7
The Interface with Biology and Medicine1
|
Some Challenges for Chemists and Chemical Engineers
|
|
GOALS
The goal of fundamental science at the interface of chemical sciences and biology is to understand life in chemical terms. Our ability to accomplish the translation of biology into chemical terms is closely related to our fundamental understanding of life’s machinery. Decoding these mysteries and producing a detailed molecular picture of how things work is a critical step. However, for most people, the real impact comes when we can use that understanding to improve our lives. Nowhere has the role of chemical sciences been better illustrated than in the discovery, development, and production of new medicines and ways in which we can more selectively deliver these medicines to the organ or tissue where they are needed. This fundamental understanding of what chemical transformations occur in living creatures, how these chemical transformations are regulated, and how they respond to extracellular stimuli is also critical to developing semisynthetic tissues and organs as replacements for damaged organs, to gene therapy, and to solving a host of clinical problems. In addition, these chemical tools will provide the basis for improvements in the productivity of agriculture in both animal and plant production systems. Most directly, advances in the chemical sciences will be essential for the prevention, diagnosis, and treatment of disease.
Underlying Science
The mechanisms of life are revealed when we understand their molecular details. Moreover, molecular understanding of biology plays the major role in
guiding drug discovery, and molecular-level understanding of biology is fundamental in developing diagnostic methods.
Nature is a highly innovative chemist, and we know only a small fraction of the universe of natural products. Currently, about half of the most widely prescribed drugs in the United States are either small-molecule natural products, such as penicillin, or synthetic molecules that are based on natural products but have better properties. The other half are products of the creative imagination of medicinal chemists. Nature’s genetic and chemical diversity is a treasure that should not be squandered. One reason to maintain ecosystems such as rain forests intact is that less than 10% of plant species have been systematically investigated for their chemical products. Also, many microbes, insects, and other species have not been explored for useful products.
Discovering which natural product is useful has historically been a slow, laborious effort. Our improved understanding of the molecular basis of life and of disease has allowed us to develop methods for rapid screening of libraries of natural products for compounds that interact with specific molecular targets. Synthetic chemists have used the techniques of combinatorial chemistry to generate large numbers of chemical compounds that also can be screened in a similar manner to identify promising drug candidates. Tremendous advances have been made in our capabilities to rapidly make a wide variety of complex new chemicals for such testing.
Although we do not know the identity of all chemical species in a living cell, we do know many of the most essential components and have determined their molecular structures. Molecular analysis of living cells has led to a flow plan of information in molecular biology, known as the Central Dogma: The information necessary for a cell is encoded in the double helix of DNA. DNA acts as a template for its own replication, and segments of DNA that encode information for the primary structure (i.e., amino acid sequence) in a protein are called genes. The information for production of a protein is first transcribed from DNA through synthesis of a messenger-RNA molecule by the process of transcription, and regulatory elements on the DNA help determine when and how often a particular gene is transcribed. The information on the messenger RNA is converted to a protein through the process known as translation; if the physiological conditions in the cell are correct, the nascent protein folds into its proper three-dimensional shape. Proteins act as catalysts (called enzymes), transporters, receptors of information, regulators, and structural elements in the cell. Their three-dimensional shape is critical to biological activity.
The language of biology uses four letters (A, T, C, and G corresponding to four different nucleotides) in DNA, and three-letter words. Each three-letter word specifies a particular amino acid (or signals to start or stop translation). Other combinations of DNA letters (of various lengths) combine with proteins in the cell to block or encourage transcription of genes; these combinations of letters constitute regulatory elements. The language of biology is universal; the same
language applies to humans, plants, and microbes. This universality makes genetic engineering and gene therapy feasible.
Recent advances in technology, using modern chemical techniques for structure determination, have allowed us to sequence or read the letters in the DNA, or genome, of a cell. We have genomic sequence data for humans, an insect, a plant, simple multicellular organisms, and many microbes; more are constantly being produced. These sequences represent the total genetic and biochemical blueprint for each of these organisms. This information is useless unless we learn to read it intelligently; that is, to relate linear sequence information to cell and organismal function. Functional genomics is a term used to describe that relationship, and it will be a primary challenge for the next 50 years.
The simple sequence of letters in the genome tells us only a little. The details of three-dimensional structure are important to understanding the chemical processes of life. For example, the discovery that DNA forms a double helix made it clear how genetic information is passed on and utilized. The biological activity of the proteins encoded on the DNA is dependent on their specific three-dimensional structure. Many chemists are concerned with how best to determine the structure of such proteins. Ideally, that structure could be predicted from the sequence of amino acids that correspond to the code in the gene on the DNA. This remains a challenging problem in computational chemistry (Chapter 6), but a combination of experimental and theoretical techniques have advanced our understanding of structure and function in proteins. For example, x-ray methods can be used when a protein can be crystallized (although this is often difficult). Nuclear magnetic resonance (NMR) techniques can be used to probe the structure of proteins in solution. Many of the computational techniques are related to recent advances in sequencing of DNA. Bioinformatics includes computational chemistry with the goals of predicting function and three-dimensional shape directly from the amino acid sequence—by comparison with sequence, function, and structural information for other proteins (often from other organisms).
Knowing protein structure can provide direct benefits to human health. The precise molecular structure of a protein gives it great selectivity in distinguishing among substrates. Sometimes that activity can be blocked by molecules that are similar to the substrate but do not cause a reaction or response when bound to the protein.
For example, with the crystal structure of the aspartyl protease from human immundeficiency virus (HIV-1) in 1989 came the opportunity to design molecules to block this important enzyme that acts as a molecular scissors. HIV is the virus responsible for AIDS. Essential to viral replication, the HIV protease cuts long strands composed of many proteins into the functional proteins found in mature virus particles. This proteolysis occurs at the very end of the HIV replication cycle (Figure 7-1). The three-dimensional structural information derived from the x-ray crystal structure, combined with computer modeling techniques, allowed chemists to design potent, selective inhibitors of the protease enzyme (Figure 7-2).
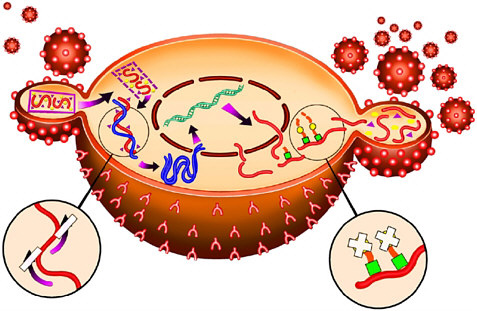
FIGURE 7-1 The HIV replication cycle. A virus particle (small sphere) attaches to a cell in the immune system such as a lymphocyte (large sphere, shown here in cross section) as the first step. Two complementary approaches to AIDS therapy utilize (a) reverse transcriptase inhibitors (enlargement, lower left) to block incorporation of the viral genome into the host cell, and (b) HIV protease inhibitors (enlargement, lower right) to block cleavage of proteins into key functional enzymes that are required for mature viral particles. Copyright Merck & Co., Inc., Whitehouse Station, New Jersey, USA, All Rights Reserved, used with permission.
Design of complex molecules that “dock” into a target is only a first step— the molecules had to be synthesized from simple building blocks. The excellent stereocontrol of reactions like the Jacobsen and Sharpless epoxidations and the Evans aldol reaction was essential to the construction of molecules with as many as five chiral centers (for which 25 or 32 possible stereoisomeric forms are possible). Target molecules had to be made in pure form and tested, and eventually the single “active” isomer had to be produced with >99.8% purity at a level exceeding 100,000 kg/year. A decade ago, production of synthetic drugs with this level of molecular complexity was inconceivable. Protease inhibitors have become an important part of the treatment for AIDS patients.
A major lesson learned from the first draft of the human genome sequence2 is that there are fewer genes (perhaps only 30,000) than originally predicted (over
100,000). This was a surprise to biologists, who had speculated that the number of genes in an organism’s genome might scale according to that organism’s level of complexity. Instead, some simple worms such as nematodes have nearly as many genes as human beings! Where are the complex functions encoded that are characteristics of higher mammals? One answer seems to be in the post-translational modification of proteins. During or after their biosynthesis, proteins can be modified with myriad added decorations, such as phosphate, sulfate, and acetyl groups; the modified proteins can have vastly different properties compared with those of their unmodified progenitors. Thus a single protein, derived from a single gene, can be transformed into numerous distinct molecular species and thereby amplify the information content of a very concise genome. Comparisons have shown that post-translational modifications are more extensive in higher organisms.
The chemistry of polysaccharides is a major frontier. Their branched and complex nature makes them far more difficult to synthesize than the linear biopolymers, proteins, and nucleic acids. For this reason, automated synthetic methods for polysaccharides were just created in 2001, several decades after com-
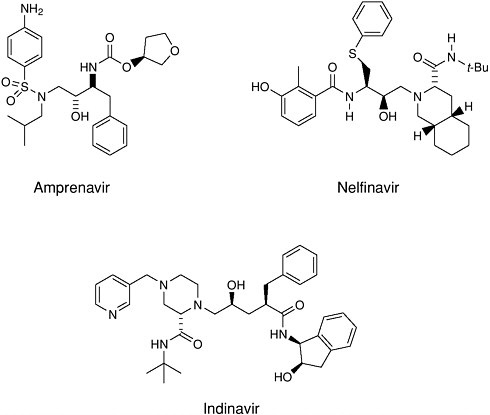
FIGURE 7-2 Examples of HIV protease inhibitors.
parable methods were perfected for the linear biopolymers. There is still a need for improved syntheses of the linkage that connects sugar residues within a polysaccharide chain, the glycosidic bond. At the moment, each specific linkage must be made with different optimal reagents, making a generalized automated synthesis very difficult.
Our increased understanding of the biology of polysaccharides has led to major breakthroughs in medicinal chemistry. The attachment of white blood cells to cells that line the walls of blood vessels is a hallmark of inflammatory conditions such as rheumatoid arthritis, psoriasis, and transplant rejection. Disruption of this cell-cell binding event would stop the inflammation and improve the patient’s health. It was discovered that the pathological binding of these two cell types is mediated by noncovalent association of a receptor on the white blood cell, termed L-selectin, with polysaccharide ligands on the blood vessel walls. Soluble versions of that polysaccharide that can bind to the receptors and thus inhibit cell binding are now in clinical trials as anti-inflammatory drugs.
In most cases, the precise functions of polysaccharides are not known; even their primary sequences are very hard to determine using current analytical techniques. Thus, a major challenge is to crack the “carbohydrate code” and determine the structures and functions of all the polysaccharides found on human cells. Terms such as “glycomics” have already been coined to describe such global efforts.
Biochemical Engineering
Biochemical engineering is both an established and an emerging field that is inherently multidisciplinary. It integrates chemical engineering with biology and biochemistry—to participate in biological discovery or to create processes, devices, or therapies.
For example, bioprocess engineering is concerned with the development of manufacturing processes based on living cells or enzymes to make pharmaceuticals, food ingredients, or chemicals. Much of biochemical engineering activity is directed toward human health and forms an important branch of biomedical engineering, particularly in activities such as drug delivery devices, artificial organs, and tissue engineering (e.g., artificial skin for burn victims).
Modern biochemical engineering has become so focused at the molecular and cellular level that the difference between molecular and cellular biology and molecular and cellular engineering can be difficult to discern. Biologists and biochemists want details of mechanisms and have successfully used a reductionist approach in which they strip away all elements except those directly under study. Biochemical engineers typically require a less detailed description of component mechanisms, but they need to understand how the cell’s components interact with each other. Thus, biochemical engineers must view the cell or collection of cells as a whole system. It is often this integrative approach that distinguishes bio-
chemical engineers from biologists and biochemists. The integrative or systems perspective of the biochemical engineer complements the traditional reductionist approach of the biologist.
Another distinction between the biochemical engineer and the biologist is that the creation of a process, therapeutic strategy, or device is the ultimate goal of the engineer, while discovery is the primary goal of the biologist. However, the engineer’s approach often leads to new discovery, and many biologists participate in creation. Since some properties of systems emerge only when component parts are integrated together, the complete discovery and understanding of living cells will require the integrative approach common in engineering, in addition to reductionist approaches.
Modern biochemical engineering began with the challenge of large-scale production of penicillin by fermentation during World War II. This challenge required the active cooperation of microbiologists, biochemists, and chemical engineers. Today many small-molecule natural products are made from microbes in large (>100,000 L) fermenters followed by a series of separation and purification operations.
Other major early contributions of biochemical engineering have been in the development of the artificial kidney and physiologically based pharmacokinetic models. The artificial kidney has been literally a lifesaver. Pharmacokinetic models divide the body of an animal or human into various compartments that act as bioreactors. These mathematical models have been used very successfully in developing therapeutic strategies for the optimal delivery of chemotherapeutic drugs and in assessing risk from exposure to toxins.
More recent accomplishments include development of bioprocesses for production of high-purity proteins, using genetically engineered cells. This challenge was particularly difficult with respect to the development of large-scale systems for production of therapeutic proteins using mammalian cells suspended (or dispersed) in a liquid medium. Examples of some of these products are tissue plasminogen activator used to treat stroke and heart attack patients, erythropoietin to treat anemia resulting from kidney damage or chemotherapy, and granulocyte colony stimulating factor as an adjuvant to chemotherapy and cancer treatment. Another accomplishment has been the development of effective devices for the controlled release of pharmaceuticals and therapeutic proteins. Localized delivery of drugs (for example, in the brain) has been a key to effective treatment of cancers and other ailments.
Commercial production of paclitaxel (better known by the commercial name Taxol), an important anticancer agent, has been accomplished using plant cell tissue cultures, where cells or small aggregates of cells are suspended in a liquid medium. This technological advance could lead to general methodology for the production of plant-based medicinals from rare and difficult-to-grow plant species. Tissue-engineered products such as artificial skin and cartilage have been recently introduced into the market. The need to combine living cells and poly-
mers under the strictest conditions of sterility challenged the ability of the engineers to mass produce and package these products..
The preceding accomplishments are applied in nature, but required tremendous amounts of basic research on mass transfer, interactions of materials with biological components, fluid dynamics, separation processes (especially chromatography and membrane separations), and biochemical kinetics.
Biochemical engineers also have made major contributions to a better understanding of cells, tissues, and organisms. These contributions fall mainly under such categories as metabolic engineering, cellular engineering, and hemodynamics.
Metabolic engineering refers to purposeful genetic manipulation of the biochemical pathways in a cell to produce a new product, to eliminate unwanted reactions, or to increase the amount of a desired product already produced by a cell. The optimal approach to effective metabolic engineering of a cell is essentially quantitative in nature; biochemical engineers have played the major role in the development and application of the appropriate mathematical models (e.g., metabolic control theory). In addition, chemists have developed a better understanding of the modular cellular machinery used to synthesize natural products and have used the techniques of metabolic engineering to harness these modular processes to create novel molecules. An excellent example is recent work to make novel polyketide antibiotics. Biochemical engineers do much of the current experimental work on metabolic engineering in collaboration with other chemical scientists.
Cellular engineering, while closely linked to metabolic engineering, is concerned not so much with metabolic pathways but instead with processes such as cell adhesion, cellular mechanics, signal transduction due to growth factors and hormones, and viral infections. By combining mathematical models of these processes with clever experiments we have far greater insight into these biological mechanisms. Such insight can assist in the design of drug candidates. As an example, both models and experiments show that the intuitive assumption that the most effective inhibitor of a receptor-based process would have the highest binding affinity is wrong. Due to receptor recycling a nonmaximal binding affinity is optimal.
|
Large-Scale Production of Proteins from Recombinant DNA Using Suspensions of Animal Cells When recombinant DNA technology was developed in the late 1970s and early 1980s, many thought that all proteins could be produced in easy-to-grow bacterial cultures. This optimism proved incorrect; bacteria were incapable of making some proteins in a therapeutically useful form. |
|
For these proteins, mammalian cells proved better hosts, as they could process the protein with intracellular machinery similar to that in humans. However, large-scale production of proteins in cell culture was problematic. Mammalian cells had to grow attached to a solid surface, such as glass in “roller bottle” culture. While the Federal Drug Administration (FDA) had approved some processes for vaccine production that used cell cultures, it required that these cells be “normal.” Normal mammalian cells can divide only a few times before they stop growing, making scale-up to large volumes difficult. The first large-scale process to circumvent these limitations was one developed by Genentech, Inc. (South San Francisco, CA) for production of recombinant tissue plasminogen activator (tPA). This protein dissolves blood clots and can be used to treat heart attacks and strokes. This process was developed in the mid-1980s, resulting in final product licensure in 1987. The process required both regulatory and technical breakthroughs. Genetic scientists chose to use a Chinese hamster ovary (CHO) cell line as a recombinant for production of human therapeutics. This cell line is “transformed,” which means it can replicate indefinitely. All cancers arise from transformed cells, so Genentech had to convince the FDA that their process, including recovery and purification, would eliminate the possibility of injecting patients with a “transforming factor” that could cause cancer in a patient. In addition, they had to adapt the CHO cells to grow in suspension culture (i.e., cells dispersed in liquid medium and unattached to any surface) and to grow in a medium without serum (which is produced from blood by the removal of cells). Serum-free suspension cultures of CHO cells greatly simplified processing problems and reduced cost. The absence of serum and the contaminating proteins it contained simplified recovery and purification of tPA, and reduced degradation as tPA was secreted from the CHO cells. By using suspensions, and thus being freed from the constraints of surface-to-volume ratios inherent in roller bottle or similar techniques, it was possible to design large (12,000 L) stirred bioreactors that could be readily scaled up and that would maintain high levels of volumetric productivity (mg tPA/L-hr). The bioreactor design issues were complex, as no one before had built a bioreactor that could supply sufficient oxygen and nutrient to the cells in such a large system while stirring gently enough not to damage the cells with high levels of fluid shear. This required novel designs of reactor shape and impeller, based on an understanding of the cellular growth kinetics, mass transfer, and fluid dynamics. The process required development of a novel large-scale, low-shear, tangential flow filtration technology for the rapid, aseptic removal of spent medium from cells. |
|
The success of the project depended on the ability of biochemical engineers, chemists, and life scientists to work together. Success depended on an integrated vision of the process and an understanding of how each component interacted with the other components. While production processes based on serum-free suspension cultures of CHO cells are now common, the integrated approach to bioprocess development remains essential to success. Chemists and biochemical engineers need sufficient knowledge in the life sciences to interact fruitfully with biologists. The cell remains the “real” bioreactor, but efficient processes required the design of macroscopic bioreactors and operating strategies that effectively couple to our knowledge of the cellular reactors housed in human-designed bioreactors. |
Other examples of important insight obtained by biochemical engineers are the effects that physiological values of fluid stress have on cells. Cells that line blood vessels are normally exposed to blood flow. This flow places a mechanical stress on a cell that not only alters cell shape but also its function. The mechanical stresses alter expression of genes. The mechanisms by which this occurs in cells are incompletely understood and a matter of active research.
PROGRESS TO DATE
When one looks back at the last decade of the 20th century it is astonishing to see what was achieved in the understanding of life’s machinery. Decoding these mysteries and producing a detailed molecular picture of how things work is a critical step. However, nowhere has the role of chemistry been better illustrated than in the creation and development of new medicines and therapies.
|
Selective Asthma Therapy In the 1930s an unknown material was hypothesized that was proposed to cause a slow and sustained contraction of smooth muscle. It was named the “slow reacting substance” (SRS). By 1940 a similar substance was reported to be found in guinea pig lungs and was called the “slow reacting substance of anaphylaxis” (SRS-A). Over the next 40 years, while no one could isolate, characterize, or synthesize this mate |
|
rial, the belief was that SRS-A played a key role in human asthma. In 1979, Bengt Samuelsson proposed a structure for SRS-A that was derived from arachidonic acid and the amino acid cysteine. He called it leukotriene C. 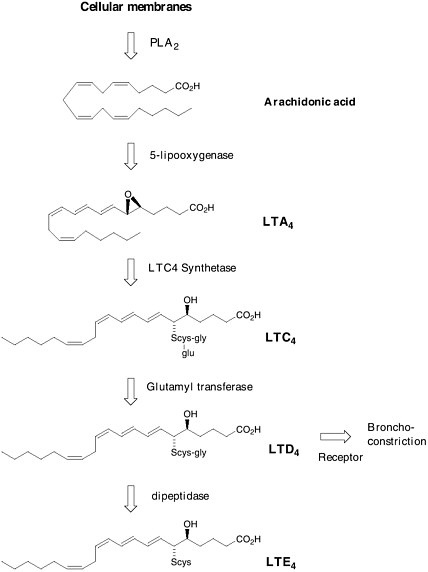 Formation of leukotrienes via the arachidonic acid cascade. Soon after, the complete structure of SRS-A was finally determined by total synthesis. SRS-A turned out to be a mixture of 3 substances now known as leukotriene C4 (LTC4), leukotriene D4 (LTD4) and leukotriene E4 (LTE4) in which LTD4 was predominant. The jump from a biological observation in 1938 to a molecular structure of LTD4 opened the door to a novel and selective treatment for asthma. The theory was that if one |
|
could find a molecule that specifically blocked the action of LTD4 on the lung, it would be possible to prevent the tightening of the airways found in asthma. Eighteen years after Samuelsson’s proposal, after the synthesis and testing of thousands of man-made compounds, Singulair (montelukast) reached the world’s pharmacies. Leukotriene modifiers are recognized to be the first important advance in asthma therapy in 25 years. 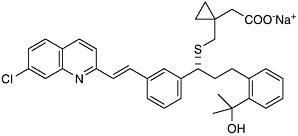 Singulair (montelukast) This is just one example of how the chemical understanding of one of life’s processes at the molecular level resulted in the solution to a 60-year-old problem. The selective LTD4 receptor antagonist was specifically designed to take the place of LTD4 on smooth muscle. Such approaches—where the drug molecule does only what it’s intended to do, without indiscriminately binding to other receptors in the body—are aimed at selective therapies that are free of side effects. It is that selectivity, designing molecules with precision based on structural information, that characterizes the state of the art of chemistry as we enter the next millennium. |
Underlying the discovery of a selective asthma therapy are numerous advances in analytical and instrumental techniques as well as synthetic methods that allow the construction of complex molecules. Practical catalytic, stereospecific, and organometallic methods that permit a high level of stereochemical control have enabled production at the multiton level of molecules previously inaccessible even at the gram scale.
|
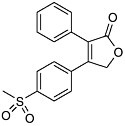
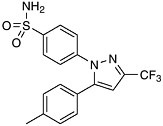
Selectivity and Anti-Inflammatory Drugs The theme of selectivity—based on detailed understanding of molecular structure and function—underlies most recent therapeutic advances. Sometimes a biochemical “revisiting” of an old discovery enables dramatic improvements in the quality of life. Among the most widely used classes of medicines are the nonsteroidal anti-inflammatory drugs (NSAIDs) such as aspirin and ibuprofen. Used for years, these pain killers and arthritis treatments work by blocking the effects of arachidonic acid on an enzyme called cyclooxygenase (COX). A major drawback to inhibiting COX is that by doing so you inadvertently block its role in protecting the gastrointestinal tract. The resulting ulcerative gastropathy is responsible for a large number of hospitalizations and deaths. Recently it was discovered that cyclooxygenase is not a single enzyme but rather a family containing at least two nearly identical members: COX-1 and COX-2. While COX-1 is responsible for the good gastroprotective effects (and shouldn’t be blocked) COX-2 is the enzyme involved with pain and inflammation (the real target). The COX-2 hypothesis stated that if one could invent a specific COX-2 inhibitor, it would be an effective anti-inflammatory and analgesic medication with substantially reduced gastrointestinal (GI) toxicity compared with the classical NSAID’s aspirin and ibuprofen. 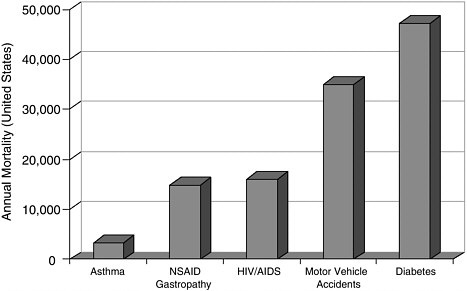 Significant mortality is associated with NSAID gastropathy. G. Singh and G. Triadafilopoulos, Epidemiology of NSAID induced gastrointestinal complications, Journal of Rheumatology, 1999, 26, 56, 18-24, by permission of Oxford University Press. |
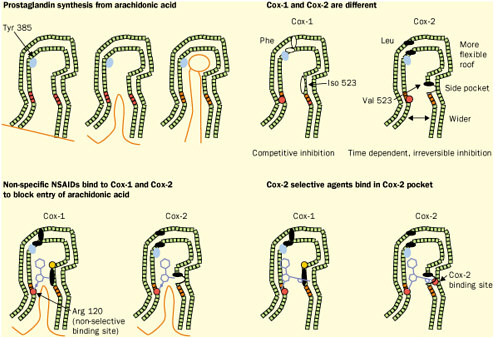 Figure reprinted with permission from Elsevier Science (The Lancet, 1999, 353, 307-314). Again, detailed structural information at a molecular level was the key. Once these “pictures” were available slight chemical differences between the COX-1 and COX-2 isoenzymes could be seen: COX-2 had a side pocket while COX-1 didn’t. This meant that a molecule that could dock into the COX-2 side pocket (binding site) but not into COX-1 would specifically block COX-2 without touching COX-1. In 1999 this hope was realized with the availability of COX-2 selective anti-inflammatories such as Rofecoxib and Celecoxib that can be as much as 50-fold selective for the target.
|
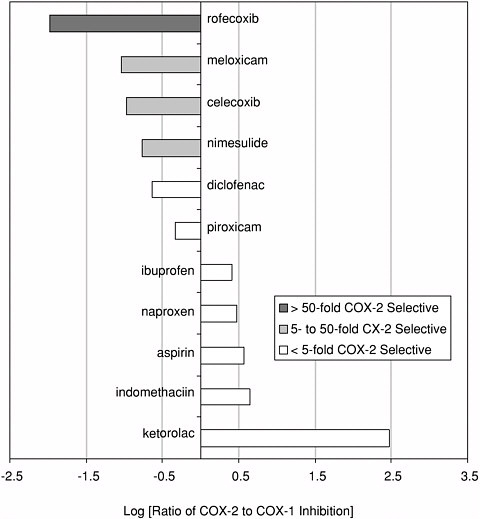 COX-2 selectivity of marketed compounds. Adapted with permission from T.D. Warner et al. Proceedings of the National Academy of Sciences, 96, 13, 7563 (1999). Copyright 1999 National Academy of Sciences, U.S.A. |
|
Specificity and Therapy for the Human Brain Nowhere is the need for specificity so great as in trying to design therapies for the human brain. Here there are numerous receptors that affect our moods, sleep, alertness, memory, and coordination. Even though the importance of serotonin (5-hydroxytryptamine, 5-HT) had been known to neuroscientists for over 100 years, it wasn’t until the discovery of drugs |
|
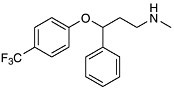
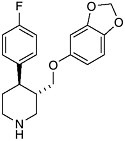
like Prozac (fluoxetine hydrochloride) and Paxil (paroxetine hydrochloride) that selective antidepressive drugs became available. The antidepressive effect results from inhibition of serotonin uptake by neurons in the brain, thus ensuring that circulating levels are adequate. The selectivity results from selective binding compared with older drugs (tricyclics) that were less discriminating in their binding to other brain receptors. The net result is fewer side effects. As the molecular basis of memory and behavior become clearer, we will see a leap in the effectiveness and specificity of drugs for the central nervous system. The arrival of the year 2000 coincided with a milestone in modern science; many believe that deciphering the human genome will provide a road map for therapeutic intervention. We are already seeing medicines that act not directly on a target tissue but on receptors that regulate the transcription of genes. The ability to “tune” the molecular signaling that continually occurs in our bodies will eventually allow for more exquisite control of the cellular processes of life. If today’s chemistry lets us turn things on or off by blocking or unblocking receptors and enzymes, tomorrow’s molecules will be able to balance complex metabolic processes like growth and aging by fine tuning the regulation of genes and their products.
|
CHALLENGES AND OPPORTUNITIES FOR THE FUTURE
The opportunities for discovery and invention at the interface of chemistry, engineering, and biology are enormous, and many examples have been described in the preceding sections. This interface represents a true research frontier—one that is critical to our ability to develop new chemistry for the prevention, diagnosis, and treatment of human disease. The continuing challenge is to discover the chemical identity of all the molecules that make up living organisms and the way they bind to each other and organize into biological structure—membranes and cell structures such as nuclei, ribosomes, etc.
|
Ribosomes The nucleotide sequences of the genes encoded in DNA are transcribed into the same sequences of messenger RNA, again carrying the genetic code, and these RNAs direct the synthesis of proteins with defined sequences of amino acids. Protein synthesis occurs in a cellular machine called a ribosome, which takes in messenger RNA and some amino acids linked to transfer RNA and uses base pairing to direct their assembly into a protein. The base pairing occurs because each kind of amino acid is attached to a different transfer RNA, one that carries a code for that particular amino acid. Thus messenger RNA plays the role of template for protein synthesis, and transfer RNA guides the amino acids to the correct spot. Assembling the protein still requires a catalyst to link the amino acids together. Essentially all biological catalysts in the modern world are themselves proteins, enzymes. However, in 1989 Sidney Altman and Thomas Cech received the Nobel prize in chemistry for showing that RNA itself could act as a catalyst for some biological reactions. This led to the idea that in an earlier time, as life was evolving, RNA may have been both the information molecule (a role usually played by the more stable DNA now) and the catalyst (the role that protein enzymes now play.) Since this idea indicates that in early times the synthesis of proteins was catalyzed by RNA, not by protein enzymes, the intriguing question is whether this is still true today. Ribosomes are complex structures consisting of one small RNA molecule and two large ones together with some 50 to 60 different proteins. Their general shape had been determined by electron microscopy some years ago, but a major breakthrough occurred in 2000. X-ray diffraction was used to determine the detailed molecular structure of a ribosomal particle that consists of almost all the molecules in a ribosome and exhibits the full catalytic and regulatory functions of a ribosome.3 The trick was to get this large particle, about 100 times as large as a simple protein enzyme, to crystallize so the x-ray technique could be applied. The results are striking, but for the real details it is important to consult the original articles). Such a detailed structure can help medicinal chemists to develop useful drugs that bind to the ribosome, but the more important result has to do with the catalytic center. The catalytic center of the ribosome, where the protein is actually made, is now seen to consist of RNA, not of a protein enzyme! The many |
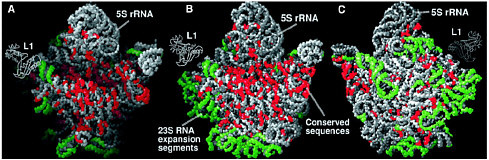 The large ribosomal subunit at 2.4 Å resolution. (A) The particle rotated with respect to the crown view so that its active site cleft can be seen. (B) The crown view. (C) The back view of the particle, i.e., the crown view rotated 180° about its vertical axis. Reprinted with permission from Ban et al., Science 289, 905 (2000). Copyright 2000 American Association for the Advancement of Science. proteins present help organize the structure, but they do not play a catalytic role. This is in line with the idea that the original process in early life forms used RNA alone. As proteins were created in processes guided and catalyzed by RNA, some proteins were incorporated into the RNA catalytic unit during evolution and improved the ribosome’s function. A follow-up paperf4 indicates a way in which the RNA could carry out the catalyzed synthesis of proteins. A particular adenosine unit in the catalytic RNA has the correct properties to be able to assist the formation of peptide bonds, which are the links in proteins. All this is basic chemical research that has wide importance, determining the molecular structure of a component of living cells. Simply, it tells us the details of how proteins are now made, but more generally it strengthens the picture of how life may have started in a world where RNA was both information molecule and catalyst. It is a major advance in scientific understanding. |
One driver for new discovery will be the completion of the human genome project. As a result of this project, the locations and sequences associated with the tens of thousands of genes of the human genome will have been determined. In the post-genomic era, then, we will know the DNA sequences and genes of a human being, but that is only the start. What are the functions associated with these sequences? We will need to isolate the proteins that are the gene products,
determine their structures, characterize their reactions and their partners in reactivity. To perturb the actions of these proteins, we will need to develop diverse arrays of small-molecule inhibitors and activators. While the human genome project gives us access to the library of the cell, we will need to learn much to exploit the information that we will have.
For example, although we designate gene sequences by a one-dimensional string of letters, the protein gene products they encode have three-dimensional architectures, and when properly folded they carry out biological function. A critical challenge in this post-genomic era will be to make the connections between protein sequence and architecture, and between protein architecture and function. We need to learn to predict how a protein folds and the relationships between a folded structure of a protein and its function. Indeed, in reaching that level of chemical understanding, we can then seek to design new functions for proteins. Both combinatorial and rational design strategies may be applied in the construction of novel biologically based catalysts.
Genomics and gene arrays will become increasingly important in strategies to prevent disease. Today, for example, we make use of simple assays such as the PSA (prostate screening antigen) test in the early detection of prostate cancer. Transcriptional profiling using gene chip technology will likely facilitate analogous tests for all cancers by simultaneously measuring all relevant mRNA levels. We will be able to determine what mRNAs or small natural products rise in concentration in association with cancers, and hence use detection methods for these molecules in cancer prevention. A challenge resulting from the enormous amount of information associated with such transcriptional profiling will be the determination of causal relationships, and of the partners and pathways associated with cellular transformation. Indeed, a major intellectual challenge to the chemical sciences is developing a systematic framework and computational tools to relate microarray data, as well as data on protein levels, to a description of the dynamic regulatory networks controlling cellular functions.
Proteomics is a combination of experimental and analytical tools to determine the total protein content of a cell or tissue. While genomics identifies the potential proteins in an organism, proteomics provides information on which proteins are actually present in a tissue under specific environmental conditions and accounts for the physiologic history of the tissue. Proteomic information is difficult to obtain due to the large amount of chemical heterogeneity displayed by proteins and our inability to amplify the amount of proteins in a sample. In contrast, polymers of nucleic acids are rather more chemically homogeneous and can be amplified using the chemical technique of polymerase chain reaction (PCR). Currently much proteomics work is accomplished using a technique called two-dimensional gel electrophoresis. This technique is slow, requires highly skilled technicians, and many proteins in a cell are too rare or too hydrophobic (water hating) to be resolved by this technique. Nonetheless, 2-D gel electrophoresis has been used successfully to monitor prion diseases (e.g., mad cow disease) and
shows promise in the diagnosis of neurodegenerative disorders such as Alzheimer’s disease. A major opportunity in the chemical sciences is the development of microarray technologies that can provide more rapid and complete proteomic information. The techniques of biotechnology applied on the nanoscale may result in miniature devices that can be used in a massively parallel fashion to do rapid separation and analysis of DNA, RNAs, or protein solutions.
Progress in genomics and proteomics offers opportunities also in the diagnosis and treatment of disease. We will soon be able to develop the chemistry necessary to routinely detect genetic variations in individuals, and to do so for a full range of genes. In fact, some pharmaceuticals that are largely effective and might represent medical advances currently fail during clinical trials as a result of an adverse reaction within a genetic subpopulation. With genetic screening, and a better understanding of the underlying biochemistry, we will be able to tailor-make therapies for patients based on their genetic dispositions.
More generally, as described earlier, medicinal chemistry has greatly contributed to the fight against disease, but there are still major challenges ahead. For example, we don’t yet have generally effective drugs to treat viral diseases, such as influenza or Ebola (although some trials of a new drug for the treatment of influenza indicate that it decreases the length of the infection). This is critical. Imagine the problem if the HIV virus that causes AIDS, or the Ebola virus that kills quickly, were able to be transmitted by the bite of a mosquito, as some other less lethal viruses can be. Until we have effective medicines to cure such viral infections, humanity is at great risk.
Another problem is bacterial resistance to antibiotics. As doctors have treated people with the available antibiotics that medicinal chemistry devised in the past, they have selected for strains of bacteria that are resistant to those antibiotics. There is now a race against time by medicinal chemists to devise new antibiotics that will work against the resistant organisms. If we do not succeed, many bacterial infections that we thought had been cured will emerge again as major threats to our health and life.
We still need much better medicines to cure cancer, heart disease, stroke, and Alzheimer’s disease. We need better drugs to deal with obesity, diabetes, arthritis, and schizophrenia. The treatments of diabetes, arthritis, and mental defects such as schizophrenia or manic depression are not yet cures, just ways to keep the symptoms under control. Cures are needed. Insights from genetics may help guide us toward elegant and rational cures, but we will also make use of screens to identify natural products and libraries of randomly generated synthetic compounds (combinatorial chemistry). A semi-empirical approach may be the best hope over the next two decades to yield drugs to alleviate these diseases.
Many of the natural products may come from “unusual” organisms and may be difficult to synthesize. In those cases, it will be necessary to develop appropriate bioprocesses to produce, recover, and purify these compounds. Both chemists and biochemical engineers will be involved in creating such processes.
But medicinal chemistry will also change in basic ways. Indeed, we are entering a completely new era of molecular medicine. We will develop technologies to screen the effects of small molecules on large arrays of gene products, from enzymes to receptors, and doing so will require advances in fields ranging from biochemistry to material design. We will develop the tools to create genomic maps of protein-protein contacts and chemical tools to decipher the hierarchy of those contacts. Our ability to digest and exploit the enormous information we obtain will also provide challenges in computation, in structure prediction, and in our quantitative understanding of molecular recognition. As a result, fundamentally new strategies will be developed to attack disease on a molecular level. For example, it is already clear that strategies to fight cancer are shifting, from those centered on maximizing toxicity in cancerous cells to those where we activate or harness different signaling pathways of the cell, depending on whether the cell has undergone transformation from normal to cancerous.
Among the fundamental new strategies, and certainly an important step to be taken by chemists in this new era of molecular medicine, will be developing a general understanding of how small molecules can be utilized to regulate gene expression and signal transduction. The goal is the design of small molecules not simply as general poisons to the cell or to some cellular function, but instead as reagents that turn off or turn on critical pathways. This challenge depends upon advances also in our understanding of molecular recognition. When a protein or small molecule binds to a particular receptor, an ensemble of weak noncovalent contacts are specifically arrayed in three-dimensional space to facilitate this recognition of one molecule by another. Chemists are now working on general strategies to achieve highly specific molecular recognition, and doing so is a first step in the rational design of new drugs as regulators of cellular processes.
Associated with this question is how to target these small molecules to sites of specific action in the cell or tissue. We have, for example, made substantial progress in delineating how some small molecules and metal ions are trafficked through the cell. Can we apply this knowledge to invent strategies for targeting small molecules to specific organelles within the cell? Our current understanding of what controls cell permeability and bioavailability is primitive, often not appreciably more advanced than “oil versus water.” As we develop a molecular perspective concerning the trafficking of molecules into and through the cell—as well as the chemistry underlying what distinguishes the surfaces of different cells—we will establish a more rational approach to targeting molecules more powerfully, and even with tissue specificity.
Because cells and the body respond not only to genetic information but also to environmental cues, any analysis must take into account the time and environment-dependent nature of the biological system. Because of their training in analysis of integrated systems, biochemical engineers should be able to contribute integrated, quantitative models of these biological systems to guide the selection of targets for intervention and the synthesis of a precise delivery system. In some
cases these devices will need to be “smart” devices to respond to a current physiological state. An example of such a device, already in research and development, is one to monitor blood glucose levels and to release insulin in response to changes in blood glucose level. This device would effectively mimic the responses of the natural pancreas. Other delivery systems may mimic viruses for DNA delivery to specific target cells as a more controllable method for gene therapy. Indeed, the controlled delivery of macromolecular therapeutics with temporal and spatial control of therapeutic distribution is an important goal for chemists and chemical engineers.
|
Controlled Delivery of Therapeutics Many pharmaceuticals are designed to effect change in a single organ or tissue. Traditional methods of drug delivery using pills or injections require the pharmaceutical to enter the blood stream and to be dispersed throughout the body (systemic delivery). Often undesirable side effects occur in nontarget organs before a therapeutically useful level of the agent is achieved at the target organ or tissue. Alternative methods of drug delivery are needed to deliver the drug to the right tissue, at the right time, and at the right amount. As an example, consider treatment of a brain tumor. The brain protects itself from the entry of potentially toxic substances through a blood-brain barrier, which is a highly organized cellular barrier to the transport of such compounds from the blood into the brain. To administer a chemotherapeutic agent to the brain through injection into the blood stream may be impractical since a very high concentration of the drug may be necessary for the drug to cross the blood-brain barrier, and the side effects of the drug on other organs in the body may be toxic. Robert Langer developed a solution to this problem and related ones through the construction of polymeric devices to release drugs at a predetermined rate for extended periods. In this particular disease, the brain tumor is removed and polymeric disks filled with a chemotherapeutic drug are inserted. The polymers have been carefully synthesized to be biocompatible and to break down at a known rate in bodily fluids. As the polymer matrix is dissolved, the drug is released slowly at high local concentrations for many weeks, killing residual cancer cells. Since the polymer disks are in the brain, the drug does not have to diffuse across the blood-brain barrier. Thus, the target, the brain, receives a high dose of drug in the area near the tumor, and the rest of the body experiences only low levels of the drug. This therapy is currently in use and can significantly extend the symptom-free lives of patients. |
|
Langer received the Draper Prize from the National Academy of Engineering in 2002 in recognition of his work on development of modern biomaterials. The polymer used for controlled release of chemotherapeutic agents in the brain is only one example. Other examples include the use of such biomaterials for controlled release of large molecules (proteins or DNA for gene therapy), as scaffolds for tissue engineering where they release growth-promoting signals, and porous aerosol particles for inhalation drug delivery. The controlled-release drug delivery industry is estimated to have revenues of $20 billion a year with excellent prospects for continued growth. This industry is based on combining a knowledge of polymer synthesis, polymer interaction with biological molecules, the kinetics of the reaction of specific chemical bonds in the polymer with water or biological fluids, and the rate of mass transfer of molecules in a polymer matrix and in tissue. |
We will also look to the chemical details of biology for lessons in how to carry out complex and important reactions under mild conditions. Today, the chemical industry produces ammonia from nitrogen through a high-temperature, high-pressure reaction that consumes lots of energy, yet microorganisms are capable of carrying out the same reaction at normal pressures and temperatures within the environment of the cell, using a metalloprotein catalyst called nitrogenase. Structural insights into this and other remarkable metalloprotein catalysts have recently become available. But can we now harness that understanding to develop new methods and new small molecular catalysts that incorporate the key attributes of the natural enzymes? Many of these enzymes, nitrogenase in particular, are capable of activating small, abundant, basically inert molecules through multielectron reactions. A tremendous challenge to the chemist lies in the design and application of catalysts, whether small molecules or materials, that can carry out such multielectron transfers to activate small molecules such as nitrogen, oxygen, and methane. Imitating some aspects of life, biomimetic chemistry is not the only way to invent new things, but it is one of the ways.
We will look to chemical biology for guidance not only in designing new smaller catalysts but also in devising methods to assemble large molecular machines. Replication, transcription, and translation, as well as other critical cellular functions, appear to be carried out through the function of multiprotein/nucleic acid particles. Advances in x-ray crystallography coupled with other imaging methods such as NMR and electron microscopy are now providing our first snapshots of these macromolecular machines, such as the ribosome in which proteins are synthesized in the cell. A spectacular recent advance is the determination of
the three-dimensional chemical structure of the multimolecular photosynthetic reaction center, for which Johann Deisenhofer, Robert Huber, and Hartmut Michel received the Nobel prize in 1988. New advances in imaging will be needed to delineate these and still larger macromolecular assemblies at atomic resolution. These structural pictures provide a critical foundation for understanding how they function. But how do these machines assemble? Are they remarkable examples of spontaneous supramolecular assembly or are they guided in some way in coming together? How do the parts of these assemblies function in concert? Are macromolecular assemblies of such complexity required to carry out these functions? Indeed, can we next begin to design novel macromolecular machines to carry out new, still more complex functions? The construction and assembly of such machines would represent the first step in compartmentalizing chemical reactions. As such, it would represent the very first steps in a tremendous challenge to the chemist and chemical engineer, the design of a synthetic cell.
|
Sequencing the Human Genome The year 2000 marked the completion of the Human Genome Project’s primary goal. Through intensive efforts of both private and public agencies, the sequence for the three billion base pairs that encodes the instructions for being human has now been determined.5 As a result of the Human Genome Project, we have determined the complete chemical structure, nucleotide by nucleotide, of the DNA within each of these chromosomes, the chemical structures that encode our lives. It is an extraordinary accomplishment in chemistry. Completing this sequencing of the human genome could only be accomplished by building upon discoveries in chemistry made over the past 30 years. It was about 20 years ago that W. Gilbert and F. Sanger showed that small segments of DNA could be sequenced directly using chemical methods. About 10 years ago, instrumentation for automated sequencing was engineered. And building upon all the advances in biotechnology of the last decades, from oligonucleotide synthesis to the polymerase chain reaction and shot-gun sequencing, biochemists in the last 2 years have been able to increase the pace of analysis, so that full genomic maps can be deciphered in months. In this post-genomic era, what can we expect? Mapping the human genome brings not only a high-resolution picture of the DNA within our |
|
cells but also the promise of molecular-based diagnosis and treatment of disease. Therapeutics could be tailor-made to take into account our individual genetic propensities. The cost of such pharmaceuticals will surely decline as clinical trials take such genetic information into account. And preventive medicine will certainly flourish as we begin to catalogue and diagnose our individual genetic predispositions to disease. But none of this is likely to happen tomorrow or even next year. There are many more conceptual advances in the chemistry of life that we need to achieve. At the start of the 21st century, we may know the sequence of bases of the human genome, but we can’t yet read this sequence to know what reactions, what chemical functions, actually make us human. |
|
Post-Genomic Therapies We have already had a glimpse into the future with the recent approval of a new approach for the treatment of adult onset, noninsulin dependent diabetes (NIDD). In this disease, also known as Type 2 diabetes, the body becomes increasingly resistant to insulin and loses its ability to control sugar levels. The glitazone family of drugs acts to increase insulin sensitivity and thus increase glycemic control. They do this by acting upstream of a gene! As selective activators (agonists) of a nuclear receptor called PPAR-gamma (found in key tissues like fat and liver) they regulate transcription of the insulin responsive genes involved in the control of glucose production, transport, and utilization. Using the cell’s signaling machinery, the drugs help the body compensate by getting the cells back to where they should be: sensitive to insulin.
|
|
The explosive growth in our understanding of the chemical basis of life and its processes couldn’t come at a better time. The demographic shift of an aging generation of baby boomers will sorely challenge the nation’s resources (both financial and human capital). The need for more effective and cost-efficient therapies will become paramount. As our understanding of the chemistry of the brain grows, intervention will be based on new insights at the molecular level. Selective therapies for problems of memory and cognition, vision and hearing, pain, addiction, and sleep disorders will be designed. The subtle control needed to balance metabolic processes—like weight gain or loss and even aging—is already on the horizon. At the same time the revolution in microprocessors, telecommunications, and materials will enable the production of human “spare parts”: joints and valves, eyes and ears linked to the brain. Even implantable endocrine systems—miniature chemical factories that combine real-time analysis with the synthesis or release of therapeutic agents—are becoming a reality. |
WHY ALL THIS IS IMPORTANT
What are the details of all the chemical transformations that occur in a living cell? How are these details affected by the physical organization of the cell, with various components such as membranes and ribosomes. How do these details differ among cells of different types (liver versus brain, human versus bacterial)? We can also begin to ask what chemical signals direct the development of a single fertilized egg into the different organized tissues in a human being, and how such signals work. What is the chemistry of aging? What special chemistry operates in the brain to store memories? Underlying these functions—functions that make humans what they are—are chemical processes that remain to be discovered and harnessed.
Other opportunities will drive both basic discovery and invention. New and improved drugs will be needed to fight disease and improve quality of life. At the same time, better methods will be needed for delivery of these drugs, and a variety of new medical devices will depend on the work of chemists and chemical engineers. While the engineering of tissue constructs has had commercial success, highly perfused or vascularized tissue remains problematic. Artificial organs, such as an artificial liver, remain objects of intense research. Commercially viable replacement organs are certainly more than a decade away. The primary challenges are understanding the signals that tissues need to control proliferation and differentiation and constructing bioreactors and scaffolds that will provide signals in appropriate sequences.
A closely related challenge is the design of materials that interact with cells or living tissues to promote desired biological responses. Such responses might be cell attachment, cellular differentiation and organization into functional tissue, or promotion of in-growth of bone into an artificial prosthesis such as an artificial hip.
The coupling of the techniques of microfabrication on silicon with cells or biological molecules also offers great promise. Devices at the natural length scales of biological systems will facilitate the use of biosensors that can be implanted. They will also aid the development of models having biological components that can be used to gain predictive insight into in vivo systems. Such nanoscale devices may mimic the biochemical interactions in the body by connecting “tissue” compartments in ways that mimic the body’s circulatory system. A particular model that would be useful would be of the blood-brain barrier, to predict which drugs or chemicals may enter the brain.
While we have very functional processes for manufacture of therapeutics, there are significant challenges left to the process chemical sciences. The United States faces a near-term crisis in the production of therapeutic proteins from mammalian cell culture due to the absence of sufficient facilities. Mammalian cell culture is both expensive and has low yields; it is used to ensure that all of the post-translational protein-processing steps (e.g., glycosylation or specific addition of certain sugar complexes to predetermined sites on the protein) are humanlike. Can we find ways to alter cellular machinery in other, more productive host cells, to produce large amounts of proteins with humanlike post-translational processing? Another challenge comes from the need to respond to bioterrorism (Chapter 11). We need new methods to produce large amounts of protective antibodies or vaccines in a matter of weeks rather than years. In addition, ways are needed to integrate better genomic, proteomic, and advanced computational methods with metabolic engineering to inexpensively produce large amounts of nonprotein products.
This listing of challenges for the future is not exhaustive, but it should provide the reader with a sense of vast possibilities for the interface of the chemical sciences and engineering with biology. These are complex scientific problems requiring multidisciplinary research. Research at the interfaces of many disciplines requires greater understanding of neighboring fields. We are left with a training paradox—we need highly skilled specialists who are also generalists. In addition, we have a funding paradox—success requires support of both fundamental research and cross-disciplinary research. With collaborative efforts these paradoxes will disappear, and we will realize the incredible potential that lies before us. The early part of the 21st century will be known as the Golden Age of the Chemistry of Life.