1
Scientific Session I: Human Genetic Variation and Nutrition
Moderated by Nicholas J. Schork, Director of Research, Scripps Genomic Medicine, and Professor, Department of Molecular and Experimental Medicine, The Scripps Research Institute, La Jolla, CA
NEW TOOLS FOR UNDERSTANDING THE ROLE OF GENETIC VARIATION IN HEALTH AND DISEASE
Presented by Francis S. Collins, M.D., Ph.D., Director, National Human Genome Research Institute, National Institutes of Health
Over the past decade the field of genomics has seen spectacular and exciting progress. The human genome has been sequenced; and a host of new fields, including genomics, proteomics, and metabolomics, have been developed to use the information made available from the identification of genes and related products. The tools developed in these areas have wide applicability, from medicine to evolutionary biology. However, as Francis Collins noted in the opening session of the workshop, few fields stand to gain more from the application of genomics than the study of human nutrition.
The paradigm for the field of nutrigenomics (see Box 1-1) was established by earlier experiences with nutrient-related diseases, such as phenylketonuria (PKU) and other metabolic disorders. Children with PKU, for example, are unable to metabolize the amino acid phenylalanine into tyrosine, and, if they are left untreated, can develop mental retardation. The discovery that PKU resulted from a genetic mutation led to an effective treatment: a diet that eliminates foods high in phenylalanine, such as dairy products, meats, nuts, and starchy foods. Today infants are routinely
|
BOX 1-1 Nutrigenomics Defined “Nutritional genomics or nutrigenomics is the application of high-throughput genomics tools in nutrition research. Applied wisely, it will promote an increased understanding of how nutrition influences metabolic pathways and how this regulation is disturbed in the early phase of a diet-related disease and to what extent individual genotypes contribute to such diseases.” SOURCE: Nature Review Genetics, 4:241 (April 1, 2003). |
screened for PKU, and those identified as having the disease-causing variant gene are able to live a normal life with dietary modification.
Nevertheless, Collins remarked, “The field of nutrition has not always come across to us hard-nosed scientists as being based on a very rigorous set of scientific findings. The evidence has not impressed us as being as solid as it might be.” The tools of genomics and related fields thus offer the promise of putting the field of nutrition on a firmer scientific footing, supported by both experimental evidence and theoretical understanding. Ultimately, genomics has the potential to dramatically expand comprehension of how nutrients affect the human body and to personalize nutrition, making possible individualized nutritional recommendations. Although nutrigenomics is still an emerging field, many scientists already believe that it will revolutionize the science of nutrition.
The Search for Nutrition-Related Genes
Nutrigenomics can be used in two different ways. The first way is to provide a better understanding of nutrition as it applies to the general population. The second way is to provide an understanding of nutrition at the level of the individual, exploring how nutrients affect people differently, depending on genetic variation.
The first step toward realizing that vision is finding the genetic variants involved in human disease, or what Collins termed “those ticking time bombs that are lurking within our genomes.” The problem is that until recently researchers have not had at their disposal efficient and cost-effective tools for finding the genetic variants that increase the risk of common diseases or identifying the environmental triggers, such as diet, that may set them off.
“We have done very well with Mendelian conditions (for example, PKU) and not so well with the genetic variants that contribute to things
like diabetes or heart disease.” The reason is that unlike Mendelian disorders, which are caused by mutations in a single gene, most common diseases arise from the interaction of many different genetic variants, each of which contributes only a small amount to the total effect. For example, heart disease generally results from a subtle interplay between many different genes and a variety of environmental components, such as diet, none of which has a major influence by itself.
The Genomics Revolution
There is comparatively little variation in genetic makeup between individuals. When the genetic sequences of two individuals are aligned and compared, only about 1 of every 1,000 base pairs of the nucleotide sequence of human DNA exhibits variance. Many of the variations occur as differences in just a single base pair, or “letter” in the DNA code, for example, a cytosine (C) in place of a guanine (G). Scientists refer to a variation involving a single base pair as a single-nucleotide polymorphism (SNP) (Figure 1-1).
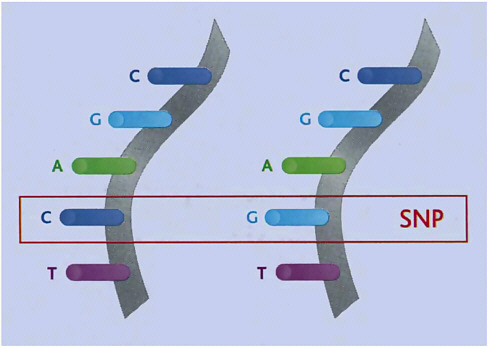
FIGURE 1-1 An SNP is a variant in the genetic code that consists of a single-letter difference in the nucleotide sequence that makes up, for example, DNA.
“Still, when you consider that the size of the human genome is 3 billion base pairs, even a 0.1 percent difference between two people adds up to a lot of genetic variation; some 3 million base pairs” Collins explained. “Most SNPs occur in parts of the genome that aren’t doing very much and therefore don’t have a lot of consequences. But, some of those differences, probably a couple hundred thousand or so, do have important phenotypic effects. It is those that we are most interested in learning about, even though their effects are likely to be subtle.”
To understand a complex disease, particularly one for which nutrition is important, researchers must identify which of the approximately 10 million common SNPs in the human genome are associated with an increased risk of that disease. However, such studies, called “whole-genome association studies,” have been a practical impossibility to conduct, until recently. Because the effects that one is looking for are small, such studies need to look at large numbers of people, ideally, at least 1,000 case patients and 1,000 control subjects, for a total of 2,000 subjects. If one were to analyze, or genotype, all 10 million SNPs in each study participant, a total of 20 billion SNPs would have to be genotyped in such a study. Just a few years ago the cost of genotyping was about 50 cents per SNP, which meant a whole-genome association study involving 2,000 subjects would have carried a staggering price of about $10 billion.
“But within the space of just four years, that has all changed,” said Collins, noting that two factors have come together to make whole-genome association studies a reality. The first is the completion of the International HapMap Project. That effort relied on understanding how DNA behaves during the process of meiosis (division of the cell nucleus) and recombination. When the strands of DNA break during this process, they tend to break only at certain “hot spots” along the DNA, which means that long stretches of DNA move from generation to generation without ever being broken. These stretches, called haplotypes, generally contain many SNPs that travel together in a neighborhood. Consequently, by identifying just one SNP on a haplotype, it is often possible to predict the other SNPs that reside on that stretch of DNA.
On average, haplotypes span about 20 kilobases (20,000 base pairs) of DNA and contain about 30 to 40 SNPs. “The trick is that these haplotypes are not all 20 kilobases,” Collins said. “Some of them are only 1 or 2 kilobases and some are 100 or 150 kilobases. The only way to figure out the boundaries of these neighborhoods is to do the experiment on a certain number of DNA samples.”
That is what the International HapMap Project did. It examined 270 samples from four different populations of people whose ancestors were from northern and western Europe; Yoruba in Ibadan, Nigeria; Tokyo, Japan; and Beijing, China (Han Chinese). The International HapMap
Consortium (2003)1 reported that the Phase I map, a public database of more than 1 million SNP genotypes from the four populations, had been successfully completed and that an even denser Phase II map is now available.
“So, instead of genotyping all 10 million SNPs in each study participant’s genome, researchers can pick a carefully chosen set of about 300,000 marker SNPs for European ancestry or Asian populations, and these serve as very effective proxies for the rest,” said Collins, adding that African populations require slightly more SNPs because their longer population histories have led to somewhat less linkage between the SNPs.
Thanks to the data generated by the HapMap Project, a whole-genome study of 2,000 subjects today requires a total of only 600 million genotypes instead of 20 billion. Yet, the costs of such a study would still be too expensive, except for the fact that over the past 4 years the cost of genotyping has plummeted from about 50 cents per genotype to about one-third of a cent. Thanks to these dramatic improvements in tools and technologies, researchers today can genotype 2,000 subjects for about $2 million, a relatively affordable cost that has made possible a wide range of whole-genome association studies focused on common complex diseases.
This new genomics capability holds tremendous promise for a number of fields but perhaps none more than nutrigenomics. Researchers would begin by collecting information about an individual’s genetic makeup, along with information about what the person eats. They would then compare those data with various health outcomes, for example, blood pressure, cholesterol levels, and rates of cancer and other diseases. With a solid base of research on how genes and diet interact to influence the risk of disease, it should eventually be possible to develop a personalized science of nutrition that would provide an understanding of why individuals respond differently to the same diets and that would allow nutritional recommendations to be tailored to individuals on the basis of their genetic inheritance.
Looking to the Future
If nutrigenomics is to fulfill this promise, the effort will require a multidisciplinary approach involving scientists from a number of areas other than genomics. Research will be needed, for instance, to develop better ways of determining food and nutrient intakes. Collins observed, “We have to get beyond some of the questionnaire-based methods of assessing intake that have been the mainstay of epidemiologists for a long time.” These food-frequency questionnaires are now known to be “hopelessly
inaccurate,” and nutrition researchers need methods of assessing food intake that will measure what people actually eat instead of what they remember eating.
Fortunately, a number of new high-tech methods of assessing dietary intake are now being developed. One method, for example, depends on a network of cell phones that people in a survey would use, at no cost to themselves, to report what they are eating on a real-time basis, perhaps even using the camera feature of the cell phone to take a picture of the meal.
According to Collins, researchers interested in applying nutrigenomics will face another, more familiar hurdle: finding funding. Even though the cost of whole-genome association studies has dropped by a factor of 5,000 over the past several years, nutrigenomics studies will still require significant resources. Fortunately, though, funding sources are appearing. One of them is a public-private partnership of the Foundation for the National Institutes of Health called the Genetic Association Information Network (GAIN). The partnership is making funds available for whole-genome association studies that will use data from existing case-control studies of patients with common diseases. GAIN has funds on hand for six studies with 1,000 case patients and 1,000 control subjects each. The deadline for submitting proposals for that round of funding was May 9, 2006. However, other rounds will be announced as funding becomes available.
A second possibility is the Genes and Environment Initiative, which was in President George W. Bush’s fiscal year 2007 proposed budget and which has been strongly promoted by the secretary of the U.S. Department of Health and Human Services. The initiative’s aim is to increase the understanding of the complex interplay of genes and the environment in the development of common diseases, and diet is one of the major environmental factors under consideration. The initiative has two main components. “One is to encourage yet more genotyping of case-controlled studies of common disease. In this initiative, we are particularly looking for studies where good data is available on diet, physical activity, and environmental exposures in order to be able to draw conclusions about interactions,” Collins said. “There is also $14 million a year for the next 4 years to develop innovative technologies to measure those environmental exposures, including dietary intake and physical activity, in a more rigorous fashion than what has often been the case in these kinds of studies.”
Even as scientists are beginning to appreciate the potential of nutrigenomics, the public is already fascinated. For instance, a number of Internet sites offer products based on the idea that one’s diet should be dictated by one’s DNA. In general, companies analyze a few genes that are known to be involved in metabolism or that are influenced by certain nutrients,
determine which variants of these genes that a person has, and then make dietary recommendations based on that information. The concern, according to Collins, is that most of these recommendations are based on very slim evidence. The gene variants analyzed may have been found to be associated with particular outcomes, for example, higher cholesterol levels in people who also eat a high-fat diet, but there have been no prospective studies showing that people taking such dietary advice actually benefit from it.
“So, we already have a problem here,” Collins said. “Before the field of nutrigenomics has barely gotten itself defined, we have a lot of activity out there marketing products to the public that are, at best, sketchy in terms of their scientific basis. “We need to move quickly. The easiest way to kill a field is to overpromise, and there is a lot of overpromising going on.”
Nutrigenomics has tremendous potential to revolutionize the understanding of nutrition, particularly nutrition on the individual level, and to help move the focus of medicine from treatment to prevention. There is also a risk, however, that nutrigenomics could come to be seen as just one more food fad in a long line of food fads. To avoid that fate, Collins said, “We need to give it some good science to grow on.”
IMPLEMENTATION OF THE HUMAN HAPMAP INITIATIVE AND LARGE-SCALE POLYMORPHISM STUDIES
Presented by David Cox, Chief Scientific Officer, Perlegen Sciences, Inc.
Over the past 20 years researchers have learned that most complex traits involve probably 20 or more individual genetic changes, according to David Cox. Each of those changes is responsible for only a small contribution on its own, so to understand the genetics of a particular complex trait, it is necessary to identify at least 20 or so individual genes. Many more than 20 may influence the trait, Cox said, but generally 20 or so genes in aggregate will account for enough of the variation in the trait to be practically useful.
According to Cox, once the various genes that influence a trait have been identified, it is generally possible to estimate a person’s risk for disease by adding the risks from the individual genes. In statistical terms, that property is called “additive variance.” “What we are doing,” Cox said, “is just slamming them all together and saying that it doesn’t really matter which one does what. You can just add them together without understanding the interactions at all, without understanding the mechanisms, and you can use it in a very useful predictive way.”
The key is to identify the individual genes that influence a particular trait, and the smaller the contribution is, the more difficult it is to pinpoint the gene. A hypothetical example can be used to describe the situation: a gene may have two variants, referred to as “Variant A” and “Variant B,” and a person with two copies of Variant B has a 50 percent greater chance of developing a trait such as heart disease than a person with two copies of Variant A. The gene would then explain a relatively small part, for example, 2 percent, of the total variation in the population, that is, which people get heart disease versus which people do not; but the risk would still be significant enough to pay attention to the gene, particularly in combination with other genes with similar contributions.
For a gene with this level of effect, one would need several hundreds of subjects of each variant type, that is, those with two copies of Variant A and those with two copies of Variant B, to achieve sufficient statistical power to identify the gene’s contribution with a reasonable level of certainty and accuracy. If the contribution of the gene were smaller, a larger subject pool would be needed to determine significance.
The traditional way to identify genes that influence a particular trait has been to use the candidate gene approach. Genes that are likely to be involved, based on previously identified traits, are selected and subjects, both carriers of the trait and noncarrier controls, are tested. The problem with this approach is that it requires that one be able to identify ahead of time most of the candidate genes, and that is seldom the case.
As an example, Cox described a study performed to identify the genes involved in determining the level of high-density lipoprotein (HDL) production, an indicator of risk for cardiovascular disease. Individuals with high HDL levels are at lower risk for coronary heart disease than those with low HDL levels. Although a number of candidate genes that potentially explain a large percentage of the variation in HDL levels among individuals were selected for genetic study, only one of these genes was found to be significantly associated with variations in HDL levels. Individuals with one copy of Variant A and one copy of Variant B of the gene were twice as likely to have high levels of HDL than people with two copies of Variant B. Nevertheless, the gene accounted for relatively little of the total variation among people with regard to their HDL levels. The findings suggest that a large fraction of the genetic variation that leads to differences in HDL levels is in genes other than the “traditional” HDL candidate genes.
Cox referred to the example of “looking under the lamppost,” because “that is where the light is. So, we clearly have to light up the street, the entire genome, if we are going to have the power to do this more systematically; and the good news is we have arrived at that ability.”
CONTEMPORARY NUTRIGENETICS STUDIES
Presented by Jose M. Ordovas, Senior Scientist and Director of the Nutrition and Genomics Laboratory, Jean Mayer U.S. Department of Agriculture Human Nutrition Research Center on Aging at Tufts University
Jose Ordovas offered an example of how nutrigenomics can be used to make sense of what otherwise seem to be confusing and contradictory findings in the field of nutrition. As Ordovas pointed out, a variety of large-scale studies have failed to find a consistent effect of diet on health, despite expectations to the contrary. A study published in the Journal of the American Medical Association2 found no evidence that eating fruits and vegetables lowered the risk of breast cancer, for example. Another study3 found no effect of either vitamin E or aspirin on the incidence of breast cancer in women. Still others have raised questions about the usefulness of low-fat diets in preventing heart disease.
To show what those studies may be missing, Ordovas focused on a particular question: how levels of HDL, the “good cholesterol,” are affected by having a particular version of the gene for apolipoprotein A-1 (APOA1). In analyzing data from the large-scale Framingham Heart Study, Ordovas said, it would appear at first glance that the different versions of APOA1 make no difference to the levels of HDL in a person’s bloodstream, as the average HDL levels are the same among the groups with the different genes. He said that the analysis, however, does not take diet into account and that when diet is taken into account, a totally different picture emerges.
The Ordovas laboratory reanalyzed the data and further divided the groups according to how much polyunsaturated fat that they had in their diet. The APOA1 gene has either the nucleotide guanine (G) or adenine (A) in their DNA at a particular locus, or position, in the APOA1 gene sequence, depending on which version of the gene a person inherits, so a person’s two copies of the gene can be GG, GA, or AA. Ordovas found that for people with the GG genotype, the more polyunsaturated fat that they had in their diets, the lower their levels of HDL were; but the pattern was exactly the opposite for the groups with GA and AA: their HDL levels went up with higher levels of consumption of polyunsaturated fat.
Ordovas summarized by indicating that “it is particularly important for people of GG type to minimize their consumption of polyunsaturated fat in order to keep their levels of the good cholesterol as high as possible.” Conversely, people with the GA or the AA form of the gene can achieve increases in good cholesterol levels by increasing their dietary intake of polyunsaturated fat. It is thus important for both doctors and researchers to take into account the genetic makeups of their patients when they try to understand and predict the effects of diet on health.










