4
Alternatives to Terran Biochemistry in Water
All that we know about terran life suggests that it exists in water at nearly neutral internal pH and at a range of temperatures that permits water to be in the liquid state. That range includes temperatures higher than 100°C, the boiling point of pure water at sea level, if pressures higher than 1 atm prevent boiling.
The committee found that it has proved difficult to find extant life in environments that do not contain liquid water, although searches for life in very dry, very cold, and very hot places continue. That suggests that water is key to the life that we know. Furthermore, the exploitation of the polar versus nonpolar, hydrophilic versus hydrophobic, and water-soluble versus water-insoluble dichotomies in the construction of terran metabolites, macromolecules, and supramolecular structures appears to be an elegant way to solve problems that are presumably presented to life generally.
The evident compatibility between terran biochemistry and water makes it difficult to distinguish cause and effect. First, the global features of terran biochemistry may indicate that the biochemistry we know is the only chemical solution to particular challenges posed by life in general. Second, the compatibility may reflect that terran biochemistry is the only chemical approach to particular challenges posed by life in water. Third, it is possible that alternative chemistry could support life, even in water, but did not originate on early Earth or did not survive in competition with the life we know. Finally, the observed facts might indicate nothing about what environments might support life but only that all life on Earth is related by a common ancestry; that is, the compatibility might reflect neither the optimal nor the universal, but simply a shared historical accident.
If the first is the case, then our observations of terran life should determine our view of life generally. In particular, if features of terran biochemistry are presumed to be the only possible solutions to problems that must be solved for life to exist and if those solutions are possible only in water, we would have to conclude that life can exist only in water. The “follow the water” strategy would thus be uniquely suited for the search for life in the cosmos.
If the second is the case, we would be called on to creatively define chemistries that might support life in nonaqueous solvents. They might be used to support NASA missions to locales, such as Titan and Venus, where liquids other than water are abundant.
If the global structures of terran life reflect origins, we must turn to models of early Earth and consider its chemistry in an effort to guess the potential for life in the solar system. Other planets and moons in our solar system have had histories different from Earth’s. We would be called on to creatively define chemistries that might have originated in the early histories of those other bodies.
Last, if the global features of terran life reflect nothing more than the common ancestry of the life that we know, we could infer little from a study of life on Earth about the nature of alien life that does not share an ancestry. Indeed, as Cleland and Copley have recently discussed,1 it is conceivable that Earth harbors yet undiscovered forms of life that are not related by common ancestry to the life that we know, have quite different biochemistry, and may have been overlooked for precisely that reason.
4.1
SYNTHETIC BIOLOGY AS A STRATEGY FOR UNDERSTANDING ALTERNATIVES TO TERRAN BIOMOLECULES
The goal of synthetic biology is to create a more comprehensive understanding of life by integrating different areas of research, such as engineering, physics, and chemistry, so as to design and construct novel biological and biochemical functional systems. Since biological systems are composed of organic compounds, synthetic biology has become more connected to synthetic chemistry (the shift from studying nature’s chemistry to the design and synthesis of new chemistry). The result is novel biochemistry. One area of current emphasis in synthetic biology that is germane to this report is the design and synthesis of new biochemicals that can lead to the synthesis of novel but functional structural, informational, and catalytic biochemical systems. Significant progress has also been made using synthetic biology approaches to origin of life studies and particularly in designing biochemical systems that might better reflect early stages in the synthesis of information macromolecules, replicators, and cell-like structures. One of the goals is to find alternative biochemical systems that undergo evolution.
The committee considered a variety of approaches to determine whether the biochemical structures found in terran life are unique. One was derived from synthetic organic chemistry and is sometimes referred to as synthetic biology.2,3 Much of contemporary biological research deconstructs living systems, but the ability of chemists to synthesize new forms of matter (i.e., new arrangements of atoms in new molecules) offers an alternative approach, especially if the aim is to ask whether alternative chemistries can support biomolecular function. It is possible for chemists to synthesize alternative chemistries, to ask Why not? and What if? questions about biomolecular structure, and to determine whether the alternative structures might function as alternative genetic molecules, membrane components, catalytic species, or metabolites. That is directly related to the question, Are the biomolecular structures that we know in terran life the only structures that can possibly meet the functional demands of living systems?
4.1.1
Terran Nucleic Acids Are Not the Only Structures That Can Support Genetic-like Behavior
In some cases, direct experimental evidence shows that the molecules found globally in terran biochemistry are not the only structures that can perform the functions that they perform in the life that we know. For example, in terran DNA, the Watson-Crick nucleobase pairs obey two rules of complementarity: size complementarity (large purines pair with small pyrimidines) and hydrogen-bonding complementarity (hydrogen-bond donors from one nucleobase pair with hydrogen-bond acceptors from the other). Those rules enable the specificity that gives rise to the simple rules for base pairing (A pairs with T, G pairs with C) that underlie terran genetics and molecular biology.
It is possible through synthesis to show that the DNA alphabet is not limited to the four standard nucleotides known in terran DNA.4,5 Rather, 12 nucleobases forming six base pairs joined by mutually exclusive hydrogen-bonding patterns are within the geometry of the Watson-Crick base pair. Figure 4.1 shows some of the standard and nonstandard nucleobase pairs and the nomenclature to designate them. Those nucleobase analogues presenting nonstandard hydrogen-bonding patterns are part of an artificially expanded genetic information system (AEGIS).
On the basis of simple binding studies, it is clear that the AEGIS components work as well as the natural nucleotides. Each nucleotide pairs with its partner, and mismatches between the 12 destabilize the duplex about as much as mismatches between the standard four nucleotides. Indeed, the specificity of the synthetic biological genetic molecules is so good that they are incorporated into diagnostic tools that first received Food and Drug Administration approval in 2002. Each year, artificial genetic molecules exploiting an expanded genetic alphabet improve the health care of some 400,000 patients infected with HIV or the hepatitis B or hepatitis C virus.
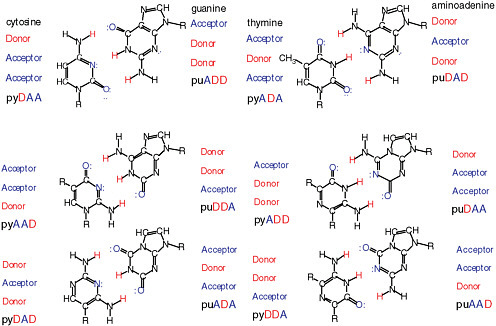
FIGURE 4.1 Twelve possible nucleobases in a DNA- or RNA-based “alphabet” that can form specific base pairs within the constraints of the Watson-Crick base-pair geometry and artificially expanded genetic information system (AEGIS). Pyrimidine base analogues are designated by “py,” purine by “pu.” The upper case letters following the designation indicate the hydrogen-bonding pattern of acceptor (A) and donor (D) groups. Thus, the standard nucleobase cytosine is pyDAA, and guanosine is puADD.
Artificial chemical systems capable of Darwinian evolution have also been prepared from artificial laboratory genetic systems. Such systems were created in the laboratory by using an artificial DNA that contained six nucleotide letters rather than the four in standard terran DNA.6,7 These were chosen from the structures shown in Figure 4.1. The artificial systems can support the basic elements of Darwinian evolution (reproduction, mutation, and inheritance of mutated forms) even if the enzymes that support the evolution of artificial genetic systems are the natural terran enzymes that have evolved for billions of years to handle standard nucleobases.
Standard tools to detect genetic molecules are designed to detect standard DNA, containing the four nucleosides adenosine, guanosine, cytidine and thymidine. They cannot detect DNA built from nonstandard building blocks. This creates a special challenge for those seeking to design instruments to detect molecules on other bodies, such as Mars or Europa. Indeed, it is even possible that a form of life based on nonstandard genetic molecules might be present on Earth, undetected by standard tools that detect standard DNA.
Similar efforts in synthetic biology have shown that the ribose and deoxyribose sugars are not unique solutions to the need for a scaffolding in a linear genetic biopolymer. Starting in the 1980s, researchers in groups across the world—including the Benner and Eschenmoser groups in Switzerland, the Herdewijn group in Belgium, the Wengel group in Denmark, and most recently the Krishnamurthy group in La Jolla—have joined researchers in industry attempting to make nucleic acid analogs that might serve as drugs to determine what other types of sugars
might function in the backbone of nucleic acids. Summaries of the work can be found in Freier and Altmann,8 Eschenmoser,9 and Benner.10
The results were surprising. Replacing the ribose with another sugar generally gave a DNA analogue that formed less stable duplexes. In many cases, the replacement sugar generally diminished the stability of the duplex. For example, a variety of hexopyranosyl-(6′-4′) oligonucleotide analogues of RNA derived from the hexose sugars known as allose, altrose, and glucose displayed Watson-Crick nucleobase pairing that was inferior to that displayed by RNA with respect to both pairing strength and pairing mode specificity.
Other backbone replacements displayed pairing strengths comparable with that seen in RNA but did not cross-bind with natural RNA and DNA. The exceptional cases are the locked nucleic acids of Wengel et al.11 and the threose nucleic acid analogues of Krishnamurthy and Eschenmoser.12
4.1.2
Terran Amino Acids Are Not the Only Structures That Can Be Incorporated into Proteins
An analogous series of experiments has shown that unnatural amino acids function in proteins as well as the standard 20 amino acids found globally in terran proteins. The natural ribosome found globally on Earth was able to incorporate these into proteins. In that way, synthetic biologists have expanded the amino acid repertoire of proteins.13 The experiments showed no reason to exclude alternative sets of amino acids from hypothetical proteins in hypothetical alien life forms.
The synthetic genetic system has even been coupled to unnatural protein synthesis. Adding extra letters to the genetic alphabet has been shown to increase the number of triplet codons that are accessible to a messenger RNA. In 1993, Bain et al. showed that additional triplet codons made possible by extra nucleobases and delivered by synthetic biological efforts could encode extra amino acids.14 That result was obtained with the terran ribosome.
4.1.3
Implications of Synthetic Biology for Our View of the Universality of Global Terran Proteins and Nucleic Acids
Work with synthetic biology makes it clear that the core set of nucleotides and the core set of amino acids found in all terran life inspected to date are not the only nucleotides and amino acids that can function in genetic and catalytic systems. Indeed, synthetic alternatives to standard terran biochemistry function in the terran machinery that has evolved to handle the core sets.
It is not clear whether alterative nucleotides and amino acids were available prebiotically. Studies of the Murchison meteorite (see Chapter 5) have revealed a large number of nonstandard amino acids; many are accepted by terran machinery. It is clear that advanced organisms, such as those found in modern life, are fully capable of synthesizing extra nucleotides for RNA and DNA and extra amino acids for proteins.
Those facts mean that alternative nucleotides and amino acids could have been accessible to terran life. The observation that terran life did not exploit them can be explained in various ways. It may be that some terran life forms do exploit alternative nucleotides or amino acids but have not yet been discovered; indeed, hypothetical terran life that uses alternative nucleotides or amino acids would not necessarily be detected by probes designed to look for terran life. It is conceivable that some terran life forms did use alternative nucleotides or amino acids but were less fit than life forms that used the standard set and thus did not survive in Darwinian competition for us to discover them. Alternatively, we might assume that chemical processes needed to make the extra nucleotides or amino acids did not happen in the 4 billion years of life on Earth.
Regardless of which explanation is favored, we have no immediate grounds to exclude alternative nucleotides or amino acids in nonterran life if we encounter it. Somewhat complex arguments based on the chemistry of some of the alternative nucleotides were proposed in the committee’s discussion to disfavor them relative to the standard four nucleotides, but it is not clear that equally compelling counterarguments would not be offered if human-like genetics exploited the nonstandard alternatives.
4.2
WHAT FEATURES OF TERRAN GENETIC MOLECULES MIGHT BE UNIVERSAL IN GENETIC MOLECULES ACTING IN WATER?
Synthetic biological experiments are available to help in identifying universal structures in terran biomolecules.
4.2.1
A Repeating Charge May Be Universal in Genetic Polymers in Water
As discussed above, it is important for metabolites to be soluble in water if life is adapted to water. RNA and DNA, by virtue of their repeating charge, are very soluble in water.15 The DNA duplex in which a polyanion binds another polyanion, appears to disregard Coulomb’s law, however. One might think (indeed, many have thought16,17) that the duplex would be more stable if one strand were uncharged, or polycationic.18
Synthetic biologists have undertaken many efforts to create nonionic analogues of DNA and RNA. For example, replacing the anionic phosphate diester linker with the uncharged dimethylenesulfone linker generated DNA and RNA analogues that were rough isosteres of the phosphate analogue (Figure 4.2).19 Short sulfone-linked DNA analogues (SNAs) displayed molecular recognition of the Watson-Crick type. In longer species, however, the loss of the repeating charge damaged the capacity for rule-based molecular recognition. Furthermore, SNAs differing by only one nucleobase displayed different levels of solubility, aggregation, folding, and chemical reactivity, a characteristic displayed by many polyfunctional molecules. That implies that SNAs of any length could not support genetics.
Those results suggest three hypotheses for why charged phosphate linkages are important for molecular recognition in DNA. First, the repeating charges in the backbone force interstrand interactions away from the backbone, causing the strands to make contact at the Watson-Crick edge of the heterocycles (Figure 4.3). Without the polyanionic backbone, interstrand contacts can be anywhere.20
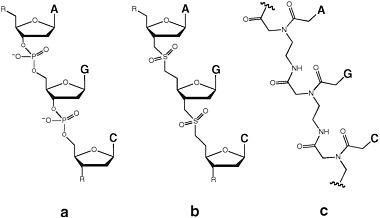
FIGURE 4.2 Replacing the anionic phosphate diester linker (a) with the uncharged dimethylenesulfone linker (b) generates DNA and RNA analogues that are rough isosteres of the phosphate analogue. (c) PNA is a DNA or RNA mimic in which the phosphate-sugar backbone has been replaced with uncharged N-(2-aminoethyl)glycine linkages in which the nucleobases are attached through methylene carbonyl linkages to the glycine amino group. The markings at the ends of the chain indicate that the chain continues.
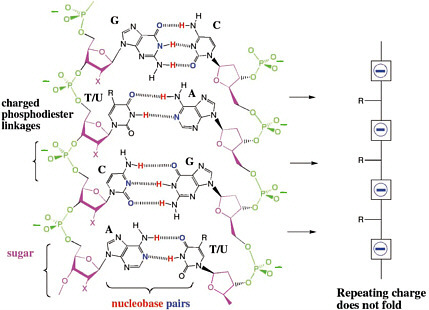
FIGURE 4.3 The repeating charges in the backbone force interstrand interactions away from the backbone, causing the strands to make contact at the Watson-Crick edge of the heterocycles. Without the polyanionic backbone, interstrand contacts can be anywhere. “X” is H in DNA and OH in RNA.
Furthermore, the repeating charges in the backbone keep DNA strands from folding. A flexible polyanion is more likely to adopt an extended conformation suitable for templating than is a flexible neutral polymer, which is more likely to fold.21 The Flory dimension of a polyelectrolyte (like DNA), other things being similar, is much larger than that of a biopolymer with, for example, a repeating dipole.22 As noted above, replication is not sufficient for a genetic molecule to support Darwinian evolution. The Darwinian system must also generate inexact replicates—descendants whose chemical structures are different from those of their parents—and the differences must then be replicable themselves. Self-replicating systems are well known in chemistry, but ones that generate inexact copies, with the inexactness itself replicable, are not.23 Indeed, small changes in molecular structure often lead to large changes in the physical properties of a system. That means that inexact replicates need not retain the general physicochemical properties of their ancestors, especially properties that are essential for replication.
In DNA, the polyanionic backbone dominates the physical properties of DNA. Replacing one nucleobase with another, therefore, has only a second-order effect on the physical behavior of the molecule. That allows nucleobases to be replaced during Darwinian evolution without losing properties essential for replication. Therefore, a repeating charge may be a universal structural feature of any genetic molecule that supports Darwinian evolution in water. Polycationic backbones may be as satisfactory as polyanionic backbones, however. Thus, if NASA missions detect life in water on other planets, its genetic system is likely to be based on polyanionic or polycationic backbones even if its nucleobases differ from those found on Earth. That structural feature can be easily detected with simple analytical devices.
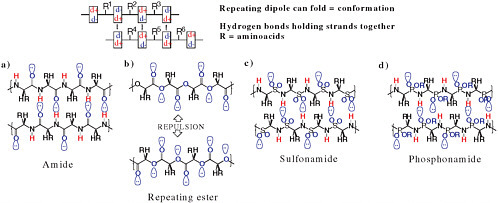
FIGURE 4.4 Some alternative backbone biopolymers with and without repeating dipoles. Red hydrogens can serve as hydrogen-bond donors because of their partial positive charge. Hydrogen-bond acceptors, atoms bearing a partial negative charge, are shown in blue. The polyesters (b) do not have a repeating dipole and therefore cannot form self-structure as easily. The lobes on the oxygen atoms represent unshared pairs of electrons.
4.2.2
A Repeating Dipole May Be Universal in Polymeric Catalytic Molecules in Water
As noted above, the specific 20 amino acids that are common in standard terran proteins need not be universal. But what about the amide (the unit that links amino acids in a polypeptide, a short protein) backbone? Unlike DNA and RNA, in which the repeating element is a monopole (a charge), a polypeptide chain has a repeating element that is a dipole. That is ideal for folding: The positive end of one dipole has a favorable energy of interaction with the negative end of another dipole. In contrast, a biopolymer based on a repeating ester linkage would not fold via backbone-backbone interactions (Figure 4.4). Folding is almost certainly required for efficient catalysis, so one might expect repeating dipoles, as found in polyamides, to be found throughout the galaxy in biospheres that are based on water.
Exobiological research offers reasons to expect that amino acids might be universal. Amino acids appear to be products of prebiotic synthesis. They are famously the products of the Miller process by which methane, ammonia, and water were subjected to electrical discharge. And amino acids are found in carbonaceous meteorites both as alpha amino acids that are standard in terran life and as the nonstandard alpha-methylamino acids found by Cronin and Pizzarello.24
The uniqueness of bonding between carbon, oxygen, and nitrogen makes it difficult to conceive of an alternative backbone that contains a repeating dipole. If one selects from the third row of the periodic table, repeating sulfonamide would be one possibility.25 Repeating phosphonamides,in which the negative charge is blocked, would be another.26
Thus, whereas a repeating charge is important for a genetic molecule in water, a repeating dipole is important for the folding of a catalytic molecule, such as a protein enzyme, in water.
4.3
IS WATER UNIQUELY SUITED AS A BIOSOLVENT?
It is clear from abundant experiments in organic chemistry that assembly involving the interaction of dipoles is a good way to create supramolecular structure in many solvents that are not water. Indeed, dipolar interactions are generally stronger in nonaqueous, nonpolar solvents than they are in water. In water, the solvent molecules
interact with dipoles, competing with supramolecular assembly. Thus, even if repeating dipoles are a universal requirement for catalytic molecules, this does not exclude nonaqueous liquids as biosolvents.
That is not the case for a repeating charge. A repeating charge generally confers water solubility on a polymer, but it nearly always makes the polymer insoluble in nonpolar solvents. That means that the repeating charge cannot be used to manage the difficult combination of Darwinian evolution and general chemistry. As noted above, small changes in molecular structure generally create large, often chaotic, changes in molecular behavior. That is not tolerable in an encoding biopolymer that must support Darwinian evolution, in which case, the physical properties of the molecule must remain relatively constant when the informational content changes. The repeating charge in RNA and DNA allows that. The repeating charge (the monopole) dominates the physical properties so greatly that one can change the nucleobases, including their hydrogen-bonding dipolar units, with very little effect on the physical properties of the biopolymer. That strategy cannot be exploited in nonpolar solvents in which a polycharged polymer is insoluble.
Thus, if we regard the repeating charge as the only approach to enabling Darwinian evolution of an encoding genetic molecule, molecular genetics is possible only in polar solvents, such as water.
4.4
OPPORTUNITIES FOR RESEARCH
It is clear that the potential diversity of backbones, recognition elements, and linker groups that are structurally quite different from those known so far, but may nevertheless support genetic-like behavior, has only begun to be explored. There are abundant opportunities for research following the paradigm set forth in the emerging field of synthetic biology. The application of synthetic biology to construct biopolymers that could be suitable to planetary environments radically different from Earth’s, such as at Titan, will require NASA missions that emphasize measurements of environmental properties that narrow the options for biopolymers.
The principal challenge in designing research programs in this field is in the number of possible alternative structures that can be drawn on paper. Chemical theory only imperfectly constrains the behavior of molecules whose structures can be drawn and therefore only modestly constrains the array of structures that can be studied. Clearly, high priority should be placed on exploring polyelectrolytes as alternative genetic materials and in developing instruments to detect these that might fly in the future to locales in the solar system where liquid water may be present. Polyelectrolytes detected there would be considered strong biosignatures, especially if they are thermodynamically unstable in their environment.
The rules for potential nucleobases have become better defined by synthetic biologists.27 It is appropriate to couple design to experimental and theoretical considerations related to heterocycles that might be generated under prebiotic conditions. NASA should place high priority on Earth-based experiments that define what heterocycles might have been formed, especially in cometary bodies and on early Earth.
The same is appropriate for the exploration of potential alternative sugars. The prebiotic origin of sugars is still poorly defined. Recommended research activities focus on constraining possible sugar backbones in alien genetic material to the ones that are more plausibly generated under prebiotic conditions. Synthetic biologists are advised to explore alternative genetic systems that combine the heterocycles and sugars.
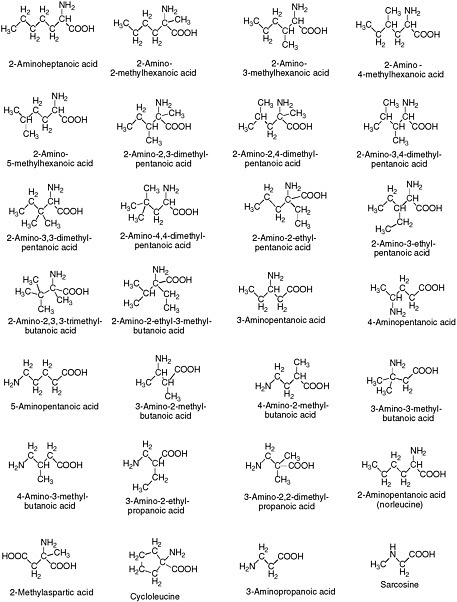
Alternative amino acids are easily conceived, both in theory and from experiment. Many alternative amino acids are known in meteorites (Figure 4.5). Several classes, including alpha-methylamino acids, form secondary structures more easily than do standard terran amino acids. Little is known, however, about the ability of polymers built from them to support catalysis.
4.5
REFERENCES
1. Cleland, C., and Copley, S. 2006. The possibility of alternative microbial life on Earth. Int. J. Astrobiology 4(3):165-173.
2 Ball, P. 2004. Starting from scratch. Nature 431:624-626.
3 Benner, S.A., and Sismour, A.M. 2005. Synthetic biology. Nature Rev. Genetics 6:533-543.
4 Switzer, C.Y., Moroney, S.E., and Benner, S.A. 1989. Enzymatic incorporation of a new base pair into DNA and RNA. J. Am. Chem. Soc. 111:8322-8323.
5 Piccirilli, J.A., Krauch, T., Moroney, S.E., and Benner, S.A. 1990. Extending the genetic alphabet. Enzymatic incorporation of a new base pair into DNA and RNA. Nature 343:33-37.
6 Sismour, A.M., Lutz, S., Park, J.-H., Lutz, M.J., Boyer, P.L., Hughes, S.H., and Benner, S.A. 2004. PCR amplification of DNA containing non-standard base pairs by variants of reverse transcriptase from human immunodeficiency virus-1. Nucl. Acids. Res. 32:728-735.
7 Sismour, A.M., and Benner, S.A. 2005. The use of thymidine analogs to improve the replication of an extra DNA base pair: A synthetic biological system. Nucl. Acids Res. 33:5640-5646.
8 Freier, S.M., and Altmann, K.H. 1997. Nucleic acid analogs. Nucl. Acids Res. 25:4429-4443.
9 Eschenmoser, A. 1999. Chemical etiology of nucleic acid structure. Science 284:2118-2124.
10 Benner, S.A. 2004. Understanding nucleic acids using synthetic chemistry. Accounts Chem. Res. 37:784-797.
11 Wengel, J., Koshkin, A., Singh, S.K., Nielsen, P., Meldgaard, M., Rajwanshi, V.K., Kumar, R., Skouv, J., Nielsen, C.B., Jacobsen, J.P., Jacobsen, N., and Olsen, C.E. 1999. LNA (locked nucleic acid). Nucleosides Nucleotides 18:1365-1370.
12 Schöning, K.U., Scholz, P., Guntha, S., Wu, X., Krishnamurthy, R., and Eschenmoser, A. 2000. Chemical etiology of nucleic acid structure. The α-threofuranosyl-3′→2′ oligonucleotide system. Science 290:1347-1351.
13 Chin, J.W., Cropp, T.A., Anderson, J.C., Mukherji, M., Zhang, Z.W., and Schultz, P.G. 2003. An expanded eukaryotic genetic code. Science 301:964-967.
14 Bain, J.D., Chamberlin, A.R., Switzer, C.Y., and Benner, S.A. 1992. Ribosome-mediated incorporation of non-standard amino acids into a peptide through expansion of the genetic code. Nature 356:537-539.
15 Westheimer, F.H. 1987. Why nature chose phosphates. Science 235:1173-1178.
16 Jayaraman, K., McParland, K., Miller, P., and Tso, P. O. P. 1981. Non-ionic oligonucleoside methylphosphonates. 4. Selective inhibition of Escherichia coli protein synthesis and growth by non-ionic oligonucleotides complementary to the 3′ end of 16S ribosomal-RNA. Proc. Natl. Acad. Sci. U.S.A. 78:1537-1541.
17 Miller, P.S., McParland, K.B., Jayaraman, K., Tso, P.O.P. 1981. Biochemical and biological effects of nonionic nucleic acid methylphosphonates. Biochemistry 20:1874-1880.
18 Reddy, P.M., and Bruice, T.C. 2003. Solid-phase synthesis of positively charged deoxynucleic guanidine (DNG) oligonucleotide mixed sequences. Bioorg. Med. Chem. Lett. 13:1281-1285.
19 Benner, S.A., and Hutter, D. 2002. Phosphates, DNA, and the search for nonterran life. A second generation model for genetic molecules. Bioorg. Chem. 30:62-80.
20 Steinbeck, C., and Richert, C. 1998. The role of ionic backbones in RNA structure: An unusually stable non-Watson-Crick duplex of a nonionic analog in an apolar medium. J. Am. Chem. Soc. 120:11576-11580.
21 Flory, P.J. 1953. Principles of Polymer Chemistry. Cornell University Press, Ithaca, N.Y.
22 Brant, D.A., and Flory, P.J., 1965. The configuration of random polypeptide chains. J. Am. Chem. Soc. 87:2788.
23 Lee, D.H., Granja, J.R., Martinez, J.A., Severin, K., and Ghadiri, M.R. 1996. A self replicating peptide. Nature 382:525-528.
24 Cronin, J.R., and Pizzarello, S. 1986. Amino acids of the Murchison meteorite. III. Seven carbon acyclic primary alpha-amino alkanoic acids. Geochim. Cosmochim. Acta 50:2419-2427.
25 Ahn, J.-M., Boyle, N.A., MacDonald, M.T., and Janda, K.D. 2002. Peptidomimetics and peptide backbone modifications. Mini. Rev. Med. Chem. 2:463-473.
26 Yamauchi, K., Mitsuda, Y., and Kinoshita, M. 1975. Peptides containing aminophosphonic acids 3. Synthesis of tripeptide containing aminomethylphosphonic acid. Bull. Chem. Soc. Jpn. 48:3285-3286.
27 Geyer, C.R., Battersby, T.R., and Benner, S.A. 2003. Nucleobase pairing in expanded Watson-Crick-like genetic information systems. The nucleobases. Structure 11:1485-1498.