled through the biosphere.
IT IS AS EASY TO MOVE ACROSS THE BOUNDARIES OF OUR TRADITIONAL DISCIPLINES AS IT IS TO MOVE WITHIN THE BOUNDARIES OF OUR TRADITIONAL DISCIPLINES
As more and more of our exploration is in knowledge spaces like this, the divisions between disciplines are going to be blurred. We have talked a little bit about specialization and generalization. It certainly is true that there is a long tradition of both interdisciplinary work and of scientists being generalists. The relative importance of interdisciplinary work has waxed and waned. It seems to me that after World War II there was a tremendous move toward specialization in science, which began with big federal funding of research. Perhaps this was the beginning of the various pressures that we have talked about in terms of being in the grant rat race.
I believe that information technology will start to reverse this trend; in fact, it already has started to make it possible for individuals to find out about things more efficiently, for individuals to become multiple experts. Indeed, because of the pace of change and the various tasks that people are going to do during their lifetimes, it will be almost mandatory for people to become multiple experts.
INFORMATION TECHNOLOGY WILL BLUR THE DISTINCTION BETWEEN HERE AND THERE
I certainly don’t mean that interacting with people over the Web should replace mentoring. As collaboration tools get better and better, they are not a way for people to avoid communicating with each other, or avoid mentoring each other—they are other ways for people to communicate and mentor. I can tell you the innermost contents of my mind as well over the Internet as I can in person. In fact, I may be even less inhibited about doing so.
THE BIOLOGY WORKBENCH
I would like now to show you what I am talking about in terms of the Biology Workbench. Figure 5.1 shows what we call the National Computational Science Alliance (Alliance) Information Workbench, for which the Biology Workbench is the prototype. The user interface in this diagram is the Web browser. The Web browser connects to the guts of the Workbench, which is the Workbench server. The Workbench server translates formats, creates queries that databases can understand, and drives application programs. The application programs have various interfaces, as can the information sources, and can be written in different languages. All of these can be readily tied together by a powerful scripting language such as Practical Extraction and Report Language (PERL), which is the easiest to use. But you can script in C as well as in PERL.
The key is that we now have the ability to take, for example, a whole series of databases in varying formats, such as various molecular biology databases, and a whole series of application programs, such as programs to visualize molecules, align molecular sequences, and construct phylogenetic trees of relationships based on those sequences, and so forth, and make them all look like one program. You interface through them and point and click as though you were on a Macintosh.
I know that this is not a Macintosh, but the point is to make everything in the world look like one. This is what we are really about. I tend to think in terms of biology, but basically the challenge is the same for chemistry. In chemistry you have databases that give you physical properties of chemicals; you have databases that give you structures, and so forth. For each discipline, what you really want to
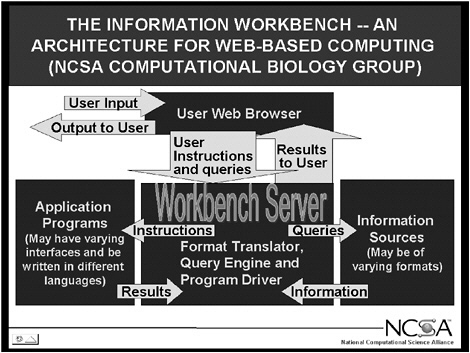
FIGURE 5.1 The National Computational Science Alliance Information Workbench, the prototype for the Biology Workbench.
have is a single, seamless computer interface providing access to all the data and visualization and analysis programs.
The next series of figures (Figures 5.2 through 5.12) are screen shots that show this type of interface for biology, the Biology Workbench, at <www.ncsa.uiuc.edu>. Figure 5.2 shows the interface for the Biology Workbench, with a picture of my colleague Shankar Subramaniam, the primary inventor and driving force for development of the Workbench. Although he has now moved to the University of California, San Diego, we still do some collaboration.
The Biology Workbench has multiple functionalities that can be accessed by simply highlighting and clicking on the desired function (Figure 5.3). To illustrate the capabilities of the Workbench, I will perform a search for human myoglobin. Once entered (Figure 5.4), the Workbench searches through multiple databases simultaneously, with thousands of sequences. The results returned by the search include the number of database entries found relating to “human,” the number relating to “myoglobin,” and the number that related to both (Figure 5.5).
One of the four databases returned from the human myoglobin search is illustrated in Figure 5.6. The database contains hyperlinks to such things as the original paper (Figure 5.7) and the structure
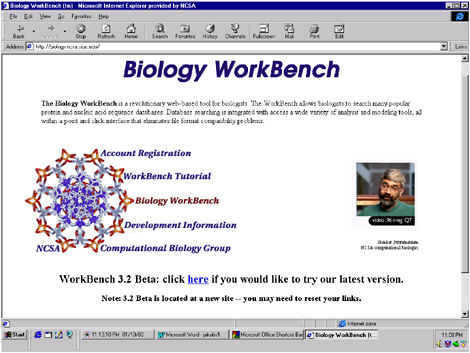
FIGURE 5.2 The Biology Workbench interface. Shankar Subramaniam, the primary inventor and driving force for development of the Workbench, is pictured on the right side.
(Figure 5.8). The structures can then be manipulated, and various measurements can be done on them with a plug-in called Chime, which is familiar to many chemists as well. It is really a computational chemistry tool.
The Biology Workbench can also be used to search the databases of amino acid sequences to find a particular sequence (Figure 5.9). The Workbench then allows you to take that sequence and use a program called BLAST to find closely related sequences. For human myoglobin, BLAST returned several “hits” of myoglobin sequences from a chimpanzee, another kind of ape, a gibbon, a gorilla, and an orangutan (Figure 5.10). The hit sequences can be aligned to provide a clear picture of sequence homogeneity (Figure 5.11). The regions that are highly conserved tell us one thing, and parts that are variable tell us another. The parts that are highly conserved must be critical for defining the basic structure and function of the molecule. The pattern of variability in those regions that are not conserved tells us the evolutionary history of the molecule and, by extension, of the organisms.
The Workbench can further be used to create a phylogenetic tree based on the pattern of differences in the amino acid sequence alignment (Figure 5.12). For our sample case, it is seen that the human (myhu) is grouped with the African apes (the chimpanzee, mycz, and the gorilla, mygo), and this cluster
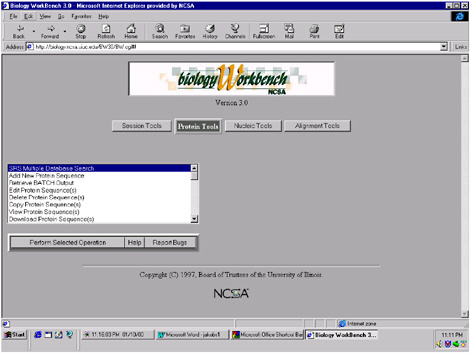
FIGURE 5.3 This picture illustrates the highlight-and-click process involved in selecting the functionality of a database search in the Biology Workbench.
is a short distance from the Asian ape (orangutan, myog), with the other primates farther away. This pattern is confirmed in the analysis of many genes.
All of the calculations shown in the above sequence can be done in point-and-click fashion in the Biology Workbench in 15 to 20 minutes. All the user needs is a networked desktop machine with a Web browser. The universal access, plus the ease and speed of accessing the data sources and doing the analysis, clearly makes this tool potentially useful in the classroom. But students and teachers need some introductory training in the use of the tool. Instead of trying, ourselves, to introduce the Biology Workbench to all of the faculty in the country, we started a process this past summer, in collaboration with the BioQUEST Curriculum Consortium at Beloit College, of building a community of biology instructors nationwide, working with them, and having them work with each other, to build curricular materials around the Biology Workbench and, in this fashion, to build a community of people who are building curricular materials.
We believe that the Workbench architecture is a prototype for the use of computing in education, and for much of scientific computing in general. It is the basis of the National Computational Science
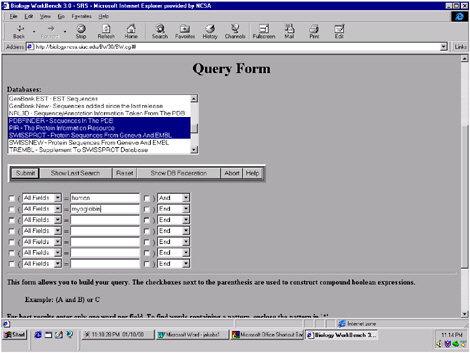
FIGURE 5.4 Screen from the Biology Workbench showing how to submit a search for human myoglobin to several databases at the same time.
Alliance Common Portal Architecture Project, which is summarized in Figure 5.13. A major part of further extensions of this kind of computer architecture is to build in collaboration tools so that people can work with each other remotely, and to extend the Workbench environment to more intensive computing than we now do on the Biology Workbench. The jobs that we do on the Biology Workbench are mostly not terribly computer intensive. It is reasonable for them to be done interactively.
We are currently extending the Biology Workbench capabilities to a variety of computational chemistry functionalities. We are building Web interfaces for molecular dynamics, stochastic dynamics, and electrostatic programs. These applications are computer intensive and will require us to do more extensive things with computing architecture to ensure that resources are set aside in advance for doing this type of work.
For many years it has been speculated that computers would have a big impact on education. Until very recently, this promise has not been fulfilled to a significant degree. Now, because of the extraordinary versatility of the Web for communication and data access, the enormous increase in underlying computer power, and the ease of use of the Web, we believe that high-performance, versatile, and user-
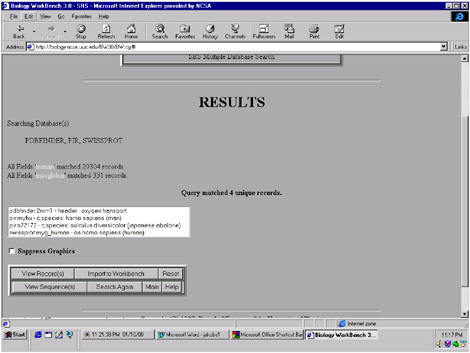
FIGURE 5.5 Results of the search for human myoglobin showing that it found 20,304 database entries related to “human” and 331 related to “myoglobin,” with just 4 entries relating to both.
friendly computing and information environments are poised to have a profound impact on chemistry and all scientific education.
DISCUSSION
Robert L. Lichter, The Camille & Henry Dreyfus Foundation: A question that arises repeatedly, as we move into the technology of information access, is the value of the information obtained and the need for arbiters, referees, and other gatekeepers. What do you foresee as some of the challenges that arise in that context, as we proceed along this track, through all the portals, collaboratories, and the rest?
Eric Jakobsson: It is an enormous challenge, and there are of different ways to deal with it. For example, the physics community, which has a big repository of papers in Los Alamos, doesn’t vet anything before it goes in and prints everybody’s comments. So, if somebody publishes a paper about
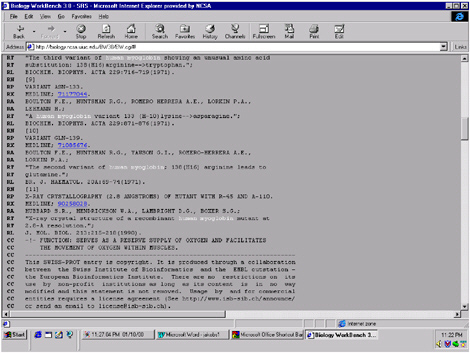
FIGURE 5.6 Part of one of the database entries given for human myoglobin. The entry includes hyperlinks to the original paper as it is indexed in the National Institutes of Health PubMed query page.
a perpetual motion machine, they count on other people heaping scorn upon it; the comments are attached to that paper, and nobody will pay any attention to it.
Biologists tend to be more conservative than that; they are very concerned about the accuracy of information. In fact, there is a growing amount of literature about people finding mistakes in the sequence databases.
My colleague Shankar Subramaniam has developed a very powerful algorithm called PDS, Potential Density Functions. It assigns parallel probabilities of interatomic distances between different kinds of atoms in a protein structure, based on statistics derived from known structures. It turns out that this is an extremely good filter for finding mistakes in structures in the protein data bank. As this data proliferates and more and more people depend on the data bank, we are going to need to pay more attention than we have to the accuracy of the data.
I should say that currently the big emphasis is on dealing with the volume of information. Mostly it is really good, but there is enough that is not that has become a nuisance, especially when you build
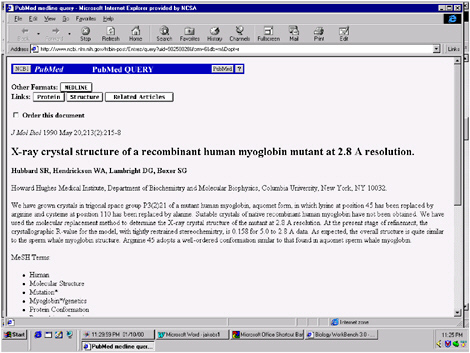
FIGURE 5.7 Original paper for the structure of myoglobin obtained for the hyperlink in the database illustrated in Figure 5.6.
something that you are doing on it. So, people really do need to be guided through using these environments. Of course, this is the great overall problem in the information flood—separating the junk from the good stuff.
People talk a lot about democratization of the Internet. They talk about two tendencies: on the one hand, we should use the Internet to democratize information; on the other hand, there is a big difference in how people use this information—and it depends on economic class, which cuts across the same fault lines as everything else in our society. I think the technology is getting so cheap that the real fault line is not going to be between people who physically have access to it or not; it will be between people who have access to the right guidance in using it or not.
James D. Martin, North Carolina State University: I would like to comment on using this kind of technology in an educational setting. By all means, I use it in the classroom, and I find it quite helpful. But, I also am feeling an increasing distaste for Web evangelists. Maybe it is because I am from the Bible Belt. There seems to be an almost evangelical flair to Web proponents. As one of my colleagues at NCSU suggested, we seem to be creating a click and drool population: we click on it, it looks cool,
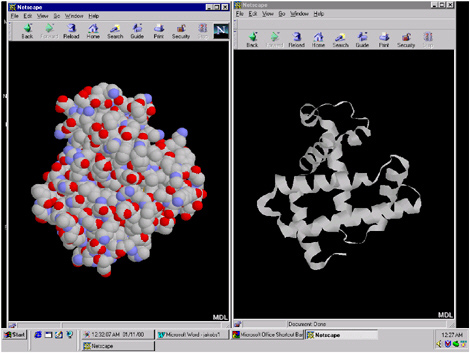
FIGURE 5.8 Illustration of the myoglobin structure. The space-filling structure is illustrated on the left and the ribbon structure on the right. The structures were obtained from a hyperlink from the database seen in Figure 5.6.
and therefore it is good. I also see a lot of research proposals that require an educational component, such as for the Research Corporation or for a NSF CAREER award, for which the educational innovation boils down to putting something else onto the Web. In my teaching of both freshman chemistry and advanced inorganic chemistry at the graduate level, I have two experiences that I think are important to relate.
At NCSU, we have redesigned our freshman chemistry curriculum and have computer support that goes with our lectures that has animations and demonstrations that I can’t do on the blackboard. They are extremely useful. But at the end of the semester I had a number of students crying for me to go back to the blackboard, for a simple reason: what I wrote on the blackboard could be copied into their notes. This is an accessibility issue that affects a student’s belief that “one day I am actually going to be able to do this myself.” By contrast, click and drool demonstrations tend to be so beautiful that we become users only and don’t really believe that we could ever do them ourselves.
In my graduate course, advanced inorganic chemistry, structural aspects are extremely important. And thus, I try to teach visualization techniques. I start with point groups, then teach space groups and
FIGURE 5.9 The database of the sequence of amino acids is also available from the National Computational Science Alliance Biology Workbench, as illustrated here.
the whole works. I try to get the students to learn how to take a crystal structure that is reported in a journal, draw a picture of it, rotate it, and to get a feel for what structure is like. I wasn’t succeeding in getting many people to get a real grasp of it. Then I asked myself what it was in my training that gave me this ability to connect 2D to 3D, which we do, whether it is on a computer screen or in a textbook.
What suddenly dawned on me was that my wood shop classes in the seventh, eighth, ninth, and tenth grades were probably the greatest asset to my education in terms of my being able to connect two dimensions to three dimensions, a skill that is absolutely important to chemistry. My seventh-grade shop teacher would not let us touch a tool until we had drawn a set of blocks to learn drafting skills, so that we could perceive two and three dimensions. I said, what the heck, and I got the guys in the shop to create a set of blocks. For advanced inorganic chemistry, students had to draw two perspectives of a set of blocks. I was amazed. They had a much better grasp of dealing with the computer screen after using the blocks. And on exams—I also teach molecular orbitals—I actually got pictures that I could interpret, as opposed to some kind of a scrawl.
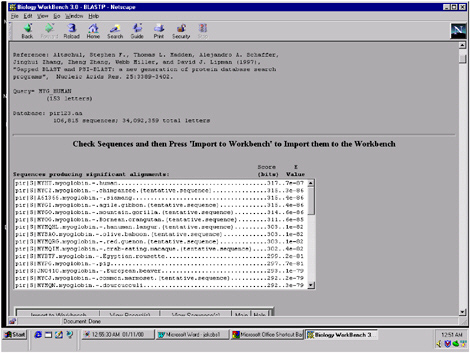
FIGURE 5.10 The results of an operation called BLAST on the sequence in Figure 5.9 are seen here. This operation involves sliding the given sequence across all the thousands of sequences in the database of Protein Information Resource, and finding the ones that are most closely related. This figure shows the “hits”—the most closely related sequences.
I guess I am calling for a bit more moderation. Let’s move away from the evangelical Web and recognize where it is and is not useful. But also remember that the “traditional university” is this amazing beast, that by definition will be changing all the time, but that has also lasted in an amazingly constant format for thousands of years. I am not sure that we have to reinvent things quite as much as we sometimes hear. Let’s take advantage of what is there, an institution that has weathered time and many changes and is going to stay. I think Web moderation as opposed to evangelism is going to be the key.
James S. Nowick, University of California, Irvine: Let me start by saying that the example you presented was of the highest caliber as a pedagogic exercise for students. One of the problems, as we develop sources of information for students, is similar to that of drinking from a fire hose. As educators, we need to teach them how to sip from the fire hose. This is what I try to teach them in my courses. One thing that can impede the portability of what you have developed is that the instructor must be involved in the creative process to do a really good job. If I simply take someone’s curriculum resources and try
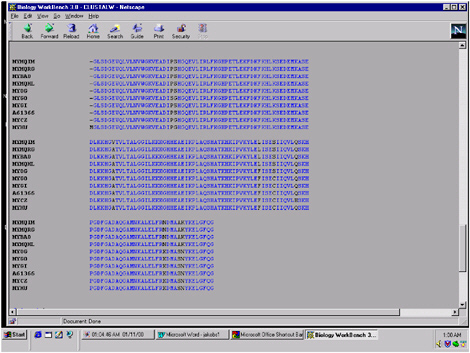
FIGURE 5.11 The sequences of all of the primary hits can be imported into the Biology Workbench workspace and aligned with each other as is illustrated here. The alignment is shaded more lightly when all the sequences are identical with each other and printed darkly when there is variation from one to the other.
to implement them in my classroom, without being personally involved in the creative process, I will do a mediocre job. If I have an intimate role in the creative process, then I will do a great job. As this stuff propagates to your first generation of biology instructors, and then to their peers, part of what will determine the success is how much they embrace this as something that they have put their own brains into.
Eric Jakobsson: Let me comment on that and reinforce what you are saying. That is exactly why we wanted to engage a substantial community of people in developing materials, as opposed to doing that ourselves. I agree with that. I also agree with the call from the previous speaker, to be as interested in what the Web can’t do, and what it needs to be supplemented by, as what it can do.
Frankie Wood-Black, Phillips Petroleum: I want to make a comment with regard to the fire hose aspect of the Web and the ability of the Web to do some interesting operations. One is that the Web
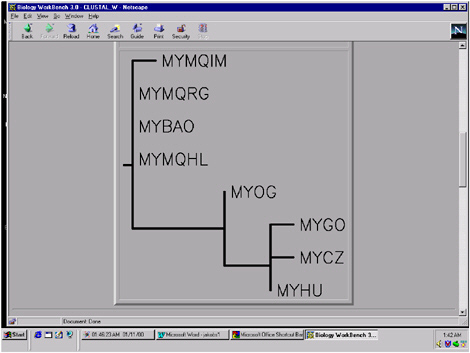
FIGURE 5.12 A phylogenetic tree can then be constructed from the pattern of differences in the alignment above in the Biology Workbench. The total length of the branches between each pair of species is proportional to the evolutionary distance between them.
provides access for everyone to get information. Increasingly, we are passing the point where it is the haves and the have-nots in terms of access but rather having the mentor or guide on how to use it. You have got to build on the fundamentals of where the understanding comes from, and they are doing it now; for example, you have a third-grade class that is sent to the library to figure out what a resource is. The Web is only one resource that can be built upon. The user needs to develop the capabilities of sorting through what is good and what is not good for an overall analysis. We have got to be sure that we are providing the guidance as well as the information.
Eric Jakobsson: I don’t intend this to be evangelic, but I think the power of the Web is overwhelming. We talk all the time about the bottom line. The bottom line is that, in terms of how many bytes of information you can transmit per dollar, including dollars for infrastructure, this is a much more powerful technology than, for example, the printed word, in addition to the various things that it can do.
It is like a fire hose. We are all in the stream of this fire hose, and we are not going to turn it off. It
FIGURE 5.13 Executive summary of the National Computational Science Alliance Common Portal Architecture Project, taken from the Web site of Dennis Gannon of Indiana University, the “roadmaster” of the Alliance effort in this area.
is a question of diverting the “stream” to a useful problem—it is an engineering problem analogous to how you are going to get hydroelectric power from a waterfall. You are not going to turn the waterfall off, but you figure out instead some system of turbines and generators, and so forth. We have got an information flow that we have to figure out how to engineer into a useful information technology.
Ernest L. Eliel, University of North Carolina: I find myself somewhat in tune with the previous speakers. I think one of the problems is that we haven’t yet managed to map a computer network on the neuronal network in our minds. I think that causes a certain amount of problems, and let me give one specific example. You replaced, in one of your early slides, a disciplinary staircase with a disciplinary framework. My experience is that when I go one step to the right or one step to the left, I have serious vocabulary problems. When I go two steps to the right or two steps to the left, I have very serious conceptual problems. I am not sure that the computer could help me with that. Certainly, I could push the help button, but I am not sure that the right answer would come up.
Eric Jakobsson: Actually, we had a dialog on this, at one point, in Physics Today. The question was about what physicists could contribute to biology. My former colleague Gregorio Weber, who has passed away, said a wonderful thing about biology and physics. He said that if you get a physicist with N good ideas together with a biologist with N good ideas, then between them you will get 2N. If you can get all those good ideas and understanding in one brain, then you are going to have N factorial good ideas.
We are beginning to do this, but we increasingly have to train people earlier, at least some of our young scientists, in an interdisciplinary mode, as opposed to expecting physicists and biologists to get together later in their careers, or chemists and physicists and biologists.
Timothy A. Keiderling, University of Illinois at Chicago: I would like to speak from this morning’s point of view and note how this presentation might reflect back on it. We talked about the need to speed things up and the need to be multidisciplinary in graduate education. It is the kinds of tools made available by information technology that can allow us to maintain a shorter time frame and let the students take their physical or chemical backgrounds and address the biological problem.
This kind of structure visualization tool allows my students to get to the real science more quickly. They used to dig up a structure on the Protein Data Bank and then run around finding the paper, then run around trying to see if they could find a program that could tell them the secondary structure, and finally, they would try to find some answers to a structural question. Now, if they can push some buttons in a unified software package, they can ask the scientific questions that they are trying to answer, and they can stay on theme better. That will allow them to meet these requirements that we were asking for this morning; that is, to keep on a reasonable time scale and yet still be multidisciplinary. I think these technologies have a powerful role. At the same time, like many of you, I am a bit frightened, since I worry whether students learn anything with this quick process. We must make them learn the basic skills so they can utilize the technology meaningfully.
Eric Jakobsson: You know, all new information technologies are frightening. From my reading—I am not a historian—my understanding is that there was a tremendous issue around the invention of the printing press, and how socially dangerous it would be to let everybody read the Bible for themselves, without the proper vetting by the ecclesiastical authorities. This is exactly the kind of question that has been dealt with before.
Dale Poulter, University of Utah: I happen to come from one of those western states that is very conservative and careful with their money. Our governor is the one who instituted the Western Governors University. For those of you who have not heard of it, this is a way to get an undergraduate degree on the Web. I think there is going to be a push into the graduate area as well at some point. Clearly, it is a way to transmit information, and to correlate information, and there is no denying that. The real concerns I have are about socialization, because although chat rooms are nice, I think humans need the one-on-one visual contact with another person to become properly socialized.
I have the same concern about mentoring. Effective mentoring has to be done up close and personal. The other problem that may not be recognized is that when you deliver a Web-based class, you don’t save time. It is almost like a British tutorial in that you can’t answer or address a single question to 50 people simultaneously and get instant feedback. You have to do it one by one by e-mail. It takes a tremendous amount of time.
Eric Jakobsson: I second the motion. Getting involved with these types of issues is not a way to work less hard.
John Schwab, National Institute of General Medical Sciences: I have heard people here comparing the Internet-driven information glut to a fire hose. This is a valid metaphor in terms of quantity, but not in terms of quality. The water that comes out of a fire hose is of uniform quality. What comes off the Internet is quite variable; there are good data and lousy data. There are good applications and flawed applications.
Unfortunately, there has long been a tendency, even in pre-Internet days, to think that if data were obtained from a machine, that those data were somehow validated. Clearly, that is naive. The utility of the Internet as a learning tool depends ultimately on how knowledgeably it’s being used. This leads me back to Dale Poulter’s point regarding the importance of mentoring. It’s absolutely critical that Internet users be taught to question and evaluate the quality of the information that they are accessing.
Eric Jakobsson: I would even go further than that. There are not only bad data and bad applications, but there is also evil. There are hate groups. We learned a couple of generations ago that there is nothing incompatible between evil and technical competence. That extends to the Web as well. It is up to people who want to, and are capable of doing good things with it, to take it over and use it as well as possible.

















