7
GIScience, Remote Sensing, and Epidemiology: Essential Tools for Collaboration
Geospatial relationships lie at the core of many public health issues, and the integration of remote sensing data, epidemiology, and geographic analysis of disease presents a rich opportunity for collaborative activity at the interface of earth science and public health. Hypotheses generated by the analysis of such geospatial relationships can be tested and refined by analytical and experimental research as the basis for identifying causal relationships. The tools and methods that facilitate analysis include conventional epidemiology, with its subspecialties of genetic, occupational, and environmental epidemiology, as well as remote sensing, Geographic Information Science (GIScience1), and the broad field of geospatial analysis that includes spatial statistics and spatial modeling.
GEOSPATIAL ANALYSIS AND EPIDEMIOLOGY
Geographic Information Systems (GISs) used in an epidemiological context are “a simple extension of statistical analyses that join epidemiological, sociological, clinical, and economic data with references to space. A GIS system does not create data but merely relates data using a system of references that describe spatial relationships” (Ricketts, 2003, p. 3).
GIScience is the science of collecting, analyzing, and theorizing about geographic information through GISs and geospatial analysis.
There has been increased attention in recent years to the public health applications of GIS, GIScience, and geospatial analysis (e.g., Melnick, 2001; Cromley and McLafferty, 2002; Cromley, 2003). One of the most useful applications of GIS in the public health arena is as a component of exposure and dose assessment. Ideally, exposure to a potential toxin, environmental contaminant, or pollutant would be measured directly at the individual level through monitors that the individual would carry or wear, allowing direct measurement of exposure and subsequent calculation of exposure-response curves. However, such studies are extremely costly, inconvenient, and infrequently carried out for large populations. Instead, exposures may be estimated based on models, and since exposure frequently varies spatially and temporally, a combination of GIS and spatial/temporal models has become an indispensable component of exposure assessment (Nuckols et al., 2004).
A GIS involves the merging of spatially based data—coordinates corresponding to latitude and longitude—with a graphical user interface (GUI). GISs that use data from remote sensing instruments such as aerial photographs and satellite images have become indispensable tools in the development of causal models linking environmental factors and both infectious and noninfectious disease. The spatially referenced database in an epidemiological context usually consists of geocoded (i.e., geo-spatially located) health information, such as the residential locations of people who have contracted a specific cancer, the location of traffic fatalities, or the location of incident cases of myocardial infarction. These data are then superimposed on other data layers, usually geocoded to the same unit as the health data.
Unfortunately, the power of GIS is not always realized in public health applications, or it is misused, because of a lack of understanding of the underlying geographic principles. Just as a person who learns to use statistical software (e.g., STATA, SAS, or SPSS) does not necessarily understand statistics, learning to use GIS software (e.g., ArcGIS, ArcView, MapInfo) does not necessarily ensure an understanding of the underlying principles of geospatial analysis.
CONCEPTS AND COMPONENTS OF GEOSPATIAL ANALYSIS
A GIS is a powerful tool for the analysis of relationships, including causal relationships, between a broad range of measurable variables from the natural sciences—climatic and weather conditions, surface water characteristics, vegetation and land cover, soil geochemistry, and many others—and public health. It is thus a tool and a set of concepts that bring the
earth sciences and epidemiology/environmental health into a cause-and-effect relationship with one another. The spatial distribution of Lyme disease can be modeled accurately using GIS and remote sensing at a range of scales to model tick dispersion with reference to a series of environmental variables (e.g., Cortinas et al., 2002; Guerra et al., 2002). The same is true of modeling the effects of climate change on disease distribution, although there is some debate about the accuracy of such models (e.g., Hay et al., 2002; Patz et al., 2002; Tanser et al., 2003; Pascual et al., 2006). Geospatial analysis, or more simply “spatial analysis,” uses mathematics and statistics to analyze data patterns that underlie GIS. Many spatial measures and spatial models are available to help summarize and understand complex spatial distributions, including central tendency, dispersion, and clustering (Cromley and McLafferty, 2002; Rushton, 2003).
Remote Sensing
Remote sensing encompasses the full array of technologies for data collection using aircraft or satellites and includes visible wavelength data as well as a broad range of other types of sensors. It is particularly useful for data describing land use, soil, and hydrological features. Satellite imagery is available over an increasing number of wavelengths and at increasing levels of resolution. Remote sensing, coupled with GIS, has been used widely to describe the environmental conditions associated with disease and to model the occurrence of disease, particularly infectious diseases that are sensitive to environmental conditions such as vectorborne and waterborne diseases.
Data Layers
Data layers are a basic element of GIS. A layer of population data may be superimposed on geological data for determining, for example, whether there is a relationship between bedrock type and population characteristics. Or earthquake vulnerability coefficients may be overlaid on layers showing the distribution of elderly or handicapped people for scenario planning for disaster response. Similarly, maps of land use may be overlaid on digital terrain maps in coastal areas, for example, to aid in efforts to mitigate the salinization problems that have been experienced in Sri Lanka and Malaysia following the flooding of rice paddies by the December 2004 tsunami. Njemanze et al. (1999) used a series of “probability layers” to assess the risk of diarrheal disease from water in rural Nigeria (see Box 7.1). The aggregate risk is a product of geological, hydrological, population, and pollutant characteristics, all of which vary spatially.
|
BOX 7.1 GIS Data Layers and Diarrheal Disease in Nigeria Diarrheal diseases are a major cause of mortality and morbidity in developing countries. The spatial distribution of severe diarrhea can be predicted, in part, as a function of the spatial distributions of geological features, population density, and environmental pollution (see Figure 7.1). Population density is important because, other parameters being equal, higher population density tends to increase the rapidity of the spread of disease and also causes an increased number of people to be infected (Halloran, 2001). 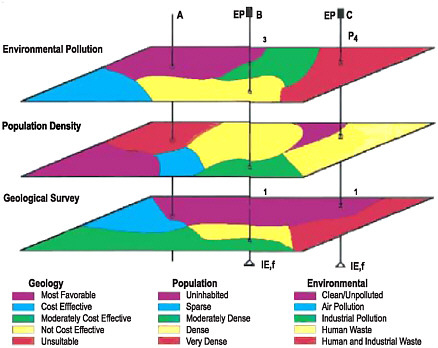 FIGURE 7.1 An example of GIS data layers showing environmental pollution and population density data superimposed on geological features to provide information for understanding the distribution of diarrheal disease in Nigeria. SOURCE: Njemanze et al. (1999). |
Spatial Epidemiology
Any spatially distributed data can be analyzed using spatial statistics, and “spatial epidemiology” has developed as a subfield of epidemiology. Spatial approaches to understanding disease are now feasible because “the availability of geographically indexed health and population data, and advances in computing, geographic information systems, and statistical methodology, have enabled the realistic investigation of spatial variation in disease risk, particularly at the small-area level. Spatial epidemiology is concerned both with describing and with understanding such variations” (Elliott et al., 2000, p. 3). These authors suggest that there are four types of largely statistical and mathematical studies that fall under the rubric of spatial epidemiology—disease mapping, spatial correlation studies, risk assessment relative to point and line sources, and disease cluster detection. In addition, causal modeling could be added to this list.
TYPES AND AVAILABILITY OF EPIDEMIOLOGICAL DATA
Research at the interface of public health and the earth sciences is only as good as the data used to integrate the two. A wide range of geographical and geological data types, particularly remotely sensed data from earth observation satellites (e.g., Guptill and Moore, 2005), are readily available. Such data are, by definition, spatially distributed, and these data are geocoded to enable spatial modeling, geospatial analysis, and the use of GIS. The same cannot be said of most readily available epidemiological data.
The use of spatial techniques, including GIS and spatial analysis, requires that health data be available with their spatial coordinates. Although health data could be geocoded using either Universal Transverse Mercator spatial coordinates or simply the patient’s address, this has implications for the maintenance of privacy of an individual’s health status. In addition, a disease with a long latency or highly specific spatial data invites spatial artifacts (e.g., associating a disease with a residential location would be misleading for a work-related illness). In the reasonable absence of such specific information, data could be made available at the census block group or census tract levels. In reality, most health data—if available at all by location—are usually released by zip code (e.g., CHARS data for Washington State, which include detailed diagnostic and procedural information for each patient discharged from a hospital in the state) or by county (e.g., HIV/AIDS data from the Centers for Disease Control and Prevention). The CDC is very concerned with confidentiality, and the
concern is that if data are released at a finer scale it may be possible to identify individual patients.
Restricted access to individual health data necessarily makes detailed analyses of spatial patterns of disease more challenging, but it is an almost unavoidable consequence of privacy concerns. This restriction can be addressed in different ways. Seiler et al. (1999) evaluated the opportunity for septic contamination of groundwater by pharmaceuticals by using the specific spatial locations of groundwater wells and linking specific health conditions to individuals through their prescriptions. These authors were able to maintain individuals’ privacy by simply refraining from publishing well location data. In other cases, the spatial location (i.e., home address) of an individual may be known by the treating physician or responsible health official, but they would be prohibited from releasing the information (although curiously enough, addresses and even dispatch type for public safety responses involving ambulance or law enforcement are often published in community newspapers). Undoubtedly, making spatial attributes of epidemiological data available for research at appropriate scales and with patient privacy safeguards will continue to pose a challenge. One solution may involve the definition of a new data block of sufficiently small geographic size to be able to associate disease with geological phenomena while providing a sufficiently large error ring around an individual’s residence.
Federal Health Datasets
Federal agencies are increasingly using GISs at the interface of the earth sciences and public health. Examples include the Agency for Toxic Substances and Disease Registry (ATSDR), which was an early adopter of GIS (Cromley, 2003), and the Environmental Protection Agency(EPA) with its Toxics Release Inventory2 (TRI). Although the TRI is not linked to disease data, there is potential to link to cancer registry data, asthma incidence and prevalence data, and other disease data that are spatially distributed. GIS has also been widely used for describing the distribution of natural hazards, for infectious disease modeling and outbreak investigations, for the detection of communicable disease clusters, and—with the recent concern about biowarfare—in new syndromic surveillance systems.3 Standard datasets collected by the National Center for Health Sta-
|
2 |
|
|
3 |
For example, see http://www.syndromic.org/index.php. |
tistics (NCHS) include the National Health Interview Survey4 (NHIS), the Behavioral Risk Factor Surveillance System,5 the National Health and Nutrition Examination Surveys6 (NHANES I, II, etc.), and the National Ambulatory Care Survey.7 In general, these datasets are available with very poor spatial resolution, although under certain stringent conditions these data may be provided by the NCHS Research Data Center at the county level or, occasionally, the individual level.
Ver Ploeg and Perrin (2004) present a tabulation of available health survey data, listing health outcome data with particular application to social disparities in health but also including most data that are available. For example, the NHANES datasets, representing multiple cross-sectional household surveys, have yielded a great deal of valuable nutritional, cardiovascular, dental, and other data. However, these studies are not available with any geographic specificity and are thus of limited use for understanding any earth science relationship with public health issues. Although NHANES could potentially address the question of whether there is a link between water hardness and cardiovascular disease, this is not possible in the absence of geographically specific data. Another example is the NHIS, which surveys the population comprehensively for major self-reported health conditions. Although the relationship between elevation and hypertension is a potentially interesting question, this dataset cannot be used to address the question because the data are not geographically specific.
A final example is the Adult Blood Lead Epidemiology and Surveillance Program8 (ABLES), administered by the CDC and the National Institute for Occupational Safety and Health, which measures lead concentrations to estimate risk. Research is currently being carried out to determine the relationship between soil lead levels and lead levels in individuals. However, because the ABLES system does not record geographic data, it cannot be used to address this important research objective.
These surveys and datasets, some of which have large sample sizes and are publicly available, contain extremely valuable data describing health status and diagnoses, health behaviors, diet, risk factors, and other information. However, they are released using the Census Bureau’s regional designations (e.g., Northeast, West, South, Midwest), which are
|
BOX 7.2 Availability of Cancer Data for Spatial Epidemiology There are a number of problems associated with attempts to establish causality between environmental exposure and cancer data. The National Cancer Atlas includes only mortality data, rather than incidence data, and these data are available only over long time periods and at large units of aggregation. Because environmental exposure to carcinogens occurs at the local level, the National Cancer Atlas is of little use for linking cancer mortality to environmental parameters. The latency period between exposure and detection of the cancer also presents problems, as available data do not permit the reconstruction of life histories and migration histories to trace where exposures may have occurred decades earlier. The “cancer clusters”reported to local and state health departments, the CDC, and the press are often questionable because of the long exposure and latency periods and the absence of life history and migration data. Another source of cancer data is cancer registries, which contain datasets of individuals in each state together with tissue diagnoses of each histological type of cancer. Incident cases are reported to cancer registries by hospital pathologists based on their tissue diagnoses, so the registries represent the highest degree of diagnostic accuracy that is available. To maintain anonymity, cancer registry data are usually released only at the county level, although it is sometimes possible to obtain data at a more local scale by cancer registry staff or established cancer researchers under strict controls. Gaining access to cancer data usually means sacrificing geographic specificity, and it may also involve including staff members from the cancer registry as coinvestigators. It may then be possible to use geocoded data, at least to the city block, with appropriate safeguards to ensure patient anonymity. |
geographically meaningless (e.g., Oklahoma, Indiana, and South Dakota are all included in the “Midwest” despite their lack of similarity in environmental and public health characteristics). Thus, it is impossible to conduct meaningful spatial analysis or mapping of the data from these otherwise very valuable data sources.
Health data in the United States, then, are available from a patchwork of sources and at a variety of nonuniform scales (see Box 7.2). Much health data are available at scales that make it extremely difficult to link to environmental exposures or to conduct spatial analyses. Therefore, researchers are frequently faced with the compelling need to generate primary data at considerable cost.
Scale Issues in Spatially Referenced Health Data
The appropriate spatial scale will vary with the frequency of the disease or health event being analyzed. It is far easier to collect data at a fine scale and aggregate upward than it is to collect data at a large scale and then be forced to infer rates at a smaller scale. Because most health data are available only at a high level of spatial aggregation (e.g., county, zip code, or census tract level) and a great deal of within-unit spatial variation is typically present in data attributes, full spatial analysis is not feasible. For example, the National Cancer Atlas9 allows queries by state, county, or State Economic Area based on cancer location and occurrence interval. Although there is substantial geographic variation in cancer mortality within states and within counties, this is not reflected in National Cancer Atlas data. Cancer registries in the United States release data only at the zip code level, and because zip codes are arbitrary units with no inherent geographic or geological significance, they are inferior to census tracts or census block groups for demonstrating spatial variation and drawing conclusions with respect to social and economic variations in health disparities (e.g., Krieger et al., 2002). Although staff members at individual cancer registries and some researchers may—under very restrictive conditions—be able to gain access to spatially more specific locations of patient residences, such access is highly variable and depends on study protocols and Institutional Review Board10 (IRB) restrictions.
Data Access Issues
Why are data so fragmentary and why is it so difficult to obtain data at a fine scale? The fragmentation of data has its roots in government structure, with responsibilities for data collection divided among local health departments, state health departments, and the federal CDC. For conditions—such as cancers—that do not need to be reported to the CDC via local and state health departments, reporting is to local cancer registries. Similarly, trauma cases are reported voluntarily to local trauma registries. Consequently, there is no central repository of health data in the United States and there is considerable variation in the formats and location requirements of the data that are reported.
The reason that the location of incident cases is so difficult to obtain
stems partly from the fragmentation of data, partly from the fact that many conditions are never reported, and partly from the responsibility of government entities to protect patient confidentiality. There is a fear that revealing specific locations, even to reputable researchers under IRB scrutiny, could compromise the privacy of individual patients. In the case of HIV/AIDS, at least early in the epidemic, the CDC expressed concern that identifying a town as a “hotspot” could result in stigmatization of that town. However, it has never been demonstrated—and is, in fact, implausible—that individuals would be identifiable from data collected at the block scale or the census tract scale. This is particularly the case if data are released only to qualified researchers who have passed appropriate training courses and have no inherent interest in identifying individuals. The Health Insurance Portability and Accountability Act (HIPAA) was designed, in part, to ensure that mandatory standards are established to safeguard the privacy of individually identifiable health information (Hobson, 1997; HHS, 2006)—so far, HIPAA seems to have imposed little constraint on biomedical and epidemiological research. The analytical focus for GIS analysis is on aggregate data patterns rather than on a single data point at a specific location.
The lack of available data and a concern for the environmental sources of disease led to an important report by the Pew Environmental Health Commission (Pew, 2000) that made a strong case for a national environmental health “tracking network” to link environmental sources of disease with resulting health conditions (see Box 7.3). The EPA and the Department of Homeland Security signed a Memorandum of Understanding in 2004 to move in the direction of coordinating data to establish such a system. It is crucial that such a system include geographically referenced health data.
OPPORTUNITIES FOR RESEARCH COLLABORATION
Both infectious and noninfectious diseases vary geographically at scales ranging from very local to global. Some of this variation may be random, and there are inferential tests of spatial randomness. For the variation that is not random, the reasons for that variation include environmental factors. One of the major purposes of GIS, remote sensing, and spatial analysis is not only to describe the variation but also to explain it in terms of environmental variables. This requires that earth and public health scientists collaborate to develop spatially and temporally accurate models for predicting disease distribution that incorporate layers of geological, geographic, and socioeconomic data.
Research to link earth science and public health in the United States is
|
BOX 7.3 Data Access and Spatial Analysis of Environmental Contamination, Kolding Town, Denmark Poulstrup and Hansen (2004) used GIS and exposure assessment to investigate the spatial relationship between malignant cancer incidence and exposure to airborne dioxin (Figure 7.2), as a test of the utility of using spatial analysis techniques to assess health effects in a population exposed to environmental contamination. The ability to apply such techniques was dependent on the availability of health and demographic data at the appropriate scale. Health data were derived from the Danish Cancer Registry on an individual basis. The demographic data described each address location (with accuracy to a few meters) and the date of birth, sex, migration (into, out of, and around the area), and date of death for individuals at these addresses. 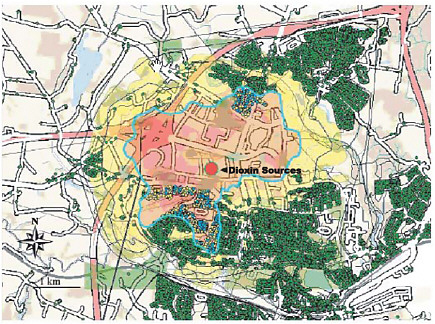 FIGURE 7.2 GIS output showing spatial relationship between three dioxin sources (red dot), airborne exposure model results (yellow/pink shading), and cancer occurrences (yellow dots) in Kolding Town, Denmark. The green dots are addresses that have been geocoded with Universal Transverse Mercator coordinates with a precision of a few meters and which are associated via Denmark’s Central Population Register with each individual’s date of birth, sex, migration (into, out of, and around the study area), and date of death. SOURCE: Poulstrup and Hansen (2004). |
severely hampered by the limited availability of geographically referenced, geocoded health data. It is further hampered by the fragmentary nature of many of the available datasets, which are not coordinated, collated, or concatenated. These issues threaten progress in this area of science and may, in the long run, exacerbate disease that results from human-environment interactions. Accordingly, the committee suggests that:
-
There should be improved coordination between agencies that collect health data, and health data should be merged to the greatest degree possible and made available in formats that are compatible with GIScience analysis.
-
Creative solutions to existing restrictions on obtaining geographically specific health data should be investigated, with the goal of defining a geospatially relevant pixel definition that allows predictive and causal analysis while maintaining individual patient privacy. Data made available by federal, state, and county agencies should be geocoded and geographically referenced to this scale. Legitimate concerns over confidentiality could be further addressed by restricting the release of data to investigators operating under the oversight of Institutional Review Boards.














