1
Why Metagenomics?
Microbes run the world. It’s that simple. Although we can’t usually see them, microbes are essential for every part of human life—indeed all life on Earth. Every process in the biosphere is touched by the seemingly endless capacity of microbes to transform the world around them. The chemical cycles that convert the key elements of life—carbon, nitrogen, oxygen, and sulfur—into biologically accessible forms are largely directed by and dependent on microbes. All plants and animals have closely associated microbial communities that make necessary nutrients, metals, and vitamins available to their hosts. Through fermentation and other natural processes, microbes create or add value to many foods that are staples of the human diet. We depend on microbes to remediate toxins in the environment—both the ones that are produced naturally and the ones that are the byproducts of human activities, such as oil and chemical spills. The microbes associated with the human body in the intestine and mouth enable us to extract energy from food that we could not digest without them and protect us against disease-causing agents.
These functions are conducted within complex communities—intricate, balanced, and integrated entities that adapt swiftly and flexibly to environmental change. But historically, the study of microbes has focused on single species in pure culture, so understanding of these complex communities lags behind understanding of their individual members. We know enough, however, to confirm that microbes, as communities, are key players in maintaining environmental stability.
By making microbes visible, the invention of microscopes in the late 18th century made us aware of their existence. The development of labora-
tory cultivation methods in the middle 1800s taught us how a few microbes make their livings as individuals, and the molecular biology and genomics revolutions of the last half of the 20th century united this physiological knowledge with a thorough understanding of its underlying genetic basis. Thus, almost all knowledge about microbes is largely “laboratory knowledge,” attained in the unusual and unnatural circumstances of growing them optimally in artificial media in pure culture without ecological context. The science of metagenomics, only a few years old, will make it possible to investigate microbes in their natural environments, the complex communities in which they normally live. It will bring about a transformation in biology, medicine, ecology, and biotechnology that may be as profound as that initiated by the invention of the microscope.
WHAT IS METAGENOMICS?
Like genomics itself, metagenomics is both a set of research techniques, comprising many related approaches and methods, and a research field. In Greek, meta means “transcendent.” In its approaches and methods, metagenomics circumvents the unculturability and genomic diversity of most microbes, the biggest roadblocks to advances in clinical and environmental microbiology. Meta in the first context recognizes the need to develop computational methods that maximize understanding of the genetic composition and activities of communities so complex that they can only be sampled, never completely characterized. In the second sense, that of a research field, meta means that this new science seeks to understand biology at the aggregate level, transcending the individual organism to focus on the genes in the community and how genes might influence each other’s activities in serving collective functions. Individual organisms remain the units of community activities, of course, and we anticipate that metagenomics will complement and stimulate research on individuals and their genomes. In the next decades, we expect that the top-down approach of metagenomics, the bottom-up approach of classical microbiology, and organism-level genomics will merge. We will understand communities, and the collection of communities that forms the biosphere, as a nested system of systems of which humans are a part and on which human survival depends. In some situations, it will be possible to apply the new understanding to problems of urgency and importance.
Metagenomics in either sense will probably never be circumscribed tightly by a definition, and it would be undesirable to attempt to so limit it now, but the term includes cultivation-independent genome-level characterization of communities or their members, high-throughput gene-level studies of communities with methods borrowed from genomics, and other “omics” studies (see Box 1-1), which are aimed at understanding transorganismal
|
BOX 1-1 The Other “Omics” Sciences The term genome was first proposed by Hans Winkler, a professor of botany at the University of Hamburg, Germany, in 1920 (Winstead 2007). It was coined to describe the total hereditary material contained in an organism long before it was known that genetic information is encoded by DNA. Today genome is used to describe all the DNA present in a haploid set of chromosomes in eukaryotes, in a single chromosome in bacteria, or all the DNA or RNA in viruses. The suffix ome is derived from the Greek for “all” or “every.” In the past several years, many related neologistic omes have come into use to describe related fields of study that encompass other aspects of large-scale biology. Some of them are:
The list of “omes” and “omics” is growing longer as scientists develop new tools and approaches for carrying out large-scale studies of biological systems. |
behaviors and the biosphere at the genomic level. Although in its current early implementation (and for the purposes of this report) metagenomics focuses on non-eukaryotic microbes (see Box 1-2), there is no doubt that its concepts and methods will ultimately transform all biology. In just this way has genomics, a science developed to aid the advancement of biomedicine and the understanding of our own species, transformed the science of all organisms and the application of that science in epidemiology, clinical microbiology, virology, agriculture, forestry, fisheries, biotechnology, microbial forensics, and many other fields.
In conceptualizing metagenomics, we might simply modify Leroy Hood’s definition of systems biology as “the science of discovering, modeling, understanding and ultimately managing at the molecular level the dynamic relationships between the molecules that define living organisms” (Hood 2006). We need only replace the last word, organisms, with the phrase “communities and the biosphere.”
|
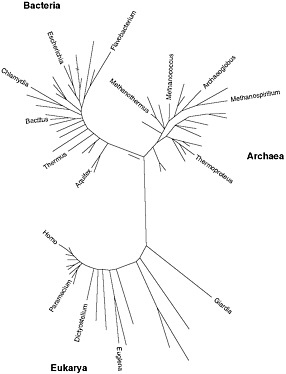
BOX 1-2 A Note on Terminology What is a microbe? In practice, the term microbe is used to describe living things invisible to the human eye, that is, generally less than about 0.2 mm. The terms microbe, microorganism, bacteria, germ, and even bug are often used interchangeably by nonscientists to describe these small organisms. Microbiologists have specific names for the various microbes, which include Bacteria, Archaea and some members of the Eukarya. The first two groups (domains), although unlike in many ways, share a type of cellular organization known as prokaryotic. They lack membrane-enclosed organelles, such as mitochondria, chloroplasts and, most notably, a nucleus. The genomes of Bacteria and Archaea typically contain little non-coding DNA and range in size from 0.5 to 10 million base pairs. By contrast, members of life’s third domain, Eukarya, which comprises animals, plants, fungi, algae, and protozoa have larger genomes with substantially more non-coding DNA. Some eukaryotes are also too small to be seen individually except under a microscope and thus have been traditionally studied by microbiologists. Included among these small eukaryotes are many fungi, such as baker’s yeast and the human pathogen Candida, and many of the algae and protozoa (harmless paramecia, for instance, and the malaria parasite Plasmodium). Viruses, although arguably not alive, in that they can replicate only inside cells and have no metabolism or cell structure of their own, are also encompassed in the science of microbiology. In this report, we address primarily metagenomics projects that focus on Bacteria, Archaea and viruses. Because of their larger genomes, microbial eukaryotes have received less attention, a situation which should be remedied as sequencing becomes less expensive and bioinformatic methods become more powerful. |
WHAT MICROBES CAN DO: FOUR EXAMPLES
We start with examples. There are countless ways in which microbes influence daily life. Earth is a biological entity as much as it is a physical one, and most of the vital biology, on which all life depends, is microbiology (see Box 1-2). But because microbes are individually invisible, we (even microbiologists) need to be reminded of our debt to them. Here are four of the thousands of reasons.
Microbes Modulate and Maintain the Atmosphere
Carbon is the most abundant chemical element in all living things, including humans (excluding the hydrogen and oxygen in the water, which makes up the bulk of our weight). Carbon dioxide (CO2) in the atmosphere
is the most abundant source of carbon on Earth, but in this form it is inaccessible to animals and most bacteria. Plants and some bacteria “fix” carbon through photosynthesis, a light-driven conversion of CO2 to sugars that generates the oxygen that fuels all aerobic forms of life. Although plants tend to get most of the credit, bacteria are responsible for about half of the photosynthesis on Earth (Pedros-Alio 2006).
Ocean microbes, collectively present at billions of cells per liter, grow at rates of about one doubling per day in surface waters and are consumed at about the same rate (Whitman et al. 1998). The organisms that carry out photosynthesis turn over rapidly in the ocean as well, on the average about once per week. Net primary productivity in the global ocean is estimated to fix 45-50 billion tons of CO2 per year (Falkowski et al. 1998). Chemical transformations mediated by marine microbes play a critical role in global biogeochemical cycles (see Figure 1-1). The collective metabolism of marine microbial communities has global effects on fluxes of energy and matter in the sea, on the composition of Earth’s atmosphere, and on global
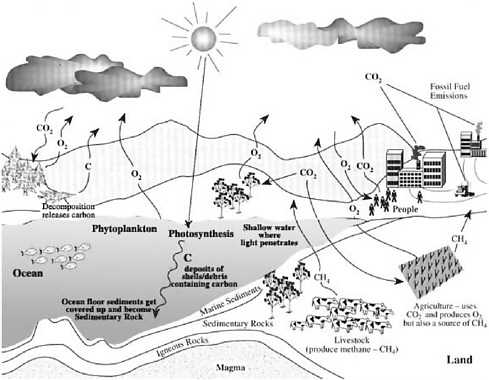
FIGURE 1-1 The global carbon cycle. SOURCE: http://www.bigelow.org/foodweb/carbon_cycle.jpg.
climate. In essence, the combined activities of microbial communities affect the chemistry of the entire ocean and maintain the habitability of the entire planet. Hidden within the population dynamics of these complex communities are fundamental lessons of environmental response and sensing, species and community interactions, gene regulation, and genomic plasticity and evolution. Microbes are the stewards of Earth’s biosphere and are Nature’s biosensors par excellence.
Perhaps most obviously today, the living oceans play a critical role in the global carbon cycle (Falkowski et al. 1998). The coupling of the upper ocean and the atmosphere results in higher concentrations of dissolved CO2 in surface seawater than in the rest of the ocean. Much of the elevated carbon input can move through the action of the ocean’s “biological pump,” which depends on microbial communities in the surface water that transform inorganic CO2 into organic carbon. The organic carbon can either be respired and recycled back to the upper ocean-atmosphere system or sink out of the surface water and be sequestered in the deep ocean. Complex microbial community interactions help to regulate the proportion of recycled versus sequestered carbon. The structure of the phytoplankton community, the rates at which phytoplankton are attacked and destroyed by viruses, and the capacity of other microbes to turn organic carbon back into CO2 all influence the fate of carbon, and the ability of the ocean to act as a source of, or a sink for, CO2. CO2 is a very important greenhouse gas, so photosynthetic bacteria serve the planet in two ways: they convert carbon into biologically accessible forms and they remove CO2 from the atmosphere, thereby mitigating some of the anthropogenic release of CO2 and other greenhouse gases.
Microbes Keep Us Healthy
It should come as no surprise that in the microbe-dominated biosphere, close relationships between microbes and animals are an ancient theme. Humans are no exception. The numbers are staggering. The microbes that reside on the surface of the human body alone outnumber human cells by about a factor of 10. The genomes of members of our indigenous microbial communities (the human metagenome) contain thousands of times more genes than the human genome (Gill et al. 2006). Microbial communities also inhabit the human mouth, skin, and respiratory and female reproductive tracts. The compositions of these communities change over time and, for some body sites, like the oral cavity, there is already evidence that certain community compositions are associated with periodontal disease. Understanding how microbial community structure affects health and disease may contribute to better diagnosis, prevention, and treatment of disease. The vast majority of these microbial partners live in the intestine,
where a diverse community of microbes, 10 to 100 trillion in number, perform functions that humans have not had to evolve, including the extraction of calories from otherwise indigestible components of our diet and the synthesis of essential vitamins and amino acids. The complex communities of microbes that dwell in the human gut shape key aspects of postnatal life, such as the development of the immune system, and influence important aspects of adult physiology, including energy balance. Gut microbes serve their host by functioning as a key interface with the environment; for example, they defend us from encroachment by pathogens that cause infectious diarrhea, and they detoxify potentially harmful chemicals that we ingest (intentionally or unintentionally). In light of the crisis in management of infectious pathogens due to emergence of antibiotic resistance, we would be well served to understand the role of microbial communities in protecting us from infectious agents. Our microbes are master physiological chemists: identifying the chemical entities that they have learned to manufacture and characterizing the functions of human genes and gene products that they manipulate should lead to valuable additions to our 21st-century medicine cabinet (pharmacopeia).
Microbes Support Plant Growth and Suppress Plant Disease
The microbial communities on and around plants play a central role in the health and productivity of crops. The most complex of these communities reside in the soil, which is a composite of mineral and organic materials teeming with bacteria and archaea. Some functions of these microbes are well known. Some bacteria fix atmospheric nitrogen, converting it from dinitrogen gas—a form unusable by plants and animals—to ammonia, which is readily used. Other soil microbes recycle nutrients from decaying plants and animals, and others convert elements, such as iron and manganese, to forms that can be used for plant nutrition. Soil microbial communities determine whether plants will become infected by pathogens. A lingering mystery is the “suppressive soil” phenomenon (Mazzola 2004). In some soils, plants stay healthy even when pathogens are present at high density; when the soil is sterilized, the disease suppression disappears, suggesting a biological basis of the phenomenon. However, in only very few cases has a single microbe isolated from a soil been able to duplicate the suppression. After decades of wrestling with the enigma of suppressive soils, plant pathologists have concluded that in many cases a complex community is responsible for the suppressive activity, which is hugely beneficial to agriculture. No organism has been found to provide the same effect in isolation, because the community members modify each other’s behavior.
Microbes Clean Up Fuel Leaks
There are hundreds of thousands of underground storage tanks in this country, most of which are used for storing gasoline. In fact, almost every corner gasoline station in the United States uses three or more of these tanks to dispense regular, premium, and super-premium versions of gasoline. The sad truth about these underground tanks is that the vast majority of them are already leaking or will leak and send gasoline into the subsurface, where it has the potential to contaminate the groundwater. Given the ubiquity and magnitude of the gasoline leaks and the fact that 50% of the US population relies on groundwater as a drinking-water source, one must wonder how it is that we are not all drinking water contaminated with gasoline!
The answer is that we are being protected by the omnipresent and vastly adaptable subsurface microbial community (Mazzola 2004). As gasoline is released into the subsurface, relatively dormant members of the microbial community are triggered to become active and biodegrade the gasoline constituents. Gasoline is composed of thousands of organic chemicals and a variety of microbes containing complementary metabolic systems are required to degrade them all. Furthermore, because there is too little of any single electron acceptor in the subsurface to react with all the electron donors of gasoline, different bacteria with different respiratory capabilities are required to complete the gasoline remediation. For example, when oxygen is depleted in the groundwater in the vicinity of a gasoline spill, bacteria that can respire nitrate take over, followed by bacteria that respire iron, manganese, sulfate, and, eventually, CO2. This complicated community of microbes works together in a self-organized pattern triggered by the movement of the leaking gasoline until the contaminants have been transformed into harmless CO2 and water. The microbial community then becomes dormant again, awaiting the next influx of substrate (either natural or anthropogenic) to return to activity.
INVISIBLE COMMUNITIES: GLOBAL IMPACT
Modulating the atmosphere, keeping humans and plants healthy, and cleaning up leaking gasoline are just a few examples of the many things that microbial communities can do. The combined activities of microbial communities shape the face of the biosphere on a global scale. The power of these communities lies hidden in the metabolic versatility of their component species that, acting together, regulate the vast majority of matter and energy transformations on Earth. In a loose analogy, the entire biosphere can be imagined as a sort of “superorganism.” Its many systems for the recycling of carbon, oxygen, nitrogen, and phosphorus can be compared with the organs of the human body working in unison to facilitate circulation,
nutrient acquisition, respiration, waste processing, and so forth. Unquestionably, humans depend on these global geochemical cycles, and microbes are vital players in the cycles’ operation and stability. Microbes can “eat” rocks, “breathe” metals, transform the inorganic to the organic, and crack the toughest of chemical compounds. They achieve these amazing feats in a sort of microbial “bucket brigade”—each microbe performs its own task, and its end product becomes the starting fuel for its neighbor. For complex transformations, no microbe can do it alone—it takes a community. For example, no microbial species is capable of completely oxidizing ammonia to nitrate, but teams of microbes do it efficiently. One microbial group oxidizes ammonia to nitrite, and its waste becomes the fuel for another species that transforms nitrite to nitrate, completing the “bucket brigade.” Virtually all elemental cycles—including the generation, consumption and flux of greenhouse gases (or, as noted above, the remediation of spilled gasoline)—involve similar sorts of microbial collaborations that are tightly regulated and coupled through microbial community interactions. So the bucket brigades are themselves interconnected laterally—an interwoven web of chains. In this way, microbial communities play essential roles in the transformations of energy and matter, producing the air we breathe and shaping the biosphere and climate that we enjoy on Earth today.
Larger organisms play key roles, too, of course: about half of all carbon is fixed and half of all oxygen produced by trees, grasses, and other macroscopic plant life. But these larger organisms also depend on microbes; for example, plants depend on the nitrogen fixation carried out by symbiotic microbes in the roots of legumes and other plants that form symbiotic associations. Humans might survive in a world lacking other macroscopic life forms, but without microbes all higher plants and animals, including humans, would die. Not only can many individual systems—for example, the human gut or such processes as the bioremediation of toxic hydrocarbons—be seen to be the tasks of complex and dynamic microbial communities, but these communities are themselves constituents of even larger systems, predominantly microbial, that collectively make up the biggest and most complex functioning system we know: the biosphere. Whatever the causes, extent, and consequences of the global climate change now upon us, the biosphere’s response to the changes—and human survival—will depend on its microbes and their activities.
We live in a time of unprecedented and dramatic global change, in which the effects of human activities challenge the ability of natural ecosystems to buffer them. The industrial revolution marked the beginning of rapid environmental transformation. For example, until the early 20th century, all nitrogen entering the biosphere was produced from atmospheric nitrogen by microbes, providing the organic nitrogen required for new plant growth. In the early 1900s, the Haber-Bosch process was invented to
perform the same job to produce vast amounts of nitrogenous plant fertilizer from atmospheric nitrogen; this industry now produces more organic nitrogen than all biological processes combined (Socolow 1999). Another obvious and dramatic change in the global environment is the enormous amount of CO2 released by the burning of fossil fuels, previously stored as relatively inert reservoirs deep in Earth. Present concentrations of atmospheric CO2 are higher than they have been in 420,000 years and, given current trajectories, will continue to rise dramatically (Petit 1999).
Understanding the dynamic role of microbial communities in this rapidly changing environment is a critical and currently unmet challenge. How resilient are microbial communities in the face of such rapid global change? Can microbial communities, versatile as they are, help to buffer and mediate key elemental cycles now undergoing rapid shifts? Can changes in microbial communities serve as sensors and early-alarm systems of environmental perturbation? To what extent can we “manage” microbial communities to modulate the effects of human activities on natural elemental cycles sensibly and deliberately? Never before have such questions had such urgency.
UNDERSTANDING MICROBIAL COMMUNITIES
Given that the microbial collective profoundly influences geochemical and greenhouse-gas cycles, as well as climate and environmental change, it is relevant to ask how well we understand microbial communities. In the past, it was difficult to study microbes in their own environments; microbiologists studied individual species one by one in the laboratory. It now appears that many microbes function in nature as multicellular, often multi-species, entities, sometimes even physically connected (as in biofilms) and often metabolically connected.
The Limits of Pure Culture
Even into the 19th century, some scientists believed that microbes were generated spontaneously from nonliving matter or from other organisms. Establishing that such tiny entities were organisms that belonged to definable, fixed species was difficult. Fixity of species was especially important in theories of disease causation; fixed species were essential if a single bacterial species was to be held responsible for a single infectious disease. Agriculturalists and botanists had long suspected that some sort of unseen organisms were associated with plant disease; in 1726, for example, the association farmers saw between barberry rust and wheat rust led the Connecticut colonial legislature to ban the bushes (Campbell et al. 1999). Over a century later, the German botanist Anton de Bary demonstrated the
correlation between the life cycle of Phytophthora infestans and the disease cycle of late blight of potato. In a series of experiments conducted in the late 1850s and early 1860s, he built on the previous work of J. Speerschneider and Marie-Anne Libert and established that P. infestans was indeed the cause of the disease (Matta 2007).
Demonstrating that microorganisms were not spontaneously generated and had distinct species was fundamental to bacteriology as well. Robert Koch published his description of the life cycle of Bacillus anthracis (the cause of anthrax) in 1876 and then published a series of papers in which he established an experimental method for confirming the specific causes of various infectious diseases. In an 1884 paper on tuberculosis, he outlined his four “postulates” for proof of microbial causation: an organism must be found in all cases of the disease but not in healthy hosts, the organism must be isolated from the host and grown in pure culture, reintroduction of the organism from such cultures must cause disease in healthy hosts, and the organism must again be isolatable from such infected hosts (Munch 2003). That rigorous approach, particularly the emphasis on pure cultures (a culture that contains organisms of only one type) set the standards for microbiology as a whole. By the middle of the 20th century, even with “environmental microbes” (the vast majority of harmless and beneficial bacteria, archaea and microbial eukaryotes), pure cultures became a gold standard for experimentation and the basis of almost all recent knowledge of medical bacteriology, biochemistry, and molecular biology.
In the pure-culture paradigm, the presence of multiple species in the same culture medium means “contamination,” and species whose growth requires metabolic products of other species are impossible to detect, study, or even name. Not surprisingly, microbes that grow well as single cells suspended in a liquid medium and that can easily form discrete colonies on Petri plates became the model for much of modern biology. Indeed, many microbiologists came to view the “planktonic” state as the natural condition of microbes—complex communities and slimy biofilms being somehow an aberration and unworthy of serious scientific attention. On the contrary, it is now becoming clear that many microbes live in communities whose members interact and communicate in complex ways. Microbial communities often interact through the medium (water or soil) in which they grow, exchanging nutrients, biochemical products, and chemical signals without direct cell-to-cell contact. Some grow on surfaces (on suspended particles, on the walls of pipes, on teeth) where they are in physical contact with others of their own kind and with other species. Biofilms, which are aggregates of microbial cells embedded in an extracellular polysaccharide matrix, exhibit a great diversity of complex structures. The composition of such communities is far from accidental. Many microbes have evolved to grow together in surface communities and many of their collective activi-
ties, whether vital to the biosphere or detrimental to human health, reflect the physical structure and division of labor within the communities.
The study of microbes in culture will continue to be important, but it falls short of telling us about environmental processes, biofilms, microbial bucket brigades of energy and matter flux, and the future trajectory of biogeochemical cycles. Understanding microbial communities will require that the traditional techniques of pure culture be supplemented with new approaches.
The Genomics Promise
One approach that has contributed greatly to understanding all organisms is genomics—learning about the evolution and capabilities of organisms by deciphering the sequence of their DNA. Genomics has also greatly advanced microbiology, but, like pure culture, traditional genomics is limited in its ability to elucidate the dynamics of microbial communities.
The precipitous decline in the cost of gene sequencing, spurred in part by the Human Genome Project, has made it possible to generate genomic sequences for a great variety of organisms. The first microbial genome sequenced, that of the pathogen Haemophilus influenzae, was published in 1995 (Fleischmann et al. 1995). Microbial genome sequences have since appeared at an exponentially increasing rate: the genome sequences of 399 bacteria, 29 archaea, and almost 30 eukaryotic microbes are publicly available at the time of this writing. Pathogenic bacteria and eukaryotes—such as the causative agents of plague, anthrax, tuberculosis, Lyme disease, candidiasis, malaria, and sleeping sickness—have received much attention. But many nonpathogenic archaea and bacteria have also been sequenced, including such beneficial organisms as several species of Prochlorococcus and Synechococcus, major producers of oxygen in the ocean; Dehalococcoides ethenogenes, effective in the bioremediation of soils contaminated with chlorinated hydrocarbons; Lactobacillus acidophilus, used in making yogurt; Bradyrhizobium japonicum, a nitrogen-fixing symbiont of soybeans; and Saccharomyces cerevisiae (baker’s yeast).1
When attention turned to sequencing the genomes of microbes, the preference for working in pure culture was reinforced. No one knew how difficult it might be to sequence an entire genome, but it was obvious that assembly (using a computer to put the sequenced fragments together in complete genomes) would be vastly more complicated if the pieces belonged to several different organisms (see Box 1-3). Until recently, all microbial genome sequences were determined from pure cultures. But in the last few years, more than a dozen microbes that can be physically separated
|
1 |
See http://www.ncbi.nlm.nih.gov/Genomes/ for more information. |
|
BOX 1-3 Blueprints for the Living World: Genes, Genomes, and Genomic Sequences Genes are made of DNA, and the exact sequence of the four canonical DNA bases (designated A, T, C, and G) in any gene specifies the product (usually a protein) that it encodes. In bacteria and archaea, genes are about 1,000 base pairs long. These microbes have 500-10,000 genes, usually arrayed on a single circular DNA molecule (a chromosome), some 600,000-12 million base pairs long (there is some space between genes for regulatory signals). Eukaryotic microbes typically have more and longer genes and multiple chromosomes. Together, all the genes in a microbe’s chromosome or chromosomes and any in accessory genetic elements, such as plasmids, make up its genome. For complete genome sequencing, the whole genome shotgun approach has proved effective. All the DNA from a pure culture is fragmented randomly into pieces of one to a few thousand base pairs. Fragments totaling some 6-10 times the genome’s length are sequenced so that overlaps between them can be used to establish the order of the fragments in the intact genome and verify the accuracy of the sequencing. This step, called assembly, is computationally intensive. So is the next step, annotation, which is the prediction of gene boundaries, regulatory regions, and the properties and function of the proteins (or sometimes RNAs) that the genes encode. Annotation usually involves finding a similar gene sequence for which a function has already been determined in another organism, although at present typically one-third of the genes in any newly sequenced microbe will not have any obvious similarity to genes with known or proposed functions. Finally, the data are released to a public data repository, such as GenBank, maintained by the National Center for Biotechnology Information (National Library of Medicine) in Bethesda, Maryland. |
from other major sources of DNA or that greatly predominate where they are found in nature have also been sequenced. Treponema pallidum and Mycobacterium leprae (which cause syphilis and leprosy, respectively) are among the former, and two species predominant among acid-mine drainage site biofilms (Ferroplasma acidarmanus and a species of Leptospirillum) are examples of the latter. Sequencing such physically purified or environmentally concentrated (and thus naturally “pure”) microbes crosses the boundary between genomics and metagenomics as far as methods are concerned.
Soon, there will be thousands of sequenced microbial genomes. If all microbial species were culturable and if such species were easily defined and limited in number (even a number in the tens of thousands), the ultimate goal of microbial genomics might be to determine all these genome sequences once the per-genome cost fell far enough. Then the meta in
metagenomics might parallel its use in meta-analysis and mean bringing together individual databases in search of a common set of truths about nature. But not all species are culturable, few are easily defined at the genomic level, and indeed the number of different genomes in nature turns out to be uncountably large. We discuss these problems in turn.
WHY GENOMICS IS NOT ENOUGH
Most Microbes Cannot Be Cultured
In 1985, Staley and Konopka reviewed data on scientists’ ability to bring microbes from the environment into laboratory cultivation. The “great plate-count anomaly” they identified was this: the vast majority of microbial cells that can be seen in a microscope and shown to be living with various staining procedures cannot be induced to produce colonies on Petri plates or cultures in test tubes. It is estimated that only 0.1-1.0% of the living bacteria present in soils can be cultured under standard conditions; the culturable fraction of bacteria from aquatic environments is ten to a thousand times lower still. The application of genomics-inspired moderate- to high-throughput nutrient screening methods and nontraditional approaches to monitoring growth responses will no doubt bring many recalcitrant organisms into culture. Indeed, two recent successes are the cultivation (and genome sequencing) of Pelagibacter ubique, a bacterium representative of one of the most common microbial phylogenetic groups found in the open ocean, and the isolation of several acidobacteria, the most abundant organisms in soil (Sait et al. 2002; Field et al. 1997; Martinez and Rodriguez-Valera 2000; Brown and Fuhrman 2005; Rappe et al. 2002). Both successes depended on the nontraditional molecular (rRNA-based) method discussed below for monitoring growth. But the fraction of organisms cultivatable in isolation will likely always be low, and for most the reason will be that, for growth, it takes a community. Culturing always favors the recovery of organisms that are best able to thrive under laboratory conditions (colloquially “lab weeds”), not necessarily the dominant or most influential organisms in the environment.
Given the evidence that many microbes resist being cultured, culture-independent methods for identifying and enumerating microbes in the environment have come to play a larger and larger role over the last several decades. Predominant among them is ribosomal RNA (rRNA) phylotyping, a powerful technique—indeed, an independent research paradigm—developed by Pace and his colleagues (Pace 1997). This method is based on the enormous database of rRNA gene sequences (more than 200,000) that have been collected for the purpose of reconstructing the universal Tree of Life (see Box 1-4). By determining the sequence of an organism’s rRNA genes,
one can position it on the appropriate branch of the Tree of Life and infer that its biology and ecology are likely to be similar to those of its closest relatives, the nearest branches on the tree. An organism does not have to be culturable to determine its phylotype. The polymerase chain reaction (PCR) allows rRNA (or other) genes to be detected and copied directly from environmental samples, then cloned and sequenced. If the environmental sample contains many types of organisms, there will be many different rRNA sequences, the diversity of which will be a measure of the complexity of the community and which, in the context of the Tree, will tell us “who is there.” Phylotyping has revolutionized the field of microbial ecology, and hundreds of environments—from dry Antarctic valleys to deep-sea hydrothermal vents (“black smokers”) to sewage-treatment plants and methane-producing reactors—have been studied in this way. Very often, new lineages whose rRNA gene sequences are little like anything that has been cultured are discovered. Indeed, the majority of the 50-plus major divisions of Bacteria that have been delineated through their rRNA genes do not yet have any cultured representatives. Community rRNA sequencing and phylogenetic analysis, in itself, is not considered metagenomics (because it focuses on only one gene, not entire genomes), but it can be a useful preliminary step in a metagenomics project because it provides a phylogenetic assessment of the diversity of a community.
Microbial Diversity and Variation Have No Limits
When genetic information from macroscopic organisms (animals or plants) is organized into phylogenetic trees to examine how they are related to one another, one can assume that all the individuals of a given species have virtually identical genomes. For example, the genomes of humans differ from one another by only 0.1%. In contrast, microbial phylotyping coupled with genome sequencing has shown that even if culturability ceased to be a problem, diversity will always be a challenge; indeed, it is a greater challenge than might have been imagined. Hundreds of thousands or even millions would be too low an estimate of the number of genomes that would have to be sequenced in any kind of whole-genome-based metagenomics program. This is due in part to the large numbers of species of microbes in most environments. But also it reflects genomic diversity within what scientists had been calling species. Almost all phylotyping surveys of almost all environments yield not a single phylotype for each likely microbial species contributor to community dynamics, but dozens or hundreds of very close but unquestionably nonidentical phylotypes that form microdiverse clusters (see Figure 1-2). In addition to differing slightly in the sequences of the marker genes used in phylotyping, these organisms—supposedly members
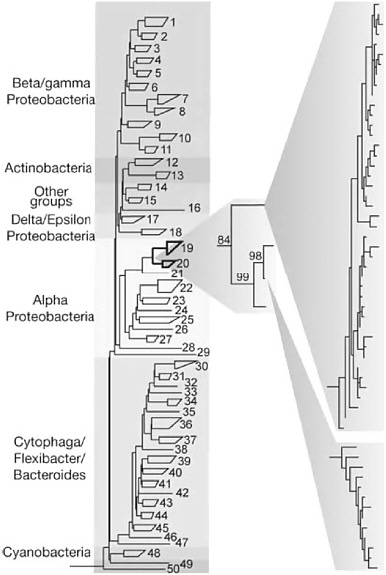
FIGURE 1-2 Microdiversity of environmental 16S rRNA sequences. PCR-amplified 16S rRNA gene sequences from an environmental DNA sample, showing a pattern of clustering often interpreted to be indicative of species divisions. Reprinted by permission from Macmillan Publishers Ltd: Nature 430:551, copyright 2004.
of the same species—differ substantially (by up to 30%) in the genes that their genomes contain.
In a recent survey of the diversity and genome sizes (gene contents) of strains of the environmental bacterium Vibrio splendidus, Polz and coworkers documented astonishing diversity (up to 25% difference in apparent gene content) in a small area (a single site off a beach in Massachusetts) (Thompson et al. 2005). They were forced to conclude that “this group consists of at least a thousand distinct genotypes, each occurring at extremely low environmental concentrations (on average less than one cell per milliliter).” All this means that no single collection of genes can be said to be “the V. splendidus genome” or “the E. coli genome” or indeed the genome of almost any designated bacterial or archaeal species, and no amount of complete genome sequencing will be enough to map the genomic diversity of the microbial world. Perhaps the biggest challenge faced by microbial ecology as a science today is to understand the ecological significance of such phylotypic microdiversity and genomic variability, and this challenge cannot be met with a traditional genomic approach.
METAGENOMICS OFFERS A WAY FORWARD
The pure culture paradigm has not only limited what microbiologists have studied; it has also limited how they think about microbes. Microbes have been studied as sovereign entities and examined only for their responses to the simple chemicals that can be added to their media. We know little about their behavior as partners in the strategic alliances that are metabolic consortia, such as the consortia that decontaminate drinking water or that make up the complex structured biofilms that keep dental hygienists busy. The invisible members of a microbial community can differ vastly in their biochemical activities and interactions, not only between species but also within species. Phylotyping gives some reliable information about “Who is there?” but because of within-species genomic diversity, only imperfect guesses as to “What are they doing?” Metagenomic methods, which will be discussed later, go a long way toward answering the second question. In the end, it may be possible to view ecosystems themselves as biological units with their own genetic repertoires and to sidestep consideration of individual species. Then, both “Who is there?” and “What are they doing?” could be replaced with “What is being done by the community?”
Such understanding can be achieved only with methods that go beyond the pure-culture and single-whole-genome approaches that have dominated microbial genomics. We must move directly to the genes, to defining environments by the potential and realized biochemical and geochemical activities of the genes that are there, and the complex patterns of interactions within and between cells that regulate their responses to changes in their
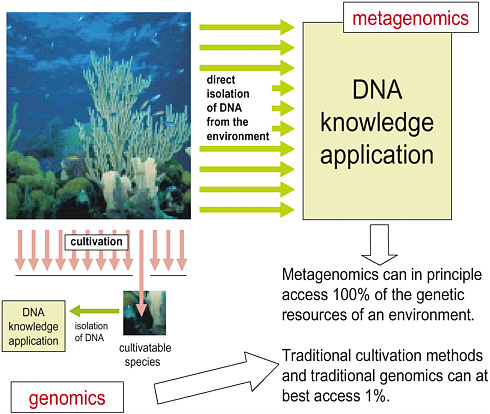
FIGURE 1-3 How metagenomics differs from microbial genomics. Image provided by W. Ford Doolittle.
physical and biological surroundings. We must do this while recognizing that—except in restricted environments and specialized consortia with limited numbers of genetically homogeneous constituents—we will be dealing with enormous amounts of data that will represent an incomplete sampling of the genetic diversity present. In short, we must adopt the methods of metagenomics (see Figure 1-3).
Pioneering steps in this direction, which illustrate the character and range of such methods, are described later in this report; but in metagenomics, necessity not only is the mother of invention but will be the grandmother of a paradigm shift. It will refocus us one level higher in the biological hierarchy (molecules, cells, organisms, species, populations, communities, the biosphere). It will shift the emphasis from individuals to interactions, from parts to processes—a change that would be timely and highly desirable even if it were not also technologically necessary.
Not coincidentally, this shift will parallel the new focus of organismal genomics on interactions between cellular components and how they are coordinated within the complex systems called organisms. This new focus is called systems biology. Metagenomics will be the systems biology of the biosphere.
Metagenomics provides a means for studying microbial communities on their own “turf.” Complex ecological interactions—including lateral gene transfer, phage-host dynamics, and metabolic complementation—can now be studied with the lens of metagenomics. Community composition, function, and dynamics can now be measured and modeled in the environment with universal microbial-community genomic approaches. These approaches have the potential to provide new insights into the environmentally relevant microbial communities and activities that control matter and energy flux on Earth. With such information in hand, it will become possible to interpret the interplay between natural cycles and human activities that together shape the future of the planet.
METAGENOMICS CAN CONTRIBUTE TO ADVANCES IN MANY FIELDS
Metagenomics offers a means of solving practical problems facing humanity. Cracking the secrets of some of Earth’s countless microbial communities will reveal ways to meet myriad challenges in biomedicine, agriculture, and environmental stewardship. These are among the most important potential contributions:
-
Earth Sciences: the development of genome-based microbial ecosystem models to describe and predict global environmental processes, change, and sustainability.
-
Life Sciences: the advancement of new theory and predictive capabilities in community-based microbial biology, ecology, and evolution.
-
Biomedical Sciences: the description, on a global scale, of the role of the human microbiome (the collective genome of our symbionts) in health and disease in individuals and populations, and the development of novel diagnostic and treatment strategies based on this knowledge.
-
Bioenergy: the development of microbial systems and processes for new bioenergy resources that will be more economical and environmentally sustainable and less vulnerable to disruption by world politics.
-
Bioremediation: the development of tools for monitoring environmental damage at all levels (from climate change to leaking gas-storage tanks) and microbe-based (green) methods for restoring healthy ecosystems.
-
Biotechnology: the identification and exploitation of the remarkably versatile and diverse biosynthetic capacities of microbial communities to generate beneficial industrial, food, and health products.
-
Agriculture: the development of more effective and comprehensive methods for early detection of threats to food production (crop and animal diseases) and food safety (monitoring and early detection of dangerous microbial contaminants) and the development of management practices that maximize the beneficial attributes of microbial communities in and around domestic plants and animals.
-
Biodefense and Microbial Forensics: the development of more effective vaccines and therapeutics against potential bioterror agents, the deployment of genomic biosensors to monitor microbial ecosystems for known and potential pathogens, and the ability to precisely identify and characterize microbes that have played a role in war, terrorism, and crime events, thus contributing to discovering the source of the microbes and the party responsible for their use.