11
The Urge to Commercialize: Interactions Between Public and Private Research Development
Robert Cook-Deegan1
Almost everything that I am going to present will be blindingly obvious. I see my job as synthesizing some of what has been talked about previously and to look at some overall trends. The reason we are here is because we like what science and technology produce. There has been a lot more spending by governments in research and development (R&D) and even faster growth in R&D spending by private entities in the postwar era.
Why are we doing that? Because we buy the products and services that come at the end of that process and we have been buying a lot of them. R&D has been a source of economic growth. Governments like it because it creates wealth, and people who have more money are happier voters and it feeds back on itself. It is a virtuous cycle. We have a robust system of innovation. We are coming to the end of a decade when it seemed like that growth was never going to end.
I am going to go through some of these overall trends, and then I will look at biomedical research as a particular sector of interest. I will spend most of my time talking about genomics because genomics is a poster child, representing an area where intellectual property and the public domain are intersecting, colliding, and causing conflict constantly, and quite conspicuously. The amount of funding going into biomedical research has increased by three to four orders of magnitude in real dollars since World War II. That is an unbelievable amount of growth in five decades. The scale of effort, the number of people doing it, the amount of money, and the commitment of social resources have all mushroomed in a relatively short period. It has happened in both the public and the private sectors, led by government funding, at least in the case of life sciences, and it has been followed with a time lag of several years by investment in the private sector.
The R&D growth in both public and private sectors has led to conflict. The Human Genome Project is especially good at generating conflict. When it does so, it usually is on the front pages of Time, Newsweek, The New York Times, and The Washington Post. You all have heard the stories I am going to recount. I am just going to try to tease out some of the structures underneath the surface.
What drives the growth of R&D—in science, in academia, in government, and in industry? The practice of generating new knowledge has become more capital intensive. Research costs a lot of money, and it takes a lot of people. The scale and the complexity have increased. We need machines to generate the data. We need computers to keep track of data. This has been increasing on a large scale since World War II in almost every discipline.
In the political arena, drawing lessons from World War II, governments woke up to the fact that universities and academic centers seem to be the focal front end of a big process that generates a lot of dollars at the back end. They realized that universities are valuable resources and began to pay attention to the policies that foster their development. In particular, they thought about their role in economic growth, in addition to the traditional university roles of creating knowledge and disseminating knowledge. At the same time, we have seen intellectual property grow in strength and quantity; this has been happening historically during this postwar period, most prominently in the United States.
I am going to shift for a moment to the life sciences, specifically on drug development and biotechnology, the products and therapeutics where most of the money is made. It is about a $200 billion a year enterprise, much larger than at the end of World War II. The overall policy framework has been a simple one that was crafted in the immediate postwar era: The government funds basic research, which spills over and people make useful things out of the new knowledge and techniques, which are turned into products and services, the so-called pipeline model. That scenario actually is not too different from how many drugs have been discovered.
The government has funded a lot of science. The past five decades have seen spectacular budget growth at the National Institutes of Health (NIH). I do not know what is going to happen after this year, but we are at the end of another doubling. There have been many doublings of the NIH budget since World War II.
We have also encouraged patenting. The Bayh-Dole Act solidified policies that were already falling into place in the 1980s. The federal court system was reorganized, creating a single appeals court for patent cases. The presumption in favor of patents increased. The structure of the judicial system presiding over intellectual property decisions reinforced the technology transfer statutes addressed earlier today. Moreover, there have been many other policies that try to foster—at the state level, at the national level, and in private industry—the private development of these public resources to bolster the public domain in science. What have the results of these policies been?
There are many start-up firms. In biotechnology, I do not think anybody knows how many firms there are, but there are more than 2,000. In 1992 you could not have called any firm in the world a genomics firm. There are now approximately 400 plus genomics firms, at least 300 of which are still operating—in less than a decade, all of these companies were created.
There have been many patents issued to academic institutions, 3,000 of them last year. Private R&D investment in universities has grown, as has licensing income from academic intellectual property. Last year, the most recent survey from the Association of University Technology Managers found that universities received about $1.1 billion. That is about 3 percent of the total that they spend on R&D. The patent logjams and intellectual property strictures on research that we are talking about, however, are some of the unintended consequences of the policies that have been largely successful (see Figure 11-1).

FIGURE 11-1 Unintended consequences.
BIOMEDICAL RESEARCH MODELS
I now want to focus on genomics. My background is in human genetics. I did my first research on the genetics of Alzheimer’s disease. A long lineage of human clinical research goes all the way back to Pasteur and beyond. Many areas of human clinical research, including human genetics, had only glancing acquaintance with the public domain. Data hoarding, keeping things secret, using your data so that you could get the next publication, and making sure that nobody else got access to data were common behaviors. Within science, particularly within human genetics, there were not strong open disclosure norms. There were other communities doing molecular biology at the same time on yeast and Drosophila that had “open-science” norms. Those norms were the ones adopted as the models for the Human Genome Project.
Within biomedical research, different norms for disclosure pervade different fields, so that open science and secretive science are often working in parallel. I merely want to point out that going all the way back to Pasteur, there have been norms of secrecy. Geison has shown that Pasteur was highly secretive, particularly regarding human experimentation (one provision of his will was to keep his laboratory records secret). Given this history, we cannot think of some golden era that we are trying to return to, at least not in human genetics. One of the goals of the Human Genome Project, as articulated by the 1988 National Research Council Report Mapping and Sequencing the Human Genome,2 was to tilt in favor of open science over the more territorial norms of human genetics and clinical research.
Pharmaceutical development is one of the most patent-dependent sectors in the whole economy. There was a tremendous amount of government and nonprofit funding flowing into genetics and genomics for the better part of a decade, before the private genomics effort began around 1992-1993. There is also an intricate mutualism between the public domain and the private domain in genomics that creates databases, products, and sequence information. Genomics attracted many players, including national and state governments, as well as private firms, because genomics was “hot.” It was in the news and everybody wanted a piece of the action. But action toward what end? The policies that we are talking about focus on information flow.
Three different models drive commercial genomics, which were created in that first wave of genomic startups from 1992 to 1994. One business plan is represented by Human Genome Sciences (HGS). It started from owning the intellectual property from a nonprofit organization created in 1992 called the Institute for Genomic Research. The Institute and HGS were initially looking for human genes by picking out the parts of the genome that were known to code for proteins and then looking for those proteins that were most likely to be thrown outside the cell, to span cell surface membranes, to bind DNA, or to serve other known functions. HGS would look for DNA sequences corresponding to proteins that might be valuable to pharmaceutical companies or to themselves to develop into pharmaceuticals, and they focused on characterizing the whole gene, sequencing it, and then doing some biology to figure out what it did. The HGS strategy was very focused on intellectual property. There was a patent lawyer, or somebody with legal training in patents, associated with every project team.
HGS was very careful about how they set up walls of nondisclosure around their arrangements. There was some outlicensing of their technologies to major firms so that they could get money, but there was not much in the way of publications. The main output and the main contribution to the public domain happens when HGS gets a patent because then it is published. Then the sequence information that was part of their patent application is published and it becomes part of the public domain, except that it is constrained by the patent rights that are associated with that work.
Incyte turned seriously to DNA sequencing of genes about a year before HGS. It had a somewhat similar scientific strategy, but a different business plan. Incyte was also looking for the juicy bits of the genome to sequence and characterize genes. Incyte had a somewhat different business model in that it was working with multiple, large pharmaceutical players, and it did not appear to have focused on exactly the same kinds of sequences as HGS. Incyte was creating a database that high-paying customers could access. Through Incyte, large pharmaceutical companies could avoid having to do all the sequencing themselves. They could license access to DNA sequence information on genes from Incyte. Incyte’s business plan has changed several times since the early
years (as has the company’s full name), but gene sequencing was the original core idea. Incyte pursued a model that allowed multiple players to get access to its data. It also had looser boundaries and there was a little bit more leakage into the public domain, and a few more publications.
After these two companies had been doing genomics for 5 years, Celera Genomics Corporation came along in 1998 and proposed to sequence the whole human genome. Celera was going to do it in a slightly different way from the way the public-domain project was sequencing the human genome. Celera’s initial goal was to create a database of genomic sequence (that is, of the genome in its native state on the chromosomes before being edited into shorter and more compact known genes). Celera needed powerful informatics, and indeed spent more on computers and programming than on DNA sequencing and laboratory biology. Because of a natural asymmetry, Celera would always have more data than was in the public domain—that is, Celera could draw on the public domain and at the same time create its own proprietary sequencing data. Celera would always have one leg up on the public genome project and they could sell a service, which was access to their data. They were doing that nonexclusively, but even more nonexclusively than Incyte, and they had more academic collaborators. Celera was selling licenses to get access to their data using discriminate pricing—a lower pricing level to academic institutions, to the Howard Hughes Medical Institute, and other academic research and higher prices (and more intensive services) for pharmaceutical companies.
Celera’s intellectual property strategy was somewhat different from Incyte and HGS. Celera was filing provisional patent applications that could be converted to patent applications, and a few dozen had ripened into patents (52 as of December 2002 compared with 711 for Incyte and 277 for HGS). The interesting thing about the Celera model is that publication in scientific journals was an inherent part of the business plan. Celera planned to publish in scientific journals, and the information they publish or that they post on their Web site is available for other people to use. I do not think we could use the word “public domain” for these data and information because there are restrictions on their use.
These three companies have three different models. If you think of a continuum with the public domain on one end and the private domain on the other, these companies are at intermediate points along a continuum of contributing data into sources where scientists can use them.
There was a strong ideology that grew out of the nematode and the yeast research communities that access to the data should be free and rapid. The sequencing centers in the Human Genome Project are getting paid by the government or nonprofit organizations (e.g., the Wellcome Trust) and are making the data available so everybody can get access to them. The scientific community wanted policies to ensure that the high-throughput sequencing centers did not get unfair advantage because of sole access. My reading is that the rapid disclosure policy was not so much a reaction against commercial practices but rather a concern about hoarding data. The idea was that the yeast and nematode genetics science communities were healthier because of the way they shared data. They are more spontaneous. They are more creative and do better science, in part because they share their data at an earlier stage. Out of that movement grew some very concrete policies, including the so-called Bermuda Rules or Bermuda Principles, which grew out of a meeting that the Wellcome Trust and other funders held in Bermuda. The policy was an agreement to dump data quickly into public sequence databases. As such, the high throughput sequencing centers that were funded as part of big human genome projects agreed to provide their data very quickly, usually on a 24-hour basis, a remarkable forcing out of information into the public domain. That is the open extreme of public-domain policy making, defining the opposite pole from HGS.
However, within the academic sector, many intermediate points on the continuum are represented. Many university laboratories have “reasonable delays” for publication of some research. There are thousands and thousands of labs that do sequencing. Most of them do not behave according to the Bermuda Principles. They release data in “publishable units” after a gene has been fully sequenced and partially characterized and once the sequence data are verified. Researchers in academic genetics, that is, who identify themselves as geneticists, report withholding data to honor an agreement with a commercial sponsor or to protect the commercial value of the data. However, I will point out that the big reason for nonsharing or withholding of data is the effort required. So in many university molecular biology laboratories, data and materials may be unpublished for protracted periods.
Yet in private industry, there may be rapid open disclosure of data. Here is the topsy-turvy part of the Human Genome Project. Pharmaceuticals are very patent dependent; it is famous in business schools for being an information-intensive, secrecy-intensive business. However, in 1994, Merck, a private firm, funded the human expressed sequence
tag and cDNA sequencing effort at Washington University, the results of which were added to the public domain. Merck did not get to see the data before anybody else. Why did a private firm pay for information to be generated and dumped into the public domain? I believe it is because of what Incyte, HGS, and other genomics startups were doing. Merck pursued one scientific strategy to counter the appropriation of DNA sequence data by funding research to put data into the public domain to defeat the strategies being pursued by some start-up genomics firms.
I have already talked about the Bermuda Principles. They were adopted by the major genome sequencing centers. The SNP Consortium is another interesting model. A “SNP” is a single nucleotide polymorphism. There is a difference in the A, G, T, or C at one place in the genome, which differs among individuals. If you can find a sequence difference, it is a SNP; polymorphism merely means “difference” to geneticists. SNPs are useful as markers on the chromosomes of humans or any other organism. SNPs are particularly valuable for looking for genes of unknown function and location, or for studying the whole genome at once. Some companies were established to find and patent SNPs. As a result, some academic institutions and 13 private firms formed a consortium to make sure that this stays in the public domain. However, the SNP Consortium did not just dump the data. They filed patent applications and then characterized the SNP markers enough so that they could be sure that nobody else could patent them.3 At that point, they would abandon the patent. It is a very sophisticated intellectual property strategy that in the end was intended to bolster the public domain. It requires coordination, lots of paperwork, and it costs money to file and process applications, but it appears to be an effective defensive patenting strategy.
The publicly funded Mammalian Gene Collection and its parallel program, the Cancer Genome Anatomy Program, pursue policies to promote rapid data disclosure into the public domain. Under Mammalian Gene Collection government contracts, groups do the same thing that Incyte and HGS were doing, which was to sequence identified genes—the juicy bits of the genome that are translated into proteins. In this case, NIH had to go to the Department of Commerce to declare “exceptional circumstances” under the Bayh-Dole Act. As a condition under those contracts, the government gets to keep all the patent rights. Because the government is not filing patent applications, it is basically a de facto nonpatenting strategy. That is the only case that I know of where the exceptional circumstance clause of the Bayh-Dole Act has been invoked, although I know others at NIH are discussing other possible uses.
Remember that when the genome project started out, it was supposed to be a public-domain infrastructure project so we could all do human genetics faster and at less cost. Those of us who thought about the Human Genome Project in the early years were not thinking primarily of commercial potential, but look at what happened to the funding streams in the year 2000. It looks like over $1.5 billion of public and nonprofit funding went into genomics in the year 2000 (see Figure 11-2). The aggregate R&D spending of “genomics” firms is in the
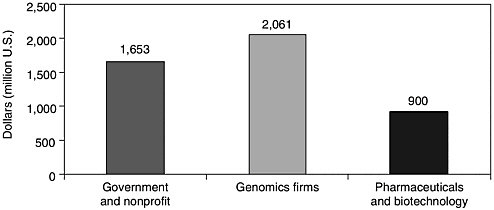
FIGURE 11-2 Private and nonprofit genomics funding, 2000. Source: World Survey of Funding for Genomics Research Stanford in Washington Program at http://www.stanford.edu/class/siw198q/websites/genomics/.
|
3 |
For more information, see Chapter 28 of these Proceedings, “The Single Nucleotide Polymorphism Consortium,” by Michael Morgan. |
neighborhood of $2 billion (genomics firms devote some substantial part of their R&D to genomics). Major pharmaceutical firms spent another $800 million to $1 billion on genomics in 2000, based on an estimate of 3 to 5 percent of R&D of members of the Pharmaceutical Research and Manufacturers’ Association. That does not include the privately held genomics firms that tend to be smaller but are much more numerous. These figures are quite uncertain, but if they can be used as a rough indicator, about a third of genomics funding comes from government and nonprofit organizations, and two-thirds of the funding, at least in 2000, was spent in private R&D.
Figure 11-3 shows government and nonprofit genomics research funding for 2000 by country. The United States, not surprisingly, is number one; the genome project in many ways originated here, even if the science did not. If you normalize for gross domestic product, you see that many countries are investing more in genomics as a fraction of their R&D and as a fraction of their economy than the United States. Estonia, the United Kingdom, Sweden, the Netherlands, Japan, and Germany are all higher than the United States. This happened very fast. In 1994 there were eight publicly traded genomics companies; in 2000 there were 73 publicly traded firms. In 2000, commercial genomics grew to about $94 billion or $96 billion capitalization (before declining precipitously in 2001 and 2002).
What was going on under the surface? Let me return to this private-public mutualism. The main target that everybody was aiming at was that $200 billion market for therapeutic pharmaceuticals, expected to grow much larger in future years. Genomics was thought of as a way to develop those products faster at the front end of the discovery process. Pharmaceutical firms were interested but they came late to the game.
How do I know that? Well, let us look at the patent holdings. Figure 11-4 is only a thousand of the DNA-based patents that were issued from 1980 to 1999. Initially, we read every patent and coded them to be able to get these data, and then we went back to figure out how many patents were owned by whom through 1993. We then used the same patent search algorithm to identify DNA-based U.S. patents and augmented the database through 1999, the basis for the bar charts on which institutions own DNA-based U.S. patents (see Figure 11-4). The U.S. government is the number one patent holder followed by the University of California. Incyte, a genomics company, was number three. Chiron, which is a first-generation biotech firm, was number four, and most of their patents actually
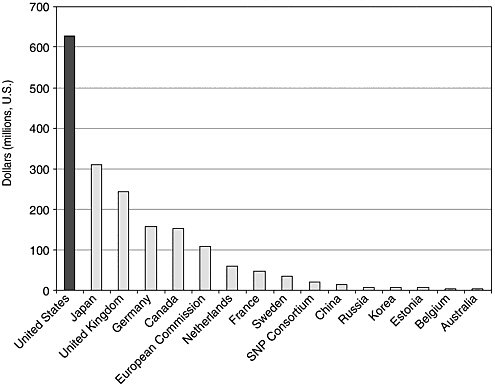
FIGURE 11-3 Government and nonprofit genomics research funding, 2000.
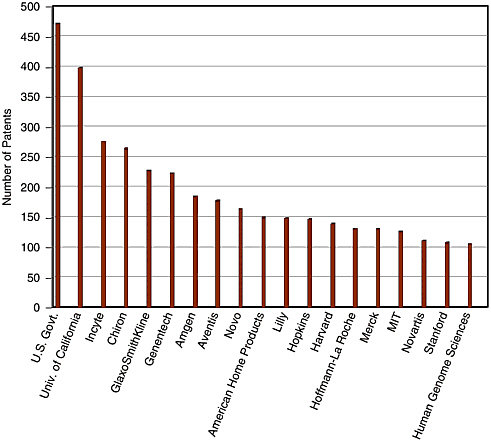
FIGURE 11-4 Number of patents in the DNA patent database, 1980-1999. NOTE: Data through the end of December 1999.
came from a firm called Cetus, which they acquired. You have to get to number five before you get to a company that is in the business of creating the end products that everybody is aiming at: GlaxoSmithKline. This represents five different kinds of players that own the intellectual property here.
There were no DNA-based patents in 1970. There were some DNA and RNA-based patents issued before 1980, but not many. A DNA sequencing method was patented in 1973; I do not think anybody uses it, but it is patented and it is in the DNA Patent Database. Starting in the mid-1990s, there was the beginning of an exponential rise. It kept that way through 1996 or 1997. The growth dipped in 1999, which may be a policy dip, because the patent office that year began to change the rules for DNA sequence patents. They raised the threshold basically, changing to a higher utility standard for examining DNA sequence patents—requiring applicants to show a “credible, substantial, and specific” utility in the patent. The U.S. Patent and Trademark Office also demanded a higher standard of written description of the invention in gene-based patents. The growth increases again in 2001 and it looks like we will be below the 2001 number for 2002, although there are a few months of patents we have not examined. The years of exponential growth may be over, but the bottom-line message here is there are 25,000 pieces of intellectual property that have already been created.
A pattern of ownership can also be observed. Of those patents that were issued between 1980 and 1993 that we read through and coded, companies only own about half of these patents. If you add up the others, which are mainly academic research institutions, almost half of the patents are owned by “public” organizations such as universities, nonprofit research centers, and government. That is a very unusual ownership pattern. Overall, academic institutions own only 3 percent of U.S. patents. Figure 11-5 illustrates ownership of 1,078 DNA-based patents from 1980 through 1993.
What is going on here? Molecular genetics appears to be a field in which private-sector actors step in to bolster the public domain. The public domain or the public funders create intellectual property subject to the Bayh-Dole
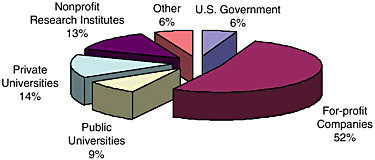
FIGURE 11-5 Ownership of DNA patents, 1980-1999.
Act, which gets protected by intellectual property rights. Private firms draw on public science and also pursue private R&D, which produces more intellectual property owned by private firms.
A couple of generalizations do flow from that. One interpretation is that, the more money you make, the more you are going to plow into R&D. That is the argument that is made by the pharmaceutical and the biotech firms, and it is probably true. If firms make more money, they are going to spend more on R&D, particularly if R&D is the way they believe they made their initial profit. As a result, there will be more innovation, there will be more of the products and services that we like, and, of course, we are going to have to pay more for them. We have been doing that for two decades. One thing to consider is whether double-digit rates of growth are sustainable. We may be beginning to encounter resistance to growth in drug and device expenditures in the early part of the twenty-first century. It is very clear that at least for these genomic startups, private firms believed that their patent portfolio mattered. Their intellectual property mattered and that partially drove this high level of private investment in genomics and in biotechnology more generally.
One of the really interesting things about genomics as a case study is that, in fact, we have a scenario that feels like a race between the public sector and the private sector. I am not sure competition is the right word because you have one group of people that are dumping data into the public domain and another group that are developing data to make a profit. But they are doing the same things in their labs. In the private sector, they have done it in a very capital-intensive way that tends to be fairly centralized.
Public and private sectors are pursuing similar lines of research, and we have a natural experiment that has been going on now for almost a decade, the outcome of which we do not really know. We can say that the academic sector has been a more important part of this story than it has been in most other technologies, such as informatics and computing, although universities were important there too. But it is not possible to compare whether more public good will come from the HGS and Incyte’s private sequencing, Celera’s quasi-public sequencing, or the strong public-domain policies under the Human Genome Project and Mammalian Gene Collection. We may never know, as products, services, and discoveries are apt to draw from many streams.
The big fight over the publication in February of the 2001 sequence data is how much is going to be put into the public domain where everybody can use it without restriction, how much of it will be publicly accessible with some restrictions, and how much of it is kept behind closed doors. That is not a resolved debate, but it is very rich. In 15 years we may be able to make more educated guesses about what our policy should have been over the past four or five years.
We have races for money, but we also have races for credit, and they are inextricably intertwined. It is very clear that part of Celera’s business strategy was to be well known and famous, as well as to sell database subscriptions. Therefore, is not just about money and it is not just about credit, it is about both. That is true on both sides of the academic and industrial divide.








