6
Opportunities for Commercial Exploitation of Networked Science and Technology Public-Domain Information Resources
Rudolph Potenzone
I would like to address, the importance of having access to data, both data that are proprietary and data that are in the public domain. I certainly agree with Paul David’s comments that it is a mixture of relationships that is vital to the good of increasing our scientific knowledge.1
Why are data so important to us, and why do we worry so much about them? Clearly, when you start a research program, no matter what the topic or field, you have to know what information is available on the subject. You have to know all the data that have been accumulated on a particular subject to do good research. You have to be able to get your hands on them, access them, and actually use them. Ultimately, the process of research is really about generating new data, and so as you study the existing information, you build your own concepts and ideas and generate new data. The research process is one of data generation.
Hopefully, if the research is successful, and you are able actually to accomplish something, the data turn into some new knowledge. Depending on your particular area or affiliation, you may want to invent the greatest new thing since sliced bread, maybe a new pharmaceutical or you have discovered some whole new unifying theory of whatever. Yet it is all based on the ability to get access to the right information at the right time.
Information resources are vitally important to every project, and it does not matter as you work on your project where relevant data come from. In fact, it is hard to predict where the best data will be found for any particular project. Even at the time the new data are generated, it is not certain what those data will be used for. How often do you find a bit of information or data that are in the most obscure place, not related to the topic you are studying, but in fact are relevant to the new project that you are working on? The availability of data is ultimately important to the ability to generate good science.
What happens to these good data once they are generated? The reality is that most data never enter a database that is available publicly, whether it is a fee-based database or a free database. Data are often locked in papers, reports, or lab notebooks. Maybe they get published and are available in a peer-reviewed journal or are archived electronically. However, by and large, those data are not accessible and are not able to be found because they actually have never been brought into a database. In particular, if the data have a particular commercial interest, and the pharmaceutical industry has been quite aggressive about this, they may find their way into a commercial
|
1 |
See Chapter 4 of these Proceedings, “The Economic Logic of ‘Open Science’ and the Balance between Private Property Rights and the Public Domain in Scientific Data and Information: A Primer,” by Paul David. |
database and become quite accessible. Otherwise, it really depends on someone’s personal initiative either to get funding to create a database through one of the government sources of funding or finding some support so that the information can be put into a form that is actually accessible.
I will describe two different scenarios for the pharmaceutical industry. One is chemical information (which is where I have spent much of my career), compared with what I call the bioinformatics movement; they differ in how the data have been handled and collected. Historically, chemical informatics has been a very commercialized activity. Three of the larger data repositories—Beilstein, Derwent, and Chemical Abstracts Service (CAS)—are organizations that all started out building printed repositories and then ultimately turned into electronic sources. They are highly commercialized, profitable activities that have served a truly critical role in the preservation and availability of chemical information.
Beilstein Institute, founded in 1881, first published its handbook with 1,500 compounds. The final version was printed in 1998, with the oldest references going back to 1771. Interestingly, this was converted in electronic form with some assistance by the German government and today holds 9 million compounds, a lot of information and data, and is distributed on a commercial basis by a Reed Elsevier company.
Derwent, which is today a Thompson company, was founded from ideas initiating from Monty Hyams in 1948. He was trying to make some sense out of the patent literature and began writing some simple abstracts about what was being published at the time. This information became useful. People started getting more interested, and it turned into a commercial operation. Today, Derwent’s world patent index is global in scope, covering 40 different patent-issuing authorities and details in over 8 million separate inventions. It is a very large repository. If you work in the area of pharmaceutical research and development, you have to go to the Derwent database to understand what the patents are about.
CAS has a similar history. They were founded in 1907, with the goal of monitoring and abstracting the world’s chemical-related literature. Today, with the Internet and all of the information that we have to deal with, it is mind-boggling to think that in 1907 this operation was formed because there was too much information to handle. CAS is a subsidiary of the American Chemical Society. There are 20 million organic and inorganic compounds registered at CAS; 21 million biological sequences; and almost 42 million separate and unique chemical entities registered in their system, all of them complete with names and references to the published literature, allowing scientists to find more information about these items. It is an incredible amount of information.
My observations are that chemical informatics has been quite commercialized and brings in quite a bit of money. In the area of pharmaceuticals, this has all been organized and put out there to be used because of its high commercial interest. These three companies and others look at scientific journals, books, patents, conferences, and dissertations; they do the work, and they extract a significant fee for it. The data are organized and then made available to the community, but at a price. The reason these data are not free is not because the underlying information is not free, because in many cases it is. But the fact that these companies have organized it and brought this together in a searchable (i.e., more useful) fashion gives these databases a very high value. This makes life much easier in terms of the chemical information community.
There is clearly value in that much of the original research was publicly funded research. There has also been a significant cost of creating these data sources. Although the pricing is fairly significant for these groups, they certainly have provided a service to the community. Frankly, whether these operations could have continued to exist in an electronic form without the current funding support is doubtful.
There are certainly hundreds, maybe thousands, of other databases and data repositories in chemistry that do not get picked up by these services. Are these less important to us as a scientific society? There are certainly cost barriers that prevent all the good data from being collected and organized in a reasonable fashion. The public funding has not been available to make these data generally available. I think that the funding tends to go toward collecting more data, and yet the funding for making the repositories of these data and making them available has not been there. My personal opinion is that funding authorities should consider the utility of the resulting data they pay to have generated from the start of these projects.
An interesting contrast to this illustration is in the bioinformatics arena, which in some ways is the antithesis of the chemical data franchise. Here, largely publicly funded projects have been formed by different highly motivated groups to put together what have become literally hundreds of sequence databases. A few of these are
being commercialized, and so there are some annotation systems in place, and there are a number of companies who provide commercial databases. Yet the vast majority are still collected and made available for free, and with little support.
Figure 6-1 provides an illustration. If you look at the increase in information and amount of the sequencing speed, coupled to how much information is out there in terms of known sequences, and the growing complexity of information as we get into systems biology and expression information, there is absolutely an astounding amount of information that is suddenly becoming available in this arena.
From LION Biosciences’ perspective, we have been the beneficiary of some of the public work. The Structure Retrieval System has been in the community for over 10 years, largely driven by scientists in terms of its capability, with over 500 parsers to search different kinds of databases. It was developed at the European Molecular Biology Laboratory (EMBL). This has now been turned over to LION Bioscience to commercialize and to keep the product growing and maturing. In terms of differential pricing, we continue to make this freely available to academic institutions, but are charging a fair price to the industrial community. We have merged this with our new technology called Discovery Center.
There are over 800 data sources that are relevant in bioinformatics research, which interrelate to each other. Examples include GenBank, SWISS-PROT, EMBL, and all the various pieces. Even if we believe that the cost of disseminating information on the Internet is free, which it really is not, the working scientist has to navigate through this network of data. He or she must sort through these many sources to find out the particular kinds of information they are looking for. This complexity suggests that they are most likely missing something in terms of the kinds of capabilities that we are making available through the proliferation of databases.
These databases are the fuel for the research projects in genomics and proteomics. These data sources came from the broader community, but they lack the financial incentives because they are all free to motivate the commercial services in terms of bringing these together. The users are individual scientists, who have to look at the local unpublished data that they are generating in their laboratory or calculations that they are doing. They have to
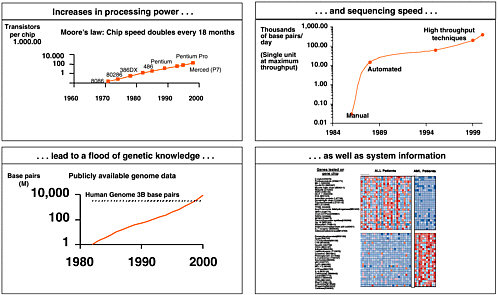
FIGURE 6-1 Genomics generates a flood of information. Sources: Genbank; Sequenom; 3700.com; Applied Biosystems; Human Genome Project; Food and Agriculture Organization of the United Nations; Database on Genome Size; and Intel Corp.
look at the other data around the company or research institution. These data could be at other sites around the world. They have to look at the Internet and all the various databases that are out on the Internet. They have to look at commercial databases, where they are loaded internally or externally. These researchers need to be able to have all of this synthesized for them in some fashion so that it ultimately facilitates their work, which is the research and creation of the new knowledge that we all like to talk about.
Having access to all these data is essential, whether they come from one’s own lab or from some small college. If those particular data are relevant to the project you are working on, it is absolutely critical that all these data continue to be collected and made available in some reasonable form. Commercial databases provide an essential part of the information chain that we have to consider. However, noncommercial sources of data are also vital for the scientific community. These often fill huge gaps that, for whatever the reason in the commercial sector, have not been funded and have not been supported and actually fill out what I like to call our “data portfolio.”
It is the integration of all this information that ultimately will enable us to continue to assist the working scientific community, to push back the frontiers of science, and to expand human knowledge into the future.




