THE EMERGENCE OF CRYPTOCOCCUS GATTII IN BRITISH COLUMBIA AND THE PACIFIC NORTHWEST1
Karen H. Bartlett, Sarah E. Kidd, and James W. Kronstad2
An unprecedented emergence of cryptococcal infections in animals and otherwise healthy humans was recognized in 1999 on the east coast of Vancouver Island, British Columbia. Unexpectedly, these infections were caused by Cryptococcus gattii, a species closely related to the AIDS-associated fungal pathogen Cryptococcus neoformans. Human cases have continued over the past 8 years and now total approximately 170 with eight deaths. Extensive environmental
__________________
1 Reprinted with kind permission from Springer Science+Business Media: Current Infectious Diseases Reports, The emergence of Cryptococcus gattii in British Columbia and the Pacific Northwest, 10, 2008, p. 108–115, Karen H. Bartlett, Sarah E. Kidd, and James W. Kronstad.
Current Infectious Disease Reports 2008, 10:58-65
Current Medicine Group LLC ISSN 1523-3847
Copyright © 2008 by Current Medicine Group LLC
Papers of particular interest, published recently, have been highlighted as:
+ Of importance
++ Of major importance
2Karen H. Bartlett, PhD, Sarah E. Kidd, PhD, and James W. Kronstad, PhD. Corresponding author: James W. Kronstad, PhD, The Michael Smith Laboratories, University of British Columbia, 2185 East Mall, Vancouver, BC, V6T 1Z4, Canada. Email: kronstad@interchange.ubc.ca.
sampling, coupled with detailed molecular typing of isolates, revealed areas of permanent and transient colonization with primarily three genotypes of the fungus. C. gattii was found in air, soil, water, and in association with numerous tree species. Importantly, there is solid evidence for human-mediated dispersal of the pathogen, and C. gattii has now been detected in the environment on the mainland of British Columbia and in the Pacific Northwest. Associated animal and human cases are now being reported and further spread of the pathogen may be inevitable.
Introduction
The basidiomycetous yeast Cryptococcus neoformans has a global distribution and has achieved prominence in recent decades because of its propensity to infect immunocompromised people (Casadevall and Perfect, 1998). In fact, cryptococcosis is recognized as an AIDS-defining illness, and in the absence of highly active antiretroviral therapy, the disease is a significant cause of death in individuals with HIV infection (Bicanic and Harrison, 2005; Bicanic et al., 2005). People and animals acquire the fungus via the inhalation of desiccated yeast cells or basidiospores from environmental sources such as avian guano, soil, and trees. Pulmonary infection often results in dissemination to the central nervous system and C. neoformans is the leading cause of fungal meningitis (Casadevall and Perfect, 1998).
Isolates of C. neoformans have previously been divided into three varieties known as grubii, neoformans, and gattii and into serotypes (A–D and hybrids such as AD) defined by antigenic differences in the capsular polysaccharide that is the major virulence factor (Casadevall and Perfect, 1998). The gattii variety is now recognized as a separate species based on phenotypic and molecular traits, and mating (Kwon-Chung et al., 2002). Thus the current view is that the species C. neoformans (var grubii and neoformans) contains strains of serotypes A, D, and AD, and the distinct species C. gattii contains isolates of the B and C serotypes (Kwon-Chung and Varma, 2006). An excellent review of the differences between C. gattii and C. neoformans has been published by Sorrell (Sorrell, 2001).
Extensive surveys have been performed over the past 10 years to characterize the genotypes and distribution of C. neoformans and C. gattii isolates (Barreto de Oliveira et al., 2004; Boekhout et al., 2001; Boukhout et al., 1997; Fraser et al., 2005+; Kidd, 2003; Kidd et al., 2004 ++; Kidd et al., 2005+; Meyer et al., 1999; Meyer et al., 2003). These surveys used a variety of DNA-based typing methods to provide detailed classifications of isolates into molecular types. Thus, isolates of C. neoformans var grubii (serotype A) are represented by the VNI, VNII, and VNB (Litvintseva et al., 2006) molecular types, var neoformans (serotype D) is represented by the VNIV type, and isolates of the AD hybrid serotype are the VNIII type. Four molecular types are recognized for C. gattii isolates (designated VGI–VGIV) and further divisions within the molecular types have been identified
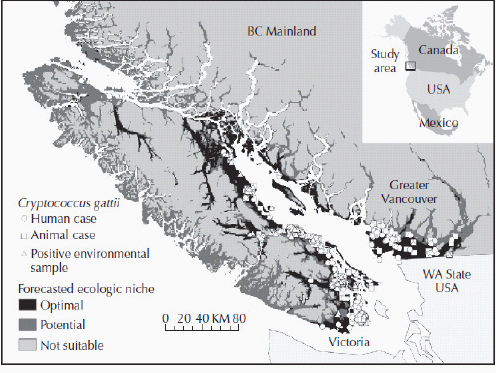
Figure A1-1 Map of the forecasted ecologic niche and region of emergence of C. gattii in British Columbia (BC). The optimal, potential, and unsuitable ecologic niches of C. gattii in BC are indicated based on biogeoclimatic data for the region (Mak, 2007). Note that the distribution of human and animal cases and the locations of positive environmental samples coincide primarily with the optimal ecologic niche. The information on human and animal cases, and environmental sampling, from Washington (WA) is not included.
(Fraser et al., 2005+; Kidd et al., 2005+; Kidd et al., 2007++). For example, VGII strains can be further classified into VGIIa and VGIIb subtypes, as well as other less-well characterized subtypes (Kidd et al., 2004; MacDougall et al., 2007++).
There is currently an intense focus on C. gattii due to the unprecedented emergence of the VGI, VGIIa, and VGIIb molecular types as primary pathogens of humans and animals on Vancouver Island in British Columbia (BC) (Kidd et al., 2004; MacDougall et al., 2007++) (Fig. A1-1). Remarkably, the majority of human cases have occurred in people without recognized immunologic defects, thus highlighting the unusual pathogenicity of C. gattii relative to C. neoformans. The purpose of this review is to summarize recent progress in the investigation of this fascinating emergence with regard to human and animal exposure, environmental colonization, isolate characterization, and the potential for further dispersal.
Overview of Veterinary and Clinical Aspects of the Emergence of C. gattii in BC
Animal sentinels played a key role in the study of the emergence of C. gattii in BC and in particular contributed to our understanding of the range of environmental niches for the pathogen. A single veterinary pathology laboratory handled clinical specimens from the majority of southern BC veterinary practices, and this allowed early detection and monitoring of C. gattii in the animal population. In addition, the BC Provincial Animal Health Branch Laboratory was able to perform necropsies on porpoises that were found stranded and dead on Vancouver Island and nearby islands, and these became index cases (Stephen et al., 2002). Beginning prior to the first documented human case in 1999 and continuing to the present, veterinary cases have been diagnosed two to three times more frequently than human cases (Lester et al., 2004); this disparity is likely an underestimate given that only those animals seen by a veterinarian are diagnosed and that infections in wildlife are not considered. The diagnosed cases have primarily been in companion animals (dogs, cats, and ferrets) but also include other domesticated species such as llamas, horses, mink, and psittacine birds (Duncan et al., 2006b; Lester et al., 2004; Stephen et al., 2002). Sampling in the environs of these animal cases has been particularly productive for identifying sources of C. gattii (Kidd et al., 2007a++; MacDougal et al., 2007++).
Unlike the colonized koalas of Australia (Krockenberger et al., 2002), no significant wild animal host or reservoir has been identified in BC. Limited surveys of wild animals were performed between 2003 and 2007 with the examination of necropsy samples of nares, lung, anus or cloacae, and brain for C. gattii. In two surveys, all fatally injured animals turned into rescue facilities were studied. In the first study, 91 animals (14 species) were examined, and only two eastern gray squirrels were positive (Duncan et al., 2006a). In the second study, only one great blue heron was found to have a pulmonary C. gattii infection of 226 animals necropsied (Bartlett, unpublished data). Additionally, 18 river otters were trapped in early spring 2007, but none showed signs of disease or colonization with C. gattii (Bartlett and Balke, unpublished data). Duncan et al. (2005b) established sentinel veterinary practices in areas known to have exposure to airborne C. gattii and found positive C. gattii cultures from nasal swabs of asymptomatic animals in 4.3% of 94 cats, 1.1% of 280 dogs, and 1.5% of 351 horses. Additionally, six cats and two dogs were found to have cryptococcal antigen titers of greater than 1:2. Of seven cats and five dogs that were selected from the asymptomatic but culture- or antigen-positive cohorts and followed over 27 months, only two cats progressed to clinical disease, suggesting that the majority of animals exposed to C. gattii may naturally clear the organism (Duncan et al., 2005a).
In the first years of recognition of both the emergence of C. gattii disease and the stability of the pathogen’s environmental niche, it appeared that all human and animal cases had some contact with Vancouver Island. MacDougall and Fyfe (MacDougall and Fyfe, 2006) were able to identify human cases of disease
with historic travel to Vancouver Island and to determine a likely incubation period (median 6–7 months) based on isolated exposure. In addition, Hoang et al. (Hoang et al., 2004) performed a retrospective chart review examining all cases of cryptococcosis identified between 1997 and 2002 at the largest teaching hospital located on the BC mainland. They discovered that there had been a sudden increase in cryptococcal cases of all origins (C. neoformans var grubii, C. n. var neoformans, C. gattii, and C. laurentii), but all C. gattii cases (3/26 charts) reported travel history to Vancouver Island (Hoang et al., 2004). The first cases of mainland-acquired C. gattii infection were identified in animals (ferret, llama, and cats) in 2003, and three cases in cats in Washington were reported in 2005. Eight off-island human cases with no travel history to an endemic area were documented (five in BC and two in Oregon) in 2004 to 2005 (MacDougall et al., 2007++). Upton et al. (Upton et al., 2007+) recently reported the first confirmed human case in Washington presenting in 2006, and the Whatcom County Public Health Department has now identified four additional cases diagnosed in 2007 (Stern, personal communication). Unlike in BC, cryptococcosis is not yet a reportable disease in Washington, although public health officials are actively soliciting case studies. The VGIIa genotype accounted for 78% of the examined veterinary cases and 87% of the human cases; all off-island veterinary cases to date had the VGIIa genotype (Bartlett, unpublished data) (MacDougall et al., 2007++).
Environmental and Dispersal Studies on Vancouver Island
Competing theories have been proposed regarding the origin of C. gattii on Vancouver Island (eg, recent introduction, long-term colonization, specific imported vectors). Suffice it to say, the colonization pattern and dispersal of the organism argues against a one-time introduction to Vancouver Island, particularly if the timeline extends only to the first animal and human cases (1998–1999). The first systematic sampling performed on Vancouver Island in 2002 mapped the colonization of C. gattii along a 200 km north-south and a 40 km east-west corridor. This study revealed that C. gattii is not homogeneously spread in the environment, with central Vancouver Island having a higher percentage of colonized trees and higher concentration of the organism in soil. The heterogeneous pockets of colonization could explain why limited-sampling strategies may miss the organism. Additionally, even though C. gattii has been found to be permanently colonized in some areas, it appears to be transiently colonized in others. The permanently colonized sites have yielded C. gattii repeatedly over the last 5 years, although transiently positive results may be due to limits of detection or failure of the organism to establish true colonization (Kidd et al., 2007a++). As well, sites that initially appeared to be negative for C. gattii have more recently yielded positive environmental samples (Bartlett, unpublished data). It has been shown that in addition to the airborne spread of propagules, wood products,
soil, water, vehicles, and shoes can act as dispersal mechanisms for the organism (Kidd et al., 2007a++). These mechanisms are consistent with the findings of a veterinary case-control study, where statistically significant risk factors for disease in cats and dogs related to soil disturbance within 10 km of cases, logging within 10 km, travel to Vancouver Island, or owner hiking within 6 months of diagnosis (Duncan et al., 2006c). Although limited environmental sampling in the San Juan Islands, Olympic Peninsula, and Oregon has not yielded C. gattii (Fraser et al., 2006; Kidd et al., 2007b++; Upton et al., 2007+). Kidd et al. (2007a++,2007b++) reported finding positive environmental samples from islands in the Georgia Strait and in northern Washington.
A rather surprising finding was that co-isolated C. gattii strains are heterogeneous. The first isolates distributed to the research community were mostly from one sampling site (central Vancouver Island) and may have unduly influenced our thinking about the composition of the BC outbreak strains (Kidd et al., 2004++; Fraser et al., 2005+; Fraser et al., 2003). In the initial analysis of the C. gattii isolates from this site, Kidd et al. (Kidd et al., 2004++) used polymerase chain reaction (PCR)-fingerprinting to demonstrate that 5% represented the VGI molecular type and 95% belonged to VGII (90% of these were VGIIa and 10% were VGIIb based upon a one polymorphic band in the PCR-fingerprint profiles). Subsequent work revealed that the composition of the C. gattii population varies in different regions where detailed molecular subtyping of isolates has been undertaken. In the southern extreme of Vancouver Island, VGIIa accounts for 91% of the isolates and the remainder are VGIIb, whereas at another site VGIIa accounts for only 66% of the isolates, with VGIIb and VGI at 19% and 15%, respectively (Bartlett and Kidd, unpublished data). Of course, the genotype frequencies are likely to be dynamic, and repeated sampling is important. Also, additional diagnostic tools sensitive enough to detect and differentiate isolates directly in environmental samples (eg, PCR on soil samples) would facilitate a better understanding of the population structure and mechanisms of spread of the organism. Already heightened awareness of changing ecologic niches has resulted in an expansion of knowledge of the environmental origins of other cryptococcal species (Filion et al., 2006).
Molecular Characterization of Isolates from BC and the Pacific Northwest
Following the initial analyses of genotype frequency described above, Kidd et al. (2005+) used multilocus sequence typing (MSLT) and gene genealogy analyses with four genes to examine patterns of molecular variation as well as population structure of the isolates from Vancouver Island compared with a worldwide sample of C. gattii strains. This work demonstrated that the VGIIa and VGIIb genotypes originally established by PCR-fingerprinting (Kidd et al., 2004++) corresponded to specific MLST profiles. Similar MLST results with additional genes were obtained by Fraser et al. (Fraser et al., 2005+). Of specific
interest from these studies was the identification of isolates from other areas of the world with identical or similar genotypes to the VGIIa (as represented by isolate A1MR265) and VGIIb (represented by isolate A1MR272) strains from Vancouver Island. For example, the VGIIa genotype was also shared by the NIH444 strain (from a patient in Seattle, ca 1971), CBS7750 (from a Eucalyptus tree in San Francisco, ca 1990) and with isolates from other parts of North America (KB10455 and KB9944) (Fraser et al., 2005+; Kidd et al., 2005+). A Brazilian clinical isolate, ICB107, differed from the VGIIa genotype at only one of 22 loci (Fraser et al., 2005+). The VGIIb genotype was also observed among environmental isolates from Australia (eg, Ram002, Ram005, WM1008), clinical isolates from Australia (eg, NT-6, NT-13), as well as a clinical isolate from Thailand (MC-S-115) (Fraser et al., 2005+; Kidd et al., 2005+). A Caribbean strain 99/473 of the VGIIb type was also found to differ at only one of 22 loci (Fraser et al., 2005+). Intriguingly, two isolates from human cases in Oregon (2004) were recently found to represent subtypes within the VGII genotype that have not identified among any other strains to date (MacDougall et al., 2007++).
The VGIIa and VGIIb isolates from Vancouver Island have been obtained from both clinical and environmental sources. However, the situation is more complex for strains of the VGI genotype from clinical and environmental sources. Specifically, Kidd et al. (2005+) characterized six VGI isolates from Vancouver Island and identified four different genotypes by MLST analysis. Two of these were environmental isolates with a different genotype from the clinical isolates. Thus, in contrast to the VGII types, it was not possible to establish an epidemiologic link between environmental and clinical isolates of the VGI type. However, recent analysis of further environmental VGI isolates from Vancouver Island indicated that they were highly similar to a porpoise isolate (A1MF2863), being identical at four MLST loci (Kidd and Bartlett, unpublished data). It is possible that the clinical isolates of the VGI type represent strains acquired during travel outside of Vancouver Island.
Overall, Kidd et al. (2005+) found that the Vancouver Island isolates were part of a predominately clonal population with little evidence of sexual recombination occurring between them. Fraser et al. (2005+) also presented evidence that the VGIIa and VGIIb strains from Vancouver Island were related in that they shared 14 identical loci out of the 30 examined and proposed that the genotypes represent either siblings arising from a past mating event, or that one may be the parent of the other, perhaps as the result of same-sex mating between MATα parents. Selected isolates from Vancouver Island and other parts of the world have been tested for mating competence. These studies revealed that the VGII isolates are generally fertile whereas VGI strains are not (Campbell et al., 2005; Fraser et al., 2003; Kidd et al., 2004++). In general, the ability of C. gattii isolates to mate has implications for recombination events that might generate strains with different virulence properties and environmental adaptability.
The Global Distribution of C. gattii
Prior to the emergence of C. gattii on Vancouver Island, it was commonly accepted that this species was restricted to tropical and subtropical regions of the world, and that infection was associated with exposure to Eucalyptus trees (Ellis and Pfeiffer, 1990; Kwon-Chung and Bennett, 1984; Sorrell et al., 1996). The idea of a limited geographic distribution came from a study that surveyed a worldwide collection of clinical isolates (Kwon-Chung and Bennett, 1984). This survey revealed that C. gattii was prevalent only in regions with tropical and subtropical climates (22%–50% of isolates) relative to C. neoformans (50%–71% of isolates). However, this study also reported that 13% of the strains from North America, and 3.3% of the strains from Europe were C. gattii (without reference to travel histories). More recent surveys have focused on identifying the molecular types of C. gattii found in collections from various regions. In this regard, VGI appears to be the most widely distributed type worldwide (Kidd, 2003; Meyer et al., 2003), and this type is also found most frequently among clinical and environmental isolates in Australia (Campbell et al., 2005). Strains of the VGII type are also found in parts of Australia as well as in North and South America (Fraser et al., 2005+; Kidd, 2003; Kidd et al., 2004++; Kidd et al., 2005+; Meyer et al., 2003). In a recent, large-scale study of IberoAmerican isolates, VGIII predominated, and this type has also been found in India and the United States (Kidd, 2003; Meyer et al., 2003). The VGIV type has been found in Central America and South Africa (Kidd, 2003; Meyer et al., 2003). Notably, the VGIII and VGIV types were not found in the collections from Vancouver Island suggesting that these genotypes may have a more limited distribution.
More recently, Meyer et al. (2007) have surveyed 160 VGII strains recovered globally since 1986 using PCR-fingerprinting, amplified fragment length polymorphism analysis and MLST with eight loci. This work revealed that the VGIIa genotype from Vancouver Island is also found among Brazilian isolates and that Colombian isolates are closely related. Interestingly, the majority of the latter isolates are mating type a in contrast to mating type α for the Vancouver Island strains (Escandon et al., 2006), and mating was demonstrated between the Colombian MATa strains and VGIIa MATα strains from Brazil and Vancouver Island. This work suggests that the VGIIa genotype was present in South America as early as 1986 and it sheds additional light on the potential mating interactions for VGII types of C. gattii that may be relevant for the situation on Vancouver Island.
Overall, these surveys provide an interesting view that the genotypes of C. gattii (at least for VGI and VGII) are likely to have a worldwide distribution and the concomitant potential for permanent colonization of suitable environments. This view highlights the need for more extensive environmental sampling globally to generate a detailed picture of genotype frequency over time and location. The most extensive view is now available from the work on Vancouver Island and the lessons learned from this work can be applied in other locations (Kidd
et al., 2007a++), especially with regard to the need for extensive multisource sampling over many years. The wide distribution of C. gattii genotypes should also be considered in light of recent reports that infections with this species are occurring in patients with AIDS (South Africa [Morgan et al., 2006], Southern California [Chaturvedi et al., 2005a]). Therefore, it will be important to identify the endemic areas for specific C. gattii genotypes in order to monitor human and animal disease.
Origin of the C. gattii in BC and the Pacific Northwest:
Aboriginal Species or Landed Immigrant?
It is fun to speculate about the origin of the genotypes on Vancouver Island, and this activity has consumed much energy in the research community. However, the extent of global strain dispersal has been demonstrated to be significant (Kidd et al., 2005+, Xu et al., 2000), making it difficult to accurately determine a specific origin of any given genotype. It is possible that the species has been a long-term resident of BC and that changing conditions (eg, climate or land use) or improved surveillance are responsible for the current level of awareness. Alternatively, it has been suggested that the emergence is due to the recent introduction of a particularly virulent genotype that may be well adapted to the local conditions such that large numbers of infectious cells are propagated (Fraser et al., 2005+). Although it may be difficult to garner strong evidence for a given theory, it is clear that much more information is needed about the C. gattii genotypes on Vancouver Island and worldwide and about the disease caused by C. gattii in immunocompetent hosts. Below, we discuss some of the studies that are needed to generate a more detailed view of C. gattii that may help in infection control.
Ecologic adaptability, colonization, and dispersal
The environmental sampling revealed a high level of soil colonization on Vancouver Island, and it would be interesting to examine soil persistence and competition in laboratory and field settings. These types of experiments may be relevant to addressing how the fungus becomes aerosolized and the nature of the infectious particle. An investigation of conditions required for the propagation of the infectious particles in soil/trees would also be highly relevant to understanding the factors that influence exposure of humans/animals.
It is likely that no one factor can explain the dramatic emergence of C. gattii on Vancouver Island, and there may be interplay between soil conditions, temperature, and moisture. Current weather station data are insufficient to adequately describe the microclimates in areas colonized by the pathogen. Climate oscillations driven by alternating El Niño and La Niña currents have produced both drier and wetter than normal summer conditions in BC over the last few decades. Outbreaks of another fungal disease, coccidioidomycosis, have been
shown to follow soil disruption in California (Zender and Talamantes, 2006). Data gathered from the BC environment conclusively show that C. gattii is well adapted to survive in dry, low nutrient soil and is more likely to be airborne during dry summer weather (Kidd et al., 2007a++). The stability of the colonization of soil and trees at permanently colonized sites suggests that the pathogen can effectively compete with resident soil microflora. Longer cycles of meteorology patterns and finer tools of climate measurement will be needed to understand the complex relationship of microbe, climate, and ecologic niche.
Additional sampling around the world is needed to investigate predicted favorable climate/soil/water conditions that might allow colonization by C. gattii. Mak (2007) has recently developed ecologic niche models that predict the probable extent of environmental colonization of C. gattii based on human, animal, and environmental data and climate projections for the Pacific Northwest (Fig. A1-1). Areas that may eventually be impacted include the Lower Mainland of BC with a population base of approximately 2 million people. These projections could be used by public health officials on both sides of the US-Canada border to plan strategies for risk communication and anticipated morbidity and mortality (Mak, 2007).
Clinical considerations
Perhaps the most relevant topics regarding the emergence of C. gattii have to do with identifying risk factors for people, designing ways to limit exposure, and developing effective methods to treat the infections that do occur. It is common to see statements in the literature that C. gattii is a primary pathogen that infects immunocompetent people, and that C. neoformans is an opportunistic pathogen that infects immunocompromised people. The distinction may be less clear given that C. gattii is now being found in AIDS patients and C. neoformans can infect seemingly immunocompetent people (Chaturvedi et al., 2005a; Morgan et al., 2006; Speed and Dunt, 1995). There is clearly a need for retrospective studies of patients to determine host risk factors as well as prospective case studies to determine efficacy of treatments. The number of cases continuing to occur on Vancouver Island (and among tourists [Lindberg et al., 2007]) would allow this type of investigation.
An interesting consideration in terms of exploring possible virulence differences for C. gattii versus C. neoformans is whether mouse virulence studies have relevance for human disease. For example, the strains with the VGIIa and VGIIb genotypes from Vancouver Island both cause disease in humans, but laboratory studies revealed virulence differences between the two strains tested (Fraser et al., 2005+). The more virulent strain, A1MR265, of the VGIIa genotype showed equal virulence in the mouse model to strain H99 that is representative of the most common VNI type of C. neoformans (var grubii). It is possible that these results reflect the fact that only one isolate of each genotype from Vancouver Island
was tested and the isolates selected may not be representative. It is clear, however, that strains of C. gattii show virulence differences (Kronstad, unpublished data) (Chaturvedi et al., 2005b; Fraser et al., 2005+) and that multiple isolates from Vancouver Island and worldwide collections need to be tested. The same is true for C. neoformans as demonstrated by the range of virulence detected by Clancy et al. (2006). Thus, we need to develop better models to assess differences in virulence and to explore possible differences that may be relevant to infection of immunocompetent versus immunocompromised hosts.
Applications of genomic approaches to develop a detailed understanding of C. gattii
The emergence of C. gattii provided the impetus to sequence the genomes of isolates representing the VGI (WM276) and VGIIa (A1MR265) genotypes (Michael Smith Genome Sciences Center, 2007++; The Broad Institute, 2007++). These are important resources for the next steps in characterizing the virulence of C. gattii, the genetic diversity of the species and the interactions of the fungus with the environment. One can imagine, for example, using the genomes for transcriptome and proteome studies to identify differences in expression for C. gattii relative to C. neoformans. Some of these differences may reveal factors that contribute to the primary pathogenesis of C. gattii relative to C. neoformans. The two C. gattii genomes also provide a platform for more detailed analyses of genotypes and comparative studies of genome variability. In the latter case, comparative hybridization or genome resequencing approaches can be used to study the microevolution of genomes in strains in the environment and clinical strains during passage through human and animal hosts (eg, during relapse or drug therapy). Comparative genome hybridization experiments with the VGI and VGIIa genomes have been initiated to identify genomic changes in mutants that have lost virulence and to examine genome variation in strains representing the VGI, VGIIa, and VGIIb genotypes (Kronstad, unpublished data). The declining cost of sequencing will also allow further genome-sequencing projects to provide a deeper view of genome content and variability. The more detailed information may eventually lead to the separation of the molecular types of C. gattii into distinct varieties or species.
Media Coverage of the Emergence of C. gattii
Any emerging infectious disease represents a challenge to the public health system. The system must respond to educate caregivers about appropriate interventions while balancing the message to allow the public to make informed choices. For example, the lay press recently reported concern by members of the public in Alabama where experimental plots of genetically engineered Eucalyptus trees will be grown; the fear being that C. gattii will be imported into the environment
through the Eucalypts (United Press International, 2007), even though no link to Eucalyptus was shown in the BC experience (Kidd et al., 2007a++). In an examination of press coverage of C. gattii as an emerging infectious disease agent, researchers at the University of BC Centre for Health and Environment Research found that during the period 2001 to 2006, BC newspapers carried 422 articles warning the public about West Nile Virus (although no West Nile Virus cases have been reported in BC) compared with 79 articles about C. gattii (170 human cases, eight deaths) (Nicol et al., unpublished data). The research group concluded that because West Nile Virus is a public health risk with identifiable precautionary actions in central Canada, newspapers were more likely to print stock West Nile Virus stories. C. gattii was seen to be a local phenomenon with no identifiable risk aversion strategies and to have potential economic repercussions to the areas affected and so was less reported. There also seemed to be confusion by news writers about the biology of Cryptococcus because the term “virus” seems to be better understood as a pathogen compared to “yeast” (Nicol et al., unpublished data). Similarly, some news items labeled C. gattii as an “Australian” fungus despite the body of literature cited above on the global distribution of the pathogen. Overall, these observations demonstrate that effective education of the media and the public is a critical component of the management of an emerging infectious disease.
Conclusions
A great deal has been learned about the emergence of C. gattii in BC over the past 8 years. We now have a clear picture of the environmental sources of the pathogen and mechanisms of dispersal, we have an understanding of the genotypes that are causing disease in humans and animals, and we have some information about clinical presentation and treatment. Certainly, there is a great deal more to investigate in terms of risk factors for the human population and treatment outcomes. In this regard, the situation on Vancouver Island presents an opportunity to develop a detailed view of an emerging infectious disease with regard to environmental exposure, the role of sentinel animals in monitoring risk, and the underlying factors that influence human susceptibility. This information may prove useful for other emerging diseases and provide methods to manage both the ongoing situation in BC and the apparent emergence of the disease in the Pacific Northwest.
Acknowledgments
The authors thank the members of the BC Cryptococcal Working Group (http://www.cher.ubc.ca/cryptococcus/) and the BC Centre for Disease Control (http://www.bccdc.org/) for helpful discussions and Sunny Mak for the preparation of Figure A1-1. The authors are supported in part by grants from the US
National Institute of Allergy and Infectious Disease (Dr. Kronstad, award RO1AI-053721), the Canadian Institutes of Health Research (Drs. Kronstad and Bartlett), British Columbia Lung Association (Dr. Bartlett), and WorkSafe BC (Dr. Bartlett). Dr. Kronstad is a Burroughs Wellcome Fund Scholar in Molecular Pathogenic Mycology, and Dr. Bartlett is a Michael Smith Foundation for Health Research Scholar.
References and Recommended Reading
Barreto de Oliveira MT, Boekhout T, Theelen B, et al.: Cryptococcus neoformans shows a remarkable genotypic diversity in Brazil. J Clin Microbiol 2004, 42:1356–1359.
Bicanic T, Harrison TS: Cryptococcal meningitis. Br Med Bull 2005, 72:99–118.
Bicanic T, Wood R, Bekker LG, et al.: Antiretroviral roll-out, antifungal roll-back: access to treatment for cryptococcal meningitis. Lancet Infect Dis 2005, 5:530–531.
Boekhout T, Theelen B, Diaz M, et al.: Hybrid genotypes in the pathogenic yeast Cryptococcus neoformans. Microbiol 2001, 147:891–907.
Boekhout T, van Belkum A, Leenders ACAP, et al.: Molecular typing of Cryptococcus neoformans: taxonomic and epidemiological aspects. Int J Sys Bacteriol 1997, 47:432–442.
Campbell LT, Fraser JA, Nichols CB, et al.: Clinical and environmental isolates of Cryptococcus gattii from Australia that retain sexual fecundity. Eukaryot Cell 2005, 4:1410–1419.
Casadevall A, Perfect JR: Cryptococcus neoformans. Washington, DC: American Society for Microbiology Press; 1998.
Chaturvedi S, Dyavaiah M, Larsen RA, Chaturvedi V: Cryptococcus gattii in AIDS patients, southern California. Emerg Infect Dis 2005a, 11:1686–1692.
Chaturvedi S, Ren P, Narasipura SD, Chaturvedi V: Selection of optimal host strain for molecular pathogenesis studies on Cryptococcus gattii. Mycopath 2005b, 160:207–215.
Clancy CJ, Nguyen MH, Alandoerffer R, et al.: Cryptococcus neoformans var. grubii isolates recovered from persons with AIDS demonstrate a wide range of virulence during murine meningoencephalitis that correlates with the expression of certain virulence factors. Microbiol 2006, 152:2247–2255.
Duncan C, Schwantje H, Stephen C, et al.: Cryptococcus gattii in wildlife of Vancouver Island, British Columbia, Canada. J Wildl Dis 2006a, 42:175–178.
Duncan C, Stephen C, Campbell J: Clinical characteristics and predictors of mortality for Cryptococcus gattii infection in dogs and cats of southwestern British Columbia. Can Vet J 2006b, 47:993–998.
Duncan C, Stephen C, Lester S, Bartlett KH: Follow-up study of dogs and cats with asymptomatic Cryptococcus gattii infection or nasal colonization. Med Mycol 2005a, 43:663–666.
Duncan C, Stephen C, Lester S, Bartlett KH: Sub-clinical infection and asymptomatic carriage of Cryptococcus gattii in dogs and cats during an outbreak of cryptococcosis. Med Mycol 2005b, 43:511–516.
Duncan CG, Stephen C, Campbell J: Evaluation of risk factors for Cryptococcus gattii infection in dogs and cats. J Am Vet Med Assoc 2006c, 228:377–382.
Ellis DH, Pfeiffer TJ: Natural habitat of Cryptococcus neoformans var gattii. J Clin Microbiol 1990, 28:1642–1644.
Escandon P, Sanchez A, Martinez M, et al.: Molecular epidemiology of clinical and environmental isolates of the Cryptococcus neoformans species complex reveals a high genetic diversity and the presence of the molecular type VGII mating type a in Colombia. FEMS Yeast Res 2006, 6:625–635.
Filion T, Kidd S, Aguirre K: Isolation of Cryptococcus laurentii from Canada goose guano in rural upstate New York. Mycopathologia 2006, 162:363–368.
+ Fraser JA, Giles SS, Wenink EC, et al.: Same-sex mating and the origin of the Vancouver Island Cryptococcus gattii outbreak. Nature 2005, 437:1360–1364.
An extensive MLST analysis of C. gattii isolates from Vancouver Island and from around the world. The authors found shared genotypes between the VGIIa and VGIIb strains from BC and strains of these molecular types from other parts of the world. This study presents interesting hypotheses about the origin of the VGIIa genotype in BC and reports the first virulence tests of VGIIa and VGIIb strains from Vancouver Island.
Fraser JA, Lim SM, Diezmann S, et al.: Yeast diversity sampling on the San Juan Islands reveals no evidence for the spread of the Vancouver Island Cryptococcus gattii outbreak to this locale. FEMS Yeast Res 2006, 6:620–624.
Fraser JA, Subaran RL, Nichols CB, Heitman J: Recapitulation of the sexual cycle of the primary fungal pathogen Cryptococcus neoformans var. gattii: implications for an outbreak on Vancouver Island, Canada. Eukaryot Cell 2003, 2:1036–1045.
Hoang LM, Maguire JA, Doyle P, et al.: Cryptococcus neoformans infections at Vancouver Hospital and Health Sciences Centre (1997–2002): epidemiology, microbiology and histopathology. J Med Microbiol 2004, 53:935–940.
Kidd SE: Molecular epidemiology and characterization of genetic structure to assess speciation within the Cryptococcus neoformans complex [PhD thesis]. Sydney: University of Sydney; 2003.
++ Kidd SE, Chow Y, Mak S, et al.: Characterization of environmental sources of the human and animal pathogen Cryptococcus gattii in British Columbia, Canada, and the Pacific Northwest of the United States. Appl Environ Microbiol 2007a, 73:1433–1443.
This important study describes a systematic and thorough investigation of the environmental colonization of C. gattii on Vancouver Island and the Pacific Northwest. Key findings include the isolation of the pathogen from air, trees, soil, freshwater, and seawater, and the identification of colonization hotspots. Additionally, this study identified characteristics of soil that may favor C. gattii colonization.
++ Kidd SE, Bach PJ, Hingston AO, et al.: Cryptococcus gattii dispersal mechanisms, British Columbia, Canada. Emerg Infect Dis 2007b, 13:51–57.
This study employed systematic environmental sampling strategies to document patterns of C. gattii colonization on Vancouver Island and to obtain evidence for human-mediated dispersal of the fungus.
+ Kidd SE, Guo H, Bartlett KH, et al.: Comparative gene genealogies indicate that two clonal lineages of Cryptococcus gattii in British Columbia resemble strains from other geographical areas. Eukaryot Cell 2005, 4:1629–1638.
This study employed MLST analysis and gene genealogy to reveal a predominantly clonal population among the Vancouver Island isolates and to demonstrate that the genotypes of isolates from BC resembled those of strains from other parts of the world.
++ Kidd SE, Hagen F, Tscharke RL, et al.: A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proc Natl Acad Sci USA 2004, 101:17258–17263.
This paper describes the results of the first marshaling of the expertise of the international research community to tackle the analysis of the emergence of C. gattii in BC. The investigators described initial studies on the environmental source of the pathogen and identified the molecular types of C. gattii that were responsible for the human and animal cases.
Krockenberger MB, Canfield PJ, Malik R: Cryptococcus neoformans in the koala (Phascolarctos cinereus): colonization by C n var gattii and investigation of environmental sources. Med Mycol 2002, 40:263–272.
Kwon-Chung KJ, Bennett JE: Epidemiologic differences between the two varieties of Cryptococcus neoformans. Am J Epidemiol 1984, 120:123–130.
Kwon-Chung KJ, Boekhout T, Fell JW, Diaz M: (1557) Proposal to conserve the name Cryptococcus gattii against C. hondurianus and C. bacillisporus (Basidiomycota, Hymenomycetes, Tremellomycetidae). Taxon 2002, 51:804–806.
Kwon-Chung KJ, Varma A: Do major species concepts support one, two or more species within Cryptococcus neoformans? FEMS Yeast Res 2006, 6:574–587.
Lester SJ, Kowalewich NJ, Bartlett KH, et al.: Clinicopathologic features of an unusual outbreak of cryptococcosis in dogs, cats, ferrets, and a bird: 38 cases (January to July 2003). J Am Vet Med Assoc 2004, 225:1716–1722.
Lindberg J, Hagen F, Laursen A, et al.: Cryptococcus gattii risk for tourists visiting Vancouver Island, Canada. Emerg Infect Dis 2007, 13:178–179.
Litvintseva AP, Thakur R, Vilgalys R, Mitchell TG: Multilocus sequence typing reveals three genetic subpopulations of Cryptococcus neoformans var grubii (serotype A) including a unique population in Botswana. Genetics 2006, 172:2223–2238.
MacDougall L, Fyfe M: Emergence of Cryptococcus gattii in a novel environment provides clues to its incubation period. J Clin Microbiol 2006, 44:1851–1852.
++ MacDougall L, Kidd SE, Galanis E, et al.: Spread of Cryptococcus gattii in British Columbia, Canada, and detection in the Pacific Northwest, USA. Emerg Infect Dis 2007, 13:42–50.
This paper describes the detection of C. gattii in three people and eight animals without a travel history to Vancouver Island, and the detection of the pathogen in air, soil, water and on trees from sites off the island. The study also reported locally acquired C. gattii infections in three cats in Washington and two people in Oregon; interestingly, the genotypes of the strains from the Oregon cases were VGIIa- and VGIIb-like, but MLST results indicated differences from the isolates of the corresponding subtypes from Vancouver Island.
Mak S: Ecological niche modeling of Cryptococcus gattii in British Columbia [MSc thesis]. Vancouver: University of British Columbia; 2007.
Meyer W, Castaneda A, Jackson S, et al.: Molecular typing of IberoAmerican Cryptococcus neoformans isolates. Emerg Infect Dis 2003, 9:189–195.
Meyer W, Kaocharoen S, Trills L, et al.: Global molecular epidemiology of Cryptococcus gattii VGII isolates traces the origin of the Vancouver Island outbreak to Latin America [abstract]. Presented at the 24th Fungal Genetics Conference. Pacific Grove, CA; March 20–25, 2007.
Meyer W, Marszewska K, Amirmostofian M, et al.: Molecular typing of global isolates of Cryptococcus neoformans var neoformans by PCR-fingerprinting and RAPD—a pilot study to standardize techniques on which to base a detailed epidemiological survey. Electrophoresis 1999, 20:1790–1799.
++ Michael Smith Genome Sciences Center: Cryptococcus Neoformans Summary. http://www.bcgsc.ca/project/cryptococcus/summary/. Accessed July 9, 2007.
The sequences of the genomes of VGI and VGIIa strains are exceptional resources for detailed investigations of the virulence properties of C. gattii. In addition, the sequences allow genome-wide comparative studies with the genomes of C. neoformans var neoformans strains and a var grubii strain.
Morgan J, McCarthy KM, Gould S, et al.: Cryptococcus gattii infection: characteristics and epidemiology of cases identified in a South African province with high HIV seroprevalence, 2002–2004. Clin Infect Dis 2006, 43:1077–1080.
Sorrell TC, Brownlee AG, Ruma P, et al.: Natural environmental sources of Cryptococcus neoformans var gattii. J Clin Microbiol 1996, 34:1261–1263.
Sorrell TC: Cryptococcus neoformans variety gattii. Med Mycol 2001, 39:155–168.
Speed B, Dunt D: Clinical and host differences between infections with the two varieties of Cryptococcus neoformans. Clin Infect Dis 1995, 21:28–34.
Stephen C, Lester S, Black W, et al.: Multispecies outbreak of cryptococcosis on southern Vancouver Island, British Columbia. Can Vet J 2002, 43:792–794.
++ The Broad Institute: Cryptococcus neoformans Serotype B Database. http://www.broad.mit.edu/annotation/genome/cryptococcus_neoformans_b. Accessed July 9, 2007.
The sequences of the genomes of VGI and VGIIa strains are exceptional resources for detailed investigations of the virulence properties of C. gattii. The genome sequence of a C. neoformans var grubii strain is also available at the Broad Institute.United Press International: GE eucalyptus tree investigation urged. http://www.sciencedaily.com/upi/index.php?feed=Science&article=UPI-1-20070614-13565200-bc-us-eucalyptus.xml. Accessed June 17, 2007.
+ Upton A, Fraser JA, Kidd SE, et al.: First contemporary case of human infection with Cryptococcus gattii in Puget Sound: evidence for spread of the Vancouver Island outbreak. J Clin Microbiol 2007, In press.
This report and MacDougall et al. (2007) document the recent emergence of C. gattii outside of BC.
Xu J, Vilgalys R, Mitchell TG: Multiple gene genealogies reveal dispersion and hybridization in the human pathogenic fungus Cryptococcus neoformans. Mol Ecol 2000, 9:1471–1481.
Zender CS, Talamantes J: Climate controls on Valley Fever incidence in Kern County, California. Int J Biometeorol 2006, 50:174–182.
THE GOOD, THE BAD, AND THE UGLY:
FUNGI MOLD YOUR WORLD
Meredith Blackwell3
Fungi4 are important members of many ecosystems. As heterotrophs they are involved in nutrient cycles, especially of carbon, nitrogen, and phosphorus. The effects of fungi were observed in prehistoric times, and their part in causing plant disease was understood before the germ theory was advanced. Today fungi are featured in the popular press and science Internet postings, indicating that they are of increasing interest and importance. Molecular methods have helped to popularize fungi by bringing rapid progress to fungal classification and discovery and have enhanced understanding of their biology. Fungi are associates of all major groups of organisms and are especially well known for their interactions with plants and insects. Fungi also are economically important and provide drugs, foods, and fermented beverages. The value of fungal activities and products far exceeds the costs of the diseases they cause.
Introduction
Human beings were aware of fungal fruiting bodies in prehistoric times, and the sudden appearance of mushrooms after rain awed those who did not comprehend the fungus lifecycle. Lowy (1974) wrote that the sudden appearance of mushrooms of Amanita muscaria was believed to have been caused by thunder-
_____________________
3 Louisiana State University.
4 In addition to members of Kingdom Fungi, several other organisms of the fungus-like group Oomycota (Phytophthora) are included.
bolts as they struck the ground, a belief held independently in Roman, Hindu, and Mayan cultures. Humans endowed mushrooms with magical properties (Wasson, 1968), and evidence of early fungal use exists in many parts of the world. Grave guardians, masks, clothing ornaments, and other artifacts were made from the fruiting bodies of wood-decaying basidiomycetes such as Fomitopsis officinalis and Haploporus odorus (Blanchette, 1997; Blanchette et al., 1992a, 2002). A surviving mushroom culture centered on magic mushrooms existed in Oaxaca for many years, and the celebrated curandera, Maria Sabina, was visited by a number of prominent individuals and notable musicians who sought her spiritual guidance (Wasson, 1957, 1976). Although yeasts themselves were not known, evidence of their activity comes from residues in nine millennia-old Neolithic vessels (Vouillamoz et al., 2006).
Plant pathogenic fungi also were known in ancient times. Three centuries BC, Theophrastus recognized fungi as the cause of certain diseases of crops, but by the first century, the knowledge had been lost, and Pliny attributed lost yields to the gods or stars (Carefoot and Sprott, 1969). Fungal effects such as disease were not understood by many until the observations and experiments of Miles Joseph Berkeley and Anton de Bary around the time of the Irish potato famine of 1845–1846. This work actually came before the general acceptance of the germ theory. The contribution of de Bary also argued strongly against a lingering belief in spontaneous generation (Matta, 2010). Fungi continue to appear suddenly as they invade natural landscapes to cause diseases of plants and animals. The invading organisms often are not noticed until they encounter naïve hosts in new regions where they cause devastating diseases. Earlier invasions included the chestnut blight fungus and several waves of Dutch elm disease fungi (Alexopoulos et al., 1996). The papers in this volume, Fungal Diseases: An Emerging Threat to Human, Animal, and Plant Health (IOM, 2011), cover the newest waves of invasive fungal diseases and their attack on naïve hosts.
More important, however, is the realization that fungi are essential for life on Earth. Fungi are decomposers that destroy plant and animal bodies and return carbon, nitrogen, phosphorus, and other minerals to nutrient cycles. Compatible with their primary role in decomposition, fungi interact with other living organisms in nutritional relationships, and their secondary metabolites and enzymes supply medicines, food and drink, and industrial products for profitable enterprises. Fungi appear regularly in newspapers and magazines. Over the past year, the New York Times featured fungi prominently. Articles have included reports of the identification of a microsporidian fungus partly responsible for colony collapse disorder of bees, a chytrid responsible for global amphibian decline, Geomyces destructans of bats, and pathogens of home garden vegetables. Ecological topics included interactions between bark beetles and fungal symbionts, mycorrhizal associations, sexual reproduction in truffles, a fungus that exerts selective pressure on rotifers, and fungal function in the environment. Fungi also have been covered in the Wall Street Journal as food items, inhabitants of saunas, and the “Torula
of Cognac,” Baudoinia compniacensis, the fungus that grows on walls of wine cellars in mists of alcoholic vapors. One fungus was reported widely because it prompted a murder investigation in a German forest when its sulfurous odor of decay was mistaken for that of a dead body (Anonymous, 2005). Coverage of a broad range of fungal topics also can be found in science blogs and Internet postings with reports of jet lag expressed in circadian rhythms of fungi; wood decay; the evolutionary arms race between a smut fungus and maize; a new species of introduced, beetle-associated fungus that kills plants in the Lauraceae; and yeast genome sequencing leading to improved bioethanol production. National Geographic News also reviews interesting fungal topics, including stories on endophyte biology and “bringing order to the fungus among us,” describing the Assembling the Fungal Tree of Life project (see below). Only 2 days before this meeting (December 12, 2010), USA Today published an article by Elizabeth Weise, “Why it’s cool to have a fungus among us.” The informative article could have been the basis for this talk—if only it had appeared earlier. The range of examples cited indicates a growing interest in and knowledge of fungi.
Fungi influence our daily lives in ways we seldom appreciate. Several entrenched fungal-influenced cultural practices are the result of fungal plant diseases. These include tea drinking in the United Kingdom, a switch imposed by devastation of coffee plants in Ceylon (present-day Sri Lanka) by the coffee rust fungus (Hemileia vastatrix) in the late 1800s (Horsfall and Cowling, 1978); consumption of cornbread as a staple in the southern United States colonies was imposed because the wheat rust fungus prevented wheat cultivation in the humid South (Horsfall, 1958); and the enjoyment of gingerbread comes from the time when the effects of stinking smut of wheat were masked by molasses and ginger (Carefoot and Sprott, 1969). We rely on fungi for clothing fads such as use of cellulase enzymes of species of Trichoderma to speed the “stone washing” of our blue jeans (Bhat, 2000). Perhaps, fungi may make us more beautiful when certain “integrative approaches to better skin”5 are followed using a blend of fungi that includes Cordyceps, reishi mushrooms, and other ingredients.
The Classification and Discovery of New Species of Fungi
Early phylogenetic studies based on DNA sequences defined a monophyletic6 group of Fungi. Oomycota and relatives, various slime mold clades, and several other groups previously considered as zygomycetes have been excluded from the monophyletic Fungi. Asexual and sexual fungi could be combined on the basis of their genetic relationships, and asexual groupings of asexual fungi were abandoned (Alexopoulos et al., 1996). More recently, mycologists have increased
_____________________
5 Several brands of skin creams include a variety of basidiomycete fruiting bodies as ingredients that are said to provide for skin relief and other effects (e.g., Dr. Weil’s Mega-Mushroom lotions, cleansers, and serums).
6 A group of taxa containing an ancestor and all its descendants.
the number of DNA markers and taxa in diverse clades to produce increasingly well-resolved phylogenies,7 the basis of predictive classifications (Figure A2-1) (Hibbett et al., 2007; James et al., 2006; White et al., 2006). An issue of the journal Mycologia (98:829–1103, 2006) was devoted to the phylogenetics of many major groups of fungi. Recent phylogenetic studies have provided new insights into fungal relationships and show that the earliest diverging lineages of zoosporic8 and zygosporic9 groups are not monophyletic as previously assumed on the basis of morphological characters and that they are more diverse than previously understood. Other findings provide data to include microsporidia within or very near the fungi (Lee et al., 2010). The new phylogenetic studies are largely the result of several National Science Foundation projects (Research Coordination Networks: A Phylogeny for Kingdom Fungi [Deep Hypha] and Assembling the Fungal Tree of Life 1 and 2) that involved more than 100 biologists from about 20 countries (Blackwell et al., 2006; Hibbett et al., 2007). Current projects under way include adding taxa to expand the fungal tree of life and pursuing an increasing number of genomics projects.
About 100,000 species of fungi have been described, but a conservative estimate suggests that there are 1.5 million fungi on Earth (Hawksworth, 1991, 2001). The estimate has spurred exploration for the million fungi that remain undiscovered (Figures A2-2A through A2-2D).
More recently the 1.5 million estimate was surpassed by a higher estimate of 3.1 to 5.1 million species based on the use of molecular methods, including highthroughput sequencing (O’Brien et al., 2005). Because of the great discrepancy between known and estimated fungal species numbers, mycologists have a renewed interest in fungal discovery. Many have wondered, where are the missing fungi (Hyde, 2001)? If the higher estimates are realistic, the number of fungi is equal to the number of animal species and may exceed the number of plants by 10:1. Abundant evidence shows that many tropical fungi remain to be discovered based on species accumulation curves of fungi collected in plots (Aime et al., 2010). Other habitats reporting large numbers of fungi include living leaves of tropical trees (Arnold, 2007), soil fungi (O’Brien et al., 2005; Taylor et al., 2010), and even the fungi in the buildings in which we spend most of our time (Amend et al., 2010). Many fungi, however, remain to be discovered in northern temperate regions, including far northeastern Asia (Petersen and Hughes, 2007). We do not have to look for undescribed fungi in completely new places or tropical regions, however, because they may be in our backyards. My colleagues and I look for new species among the yeasts and other microscopic fungi that are difficult to see
_____________________
7 A phylogeny is an inferred history of evolutionary relationships of organisms; often depicted in a tree diagram.
8 Zoospores are flagellated cells of certain fungi (see Figure A2-1) that are produced in sporangia in asexual reproduction.
9 Zygospores are thick-walled spores produced in some fungi (see Figure A2-1) resulting from the fusion of like gametes.
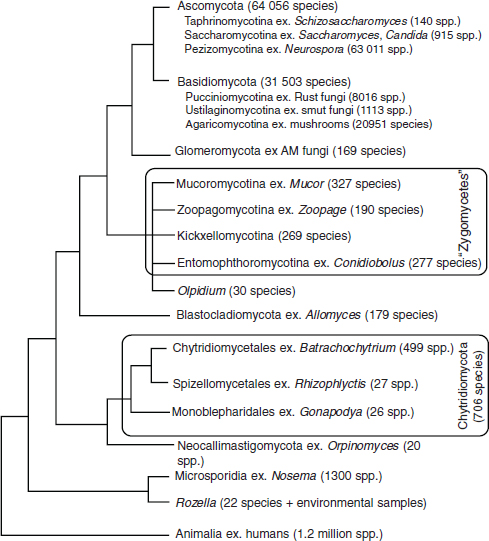
Figure A2-1 Diagrammatic representation of relationships of fungal taxa, examples (ex.), and approximate number of species in each group. Zoosporic and zygosporic fungi are more diverse than previously recognized on the basis of morphological traits, and they are not monophyletic. Two flagellated taxa, Rozella and Olpidium, are of uncertain taxonomic placement. Evidence from multilocus sequencing and genomics reveals that microsporidians branch within or near fungi. Ascomycota and Basidiomycota, the most speciose phyla, are each divided into three subphyla. The largest number of fungal species are classified in the subphyla Pezizomycotina and Agaricomycotina.
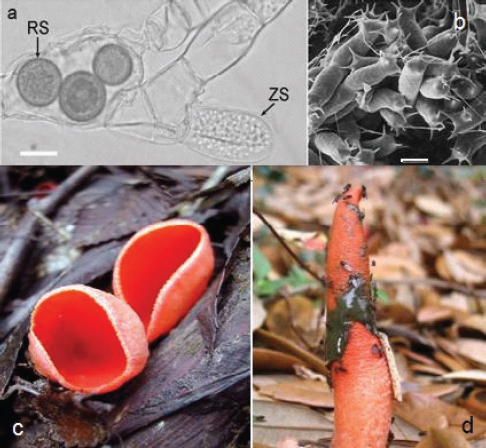

Figure A2-2 Images of representative fungal groups. (A) Hyphae of the blastocladialean fungus, Allomyces sp. Note a terminal zoosporangium (ZS) containing zoospores. The spiny, dark, thick-walled resting spores (RS) within the hyphae are those of a zoosporic fungal parasite, Rozella allomycis, of uncertain taxonomic placement. Bar = 10 µm. (B) Lobosporangium transversale. The zygosporic fungus in the Mortierellales has unusual spiny lobed sporangia (Benny and Blackwell, 2004). Bar = 50 µm. (C) Sarcoscypha coccinea. The several cm diameter fruiting body of the scarlet cup ascomycete, Sarcoscypha coccinea. Ascospores are formed within asci on the inner surface of the cup. (D) Mutinus sp. A stinkhorn similar to one mentioned in the text that caused a search for a dead body (Anonymous, 2005). Stinkhorns produce noxious compounds that attract insect spore dispersers. The dark slimy mass of spores has been partially removed by flies.
SOURCES: (A) photo courtesy of Timothy Y. James, provided by Meredith Blackwell (2009). (B) micrograph courtesy of Kerry O’Donnell, provided by Meredith Blackwell (2004). (C) photo courtesy of eriotropus/coqui, provided by Meredith Blackwell (2002). (D) photo courtesy of Nhu H. Nguyen, provided by Meredith Blackwell (2005).
with the unaided eye (Boekhout, 2005; Suh et al., 2005), and members of early diverging lineages that often are difficult to isolate and culture. Ascomycetes and basidiomycetes are expected to provide the greatest diversity of additional taxa based on numbers of currently known fungi, but certainly the developing methods using high-throughput sequencing of DNA will lead to the discovery of more of the early diverging groups (Figure A2-1) (Kirk et al., 2008).
Examples of large numbers of species isolated into culture from certain substrates include the finding of 418 unique morphotypes of endophytic fungi from 83 leaves in Panama (Arnold, 2007), 257 fungal endophyte genotypes in coffee plants (Vega et al., 2010), and 650 yeast isolates representing 290 genotypes of nearly 200 undescribed taxa from the gut of beetles (Suh et al., 2005). Acquiring cultures and specimens will remain important in cases when fungi and cultures are needed for certain purposes, including population studies, environmental remediation, and secondary metabolites. Taylor and his colleagues (2010) used high-throughput sequencing to estimate the presence of more than 200 taxa in a 0.25 g soil sample with only 14 percent overlap in taxa in a sample taken a meter away. If we are to determine the number of fungi on Earth, environmental sequencing will be necessary to speed fungal exploration and discovery. In addition to new species, entire lineages, some probably at the level of subphylum, may be recognized by DNA sequences such as Soil Clone Group 1 (Porter et al., 2008; Rosling et al., 2010). More work will be needed to determine geographical and substrate ranges in order to obtain more accurate estimates of species numbers.
Species discovery is relevant to the topic of this workshop because previously unknown plant and animal pathogenic fungi have been introduced into the United States many times. These fungi probably caused few symptoms and went unnoticed in their native hosts. Devastation of naïve hosts, however, led to their recognition and subsequent description as new species. This scenario certainly is repeated by the fungi discussed in this meeting, including Batrachochytrium dendrobatidis, the pathogenic chytrid of amphibians spread around the world; Geomyces destructans, the pathogen of bats in North America; and Phytophthora ramorum, causing declines of certain plants in North America and Europe. Prior invasions have included several fungal agents of Dutch elm disease; the chestnut blight fungi; the newly arrived agent of the laurel wilt delivered within the mycangia10 of its ambrosia beetle vector; and Discula destructans, a pathogen of North American dogwoods (Alexopoulos et al., 1996; Harrington and Fraedrich, 2010; Zhang and Blackwell, 2001). Recently, a new approach to discovering the native ranges of certain fungi has been profitable. Ning Zhang (Personal communication, Rutgers University, December 10, 2010) designed an efficient assay method using specific primers to detect the dogwood pathogenic fungus in herbarium specimens. The method promises to greatly reduce the time involved in determining geographical and host ranges and is ideal for working with col-
_____________________
10 Mycangia are pouch-like invaginations in the cuticle of certain insects used to transport cells and spores of symbiotic fungi, found especially in some species of bark and ambrosia beetles as well as a few other groups of insects.
laborators at herbaria throughout the world. Because patterns of introduction of pathogens may exist, determination of native ranges is essential in combating invasive organisms.
Supermodels
Fungi are important as model systems in research. Saccharomyces cerevisiae (Figure A2-3) is a supermodel known for its baking and brewing prowess and as the first eukaryote to have its entire genome sequenced.
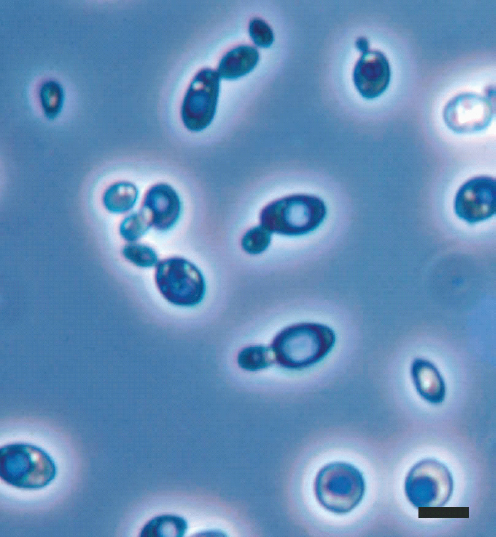
Figure A2-3 Saccharomyces cerevisiae (Y-2235), baker’s yeast and model organism. Note the many budding cells in the stained preparation. Bar = 5 µm.
SOURCE: Photo courtesy of Cletus P. Kurtzman, provided by Meredith Blackwell (2008).
In addition to S. cerevisiae, three other fungi that have been important in research and were the subjects of Nobel Prize–winning research are Schizosaccharomyces pombii, another fast-growing organism with a yeast growth form; Penicillium crysosporium, producer of the first effective antibiotic; and Neurospora crassa. In his Nobel Prize acceptance speech, Tatum (1958) acknowledged, among others, “B.O. Dodge for his establishment of this Ascomycete as a most suitable organism for genetic studies.” Beadle (1958) also spoke of Neurospora crassa and pointed out that “Dodge was an enthusiastic supporter of Neurospora as an organism for genetic work. ‘It’s even better than Drosophila,’ he insisted to Thomas Hunt Morgan, whose laboratory he often visited. He finally persuaded Morgan to take a collection of Neurospora cultures with him from Columbia University to the new Biology Division of the California Institute of Technology, which he established in 1928.” This was the beginning of the development of Neurospora crassa in genetics research.
As mentioned above, S. cerevisiae was the first eukaryotic organism to have its entire genome sequenced. This yeast and other species in the Saccharomycotina have relatively small genomes that make them economical candidates for sequencing (Mewes et al., 1997). In addition, yeasts and other model fungi are easy to grow and complete their lifecycles in culture in a few days; because they are haploid throughout most of their lifecycle, induced mutations are expressed rapidly. Many fungi, including some yeasts, also have a sexual state from which all products of meiosis can be isolated in addition to asexual spores and somatic cells from which uniform populations can be established. They also are excellent organisms for population studies (Anderson et al., 2010). Some fungi, including S. cerevisiae, have morphological cues that indicate the occurrence of certain cell cycle events, and a large body of background information is available for previously established model fungi studies. Improvements in genome sequencing have made it possible to develop many new “models,” including plant and animal pathogens and their hosts. For example, yeasts from the gut of wood-feeding beetles have been of particular interest because many of them ferment xylose, a requirement for efficient digestion of lignocellulose in biofuel production. These species have undergone biochemical and metabolic engineering to obtain more information on xylose fermentation pathways, and genome sequencing is important toward this end (Jeffries et al., 2007; Joint Genome Institute, 2007; Van Vleet and Jeffries, 2009).
Fungi Make Money: Useful Fungal Products
Humans have used a variety of fungal products for different purposes, including cures. In fact some of the magical fungi mentioned above also have been used for their medicinal properties, which may have been known since prehistoric times. Evidence exists for the use of fungi by early humans. Ötzi the Iceman lived about 5,300 years ago, and his mummified body was discovered in 1991
on the border of Italy and Austria. He carried pieces of the fruiting bodies from two species of wood-rotting basidiomycetes, Piptoporus betulinus and Fomes fomentarius, perhaps for medicinal uses (Peinter et al., 1998). Other writers have suggested that one of the fruiting bodies was used as a strop for sharpening knives and tools, but whatever their use, fungi appear to have been important to Copper Age Europeans.
Some basidiomycetes have been used medicinally in more recent times. Extracts of Inonotus obliquus was used in Europe as a treatment for cancer, and the fruiting bodies of Fomitopsis officinalis (the quinine conk), mentioned earlier as grave guardians in the Pacific Northwest, were also harvested for medicinal properties. A different kind of medicinal use by foresters was the application of sheets of mycelium on ax injuries to stop bleeding (Gilbertson, 1980). The spore masses of giant puffballs that were discovered stockpiled along Hadrian’s Wall (in Northern England) also have been used as a styptic (Personal communication, Roy Watling, former Head of Mycology and Plant Pathology, Royal Botanic Garden Edinburgh, August 27, 1977), and spores of unspecified puffballs also were widely used as a styptic by natives of North America as well (Blackwell, 2004).
Certain ascomycete fungi, previously known as species of Cordyceps, have been used in Asian traditional medicine for several centuries (Spatafora et al., 2007). One of these fungi, a parasite of caterpillars, known as Cordyceps sinensis since 1878, now is Ophiocordyceps sinensis based on a phylogenetic study (Sung et al., 2007). Recent interest in the fungus has provided evidence that it may be effective in the treatment of certain tumors (Spatafora et al., 2007). The revision of the entire group of insect–pathogenic fungi previously placed in the genus Cordyceps has resulted in the placement of species in three different families (Sung et al., 2007). This is an important development because phylogenies are predictive of traits common to closely related fungi, and other Ophiocordyceps species may be targeted for the mining of metabolites. The efforts to develop penicillin for the treatment of bacterial infections at the beginning of World War II resulted in the discovery of a long-sought magic bullet and hastened the rise of the modern pharmaceutical industry. In addition to the fungus-derived drug penicillin, three statin drugs for lowering cholesterol levels (e.g., Lipitor®) and the immune suppressant cyclosporine each have earned more than a billion dollars annually. Cyclosporine, once critical to transplant surgery, is today used to treat dry eye as well as more serious conditions (Blackwell, 2011).
Fungi also are big business in the food and beverage industries. In addition to the usual fresh fruiting bodies of basidiomycetes (mushrooms) and a few highly favored ascomycetes (truffles and morels), other fungi, such as cuitlacoche (corn smut) and rice smut, are eaten in Mexico and Asia, respectively. Processed foods also are made from fungi. These include yeast extract spreads such as marmite and vegemite and the meat substitute, Quorn™, a product of hyphae of an ascomycete, a species of Fusarium. Several species of Aspergillus are used in the processing of soy sauce, and fungi play a part in the flavoring process of cheeses.
Throughout the world many fermented foods rely on fungi at least in part to increase nutritional value, improve texture and flavor, and preserve the foodstuff. In one short street block in Brussels, I examined shop windows to count the many products that had been touched by fungi: coffee, certain teas, chocolate, cheeses, bread, salami and dry-cured hams, and numerous fermented beverages (Tamang and Fleet, 2009). Many African and Asian foods, including miso, ontjom, and tempeh, are the products of fermentation (Nout, 2009; Rodríguez Couto and Sanromán, 2006).
As in the case of other fungal products, the making of alcoholic beverages almost certainly was discovered millennia ago, found accidently in prehistoric times when wild yeasts settled into a sugary beverage. Yeasts are essential to the multibillion-dollar alcoholic beverage industry. In the United States, sales of beer, spirits, and wine were $116 billion in 2003 (Library Index, 2011). The yeasts involved in brewing were first isolated into pure culture by Emil Hansen at the Carlsberg Brewery in Copenhagen, and the brewery lab became an important site of classic yeast genetics and biotechnology research (Hansen and Kielland-Brandt, 2003). Pretorius (2000) suggested that many additional yeast species might be used in winemaking. In this context my colleagues and I have discovered nearly 300 previously unknown yeasts, many of which have the ability to ferment a variety of sugars, yet are untried for making beverages (Suh et al., 2005; Urbina and Blackwell, unpublished). In addition to its significance in brewing and bread making, S. cerevisiae, of course, has been extremely important in industrial biotechnology because of the development of efficient transformation methods and specialized expression vectors, and for a variety of other genetics tools (Nevoigt, 2008).
Fungi Interact with Other Organisms
Fungi interact with all major groups of organisms. Specific interactions with photosynthetic organisms are generally well known (Table A2-1). About 80 percent of all plant species and 92 percent of plant families form close associations with fungi known as mycorrhizae (Smith and Read, 2008; Trappe, 1987). Fungi and plant roots or underground stems form several kinds of mycorrhizae that are classified by the morphology of the interacting fungus in relation to the root. The associations are important for carbon, mineral, and water exchange, with carbon generally transferred from the plant to the fungus.
Arbuscular mycorrhizal (AM) fungi are known from the 400 million-year-old Rhynie chert. The fungi penetrate the plant cell wall and form a highly branched arbuscule that invaginates the plasma membrane of the root cortex cells. The 200 members of the asexually reproducing phylum Glomeromycota are obligate fungal partners of about 60 percent of all plant species. Hosts include a variety of crop and forage plants such as maize, rice, alfalfa, and citrus, as well as many non-cultivated plants. Molecular methods have detected previously unknown host
TABLE A2-1 Examples of Fungal Associations with Plants
| Association | Plants | Fungus | Reference |
| AM Mycorrhizae | 60% of all species | Glomeromycota | Selosse et al. (2006) |
| Ectomycorrhizae | 2,000 species | ~5,000 species of basidiomycetes, ascomycetes, Endogenales | Smith and Read (2008) |
| Endophytes | 95% of all plants | Many groups of ascomycetes and some basidiomycetes | Rodriguez et al. (2009) |
| Lichens | ~100 species of photobionts (green algae, blue/green bacteria) | ~32,000 ascomycetes (Leotiales, Dothideales, and Pezizales), a few basidiomycetes | Schoch et al. (2009) |
specificity in some cases (Selosse et al., 2006). Ectomycorrhizal fungi (FIGURE A2-4) are associated with fewer hosts, including certain dominant forest trees such as birch, dipterocarp, eucalyptus, oak, and pine. Greater ectomyccorhizal fungal diversity is evident, and basidiomycetes, ascomycetes, and a few zygomycetes are involved in these associations. Many of the fungi are generalists, but more specificity occurs than among AM associates. The fungi produce an external mantle over young roots and often cause dramatic shortening and dichotomous branching of the mycorrhizal root (Smith and Read, 2008).
Endophytes are fungi that usually grow within above-ground plant parts without causing disease symptoms in about 95 percent of all plants examined (Arnold, 2007). The fungi that form the associations have been placed in four groups, depending on host specificity, tissues colonized, and amount of colonization within the plant (Rodriguez et al., 2009). Hypocrealean endophytes of grasses and sedges produce alkaloids that have been suggested to deter feeding by insects and vertebrates. Endophyte-infected grasses have enhanced growth and drought resistance (Rodriguez et al., 2009). A different group of endophytes is more taxonomically diverse and has broad plant host range with restricted growth within the plant, often occupying only a single cell. Some of these horizontally transmitted endophytes convey protection from plant pathogens (Arnold et al., 2003; Rodriguez et al., 2009). An endophyte was reported to convey heat tolerance to its grass host near a hot springs in Yellowstone National Park, but additional research has shown that only virus-infected endophytes convey thermal tolerance, a sign of the complexity of such associations (Márquez et al., 2007).
About half of the estimated 64,000 ascomycetes (e.g., Leotiales, Dothideales, and Pezizales) and a few basidiomycetes are the fungal associates (mycobionts) of about 100 species of photosynthetic organisms (photobionts) to form lichens
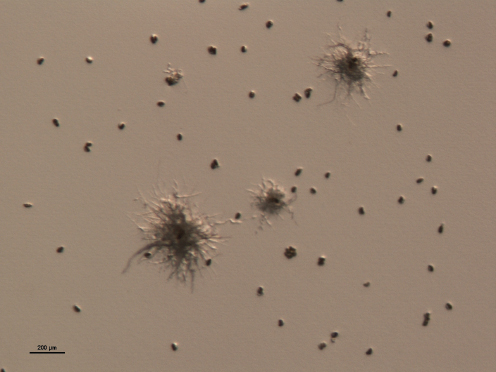
Figure A2-4 Anaptychia ciliaris. Small colonies of the lichen-forming fungus on agar medium after 3 months of growth. Bar = 200 µm.
SOURCE: Photo courtesy of Ning Zhang, provided by Meredith Blackwell (2010).
(Schoch et al., 2009). Lichens have been used as indicators of pollution. In addition to the photosynthetic partner, usually a green alga, a photosynthetic, nitrogen-fixing blue/green bacterium also may occur in a tripartite association in the lichen. Although the fungal associate can be grown on artificial media, they usually grow very slowly (Figure A2-5). Lichens are hosts for pathogenic fungi as well as endolichenic fungi, the lichen equivalent of endophytes. Each partner in the lichen has a scientific name, but the name of the lichen as a whole is that of the fungus (Ahmadjian, 1993; Nash, 2008).
Wood-Decaying Fungi
Fungi are heterotrophic and their ability to degrade organic materials and return them to nutrient cycles is an essential activity in almost all ecosystems. The ability of a fungus to degrade specific substrates depends on the enzymes it produces, and certain fungi are especially important in forest ecosystems where they are the primary decomposers of wood. Basidiomycetes and some ascomycetes are the primary decomposers of plant cell wall carbohydrates (cellulose and
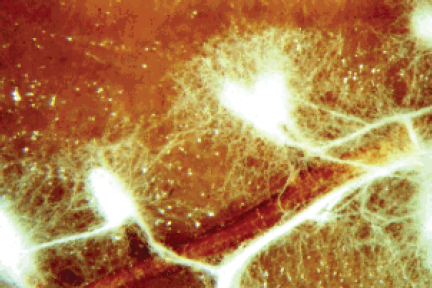
Figure A2-5 Ectomycorrhizal root. The hyphae of Rhizopogon rubescens enveloping the young roots of a Virginia pine seedling. The mycelium extends from the roots into the surrounding environment.
SOURCE: Photo by J. B. Anderson, provided by Meredith Blackwell (1996).
hemicellulose) and lignin polymers (Gilbertson, 1980). Some wood-decaying fungi invade living trees and attack non-functional tissues, especially heartwood, the non-conducting vascular tissue in the center of a cross section of the trunk. Few wood-decaying fungi actually cause diseases and most of the damage comes from the weakening of tree trunks so that they fall in wind or ice storms. The loss of weakened trees is a natural process that culls branches and entire trees to create clearings in older forests (Gilbertson, 1980). Aldo Leopold recognized the value of wood decay for wildlife in the chapter “November” of A Sand County Almanac and Sketches Here and There. He referred to his woodlot as “a mighty fortress that fell heir to all the diseases of plants” known to humankind. The importance of wood-decaying fungi in the formation of nesting holes for wildlife is well known (Gilbertson, 1980). The red-cockaded woodpecker prefers to nest in mature pines about 60 years old that have been rotted by the basidiomycete Phellinus pini. Old pine stands are a diminishing habitat in regions where pines are grown in plantations on a 15-year rotation or less for commercial use. The ivory-billed woodpecker may be extinct because the extensive old-growth, bottomland hardwood forests the species inhabited have been lost (Gilbertson, 1980).
A less significant but interesting use of wood decay is the creation of wooden objects that have been modified by wood-decaying fungi. Spalted wood is distinguished by zone lines, the dark lines formed by oxidation at the points of contact between closely related fungal colonies. The patterned wood is often favored by
collectors and increases the cost of hand-turned bowls at craft fairs. These fungal effects include the deep blue/green stain of an ascomycete fungus that remains green in intarsia of fine Italian furniture and the inlay of Tunbridge Ware objects (Blanchette et al., 1992b). Even Stradivarius violins may have been made more resonant by the partial decay of the wood (Schwarze et al., 2008).
Insects Associated with Fungi and Vertebrates
The importance of many insects in the ecosystem is overlooked, but many of them are important in degradation of course particles, dispersal of bacteria and fungi, and, as is well known, as agents of fungal fertilization. Fungi clearly provide benefits for insects, although the exact advantages to the fungi beyond providing habitat and a means of dispersal often are not clear (Buchner, 1965; Gilbertson, 1984; Mueller et al., 2005). Few animals have the enzymes necessary to digest refractory plant cell wall materials or to synthesize vitamins. Fungi also may detoxify plant toxins and produce pheromones for insects (Table A2-2) (Dowd, 1991; Vega and Dowd, 2005; Wheeler and Blackwell, 1984; Wilding et al., 1989). The best known fungus–insect associations include the farming interactions of basidiomycetes with Old World termites (Macrotermitinae) (Aanen
TABLE A2-2 Examples of Fungal Associations with Insects
| Insect Group | Fungi | Reference |
| Macrotermitinae | Termitomyces spp. | Aanen et al. (2002) |
| Formicidae: Attini (derived clades) | Leucocoprinus spp. | Mueller et al. (2005) |
| Formicidae: Attini (most in Apterostigma pilosum clade) | Pterulaceae spp. | Munkacsi et al. (2004) |
| Scolytinae and Platypodinae | Ophiostomatoid ascomycetes | Farrell et al. (2001); |
| (Bark and ambrosia beetles) | Harrington (2005) | |
| Siricidae (wood wasps) | Certain species of Amylostereum, Stereum, and Daedalea | Martin (1992) |
| Passalidae (bess beetles) | Several clades of xylose-fermenting yeasts | Suh et al. (2003) |
| Mushroom-feeding beetles | Candida tanzawaensis clade yeasts | Suh et al. (2005) |
| Drosophila in cacti | Various yeasts | Starmer et al. (2006) |
| Nectar feeding beetles | Various yeasts | Lachance et al. (2001) |
| Coccidae | Septobasidium | Henk and Vilgalys (2007) |
| Certain insects, especially aquatic larvae | Harpellales, Asellariales | Lichtwardt et al. (2001) |
et al., 2002) and attine ants (Figure A2-6) (Formicidae: Attini) (Mueller et al., 2005) and of ascomycetes by bark and ambrosia beetles (Scolytinae and Platypodinae) (Harrington, 2005). The females of another insect group, siricid wood wasps (Siricidae), are less well studied, but they have been considered by some to form farming interactions with fungi (see Gilbertson, 1984). The interaction, however, does not meet all the criteria established for what has been defined as “agriculture” (Mueller et al., 2005).
The farming association of the basidiomycete Termitomyces with Old World macrotermitine termites arose once in Africa. Since that event no additional fungal lineages have been domesticated and no reversals of the fungus to a free-living state have been found. Repeated host switching, however, has occurred within termite clades as reflected in the phylogenetic trees of termites and associated fungi (Aanen et al., 2002). Nest initiation by both males and females of certain species has been suggested to have influenced the mode of transmission of the fungus, usually acquired from the environment or some source other than a parent (horizontal transmission) (Aanen et al., 2002). In the New World it is not termites, but attine ants that are involved with basidiomycetes in farming interactions, and Aanen and his colleagues (2002) compared the associations. The attines have become associated with several clades of fungi, and in contrast to termite transmission, transmission of the fungi is usually directly from parent to offspring (vertical) except in the early diverging ant lineages. Another important difference is that the ant-associated fungi apparently do not reproduce sexually. The work on the fungus–attine ant associations have revealed that ants have evolved with several groups of fungi on several different occasions. Although the best-known fungal mutalists are species of Leucocoprinus, other fungal groups, including certain species of Pterulaceae, have an association with ants in the Apterostigma pilosum clade (Munkacsi et al., 2004). The intensive studies of the fungi and attine ant associations have led to the discovery of other organisms that participate in the complex interactions. Species of hypocrealean ascomycetes in the genus Escovopsis are parasites of the cultivated fungus. Actinomycete associates of the ants produce antibiotics that have been reported to be specific in inhibiting Escovopsis (Currie et al., 1999), but more recently Sen et al. (2009) found that the bacteria they isolated had more generalized antibiotic activity, including activity against the cultivated fungus. The association of a fourth component of the association is black yeasts that apparently reduce the efficiency of the antibiotics (Little and Currie, 2008). This attine and—cultivated fungus—Escovopsis parasite associations provide the best example of coevolution, in this case tripartite association, among fungi and associates (Currie et al., 2003).
Unlike the termite and ant interactions, fungus-beetle associations have arisen multiple times. Some bark and ambrosia beetles have mycangia already mentioned above in which they carry inoculum of certain fungi (Malloch and Blackwell, 1993). The fungi, often Ceratocystis and Ophiostoma or relatives, may be the agents of plant diseases, and some of the fungi have been introduced

Figure A2-6 Excavation of deeply entrenched nest of the ant Atta texana requires heavy equipment or, alternatively, ground-penetrating radar to map such nests. The ant is native to adjacent parts of Texas and Louisiana, and the nests are said to be able to contain a three-story house. Visual materials based on a ground-penetrating radar nest model are available on the Internet from Carol LaFayette, Department of Visualization, Texas A&M. http://www.viz.tamu.edu/faculty/lurleen/main/attatunnel/tunnel.html.
SOURCE: Photo courtesy of John Moser, provided by Meredith Blackwell (2009).
with the beetles as in the case of Raffaelea laurelensis, the agent of laurel wilt disease (Harrington, 2005; Harrington and Fraedrich, 2010). Ophiostoma ulmi and similar fungi have been introduced into the United States, where they are virulent pathogens of trees, including American elms. The most efficient dispersers of some of these fungi actually were introduced before the fungus, Ophiostoma ulmi (Alexopoulos et al., 1996). In this discussion of beneficial fungi, these interactions benefit the insects and call attention to potential devastating effects of efficient insect dispersal in the context of emerging plant diseases.
Other beneficial fungal associates of insects involve siricid wood wasps and wood-decaying basidiomycetes, species of Amylostereum, Stereum, and Daedalea. The wasps lay their eggs through long ovipositors, tube-shaped organs at the posterior of the abdomen, and the larvae probably rely on fungal enzymes to decompose and detoxify the wood they ingest (Gilbertson, 1984; Martin, 1992). Many more fungi are associated with insects as necrotrophic parasites (FIGURE A2-7), and some of these deadly fungi have potential for development as biological control agents (Vega et al., 2009). In addition, many of about 1,000 described yeast species have close associations with insects (Table A2-2), and the yeasts provide important services to the insects (Vega and Dowd, 2005). Cer-

Figure A2-7 Hirsutella citriformis (Ophiostomataceae) on a delphacid planthopper. The asexual fruiting structure of this fungus erupted through the cuticle of the parasitized insect soon after its death.
SOURCE: Photo courtesy of Jennifer Luangsa-ard, provided by Meredith Blackwell (2010).
tain clades of gut yeasts appear to have diversified with insect hosts into certain habitats, and the yeasts provide basic resources for the insects to survive when subjected to new nutritional situations (Suh et al., 2003, 2006). About 200 species of Septobasidium in the Septobasidiales are known as associates of scale insects; only a few related species of Pachnocybe grow on wood (Henk and Vilgalys, 2007). The use of insect hosts is unusual for fungi that are related to the plant pathogenic rust fungi. The fungi are parasites of a few of the scale individuals, but in general benefit the entire insect colony by providing a protective covering against parasitic wasps (Henk and Vilgalys, 2007). Two orders of zygomycetes, Harpellales and Asellariales, were previously placed in a polyphyletic group known as Trichomycetes. The results of several studies indicate that these gut fungi produce vitamins and perhaps other benefits for their aquatic insect hosts (Lichtwardt et al., 2001). One species is known to parasitize simulid black flies (Lichtwardt et al., 2001), potentially a benefit to those who engage in outdoor activities.
Another nutritional interaction between fungi and animals is only briefly noted here, but is extremely important. An early diverging lineage of obligately anaerobic multiflagellated fungi, the Neocallimastigomycota, and vertebrate herbivores are closely associated (Griffith et al., 2010). The fungi reside in the host rumen or another anaerobic part of the gut, where they are important in supplying cellulases and other enzymes for the degradation of the large quantities of cellulose ingested by the herbivore (James et al., 2006).
Conclusion
Many fungi are obligate, beneficial associates of other groups of organisms. These are the “good fungi” of this article, and we often fail to appreciate their value because the fungi usually are unseen within their substrates unless they form macroscopic fruiting bodies. More often it is the effects of the fungi that we observe when they ferment fruit juice, or fitting to this volume, cause dramatic new outbreaks of disease. The Robert Frost poem quoted in the prologue of this publication describes the costs of the introduction of the disease caused by the chestnut blight fungus, Cryphonectria parasitica. The poem predicts that the disease will ravage until a new pathogen comes to kill the fungus, and in fact a virus did appear to suppress the fungus. In 1974, however, yet another pathogen, the oriental chestnut gall wasp, was introduced to attack the trees, an additional turn not predicted by the verse.
Today, as one out of every six or seven humans on Earth is reported to be malnourished or hungry (FAO, 2010), the war against pathogenic diseases of plants and animals is as important as ever. An earlier writer, Jonathan Swift (1667–1745) addressed the topic of hunger with his essay, A Modest Proposal, written to bring attention to the starvation of Irish tenant farmers during the potato famine. In Gulliver’s Travels he wrote directly of the importance of increasing agriculture yields:
And he gave it for his opinion, “that whoever could make two ears of corn, or two blades of grass, to grow upon a spot of ground where only one grew before, would deserve better of mankind, and do more essential service to his country, than the whole race of politicians put together.”
—Jonathan Swift, Gulliver’s Travels, Part II, Voyage to Brobdingnag,
first published in 1726–1727.
This volume, Fungal Diseases: An Emerging Threat to Human, Animal, and Plant Health, provides a discussion of new fungal diseases of plants and the animals that we strive to overcome at a time when introduced diseases contribute to hunger.
Acknowledgments
I am grateful to Dr. Fernando Vega, who improved the original manuscript through his careful editing. Several colleagues provided images, and Dr. Matthew Brown kindly prepared the plate. I acknowledge support from the National Science Foundation (NSF-0732671 and DEB-0417180) and the Louisiana State University Boyd Professor support fund.
References
Aanen, D. K., P. Eggleton, C. Rouland-Lefèvre, T. Guldberg-Frøslev, S. Rosendahl, and J. J. Boomsma. 2002. The evolution of fungus-growing termites and their mutualistic fungal symbionts. Proceedings of the National Academy of Sciences, USA 99:14887–14892.
Ahmadjian, V. 1993. The lichen symbiosis. New York: John Wiley and Sons.
Aime, M. C., D. L. Largent, T. W. Henkel, and T. J. Baroni. 2010. The entolomataceae of the Pakaraima Mountains of Guyana IV: New species of Calliderma, Paraeccilia and Trichopilus. Mycologia 102:633–649.
Alexopoulos, C. J., C. W. Mims, and M. Blackwell. 1996. Introductory mycology. New York: John Wiley and Sons.
Amend, A. S., K. A. Seifert, R. Samson, and T. D. Bruns. 2010. Indoor fungal composition is geographically patterned and more diverse in temperate zones than in the tropics. Proceedings of the National Academy of Sciences, USA 107:13748–13753.
Anderson, J. B., J. Funt, D. A. Thompson, S. Prabhu, A. Socha, C. Sirjusingh, J. R. Dettman, L. Parreiras, D. S. Guttman, A. Regev, and L. M. Kohn. 2010. Determinants of divergent adaptation and Dobzhansky-Muller interaction in experimental yeast populations. Current Biology 20:1383–1388.
Anonymous. 2005. “. . . while in Germany, a search for a body turns up mushrooms.” The Mycophile 46 (6):5 http://www.namyco.org/images/pdf_files/MycophileNovDec05.pdf [reprint from Reuters Limited web site, Updated: 3:45 p.m. ET Aug. 2, 2005].
Arnold, A. E. 2007. Understanding the diversity of foliar endophytic fungi: Progress, challenges, and frontiers. Fungal Biology Reviews 21:51–66.
Arnold, A. E., L. C. Mejía, D. Kyllo, E. I. Rojas, Z. Maynard, N. Robbins, and E. A. Herre. 2003. Fungal endophytes limit pathogen damage in a tropical tree. Proceedings of the National Academy of Sciences, USA 100:15649–15654.
Beadle, G. W. 1958. Nobel lecture. http://nobelprize.org/nobel_prizes/medicine/laureates/1958/beadle-lecture.html (accessed February 24, 2011).
Benny, G. L., and M. Blackwell. 2004. Lobosporangium, a new name for Echinosporangium Malloch, and Gamsiella, a new genus for Mortierella multidivaricata. Mycologia 96:143–149.
Bhat, M. K. 2000. Cellulases and related enzymes in biotechnology. Biotechnology Advances 18: 355–383.
Blackwell, M. 2011. The fungi: 1, 2, 3, … 5.1 million species? American Journal of Botany: 98:426–438.
Blackwell, M., D. S. Hibbett, J. W. Taylor, and J. W. Spatafora. 2006. Research coordination networks: A phylogeny for kingdom Fungi (Deep Hypha). Mycologia 98:829–837.
Blackwell, M., C. P. Kurtzman, M.-A. Lachance, and S.-O. Suh. 2009a. Saccharomycotina. Saccharomycetales. Version 22 January 2009. http://tolweb.org/Saccharomycetales/29043/2009.01.22 (accessed March 30, 2011).
Blackwell, M., R. Vilgalys, T. Y. James, and J. W. Taylor. 2009b. Fungi. Eumycota: mushrooms, sac fungi, yeast, molds, rusts, smuts, etc. Version 10 April 2009. http://tolweb.org/Fungi/2377/2009.04.10 (accessed March 30, 2011).
Blackwell, W. H. 2004. Puffballs: Overlooked medicinals? Mushroom, the Journal Fall 2004:1–5.
Blanchette, R. A. 1997. Haploporus odorus: A sacred fungus in traditional Native American culture of the northern plains. Mycologia 89:233–240.
Blanchette, R. A., B. D. Compton, N. J. Turner, and R. L. Gilbertson. 1992a. Nineteenth century shaman grave guardians are carved Fomitopsis officinalis sporophores. Mycologia 84:119–124.
Blanchette, R. A., A. M. Wilmering, and M. Baumeister. 1992b. The use of green-stained wood caused by the fungus Chlorociboria in intarsia masterpieces from the 15th-century. Holzforschung 46:225–232.
Blanchette, R. A., C. C. Renner, B. W. Held, C. Enoch, and S. Angstman. 2002. The current use of Phellinus igniarius by the Eskimos of Western Alaska. Mycologist 16:142–145.
Boekhout, T. 2005. Gut feeling for yeasts. Nature 434:449–451.
Buchner, P. 1965. Endosymbiosis of animals with plant microorganisms. New York: John Wiley and Sons.
Carefoot, E. R., and G. L. Sprott. 1969. Famine on the wind: Man’s battle against plant disease. Chicago, IL: Rand McNally and Company.
Currie, C. R., J. A. Scott, R. C. Summerbell, and D. Malloch. 1999. Fungus-growing ants use antibiotic-producing bacteria to control garden parasites. Nature 398:701–704.
Currie, C. R., B. Wong, A. E. Stuart, T. R. Schultz, S. A. Rehner, U. G. Mueller, G.-H. Sung, J. W. Spatafora, and N. A. Straus. 2003. Ancient tripartite coevolution in the attine ant-microbe symbiosis. Science 299:386–388.
Dowd, P. F. 1991. Symbiont-mediated detoxification in insect herbivores. In Microbial mediation of plant–herbivore interactions, edited by P. Barbosa, V. A. Krischik, and C. G. Jones. New York: John Wiley and Sons. Pp. 411–440.
FAO (Food and Agriculture Organization). 2010. The state of food insecurity in the world 2010. http://www.fao.org/docrep/013/i1683e/i1683e.pdf (accessed June 13, 2011).
Farrell, B. D., A. S. Sequeira, B. C. O’Meara, B. B. Normark, J. H. Chung, and B. H. Jordal. 2001. The evolution of agriculture in beetles (Curculionidae: Scolytinae and Platypodinae). Evolution 55:2011–2027.
Gilbertson, R. L. 1980. Wood-rotting fungi of North America. Mycologia 72:1–49.
———. 1984. Relationships between insects and wood-rotting basidiomycetes. In Fungus-insect relationships, perspectives in ecology and evolution, edited by Q. Wheeler and M. Blackwell. New York: Columbia University Press. Pp. 130–165.
Griffith, G., S. Baker, K. Fliegerova, A. Liggenstoffer, M. van der Giezen, and G. Beakes. 2010. Anaerobic fungi: Neocallimastigomycota. IMA Fungus 1:181–185.
Hansen, J., and M. C. Kielland-Brandt. 2003. Brewer’s yeast: Genetic structure and targets for improvement. In Functional genetics of industrial yeasts, edited by J. H. de Winde. Berlin, Germany: Springer. Pp. 143–170.
Harrington, T. C. 2005. Ecology and evolution of mycophagous bark beetles and their fungal partners. In Insect–fungal associations: Ecology and evolution, edited by F. E. Vega and M. Blackwell. New York: Oxford University Press. Pp. 257–291.
Harrington, T. C., and S. W. Fraedrich. 2010. Quantification of propagules of the laurel wilt fungus and other mycangial fungi from the redbay ambrosia beetle, Xyleborus glabratus. Phytopathology 100:1118–1123.
Hawksworth, D. L. 1991. The fungal dimension of biodiversity: Magnitude, significance, and conservation. Mycological Research 95:641–655.
———. 2001. The magnitude of fungal diversity: The 1.5 million species estimate revisited. Mycological Research 105:1422–1432.
Henk, D. A., and R. Vilgalys. 2007. Molecular phylogeny suggests a single origin of insect symbiosis in the Pucciniomycetes with support for some relationships within the genus Septobasidium. American Journal of Botany 94:1515–1526.
Hibbett, D. M., M. Binder, J. F. Bischoff, M. Blackwell, P. F. Cannon, O. Eriksson, S. Huhndorf, T. Y. James, P. M. Kirk, R. Lücking, T. Lumbsch, F. Lutzoni, P. B. Matheny, D. J. McLaughlin, M. J. Powell, S. Redhead, C. L. Schoch, J. W. Spatafora, J. A. Stalpers, R. Vilgalys, M. C. Aime, A. Aptroot, R. Bauer, D. Begerow, G. L. Benny, L. A. Castlebury, P. W. Crous, Y.-C. Dai, W. Gams, D. M. Geiser, G. W. Griffith, D. L. Hawksworth, V. Hofstetter, K. Hosaka, R. A. Humber, K. Hyde, U. Kõljalg, C. P. Kurtzman, K.-H. Larsson, R. Lichtwardt, J. Longcore, A. Miller, J.-M. Moncalvo, S. Mozley Standridge, F. Oberwinkler, E. Parmasto, J. D. Rogers, L. Ryvarden, J. P. Sampaio, A. Schuessler, J. Sugiyama, J. W. Taylor, R. G. Thorn, L. Tibell, W. A. Untereiner, C. Walker, Z. Wang, A. Weir, M. Weiss, M. White, K. Winka, Y.-J. Yao, and N. Zhang. 2007. A higher-level phylogenetic classification of the fungi. Mycological Research 111:509–547.
Horsfall, J. G. 1958. The fight with the fungi: The rusts and rots that rob us, the blasts and blights that beset us. In Fifty years of botany: Golden jubilee volume of the Botanical Society of America, edited by W. C. Steere. New York: McGraw-Hill. Pp. 50–60.
Horsfall, J. G., and E. B. Cowling. 1978. Some epidemics man has known. In Plant pathology: An advanced treatise. Vol. 2. The diseased plant, edited by J. G. Horsfall and E. B. Cowling. New York: Academic Press. Pp. 17–32.
Hyde, K. D. 2001. Where are the missing fungi? Mycological Research 105:1409–1412.
IOM (Institute of Medicine). 2011. Fungal diseases: An emerging challenge to human, animal, and plant health—a workshop summary. Washington, DC: The National Academies Press.
James, T. Y., P. M. Letcher, J. E. Longcore, S. E. Mozley-Standridge, D. Porter, M. J. Powell, G. W. Griffith, and R. Vilgalys. 2006. A molecular phylogeny of the flagellated Fungi (Chytridiomycota) and description of a new phylum (Blastocladiomycota). Mycologia 98:860–871.
Jeffries, T. W., I. V Grigoriev, J. Grimwood, J. M. Laplaza, A. Aerts, A. Salamov, J. Schmutz, E. Lindquist, P. Dehal, H. Shapiro, Y.-S. Jin, V. Passoth, and P. M. Richardson. 2007. Genome sequence of the lignocellulose-bioconverting and xylose-fermenting yeast Pichia stipitis. Nature Biotechnology 25:319–326.
Joint Genome Institute. 2007. Super-fermenting fungus genome sequenced. To be harnessed for improved biofuels production. http://www.jgi.doe.gov/News/news_3_5_07.html (accessed March 25, 2011).
Lachance, M. A., W. T. Starmer, C. A. Rosa, J. M. Bowles, J. S. F. Barker, and D. H. Janzen. 2001. Biogeography of the yeasts of ephemeral flowers and their insects. FEMS Yeast Research 1:1–8.
Lee, S. C., N. Corradi, S. Doan, F. S. Dietrich, P. J. Keeling, and J. Heitman. 2010. Evolution of the sex-related locus and genomic features shared in Microsporidia and Fungi. PLoS ONE 5:e10539.
Library Index. 2011. U.S. alcohol sales and consumption. http://www.libraryindex.com/pages/2127/Economics-Alcohol-Tobacco-U-S-ALCOHOL-SALES-CONSUMPTION.html (accessed March 28, 2011).
Lichtwardt, R. W., M. J. Cafaro, and M. M. White. 2001. The Trichomycetes, fungal associates of arthropods. Revised edition. http://www.nhm.ku.edu/~fungi/monograph/text/mono.htm (accessed March 22, 2011).
Little, A. E. F., and C. R. Currie. 2008. Black yeast symbionts comprise the efficiency of antibiotic defenses in fungus-growing ants. Ecology 89:1216–1222.
Lowy, B. 1974. Amanita muscaria and the thunderbolt legend in Guatemala and Mexico. Mycologia 66:188–191.
Malloch, D., and M. Blackwell. 1993. Dispersal biology of ophiostomatoid fungi. In Ceratocystis and Ophiostoma: Taxonomy, ecology and pathology, edited by M. J. Wingfield, K. A. Seifert, and J. F. Webber. St. Paul, MN: APS. Pp. 195–206.
Márquez, L. M., R. S. Redman, R. J. Rodriguez, and M. J. Roossinck. 2007. A virus in a fungus in a plant—three-way symbiosis required for thermal tolerance. Science 315:513–515.
Martin, M. M. 1992. The evolution of insect–fungus associations: From contact to stable symbiosis. American Zoologist 32:593–605.
Matta, C. 2010. Spontaneous generation and disease causation: Anton de Bary’s experiments with Phytophthora infestans and late blight of potato. Journal of the History of Biology 43:459–491.
Mewes, H. W., K. Albermann, M. Bähr, D. Frishman, A. Gleissner, J. Hani, K. Heumann, K. Kleine, A. Maier, S. G. Oliver, F. Pfeiffer, and A. Zollner. 1997. Overview of the yeast genome. Nature 387:7–8.
Mueller, U. G., N. M. Gerardo, T. R. Schultz, D. Aanen, and D. Six. 2005. The evolution of agriculture in insects. Annual Review of Ecology and Systematics 36:563–569.
Munkacsi, A. B., J. J. Pan, P. Villesen, U. G. Mueller, M. Blackwell, and D. J. McLaughlin. 2004. Convergent coevolution in the domestication of coral mushrooms by fungus-growing ants. Proceedings of the Royal Society of London, Series B, Biological Sciences 271:1777–1782.
Nash, T. H. 2008. Lichen biology, 2nd ed. Cambridge, U.K.: Cambridge University Press.
Nevoigt, E. 2008. Progress in metabolic engineering of Saccharomyces cerevisiae. Microbiology and Molecular Biology Reviews 72:379–412.
Nout, M. J. R. 2009. Rich nutrition from the poorest: Cereal fermentations in Africa and Asia. Food Microbiology 26:685–692.
O’Brien, B. L., J. L. Parrent, J. A. Jackson, J. M. Moncalvo, and R. Vilgalys. 2005. Fungal community analysis by large-scale sequencing of environmental samples. Applied and Environmental Microbiology 71:5544–5550.
Peinter, U., R. Pöder, and T. Pümpel. 1998. The iceman’s fungi. Mycological Research 102:1153–1162.
Petersen, R. H., and K. W. Hughes. 2007. Some agaric distributions involving Pacific landmasses and Pacific Rim. Mycoscience 48:1–14.
Porter, T. M., C. W. Schadt, L. Rizvi, A. P. Martin, S. K. Schmidt, L. Scott-Denton, R. Vilgalys, and J. M. Moncalvo. 2008. Widespread occurrence and phylogenetic placement of a soil clone group adds a prominent new branch to the fungal tree of life. Molecular Phylogenetics and Evolution 46:635–644.
Pretorius, I. S. 2000. Tailoring wine yeast for the new millennium: Novel approaches to the ancient art of winemaking. Yeast 16:675–729.
Rodriguez, R. J., J. F. White, Jr., A. E. Arnold, and R. S. Redman. 2009. Fungal endophytes: Diversity and functional roles. New Phytologist 182:314–330.
Rodríguez Couto, S., and M. A. Sanromán. 2006. Application of solid-state fermentation to food industry—a review. Journal of Food Engineering 76:291–302.
Rosling, A., K. Cruz Martinez, A. Menkis, K. Ihrmark, S. Holmström, S. Norström, A. Broberg, and B. D. Lindahl et al. 2010. Getting to know the fungi in Soil Clone Group 1. Abstract. International Mycological Congress, Edinburgh, Scotland. August 4, 2010.
Schoch, C. L., G.-H. Sung, F. L. López-Giráldez, J. P. Townsend, J. Miadlikowska, V. Rie Hofstetter, B. Robbertse, P. B. Matheny, F. Kauff, Z. Wang, C. Gueidan, R. M. Andrie, K. Trippe, L. M. Ciufetti, A. Wynns, E. Fraker, B. P. Hodkinson, G. Bonito, J. Z. Groenewald, M. Arsanlou, G. S. De Hoog, P. W. Crous, D. Hewitt, D. H. Pfister, K. Peterson, M. Grysenhout, M. J. Wingfield, A. Aptroot, S.-O. Suh, M. Blackwell, D. M. Hillis, G. W. Griffith, L. A. Castlebury, A. Y. Rossman, H. T. Lumbsch, R. L. Lücking, B. Büdel, A. Rauhut, P. Diederich, D. Ertz, D. M. Geiser, K. Hosaka, P. Inderbitzin, J. Kohlmeyer, B. Volkmann-Kohlmeyer, L. Mostert, K. O’Donnell, H. Sipman, J. D. Rogers, R. A. Shoemaker, J. Sugiyama, R. C. Summerbell, W. Untereiner, P. R. Johnston, S. Stenroos, A. Zuccaro, P. S. Dyer, P. D. Crittenden, M. S. Cole, K. Hansen, J. M. Trappe, R. Yahr, F. Lutzoni, and J. W. Spatafora. 2009. The Ascomycota tree of life: A phylum-wide phylogeny clarifies the origin and evolution of fundamental reproductive and ecological traits. Systematic Biology 58:224–239.
Schwarze, F. W., M. Spycher, and S. Fink. 2008. Superior wood for violins—wood decay fungi as a substitute for cold climate. New Phytologist 179:1095–1104.
Selosse, M.-A., F. Richard, X. He, and S. W. Simard. 2006. Mycorrhizal networks: des liaisons dangereuses? Trends in Ecology and Evolution 21:621–628.
Sen, R., H. D. Ishak, D. Estrada, S. E. Dowd, E. Hong, and U. G. Mueller. 2009. Generalized antifungal activity and 454-screening of Pseudonocardia and Amycolatopsis bacteria in nests of fungus-growing ants. Proceedings of the National Academy of Sciences, USA 106:17805–17810.
Smith, S. E., and D. J. Read. 2008. Mycorrhizal symbiosis. San Diego, CA: Academic.
Spatafora, J. W., G.-H. Sung, J.-M. Sung, N. Hywel-Jones, and J. F. White. 2007. Phylogenetic evidence for an animal pathogen origin of ergot and the grass endophytes. Molecular Ecology 16:1701–1711.
Starmer, W. T., R. A. Schmedicke, and M. A. Lachance. 2006. The origin of the cactus-yeast community. FEMS Yeast Research 3:441–448.
Suh, S.-O., C. J. Marshall, J. V. McHugh, and M. Blackwell. 2003. Wood ingestion by passalid beetles in the presence of xylose-fermenting gut yeasts. Molecular Ecology 12:3137–3145.
Suh, S.-O., J. V. McHugh, D. Pollock, and M. Blackwell. 2005. The beetle gut: A hyperdiverse source of novel yeasts. Mycological Research 109:261–265.
Suh, S.-O., M. Blackwell, C. P. Kurtzman, and M.-A. Lachance. 2006. Phylogenetics of Saccharomycetales, the ascomycete yeasts. Mycologia 98:1008–1019.
Sung, G.-H., N. L. Hywel-Jones, J.-M. Sung, J. Luangsa-ard, B. Shrestha, and J. W. Spatafora. 2007. Phylogenetic classification of Cordyceps and the clavicipitaceous fungi. Studies in Mycology 57:5–59.
Tamang, J. P., and G. H. Fleet. 2009. Yeast diversity in fermented foods and beverages. In Yeast biotechnology: Diversity and application, edited by T. Satyanarayana and G. Kunze. Berlin, Germany: Springer. Pp. 169–198.
Tatum, E. L. 1958. Nobel lecture. http://nobelprize.org/nobel_prizes/medicine/laureates/1958/tatumlecture.html (accessed June 13, 2011).
Taylor, D. L., I. C. Herriott, K. E. Stone, J. W. McFarland, M. G. Booth, and M. B. Leigh. 2010. Structure and resilience of fungal communities in Alaskan boreal forest soils. Canadian Journal of Forest Research 40:1288–1301.
Trappe, J. M. 1987. Phylogenetic and ecologic aspects of mycotrophy in the angiosperms from an evolutionary standpoint. In Ecophysiology of VA mycorrhizal plants, edited by G. R. Safir. Boca Raton, FL: CRC Press. Pp. 2–25.
Urbina, H. and M. Blackwell. 2010 (unpublished). Yeasts associated with wood-ingesting beetles. Baton Rouge, LA: Louisiana State University.
Van Vleet, J. H., and T. W. Jeffries. 2009. Yeast metabolic engineering for hemicellulosic ethanol production. Current Opinion in Biotechnology 20:300–306.
Vega, F. E., and P. F. Dowd. 2005. The role of yeasts as insect endosymbionts. In Insect–fungal associations: Ecology and evolution, edited by F. E. Vega and M. Blackwell. New York: Oxford University Press. Pp. 211–243.
Vega, F. E., M. S. Goettel, M. Blackwell, D. Chandler, M. A. Jackson, S. Keller, M. Koike, N. K. Maniania, A. Monzón, B. H. Ownley, J. K. Pell, D. E. N. Rangel, and H. E. Roy. 2009. Fungal entomopathogens: New insights on their ecology. Fungal Ecology 2:149–159.
Vega, F. E., A. Simpkins, M. C. Aime, F. Posada, S. W. Peterson, S. A. Rehner, F. Infante, A. Castillo, and A. E. Arnold. 2010. Fungal endophyte diversity in coffee plants from Colombia, Hawai’i, Mexico, and Puerto Rico. Fungal Ecology 3:122–138.
Vouillamoz, J. F., P. E. McGovern, A. Ergul, G. Söylemezoğlu, G. Tevzadze, C. P. Meredith, and M. S. Grando. 2006. Genetic characterization and relationships of traditional grape cultivars from Transcaucasia and Anatolia. Plant Genetic Resources 4:144–158.
Wasson, R. G. 1957. Seeking the magic mushroom. Life magazine, May 13, 1957:100–120.
———. 1968. Soma: Divine mushroom of immortality. New York: Harcourt Brace Jovanovich.
———. 1976. Maria Sabina and her Mazatec mushroom velada. New York: Harcourt.
Wheeler, Q. D., and M. Blackwell. 1984. Fungus–insect relationships: Perspectives in ecology and evolution. New York: Columbia University Press.
White, M. M., T. Y. James, K. O’Donnell, M. J. Cafaro, Y. Tanabe, and J. Sugiyama. 2006. Phylogeny of the Zygomycota based on nuclear ribosomal sequence data. Mycologia 98:872–884.
Wilding, N., N. M. Collins, P. M. Hammond, and J. F. Webber. 1989. Insect–fungus interactions. New York: Academic Press.
Zhang, N., and M. Blackwell. 2001. Molecular phylogeny of dogwood anthracnose fungus (Discula destructiva) and the Diaporthales. Mycologia 93:356–364.
THE FUNGI: 1, 2, 3 … 5.1 MILLION SPECIES?11,12,13
Premise of the Study
Fungi are major decomposers in certain ecosystems and essential associates of many organisms. They provide enzymes and drugs and serve as experimental organisms. In 1991, a landmark paper estimated that there are 1.5 million fungi on the Earth. Because only 70000 fungi had been described at that time, the estimate has been the impetus to search for previously unknown fungi. Fungal habitats include soil, water, and organisms that may harbor large numbers of understudied fungi, estimated to outnumber plants by at least 6 to 1. More recent estimates based on high-throughput sequencing methods suggest that as many as 5.1 million fungal species exist.
Methods
Technological advances make it possible to apply molecular methods to develop a stable classification and to discover and identify fungal taxa.
_____________________
11 Reprinted with kind permission from the Botanical Society of America, www.amjbot.org.
12 Manuscript received 10 August 2010; revision accepted 19 January 2011.
13 Key words: biodiversity; fungal habitats; fungal phylogeny; fungi; molecular methods; numbers of fungi.
14 Department of Biological Sciences; Louisiana State University; Baton Rouge, Louisiana 70803 USA.
15 Author for correspondence (e-mail: mblackwell@lsu.edu) doi:10.3732/ajb.1000298.
Key Results
Molecular methods have dramatically increased our knowledge of Fungi in less than 20 years, revealing a monophyletic kingdom and increased diversity among early-diverging lineages. Mycologists are making significant advances in species discovery, but many fungi remain to be discovered.
Conclusions
Fungi are essential to the survival of many groups of organisms with which they form associations. They also attract attention as predators of invertebrate animals, pathogens of potatoes and rice and humans and bats, killers of frogs and crayfish, producers of secondary metabolites to lower cholesterol, and subjects of prize winning research. Molecular tools in use and under development can be used to discover the world’s unknown fungi in less than 1000 years predicted at current new species acquisition rates.
What are Fungi?
Fungal biologists debated for more than 200 years about which organisms should be counted as Fungi. In less than 5 years, DNA sequencing provided a multitude of new characters for analysis and identified about 10 phyla as members of the monophyletic kingdom Fungi (Fig. A3-1). Mycologists benefited from early developments applied directly to fungi. The “universal primers,” so popular in the early 1990s for the polymerase chain reaction (PCR), actually were designed for fungi (Innis et al., 1990; White et al., 1990). Use of the PCR was a monumental advance for those who studied minute, often unculturable, organisms. Problems of too few morphological characters (e.g., yeasts), noncorresponding characters among taxa (e.g., asexual and sexual states), and convergent morphologies (e.g., long-necked perithecia producing sticky ascospores selected for insect dispersal) were suddenly overcome. Rather than producing totally new hypotheses of relationships, however, it is interesting to note that many of the new findings supported previous, competing hypotheses that had been based on morphological evidence (Alexopoulos et al., 1996; Stajich et al., 2009). Sequences and phylogenetic analyses were used not only to hypothesize relationships, but also to identify taxa rapidly (Kurtzman and Robnett, 1998; Brock et al., 2009; Begerow et al., 2010).
Most fungi lack flagella and have filamentous bodies with distinctive cell wall carbohydrates and haploid thalli as a result of zygotic meiosis. They interact with all major groups of organisms. By their descent from an ancestor shared with animals about a billion years ago plus or minus 500 million years (Berbee and Taylor, 2010), the Fungi constitute a major eukaryotic lineage equal in numbers to animals and exceeding plants (Figs. A3-2–10). The group includes molds, yeasts, mushrooms, polypores, plant parasitic rusts and smuts, and Penicillium chrysoge-
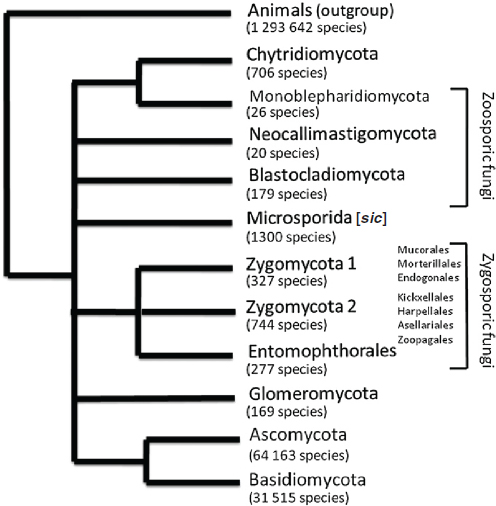
Figure A3-1 Fungal phyla and approximate number of species in each group (Kirk et al., 2008). Evidence from gene order conversion and multilocus sequencing indicates that microsporidians are Fungi (see below; Lee et al., 2010). Note also that zoosporic and zygosporic fungal groups are not supported as monophyletic. Tree based on Hibbett et al. (2007), White et al. (2006), and James et al. (2006).
num, Neurospora crassa, Saccharomyces cerevisiae, and Schizosaccharomyces pombe, the important model organisms studied by Nobel laureates.
Phylogenetic studies provided evidence that nucleriid protists are the sister group of Fungi (Medina et al., 2003), nonphotosynthetic heterokont flagellates are placed among brown algae and other stramenopiles, and slime mold groups are excluded from Fungi (Alexopoulos et al., 1996). Current phylogenetic evidence suggests that the flagellum may have been lost several times among the early-diverging fungi and that there is more diversity among early diverging
zoosporic and zygosporic lineages than previously realized (Bowman et al., 1992; Blackwell et al., 2006; Hibbett et al., 2007; Stajich et al., 2009).
Sequences of one or several genes are no longer evidence enough in phylogenetic research. A much-cited example of the kind of problem that may occur when single genes with different rates of change are used in analyses involves Microsporidia. These organisms were misinterpreted as early-diverging eukaryotes in the tree of life based on their apparent reduced morphology (Cavalier-Smith, 1983). Subsequently, phylogenetic analyses using small subunit ribosomal RNA genes wrongly supported a microsporidian divergence before the origin of mitochondria in eukaryotic organisms (Vossbrinck et al., 1987). More recent morphological and physiological studies have not upheld this placement, and analyses of additional sequences, including those of protein-coding genes, support the view that these obligate intracellular parasites of insect and vertebrate hosts are members of the Fungi (Keeling, 2009; Corradi and Keeling, 2009). Additional evidence from genome structure as well as phylogenetic analyses, supports the inclusion of microsporidians within the Fungi and indicates that comparison of whole genomes contributes to the solution of challenging phylogenetic problems (Lee et al., 2010).
The level of resolution and sophistication of systematics studies made possible by molecular markers and phylogenetic analyses put mycologists on equal footing with other biologists for competitive funding, and they joined in several community-wide efforts to organize fungal diversity within a phylogenetic classification. Three projects funded by the National Science Foundation were initiated, including the Research Coordination Network: A Phylogeny for Kingdom Fungi (Deep Hypha) and successive Tree of Life projects, Assembling the Fungal Tree of Life (AFTOL-1) and a second ongoing project (AFTOL-2) (Blackwell et al., 2006). A major product of the Deep Hypha project was the publication of 24 papers on fungal phylogeny in a single journal issue (Mycologia 98: 829–1103). The papers included an introduction to progress in fungal phylogeny, a paper on dating the origin of Fungi, one on the evolution of morphological traits, and 21 articles with multilocus phylogenies of most major groups. Participants included 156 authors with some involved in more than one paper; only 72 of the authors were originally from North America. The multi-investigator AFTOL-1 publication (Hibbett et al., 2007) included a widely used and often cited phylogenetic classification to the level of order (e.g., Kirk et al., 2008; The NCBI Entrez Taxonomy Home-page, http://www.ncbi.nlm.nih.gov/taxonomy; Science Watch, http://sciencewatch.com/dr/nhp/2009/09jannhp/09jannhpHibb). The paper included 68 authors from more than 20 countries.
It is important to note that there was broad participation and, essentially, global involvement on these projects, emphasizing that studies of biodiversity are indeed global endeavors. Additional pages were contributed to the Tree of Life web project (http://www.tolweb.org/Fungi/2377) to make information on fungi more accessible to students and the general public. Two objectives of the ongoing AFTOL-2 project include increased taxon sampling of fungi for molecular
FIGURES A3-2–10 Examples of fungal diversity. 2. Lemonniera sp. Tetraradiate conidia developed on a submerged leaf in a well-aerated freshwater stream surrounded by lush vegetation. This type of aquatic species, an Ingoldian ascomycete, is named for C. T. Ingold, who pioneered the study of these fungi, that are characterized by highly branched conidia. Photo courtesy of H. Raja. 3. The aero-aquatic ascomycete Helicoon gigantisporum produces distinctive tightly coiled conidia. As the spore develops air is trapped in the coil and causes it to be buoyant. This feature is an adaptation for the polyphyletic aero-aquatic fungi that grow on leaves in slow-moving or stagnant freshwater. Photo courtesy of H. Raja. 4. The smut Testicularia sp. develops in the ovary of grasses and (as shown here) sedges. The spores mature sequentially, with the dark spores being more mature. A plant taxonomy student once thought he had discovered a new species of Leersia, distinguished by large ovaries of ca. 1 cm, only to be disappointed that the enlargement was caused by a fungus. It is helpful to mycologists when plant taxonomists collect and accession fungal diversity by selecting some diseased plant specimens, an activity that should be encouraged. 5. Perithecia of Pyxidiophora sp. (Laboulbeniomycetes) developed in moist chamber on moose dung from Meredith Station, New Brunswick, Canada. The 150 µm long ascospores are seen at the tip of the perithecium neck in the center. Spores adhere to phoretic mites that are carried by dung beetles to fresh dung piles. Some fungi have complex animal dispersal systems. Pyxidiophora species are usually mycoparasites that grow on fungi in dung or other substrates including wrack washed up on beaches. The genus is a “missing link” and provided clues to confirm that Laboulbeniomycetes are ascomycetes and not other kinds of fungi or floridian red algae. 6. The ca. 8 cm wide basidiomata of Pycnoporus sp., a wide-ranging, brightly colored, wood-decaying polypore, photographed at Barro Colorado Island, Panama. Some collectors have referred to basidiomycetes that produce colorful basidiomata as charismatic megamycota of the fungus world. 7. Peniophorella baculorubrensis, a bark-decaying basidiomycete common on and restricted to living live oak (Quercus virginiana), decays the bark and changes its water-holding capacity. The effect of decay on bryophyte communties by this fungus was first studied by ecologists (Penfound and Mackaness, 1940) more than 70 yr ago but was not described until a specialist on wood-decaying fungi happened to notice it on the Louisiana State University campus, Baton Rouge (Gilbertson and Blackwell, 1984). The inconspicuous basidiomata are shown growing on the lower side of a 7 cm long bark segment aimed downward for basidiospore discharge in response to gravity. 8. Basidiomata of Perenniporia phloiophila on the bark of living Quercus virginiana. Although the basidiomata are obvious against the darker bark, this species was not described until it was discovered at the same time and often on the same trees as Peniophorella baculorubrensis. Although the fungus usually rots only the outer bark, it will invade and decay wood whenever the vascular cambium is broached by a bird or insect. In addition to the two species on live oak, six other species have been described from the campus, illustrating the need for specialists to study noncharismatic fungi. 9. A basidioma (8 cm diameter) of the wood-decaying fungus, Favolus tenuiculus, a favorite food of several species of mushroom-feeding beetles (see Fig. A3-10). Photo courtesy of N. H. Nguyen. 10. The small (>10 mm long) brightly colored beetle, Mycotretus sp. (Erotylidae), was collected at Barro Colorado Island, Panama. Many erotylid beetles have specialized yeast-packed pouches at the anterior end of the midgut. More than 200 novel yeasts have been isolated from the gut of ca. 15 families of mushroom-feeding beetles (Suh et al., 2005). Photo courtesy of James A. Robertson.
data and the discovery of correlated morphological and biochemical characters (AFTOL Structural and Biochemical Database, https://aftol.umn.edu; Celio et al., 2006).
Known Fungal Species
The Dictionary of Fungi (Kirk et al., 2008) reported 97330 species of described fungi at the “numbers of fungi” entry. The addition of 1300 microsporidians brings the total of all described fungi to about 99000 species (Fig. A3-1). The Dictionary’s estimate of known species has almost tripled in the period between the first edition in 1943 (38000 described species) and now, amounting to an increase of more than 60000 described species over the 65-yr period (Fig. A3-11). Factors such as difficulty of isolation and failure to apply molecular methods may contribute to lower numbers of species in certain groups, but there cannot be any doubt that ascomycetes and basidiomycetes comprise the vast majority of fungal diversity (Fig. A3-1).
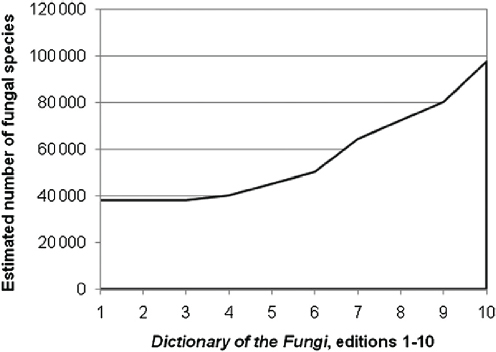
Figure A3-11 Numbers of known fungi from the Dictionary of the Fungi (editions 1–10, 1950–2008). Authors state that the large increase in species numbers in the 10th edition may be inflated because asexual and sexual forms were counted separately and molecular techniques that distinguish close taxa have been used.
Estimated total fungal numbers
In 1991, a landmark paper provided several qualified estimates of the number of fungi on the Earth based on ratios of known fungi to plant species in regions where fungi were considered to be well-studied (Hawksworth, 1991). “Estimate G” of 1.5 million species was accepted as a reasonable working hypothesis based on a fungus to plant ratio of 6:1, in contrast to the much lower 50–60-yr-old estimates by Bisby and Ainsworth (1943) of 100000 fungal species and by Martin (1951) of 250000 species based on one fungus for every phanerogam known at the time. A more recent estimate of the total number of fungi, 720 256 (Schmit and Mueller, 2007), is also low compared to present estimates that include environmental samples.
Hawksworth’s (1991) estimate now is considered to be conservative by many, including Hawksworth (Hawksworth and Rossman, 1997), because numerous potential fungal habitats and localities remain understudied (Hawksworth, 2001). Furthermore, the use of molecular methods had not yet been considered as a means of species discovery. For example, analysis of environmental DNA samples from a soil community revealed a high rate of new species accumulation at the site, and these data supported an estimate of 3.5 to 5.1 million species (O’Brien et al., 2005). Using the present discovery rate of about 1200 fungal species per year based on the last 10 years, Hibbett and his colleagues (in press) estimated that it would take 1170 years to describe 1.4 million fungi (based on Estimate G of Hawksworth [1991]) and 2840 to 4170 yr to describe 3.5 to 5.1 million (based on O’Brien et al., 2005).
Using present higher estimates of land plant numbers as somewhat under 400000 species (Paton et al., 2008; Joppa et al., 2010) fungal species numbers now are expected to outnumber land plants by as much as 10.6:1 based on O’Brien et al. (2005). Even higher ratios have been predicted using data from highthroughput sequencing of clone libraries, although individual ecosystems will vary (L. Taylor, University of Alaska, Fairbanks, personal communication, January 2011). The large gap between known and estimated species numbers has led to a series of papers and symposia (e.g., Hawksworth and Rossman, 1997; Hawksworth, 2001; Hyde, 2001; Mueller and Schmit, 2007) attempting to answer the question “Where are the missing fungi?”
How to Discover New Fungi
Collecting and culturing fungi from the environment will remain important because of the need to identify specimens, revise taxonomy, assess the roles in the environment, and provide strains for biological control, environmental remediation, and industrial processes. A physical specimen, including an inert culture, is still required as a type specimen (but see Conclusions later), and vouchers of known fungi are used for documenting DNA sequences deposited in some databases (Nilsson et al., 2006). For example, the current AFTOL project has
a requirement that each sequence deposited as part of the project be linked to a specimen, including a culture.
All taxa biological inventories (ATBIs) attempt to survey organisms within particular geographical regions by collection of specimens and culture of substrates. One of these, Discover Life in America, All Taxa Biological Inventory, seeks to survey an estimated 50000 to 100000 species of organisms in the Great Smoky Mountains National Park. Karen Hughes and Ronald Petersen have been successful in collecting more than 3000 species of fungi, mostly agarics housed in the University of Tennessee Fungal Herbarium (http://tenn.bio.utk.edu/fungus/database/fungus-browse-results.asp?GSMNP=GSMNP), out of about 17000 species of all taxa that have been collected by others in the park (Biodiversity Surveys and Inventories: Agaric Diversity in the Great Smoky Mountains National Park, NSF DEB 0338699). All fungal specimens have been identified, and the agarics have been studied to the extent that a culture, ITS barcode sequence, and genetic analysis are available for many species. This successful project has required hours of time over a number of years and costly resources for studying the material, but it serves as an example of the commitment needed to acquire specimen-based information on fungi.
DNA methodology makes it possible to use independent sampling methods to discover the presence of organisms without ever seeing a culture or a specimen. Several new methods significantly outperform previous automated sequencing methods (e.g., Jumpponen and Jones, 2009; Metzker, 2010). Although there may be certain limitations and biases for the different methods (Amend et al., 2010a; Tedersoo et al., 2010), mycologists have been quick to embrace them in ecological and biodiversity studies. O’Brien and colleagues (2005) pointed out that collection and culture methods revealed numbers of fungi similar to those acquired by sampling environmental DNA. Hibbett et al. (in press), however, used data from GenBank to show that by 2008 and 2009 the number of environmental samples, excluding overwhelming numbers of sequences discovered by pyrosequencing, exceeded the accessions of specimen-based sequences. The rapid development of automated, high-throughput methods also has made it possible to acquire whole genome sequences for population level studies (Liti et al., 2009; Neafsey et al., 2010).
Which Regions of the Earth Harbor Fungal Diversity?
Fungi grow in almost all habitats on Earth, surpassed only by bacteria in their ability to withstand extremes in temperature, water activity, and carbon source (Raspor and Zupan, 2006). Tropical regions of the world are considered to have the highest diversity for most groups of organisms (Pianka, 1966; Hillebrand, 2004), and this is generally true for fungi as well (Arnold and Lutzoni, 2007).
A group of researchers are studying the diversity of the Guyana Shield. For the last 11 years, Terry Henkel and Cathie Aime and their colleagues have studied the fungi in six 1-km2 plots—three in a Dicymbe corymbosa-dominated
forest and three in a mixed tropical forest. Their current collections contain 1200 morphospecies, primarily basidiomycetes. Approximately 260 species were collected repeatedly only in the Dicymbe plots. Thus far, two new genera and ca. 50 new species have been described. On the basis of groups already studied, Aime estimated that ca. 120 new ectomycorrhizal taxa have been discovered. Including novel saprobes as well as ectomycorrhizal fungi, ca. 500 new species are expected among the 1200 taxa collected. It is clear, however, that these are not simply high numbers of new taxa, but biologically interesting fungi as well (Aime et al., 2010). One species is so unusual, that a reviewer of the original report called it “the find of the century” (Redhead, 2002). As Aime has quipped “if one were to compare the ratio of fungi to plants in the Dicymbe plots as did Hawksworth (1991), the ratio would be 260 to 1, obviously an overestimate but also a cautionary exercise in basing any estimate on a single ecotype” (M. C. Aime, Louisiana State University, personal communication, August 2010).
Many fungi have in fact come from temperate regions, and some studies report a high diversity of fungi. For example, in a study of indoor air from buildings using culture-independent sampling methods, diversity was found to be significantly higher in temperate sites independent of building design or use. The authors also alluded to the possibility that previous studies of certain mycorrhizal fungi showed similar trends (Amend et al., 2010b). More investigation in this area is needed, but it is clear that many undescribed fungi are present in temperate regions. Popular literature often rationalizes the need to save the rainforests, not because of their intrinsic value, but because of the potential drug-producing organisms that may be found there. Many of the commercially most successful fungal drugs, however, come from temperate fungi. Penicillium chrysogenum, producer of penicillin, was found in a northern temperate city. Another remarkable fungus, Tolypocladium infl atum from Norwegian soil, synthesizes cyclosporine, an immune-suppressant drug that revolutionized organ transplants (Borel, 2002); the sexual state of this fungus was collected in New York, USA (Hodge et al., 1996). Today the drug is commonly used to treat dry eye (Perry et al., 2008), as well as many serious conditions. Statins produced by fungi such as Aspergillus terreus from temperate regions, combat high cholesterol levels, as well as providing other benefits (Vaughan et al., 1996; Askenazi et al., 2003; Baigent et al., 2005).
In temperate deserts, mycorrhizal boletes, agarics, and rust and smut fungi, are common. A surprising number of wood-decaying basidiomycetes have been discovered on living and dead desert plants, including cacti and are in the University of Arizona, Robert L. Gilbertson Mycological Herbarium (http://ag.arizona.edu/mycoherb/herbholdings). When a noted mycologist moved to Arizona early in his career, he became excited about the new and unreported fungal diversity found in the desert. His proposed study of the wood-decaying fungi of the Sonoran Desert was poorly received with a comment that wood-decaying fungi were not present in the desert (R. L. Gilbertson, University of Arizona, personal communication, August 1979). The Sonoran Desert, however, has many plants
(e.g., cacti, ocotillo, and mesquite and other desert legumes) that are substrates for polypores and resupinate basidiomycetes (e.g., Gilbertson and Ryvarden, 1986, 1987).
Fungi also grow at low temperatures. An example involves fungal deterioration of historic huts built between 1901 and 1911 for use by Antarctic explorers including Robert Scott and Ernest Shackleton, and although there are not large species numbers, it is important not to overlook this fungal habitat in diversity studies (Held et al., 2005). Lichens have often been reported to be common in Arctic and Antarctic regions (Wirtz et al., 2008), and yeasts are active under frozen conditions in the Antarctic (Vishniac, 2006; Amato et al., 2009). In some cases, a yeast isolated from the Antarctic (based on 28S rDNA barcoding) also has been reported from varied habitats, including human infections, the gut of insects, deep seas, and hydro-carbon seeps (Kurtzman and Fell, 1998; Bass et al., 2007; personal observation). Although some fungi are specialized for cold regions, others simply occupy a wide variety of environmental conditions.
Many regions and habitats of the world need to be included in fungal discovery. In general, microscopic fungi and those that cannot be cultured are very poorly known. Parts of Africa remain to be collected for many, although not all, fungal groups (Crous et al., 2006). Fungi are important as symbionts, and they are associated with every major group of organisms, bacteria, plants and green algae, and animals including insects. Because certain under-studied symbiotic associations are known to include large numbers of fungi, these are a good place to search for new taxa. The associated organisms also allow for resampling, a quick way to obtain data about host specificity. Targeting hosts also is a productive method for discovering fungal fossils, such as those associated with plants of the Rhynie Chert (Taylor et al., 2004). Examples of diversity in particular fungal habitats are reviewed in the following sections.
Fungi and Plant Roots
Mycorrhizal plants and their fungal partners have been studied by a number of mycologists (Trappe, 1987; Smith and Read, 2008). The fungi often are essential to their plant hosts because they take up water, nitrogen, phosphorus, and other nutrients from the soil and transfer them to the plant roots. Some of these fungi may not prosper or even grow without the host. In addition to flowering plants and conifers, many bryophytes and ferns are mycorrhizal (Pressel et al., 2010). Certain mycorrhizal fungi specialize on orchids and ericoid plants, and some are known to have invaded new habitats with successful invasive plants (Pringle et al., 2009).
There are two main types of mycorrhizal fungi, arbuscular mycorrhizae (AM) and ectomycorrhizae. AM associations are more common and occur with up to 80% of all plant species and 92% of plant families. AM fungi are all members of the phylum Glomeromycota, a less diverse group than ectomycorrhizal
fungi with about 250 described species in a variety of taxa (Gerdemann, 1968; Schüssler and Walker, 2011; Wang and Qiu, 2006). Evidence from recent molecular studies, however, indicates that cryptic species with higher levels of host specificity than previously realized will increase the number of known AM fungi (Selosse et al., 2006; Smith and Read, 2008). More than 6000 species, mostly of mushroom-forming basidiomycetes, form ectomycorrhizae with about 10% of all plant families. Greater host specificity usually occurs in the ectomycorrhizal fungus–plant associations than in AM associations (Smith and Read, 2008). Vast parts of the world remain to be sampled (Mueller et al., 2007), and it is expected that barriers to inter-breeding have led to high genetic diversity among these fungi (Petersen and Hughes, 2007).
Inside Plant Leaves and Stems
Almost all plants on Earth are infected with endophytes, fungi that do not cause disease symptoms (Saikkonen et al., 1998). Endophytes occur between the cells, usually of above ground plant parts, and represent a broad array of taxonomic groups (Arnold, 2007; Rodriguez et al., 2009). The earliest studies of endophytes were of those associated with grasses (Diehl, 1950). Some grass endophytes are specialized members of the Clavicipitaceae, relatives of insect and fungal parasites in the Hypocreales, and many species produce alkaloid toxins effective against insects, other invertebrate animals, and vertebrates (Clay et al., 1993). Some grass endophytes are transmitted to the host offspring in seeds, and others inhibit sexual reproduction in the host and are dispersed within plant parts such as leaf fragments. For grass endophytes that reproduce sexually, fertilization may occur by insect dispersal. Water intake is increased in infected hosts, and these plants often grow taller than uninfected hosts.
A much more diverse group of endophytic fungi are associated with plants in addition to grasses, including a variety of dicots and conifers (Carroll, 1988; Rodriguez et al., 2009). In some tropical forests considered to be diversity hotspots for endophytes, there are extremely large numbers of the fungi, sometimes with hundreds reported from a single tree species, judged by both cultural and molecular methods of discovery and identification (Arnold et al., 2001; Arnold and Lutzoni, 2007; Pinruan et al., 2007; Rodriguez et al., 2009). In one study, more than 400 unique morphotypes were isolated from 83 leaves of two species of tropical trees. A subset of the fungi was distributed among at least seven orders of ascomycetes (Arnold et al., 2000). Leaves usually acquired multiple infections as they matured, and there was strong evidence that the endophytes protected leaves of plants, such as Theobroma cacao, from infection when they were challenged with pathogens (Arnold et al., 2003). Vega and colleagues (2010) also found high diversity of endophytes in cultivated coffee plants. Interestingly, some of these were insect pathogens and experiments are being conducted to develop endophytes as biological control agents of insect pests.
Plant Pathogens
Plant pathogens differ from endophytes in that they cause disease symptoms. Although some zoosporic and zygosporic fungi are plant pathogens, most plant pathogens are ascomycetes and basidiomycetes. A large number of ascomycetes and ca. 8000 species of basidiomycetes are plant pathogens. In addition to crop pathogens, it is important to remember that many pathogens are numerous and important in natural ecosystems (Farr et al., 1989; Burdon, 1993). Nonpathogenic phylloplane yeasts occupy leaf surfaces of many plants and are increasingly recognized for their control of potential leaf pathogens (Fonseca and Inácio, 2006). In addition to the thousands of native fungi that parasitize plants in the United States, pathologists are constantly on the lookout for introduced pathogens that often are undescribed when they arrive to decimate naïve native plant populations. For example, invasive fungi such as those grouped as Dutch elm disease fungi, chestnut blight fungus, dogwood anthracnose fungus, and redbay wilt fungus, were all unknown until they were observed soon after their introduction (Alexopoulos et al., 1996; Zhang and Blackwell, 2001; Harrington et al., 2008). Exotic localities will need to be searched for undescribed fungi that probably go largely unnoticed on their native hosts. It is important to note that although fungi may cause only minor symptoms to hosts in their native habitats, one of these may have the potential to be the next destructive disease after introduction to a new region.
Molecular methods have helped to clarify limits of closely related species and to establish host ranges (e.g., Crous et al., 2008). In a study of 26 leaf spot fungi in Australia, three genera of Myrtaceae, including Eucalyptus, were hosts for three new genera and 20 new species (Cheewangkoon et al., 2009). Although the authors acknowledged the high level of new taxa discovered, they pointed out that the potential for host shifts within plantations might lower estimates of fungal species numbers worldwide. Host or substrate specificity is a concept that can be applied to fungal groups that are closely associated with hosts such as endophytes, pathogens, and mycorrhizal fungi but not usually for saprobic species (Zhou and Hyde, 2001). In the past species of plant pathogens often were based on host identity, a practice that is not always effective because some groups are host-specific while others are not.
Lichens and Lichenicolus Fungi
About 20% of all fungi and 40% of the ascomycetes (13500 species) are lichen-forming fungi (Lutzoni and Miadlikowska, 2009). Lichenicolous fungi, parasites, and other associates of lichens are not well collected, but an estimate for the combined lichens and lichenicolous fungi is about 20000 species (Feuerer and Hawksworth, 2007). Lichens and lichenicolous fungi are polyphyletic, and several different groups of ascomycetes and a few species of basidiomycetes have become associated with green algae and cyanobacteria (Lutzoni and
Miadlikowska, 2009). Feuerer (2010) can be consulted for information on lichen diversity worldwide. This checklist also highlights the absence of collections in certain regions.
Deserts are rich in lichens. Of 1971 lichen species and associated fungi reported from the Sonoran Desert, about 25% studied since 1990 are new. Three volumes on lichens of the greater Sonoran Desert region have been published (Nash et al., 2002, 2004). Other habitats of high lichen diversity are Arctic and Antarctic regions (Feuerer, 2010).
Fungi From Arthropod and Invertebrate Animals
There is a need for more information on arthropod- and insect-associated fungi. As was mentioned earlier, estimates of global fungal diversity usually omit insect-associated species because they are so poorly known (Hawksworth, 1991; Rossman, 1994; Mueller and Schmit, 2007; Schmit and Mueller, 2007). Several post-1991 estimates of insect-associated fungi suggested that 20 000–50 000 species exist (Rossman, 1994; Weir and Hammond 1997a, b; Schmit and Mueller, 2007). Some parasites are biotrophic, associated with living insects, and many do not grow in culture. These also usually require special methods for removal and mounting, and few mycologists or entomologists have ever seen members of the Laboulbeniomycetes or the fungal trichomycetes, Asellariales and Harpellales (Lichtwardt et al., 2001; Cafaro, 2005). Laboulbeniomycetes are seta-sized, ectoparasitic ascomycetes of insects, mites, and millipedes (Weir and Blackwell, 2005). All 2000 known species have distinctive life cycles with determinate thalli arising from two-celled ascospores. About 90% of the species have been found on adult beetles (12 of 24 superfamilies) or on flies. New arthropod hosts at the level of family are still being discovered (Weir and Hammond, 1997a, b; Rossi and Weir, 2007), and there is an indication that there is some degree of host specificity (De Kesel, 1996). In the future, increased use of molecular methods will make it possible to determine the degree of species level host specificity, but the information is not available now. Septobasidiales, relatives of the basidiomycete rust fungi are associated with scale insects, and their felty basidiomata presumably protect the insects from parasitoid wasps. Many microsporidians also are parasites of a broad group of host insects.
Necrotrophic parasites of insects include some members of Chytridiomycota, Blastocladiales (Coelomomyces), Entomophthorales, and Tubeufiaceae (Podonectria) (Benjamin et al., 2004). About 5000 members of three families of Hypocreales are necrotrophic parasites of arthropods (Spatafora et al., 2007, 2010). These species show an evolutionary pattern of host shifting among plants, fungi, and insects in addition to displaying a high level of host specificity.
Fungi also occur in ancient, obligate gardening associations with bark and ambrosia beetles, attine ants, and Old World termites, and new species are still being discovered in these groups (Benjamin et al., 2004; Little and Currie, 2007;
Harrington et al., 2008; Aanen et al., 2009). Many yeasts are associated with insects, particularly insects that feed on nectar (Lachance, 2006; Robert et al., 2006).
Other insects contain gut yeasts, a habitat where few have looked for them. Isolations from the gut of mushroom-feeding beetles yielded up to 200 new species of yeasts (Suh et al., 2004, 2005; see also Lachance et al., 2010). Because only about 1500 ascomycete yeasts (Saccharomycotina) have been described, the gut yeasts represent a dramatic increase in diversity from a limited geographical range (Boekhout, 2005; C. Kurtzman, USDA-ARS, personal communication, July 2010). In fact, the estimated total number of yeast species worldwide could be increased by as much as 50% by simply recollecting in previously collected sites from the study (Suh et al., 2005). As Lachance (2006) pointed out, based on predictions of yeast numbers using data from species in slime fluxes and in associations with flower-visiting insects, it is necessary to obtain more information on specificity and geographical ranges before better estimates can be made. Although not all insects harbor large numbers of yeasts in their guts, those with restricted diets in all life history stages such as mushrooms or wood are often associated with yeasts. Host insects may acquire digestive enzymes or vitamins from the yeasts. This contention is supported by the fact that unrelated insects feeding on mushrooms (e.g., beetles in different lineages, lepidopteran larvae) all have gut yeasts with similar assimilative capabilities and vitamin production. The high rate of discovery of yeasts in under-collected habitats and localities suggests that far more taxa await discovery (Suh et al., 2005), and the gut habitat has been considered a yeast diversity hotspot (Boekhout, 2005).
Insects may be food for fungi, especially in low nitrogen environments. The mycelium of Pleurotus ostreatus, a favorite edible species for humans, secretes toxic droplets that kill nematodes. A study involving the mushroom-producing, ectomycorrhizal basidiomycete, Laccaria bicolor, was designed to determine the amount of predation by springtails on the fungal mycelium. The study led to the surprise discovery that the fungus was not insect food, but rather, it, and indirectly, the host tree benefited by obtaining substantial amounts of nitrogen from the insects (Klironomos and Hart, 2001). The predatory habit has arisen independently on several occasions in at least four phyla of fungi and oomycetes. Predaceous fungi such as species of Arthrobotrys and Dactylella lure, then trap, snare, or grip nematodes and other small invertebrate animals in soils and in wood (Barron, 1977).
Ødegaard (2000) revised global estimates of arthropods downward from 30 million to 5–10 million. Not all insects and arthropods are tightly associated with fungi, but even the revised species estimates indicate that the numbers of insect-associated fungi will be very high.
Soil Fungi
Soil is a habitat of high fungal diversity (Waksman, 1922; Gilman, 1957; Kirk et al., 2004; Domsch et al., 2007). Soil fungi and bacteria are important in biogeochemical cycles (Vandenkoornhuyse et al., 2002), and the diversity of soil fungi is highest near organic material such as roots and root exudates. Per volume, large numbers of microscopic fungi occur in pure soil, and these are largely asexual ascomycetes and some zygomycetes, including animal-associated Zoopagales. Gams (2006) estimated that 3150 species of soil fungi are known, and ca. 70% are available in culture. There presently is a high rate of new species acquisition, and the group appears to be better known than most ecologically defined groups. Molecular studies, however, are predicted to increase the total number (Bills et al., 2004). In fact a study of soil communities in several forest types at the Bonanza Creek Long Term Ecological Research site, Fairbanks, Alaska, United States, revealed not only seasonal changes in community composition but also in dominance of fungi over bacteria. The data acquired by several molecular methods including high-throughput sequencing greatly increased the total number of fungal sequences in GenBank at the time (Taylor et al., 2010). Taylor and his colleagues found more than 200 operational taxonomic units in a 0.25 g soil sample with only 14% overlap in a sample taken a meter away. This study is not directly comparable with the soil fungi reported by Gams (2006) because Gams’ figures excluded fungi such as mycorrhizal species.
Another study of soil fungi based on environmental DNA sequences showed an unexpected distribution of a group of zoosporic fungi, Chytridiomycota. The chytrids, were found to be the predominate group of fungi in nonvegetated, high-elevation soils at sites in Nepal and in the United States in Colorado, where more than 60% of the clone libraries obtained were from chytrids. A phylogenetic analysis of the sequences compared with those of a broad selection of known chytrids, indicated that a diverse group of Chytridiomycota representing three orders was present (Freeman et al., 2009).
Most major fungal lineages are known from cultures and specimens, but there have been a few surprises even in well-sampled habitats such as soil. Soil clone group I (SCGI) represents a major lineage of fungi that occurs in temperate and tropical soils on three continents, but no one has ever seen or isolated any of the species into culture (Schadt et al., 2003; Porter et al., 2008).
The phylogenetic position of this lineage, perhaps a new phylum, appeared as a sister group to the clade of Pezizomycotina–Saccharomycotina (Porter et al., 2008). Other unexpected higher taxonomic level fungal clades have been detected from environmental DNA sequences (Vandenkoornhuyse et al., 2002; Jumpponen and Johnson, 2005; Porter et al., 2008). Another lineage detected by environmental sequences was subjected to fluorescent in situ hybridization (FISH). The outline of a single-celled, flagellated organism was detected (Jones and Richards, 2009), but apparently none of these fungi has been cultured either. Higher-level
bacterial taxa have been discovered by environmental sampling, but this is a far less common occurrence for fungi (Porter et al., 2008).
Fungi form crusts that stabilize desert soils. Crusts usually are made up of darkly pigmented ascomycetes, lichens, and nitrogen-fixing cyanobacteria (States and Christensen, 2001). Rock-inhabiting fungi occur in the surface and subsurface layers of desert rocks. These darkly pigmented ascomycetes are members of the classes Dothideomycetes and Arthoniomycetes, but basidiomycetes and bacteria may occur in the associations (Kuhlman et al., 2006; Ruibal et al., 2009). Easily cultured asexual ascomycetes and other fungi also occur in desert soils, and these include an unusual zygomycete, Lobosporangium transversale (Ranzoni, 1968), known only from three isolations including Sonoran Desert soil. Yeasts are well known from American deserts in association with cacti and flies where they detoxify plant metabolites (Starmer et al., 2006).
Freshwater Fungi
Certain fungi are adapted for life in fresh water. More than 3000 species of ascomycetes are specialized for a saprobic life style in freshwater habitats where they have enhanced growth and sporulation (Shearer et al., 2007; Kirk et al., 2008; Shearer and Raja, 2010). The asci are evanescent, and ascospores have appendages and sticky spore sheaths, that anchor the spores to potential substrates in the aquatic environment. Conidia have several dispersal strategies, and these are designated as Ingoldian (Fig. A3-2) and aero-aquatic (Fig. A3-3) conidia. Ingoldian conidia are sigmoidal, branched, or tetraradiate and attach to plants and other material in the water. The conidia float on foam that accumulates at the banks of streams, especially during heavy runoff, and when the bubbles burst, the spores may be dispersed for great distances from the water and into trees, where they can be isolated from water-filled tree holes (Bandoni, 1981; Descals and Moralejo, 2001; Gönczöl and Révay, 2003). Aero-aquatic fungi have multicellular, often tightly helical conidia with air spaces to make them buoyant on the surface of slower-moving waters (Fisher, 1977).
Other, less obviously modified fungi are present in water, and some of these are active in degrading leaves in streams after the heavy autumn leaf fall. A few specialized freshwater basidiomycetes also are known, and several have branched conidia similar to those of the Ingoldian ascomycetes. Flagellated fungi occur in aquatic habitats, including Chytridiomycota, Blastocladiomycota, and Monoblepharomycota (James et al., 2006). Batrachochytrium dendrobatidis, the recently described amphibian killer, is an aquatic chytrid (Longcore et al., 1999). Members of Neocallimastigomycota also live in a specialized largely aquatic environment, the gut of vertebrate herbivores, where they are essential for digestion of cellulosic substrates.
Marine Fungi
Marine waters provide a habitat for certain specialized fungi (Kohlmeyer and Volkmann-Kohlmeyer, 1991), and Hyde et al. (1998) estimated that more than 1500 species of marine fungi occur in a broad array of taxonomic groups. Many of these fungi are distinct from freshwater aquatic species, and they may be saprobic on aquatic plant substrates. Some species have characters such as sticky spore appendages, indicators of specialization for the marine habitat (Kohlmeyer et al., 2000).
It is interesting that few fungi from early-diverging lineages have been reported from marine environments, perhaps in part because mycologists studying these groups sampled more often from fresh water habitats. More recently, an investigation of deep-sea hydrothermal ecosystems revealed not only novel species of ascomycetes and basidiomycetes, but also what may be a previously unknown lineage of chytrids (Le Calvez et al., 2009).
Most marine fungi are ascomycetes and basidiomycetes, and these include ascomycete and basidiomycete yeasts (Nagahama, 2006). Some of the yeasts degrade hydrocarbon compounds present in natural underwater seeps and spills (Davies and Westlake, 1979). Certain ascomycetes are specialists on calcareous substrates including mollusk shells and cnidarian reefs. Even a few mushroom-forming basidiomycetes are restricted to marine waters (Binder et al., 2006). Some fungi use other marine invertebrates as hosts (Kim and Harvell, 2004), including antibiotic producers that live in sponges (Bhadury et al., 2006; Pivkin et al., 2006; Wang et al., 2008). A wide variety of fungi considered to be terrestrial also are found in marine environments. Basidiomycete (i.e., Lacazia loboi) and ascomycete yeasts, and other fungi including Basidiobolus ranarum, may occur in marine waters where they infect porpoises and other vertebrates (Kurtzman and Fell, 1998; Murdoch et al., 2008; Morris et al., 2010).
Fungal Species
Currently, molecular methods provide large numbers of characters for use in phylogenetic species discrimination (e.g., Kohn, 2005; Giraud et al., 2008). In the past, biologists relied primarily on phenotype for species delimitation, and most of the formally described species known today were based on morphology. In addition, mating tests have been used to distinguish so-called biological species, especially among heterothallic basidiomycetes (Anderson and Ullrich, 1979; Petersen, 1995). The ability to mate, however, may be an ancestral character. For example, Turner et al. (2010) found evidence that fungi have evolved strong barriers to mating when they have sympatric rather than allopatric distributions. Distant populations would not have had strong selective pressure against hybridization, thereby avoiding production of progeny less fit than conspecific progeny (e.g., Garbelotto et al., 2007; Stireman et al., 2010). This phenomenon, known as reinforcement, helps to explain how fungi from different continents can mate
in the laboratory but never in nature and is an argument in favor of recognizing species by phylogenetics. A number of researchers have recognized species using “phylogenetic species recognition” criteria (Taylor et al., 2000). The operational phylogenetic method is based on a “concordance of multiple gene genealogies,” and in addition to discriminating species, the method indicates whether fungal populations actually exchange genes in nature (Taylor et al., 2000; Fisher et al., 2002; Dettman et al., 2006; Jacobson et al., 2006).
The use of phylogenetic species criteria results in recognition of more species than those delimited by morphological characters. For example, work on Neurospora species resulted in the discovery of 15 species within five previously recognized species (Dettman et al., 2006; Villalta et al., 2009). There are many such examples among other groups of fungi, and eventually these may be a significant source of new species discovery in the effort to discover 5 million fungi. Fungal species recognized in this way may be described without a phenotypic diagnosis, but it is not uncommon for distinguishing characters to be found with guidance from the phylogenetics study (e.g., Otrosina and Garbelotto, 2010).
Conclusions
Until recently, estimates of numbers of fungi did not include results from large-scale environmental sequencing methods. Newer estimates based on data acquired from several molecular methods, however, have predicted as many as 5.1 million species of fungi (O’Brien et al., 2005; Taylor et al., 2010). Mycologists also are beginning to use high-throughput methods to gain insight into questions including geographical ranges and host and substrate specificity, topics that have direct bearing on species numbers (Lumbsch et al., 2008). For example, high-throughput methods have been used to determine the amount of overlap between species within a given region by comparing soil samples a meter apart to find only 14% species overlap (Taylor et al., 2010).
A better estimate of fungal numbers also can be speeded by enlisting more biologists to accomplish the goal. When amphibian populations first were observed to be dwindling and some species were determined to have disappeared almost 20 yr earlier, a number of causes, all nonfungal, were suggested as the explanation. The revelation that a chytrid was involved brought to mind that there were probably fewer than 10 mycologists in the world who could collect, isolate, culture, and identify the novel flagellated fungus, Batrachochytrium dendrobatidis (Longcore et al., 1999). Since that time interest in and publications on chytrids have increased dramatically (e.g., Freeman et al., 2009; LeCalvez et al., 2009). The interest in amphibian disease was in part the impetus for a large number of recent publications on amphibian decline, but amphibian decline also justified other projects, including training new chytrid systematists in monographic work. This effort has resulted in the discovery of many new chytrid species and the description of five new orders between 2008 and 2010. The rise of AIDS and the
accompanying large number of fungal infections brought about a similar interest in medical mycology several decades ago.
In addition to any sudden influx of biologists to obtain better estimates of fungal numbers, a new approach clearly is needed. In a thoughtful paper, Hibbett and colleagues (in press) called for obtaining clusters of similar sequences and assigning Latin binomials to these molecular operational taxonomic units (MOTUs). The names would allow the sequences to be integrated into a specimen-based taxonomic data stream. They considered inclusion of the sequence-based taxa among all taxa to be a better alternative than the candidate taxon status used by bacteriologists. Changes in the International Code of Botanical Nomenclature would be needed if sequence-based materials were to be allowed as nomenclatorial types. This proposal seems to be a practical approach to handling the overwhelming fungal diversity being discovered.
Recent experience in working as a broadly inclusive group to plan and produce a phylogenetic classification, the development of freely accessible databases, and the use of new tools to survey fungi in ecological studies has prepared the mycological community to accomplish a number of new goals, including the discovery of millions of fungi.
References
Aanen, D. K., H. H.De Fine Licht, A. J. M. Debets, N. G. Kerstes, R. F. Hoekstra, and J. J. Boomsma. 2009. High symbiont relatedness stabilizes mutualistic cooperation in fungus-growing termites. Science 326:1103–1106.
Aime, M. C., D. L. Largent, T. W. Henkel, and T. J. Baroni. 2010. The Entolomataceae of the Pakaraima Mountains of Guyana IV: New species of Calliderma, Paraeccilia and Trichopilus. Mycologia 102:633–649.
Ainsworth, G. C., and G. R. Bisby. 1943. Dictionary of the Fungi. Imperial Mycological Institute, Kew, UK.
Alexopoulos,C. J. C. W., Mims, and M. Blackwell.1996. Introductory mycology. Wiley, New York, New York, USA.
Amato, P., S. M. Doyle, and B. C. Christner. 2009. Macromolecular synthesis by yeasts under frozen conditions. Environmental Microbiology 11:589–596.
Amend, A. S. K. A. Seifert, and T. D. Bruns. 2010a. Quantifying microbial communities with 454 pyrosequencing: Does read abundance count? Molecular Ecology 10.1111/j.1365-294X.2010.04898.x.
Amend, A. S., K. A. Seifert, R.Samson, and T. D. Bruns. 2010b. Indoor fungal composition is geographically patterned and more diverse in temperate zones than in the tropics. Proceedings of the National Academy of Sciences, USA 107:13748–13753.
Anderson, J. B., and R. C.Ullrich, 1979. Biological species of Armillaria in North America. Mycologia 71:402–414.
Arnold, A. E. 2007. Understanding the diversity of foliar endophytic fungi: Progress, challenges, and frontiers. Fungal Biology Reviews 21:51–66.
Arnold, A. E., and F. Lutzoni. 2007. Diversity and host range of foliar fungal endophytes: Are tropical leaves biodiversity hotspots? Ecology 88:541–549.
Arnold, A. E., Z. Maynard, and G. S. Gilbert. 2001. Fungal endophytes in dicotyledonous neotropical trees: Patterns of abundance and diversity. Mycological Research 105:1502–1507.
Arnold, A. E., Z. Maynard, G. S. Gilbert, P. D. Coley, and T. A. Kursar. 2000. Are tropical fungal endophytes hyperdiverse? Ecology Letters 3:267–274.
Arnold, A. E., L. C. Mejía, D. Kyllo, E. Rojas, Z. Maynard, N.Robbins, and E. A.Herre. 2003. Fungal endophytes limit pathogen damage in leaves of a tropical tree. Proceedings of the National Academy of Sciences, USA 100:15649–15654.
Askenazi, M., E. M. Driggers, D. A. Holtzman, T. C. Norman, S. Iverson, D. P. Zimmer, M. E. Boers, et al. 2003. Integrating transcriptional and metabolite profiles to direct the engineering of lovastatin-producing fungal strains. Nature Biotechnology 21:150–156.
Baigent, C., A. Keech, P. M. Kearney, L. Blackwell, G. Buck, C. Pollicino, A. Kirby, et al. 2005. Efficacy and safety of cholesterol-lowering treatment: Prospective meta-analysis of data from 90,056 participants in 14 randomised trials of statins. Lancet 366:1267–1278.
Bandoni, R. J. 1981. Aquatic hyphomycetes from terrestrial litter. In D. T. Wicklow and G. C. Carroll [eds.], The fungal community: Its organization and role in the ecosystem, 693–708. Marcel Dekker, New York, New York, USA.
Barron, G. L. 1977. The nematode destroying fungi. Canadian Biological Publishers, Guelph, Ontario, Canada.
Bass, D., A. Howe, N. Brown, H. Barton, M. DeMidova, H. Michelle, L. Li, et al. 2007. Yeast forms dominate fungal diversity in the deep oceans. Proceedings of the Royal Soceity of London, B, Biological Sciences 274:3069–3077.
Begerow, D., H. Nilsson, M. Unterseher, and W. Maier, 2010. Current state and perspectives of fungal DNA barcoding and rapid identification procedures. Applied Microbiology and Biotechnology 87:99–108.
Benjamin, R. K., M. Blackwell, I. Chapella, R. A. Humber, K. G. Jones, K. A. Klepzig, R. W. Lichtwardt, et al. 2004. The search for diversity of insects and other arthropod associated fungi. In G. M. Mueller, G. F. Bills, and M. S. Foster [eds.], Biodiversity of fungi: Inventory and monitoring methods, 395–433. Elsevier Academic Press, San Diego, California, USA.
Berbee, M. L., and J. W. Taylor. 2010. Dating the molecular clock in fungi—How close are we? Fungal Biology Reviews 24:1–16.
Bhadury, P., B. T. Mohammad, and P. C. Wright. 2006. The current status of natural products from marine fungi and their potential as anti-infective agents. Journal of Industrial Microbiology & Biotechnology 33:325–337.
Bills, G. F., M. Christensen, M. J. Powell, and G. Thorn. 2004. Saprobic soil fungi. In G. M. Mueller, G. F. Bills, and M. S. Foster [eds.], Biodiversity of fungi: Inventory and monitoring methods, 271–302. Elsevier Academic Press, San Diego, California, USA.
Binder, M., D. S. Hibbett, Z. Wang, and W. F. Farnham. 2006. Evolutionary relationships of Mycaureola dilseae (Agaricales), a basidiomycete pathogen of a subtidal rhodophyte. American Journal of Botany 93:547–556.
Bisby, G. R., and G. C. Ainsworth. 1943. The numbers of fungi. Transactions of the British Mycological Society 26:16–19.
Blackwell, M., D. S. Hibbett, J. W. Taylor, and J. W. Spatafora. 2006. Research coordination networks: A phylogeny for kingdom Fungi (Deep Hypha). Mycologia 98:829–837.
Boekhout, T. 2005. Gut feeling for yeasts. Nature 434:449–451.
Borel, J. F. 2002. History of the discovery of cyclosporin and of its early pharmacological development. Wiener Klinische Wochenschrift 114:433–437.
Bowman, B. H., J. W. Taylor, A. G. Brownlee, J. Lee, S.-D. Lu, and T. J. White. 1992. Molecular evolution of the fungi: Relationship of the Basidiomycetes, Ascomycetes and Chytridiomycetes. Molecular Biology and Evolution 9:285–296.
Brock, P. M., H. Doring, and M. I. Bidartondo. 2009. How to know unknown fungi: The role of a herbarium. New Phytologist 181:719–724.
Burdon, J. J. 1993. The structure of pathogen populations in natural plant communities. Annual Review of Phytopathology 31:305–323.
Cafaro, M. J. 2005. Eccrinales (Trichomycetes) are not fungi, but a clade of protists at the early divergence of animals and fungi. Molecular Phylogenetics and Evolution 35:21–34.
Carroll, G. C. 1988. Fungal endophytes in stems and leaves: From latent pathogen to mutualistic symbiont. Ecology 69:2–9.
Cavalier-Smith, T. 1983. A 6-kingdom classification and a unified phylogeny. In H. E. A. Chenk and W. S. Schwemmler [eds.], Endocytobiology II: Intracellular space as oligogenetic, 1027–1034. Walter de Gruyter, Berlin, Germany.
Celio, G. J., M. Padamsee, B. T. Dentinger, R. Bauer, and D. J. McLaughlin. 2006. Assembling the Fungal Tree of Life: Constructing the structural and biochemical database. Mycologia 98:850–859.
Cheewangkoon, R., J. Z. Groenwald, B. A. Summerell, K. D. Hyde, C. To-Anun, and P. W. Crous. 2009. Myrtaceae, a cache of fungal biodiversity. Persoonia 23:55–85.
Clay, K., S. Marks, and G. P. Cheplick. 1993. Effects of insect herbivory and fungal endophyte infection on competitive interactions among grasses. Ecology 74:1767–1777.
Corradi, N., and P. J. Keeling. 2009. Microsporidia: A journey through radical taxonomic revisions. Fungal Biology Reviews 23:1–8.
Crous, P. W., I. H. Rong, A. Wood, S. Lee, H. Glen, W. Botha, B. Slippers, et al. 2006. How many species of fungi are there at the tip of Africa? Studies in Mycology 55:13–33.
Crous, P. W., B. A. Summerell, L. Mostert, and J. Z. Groenewald. 2008. Host specificity and speciation of Mycosphaerella and Teratosphaeria species associated with leaf spots of Proteaceae. Persoonia 20:59–86.
Davies, J. S., and D. W. S. Westlake. 1979. Crude oil utilization by fungi. Canadian Journal of Microbiology 25:146–156.
De Kesel, A. 1996. Host specificity and habitat preference of Laboulbenia slackensis. Mycologia 88:565–573.
Descals, E., and E. Moralejo. 2001. Water and asexual reproduction in the Ingoldian fungi. Botanica Complutensis 25:13–71.
Dettman, J. R., D. J. Jacobson, and J. W. Taylor. 2006. Multilocus sequence data reveal extensive phylogenetic species diversity within the Neurospora discreta complex. Mycologia 98:436–446.
Diehl, W. W. 1950. Balansia and the Balansiae in America. USDA Agriculture Monograph 4:1–82.
Domsch, K. H., W. Gams, and T. H. Anderson. 2007. Compendium of soil fungi, 2nd ed. IHW-Verlag and Verlagsbuchhandlung, Eching, Germany.
Farr, D. F., G. F. Bills, G. P. Chamuris, and A. Y. Rossman. 1989. Fungi on plants and plant products in the United States, 2nd ed. American Phytopathological Society Press, St. Paul, Minnesota, USA.
Feuerer, T. [ed.]. 2010. The index of checklists of lichens and lichenicolous fungi [online]. Website http://www.biologie.uni-hamburg.de/checklists/lichens/portalpages/portalpage_checklists_switch.htm [accessed 30 January 2011].
Feuerer, T., and D. L. Hawksworth. 2007. Biodiversity of lichens, including a world-wide analysis of checklist data based on Takhtajan’s floristic regions. Biodiversity and Conservation 16:85–98.
Fisher, M. C., G. L. Koenig, T. J. White, and J. W. Taylor. 2002. Molecular and phenotypic description of Coccidioides posadasii sp. nov., previously recognized as the non-California population of Coccidioides immitis. Mycologia 94:73–84.
Fisher, P. J. 1977. New methods of detecting and studying saprophytic behaviour of aero-aquatic hyphomycetes. Transactions of the British Mycological Society 68:407–411.
Fonseca, Á., and J. Inácio. 2006. Phylloplane yeasts. In C. Rosa and P. Gábor [eds.], Biodiversity and ecophysiology of yeasts, 63–301. Springer-Verlag, Berlin, Germany.
Freeman, K. R., A. P. Martin, D. Karki, R. C. Lynch, M. S. Mitter, A. F. Meyer, J. E. Longcore, et al. 2009. Evidence that chytrids dominate fungal communities in high-elevation soils. Proceedings of the National Academy of Sciences, USA 106:18315–18320.
Gams, W. 2006. Biodiversity of soil-inhabiting fungi. Biodiversity and Conservation 16:69–72.
Garbelotto, M., P. Gonthier, and G. Nicolotti. 2007. Ecological constraints limit the fitness of fungal hybrids in the Heterobasidion annosum species complex. Applied and Environmental Microbiology 73:6106–6111.
Gerdemann, J. W. 1968. Vesicular arbuscular mycorrhiza and plant growth. Annual Review of Phytopathology 6:397–418.
Gilbertson, R. L., and M. Blackwell. 1984. Two new basidiomycetes on living live oak in the southeast and Gulf Coast region. Mycotaxon 20:85–93.
Gilbertson, R. L., and L. Ryvarden. 1986. North American polypores, vol. I. Abortiporus-Lindtneria. Fungiflora Press, Oslo, Norway.
Gilbertson, R. L., and L. Ryvarden. 1987. North American polypores, vol. II. Megasporoporia-Wrightoporia. Fungiflora Press, Oslo, Norway.
Gilman, J. C. 1957. A manual of soil fungi, 2nd ed. Iowa State College Press, Ames, Iowa, USA.
Giraud, T., G. Refrégier, M. Le Gac, D. M. De Vienne, AND M. E. Hood. 2008. Speciation in fungi. Fungal Genetics and Biology 45:791–802.
Gönczöl, J., and Á. Révay. 2003. Treehole fungal communities: Aquatic, aero-aquatic and dematiaceous hyphomycetes. Fungal Diversity 12:19–24.
Harrington, T. C., S. W. Fraedrich, and D. N. Aghayeva. 2008. Raffaelea lauricola, a new ambrosia beetle symbiont and pathogen on the Lauraceae. Mycotaxon 104:399–404.
Hawksworth, D. L. 1991. The fungal dimension of biodiversity: Magnitude, significance, and conservation. Mycological Research 95:641–655.
Hawksworth, D. L. 2001. The magnitude of fungal diversity: The 1.5 million species estimate revisited. Mycological Research 105:1422–1432.
Hawksworth, D. L., and A. Y. Rossman. 1997. Where are all the undescribed fungi? Phytopathology 87:888–891.
Held, B. W., J. A. Jurgens, B. E. Arenz, S. M. Duncan, R. L. Farrell, and R. A. Blanchette. 2005. Environmetal factors influencing microbial growth inside the historic huts of Ross Island, Antarctica. International Biodeterioration & Biodegradation 55:45–53.
Hibbett, D. M., M. Binder, J. F. Bischoff, M. Blackwell, P. F. Cannon, O. Eriksson, S. Huhndorf, et al. 2007. A higher-level phylogenetic classification of the Fungi. Mycological Research 111:509–547.
Hibbett, D. S., A. Ohman, D. Glotzer, M. Nuhn, P. Kirk, and R. H. Nilsson. In press. Progress in molecular and morphological taxon discovery in Fungi and options for formal classification of environmental sequences. Fungal Biology Reviews.
Hillebrand, H. 2004. On the generality of the latitudinal diversity gradient. American Naturalist 163:192–211.
Hodge, K. T., S. B. Krasnoff, and R. A. Humber. 1996. Tolypocladiuminfl atum is the anamorph of Cordyceps subsessilis. Mycologia 88:715–719.
Hyde, K. D. 2001. Where are the missing fungi? Mycological Research 105:1409–1412.
Hyde, K. D., E. B. G. Jones, E. Leaño, S. B. Pointing, A. D. Poonyth, and L. L. P. Vrijmoed. 1998. Role of fungi in marine ecosystems. Biodiversity and Conservation 7:1147–1161.
Innis, M. A., D. H. Gelfand, J. J. Sninsky, and T. J. White. 1990. PCR protocols: A guide to methods and applications. Academic Press, San Diego, California, USA.
Jacobson, D. J., J. R. Dettman, R. I. Adams, C. Boesl, S. Sultana, T. Roenneberg, M. Merrow, et al. 2006. New findings of Neurospora in Europe and comparisons of diversity in temperate climates on continental scales. Mycologia 98:550–559.
James, T. Y., P. M. Letcher, J. E. Longcore, S. E. Mozley-Standridge, D. Porter, M. J. Powell, G. W. Griffith, and R. Vilgalys. 2006. A molecular phylogeny of the flagellated Fungi (Chytridiomycota) and description of a new phylum (Blastocladiomycota). Mycologia 98:860–871.
Jones, M. D. M., and T. A. Richards. 2009. Environmental DNA combined with fluorescent in situ hybridisation reveals a missing link in the fungal tree of life. Proceedings of 25th Fungal Genetics Conference, 2009, Asilomar, California, USA, abstract 427.
Joppa, L. N., D. L. Roberts, and S. L. Pimm. 2010. How many species of flowering plants are there? Proceedings of the Royal Society of London, B, Biological Sciences 278:554–559.
Jumpponen, A., and L. C. Johnson. 2005. Can rDNA analyses of diverse fungal communities in soil and roots detect effects of environmental manipulations—A case study from tallgrass prairie. Mycologia 97:1177–1194.
Jumpponen, A., and K. L. Jones. 2009. Massively parallel 454 sequencing indicates hyperdiverse fungal communities in temperate Quercus macrocarpa phyllosphere. New Phytologist 184:438–448.
Keeling, P. J. 2009. Five questions about Microsporidia. PLoS Pathogens 5: e1000489.
Kim, K., and C. D. Harvell. 2004. The rise and fall of a six year coral fungal epizootic. American Naturalist 164:S52–S63.
Kirk, J. L., L. A. Beaudette, M. Hart, P. Moutoglis, J. N. Klironomos, H. Lee, and J. T. Trevors. 2004. Methods of studying soil microbial diversity. Journal of Microbiological Methods 58:169–188.
Kirk, P. M., P. F. Cannon, D. W. Minter, and J. A. Stalpers. 2008. Dictionary of the Fungi, 10th ed. CABI, Wallingford, UK.
Klironomos, J. N., and M. M. Hart. 2001. Animal nitrogen swap for plant carbon. Nature 410:651–652.
Kohlmeyer, J., J. W. Spatafora, and B. Volkmann-Kohlmeyer. 2000. Lulworthiales, a new order of marine Ascomycota. Mycologia 92:453–458.
Kohlmeyer, J., and B. Volkmann-Kohlmeyer. 1991. Illustrated key to the filamentous higher marine fungi. Botanica Marina 34:1–61. Kohn, L. M. 2005. Mechanisms of fungal speciation. Annual Review of Phytopathology 43:279–308.
Kuhlman,K. R.,W. G.Fusco,M. T.La Duc,L. B.Allenbach,C. L.Ball, G. M. Kuhlman, R. C. Anderson, et al. 2006. Diversity of microorganisms within rock varnish in the Whipple Mountains, California. Applied and Environmental Microbiology 72:1708–1715.
Kurtzman,C. P., and J. W. Fell. 1998. The yeasts, a taxonomic study, 4th ed. Elsevier, Amsterdam, Netherlands.
Kurtzman, C. P., and C. J. Robnett. 1998. Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie van Leeuwenhoek 73:331–371.
LaChance, M.-A. 2006. Yeast biodiversity: How many and how much? In C. Rosa and P. Gábor [eds.], Biodiversity and ecophysiology of yeasts, 1–9. Springer-Verlag, Berlin, Germany.
LaChance, M.-A., J. Dobson, D. N. Wijayanayaka, and A. M. E. Smith. 2010. The use of parsimony network analysis for the formal delineation of phylogenetic species of yeasts: Candida apicola, Candida azyma, and Candida parazyma sp. nov., cosmopolitan yeasts associated with floricolous insects. Antonie van Leeuwenhoek 97:155–170.
Le Calvez, T., G. Burgaud, S. Mahé, G. Barbier, and P. Vandenkoornhuyse. 2009. Fungal diversity in deepsea hydrothermal ecosystems. Applied and Environmental Microbiology 75:6415–6421.
Lee, S. C., N. Corradi, S. Doan, F. S. Dietrich, P. J. Keeling, and J. Heitman. 2010. Evolution of the sex-related locus and genomic features shared in Microsporidia and Fungi. PLoS ONE 5:e10539. 10.1371/journal.pone.0010539.
Lichtwardt, R. W., M. J. Cafaro, and M. M. White. 2001 The Trichomycetes: Fungal associates of arthropods, revised ed. [online]. Website http://www.nhm.ku.edu/~fungi [accessed 30 January 2011].
Liti, G., D. M. Carter, A. M. Moses, J. Warringer, L. Parts, S. A. James, R. P. Davey, et al. 2009. Population genomics of domestic and wild yeasts. Nature 458:337–341.
Little, A. E. F., and C. R. Currie. 2007. Symbiont complexity: Discovery of a fifth symbiont in the attine ant–microbe symbiosis. Biology Letters 3:501–504.
Longcore, J. E., A. P. Pessier, and D. K. Nichols. 1999. Batrachochytrium dendrobatidis gen. et sp. nov., a chytrid pathogenic to amphibians. Mycologia 91:219–227.
Lumbsch, H. T., P. K. Buchanan, T. W. May, and G. M. Mueller. 2008. Phylogeography and biogeography of Fungi. Mycological Research 112:423–484.
Lutzoni, F., and J. Miadlikowska. 2009. Lichens. Current Biology 19:R502–R503.
Martin, G. W. 1951. The numbers of fungi. Proceedings of the Iowa Academy of Science 58:175–178.
Medina, M., A. G. Collins, J. W. Taylor, J. W. Valentine, J. H. Lipps, L. A. Amaral-Zettler, and M. L. Sogin. 2003. Phylogeny of Opistokonta and the evolution of multicellularity and complexity in Fungi and Metazoa. International Journal of Astrobiology 2:203–211.
Metzker, M. L. 2010. Sequencing technologies—The next generation. Nature Reviews Genetics 11:31–46.
Morris, P. J., W. R. Johnson, J. Pisanic, G. D. Bossart, J. Adams, J. S. Reif, and P. A. Fair. 2010. Isolation of culturable microorganisms from free-ranging bottle nose dolphins (Tursiops truncatus) from the southeastern United States. Veterinary Microbiology 10.1016/j. vetmic.2010.08.025.
Mueller, G. M., and J. P. Schmit. 2007. Fungal biodiversity: What do we know? What can we predict? Biodiversity and Conservation 16:1–5.
Mueller, G. M., J. P.Schmit, P. R.Leacock, B. Buyck, J. Cifuentes, D. E. DesJardin, R. E. Halling, et al. 2007. Global diversity and distribution of macrofungi. Biodiversity and Conservation 16:37–48.
Murdoch, M. E., J. S. Reif, M. Mazzoil, S. D. McCulloch, P. A. Fair, and G. D. Bossart. 2008. Lobomycosis in bottlenose dolphins (Tursiops truncatus) from the Indian River Lagoon, Florida: Estimation of prevalence, temporal trends, and spatial distribution. EcoHealth 5:289–297.
Nagahama, T. 2006. Yeast biodiversity in freshwater, marine and deep-sea environments. In C. Rosa and P. Gábor [eds.], Biodiversity and ecophysiology of yeasts, 241–262. Springer-Verlag, Berlin, Germany.
Nash, T. H. III, B. D. Ryan, P. Diederich, C. Gries, and F. Bungartz. 2004. Lichen flora of the greater Sonoran Desert region, vol. 2, Most of the microlichens, balance of the macrolichens, and the lichenicolous fungi. Lichen Unlimited, Tempe, Arizona, USA.
Nash, T. H. III, B. D. Ryan, C. Gries, and F. Bungartz.2002. Lichenflora of the greater Sonoran Desert region, vol. 1, The pyrenolichens and most of the squamulose and marolichens. Lichen Unlimited, Tempe, Arizona, USA.
Neafsey, D. E., B. M. Barker, T. J. Sharpton, J. E. Stajich, D. J. Park, E. Whiston, C.-Y. Hung, et al. 2010. Population genomic sequencing of Coccidioides fungi reveals recent hybridization and transposon control. Genome Research 20:938–946.
Nilsson, R. H., M. Ryberg, E. Kristiansson, K. Abarenkov, K.-H. Larsson, and U. Kõljalg. 2006. Taxonomic reliability of DNA sequences in public sequence databases: A fungal perspective. PLoS ONE 1:e59. 10.1371/journal.pone.0000059.
O’Brien, B. L., J. L. Parrent, J. A. Jackson, J. M. Moncalvo, and R. Vilgalys. 2005. Fungal community analysis by large-scale sequencing of enviromental samples. Applied and Environmental Microbiology 71:5544–5550.
Ødegaard, F. 2000. How many species of arthropods? Erwin’s estimate revised. Biological Journal of the Linnean Society 71:583–597.
Otrosina, W. J., and M. Garbelotto. 2010. Heterobasidion occidentale sp. nov. and Heterobasidion irregulare nom. nov.: A disposition of North American Heterobasidion biological species. Fungal Biology 114:16–25.
Paton, A. J., N. Brummitt, R. Govaerts, K. Harman, S. Hinchcliffe, B. Allkin, and E. N. Lughadha. 2008. Towards Target 1 of the Global Strategy for Plant Conservation: A working list of all known plant species—Progress and prospects. Taxon 57:602–611.
Penfound, W. T., and F. P. Mackaness. 1940. A note concerning the relation between drainage pattern, bark conditions, and the distribution of corticolous bryophytes. Bryologist 43:168–170.
Perry, H. D., R. Solomon, E. D. Donnenfeld, A. R. Perry, J. R. WittpenN, H. E. Greenman, and H. E. Savage. 2008. Evaluation of topical cyclosporine for the treatment of dry eye disease. Archives of Ophthalmology 126:1046–1050.
Petersen, R. H. 1995. There’s more to a mushroom than meets the eye: Mating studies in the Agaricales. Mycologia 87:1–17.
Petersen, R. H., and K. W. Hughes. 2007. Some agaric distributions involving Pacific landmasses and Pacific Rim. Mycoscience 48:1–14.
Pianka, E. R. 1966. Latitudinal gradients in species diversity: A review of concepts. American Naturalist 100:33–46.
Pinruan, U., K. D. Hyde, S. Lumyong, E. H. C. McKenzie, and E. B. G. Jones. 2007. Occurrence of fungi on tissues of the peat swamp palm Licuala longicalycata. Fungal Diversity 25:157–173.
Pivkin, M. V., S. A. Aleshko, V. B. Krasokhin, and YU. V.Khudyakova. 2006. Fungal assemblages associated with sponges of the southern coast of Sakhalin Island. Russian Journal of Marine Biology 32:207–213.
Porter, T. M., C. W. Schadt, L. Rizvi, A. P. Martin, S. K. Schmidt, L. Scott-Denton, R. Vilgalys, and J. M. Moncalvo. 2008. Widespread occurrence and phylogenetic placement of a soil clone group adds a prominent new branch to the fungal tree of life. Molecular Phylogenetics and Evolution 46:635–644.
Pressel, S., M. I. Bidartondo, R. Ligrone, and J. G. Duckett. 2010. Fungal symbioses in bryophytes: New insights in the twenty first century. Phytotaxa 9:238–253.
Pringle, A., J. D. Bever, M. Gardes, J. L. Parrent, M. C. Rillig, and J. N. Klironomos. 2009. Mycorrhizal symbioses and plant invasions. Annual Review of Ecology, Evolution, and Systematics 40:699–715.
Ranzoni, F. V. 1968. Fungi isolated in culture from soils of the Sonoran Desert. Mycologia 60:356–371.
Raspor, P., and J. Zupan. 2006. Yeasts in extreme environments. In C. Rosa and P. Gábor [eds.], Biodiversity and ecophysiology of yeasts, 372–417. Springer-Verlag, Berlin, Germany.
Redhead, S. 2002. Pseudotulostoma: The find of the century? Inoculum 53:2.
Robert, V., J. Stalpers, T. Boekhout, and S.-H. Tan. 2006. Yeast biodiversity and culture collections. In C. Rosa and P. Gábor [eds.], Biodiversity and ecophysiology of yeasts, 31–44. Springer-Verlag, Berlin, Germany.
Rodriguez, R. J., J. F. White JR., A. E. Arnold, and R. S. Redman. 2009. Fungal endophytes: Diversity and functional roles. New Phytologist 182:314–330.
Rossi, W., and A. Weir. 2007. New species of Corethromyces from South America. Mycologia 99:131–134.
Rossman, A. 1994. A strategy for an all-taxa inventory of fungal biodiversity. In C. I. Peng and C. H. Chou [eds.], Biodiversity and terrestrial ecosystems, 169–194. Academia Sinica Monograph Series no. 14, Taipei, Taiwan.
Ruibal, C., C. Gueidan, L. Selbmann, A. A. Gorbushina, P. W. Crous, J. Z. Groenewald, L. Muggia, et al.. 2009. Phylogeny of rock inhabiting fungi related to Dothideomycetes. Studies in Mycology 64:123–133.
Saikkonen, K., S. H. Faeth, M. Helander, and T. J. Sullivan. 1998. Fungal endophytes: A continuum of interactions with host plants. Annual Review of Ecology and Systematics 29:319–343.
Schadt, C. W., A. P. Martin, D. A. Lipson, and S. K. Schmidt. 2003. Seasonal dynamics of previously unknown fungal lineages in tundra soils. Science 301:1359–1361.
Schmit, J. P., and G. M. Mueller. 2007. An estimate of the lower limit of global fungal diversity. Biodiversity and Conservation 16:99–111.
Schüssler, A., and C. Walker. 2010. Glomeromycota species list [online]. Website http://www.lrz.de/~schuessler/amphylo/amphylo_species.html [accessed 30 January 2011].
Selosse, M. A., F. Richard, X. He, and S. W. Simard. 2006. Mycorrhizal networks: Des liaisons dangereuses? Trends in Ecology & Evolution 21:621–628.
Shearer, C. A., E. Descals, B. Kohlmeyer, J. Kohlmeyer, L. Marvanová, D. Padgett, D. Porter, et al. 2007. Fungal diversity in aquatic habitats. Biodiversity and Conservation 16:49–67.
Shearer, C. A., and H. A. Raja. 2010. Freshwater ascomycetes database [online]. Website http://fungi.life.illinois.edu/ [accessed 30 January 2011].
Smith, S. E., and D. J. Read. 2008. Mycorrhizal symbiosis, 3rd ed. Academic Press, San Diego, California, USA.
Spatafora, J. W., G.-H. Sung, and R. Kepler. 2010. An electronic monograph of Cordyceps and related fungi [online]. Website http://Cordyceps.us [accessed 30 January 2011].
Spatafora, J. W., G.-H. Sung, J.-M. Sung, N. Hywel-Jones, and J. F.White. 2007. Phylogenetic evidence for an animal pathogen origin of ergot and the grass endophytes. Molecular Ecology 16:1701–1711.
Stajich, J. E., M. L. Berbee, M. Blackwell, D. S. Hibbett, T. Y. James, J. W. Spatafora, and J. W. Taylor. 2009. The Fungi. Current Biology 19:R840–R845.
Starmer, W. T., V. Aberdeen, and M.-A. LaChance. 2006. The biogeographic diversity of cactophilic yeasts. In C. Rosa and P. Gábor [eds.], Biodiversity and ecophysiology of yeasts, 486–499. Springer-Verlag, Berlin, Germany.
States, J. S., and M. Christensen. 2001. Fungi associated with biological soil crusts in desert grasslands of Utah and Wyoming. Mycologia 93:432–439.
Stireman, J. O. III, H. P. Devlin, T. G. Carr, and P. Abbot. 2010. Evolutionary diversification of the gall midge genus Asteromyia (Cecidomyiidae) in a multitrophic ecological context. Molecular Phylogenetics and Evolution 54:194–210.
Suh, S.-O., J. V. McHugh, and M. Blackwell. 2004. Expansion of the Candida tanzawaensis yeast clade: 16 novel Candida species from basidiocarp-feeding beetles. International Journal of Systematic and Evolutionary Microbiology 54:2409–2429.
Suh, S.-O., J. V. McHugh, D. Pollock, and M. Blackwell. 2005. The beetle gut: A hyperdiverse source of novel yeasts. Mycological Research 109:261–265.
Taylor, D. L., I. C. Herriott, K. E. Stone, J. W. McFarland, M. G. Booth, and M. B. Leigh. 2010. Structure and resilience of fungal communities in Alaskan boreal forest soils. Canadian Journal of Forest Research 40:1288–1301.
Taylor, J. W., D. J. Jacobson, S. Kroken, T. Kasuga, D. M. Geiser, D. S. Hibbett, and M. C. Fisher. 2000. Phylogenetic species recognition and species concepts in fungi. Fungal Genetics and Biology 31:21–32.
Taylor, T. N., S. D. Klavins, M. Krings, E. L. Taylor, H. Kerp, and H. Hass. 2004. Fungi from the Rhynie Chert: A view from the dark side. Transactions of the Royal Society of Edinburgh, Earth Sciences 94:457–473.
Tedersoo, L., R. H. Nilsson, K. Abarenkov, T. Jairus, A. Sadam, I. Saar, M. Bahram, et al. 2010. 454 pyrosequencing and Sanger sequencing of tropical mycorrhizal fungi provide similar results but reveal substantial methodological biases. The New Phytologist 166:1063–1068.
Trappe, J. M. 1987. Phylogenetic and ecologic aspects of mycotrophy in the angiosperms from an evolutionary standpoint. In G. R. Safir [ed.], Ecophysiology of VA mycorrhizal plants, 2–25. CRC Press, Boca Raton, Florida, USA.
Turner, E., D. J. Jacobson, and J. W. Taylor. 2010. Reinforced post-mating reproductive isolation barriers in Neurospora, an ascomycete microfungus. Journal of Evolutionary Biology 23:1642–1656.
Vandenkoornhuyse, P., S. L. Baldauf, C. Leyval, J. Straczek, and J. P. W. Young. 2002. Extensive fungal biodiversity in plant roots. Science 295:2051.
Vaughan, C. J., M. B. Murphy, and B. M. Buckley. 1996. Statins do more than just lower cholesterol. Lancet 348:1079–1082.
Vega, F. E., A. Simpkins, M. C. Aime, F. Posada, S. W. Peterson, S. A. Rehner, F. Infante, et al. 2010. Fungal endophyte diversity in coffee plants from Colombia, Hawai’i, Mexico, and Puerto Rico. Fungal Ecology 3:122–138.
Villalta, C. F., D. J. Jacobson, and J. W. Taylor. 2009. Three new phylogenetic and biological Neurospora species: N. hispaniola, N. metzenbergii and N. perkinsii. Mycologia 101:777–789.
Vishniac, H. S. 2006. Yeast biodiversity in the Antarctic. In C. Rosa and P. Gábor [eds.], Biodiversity and ecophysiology of yeasts, 419–440. Springer-Verlag, Berlin, Germany.
Vossbrinck, C. R., J. V. Maddox, S. Friedman, B. A. DeBrunner-Vossbrinck, and C. R. Woese. 1987. Ribosomal RNA sequence suggests microsporidia are extremely ancient eukaryotes. Nature 326: 411–414.
Waksman, S. A. 1922. A method for counting the number of fungi in the soil. Journal of Bacteriology 7:339–341.
Wang, B., and Y.-L. Qiu. 2006. Phylogenetic distribution and evolution of mycorrhizas in land plants. Mycorrhiza 16:299–363.
Wang, G., Q. Li, and P. Zhu. 2008. Phylogenetic diversity of culturable fungi associated with the Hawaiian sponges Suberites zeteki and Gelliodes fi brosa. Antonie van Leeuwenhoek 93:163–174.
Weir, A., and M. Blackwell. 2005. Phylogeny of arthropod ectoparasitic ascomycetes. In F. E. Vega and M. Blackwell [eds.], Insect–fungal associations: Ecology and evolution, 119–145. Oxford University Press, New York, New York, USA.
Weir, A., and P. M. Hammond. 1997a. Laboulbeniales on beetles: Host utilization patterns and species richness of the parasites. Biodiversity and Conservation 6:701–719.
Weir, A., and P. M. Hammond. 1997b. A preliminary assessment of speciesrichness patterns of tropical, beetle-associated Laboulbeniales (Ascomycetes). In K. D. Hyde [ed.], Biodiversity of tropical microfungi, 121–139. Hong Kong University Press, Hong Kong.
White, M. M., T. Y. James, K. O’Donnell, M. J. Cafaro, Y. Tanabe, and J. Sugiyama. 2006. Phylogeny of the Zygomycota based on nuclear ribosomal sequence data. Mycologia 98:872–884.
White, T. J., T. D. Bruns, S. B. Lee, and J. W. Taylor. 1990. Amplification and direct sequencing of fungal ribosomal RNA Genes for phylogenetics. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, and T. J. White [eds.], PCR protocols and applications—A laboratory manual, 315–322. Academic Press, New York, New York, USA.
Wirtz, N., C. Printzen, and H. T. Lumbsch. 2008. The delimitation of Antarctic and bipolar species of neuropogonoid Usnea (Ascomycota, Lecanorales): A cohesion approach of species recognition for the Usnea perpusilla complex. Mycological Research 112:472–484.
Zhang, N., and M. Blackwell. 2001. Molecular phylogeny of dogwood anthracnose fungus (Discula destructiva) and the Diaporthales. Mycologia 93:356–364.
Zhou, D., and K. D. Hyde. 2001. Host-specificity, host-exclusivity, and host-recurrence in saprobic fungi. Mycological Research 105:1449–1457.
BAT WHITE-NOSE SYNDROME IN NORTH AMERICA16
David S. Blehert, Jeffrey M. Lorch, Anne E. Ballmann, Paul M. Cryan, and
Carol U. Meteyer17
Since 2007, infections by a previously unrecognized, perhaps imported fungus killed an estimated 1 million bats in North America.
Summary
- The newly described fungus, Geomyces destructans, causes an invasive skin infection in bats and is the likely agent of white-nose syndrome (WNS).
_____________________
16 Reprinted with permission from the American Society for Microbiology (Microbe, June 2011, pp. 267–273).
17 David S. Blehert is the head of the diagnostic microbiology laboratory at the U.S. Geological Survey (USGS)–National Wildlife Health Center, Madison, Wis. (dblehert@usgs.gov), Jeffrey M. Lorch is a graduate student with the Molecular and Environmental Toxicology Center, University of Wisconsin–Madison, Medical Sciences Center, Madison, Wisconsin (jmlorch@wisc.edu), Anne E. Ballmann is a wildlife disease specialist at the USGS–National Wildlife Health Center, Madison, Wis. (aballmann@usgs.gov), Paul M. Cryan is a bat ecologist at the USGS–Fort Collins Science Center, Fort Collins, Colo. (cryanp@usgs.gov), and Carol U. Meteyer is a wildlife pathologist at the USGS–National Wildlife Health Center, Madison, Wis. (cmeteyer@usgs.gov).
- With immune system functions and body temperatures reduced during hibernation, bats may be unusually susceptible to a pathogenic fungus such as G. destructans.
- WNS was first observed in a popular show cave near Albany, New York, leading some investigators to suspect that a visitor inadvertently introduced G. destructans at this site, triggering a wider WNS outbreak in North America.
- Biologists trying to manage WNS within North American bat populations face major challenges, including the variety of susceptible host species, incredible dispersal capabilities of bats, difficulties in treating such populations, and persistence of the pathogen in their vulnerable underground habitats.
In 2007 bats in eastern North America began dying in unprecedented numbers from a previously undocumented disease, now called white-nose syndrome (WNS). Although the ecological and economic impacts of this disease are not fully elucidated, this severe loss of insectivorous bats threatens decreased crop yields, forest defoliation, and a rise in insect-borne diseases. The recent emergence of WNS in bats of eastern North America, its rapid spread, and the severity of the outbreak highlight the importance of wildlife disease as an integral component of ecosystem health.
Biologists with the New York State Department of Environmental Conservation first recognized WNS as a problem in late winter 2007 at five hibernation sites near Albany, N.Y. Subsequently, a recreational caver furnished a photograph from February 2006 in nearby Howes Cave depicting bats with clinical signs of WNS, implicating this location as the likely index site and suggesting disease emergence the winter before New York state biologists drew public attention to the disease. By 2011 WNS had spread south along the Appalachian Mountains into eastern Tennessee, as far west as southern Indiana and western Kentucky, and north into the Canadian provinces of Quebec, Ontario, and New Brunswick (Figure A4-1). Experts estimate that more than 1 million bats have died from WNS thus far. Modeling studies show that, if such mortality trends continue, one of the most abundant bat species in eastern North America, the little brown bat (Myotis lucifugus), could disappear from this region within 16 years. Sustained killing of this magnitude from an infectious disease is unprecedented among the approximately 1,100 species of bats known worldwide.
The Host, Pathogen, and Environment
The likely agent of WNS is a newly described fungus, Geomyces destructans, which causes an invasive skin infection that is the hallmark of this disease (Figure A4-2). G. destructans belongs to the order Helotiales within the phylum Ascomycota. Characteristics that distinguish it from other Geomyces spp. include
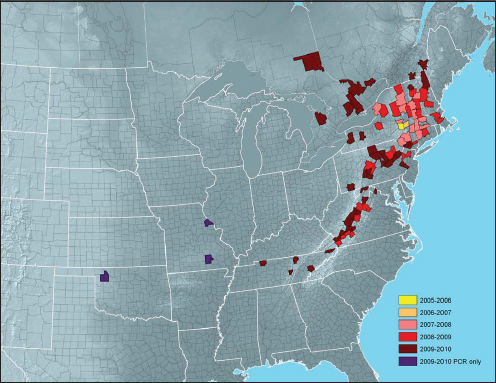
curved conidia (Figure A4-2), slow growth on laboratory medium, cold adaptation, and pathogenicity to bats. Species of Geomyces exist in soils worldwide, especially in colder regions.
Any infectious disease involves interactions among a susceptible host, pathogen, and the environment. To comprehend the ecology of WNS, we must consider the physiological and behavioral aspects of bats that make them susceptible to the disease, the characteristics of the fungus that allow it to act as a pathogen, and the role of underground sites (hibernacula) such as caves and mines in providing conditions conducive to maintaining this pathogen and enabling it to infect these hosts.
WNS appears to occur only in bats, suggesting they possess unique traits that make them a suitable host. Bats are nocturnal and the only mammals capable of powered flight. Their forelimbs are highly modified, consisting of elongated phalanges connected by a thin layer of skin to form wings. This body plan provides bats with selective advantages that allow them to dominate the night skies, making them the second most diverse group of mammals, accounting for approximately 1,100 of 5,400 mammalian species. Of 45 bat species in the United
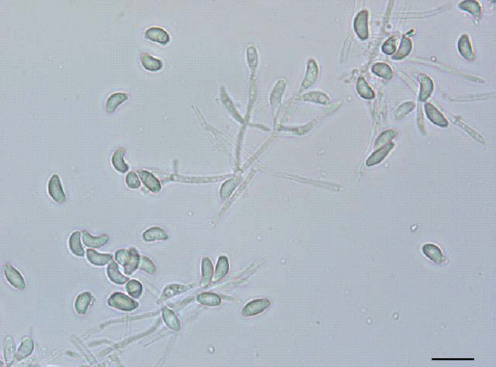
States, at least 6 of the approximately 25 that hibernate have been documented with WNS, including the little brown bat, the northern long-eared bat (M. septentrionalis), the eastern small-footed bat (M. leibii), the endangered Indiana bat (M. sodalis), the tricolored bat (Perimyotis subflavus), and the big brown bat (Eptesicus fuscus). All six of those species are insectivorous and cope with winter food shortages by hibernating in cold and humid, thermally stable caves and mines. When hibernating, the animals typically congregate in large numbers, dramatically reduce metabolic functions, and assume a body temperature close to that of their surroundings (2–7°C). These physiological adaptations and behaviors likely predispose bats to infection by G. destructans and consequent development of WNS. Because approximately half the bat species of the United States are obligate hibernators, another 19 species are at risk for infection by G. destructans if it spreads beyond its current range.
G. destructans colonizes the skin of bat muzzles, wings, and ears, then erodes the epidermis and invades the underlying skin and connective tissues. This pattern is distinctive and is more severe than that caused by typical transmissible dermatophytes. Although the disease was named for the characteristic white growth visible around an infected animal’s nose, the primary site of infection is
the wing (Figure A4-3A). Gross damage to wing membranes such as depigmentation, holes, and tears are suggestive of WNS, but these lesions are nonspecific, and histopathologic examination is necessary to diagnose the disease.
Specifically, fungal invasion of wing membranes ranges from characteristic cup-like epidermal erosions filled with fungal hyphae to ulceration and invasion of underlying connective tissue, with fungal invasion sometimes spanning the full thickness of the wing membrane (Figure A4-3B). Fungal hyphae can also fill hair follicles and destroy skin glands and local connective tissue. Bat wings play an important role in the pathogenesis of WNS by providing a large surface area for the fungus to colonize. Once infected, the thin layer of skin that composes the bat wing is vulnerable to damage that may catastrophically disrupt homeostasis during hibernation.
In North America, bat hibernacula range in temperature from approximately 2–14°C, temperatures all permissive to growth of G. destructans. Within this temperature range, G. destructans exhibits increasing growth rates with increasing temperature (Figure A4-4), but the fungus does not grow at temperatures of approximately 20°C or higher. This temperature sensitivity helps to explain why WNS is observed only among hibernating or recently emerged bats and why the disease is not diagnosed in bats during their active season when body temperatures are consistently elevated above those permissive to growth of G. destructans.
Looking for Other Host and Environmental Susceptibility Factors
Hosts with impaired immune functions tend to be susceptible to opportunistic fungi in their environments. Guided by this concept, some investigators suspected that insults such as exposure to environmental contaminants or infections by viral pathogens compromised bat immunity and made them vulnerable to G. destructans. However, neither contaminant exposure nor viral coinfections can be consistently identified in bats infected with that fungus.
Hibernating bats with WNS generally do not exhibit signs of an inflammatory response. However, severe inflammation typifies fungal skin infections of bats aroused from hibernation, providing evidence that such animals are not immunocompromised. Although studies of bat immune functions are in their infancy, studies of other mammalian species indicate that their immune functions are naturally suppressed during hibernation. Thus, rather than suggesting immune-function impairment, the lack of inflammatory response to fungal infection by hibernating bats may reflect an immune suppression that is part of hibernation physiology.
In addition, the body temperature of hibernating bats drops dramatically, providing another vulnerability to infection by G. destructans. Fatal fungal diseases are relatively rare among endothermic, or warm-blooded, animals because their tissues are too warm to support the growth of most fungal species. However,
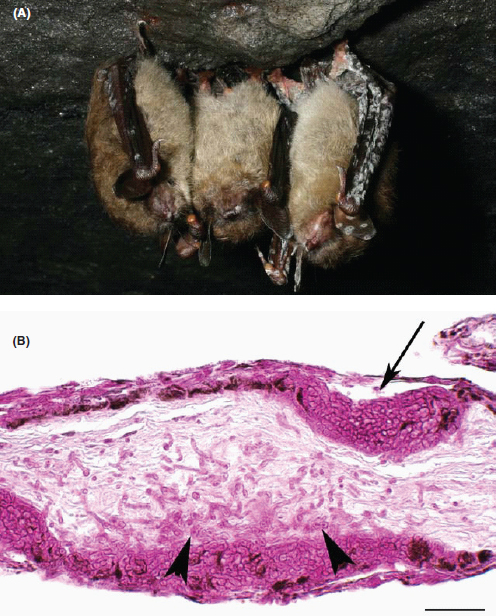
Figure A4-3 (A) Three little brown bats (Myotis lucifugus) photographed by Alan Hicks (New York State Department of Environmental Conservation) in Graphite Mine, New York, in November, 2008. Note the white fungus colonizing the muzzles and nostrils of all three bats. Also note the extensive fungal colonization of the skin of the ears and wings of the bat pictured on the right; (B) Periodic acid-Schiff (PAS) stained microscopic section of wing membrane from a little brown bat with white-nose syndrome collected in Pennsylvania in February, 2009. Dense colonies of fungal hyphae erode skin and fill the cup-shaped depressions (arrow). Ulceration of epidermis with penetration and replacement of subcutaneous tissue (arrow heads) dramatically alters the integrity of wing membrane. Bar = 25 µm.
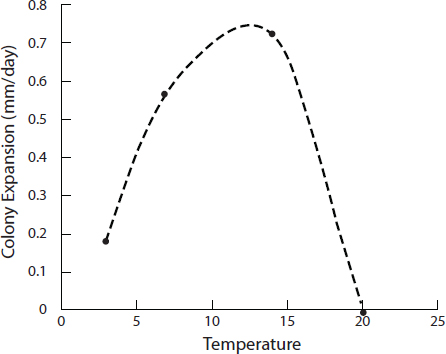
FIGURE A4-4 Colony expansion rates of Geomyces destructans when grown on cornmeal agar at 3, 7, 14, and 20°C. The trend line estimates colony expansion rates at temperatures ranging from 3–20°C.
fungi are more apt to cause fatal diseases in ectothermic, or cold-blooded, organisms such as insects, fish, amphibians, and plants. Bats and other mammals that hibernate are unique in that they are warm-blooded when metabolically active, but cold-blooded during hibernation—a period when their metabolism and body temperatures are dramatically suppressed. Although lowered body temperatures may predispose torpid bats to infection by G. destructans, the mechanism enabling this specific fungus to be a pathogen for bats while other cave-associated fungi remain innocuous is not known.
How G. destructans kills bats is under active investigation. One possibility is that fungal infection disrupts how bats behave while hibernating, leading to more frequent or longer arousals from torpor and thus accelerating usage of fat reserves. However, fat depletion is not consistently observed among all bats with WNS. Infected bats also may exhibit other aberrant behaviors midway through the hibernation season, such as shifting from thermally stable roost sites deep within hibernacula to areas with more variable temperatures near entrances.
Sometimes, they depart early from hibernacula. Thus, exposure to cold could account for some WNS-associated mortality.
Further, fungal damage to wing membranes, which can account for more than 85% of the total surface area of a bat, may increase fatality rates. In addition to the key role that wings play in flight, wing membrane integrity is essential for maintaining water balance, temperature, blood circulation, and cutaneous respiration. Disrupting any of these functions could increase WNS mortality rates.
As with so many other diseases, the environment affects the progress and transmission of WNS. Some pathogenic fungi such as Histoplasma capsulatum, Cryptococcus spp., and Batrachochytrium dendrobatidis can persist in the environment without an animal host for survival. This independence contrasts with host-requiring viruses or other pathogens for which transmission dynamics tend to moderate as infected hosts are removed from a population. G. destructans likely does not require bat hosts to survive and can persist in caves by exploiting other nutrients.
The cool and humid conditions of underground hibernacula provide ideal environmental conditions for G. destructans or other fungal growth. While most G. destructans isolates were cultured from skin or fur of bats collected in or near underground hibernacula during winter, DNA from the same fungus is found in soil samples from several hibernacula that harbor WNS-infected bats in the northeastern US. Also, G. destructans has been cultured from soil samples from hibernacula in three states where WNS occurs, supporting the hypothesis that bat hibernacula are reservoirs for this pathogen and that bats, humans, or fomites may transport G. destructans between hibernacula. How temperature and humidity differences among hibernacula influence G. destructans and WNS is not known.
Uncertainties about WNS Emergence
What caused WNS to emerge in a North American cave during the winter of 2005 to 2006? Bats with clinical signs consistent with WNS were first observed in Howes Cave, a hibernaculum connected to a popular North American show cave. Because of its high human traffic, a tourist might have inadvertently introduced G. destructans at this site.
Europe might be the source for the fungus causing WNS. Reports dating back several decades describe hibernating bats in Germany with white muzzles resembling bats with WNS in North America. Recent culture and PCR surveys indicate that G. destructans is widespread in Europe, including among hibernating bats in hibernacula in the Czech Republic, France, Germany, Hungary, Slovakia, and Switzerland. Unlike in North America, however, mortality rates and population declines remain normal among European bat species. This sharp contrast between disease manifestation among bats in Europe and North America provides an opportunity to investigate how bat species may differ in terms of their susceptibilities to fungal infection, continental variability among fungal strains, and the influence of environmental conditions and bat behavior on this fungal disease.
Challenges in Managing WNS, Conserving Bat Populations
Bat conservation efforts have historically focused mainly on reducing human causes of bat mortality, including habitat destruction, detrimental intrusions into roosts, and intentional extermination of colonies. Bat census figures prior to the emergence of WNS in North America indicate many populations of cave-hibernating bats were stable or increasing. However, the current WNS outbreak brings an even more serious threat to bat populations of North America, confronting biologists with a new set of conservation and management challenges.
Mitigating diseases in free-ranging wildlife populations requires very different approaches from those applied in agriculture for domestic animals. Once established, diseases in free-ranging wildlife are rarely, if ever, eradicated. Biologists trying to manage WNS within bat populations face multiple challenges, including the need to deal with numerous host species, long-distance migrations of infected hosts, poor access to some host populations, impracticalities associated with treating individual wild animals, infected hosts that are sensitive to being disturbed and that inhabit fragile ecosystems, and environmental persistence of the pathogen.
The guiding principle for physicians and veterinarians, “first, do no harm,” will help to prevent WNS management efforts from having unintended adverse consequences. For example: depopulating an infected colony would not be effective unless all infectious animals are removed and all hibernacula used by the population are decontaminated—conditions unlikely to be achieved among free-ranging wildlife; using disinfectants to decontaminate hibernacula could have toxic effects on other organisms reliant on those environments; treating individual bats with antifungal agents is labor intensive, is not self-sustaining, and could be toxic for treated animals or their symbionts; and careless intervention could disrupt natural selective processes that might yield behaviorally or immunologically resistant bats.
However, “first, do no harm” does not mean “do nothing.” State and federal agencies already are taking measures to combat WNS, including closing caves and mandating decontamination procedures. Such steps are intended to prevent people from disturbing hibernating bats and to reduce the chance that intruding humans will transfer G. destructans from one hibernaculum to another. For example, taking a proactive approach prior to the appearance of WNS, state wildlife officials in Wisconsin conferred threatened status on four cave bat species that hibernate within its borders and designated G. destructans a prohibited invasive species providing state resource managers with legal authorities to take disease management actions.
Since the first description of G. destructans in 2008, its genome has been sequenced, and WNS pathology has been more fully defined. Additionally, hibernacula are being surveyed internationally, and ongoing analyses are revealing much about the biodiversity of fungi associated with bat hibernacula. With these and other advances in understanding WNS, opportunities will arise to better
manage the disease cycle. The sudden and unexpected emergence of WNS exemplifies the importance of monitoring, investigating, and responding to emerging wildlife diseases and the ecological and societal threats that they present.
SUGGESTED READING
Blehert, D. S., A. C. Hicks, M. Behr, C. U. Meteyer, B. M. Berlowski-Zier, E. L. Buckles, J. T. H. Coleman, S. R. Darling, A. Gargas, R. Niver, J. C. Okoniewski, R. J. Rudd, and W. B. Stone. 2009. Bat white-nose syndrome: an emerging fungal pathogen? Science 323:227.
Casadevall, A. 2005. Fungal virulence, vertebrate endothermy, and dinosaur extinction: Is there a connection? Fungal Genet. Biol. 42:98–106.
Cryan, P. M., C. U. Meteyer, D. S. Blehert, and J. G. Boyles. 2010. Wing pathology of white-nose syndrome in bats suggests life-threatening disruption of physiology. BMC Biol. 8:135.
Desprez-Loustau, M-L., C. Robin, M. Buée, R. Courtecuisse, J. Garbaye, F. Suffert, I. Sache, and D. M. Rizzo. 2007. The fungal dimension of biological invasions. Trends Ecol. Evol. 22:472–480.
Frick, W. F., J. F. Pollock, A. C. Hicks, K. E. Langwig, D. S. Reynolds, G. G. Turner, C. M. Butchkoski, and T. H. Kunz. 2010. An emerging disease causes regional population collapse of a common North American bat species. Science 329:679–682.
Gargas, A., M. T. Trest, M. Christensen, T. J. Volk, and D. S. Blehert. 2009. Geomyces destructans sp. nov. associated with bat white-nose syndrome. Mycotaxon 108:147–154.
Kunz, T. H. and M. B. Fenton (ed.). 2003. Bat ecology. University of Chicago Press, Chicago.
Lindner, D. L., A. Gargas, J. M. Lorch, M. T. Banik, J. Glaeser, T. H. Kunz, and D. S. Blehert. 2010. DNA-based detection of the fungal pathogen Geomyces destructans in soil from bat hibernation sites. Mycologia 103:241–246.
Meteyer, C. U., E. L. Buckles, D. S. Blehert, A. C Hicks, D. E. Green, V. Shearn-Bochsler, N. J. Thomas, A. Gargas, and M. J. Behr. 2009. Pathology criteria for confirming white-nose syndrome in bats. J. Vet. Diag. Invest. 21:411–414.
Wibbelt, G., A. Kurth, D. Hellmann, M. Weishaar, A. Barlow, M. Veith, J. Prüger, T. Görföl, T. Grosche, F. Bontadina, U. Zöphel, H.-P. Seidl, P. M. Cryan, and D. S. Blehert. 2010. White-nose syndrome fungus (Geomyces destructans) in bats, Europe. Emerg. Infect. Dis. 16:1237–1242.
MAMMALIAN ENDOTHERMY OPTIMALLY RESTRICTS FUNGI AND METABOLIC COSTS18,19
Aviv Bergman20and Arturo Casadevall21
Abstract
Endothermy and homeothermy are mammalian characteristics whose evolutionary origins are poorly understood. Given that fungal species rapidly lose their capacity for growth above ambient temperatures, we have proposed that mammalian endothermy enhances fitness by creating exclusionary thermal zones that protect against fungal disease. According to this view, the relative paucity of invasive fungal diseases in immunologically intact mammals relative to other infectious diseases would reflect an inability of most fungal species to establish themselves in a mammalian host. In this study, that hypothesis was tested by modeling the fitness increase with temperature versus its metabolic costs. We analyzed the tradeoff involved between the costs of the excess metabolic rates required to maintain a body temperature and the benefit gained by creating a thermal exclusion zone that protects against environmental microbes such as fungi. The result yields an optimum at 36.7°C, which closely approximates mammalian body temperatures. This calculation is consistent with and supportive of the notion that an intrinsic thermally based resistance against fungal diseases could have contributed to the success of mammals in the Tertiary relative to that of other vertebrates.
Importance
Mammals are characterized by both maintaining and closely regulating high body temperatures, processes that are known as endothermy and homeothermy,
_____________________
18 Originally published as: Bergman, A. and A. Casadevall. 2010. Mammalian Endothermy Optimally Restricts Fungi and Metabolic Costs. mBio 1(5): e00212-10. doi:10.1128/mBio.00212-10.
19Received 17 August 2010 Accepted 11 October 2010 Published 9 November 2010 Citation Bergman, A., and A. Casadevall. 2010. Mammalian endothermy optimally restricts fungi and metabolic costs. mBio 1(5):e00212-10. doi: 10.1128/mBio.00212-10. Editor Françoise Dromer, Institut Pasteur Copyright © 2010 Bergman and Casadevall. This is an open-access article distributed under the terms of the Creative Commons Attribution-Noncommercial-Share Alike 3.0 Unported License, which permits unrestricted noncommercial use, distribution, and reproduction in any medium, provided the original author and source are credited. Address correspondence to Arturo Casadevall, arturo.casadevall@einstein.yu.edu.
20 Department of Systems and Computational Biology, Albert Einstein College of Medicine, Bronx, New York, USA.
21 Department of Microbiology and Immunology, Albert Einstein College of Medicine, Bronx, New York, USA.
respectively. The mammalian lifestyle is energy intensive and costly. The evolutionary mechanisms responsible for the emergence and success of these mammalian characteristics are not understood. This work suggests that high mammalian temperatures represent optima in the tradeoff between metabolic costs and the increased fitness that comes with resistance to fungal diseases.
Endothermy and homeothermy are fundamental aspects of mammalian physiology whose evolutionary origin remains poorly understood. Although many explanations have been suggested for the origins of endothermy and homeothermy, none are fully satisfactory given their high metabolic costs (Kemp, 2008; Ruben, 1995). Furthermore, the factors responsible for the mammalian set point remain unknown, posing the additional question of why mammals are so hot. Recently, the observation that fungal diseases are common in plants and insects but rare in mammals, combined with the thermal susceptibility of fungi, led to the proposal that mammalian endothermy and homeothermy create a thermal exclusionary zone that protects mammals against mycoses (Robert and Casadevall, 2009). Endothermy was also suggested to have provided a fitness advantage in the fungal bloom that followed the end of the Cretaceous such that it could have contributed to the success of mammals in the Tertiary (Casadevall, 2005; Robert and Casadevall, 2009).
Assuming that a relationship exists between endothermy and reduced susceptibility to certain classes of microbes, we hypothesized a tradeoff relationship whereby the high costs of endothermy were mitigated by protection against infectious diseases. In other words, we posited that increases in body temperature would protect against microbes by creating a thermal exclusionary zone but that such increases would be increasingly costly with regard to metabolic rates as the host body temperature diverged from ambient temperatures. Given that there is robust information on fungal thermal tolerances (Robert and Casadevall, 2009), we decided to test this hypothesis by attempting to identify body temperatures that confer maximal fitness for certain metabolic rates.
To address this question, we propose a first-order model wherein a tradeoff exists between the excess metabolic rates required to maintain a body temperature, T, and the benefit gained by protection against deleterious microbes because of the creation of a thermal exclusion zone. Metabolism, the exchange of energy between the organism and its environment, as well as the transformation of that energy to material within an organism, is affected by two main factors, body mass, M, and body temperature, T. Due to the fractal nature of transport networks, that is, vessel architecture and branching (Gillooly et al., 2001; Savage et al., 2008), over ontogeny, the resting metabolic rate, Brest, scales with body mass, m, as Brest = B0m3/4, where B0 is a normalization constant for a given taxon. Also, the normalization coefficient, B0, exponentially increases with body temperature B0 ~ e–E0/KT, where E0 is the average activation energy for the rate-limiting enzyme-catalyzed biochemical reactions of metabolism (ca. 0.65 eV), K is Boltzmann’s constant (8.62 × 10–5eV/K), and T is body temperature (Brown et al., 2004;
Gillooly et al., 2001). The scaling relationship between resting metabolic rate and body mass, ∝m3/4, has been predicted from allometric theories and supported by data on a diverse set of organisms, including mammals, birds, fish, and mollusks (Brody, 1964; Moses et al., 2008; Savage et al., 2004; West et al., 1997). As can be seen from the formulas above, body temperature affects the metabolic rate through its effects on rates of biochemical reaction kinetics according to Boltzmann’s factor, e–Ei/kT, where T is measured in kelvins (absolute temperature). The resting metabolic rate, Brest, is proportional to the product of these two effects and again has been shown to be well approximated, within a biologically relevant temperature range (0°C to 40°C), as B(T) ∝e–Ei/kTm3/4 (Gillooly, 2001). The first part of our analysis examined the excess cost for an organism of body mass m to maintain a body temperature T (assuming no dependence of body mass on temperature).
In the second part of our analysis, the benefit, noted here as F(T), is calculated as the reduction in the number of fungal species capable of infecting a host; this number is reduced approximately by s ≈ 6% for every degree Celsius in the temperature range of 27°C to 40°C (Robert and Casadevall, 2009). The increased benefit of the successive elimination of fungal species can thus be expressed as F(T) ∝F0[1 – (1 – s)T], where F0 is a constant scaling factor. The quantity W(T) = F(T)/B(T) can represent the balance between cost and benefit; thus, W(T) can be viewed as the total fitness of an organism as a function of its body temperature. Within the biologically relevant temperature range, the proposed fitness measure reaches its maximum at approximately 37°C (Fig. A5-1). Note that in this formulation, the optimal body temperature, where W(T) attains its maximum value, does not depend on the organism’s body mass. Furthermore, the one parameter that is determined from biological observation is the reduction in the number of fungal species capable of infecting a host; thus, to determine our model’s dependence on this parameter, we calculated the optimal temperature over a wide range of possible reduction percentages, i.e., 4% to 8%. In this range, the optimal temperature was found to remain in a tight range of less than 2°C, from 37.7°C to 35.9°C, respectively, which is still within the biologically relevant range of mammalian body temperatures. The insensitivity of the model to its only parameter further strengthens our hypothesis.
In summary, we present a minimal, parsimonious model to account for the cost of maintaining a high body temperature in mammalian organisms. A body temperature of 36.7°C maximizes fitness by restricting the growth of most fungal species relative to its metabolic cost. Our model suggests that no additional elaborations are required to explain the evolution of endothermy other than the tradeoff between protection against environmentally acquired microbial diseases and the cost of metabolism. Although we cannot rule out the possibility that this body temperature optimum arose by some remarkable coincidence, we think this highly unlikely because it emerges from considering two unrelated processes, fungal thermal tolerance and mammalian metabolic costs. Nonethe-
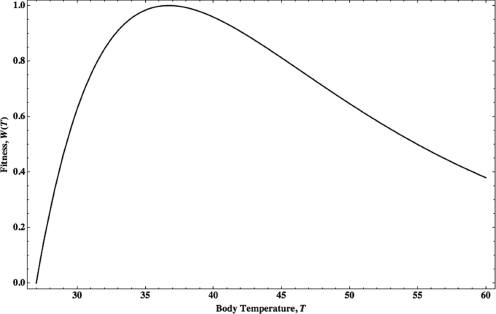
Figure A5-1 Organism fitness as a function of body temperature. We normalized fitness, W(T), to attain a maximum value of 1 and plotted body temperature in degrees Celsius over a range of 27°C to 60°C. Fitness reaches a maximum value at WMAX(T) ≈ 36.7°C.
less, we acknowledge that similar temperature optima might emerge from other considerations. For example, the specific heat capacity of water has a minimum at 36°C, and if the efficiency of metabolic processes is related to heat capacity, then using this parameter as the optimality criterion may result in a similar range of solutions. Nevertheless, we note the internal consistency in the theme that fungal diseases are rare in immunologically intact mammals and the tradeoff between increased fitness and metabolic costs closely approximates mammalian body temperatures.
Acknowledgements
Aviv Bergman is supported by 5P01AG027734-04 and 5R01AG028872-04. Arturo Casadevall is supported by AI33774-11, HL59842-07, AI33142-11, AI52733-02, and U54-AI057158-Lipkin.
References
Brody, S. 1964. Bioenergetics and growth. Hafner, New York, NY.
Brown, J. H., J. F. Gillooly, A. P. Allen, V. M. Savage, and G. B. West. 2004. Toward a metabolic theory of ecology. Ecology 85:1771–1789.
Casadevall, A. 2005. Fungal virulence, vertebrate endothermy, and dinosaur extinction: is there a connection? Fungal Genet. Biol. 42:98–106.
Gillooly, J. F., J. H. Brown, G. B. West, V. M. Savage, and E. L. Charnov. 2001. Effects of size and temperature on metabolic rate. Science 293:2248–2251.
Kemp, T. S. 2008. The origin of mammalian endothermy: a paradigm for the evolution of complex biological structure. Zool. J. Linn. Soc. 147:473–488.
Moses, M. E., C. Hou, W. H. Woodruff, G. B. West, J. C. Nekola, W. Zuo, and J. H. Brown. 2008. Revisiting a model of ontogenetic growth: estimating model parameters from theory and data. Am. Nat. 171:632–645.
Robert, V. A., and A. Casadevall. 2009. Vertebrate endothermy restricts most fungi as potential pathogens. J. Infect. Dis. 200:1623–1626.
Ruben, J. 1995. The evolution of endothermy in mammals and birds: from physiology to fossils. Annu. Rev. Physiol. 57:69–95.
Savage, V. M., E. J. Deeds, and W. Fontana. 2008. Sizing up allometric scaling theory. PLoS Comput. Biol. 4:e1000171.
Savage, V. M., J. F. Gillooly, W. H. Woodruff, G. B. West, A. P. Allen, B. J. Enquist, and J. H. Brown. 2004. The predominance of quarter-power scaling in biology. Funct. Ecol. 18:257–282.
West, G. B., J. H. Brown, and B. J. Enquist. 1997. A general model for the origin of allometric scaling laws in biology. Science 276:122–126.
VERTEBRATE ENDOTHERMY RESTRICTS MOST FUNGI AS POTENTIAL PATHOGENS22,23
Vincent A. Robert24and Arturo Casadevall25
The paucity of fungal diseases in mammals relative to insects, amphibians, and plants is puzzling. We analyzed the thermal tolerance of 4802 fungal strains from 144 genera and found that most cannot grow at mammalian temperatures. Fungi from insects and mammals had greater thermal tolerances than did isolates from soils and plants. Every 1°C increase in the 30°C–40°C range excluded an
_____________________
22 Vincent A. Robert and Arturo Casadevall, “A Vertebrate Endothermy Restricts Most Fungi as Potential Pathogens”, Journal of Infectious Diseases, 2009, Vol. 200, Iss. 10, pp. 1623–1626. Reprinted by permission of Oxford University Press.
23 Received 23 May 2009; accepted 18 June 2009; electronically published 14 October 2009. Potential conflicts of interest: none reported. Financial support: National Institutes of Health (awards 5R01AI033774, 5R01HL059842, and 2U54AI057158). Reprints or Correspondence: Dr Arturo Casadevall, Department of Medicine, Albert Einstein College of Medicine, Yeshiva University, 1300 Morris Park Ave, Bronx, NY 10461 (arturo.casadevall@einstein.yu.edu).
The Journal of Infectious Diseases 2009;200:000–000
© 2009 by the Infectious Diseases Society of America. All rights reserved. 0022-1899/2009/20010-00XX$15.00
DOI: 10.1086/644642
24 Centraalbureau voor Schimmelcultures, Utrecht, the Netherlands.
25 Department of Microbiology and Immunology and Division of Infectious Diseases, Department of Medicine, Albert Einstein College of Medicine, Yeshiva University, Bronx, New York.
additional 6% of fungal isolates, implying that fever could significantly increase the thermal exclusion zone. Mammalian endothermy and homeothermy are potent nonspecific defenses against most fungi that could have provided a strong evolutionary survival advantage against fungal diseases.
Of the 1.5 million fungal species, only a few hundred are pathogenic to mammals (Kwon-Chung and Bennett, 1992). Fungal diseases in mammals often reflect impaired immune function, and fungi did not emerge as major pathogens for humans until the late 20th century. For example, candidiasis was uncommon until the 1950s, when thrush was associated with the introduction of antibiotics that disrupted bacterial flora. Similarly, diseases such as cryptococcosis, aspergillosis, and histoplasmosis were rare until recently, when their prevalence increased with the human immunodeficiency virus epidemic and the development of immunosuppressive therapies. In contrast, the number of fungal species pathogenic to plants and insects is estimated to be 270,000 and 50,000, respectively (Hawksworth and Rossman, 1997). Amphibians are particularly vulnerable to certain fungal infections, as evidenced by the current catastrophic epidemic of chytridiomycosis in frogs.
The resistance of mammals with intact immune systems to systemic fungal diseases, coupled with their endothermic and homeothermic lifestyles, suggested that these costly physiological adaptations were evolutionarily selected because they conferred a survival advantage by protecting against environmental pathogens (Casadevall, 2005). However, testing this hypothesis was difficult because knowledge of fungal thermal tolerance is largely anecdotal. Consequently, we evaluated the thermal growth tolerances of fungal species in a reference collection and compared them to mammalian temperatures.
Methods
A total of 4802 fungal strains belonging to 144 genera in the Centraalbureau voor Schimmelcultures (Utrecht) collection were tested for growth at 4°C, 12°C, 15°C, 18°C, 21°C, 25°C, 30°C, 35°C, 37°C, 40°C, 42°C, and 45°C. Strains were grown for times ranging from a few days to a few weeks on the most suitable medium, generally glucose–peptone–yeast extract agar, potato-dextrose agar, or yeast extract–malt extract agar. Growth was considered positive when a colony was visible without magnification. The strain set included Ascomycetes and Basidiomycetes but excluded Zygomycetes, which is not in the yeast database.
The culture deposit records were reviewed to identify the isolation source. Fungi isolated from flowers, grains, and herbal exudates were grouped under plant isolates. Animal isolates were classified depending on whether they originated from endothermic (mammals and birds) or ectothermic (insects, nematodes, fishes, and crustaceans) species. Another group comprised isolates from nonliving environmental sources, which included predominantly soils; this group is referred to as soil isolates. These groups were compared for thermal tolerance at 2 tem-
peratures, 25°C and 37°C, which reflect ambient and mammalian temperatures, respectively.
To test the significance of the difference in growth patterns between fungal strains isolated from different groups, we calculated the test statistics
![]()
where p1 and p2 are the observed sample proportions for each group of fungal strains at a given temperature, n1 and n2 are the size of the 2 groups under comparison, and
P = (p1 × n1 + p2 × n2)/(n1 + n2).
The statistic z was assumed to be distributed normally. The 2-tailed probability from the absolute z score to infinity on both tails of the distribution was calculated (http://www.danielsoper.com/statcalc/calc21.aspx) and confirmed using the NORMSDIST function in Excel (Microsoft) to assess the significance of differences in growth between groups at different temperatures.
Results and Discussion
Knowledge of fungal thermal tolerance is limited to a few species because the subject has not been systematically studied. In fact, such studies may be very difficult to do, and a comprehensive prospective study of fungal thermal tolerance would require a gargantuan effort. However, culture collections provide an attractive alternative for initial explorations of this subject. Culture collections store and maintain fungal strains and record basic nutritional needs and temperature tolerances. This information, when accessed and analyzed with bioinformatics tools, provides a useful starting point for the analysis of fungal thermal tolerances.
Our results show that most strains grew well in the 12°C–30°C range, but there was a rapid decline in thermal tolerance at temperatures >35°C (Figure A6-1). A plot of the fraction of fungal strain that grew versus temperature in the 30°C–42°C range revealed a linear relationship with an equation of y = -0.0166x × 2.7911, such that for every 1° increase in temperature >30°C, ~6% fewer strains could grow.
For 3020 strains, there was information on both source isolation and temperature tolerance. This group included isolates from the environment (primarily soils), plants, ectothermic animals, and endothermic animals. The majority of these isolates grew at 25°C regardless of their source (Table A6-1). Nevertheless, the proportion growing at 25°C was significantly greater for isolates recovered from living hosts than from soils, irrespective of whether the hosts were ectothermic plants and animals or endothermic animals. At 37°C, the proportion of
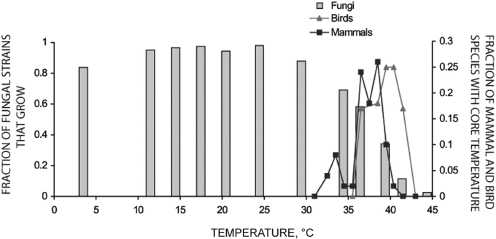
Figure A6-1 Frequency histogram of thermal growth tolerance for 4802 fungal strains (bars). Lines connect percentages for 49 mammalian (blue) and 12 bird (red) species core temperatures. Obtained from McNab (1970).
Table A6-1 Growth Tolerances for Fungi from Soils, Animals, and Plants at 2 Temperatures
| Origin, host type | Isolate Growth | Total | P valuesb | ||||
| Yes | No | Unknowna | P1 | P2 | P3 | ||
| at 25°C | |||||||
| Soils, NA | 657 | 42 | 7 | 706 | |||
| Plant, ectotherm | 1108 | 30 | 5 | 1143 | <.001 | ||
| Animal | |||||||
| Ectotherm | 490 | 0 | 6 | 496 | <.001 | .29 | |
| Endotherm | 661 | 5 | 9 | 675 | <.001 | .214 | .263 |
| at 37°C | |||||||
| Soils, NA | 146 | 535 | 15 | 706 | |||
| Plant, ectotherm | 304 | 871 | 22 | 1143 | .292 | ||
| Animal | |||||||
| Ectotherm | 193 | 284 | 19 | 496 | <.001 | .004 | |
| Endotherm | 466 | 202 | 7 | 605 | <.001 | <.001 | <.001 |
NOTE. NA, not applicable
aRefers to a small no. of isolates for which the temperature growth data was not complete.
bP1 refers to the comparison of isolates from soils, P2 refers to the comparison versus plant isolates, and P3 refers to the comparison between isolates from ectothermic and endothermic animals.
fungi that grew was much higher for isolates from endothermic animals than from ectothermic animals. The proportions of Ascomycetes and Basidiomycetes fungi in each group were comparable, except for ectothermic hosts, which yielded predominantly Ascomycetes fungi.
Isolates from ectothermic hosts (such as plants and insects) were significantly more thermotolerant than isolates from soils. A significantly greater percentage of fungal strains from insects grew at 37°C relative to those recovered from plants, possibly reflecting the fact that insects can increase their temperature through behavioral fevers that increase survival after fungal infection (Thomas and Blanford, 2003). However, this explanation is unlikely to apply to plants, which have much lower metabolic rates. Since thermal tolerance must be associated with numerous metabolic changes that mitigate fungal damage, the association between greater thermotolerance and plant pathogenicity could mirror adaptation to survival in a host with potent antifungal defenses, raising the tantalizing possibility that selection pressures by virulence may contribute to thermal stability and vice versa. In this regard, we note that Hsp90 orchestrates morphogenesis in Candida albicans (Shapiro et al., 2009), thus providing a molecular association for heat shock and a virulence-related phenotype that may be conserved in other fungi.
A survey of the fungal genera represented in our sample collection revealed differences in the percentage of isolates capable of growth at 37°C. All genera studied included some thermotolerant species, as defined by their ability to grow at 37°C, but there were large differences in the percentage of species within each genera. Thermotolerant genera included those from both Ascomycetes and Basidiomycetes, but basidiomycetous genera were disproportionately more common among the thermotolerant genera (P < .001, Fisher exact test). The strains grouped within the sexually related basidiomycetous genera Filobasidiella (a telemorph of Cryptococcus) and Cryptococcus (an anamorph of Filobasidiella) included comparable numbers of thermotolerant species (61% among 116 strains and 53% among 287 strains, respectively). These data suggest an association between phylogeny and thermotolerance.
The capacity for thermotolerance was interspersed among Ascomycetes and Basidiomycetes, suggesting that it may have emerged independently several times in evolution. Alternatively, thermotolerance may be an ancient fungal trait that was lost by those species that cannot grow at 37°C. In this regard, we note that the climate for much of Earth’s history was much warmer than in recent geologic epochs, having cooled by ~5°C during the Eocene-Oligocene transition ~34 million years ago (Liu et al., 2009). The fact that thermotolerance is a complex trait that can be lost by a single mutation, as demonstrated by laboratory-generated temperature-sensitive mutants, makes the explanation of a retained phenotype attractive.
Our results may be relevant to the ongoing debate on the origin and function of endothermy, homeothermy, and fever, each a major unsolved problem in vertebrate physiology (Kemp, 2008; Ruben, 1995). There is no consensus as to
why mammals have adopted such an energetically costly lifestyle. Endothermy is associated with certain metabolic benefits and thermodynamic efficiency, but these benefits come at a high cost since endothermic vertebrates require ~10 times more oxygen to support metabolism than do ectothermic vertebrates (Ruben, 1995). Our analysis suggests that part of the cost is mitigated by the creation of a thermal exclusionary zone that can protect against environmental microbes. Given the high metabolic cost of endothermy, the core temperatures of individual mammal and bird species are likely to be a compromise between its benefits and costs. If endothermy was selected for protection against infectious disease, then a case could be made that endothermy preceded homeothermy. Similarly, if one considers fever as a mechanism to extend the thermal exclusionary zone against environmental microbes such as fungi, increases in temperature of only 1°–3° can significantly reduce the proportion of such microbes that can inhabit the host.
The benefits of endothermy and homeothermy in protection against microbes do not appear to have been previously considered as mechanisms for evolutionary selection, possibly because most of the viral and bacterial diseases that currently plague animals are often acquired from other warm hosts, and these necessarily involve thermotolerant microbes. However, the perspective is very different when one focuses on environmentally acquired microbes and the fungi in particular. Pathogenic microbes are a very small subset of the total terrestrial microbial flora, and these can be divided as to whether they are acquired from other hosts or directly from the environment (Casadevall and Pirofski, 2007). For mammals, pathogenic microbes acquired from other hosts are usually adapted to mammalian temperatures, but microbes acquired directly from the environment would not be subject to such selection pressures. Hence, the potential benefit of endothermy and homeothermy to host defense may become apparent only when one considers the entire microbiota and that subset of pathogenic microbes that is acquired directly from the environment. In support of this notion, we note that bats become susceptible to a cold-loving fungus when hibernation greatly reduces their body temperatures (Blehert et al., 2009) and that primitive mammals (such as the egg-laying platypus, which has a body temperature of 32°C) are susceptible to fungal diseases (Obendorf et al., 1993). An epidemiological observation consistent with the protective function of endothermy comes from the observation that serotype D Cryptococcus neoformans are less thermotolerant (Martinez et al., 2001) than other varieties and are associated with cutaneous cryptococcosis (Dromer et al., 1996). Experimental support for the notion that endothermy restricts fungal infection comes from the observation that rabbits, which have core temperatures of 38°C–39°C, are notoriously resistant to cryptococcosis, and infection can be induced only in cooler organs, such as testes (Perfect et al., 1980). However, the same system also shows that mammalian immune systems also make a decisive contribution to host defense against fungi since systemic cryptococcosis can be induced in rabbits after corticosteroid administration (Perfect et al., 1980).
This study reflects the power of a bioinformatics analysis of archival data
from culture collection, which allows comparison of temperature growth data on thousands of isolates. However, there are certain limitations that should be considered in evaluating the data. Strains from plants and insects were disproportionately represented in the collection, and this may introduce certain biases. The relative paucity or absence of strains from certain sources and taxonomic groups could contribute to bias in the statistical analysis. For example, there were relatively few isolates from birds and ectothermic animals other than insects, no Zygomycetes fungi, and only a few filamentous fungi. Furthermore, the catalogued information was insufficiently detailed to distinguish between skin and systemic isolates from endotherms, which could differ in thermotolerance.
The discovery of fossilized fungal proliferation at the Cretaceous-Tertiary boundary was proposed to contribute to extinction events at the end of the Cretaceous epoch that replaced reptiles with mammals as the dominant large animals (Casadevall, 2005). Thermal tolerance is a necessary, but not sufficient, characteristic of microbes being capable of causing invasive disease in mammals. Since thermal tolerance almost certainly involves many genes and biochemical processes, it is unlikely that this trait can be rapidly acquired by any one microbial species. Consequently, new human pathogenic fungi are likely to emerge from genera that are already tolerant to higher temperatures; such species may warrant special attention given likely climatic changes in the years ahead that could alter patterns of fungal prevalence.
References
Blehert DS, Hicks AC, Behr M, et al. Bat white-nose syndrome: an emerging fungal pathogen? Science 2009; 323:227.
Casadevall A. Fungal virulence, vertebrate endothermy, and dinosaur extinction: is there a connection? Fungal Genet Biol 2005; 42:98–106.
Casadevall A, Pirofski LA. Accidental virulence, cryptic pathogenesis, martians, lost hosts, and the pathogenicity of environmental microbes. Eukaryot Cell 2007; 6:2169–74.
Dromer F, Mathoulin S, Dupont B, Letenneur L, Ronin O. Individual and environmental factors associated with infection due to Cryptococcus neoformans serotype D. Clin Infect Dis 1996; 23:91–6.
Hawksworth DL, Rossman AY. Where are all the undescribed fungi? Phytopathology 1997; 87:888–91.
Kemp TS. The origin of mammalian endothermy: a paradigm for the evolution of complex biological structure. Zool J Linn Soc 2008; 147:473–88.
Kwon-Chung KJ, Bennett JE. Medical mycology. Philadelphia: Lea & Febiger, 1992.
Liu Z, Pagani M, Zinniker D, et al. Global cooling during the eocene-oligocene climate transition. Science 2009; 323:1187–90.
Martinez LR, Garcia-Rivera J, Casadevall A. Cryptococcus neoformans var. neoformans (serotype D) strains are more susceptible to heat than C. neoformans var. grubii (serotype A) strains. J Clin Microbiol 2001; 39:3365–7.
McNab BK. Body weight and the energetics of temperature regulation. J Exp Biol 1970; 53:329–48.
Obendorf DL, Peel BF, Munday BL. Mucor amphibiorum infection in platypus (Ornithorhynchus anatinus) from Tasmania. J Wildl Dis 1993; 29:485–7.
Perfect JR, Lang SDR, Durack DT. Chronic cryptococcal meningitis. Am J Path 1980; 101:177–93.
Ruben J. The evolution of endothermy in mammals and birds: from physiology to fossils. Annu Rev Physiol 1995; 57:69–95.
Shapiro RS, Uppuluri P, Zaas AK, et al. Hsp90 orchestrates temperature-dependent Candida albicans morphogenesis via Ras1-PKA signaling. Curr Biol 2009; 19:621–9.
Thomas MB, Blanford S. Thermal biology in insect-pathogen interactions. Trends Ecol Evol 2003; 18:344–50.
SURVEILLANCE FOR EMERGING DISEASES IN WILDLIFE
Peter Daszak, Carlos Zambrana-Torrelio, and Tiffany Bogich26
Impact of Fungal Diseases on Wildlife
At this meeting, a number of presenters discussed the role of specific fungal diseases in rapid declines and even extinctions of wildlife. Getting an accurate measure of the impact of a pathogen or group of pathogens on wildlife is notoriously difficult. First, wildlife populations undergo often dramatic shifts that are difficult to distinguish from the effects of an outbreak. These may be seasonal or interannual, and occur in response to long-term climatic fluctuations, variation in predator–prey cycles, or a range of other difficult-to-measure factors (Daszak et al., 2005; Pechmann et al., 1991). Second, outbreaks of disease in wildlife may cause significant mortality, but these events may be difficult to detect due to rapid scavenging or decay of carcasses. Even when carcasses are found, they may be too decayed to conduct proper pathological investigations. Third, despite a range of infectious agents linked to recent declines, these are relatively new to ecologists and wildlife managers, so that die-offs are often attributed to other factors, and diseases may not be examined. Despite these and other issues, emerging diseases—indeed, emerging fungal diseases—have been shown to cause significant population declines and even extinctions in wildlife (Daszak and Cunningham, 1999; Frick et al., 2010; Schloegel et al., 2006).
The emerging fungal disease chytridiomycosis is a good example of this trend. As discussed elsewhere in this report, it is caused by the fungal pathogen Batrachochytrium dendrobatidis, which infects the keratin-rich cells on the skin of adult amphibians (Kilpatrick et al., 2010). This disease was discovered in the 1990s and associated with significant population declines in Australia and Central America (Berger et al., 1998). Substantial support from other studies shows it is the major cause of global amphibian declines (Crawford et al., 2010; Lips et al., 2006; Skerratt et al., 2007). However, when this disease was first reported, a debate took place in the literature over whether amphibians were undergoing
_____________________
26 EcoHealth Alliance, 340 West 34th Street, New York, NY 10001.
population declines, or simple fluctuations (Blaustein, 1994). Debate continued on the importance of chytridiomycosis and other factors. The ecological community took about a decade to accept the role of disease in these wild animals as important enough to call for large-scale global action to prevent further spread (Mendelson et al., 2006).
Despite these issues, reports of emerging diseases affecting wildlife populations have grown rapidly (Aguirre and Tabor, 2008; Cunningham, 2005; Daszak et al., 2000; Deem et al., 2001; Nettles, 1996; Wildlife and emerging disease, 2009; Williams et al., 2002). These include diseases that have caused a number of high-profile declines, such as mycoplasmal conjunctivitis of house finches in the United States (Fischer et al., 1997); trichomonosis in declining birds in the United Kingdom (Robinson et al., 2010); chronic wasting disease of cervids in the United States (Miller and Williams, 2004; Sigurdson, 2008); and white-nose syndrome (Blehert et al., 2009) or geomycocis (Chaturvedi and Chaturvedi, 2011) of bats. Among these are a surprising number of fungal diseases, which often have a high impact. In humans, fungal pathogens do not represent a major cause of emerging diseases.
Analysis of a global database of emerging pathogens (Jones et al., 2008) suggests that fungi are responsible for only 5.9 percent of the emerging infectious diseases of people in the past four decades (Figure A7-1). Yet in wildlife, fungi have been implicated in global declines of amphibians, leading to extinction of species (Schloegel et al., 2006), multistate declines of bat populations (Frick et al., 2010), the near extinction of the Florida Torreya tree (Schwartz et al., 1995, 2000), and the collapse of eel grass beds, leading to global extinction of the eel grass limpet Lottia albicans (Carlton, 1993; Carlton et al., 1991).
Challenges in the Surveillance of Wildlife for Fungal and Other Pathogens
The emergence of so many high-impact diseases in wildlife and the role of wildlife as reservoirs for human emerging infectious diseases, or EIDs (Mahy and Brown, 2000; Taylor et al., 2001), have led to expanding efforts in surveillance of wildlife populations, both for new pathogens of human significance and for potential EIDs affecting wildlife. However, a number of important factors hinder this strategy. Here, we highlight the two most important challenges to effective global surveillance in wildlife, as well as review some approaches to address these challenges; and put them into context for emerging pathogens of humans and wildlife, and for fungal diseases in particular.
Jurisdictional Problems in the Surveillance of Wildlife
The agencies, funding bodies, non-governmental organizations (NGOs), and researchers that normally work with wildlife populations are usually distinct from those involved in public health and agricultural diseases. For example, in
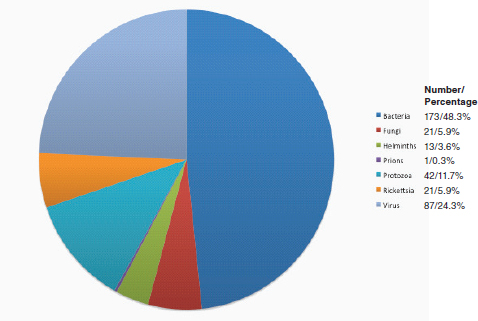
Figure A7-1 Proportion of emerging infectious diseases caused by different taxonomic groups of pathogens. Fungi are responsible for only 5.9 percent of emerging human disease, yet have a disproportionate impact on wildlife, with responsibility for a series of major population declines and extinctions. The data are taken from the database published in a recent global analysis of emerging infectious diseases of humans over the last four decades.
SOURCE: Adapted from Jones et al. (2008).
the United States, the U.S. Fish and Wildlife Service is the agency responsible for protecting wildlife, but if the threat is an emerging pathogen, this agency has little capacity for outbreak investigation and control. Similarly, the national agency responsible for funding ecological research in the United States is the National Science Foundation (NSF). The National Institutes of Health (NIH) oversees a broad range of issues, from infections to organ dysfunction to mental health and other areas. The U.S. Department of Agriculture oversees agricultural health. At the international scale, the intergovernmental agency to protect wildlife is the International Union for the Conservation of Nature (IUCN), whereas the global health agenda falls under the World Health Organization (WHO), agricultural health under the Food and Agriculture Organization (FAO), and trade-related disease issues under the World Organisation for Animal Health (OIE). This siloed approach is followed by many countries globally; they tend to have separate ministries for health, agriculture, trade, and environment/forestry/wildlife. These approaches work well until the threats to human health cross these jurisdictional boundaries. With emerging diseases, they have done so repeatedly. For example, the emergence of severe acute respiratory syndrome involved wildlife reservoir
species (Li et al., 2005), the national and international trade in hunted and farmed wildlife and livestock (Xu et al., 2004), and international travel and migration (Anderson et al., 2004). Likewise, the global emergence of amphibian chytridiomycosis has been linked to trade and climate change (Lips et al., 2008), and involves the medical industry (Weldon et al., 2004), the production of amphibians for food (Schloegel et al., 2009), and introduced or invasive species (Kilpatrick et al., 2010).
One simple approach to overcoming these challenges is to encourage cross-disciplinary, cross-agency collaboration. This approach to research and policy has been led by the fields of “conservation medicine” (Daszak et al., 2004), One Health (Karesh and Cook, 2005), and EcoHealth (Daszak, 2009; Wilcox and Daszak, 2006). In the United States, some efforts have successfully bridged the funding gap between NIH and NSF, notably the Ecology of Infectious Diseases program launched jointly by the NSF and the NIH John E. Fogarty International Center in 2000 (Scheiner and Rosenthal, 2006). Likewise, there is a unique U.S. federal agency with a specific remit to address with wildlife diseases, the National Wildlife Health Center (NWHC) (Fleischli et al., 2004; Skerratt et al., 2005). The NWHC has been conducting surveillance, monitoring, investigation, research, and response on wildlife diseases for 35 years, and is registered with the Centers for Disease Control and Prevention Select Agent Program, marking it as a laboratory of significant relevance to human and livestock as well as wildlife health. It has a sophisticated network of laboratories, including Biosecurity Level 3 biocontainment labs, necropsy suites, and isolation rooms. In addition, it publishes quarterly reports of mortality investigations, and acts as a national focal point for similar activities in universities and NGOs. At the intergovernmental scale, there has been a recent flurry of activity to bring together agencies around the One Health agenda, including formal links among the OIE, FAO, and WHO, which originated from their collaborative efforts to tackle avian influenza (Anderson et al., 2010). Additionally, wildlife health has two significant nuclei within the United Nations system.
First, in the IUCN, there is the Species Survival Commission Wildlife Health Specialist Group (http://www.iucn.org/about/work/programmes/species/about_ssc/specialist_groups/specialist_group_pprofiles/veterinary_sg_profile/), which has a network of more than 400 wildlife veterinarians and researchers globally. Second, the OIE has a Working Group on Wildlife Diseases (http://web.oie.int/wildlife/eng/en_wildlife.htm), which has operated for more than 15 years and advises the OIE on wildlife health issues.
These initiatives have begun to bring diverse disciplines together to understand the drivers and impacts of wildlife EIDs, and to conduct effective surveillance and control of wildlife as reservoirs for human EIDs. However, they could be improved significantly with some simple approaches. First, within the United States, the government has the capacity to form interagency task forces for specific issues that cross agency mandates. In a previous administration, the complex issue of amphibian declines was addressed with the formation of the Interagency
Taskforce on Amphibian Declines and Deformities. Similar task forces are likely to be useful to address the need for better surveillance of the wildlife trade for pathogens (Smith et al., 2009a), or the threat of white-nose syndrome in bats. Second, at a global scale, strengthening of laboratory capacity and personnel for wildlife diseases, and support for One Health approaches from the development community, could be extremely useful in fostering linkages among disparate ministries, universities, NGOs, and others. Recently, the U.S. Agency for International Development launched an Emerging Pandemic Threats program that specifically adopted a One Health approach to build capacity in the regions where emerging zoonoses most commonly originate (http://www.usaid.gov/our_work/global_health/home/News/ai_docs/ept_brochure.pdf) (Daszak, 2009). This includes specific collaboration among human and veterinary medical scientists; ministries of health, agriculture, and environment; and OIE, FAO, and WHO.
Limited Resources for Surveillance and Lack of Predictive Capacity
During the past decade, significant economic investment has been made in global surveillance for EIDs. Funding was provided by intergovernmental agencies, and national governments in particular, to counter the specific threat of H5N1 avian influenza. Although much of the surveillance and control efforts focused on Southeast Asia, the recent emergence of an H1N1 triple reassortant (Neumann et al., 2009; Smith et al., 2009b; WHO gets mixed reviews for H1N1 response, 2011) from a different region may have changed the public’s perception of pandemic risk, and a shift of funding priorities. At the same time, the prominence of H5N1 in the media over the past decade and its inability so far to mount a pandemic heightens the perception of emerging diseases as difficult to predict or forecast. With limited global resources for pandemic prevention or control, predictive efforts that can help target resources are greatly needed.
Recent work has shown that analyzing past trends in disease emergence and correcting data for surveillance biases allows us a strategy to identify the regions most likely to propagate a new emerging zoonosis (Jones et al., 2008; King et al., 2006; Taylor et al., 2001; Woolhouse and Gowtage-Sequeria, 2005). These approaches may provide a way to geographically target surveillance efforts. They suggest that surveillance programs should target regions with high biodiversity and dense human populations, essentially in tropical and subtropical regions (Jones et al., 2008). This approach essentially analyzes the underlying drivers of emerging diseases, which are usually socioeconomic factors (e.g., travel, trade, agricultural changes) or environmental factors (e.g., climate, land use change). If we adapt this to fungal pathogens, we see that two drivers in particular are important—climate and trade. For human EIDs, analysis of the database in Jones et al. (2008) suggests that the majority of emerging fungal diseases were associated with HIV/AIDS or antimicrobial use. For plant EIDs, analyses suggest that agricultural trade and climate are the most important drivers of new emerging diseases (Anderson et al., 2004). With the limited work that has been conducted
on emerging fungal pathogens of wildlife, trade appears to be an important driver of emergence. For example, significant evidence points to the global amphibian trade for food as a major driver of the spread of the amphibian fungal disease chytridiomycosis (Schloegel et al., 2010). The similarity of U.S. isolates of Geomyces destructans (the causative agent of bat white-nose syndrome) with those from Europe and its continued spread as if from a focal point at a tourist cave in New York state strongly suggest anthropogenic introduction.
Two important recommendations emerge: First, targeting of wildlife trade routes, wet markets,27 and international ports of entry for surveillance of wildlife may be particularly fruitful in identifying novel EIDs (Karesh et al., 2005; Pavlin et al., 2009; Smith et al., 2009a). Second, use of novel modeling approaches to geographically targeting wildlife for surveillance may be particularly useful for identifying novel pathogens. This “smart surveillance” approach (see Daszak, 2009) would help identify a pool of microbes harbored by a reservoir, some of which ultimately may become zoonotic under the right circumstances. Clearly, issues remain in that many microbes will be unable make the species jump to humans, so that distinguishing the truly potential zoonoses from these is not a simple task. However, conducting PCR-based surveillance in wildlife using conserved (or “degenerate”) primers for known pathogen groups would allow the targeting of novel agents related to known pathogens. This would enable a greater understanding of the risk of new zoonoses from each wildlife species examined, and would be a simple, cost-effective way to better evaluate the diversity of likely zoonoses. Diagnostic assays could then be developed for these pathogens, and used to screen people who live in close contact with the wildlife reservoirs.
References
Aguirre, A. A., and G. M. Tabor. 2008. Global factors driving emerging infectious diseases: Impact on wildlife populations. Annals of the New York Academy of Sciences 1149:1–3.
Anderson, P. K., A. A. Cunnigham, N. G. Patel, F. J. Morales, P. R. Epstein, and P. Daszak. 2004. Emerging infectious diseases of plants: Crop homogeneity, pathogen pollution and climate change drivers. Trends in Ecology and Evolution 19(10):535–544.
Anderson, R. M., C. Fraser, A. C. Ghani, C. A. Donnelly, S. Riley, N. M. Ferguson, G. M. Leung, T. H. Lam, and A. J. Hedley. 2004. Epidemiology, transmission dynamics and control of SARS: The 2002–2003 epidemic. Philosophical Transactions of the Royal Society of London Series B: Biological Sciences 359:1091–1105.
Anderson, T., et al. 2010. FAO–OIE–WHO Joint Technical Consultation on avian influenza at the human–animal interface. Influenza and Other Respiratory Viruses 4:1–29.
Berger, L., R. Speare, P. Daszak, D. E. Green, A. A. Cunningham, C. L. Goggin, R. Slocombe, M. A. Ragan, A. D. Hyatt, K. R. McDonald, H. B. Hines, K. R. Lips, G. Marantelli, and H. Parkes. 1998. Chytridiomycosis causes amphibian mortality associated with population declines in the rain forests of Australia and Central America. Proceedings of the National Academy of Sciences, USA 95:9031–9036.
_____________________
27 Markets which sell live wildlife (often mixed with livestock) are called “Wetmarkets,” particularly with reference to Asia.
Blaustein, A. R. 1994. Chicken little or Nero’s fiddle? A perspective on declining amphibian populations. Herpetologica 50:85–97.
Blehert, D. S., A. C. Hicks, M. Behr, C. U. Meteyer, B. M. Berlowski-Zier, E. L. Buckles, J. T. Coleman, S. R. Darling, A. Gargas, R. Niver, J. C. Okoniewski, R. J. Rudd, and W. B. Stone. 2009. Bat white-nose syndrome: An emerging fungal pathogen? Science 323:227.
Carlton, J. T. 1993. Neoextinctions of marine-invertebrates. American Zoologist 33:499–509.
Carlton, J. T., G. J. Vermeij, D. R. Lindberg, D. A. Carlton, and E. C. Dudley. 1991. The 1st historical extinction of a marine invertebrate in an ocean-basin—the demise of the eelgrass limpet Lottiaalveus. Biological Bulletin 180:72–80.
Chaturvedi, V., and S. Chaturvedi. 2011. Editorial: What is in a name? A proposal to use geomycosis instead of white nose syndrome (WNS) to describe bat infection caused by Geomyces destructans. Mycopathologia 171:231–233.
Crawford, A. J., K. R. Lips, and E. Bermingham. 2010. Epidemic disease decimates amphibian abundance, species diversity, and evolutionary history in the highlands of central Panama. Proceedings of the National Academy of Sciences, USA 107:13777–13782.
Cunningham, A. A. 2005. A walk on the wild side—emerging wildlife diseases: They increasingly threaten human and animal health. British Medical Journal 331:1214–1215.
Daszak, P. 2009. A call for “smart surveillance”: A lesson learned from H1N1. Ecohealth 6:1–2.
Daszak, P., and A. A. Cunningham. 1999. Extinction by infection. Trends in Ecology & Evolution 14:279.
Daszak, P., A. A. Cunningham, and A. D. Hyatt. 2000. Emerging infectious diseases of wildlife—threats to biodiversity and human health. Science 287:443–449.
Daszak, P., G. M. Tabor, A. M. Kilpatrick, J. Epstein, and R. Plowright. 2004. Conservation medicine and a new agenda for emerging diseases. Annals of the New York Academy of Sciences 1026:1–11.
Daszak, P., D. E. Scott, A. M. Kilpatrick, C. Faggioni, J. W. Gibbons, and D. Porter. 2005. Amphibian population declines at Savannah River site are linked to climate, not chytridiomycosis. Ecology 86:3232–3237.
Deem, S. L., W. B. Karesh, and W. Weisman. 2001. Putting theory into practice: Wildlife health in conservation. Conservation Biology 15:1224–1233.
Fischer, J. R., D. E. Stallknecht, M. P. Luttrell, A. A. Dhondt, and K. A. Converse. 1997. Mycoplasmal conjunctivitis in wild songbirds: The spread of a new contagious disease in a mobile host population. Emerging Infectious Diseases 3:69–72.
Fleischli, M. A., J. C. Franson, N. J. Thomas, D. L. Finley, and W. Riley. 2004. Avian mortality events in the United States caused by anticholinesterase pesticides: A retrospective summary of National Wildlife Health Center records from 1980 to 2000. Archives of Environmental Contamination and Toxicology 46:542–550.
Frick, W. F., J. F. Pollock, A. C. Hicks, K. E. Langwig, D. S. Reynolds, G. G. Turner, C. M. Butchkoski, and T. H. Kunz. 2010. An emerging disease causes regional population collapse of a common North American bat species. Science 329(5992):679–682.
Jones, K. E., N. G. Patel, M. A. Levy, A. Storeygard, D. Balk, J. L. Gittleman, and P. Daszak. 2008. Global trends in emerging infectious diseases. Nature 451:990–994.
Karesh, W. B., and R. A. Cook. 2005. The human–animal link, one world–one health. Foreign Affairs 84:38–50.
Karesh, W. B., R. A. Cook, E. L. Bennett, and J. Newcomb. 2005. Wildlife trade and global disease emergence. Emerging Infectious Diseases 11:1000–1002.
Kilpatrick, A. M., C. J. Briggs, and P. Daszak. 2010. The ecology and impact of chytridiomycosis: An emerging disease of amphibians. Trends in Ecology & Evolution 25:109–118.
King, D. A., C. Peckham, J. K. Waage, J. Brownlie, and M. E. J. Woolhouse. 2006. Infectious diseases: Preparing for the future. Science 313:1392–1393.
Li, W. D., Z. Shi, M. Yu, W. Ren, C. Smith, J. H. Epstein, H. Wang, G. Crameri, Z. Hu, H. Zhang, J. Zhang, J. McEachern, H. Field, P. Daszak, B. T. Eaton, S. Zhang, and L. F. Wang. 2005. Bats are natural reservoirs of SARS-like coronaviruses. Science 310:676–679.
Lips, K. R., F. Brem, R. Brenes, J. D. Reeve, R. A. Alford, J. Voyles, C. Carey, L. Livo, A. P. Pessier, and J. P. Collins. 2006. Emerging infectious disease and the loss of biodiversity in a neotropical amphibian community. Proceedings of the National Academy of Sciences, USA 103:3165–3170.
Lips, K. R., J. Diffendorfer, J. R. Mendelson, and M. W. Sears. 2008. Riding the wave: Reconciling the roles of disease and climate change in amphibian declines. PLoS Biology 6:441–454.
Mahy, B. W. J., and C. C. Brown. 2000. Emerging zoonoses: Crossing the species barrier. J Rev Sci Tech OIE 19:33–40.
Mendelson, J. R., K. R. Lips, R. W. Gagliardo, G. B. Rabb, J. P. Collins, J. E. Diffendorfer, P. Daszak, D. R. Ibáñez, K. C. Zippel, D. P. Lawson, K. M. Wright, S. N. Stuart, C. Gascon, H. R. da Silva, P. A. Burrowes, R. L. Joglar, E. La Marca, S. Lötters, L. H. du Preez, C. Weldon, A. Hyatt, J. V. Rodriguez-Mahecha, S. Hunt, H. Robertson, B. Lock, C. J. Raxworthy, D. R. Frost, R. C. Lacy, R. A. Alford, J. A. Campbell, G. Parra-Olea, F. Bolaños, J. J. Domingo, T. Halliday, J. B. Murphy, M. H. Wake, L. A. Coloma, S. L. Kuzmin, M. S. Price, K. M. Howell, M. Lau, R. Pethiyagoda, M. Boone, M. J. Lannoo, A. R. Blaustein, A. Dobson, R. A. Griffiths, M. L. Crump, D. B. Wake, and E. D. Brodie, Jr. 2006. Biodiversity: Confronting amphibian declines and extinctions. Science 313:48.
Miller, M. W., and E. S. Williams. 2004. Chronic wasting disease of cervids. Mad Cow Disease and Related Spongiform Encephalopathies 284:193–214.
Nettles, V. F. 1996. Reemerging and emerging infectious diseases: Economic and other impacts on wildlife—Transport of animals sometimes spreads infections, while other outbreaks are a mystery. ASM News 62:589–591.
Neumann, G., T. Noda, and Y. Kawaoka. 2009. Emergence and pandemic potential of swine-origin H1N1 influenza virus. Nature 459:931–939.
Pavlin, B. I., L. M. Schloegel, and P. Daszak. 2009. Risk of importing zoonotic diseases through wildlife trade, United States. Emerging Infectious Diseases 15:1721–1726.
Pechmann, J. H. K., D. E. Scott, R. D. Semlitsch, J. P. Caldwell, L. J. Vitt, and J. W. Gibbons. 1991. Declining amphibian populations: The problem of separating human impacts from natural fluctuations. Science 253:892–895.
Robinson, R. A., B. Lawson, M. P. Toms, K. M. Peck, J. K. Kirkwood, J. Chantrey, I. R. Clatworthy, A. D. Evans, L. A. Hughes, O. C. Hutchinson, S. K. John, T. W. Pennycott, M. W. Perkins, P. S. Rowley, V. R. Simpson, K. M. Tyler, and A. A. Cunningham. 2010. Emerging infectious disease leads to rapid population declines of common British birds. PLoS ONE 5(8):1–12.
Scheiner, S. M., and J. P. Rosenthal. 2006. Ecology of infectious disease: Forging an alliance. Ecohealth 3:204–208.
Schloegel, L. M., J. M. Hero, L. Berger, R. Speare, K. McDonald, and P. Daszak. 2006. The decline of the sharp-snouted day frog (Taudactylus acutirostris): The first documented case of extinction by infection in a free-ranging wildlife species? Ecohealth 3:35–40.
Schloegel, L. M., A. M. Picco, A. M. Kilpatrick, A. J. Davies, A. G. Hyatt, and P. Daszak. 2009. Magnitude of the U.S. trade in amphibians and presence of Batrachochytrium dendrobatidis and ranavirus infection in imported North American bullfrogs (Rana catesbeiana). Biological Conservation 142:1420–1426.
Schloegel, L. M., P. Daszak, A. A. Cunningham, R. Speare, and B. Hill. 2010. Two amphibian diseases, chytridiomycosis and ranaviral disease, are now globally notifiable to the World Organization for Animal Health (OIE): An assessment. Diseases of Aquatic Organisms 92:101–108.
Schwartz, M. W., S. M. Hermann, and C. S. Vogel. 1995. The catastrophic loss of Torreya-taxifolia—assessing environmental induction of disease hypotheses. Ecological Applications 5:501–516.
Schwartz, M. W., S. M. Hermann, and P. J. van Mantgem. 2000. Estimating the magnitude of decline of the Florida torreya (Torreya taxifolia Arn.). Biological Conservation 95:77–84.
Sigurdson, C. J. 2008. A prion disease of cervids: Chronic wasting disease. Veterinary Research 39–41.
Skerratt, L. F., J. C. Franson, C. U. Meteyer, and T. E. Hollmen. 2005. Causes of mortality in sea ducks (Mergini) necropsied at the USGS-National Wildlife Health Center. Waterbirds 28:193–207.
Skerratt, L. F., L. Berger, R. Speare, S. Cashins, K. Raymond McDonald, A. D. Phillott, H. B. Hines, and N. Kenyon. 2007. Spread of chytridiomycosis has caused the rapid global decline and extinction of frogs. Ecohealth 4:125–134.
Smith, G. J. D., D. Vijaykrishna, J Bahl, S. J. Lycett, M. Worobey, O. G. Pybus, S. K. Ma, C. L. Cheung, J. Raghwani, S. Bhatt, J. S. Peiris, Y. Guan, and A. Rambaut. 2009b. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459:1122–1126.
Smith, K. F., M. Behrens, L. M. Schloegel, N. Maranao, S. Burgiel, and P. Daszak. 2009a. Reducing the risks of the wildlife trade. Science 324:594–595.
Taylor, L. H., S. M. Latham, and M. E. Woolhouse. 2001. Risk factors for human disease emergence. Philosophical Transactions of the Royal Society of London B: Biological Sciences 356:983–989.
Weldon, C., L. H. du Preez, A. D. Hyatt, R. Muller, and R. Speare. 2004. Origin of the amphibian chytrid fungus. Emerging Infectious Diseases 10:2100–2105.
WHO gets mixed reviews for H1N1 response. 2011. Science 331:1371–1371.
Wilcox, B. A., and P. Daszak. 2006. Launching the International EcoHealth Association. EcoHealth 3:125–126.
Wildlife and emerging disease. 2009. Veterinary Record 165:458–459.
Williams, E. S., T. Yuill, M. Artois, J. Fischer, and S. A. Haigh. 2002. Emerging infectious diseases in wildlife. Revue Scientifique Et Technique De L Office International Des Epizooties 21:139–157.
Woolhouse, M. E. J., and S. Gowtage-Sequeria. 2005. Host range and emerging and re-emerging pathogens. Emerging Infectious Diseases 11:1842–1847.
Xu, R. H., J. F. He, M. R. Evans, G. W. Peng, H. E. Field, D. W. Yu, C. K. Lee, H. M. Luo, W. S. Lin, P. Lin, L. H. Li, W. J. Liang, J. Y. Lin, and A. Schnur. 2004. Epidemiologic clues to SARS origin in China. Emerging Infectious Diseases 10:1030–1037.
Introduction
Human disease resulting from infection by Coccidioides spp. was first recognized late in the 19th century. Since then, with more information and changing demographics, our understanding of this problem and our perception of its importance has evolved in many ways (Galgiani, 2007). First thought of as a rare
_____________________
28 Valley Fever Center for Excellence, University of Arizona College of Medicine.
29 Correspondence and current address: John N. Galgiani, M.D.; Professor, University of Arizona College of Medicine; Director, Valley Fever Center for Excellence; P.O. Box 245215, Tucson, AZ 85724; Tel.: 520-626-4968; Fax: 520-626-4971; e-mail: spherule@u.arizona.edu.
and always fatal illness, coccidioidomycosis later was appreciated as a common and frequently self-limited illness known as valley fever, in which only a small percentage of those affected suffered serious complications (Smith, 1940).
With U.S. military training within the endemic regions of California and Arizona during World War II, the potential of coccidioidomycosis as a significant problem affecting military readiness quickly became apparent, a problem that persists for the military into the present (Crum-Cianflone, 2007; Smith, 1958). With the rapid and extensive population expansion within south-central Arizona over the past decades, the impact of coccidioidomycosis has emerged from a rural to a much larger public health problem. With continued population growth within the central valley of California, in Mexico along the U.S. border, and in other endemic regions throughout the Western Hemisphere, it is expected that the numbers of infected persons will continue to increase. In addition to the medical problem for humans, a variety of other species, especially dogs, are susceptible to coccidioidomycosis and suffer considerable morbidity and mortality as a result (Shubitz, 2007).
A comprehensive assessment of coccidioidomycosis should address the fungus as it exists in the environment as well as how it creates medical problems. In both arenas, there are opportunities for better understanding which in turn could lead to improvements in public health. Hopefully, this article will be useful to call attention to where our knowledge is limited and how advances in those areas could benefit prevention and management of coccidioidal disease.
Coccidioides spp. in the Environment
Our primary source for our understanding of the relative endemnicity for regions within the United States is derived from studies conducted in the 1950s of coccidioidin skin-test prevalence of naval recruits from across the country (Palmer et al., 1957). These primarily include extensive portions of southern California, the lower deserts of Arizona, west Texas, and smaller areas of Utah, Nevada, and New Mexico. More recent information from California and Arizona (Hector et al., 2011), where coccidioidomycosis is a reportable infectious disease, is consistent with the earlier estimate. Coccidioidal infections are found in numerous other countries within the Western Hemisphere. What is known about these areas also suggests that the endemic distribution has been stable over the past several decades (Laniado-Laborin, 2007). On the other hand, the endemic regions in a longer time scale may not be constant. Recent archeological findings in Nebraska identified a Holocene bison dating back 8,500 years with coccidioidomycosis in a bone (Morrow, 2006). Also, models of wind pattern trends with global warming would predict increased westerly currents from the Coccidioides-endemic regions of west Texas toward the more populated portions of the state (Reheis and Rademaekers, 1997). Thus, with climate change we might see the endemic region expand.
Recent population studies of fungal isolates have identified genetic differences in isolates from patients in California (now classified as Coccidioides immitis) as compared to isolates from elsewhere (now classified as Coccidioides posadasii) (Burt et al., 1997; Fisher et al., 2000, 2001). This geographic separation is surprising, given the likelihood that coccidioidal spores should travel freely in and out of California, either by wind currents or as contaminants of trains, planes, or road vehicles. One possibility is that C. immitis is much better suited to its California niche than is C. posadasii and vice versa. Another explanation might relate to the biology of how soil becomes infected and endemic regions become established. For example, simply inoculating new soil may not be sufficient to create a new coccidioidal colony. Instead, a more complex interaction with rodents, plants, or other factors in the environment may be needed. Reinforcing this speculation is the sparse distribution of Coccidioides spp. within the environment. For example, in one systematic study in the central valley of California, only four genetically distinct isolates were found from 720 soil samples (Greene et al., 2000). Soil sampling in southern Arizona has also found isolates in a very small proportion of samples tested (Barker et al., 2010).
Researchers have known for some time that Coccidioides spp. are more prevalent in specimens from rodent burrows than from random subsurface soil (Elconin et al., 1957), although the reasons for this association remain unclear (Barker et al., 2010). In contrast, repeated sampling from a site known to be positive often yields positive results. One site identified as containing Coccidioides spp. in the 1960s (Converse and Reed, 1966) has again yielded C. posadasii half a century later (Personal communication, January 2011, M. J. Orbach). The sparse and stable nature of coccidioidal residence within the endemic regions affords an implication as to what factors are more likely to be associated with risk of exposure. Although there have been well-documented outbreaks associated with archeological and other dirt-disrupting sites (Pappagianis, 1983; Werner et al., 1972), in general occupational exposure does not appear to be a major risk factor (Kim et al., 2009). What might appear to be a paradox is actually consistent with the sparse prevalence of the fungus within the soil of even its most endemic regions, in which case much of the soil-disrupting activities occur at sites where the fungus is not present. In contrast, climate and especially wind patterns have been shown to affect seasonal incidence of infection (Comrie and Glueck, 2007; Hugenholtz, 1957). Taken together, the evidence would suggest that simply length of endemic exposure rather than a specific activity constitutes the more dominant effect for risk of infection.
With our current understanding of Coccidioides spp. residing in the environment, it is not possible to predict whether it exists in a specific location with any degree of precision from a physical or chemical analysis of the soil. Furthermore, high-throughput methods do not exist for microbial detection of Coccidioides spp. in soil samples. These missing tools prevent us from identifying specific locations which, if disrupted, would likely create a release of fungal spores. If
methods were available to do this, methods for treating the soil exist that would minimize or prevent this exposure from happening. Advances in this area would have practical public health benefits.
Coccidioidomycosis: The Scope of the Problem
Statistics about newly diagnosed coccidioidomycosis from California and Arizona were expected to total more than 16,000 new infections in 2010 (Figure A8-1). The sharp increase in reported cases in Arizona in 2009 is the result of an administrative change by a single large clinical laboratory to report a more sensitive coccidioidal serologic test as indicative of a new infection when positive. However, for both states there has been increased disease activity in the last half of 2010 that is unexplained.
In Arizona, the Department of Health Services conducted a telephone questionnaire of 10 percent of newly identified patients in 2007. It provides a better understanding of the impact of this disease (Tsang et al., 2010). According to the survey, illness lasted for an average of 6 months; three quarters of employed
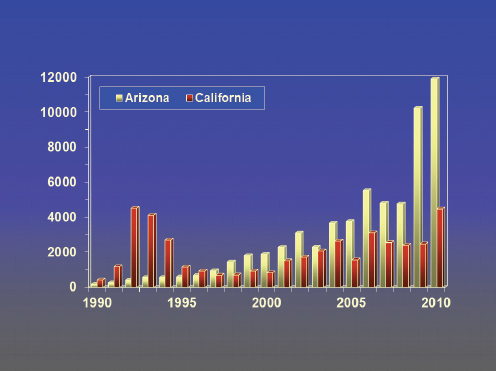
Figure A8-1 Annual cases of coccidioidomycosis.
SOURCE: Figure courtesy of John N. Galgiani (data from the California Department of Health and Arizona Department of Health Services).
patients lost more than a month of work; a quarter of patients needed 10 or more physician visits; and 40 percent of patients required hospitalization for their illness (Tsang et al., 2010). The same report shows Arizona hospital costs for coccidioidomycosis were more than $86 million. Extrapolating from these costs, estimates that include all outpatient medical care, often lasting for years if not entire lives, could easily reach a quarter of a billion dollars.
As significant as these findings are, other projections suggest that the actual number of persons seeking medical attention for coccidioidal infection is several times greater than those diagnosed and included in state public health statistics. One study found that only 3–13 percent of patients with pneumonia in Phoenix were tested for coccidioidomycosis (Chang et al., 2008). By contrast, a prospective study in Tucson in which patients with community-acquired pneumonia were tested for coccidioidomycosis demonstrated that nearly a third of these subjects had a coccidioidal infection (Valdivia et al., 2006).
State statistics show case rates for college-age persons in Pima County to be from 34 to 48 cases per 100,000 annually. However, recent surveillance of scholarship athletes at the University of Arizona, which is in Pima County, indicated 374 cases per 100,000 (Stern and Galgiani, 2010). Further analysis suggested that the most important reason for this much higher case rate was that the athletes received many more serologic tests for coccidioidomycosis. Evidently, more patients would be accurately identified as to the true cause of their illness if patients with endemic exposure to Coccidioides spp. were tested more routinely for this possibility. This is now the recommendation of the Arizona Department of Health Services and a growing number of Arizona state medical specialty societies and other professional organizations (Tsang et al., 2010).
With the commercial and recreational growth of the southwestern United States, coccidioidomycosis has become an increasing problem for the rest of the country as well. For example, persons who develop a respiratory illness within a month after returning from vacation or business conferences in south-central Arizona would have the same risk (approximately 30 percent) that their illness is due to Coccidioides spp. as would residents of the endemic regions. Using Arizona Department of Tourism statistics for 2008, the chance of an individual visitor developing any clinical illness would be expected to be small (approximately 1 in 17,000). However, because more than 22 million persons visit Arizona for an average of 4 to 5 days, the total number of illnesses occurring after leaving Arizona would add up to more than 1,300 per year. Evidence suggests that most of these illnesses would be diagnosed incorrectly in the course of routine medical care (Standaert et al., 1995).
Even if physicians obtain appropriate testing, establishing a diagnosis of early coccidioidal infection is often difficult. Coccidioidal serology is very specific when results are positive. Moreover, in progressive forms of infection, serology is very likely to be diagnostic (Fish et al., 1990; Pappagianis and Zimmer, 1990). However, these tests are not nearly as sensitive early in the course of the
acute pneumonia syndrome, the most common manifestation of a coccidioidal infection. In one study, depending on the method of analysis, false-negative results were estimated to occur from one third to two thirds of the time on first testing (Wieden et al., 1996). Although improved methods for detecting Coccidioidesspecific antibodies may improve the sensitivity, approaches such as detection of coccidioidal DNA by polymerase chain reaction (Clark and McAllister, 1996) or detection of coccidioidal antigens by enzyme-linked immunosorbent assay (ELISA) (Durkin et al., 2008; Helfrich et al., 2011) offer theoretical advantages to earlier detection. Improvements in early diagnosis would be very useful to clinicians trying to manage their patients.
Therapy for Coccidioidomycosis and Its Current Limitations
The most serious complications of coccidioidomycosis include progressive chronic pneumonia and hematogenous spread of infection to parts of the body outside of the chest. Patients with these problems have benefited greatly from the advent of orally absorbed azole antifungal agents (Galgiani et al., 2005). Fluconazole and itraconazole are commonly used for these complications (Galgiani et al., 1993, 2000; Tucker et al., 1990). The more recently available azoles, voriconazole and posaconazole, offer additional options (Catanzaro et al., 2007; Prabhu et al., 2004; Proia and Tenorio, 2004; Stevens et al., 2007). This class of antifungal drugs has been found to be relatively safe and generally well tolerated for extended courses of administration. However, approximately a quarter of patients with such infections do not respond adequately to these drugs. Even in patients who appear to respond to azole treatment, relapses occur in approximately a third of those when treatment is discontinued. Thus, for some patients, including all patients with coccidioidal meningitis, treatment is recommended to be lifelong (Dewsnup et al., 1996). The most currently available drugs offer a suppressive effect; they do not eradicate infections or cure patients.
Surprisingly, there is virtually no published experience on the use of any antifungal drug for the most common manifestations of coccidioidal infection, that of the early respiratory syndrome. In a recent prospective observational study (Ampel et al., 2009), patients treated with oral azoles (usually fluconazole) appeared neither to improve at a faster rate nor subsequently to avoid progressive complication than did patients who did not receive antifungal treatment. Although not a randomized controlled trial, this study presents the only information available and provides little encouragement for the value of early treatment.
Prospects for new drugs to treat coccidioidomycosis are limited (Ostrosky-Zeichner et al., 2010). This is due in part to the general contraction in anti-infective drug development in general (Talbot et al., 2006), but is especially problematic for coccidioidomycosis because of its status as an orphan disease, defined by the Food and Drug Administration (FDA) as having a U.S. prevalence of less than 200,000. Even for new antifungals, such as voriconazole, posacon-
and caspofungin (Gonzalez et al., 2001, 2007), which already have FDA approval for other fungal diseases, there are very limited or no controlled clinical trials conducted in patients with any form of coccidioidomycosis.
One exception to this pattern has been the persistent interest in bringing nikkomycin Z into clinical trials. Nikkomycin Z is a competitive inhibitor of chitin synthase, first discovered by German scientists in the 1970s. It was part of a fungicide discovery program at the Bayer Company (Fiedler, 1988). Its potential as a therapeutic for coccidioidomycosis was identified in the 1980s (Hector et al., 1990). In mice, nikkomycin Z treatment produced sterile lungs under conditions in which the lungs of untreated mice yielded several million viable fungal colonies. This observation raises the possibility that nikkomycin Z might offer a curative treatment for coccidioidomycosis. If so, this would provide even more incentive to diagnose coccidioidomycosis early in order to eradicate it and thereby prevent later and serious complications. Clinical development of nikkomycin Z was begun in the 1990s by Shaman Pharmaceuticals (Galgiani, 2007). However, the program became inactive when Shaman ceased to exist in 2000, and for several years nikkomycin Z development remained dormant.
In 2005, the University of Arizona acquired the program along with several kilograms of bulk nikkomycin Z that remained from Shaman’s program. Since then faculty at the University of Arizona and a small start-up company, Valley Fever Solutions, have successfully competed for research awards and small business grants from the National Institutes of Health (NIH) and the FDA Office of Orphan Products Development. With these funds as well as philanthropic support, clinical trials with nikkomycin Z were resumed with a 2-week multidose safety trial that was completed in 2009. This support is also being used to develop a more efficient manufacturing process. Supplies of nikkomycin Z made by this new process are planned to be available to begin a Phase II clinical trial in 2011 or 2012. It is hoped that this progress will advance the program sufficiently to attract pharmaceutical or investment interest to complete the commercialization process.
Prospects for a Preventative Vaccine
A large majority of the estimated 150,000 U.S. coccidioidal infections occurring annually resolve with or without symptoms and whether or not specific antifungal treatment is instituted (Galgiani et al., 2005). Healing occurs as a result of the patient’s cellular immunity, which is remarkably durable, usually lasting a lifetime (Galgiani et al., 2010). The fact that natural infection so often produces resistance to second infections has attracted a long-time interest in engendering this immunity through active immunization, leading to a clinical trial of a whole-cell killed vaccine (Pappagianis and Valley Fever Vaccine Study Group, 1993). This preparation did not produce protection. In retrospect it was surmised that the irritation at the injection site of the fungal cell wall polysaccharides prevented sufficient doses of protective immunogens to be delivered during vaccination. For
the past 15 years, a collaboration of several research groups has yielded a number of immunogenic coccidioidal proteins and vaccines prepared from recombinant proteins with adjuvants, some of which have shown excellent protection in mice and efficacy in primates (Cole et al., 2004; Cox and Magee, 2004; Herr et al., 2007; Johnson et al., 2007; Shubitz et al., 2006; Tarcha et al., 2006).
The next step for existing recombinant vaccine candidates would be for them to be moved into clinical trials. However, these candidates have met with major challenges including developing a suitable manufacturing process; identifying a suitable and available adjuvant; and compounding a suitable formulation appropriate for human experimentation (Galgiani, 2008). None of these challenges are insurmountable, but all require significant development investment. The overall impact of coccidioidomycosis within the endemic region is not so dissimilar to that caused by polio in the United States before a polio vaccine was available (approximately 10 per 100,000 population). However, the impact of coccidioidomycosis involves a much smaller population at risk as compared to the worldwide distribution of polio. This difference in market size makes it unlikely that a commercial vaccine manufacturer will invest in developing a coccidioidal vaccine even though such a vaccine, once developed, could arguably be profitable to manufacture and distribute (Barnato et al., 2001). Moving a coccidioidal vaccine into clinical trials probably requires the discovery of new, more easily formulated protective antigens; a breakthrough in vaccine technology that greatly reduces the cost of development; or a growing public health imperative to underwrite the costs needed for vaccine development.
Summary
Coccidioidomycosis is a major public health problem for a major, growing segment of the U.S. population as well as other endemic regions throughout the Western Hemisphere. A more complete understanding of its biology and ecology where it exists in the endemic environment could lead to risk abatement strategies not currently available. Improved recognition by healthcare professionals of coccidioidomycosis as a cause of community-acquired pneumonia when it occurs in their patients could improve management. This could be assisted further by developing more sensitive and clinically available diagnostic tests based on biosignatures such as DNA or proteins from the fungus itself. Curative therapies are also needed, but none exist today. Finally, eliminating problems caused by Coccidioides spp. might be possible if a preventive vaccine were developed. Even though coccidioidomycosis is an orphan disease, pursuit of these objectives is more than justified by the potential public health benefit and the reduced medical costs to society that their achievement would provide.
Acknowledgments
This presentation was supported in part by Award Number U54AI065359 from the National Institute of Allergy and Infectious Diseases (NIAID). The content is the sole responsibility of the authors and does not necessarily represent the official views of the NIAID or NIH.
Disclosure
Dr. Galgiani is chief medical officer, chair of the board, and a significant stock holder in Valley Fever Solutions, Inc.
References
Ampel, N. M., A. Giblin, J. P. Mourani, and J. N. Galgiani. 2009. Factors and outcomes associated with the decision to treat primary pulmonary coccidioidomycosis. Clinical Infectious Diseases 48:172–178.
Barker, B. M., J. Tabor, L. Shubitz, R. Perill, and M. J. Orbach. 2010. Detection and phylogenetic analysis of Coccidioides posadasii in Arizona soil samples. Fungal Ecology. In press.
Barnato, A. E., G. D. Sanders, and D. K. Owens. 2001. Cost-effectiveness of a potential vaccine for Coccidioides immitis. Emerging Infectious Diseases 7:797–806.
Burt, A., B. M. Dechairo, G. L. Koenig, D. A. Carter, T. J. White, and J. W. Taylor. 1997. Molecular markers reveal differentiation among isolates of Coccidioides immitis from California, Arizona and Texas. Molecular Ecology 6:781–786.
Catanzaro, A., G. A. Cloud, D. A. Stevens, B. E. Levine, P. L. Williams, R. H. Johnson, A. Rendon, L. F. Mirels, J. E. Lutz, M. Holloway, and J. N. Galgiani. 2007. Safety, tolerance, and efficacy of posaconazole therapy in patients with nonmeningeal disseminated or chronic pulmonary coccidioidomycosis. Clinical Infectious Diseases 45:562–568.
Chang, D. C., S. Anderson, K. Wannemuehler, D. M. Engelthaler, L. Erhart, R. H. Sunenshine, L. A. Burwell, and B. J. Park. 2008. Testing for coccidioidomycosis among patients with community-acquired pneumonia. Emerging Infectious Diseases 14:1053–1059.
Clark, K. A., and D. McAllister. 1996. Direct detection of Coccidioides immitis in clinical specimens using target amplification. In Coccidioidomycosis, edited by H. E. Einstein and A. Catanzaro. Proceedings of the Fifth International Conference. Washington, DC: National Foundation for Infectious Diseases. Pp. 129–136.
Cole, G. T., J. M. Xue, C. N. Okeke, E. J. Tarcha, V. Basrur, R. A. Schaller, R. A. Herr, J. J. Yu, and C. Y. Hung. 2004. A vaccine against coccidioidomycosis is justified and attainable. Medical Mycology 42:189–216.
Comrie, A. C., and M. F. Glueck. 2007. Assessment of climate-coccidioidomycosis model: Model sensitivity for assessing climatologic effects on the risk of acquiring coccidioidomycosis. Annals of the New York Academy of Sciences 1111:83–95.
Converse, J. L., and R. E. Reed. 1966. Experimental epidemiology of coccidioidomycosis. Bacteriological Reviews 30:678–695.
Cox, R. A., and D. M. Magee. 2004. Coccidioidomycosis: Host response and vaccine development. Clinical Microbiology Reviews 17:804–839, table.
Crum-Cianflone, N. F. 2007. Coccidioidomycosis in the U.S. military: A review. Annals of the New York Academy of Sciences 1111:112–121.
Dewsnup, D. H., J. N. Galgiani, J. R. Graybill, M. Diaz, A. Rendon, G. A. Cloud, and D. A. Stevens. 1996. Is it ever safe to stop azole therapy for Coccidioides immitis meningitis? Annals of Internal Medicine 124:305–310.
Durkin, M., P. Connolly, T. Kuberski, R. Myers, B. M. Kubak, D. Bruckner, D. Pegues, and L. J. Wheat. 2008. Diagnosis of coccidioidomycosis with use of the Coccidioides antigen enzyme immunoassay. Clinical Infectious Diseases 47:e69–e73.
Elconin, A. F., R. O. Egeberg, and R. Lubarsky. 1957. Growth pattern of Coccidiodes immitis in the soil of an endemic area. U.S. Public Health Service Publication 575:168–170.
Fiedler, H. P. 1988. The nikkomycin story. In Sekundarmetabolismus bei Mikroorganismen, edited by H. von Willi Kuhn and H.-P. Fiedler. Tubingen, Germany: Attempto Verlag.
Fish, D. G., N. M. Ampel, J. N. Galgiani, C. L. Dols, P. C. Kelly, C. H. Johnson, D. Pappagianis, J. E. Edwards, R. B. Wasserman, R. J. Clark, D. Antoniskis, R. A. Larsen, S. J. Englender, and E. A. Petersen. 1990. Coccidioidomycosis during human immunodeficiency virus infection. A review of 77 patients. Medicine 69:384–391.
Fisher, M. C., G. L. Koenig, T. J. White, and J. W. Taylor. 2000. Pathogenic clones versus environmentally driven population increase: Analysis of an epidemic of the human fungal pathogen Coccidioides immitis. Journal of Clinical Microbiology 38:807–813.
Fisher, M. C., G. L. Koenig, T. J. White, G. San Blas, R. Negroni, I. G. Alvarez, B. Wanke, and J. W. Taylor. 2001. Biogeographic range expansion into South America by Coccidioides immitis mirrors New World patterns of human migration. Proceedings of the National Academy of Sciences, USA 98:4558–4562.
Galgiani, J. N. 2007. Coccidioidomycosis: Changing perceptions and creating opportunities for its control. Annals of the New York Academy of Sciences 1111:1–18.
———. 2008. Vaccines to prevent systemic mycoses: Holy grails meet translational realities. Journal of Infectious Diseases 197:938–940.
Galgiani, J. N., A. Catanzaro, G. A. Cloud, J. Higgs, B. A. Friedman, R. A. Larsen, and J. R. Graybill. 1993. Fluconazole therapy for coccidioidal meningitis. The NIAID–Mycoses Study Group. Annals of Internal Medicine 119:28–35.
Galgiani, J. N., A. Catanzaro, G. A. Cloud, R. H. Johnson, P. L. Williams, L. F. Mirels, F. Nassar, J. E. Lutz, D. A. Stevens, P. K. Sharkey, V. R. Singh, R. A. Larsen, K. L. Delgado, C. Flanigan, and M. G. Rinaldi. 2000. Comparison of oral fluconazole and itraconazole for progressive, nonmeningeal coccidioidomycosis. A randomized, double-blind trial. Mycoses Study Group. Annals of Internal Medicine 133:676–686.
Galgiani, J. N., N. M. Ampel, J. E. Blair, A. Catanzaro, R. H. Johnson, D. A. Stevens, and P. L. Williams. 2005. Coccidioidomycosis. Clinical Infectious Diseases 41:1217–1223.
Gonzalez, G. M., R. Tijerina, L. K. Najvar, R. Bocanegra, M. Luther, M. G. Rinaldi, and J. R. Graybill. 2001. Correlation between antifungal susceptibilities of Coccidioides immitis in vitro and antifungal treatment with Caspofungin in a mouse model. Antimicrobial Agents and Chemotherapy 45:1854–1859.
Gonzalez, G. M., G. Gonzalez, L. K. Najvar, and J. R. Graybill. 2007. Therapeutic efficacy of caspofungin alone and in combination with amphotericin B deoxycholate for coccidioidomycosis in a mouse model. Journal of Antimicrobial Chemotherapy 60:1341–1346.
Greene, D. R., G. Koenig, M. C. Fisher, and J. W. Taylor. 2000. Soil isolation and molecular identification of Coccidioides immitis. Mycologia 92:406–410.
Hector, R. F., B. L. Zimmer, and D. Pappagianis. 1990. Evaluation of nikkomycins X and Z in murine models of coccidioidomycosis, histoplasmosis, and blastomycosis. Antimicrobial Agents and Chemotherapy 34:587–593.
Hector, R. F., G. W. Rutherford, C. A. Tsang, L. M. Erhart, O. McCotter, K. Komatsu, S. M. Anderson, F. Tabnak, D. J. Vugia, Y. Yang, and J. N. Galgiani. 2011. Public health impact of coccidioidomycosis in California and Arizona. International Journal of Environmental Research and Public Health 8(4):1150–1173.
Helfrich, F. S. E., L. F. Shubitz, T. Peng, K. S. Knox, N. M. Ampel, J. N. Galgiani, and V. H. Wysocki. 2011. Proteomic identification of coccidioidal antigens from lung fluid of infected mice: MRM analysis to confirm presence in biological fluids. Proceedings of the Third Annual Meeting of the Association for Mass Spectrometry Applications to the Clinical Lab, San Diego, CA, February 2011.
Herr, R. A., C. Y. Hung, and G. T. Cole. 2007. Evaluation of two homologous proline-rich proteins of Coccidioides posadasii as candidate vaccines against coccidioidomycosis. Infection and Immunity 75:5777–5787.
Hugenholtz, P. G. 1957. Climate and coccidioidomycosis. In Proceedings of Symposium on Coccidioidomycosis, Phoenix, AZ. Atlanta, GA: Public Health Service Publication 575:136–143.
Johnson, S. M., N. W. Lerche, D. Pappagianis, J. L. Yee, J. N. Galgiani, and R. F. Hector. 2007. Antigenicity, safety and efficacy of a recombinant coccidioidomycosis vaccine in cynomolgus maques (Macaca fascicularis). Annals of the New York Academy of Sciences 1111:290–300.
Kim, M. M., J. E. Blair, E. J. Carey, Q. Wu, and J. D. Smilack. 2009. Coccidioidal pneumonia, Phoenix, AZ, USA, 2000–2004. Emerging Infectious Diseases 15:397–401.
Laniado-Laborin, R. 2007. Expanding understanding of epidemiology of coccidioidomycosis in the Western Hemisphere. Annals of the New York Academy of Sciences 1111:19–34.
Morrow, W. 2006. Holocene coccidioidomycosis: Valley Fever in early Holocene bison (Bison antiquus). Mycologia 98:669–677.
Ostrosky-Zeichner, L., A. Casadevall, J. N. Galgiani, F. C. Odds, and J. H. Rex. 2010. An insight into the antifungal pipeline: Selected new molecules and beyond. Nature Reviews Drug Discovery 9:719–727.
Palmer, C. E., P. Q. Edwards, and W. E. Allfather. 1957. Characteristics of skin reactions to coccidioidin and histoplasmin with evidence of an unidentified source of sensitization. American Journal of Hygeine 66:196–213.
Pappagianis, D. 1983. Coccidioidomycosis (San Joaquin or Valley Fever). In Occupational Mycoses, edited by A. DiSalvo. Philadelphia, PA: Lea and Febiger. Pp. 13–28.
Pappagianis, D., and Valley Fever Vaccine Study Group. 1993. Evaluation of the protective efficacy of the killed Coccidioides immitis spherule vaccine in humans. American Review of Respiratory Disease 148:656–660.
Pappagianis, D., and B. L. Zimmer. 1990. Serology of coccidioidomycosis. Clinical Microbiology Reviews 3:247–268.
Prabhu, R. M., M. Bonnell, B. L. Currier, and R. Orenstein. 2004. Successful treatment of disseminated nonmeningeal coccidioidomycosis with voriconazole. Clinical Infectious Diseases 39:e74–e77.
Proia, L. A., and A. R. Tenorio. 2004. Successful use of voriconazole for treatment of Coccidioides meningitis. Antimicrobial Agents and Chemotherapy 48:2341.
Reheis, M., and J. Rademaekers. 1997. Predicted dust emission vs. measured dust deposition in the southwestern United States. U.S. Geological Survey: http://geochange.er.usgs.gov/sw/impacts/geology/dust2/ (accessed November 15, 2010).
Shubitz, L. F. 2007. Comparative aspects of coccidioidomycosis in animals and humans. Annals of the New York Academy of Sciences 1111:395–403.
Shubitz, L. F., J. J. Yu, C. Y. Hung, T. N. Kirkland, T. Peng, R. Perrill, J. Simons, J. Xue, R. A. Herr, G. T. Cole, and J. N. Galgiani. 2006. Improved protection of mice against lethal respiratory infection with Coccidioides posadasii using two recombinant antigens expressed as a single protein. Vaccine 24:5904–5911.
Smith, C. E. 1940. Epidemiology of acute coccidioidomycosis with erythema nodosum. American Journal of Public Health 30:600–611.
———. 1958. Coccidioidomycosis. In Communicable diseases transmitted chiefly through respiratory and alimentary tracts. Vol. 4, edited by J. B. Coates and E. C. Hoff. Washington, DC: Office of the Surgeon General, Medical Department, U.S. Army. Pp. 285–316.
Standaert, S. M., W. Schaffner, J. N. Galgiani, R. W. Pinner, L. Kaufman, E. Durry, and R. H. Hutcheson. 1995. Coccidioidomycosis among visitors to a Coccidioides immitis-endemic area: An outbreak in a military reserve unit. Journal of Infectious Diseases 171:1672–1675.
Stern, N. G., and J. N. Galgiani. 2010. Coccidioidomycosis among scholarship athletes and other college students, Arizona, USA. Emerging Infectious Diseases 16:321–323.
Stevens, D. A., A. Rendon, V. Gaona-Flores, A. Catanzaro, G. M. Anstead, L. Pedicone, and J. R. Graybill. 2007. Posaconazole therapy for chronic refractory coccidioidomycosis. Chest 132:952–958.
Talbot, G. H., J. Bradley, J. E. Edwards, Jr., D. Gilbert, M. Scheld, and J. G. Bartlett. 2006. Bad bugs need drugs: An update on the development pipeline from the Antimicrobial Availability Task Force of the Infectious Diseases Society of America. Clinical Infectious Diseases 42:657–668.
Tarcha, E. J., V. Basrur, C. Y. Hung, M. J. Gardner, and G. T. Cole. 2006. Multivalent recombinant protein vaccine against coccidioidomycosis. Infection and Immunity 74:5802–5813.
Tsang, C. A., S. M. Anderson, S. B. Imholte, L. M. Erhart, S. Chen, B. J. Park, C. Christ, K. K. Komatsu, T. Chiller, and R. H. Sunenshine. 2010. Enhanced surveillance of coccidioidomycosis, Arizona, USA, 2007–2008. Emerging Infectious Diseases 16:1738–1744.
Tucker, R. M., D. W. Denning, B. Dupont, and D. A. Stevens. 1990. Itraconazole therapy for chronic coccidioidal meningitis. Annals of Internal Medicine 112:108–112.
Valdivia, L., D. Nix, M. Wright, E. Lindberg, T. Fagan, D. Lieberman, T. Stoffer, N. M. Ampel, and J. N. Galgiani. 2006. Coccidioidomycosis as a common cause of community-acquired pneumonia. Emerging Infectious Diseases 12:958–962.
Werner, S. B., D. Pappagianis, I. Heindl, and A. Mickel. 1972. An epidemic of coccidioidomycosis among archeology students in northern California. New England Journal of Medicine 286:507–512.
Wieden, M. A., L. L. Lundergan, J. Blum, K. L. Delgado, R. Coolbaugh, R. Howard, T. Peng, E. Pugh, N. Reis, J. Theis, and J. N. Galgiani. 1996. Detection of coccidioidal antibodies by 33-kDa spherule antigen, Coccidioides EIA, and standard serologic tests in sera from patients evaluated for coccidioidomycosis. Journal of Infectious Diseases 173:1273–1277.
CRYPTOCOCCUS GATTII: AN EMERGING PATHOGEN IN THE UNITED STATES
Julie R. Harris30
The genus Cryptococcus comprises 37 different species, of which only 2 are relevant for clinical infection (C. gattii and C. neoformans). Cryptococcus spores are inhaled from the environment, causing a primary lung infection that may or may not be symptomatic. Disseminated disease may result in meningitis and death (Li and Mody, 2010). Intricately linked with severe immunosuppression, C. neoformans was rarely reported before the 1950s, when cancer treatments and organ transplants—conditions that often require immunosuppressive treatments—began occurring with increasing frequency (Perfect and Casadevall, 2011). The era of AIDS led to an exponential increase in the
_____________________
30 Centers for Disease Control and Prevention.
numbers of immunosuppressed persons, and a corresponding massive increase in C. neoformans infections worldwide (Mitchell and Perfect, 1995; Perfect and Casadevall, 2011). Today, cryptococcal infections due to C. neoformans are among the most common AIDS-defining infections (Park et al., 2009; Warkentien and Crum-Cianflone, 2010). In contrast, infections caused by C. gattii are reported much less frequently. Gatti and Eeckels produced the first report of C. gattii infection in 1970, from a 7-year-old boy in Congo in 1966 (Gatti and Eeckels, 1970). The boy was found to have a cryptococcal infection clinically similar to C. neoformans, but with a different pathogen morphology (Gatti and Eeckels, 1970). The newly observed pathogen was deemed to be a variant of C. neoformans called C. neoformans var. gattii.
Today, C. gattii is considered its own species (Kwong-Chung et al., 2002). During the mid-1980s, studies of geographic sources of C. gattii and C. neoformans isolates demonstrated that although C. neoformans was found from all areas of the world, C. gattii was found only in tropical and subtropical climatic zones (Kwon-Chung and Bennett, 1984a,b). The authors concluded that C. gattii was likely to be restricted to locations where the minimum winter temperatures typically remained above freezing (Kwon-Chung and Bennett, 1984a). During the next decade, limited information became available about the epidemiology of C. gattii, much of it from papers describing infections in endemic Australia and Papua New Guinea (Ellis, 1987; Mitchell et al., 1995; Seaton et al., 1996b, 1997; Speed and Dunt, 1995). These studies demonstrated that C. gattii, unlike C. neoformans, was found almost exclusively in immunocompetent persons. In agreement with the findings of Kwon-Chung and Bennett (1984a), C. gattii infections were still seen only in patients living in tropical and subtropical climates (Chen et al., 2000; Lalloo et al., 1994; Laurenson et al., 1993, 1996, 1997; Mitchell et al., 1995; Seaton et al., 1996a, 1997; Slobodniuk and Naraqi, 1980; Speed and Dunt, 1995).
Beginning in 1999, the rate of cryptococcal infections among HIV-uninfected persons living on temperate Vancouver Island, British Columbia (B.C.), Canada, began increasing rapidly (Hoang et al., 2004; Fyfe et al., 2002). Early investigations demonstrated that most of these infections were caused by C. gattii, rarely reported before from Canada (Kwon-Chung and Bennett, 1984a). Veterinarians, too, noted that a wide range of animals were becoming infected with C. gattii where they had not previously been found with non-neoformans cryptococcosis (Duncan et al., 2006). During the following years, the disease continued spreading in B.C., infecting humans and animals on the nearby mainland who had no history of travel to Vancouver Island (MacDougall et al., 2007), and by 2007, 218 human infections had been reported from B.C. Although four genetic groups of C. gattii have been identified (VGI, VGII, VGIII, and VGIV), most infections in British Columbia were caused by the relatively uncommon VGII genotype. Due to the high numbers of isolates available from the outbreak, further genetic subdivision of outbreak-associated isolates was performed, demonstrating that
approximately 90 percent of isolates in B.C. were of a “major strain” genetic subtype VGIIa, with a smaller number of “minor strain” subtype VGIIb isolates (Galanis and MacDougall, 2010).
Clinicians began noting C. gattii infections among patients in the Pacific Northwestern (PNW) states of Washington and Oregon in 2004 and 2005, respectively (MacDougall et al., 2007). In addition to the VGIIa and VGIIb infections, a new genotype of infection was noted in the United States, called VGIIc (Byrnes et al., 2010a). Beginning in October 2009, the Pacific Northwest Cryptococcus gattii Public Health Working Group, comprising the U.S. Centers for Disease Control and Prevention (CDC) and state health departments and laboratories in the Pacific Northwest, began retrospectively and prospectively collecting standardized information on C. gattii infections in the United States. As national awareness about the outbreak began to grow, other states also reported rare infections or submitted isolates for speciation and genotyping at the CDC.
Outbreak of C. gattii in the United States
To date, nearly 80 laboratory-confirmed human cases have been reported in the United States (Harris et al., 2010), most from Washington and Oregon (Figure A9-1). As awareness of the outbreak in the PNW has spread throughout the United States, infections from other states have also been reported. Case counts have increased each year, from a single case reported during 2004 to 24 cases reported during 2010 (Figure A9-2). The increase in reported cases is likely to be a result of both improved surveillance, as indicated by the recent increase in cases being reported from areas outside the PNW (Figure A9-2), and actual increases in case occurrences.
Among human patients in the United States identified through surveillance as having laboratory-confirmed C. gattii infection, approximately half are male, with a median age of 56 (range, 15–95). Patients aged 30 and older comprise the majority of cases (Table A9-1). The most commonly reported symptoms are headache, nausea, and cough, affecting more than half of all patients; more than half present with pneumonia and approximately half have meningitis. Most (73 percent) patients have an underlying immunosuppressive condition, including (but not limited to) a recent history of oral steroid use; lung, heart, kidney, or liver disease; a history of cancer; or a solid organ transplant (Table A9-2). HIV infection was more frequent among C. gattii patients than is found in the general U.S. population (5.9 percent vs. 0.6 percent) (CIA World Factbook, 2010), but was still the least commonly reported underlying condition. Of 59 patients with data, 17 (29 percent) had no detectable underlying immunocompromising condition. Cryptococcomas, or fungal masses, were found in the lungs and/or brains of substantial proportions of patients who received the corresponding scans or x-rays. Nearly all patients were hospitalized, and nearly one third with follow-up information died with or from their infections (Table A9-1).
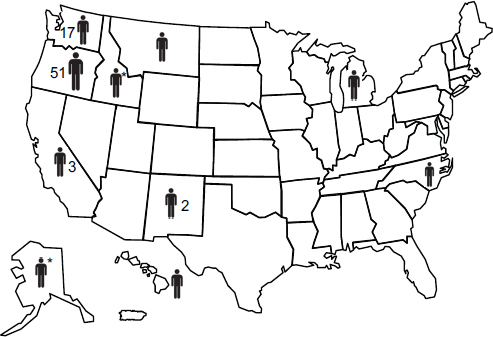
Figure A9-1 Human infections with C. gattii, United States, December 2004–January 2011 (n = 79).
NOTE: * indicates patients that reported extensive travel to Washington and/or Oregon during the year before their illness onsets.
Outbreak-Strain vs. Non-Outbreak-Strain C. gattii Infections in the United States
Currently, 77 of 79 reported U.S. infections have been genotyped. Of these 77, 38 (49 percent) were VGIIa; 19 (25 percent) were VGIIc; 6 (8 percent) were VGIIb; and 14 (18 percent) were other genotypes (VGI, VGIII, and unrelated VGII).
Clear delineations exist between C. gattii infections in Oregon and Washington (“PNW-associated infections”) and those occurring in other parts of the United States (Figure A9-2). In particular, most PNW-associated infections are genotype VGIIa, VGIIb, or VGIIc (“outbreak-strain” genotypes). Outbreak-strain infections have also been reported from humans living in states outside of Washington and Oregon; those that have been reported were linked to these states by residential or travel history (Harris et al., 2010). One case in Idaho (subtype VGIIc) (CDC, 2010; Iqbal et al., 2010) and one case from Alaska (subtype VGIIa) (Harris et al., 2010) are considered linked to the outbreak, with both patients reporting extensive travel throughout Oregon, Washington, and/or B.C. during the year before their illness onsets (CDC, 2010).
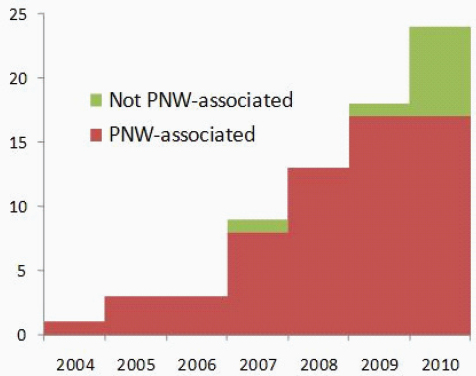
Figure A9-2 U.S. human cases of C. gattii, by year of illness onset (n = 71*).
* NOTE: Onset year is reported for 62 patients and is estimated by initial report year for 9 patients for whom onset date was not available. 2010 data are current as of January 2011; complete case data typically lag illness onset by several months. PNW = Pacific Northwest.
While a small number of non-outbreak-strain infections have occurred in the PNW, most have been reported from states outside the PNW. A patient from North Carolina was diagnosed with C. gattii (genotype VGI) in 2007 following travel to San Francisco (Byrnes et al., 2009), and two cases of C. gattii (genotypes VGI and VGIII) were reported in 2010 from patients in New Mexico who did not have any recent history of travel outside of the state (Harris et al., 2010). In 2010, a patient was reported from Hawaii with a novel subtype of infection belonging to the genotype VGII (Harris et al., 2010). Since 2004, two patients have been reported from California and one from Michigan with VGIII-type infections. Thus, at least one focus of infection involving genotypes novel in the United States appears to be ongoing in the PNW; it also appears that sporadic C. gattii infection is occurring elsewhere, with genotypes distinct from those found in the PNW.
The collection of standardized data on U.S. outbreak-strain and non-outbreak-strain infections has provided an opportunity to examine clinical
Table A9-1 Characteristics of C. gattii Patients in the United States, 2004–2010
| Characteristic | n | N | % |
| Age (mean, median, range), years | 52, 55 (15–95) | ||
| . 18 | 3 | 64 | 5 |
| 19–29 | 3 | 64 | 5 |
| 30–49 | 23 | 64 | 36 |
| 50–69 | 27 | 64 | 42 |
| 70+ | 8 | 64 | 13 |
| Symptoms | |||
| Headache | 32 | 55 | 58 |
| Nausea | 26 | 50 | 52 |
| Cough | 29 | 57 | 51 |
| Dyspnea | 24 | 51 | 47 |
| Fever | 25 | 57 | 44 |
| Vomiting | 22 | 50 | 44 |
| Fatigue | 17 | 40 | 43 |
| Weight loss | 21 | 50 | 42 |
| Loss of appetite | 15 | 46 | 33 |
| Chills | 16 | 52 | 31 |
| Muscle pain | 15 | 52 | 29 |
| Chest pain | 14 | 52 | 27 |
| Neck stiffness | 11 | 49 | 22 |
| Night sweats | 6 | 44 | 14 |
| Photophobia | 5 | 49 | 10 |
| Seizure | 4 | 45 | 9 |
| Blurry vision | 2 | 45 | 4 |
| Any respiratory | 39 | 55 | 71 |
| Any central nervous system | 19 | 44 | 43 |
| Clinical findings | |||
| Pneumonia | 27 | 49 | 55 |
| Meningitis | 26 | 51 | 51 |
| Crypto lung | 18 | 50 | 36 |
| Crypto head | 6 | 22 | 27 |
| Outcomes of infection | |||
| Hospitalized | 52 | 57 | 91 |
| Died | 16 | 52 | 31 |
differences by genotype and patient type. Although a relatively small number of non-outbreak-strain infections have been reported, significant differences between these and outbreak-strain infections have been noted in the proportion of infected patients who were immunocompromised and the propensity for respiratory and CNS symptoms (Table A9-2). It seems likely that different genotypes of C. gattii might infect different patient types; in addition, clinical manifestation of C. gattii infection might differ either by infecting strain genotype, patient immune status, or both.
| Symptoms, Patient Characteristics, Outcomes | VGIIa/b/c | Other genotypes | RR | p |
| Respiratory symptoms | ||||
| Cough | 58% | 11% | 5.3 | 0.01 |
| Dyspnea | 53% | 12% | 4.3 | 0.05 |
| Any respiratory symptom | 76% | 44% | 1.7 | 0.10 |
| Central nervous system symptoms | ||||
| Seizure | 3% | 50% | 0.05 | 0.01 |
| Blurry vision | 8% | 43% | 0.19 | 0.04 |
| Any central nervous system symptom | 34% | 100% | 0.04 | 0.00 |
| Preexisting medical condition | 78% | 33% | 7.1 | 0.01 |
| Hospitalized | 90% | 100% | 0.9 | 1.00 |
| Died from or with C. gattii | 32% | 16% | 2 | 0.65 |
NOTES: p: Fisher’s exact p-value; RR: Relative risk
Historical Isolates of C. gattii in the United States
In spite of extensive press coverage in the United States during 2010 about the “new deadly fungus” in the United States and British Columbia (Discover Magazine, 2010; Hutchison, 2010; Park, 2010), C. gattii infections are not entirely novel in the United States: in particular, the infection repeatedly has been found in California in the past. In a 1984 study, C. gattii was found in 46 of 315 clinical isolates of Cryptococcus from the United States; 30 (65 percent) of the 46 C. gattii isolates were from patients residing in Southern California. Among all 71 Cryptococcus isolates from Southern California, C. gattii represented 42 percent, compared with 6 percent in the rest of the continental United States (Kwon-Chung and Bennett, 1984b). The same group reported in a different paper that both of the two clinical Cryptococcus isolates from Hawaii were C. gattii (Kwon-Chung and Bennett, 1984a). Another study, which examined cryptococcal isolates from HIV-infected patients from Los Angeles in the early 1990s, found C. gattii in 12 percent of these isolates (Chaturvedi et al., 2005). Genotyping of the isolates showed that 28 of 30 isolates were of subtype VGIII; 1 was VGI, and 1 was of subtype VGII (Byrnes et al., 2010b).31C. gattii was also found in three of 358 genotyped isolates collected from around the United States during surveillance from 1992 to 1994; two isolates were from San Francisco (VGI and VGIII) and one was from Atlanta (VGI) (Brandt et al., 1996). Finally, two isolates of genotype VGIIa, indistinguishable by multilocus sequence typing (MLST) from the major strain in Vancouver Island, were collected from the sputum of a Seattle man in the 1970s (Diaz et al., 2000) and from a eucalyptus tree in San
_____________________
31 Also see contributed manuscript by Heitman in Appendix A (pages 226–248).
Francisco in 1992 (Fraser et al., 2005). Taken together, these data suggest that Southern California, but probably not the rest of the country, has long been an endemic area for C. gattii. In addition, sporadic infections appear to be occurring from other states, although the travel history of these patients and their potential exposure site is unknown. Recently, environmental isolates of C. gattii have also been found in Puerto Rico from a variety of cacti and tree material (Loperena-Alvarez et al., 2010).
C. gattii in the PNW vs. C. gattii in Other Areas of the World
Globally, the most common infecting strains of C. gattii and the immune status of patients infected appear to vary. In endemic Australia and Papua New Guinea, C. gattii (usually type VGI) appears to occur primarily in apparently immunocompetent patients and cause CNS disease (Seaton et al., 1996a,b; Speed and Dunt, 1995). However, in several African countries, VGIV-type infections are most frequent, and are reported exclusively from HIV-infected patients (Litvintseva et al., 2005; Steele et al., 2010). By contrast, in Venezuela (Villanueva et al., 1989), Brazil (Santos et al., 2008), Paraguay, Argentina (Castanon-Olivares et al., 2000; Kwon-Chung and Bennett, 1984a,b), Peru (Bustamante Rufino and Swinne, 1998; Kwon-Chung and Bennett, 1984a,b), and Colombia (Escandon et al., 2006), infections (most commonly genotype VGII, although different from the PNW outbreak strains) occur in both HIV-uninfected and HIV-infected patients. In Mexico, all four C. gattii genotypes have been reported (Olivares et al., 2009), primarily from HIV-uninfected persons (Castanon-Olivares et al., 2000; Lopez-Martinez et al., 1996). In Asia, C. gattii (primarily VGI and VGII) has been reported from Vietnam, Cambodia, Thailand, Korea, Japan, Malaysia, India, China, and Nepal (Chen et al., 2008; Choi et al., 2010; Kwon-Chung and Bennett, 1984a,b; Lui et al., 2006; Okamoto et al., 2010), in both immunocompetent and immunocompromised persons (Chau et al., 2010; Ngamskulrungroj et al., 2010).
The outbreak of C. gattii in B.C. and the PNW is qualitatively different in genotype, presentation, disease course, and outcome than has been seen with C. gattii in other locations. In contrast to findings from areas outside of North America, C. gattii patients in both B.C. and the PNW most commonly present with respiratory rather than CNS symptoms (Galanis and MacDougall, 2010; Harris et al., 2010). In addition, a high proportion of infections in both the PNW and B.C. occur in immunocompromised (but usually HIV-uninfected) patients. In B.C., 38 percent of patients are immunocompromised (Galanis and MacDougall, 2010); using the same case definition for immunocompromised state, in the United States, approximately 59 percent of patients are immunocompromised (CDC, unpublished data).
Interestingly, differences also might exist between C. gattii patients in the United States and in B.C. U.S. patients were younger, more likely to be hospital-
ized, and more likely to die from or with their infection than were B.C. patients (Harris et al., 2010). The reasons for these differences are unclear, but might include differences in case ascertainment in the PNW compared with B.C., which has led to capture of a lower proportion of non-hospitalized C. gattii patients in the PNW. Alternately, they could relate to the different genotypes seen in the PNW compared with B.C. A comprehensive review of patient medical charts is under way. It might help elucidate the differences and provide some insight into whether these differences are real, and if so, why they exist.
The differences in U.S. C. gattii infections in the PNW, U.S. infections outside the PNW, and infections occurring in other areas of the world might be a function of C. gattii subtype, tropism, environmental distribution, or surveillance bias. That is, C. gattii infections might not be reported completely from other areas because they have not been looked for systematically elsewhere. Regardless of the reasons for the outbreak in North America, reports of the outbreaks do not appear to be exclusively due to temporal changes in surveillance. Retrospective speciation studies of isolates from B.C. before 1999 (Fyfe et al., 2008) and from the Seattle area before 2004 (Upton et al., 2007) suggested that the increase in reported cases represents a true increase in infections in the region. It remains to be seen whether the U.S. C. gattii infections outside of the PNW are sporadic, travel associated, or linked to part of a larger emerging health issue in other areas of the United States. Below is a discussion about efforts to conduct surveillance for C. gattii infections outside of the PNW.
Surveillance for Human C. gattii Infections Outside the PNW
To address the question of how frequently C. gattii infections are occurring outside of the PNW, in October 2010 the CDC issued an alert through ClinMicroNet (Dwyer, 2003), a listserv sent to directors of U.S. clinical microbiology laboratories. The alert described the ongoing outbreak in the Pacific Northwest, and requested that any unspeciated Cryptococcus isolates (or those already speciated as C. gattii) be sent to the Mycotic Diseases Branch laboratory at the CDC, with isolation date, source city, and source state. Isolates were speciated at CDC and C. gattii isolates were genotyped. As of January 2011, 32 isolates, isolated between 2006 and 2010, from patients in seven states had been submitted to the CDC. Of these, 10 were C. gattii; half were from California (Table A9-3). Collection of isolates is ongoing and cases continue to be reported to the CDC, albeit rarely, from states outside of Washington and Oregon. The recent commercial availability of specialized culture medium (CGB medium) that enables rapid discrimination between C. neoformans and C. gattii will hopefully facilitate this process (Butler-Wu and Limaye, 2011).
| State | C. neoformans |
C. neoformans |
Total isolates |
| AL | 3 | 0 | 3 |
| CA | 1 | 5 | 6 |
| GA | 1 | 2 | 3 |
| HI | 0 | 1 | 1 |
| IL | 14 | 0 | 14 |
| OR | 0 | 1 | 1 |
| TX | 3 | 0 | 3 |
| UT | 0 | 1 | 1 |
| Total | 22 | 10 | 32 |
Clinical Aspects of C. gattii Infection and Differences from C. neoformans
Several attempts have been made to carry out head-to-head comparisons of clinical disease caused by C. gattii and C. neoformans. In 1995, Speed and Dunt (1995) published a paper comparing clinical symptoms among hospitalized C. gattii and C. neoformans patients. In addition to noting that fewer than 10 percent of C. neoformans infections in Australia occurred in otherwise healthy patients while 100 percent of C. gattii infections occurred in healthy patients, the authors also reported that C. gattii more frequently involved cerebral and meningeal sites, had neurologic sequelae, required CNS or thoracic surgery to resect cryptococcomas, and required longer periods of treatment (Speed and Dunt, 1995). In addition, C. gattii infections more frequently resulted in relapse than C. neoformans infections. The authors commented that the nearly three-fold longer therapy times required for patients with C. gattii compared with patients with C. neoformans infections stemmed from the difficulty in reducing the size of the cryptococcomas and the inability to rapidly control infection in patients with C. gattii. However, they also noted that both bloodstream infections and mortality were exclusively limited to patients with C. neoformans (Speed and Dunt, 1995).
The same year, Mitchell et al. (1995) published a report directly comparing the two cryptococcal species in immunocompetent patients with cerebral disease (Mitchell et al., 1995). Similar to the findings by Speed and Dunt (1995) and later by Chen et al. (2000), the authors reported that both lung and brain cryptococcomas were more common among patients with C. gattii than C. neoformans infections. In addition, they reported that immunocompetent patients with C. gattii infection were significantly more likely to have a “poor outcome”—defined as moderate to major sequelae or death—than immunocompetent patients with
C. neoformans infections. When they compared outcomes among patients with meningitis but normal brain imaging at initial presentation, they found no differences by infecting species; however, they did find that C. gattii patients presenting with mass lesions on initial intracranial scan were more likely to have poor outcomes than those who did not (Mitchell et al., 1995). Taken together, these data suggested that the epidemiology of C. neoformans and C. gattii in Australia was different even when controlling for patient immune status.
However, a similar paper, published in 2010, compared infection with C. gattii and C. neoformans in immunocompetent patients in Vietnam (Chau et al., 2010) and failed to demonstrate differences in clinical phenotype by infecting species. The authors of this report, in contrast to Mitchell et al. (1995), suggested that host immune status was more influential on clinical course and outcome than was infecting species. This was also a conclusion drawn by Chen et al. (2000), who compared cryptococcal infections among both immunocompetent and immunosuppressed patients in Australia and New Zealand and demonstrated that immunocompetent hosts were more likely to present with lung infections, species type notwithstanding, than were immunocompromised hosts. Chen et al. (2000) also noted that C. gattii infections were more likely to occur in the brain than C. neoformans infections, but that cryptococcomas were associated both with infection with C. gattii and with immunocompetent status.
Lui et al. (2006) also compared cryptococcal infection in immunocompetent and immunosuppressed patients in China, noting elevated proportions of immunocompetent patients presenting with meningitis compared with immunocompromised patients, more intense inflammatory responses, and a lower risk of death. Although the paper also indicated that immune status might influence clinical course more than infecting species, the small numbers of C. gattii infections made evaluation of the effect of infecting species difficult.
Only two studies have directly compared C. gattii infections to C. neoformans infections exclusively in AIDS patients. In Botswana, Steele et al. (2010) compared C. gattii and C. neoformans infections in AIDS patients with cryptococcal meningitis, and found few differences in terms of clinical presentation or in-hospital mortality (Steele et al., 2010). Morgan et al. (2006) found similar results among South African patients with cryptococcal meningitis. It is possible that, among severely immunosuppressed patients who have a low capacity to respond immunologically to infection, such as late-stage AIDS patients, the infecting cryptococcal species is irrelevant to outcomes, while infecting species have a stronger influence when patients are mildly to moderately immunosuppressed or not immunosuppressed. These studies did not carry out extensive brain or thoracic imaging to evaluate the presence of cryptococcomas.
The poor prognosis of immunocompetent patients infected with C. neoformans has been studied previously, and has been suggested to be due to the delay in diagnosis, inappropriate treatment, and potentially the presence of an intact immune system that might provoke an immune reconstitution inflammatory
syndrome (IRIS) (Ecevit et al., 2006; Lui et al., 2006). IRIS is a paradoxical clinical deterioration that is well documented during treatment of cryptococcosis following initiation of antiretroviral therapy in AIDS patients (Woods et al., 1998) and is thought to be due to an overzealous “rebound” immune response in the presence of significant amounts of infecting pathogen. An IRIS-like syndrome has also been documented in patients infected with C. gattii, where the syndrome was suggested to be due to concomitant immune rebound and decreases in IL-10 (Einsiedel et al., 2004). These same factors may contribute to the severity of C. gattii infection in immunocompetent hosts. At least one report exists of a patient whose condition improved with steroid treatment, suggesting that an overly functional immune system could confound treatment efforts in some patients (Lane et al., 2004).
Discussion
The described outbreaks of C. gattii infection in the temperate climates of B.C. and the PNW demonstrate a much less restrictive geographic range for C. gattii than previously thought, and a broader range of persons who are susceptible to infection. In particular, a compromised immune status now appears to be a significant risk factor for at least some subtypes of C. gattii infection (CDC, 2010; Galanis and MacDougall, 2010). In addition, data from patients associated with these outbreaks suggest that different C. gattii genotypes might infect different types of patients, and/or demonstrate different clinical courses resulting from infection. The mere existence of an outbreak associated with C. gattii, never previously reported, suggests that genetic components might be important for pathogen spread in ways that are still poorly understood. More than ever, collecting data is important that disease recognition and optimal treatment of C. gatiii infections can be investigated. Several existing challenges now face the field.
Existing Challenges
One existing challenge is diagnosis of infection. Although several methods exist to identify cryptococcal infections, including culture, India Ink stains of cerebrospinal fluid or sputum (Cohen, 1984), and commercially available cryptococcal antigen (CrAg) test kits (Saha et al., 2008), these methods cannot distinguish between C. neoformans and C. gattii. A simple way to confirm whether or not a cryptococcal isolate is species gattii is to plate the isolate on canavanineglycine bromothymol blue (CGB) agar (Klein et al., 2009; Kwon-Chung et al., 1982), where C. neoformans will leave the medium unaffected in color (yellow to green) due to a failure to grow, and C. gattii will turn the medium blue due to use of glycine as a carbon source. This medium is currently available from at least one commercial supplier, but is not widely used in U.S. clinical microbiology labs. Thus, many C. gattii infections likely are being misdiagnosed as
C. neoformans. Ensuring that clinicians are aware of C. gattii infection and the possible need for clinical differentiation from C. neoformans, and that their reference laboratories are able to speciate Cryptococcus isolates (and have an interest in doing so), is critical to evaluate fully the geographic spread of disease and the clinical spectrum of infections.
Investigating whether or not the most recent findings warrant modified treatment guidelines is an additional challenge. The Infectious Diseases Society of America published guidelines in 2010 (Perfect et al., 2010) that refer to differences in the treatment of C. gattii infections, compared with C. neoformans infections: specifically, C. gattii infections might require lengthier, more aggressive treatment when compared with C. neoformans infections. The increased propensity for C. gattii to form cryptococcomas is also noted (Perfect et al., 2010). However, the guidelines were largely based on data from C. gattii infections occurring in Australia and Papua New Guinea. Increasingly, our data suggest that even among C. gattii infections, not all cryptococcal infections are alike. However, it is unclear which factors—infecting species, infecting subtype, host immune status, or perhaps even host genetics—are most influential on patient presentation and infection. Data from rigorous clinical studies are of utmost importance in ensuring that clinician guidelines provide sufficient guidance to optimize patient care. To this end, a large-scale, longitudinal chart review of C. gattii infections is ongoing as a collaborative effort among Australia, B.C., and the United States, designed to address some of these questions. Results are expected sometime in 2012.
Finally, the development of prevention messages is a challenge. Unlike C. neoformans, which grows in pigeon feces, C. gattii appears to live in association with trees and soil surrounding them (Springer and Chaturvedi, 2010). The tree type appears to be less important than the presence of a wood substrate for growth, and C. gattii has to date been associated with more than 50 tree species (Randhawa et al., 2001; Springer and Chaturvedi, 2010). It has also been found in air and water samples. These findings notwithstanding, C. gattii has not been found ubiquitously around the globe in a distribution similar to C. neoformans, and thus we can postulate that at least some environmental restrictions remain in place for this organism. Environmental organisms present a specific challenge for public health prevention because infections are usually relatively rare, and difficult to avoid without draconian measures (e.g., staying indoors and purifying air). This represents a quandary for public health officials. It remains to be seen whether “hot spots” of infection exist in the PNW for which generalized recommendations can be made that would benefit patient health, perhaps for subgroups of higher risk patients. The benefits of outdoor activity would need to be weighed against any risk calculated for these patients, and such recommendations are bound to be controversial.
Where Did C. gattii Come From, and Where Is It Going?
How did C. gattii arrive in the temperate areas of North America? Kidd et al. (2007) demonstrated, during an environmental sampling study for C. gattii in B.C., that anthropogenic activities could spread the pathogen, showing the presence of C. gattii on the shoes of human samplers and wheel wells of sampling vehicles as they traveled from one sampling site to the next. In addition, they showed that the pathogen was present in highly trafficked areas of Vancouver Island; that it could be found in the air, in freshwater, and in seawater around the area; and that the spores could survive for more than a year in many of these media. Thus, it is not difficult to hypothesize several methods by which the pathogen could have found itself in temperate B.C., and easier still to imagine mechanisms by which it could be transported into the United States. However, the question of whether new, cold-tolerant strains of C. gattii were brought to B.C. or whether they were formed there, through recombination of two or more preexisting or “seeded” strains, is still unresolved. It is also possible that the pathogen was always tolerant of temperate-weather climates, but existed in caches too small in these regions to sustain human infection or to establish permanent habitats; repeated seeding of these regions through global materials trafficking (e.g., wood or trees) could have created a sustainable niche for the pathogen. Alternately, changes in the global climate could be facilitating the optimal habitat development and spread of C. gattii, providing minimum conditions under which the pathogen can successfully propagate. Whole-genome sequencing is currently being carried out with C. gattii isolates obtained from patients associated with this outbreak, as well as more historical isolates (Lockhart et al., 2010), which could shed some light on the origin of the current outbreak and provide ideas for where it might move next.
Conclusion
The increase in the number of reports during the past decade related to the occurrence of C. gattii infections outside of traditional endemic tropical and subtropical regions has provided excellent opportunities to learn more about this important pathogen. The differences between individual cryptococcal infections appear to be linked not only to patient immune status and infecting species, but also to genetic subtypes within a species. It is unclear if the species and subtypes have preferences for infection among certain patient types, possibly due to a need for host immune support (or lack thereof) for replication, or if differences in environmental colonization patterns might influence the type of patient infected. For example, a pathogen with a ubiquitous distribution and a preference for immunocompromised patients will have a much higher infection rate and a much higher immunocompromised to immunocompetent patient ratio than would a pathogen with “hot spot distribution,” which would infect fewer patients overall, but be limited to patients living in its area of environmental distribution (most of
whom are immunocompetent). Thus, the immunocompromised to immunocompetent ratio might be nothing more than a function of the degree of distribution of a Cryptococcus species in the environment and the types of patients living in its area of distribution. Determining what governs the range of distribution of C. gattii—and understanding when it reaches a stable environmental equilibrium in new areas of emergence—is critical for understanding this relationship.
In spite of the recent flood of reports about C. gattii, much remains unknown. The epidemiologic curve has not yet stabilized in the United States, and the trajectory of future infections is unknown. The lack of comprehensive surveillance, both within North America and without, and the genetic variety inherent in C. gattii, has limited our current understanding of pathogen spread and pathogenesis. The conditions that favor pathogen colonization and propagation are not known. Evidence shows that infections in the United States and particularly the PNW are qualitatively different from those occurring elsewhere, but it is unclear whether or not these differences warrant modifications to existing treatment guidelines. Continued collection of robust surveillance data will assist in answering some of these questions. The coming years should see increasing amounts of information on C. gattii infections globally, which should shed light on genotype- and subtype-specific differences among C. gattii infections.
References
Brandt, M. E., L. C. Hutwagner, L. A. Klug, W. S. Baughman, D. Rimland, E. A. Graviss, R. J. Hamill, C. Thomas, P. G. Pappas, A. L. Reingold, and R. W. Pinner. 1996. Molecular subtype distribution of Cryptococcus neoformans in four areas of the United States. Cryptococcal Disease Active Surveillance Group. Journal of Clinical Microbiology 34(4):912–917.
Bustamante Rufino, B., and D. Swinne. 1998. [Cryptococcus neoformans var. gattii isolates from two Peruvian patients.]. Revista Iberoamericana de Micología 15(1):22–24.
Butler-Wu, S. M., and A. P. Limaye. 2011. A quick guide to the significance and laboratory identification of Cryptococcus gattii. American Society of Microbiology. http://www.asm.org/asm/images/pdf/Clinical/cgattii.pdf (accessed March 28, 2011).
Byrnes, E. J., III, W. Li, Y. Lewit, J. R. Perfect, D. A. Carter, G. M. Cox, and J. Heitman. 2009. First reported case of Cryptococcus gattii in the Southeastern USA: Implications for travel-associated acquisition of an emerging pathogen. PLoS One 4(6):e5851.
Byrnes, E. J., III, W. Li, Y. Lewit, H. Ma, K. Voelz, P. Ren, D. A. Carter, V. Chaturvedi, R. J. Bildfell, R. C. May, and J. Heitman. 2010a. Emergence and pathogenicity of highly virulent Cryptococcus gattii genotypes in the northwest United States. PLoS Pathogens 6(4):e1000850.
———. 2010b. Examination of Cryptococcus gattii isolates from HIV/AIDS patients uncovers a diverse population of VGIII molecular type isolates endemic in Southern California. Paper presented at Meeting of the Infectious Diseases Society of America, Vancouver, Canada, October 24, 2010.
Castanon-Olivares, L. R., R. Arreguin-Espinosa, G. Ruiz-Palacios y Santos, and R. Lopez-Martinez. 2000. Frequency of Cryptococcus species and varieties in Mexico and their comparison with some Latin American countries. Revista Latinoamericana de Microbiologia 42(1):35–40.
CDC (Centers for Disease Control and Prevention). 2010. Emergence of Cryptococcus gattii—Pacific Northwest, 2004–2010. Morbidity and Mortality Weekly Report 59(28):865–868.
Chaturvedi, S., M. Dyavaiah, R. A. Larsen, and V. Chaturvedi. 2005. Cryptococcus gattii in AIDS patients, southern California. Emerging Infectious Diseases 11(11):1686–1692.
Chau, T. T., N. H. Mai, N. H. Phu, H. D. Nghia, L. V. Chuong, D. X. Sinh, V. A. Duong, P. T. Diep, J. I. Campbell, S. Baker, T. T. Hien, D. G. Lalloo, J. J. Farrar, and J. N. Day. 2010. A prospective descriptive study of cryptococcal meningitis in HIV uninfected patients in Vietnam—high prevalence of Cryptococcus neoformans var grubii in the absence of underlying disease. BMC Infectious Diseases 10:199.
Chen, J., A. Varma, M. R. Diaz, A. P. Litvintseva, K. K. Wollenberg, and K. J. Kwon-Chung. 2008. Cryptococcus neoformans strains and infection in apparently immunocompetent patients, China. Emerging Infectious Diseases 14(5):755–762.
Chen, S., T. Sorrell, G. Nimmo, B. Speed, B. Currie, D. Ellis, D. Marriott, T. Pfeiffer, D. Parr, and K. Byth. 2000. Epidemiology and host- and variety-dependent characteristics of infection due to Cryptococcus neoformans in Australia and New Zealand. Australasian Cryptococcal Study Group. Clinical Infectious Diseases 31(2):499–508.
Choi, Y. H., P. Ngamskulrungroj, A. Varma, E. Sionov, S. M. Hwang, F. Carriconde, W. Meyer, A. P. Litvintseva, W. G. Lee, J. H. Shin, E. C. Kim, K. W. Lee, T. Y. Choi, Y. S. Lee, and K. J. Kwon-Chung. 2010. Prevalence of the VNIc genotype of Cryptococcus neoformans in non-HIV-associated cryptococcosis in the Republic of Korea. FEMS Yeast Research 10(6):769–778.
CIA (Central Intelligence Agency). 2010. The world factbook: United States. https://www.cia.gov/library/publications/the-world-factbook/geos/us.html (accessed October 31, 2010).
Cohen, J. 1984. Comparison of the sensitivity of three methods for the rapid identification of Cryptococcus neoformans. Journal of Clinical Pathology 37(3):332–334.
Diaz, M. R., T. Boekhout, B. Theelen, and J. W. Fell. 2000. Molecular sequence analyses of the intergenic spacer (IGS) associated with rDNA of the two varieties of the pathogenic yeast, Cryptococcus neoformans. Systematic and Applied Microbiology 23(4):535–545.
Discover Magazine. 2010. A tropical, fatal fungus gains a foothold in the Pacific Northwest. http://blogs.discovermagazine.com/80beats/2010/04/23/a-tropical-fatal-fungus-gains-a-foothold-in-the-Pacific-Northwest/ (accessed October 25, 2010).
Duncan, C., H. Schwantje, C. Stephen, J. Campbell, and K. Bartlett. 2006. Cryptococcus gattii in wildlife of Vancouver Island, British Columbia, Canada. Journal of Wildlife Diseases 42(1):175–178.
Dwyer, V. 2003. ClinMicroNet—Sharing experiences and building knowledge virtually. Clinical Microbiology Newsletter 25(16):121–125.
Ecevit, I. Z., C. J. Clancy, I. M. Schmalfuss, and M. H. Nguyen. 2006. The poor prognosis of central nervous system cryptococcosis among nonimmunosuppressed patients: A call for better disease recognition and evaluation of adjuncts to antifungal therapy. Clinical Infectious Diseases 42(10):1443–1447.
Einsiedel, L., D. L. Gordon, and J. R. Dyer. 2004. Paradoxical inflammatory reaction during treatment of Cryptococcus neoformans var. gattii meningitis in an HIV-seronegative woman. Clinical Infectious Diseases 39(8):e78–e82.
Ellis, D. H. 1987. Cryptococcus neoformans var gattii in Australia. Journal of Clinical Microbiology 25(2):430–431.
Escandon, P., A. Sanchez, M. Martinez, W. Meyer, and E. Castaneda. 2006. Molecular epidemiology of clinical and environmental isolates of the Cryptococcus neoformans species complex reveals a high genetic diversity and the presence of the molecular type VGII mating type a in Colombia. FEMS Yeast Research 6(4):625–635.
Fraser, J. A., S. S. Giles, E. C. Wenink, S. G. Geunes-Boyer, J. R. Wright, S. Diezmann, A. Allen, J. E. Stajich, F. S. Dietrich, J. R. Perfect, and J. Heitman. 2005. Same-sex mating and the origin of the Vancouver Island Cryptococcus gattii outbreak. Nature 437(7063):1360–1364.
Fyfe, M., W. Black, M. Romney, et al. 2002. Unprecedented outbreak of Cryptococcus neoformans var. gattii infections in British Columbia, Canada. Paper presented at the Fifth International Conference on Cryptococcus and Cryptococcosis, Adelaide, Australia, March 3–7.
Fyfe, M., L. MacDougall, M. Romney, M. Starr, M. Pearce, S. Mak, S. Mithani, and P. Kibsey. 2008. Cryptococcus gattii infections on Vancouver Island, British Columbia, Canada: Emergence of a tropical fungus in a temperate environment. Canada Communicable Disease Report 34(6):1–12.
Galanis, E., and L. MacDougall. 2010. Epidemiology of Cryptococcus gattii, British Columbia, Canada, 1999–2007. Emerging Infectious Diseases 16(2):251–257.
Gatti, F., and R. Eeckels. 1970. An atypical strain of Cryptococcus neoformans (San Felice) Vuillemin 1894. Description of the disease and of the strain. Annales des Sociétés Belges de Médecine Tropicale, de Parasitologie, et de Mycologie 50(6):689–693.
Harris, J., S. R. Lockhart, N. Marsden-Haug, R. Wohrle, C. Free, E. DeBess, and T. Chiller. 2010. Poster 642. Cryptococcus gattii: Emergence of a novel pathogen in the United States Pacific Northwest. Paper presented at the Infectious Diseases Society of America Meeting, Vancouver, Canada, October 20–24, 2010.
Hoang, L. M., J. A. Maguire, P. Doyle, M. Fyfe, and D. L. Roscoe. 2004. Cryptococcus neoformans infections at Vancouver Hospital and Health Sciences Centre (1997–2002): Epidemiology, microbiology and histopathology. Journal of Medical Microbiology 53(Pt 9):935–940.
Hutchison, C. 2010. Fatal fungus Cryptococcus gattii: Experts say fears overblown. http://abcnews.go.com/Health/Wellness/fatal-fungus-sparks-fear-worry/story?id=10438475 (accessed October 5, 2010).
Iqbal, N., E. E. DeBess, R. Wohrle, B. Sun, R. J. Nett, A. M. Ahlquist, T. Chiller, and S. R. Lockhart. 2010. Correlation of genotype and in vitro susceptibilities of Cryptococcus gattii strains from the Pacific Northwest of the United States. Journal of Clinical Microbiology 48(2):539–544.
Kidd, S. E., P. J. Bach, A. O. Hingston, S. Mak, Y. Chow, L. MacDougall, J. W. Kronstad, and K. H. Bartlett. 2007. Cryptococcus gattii dispersal mechanisms, British Columbia, Canada. Emerging Infectious Diseases 13(1):51–57.
Klein, K. R., L. Hall, S. M. Deml, J. M. Rysavy, S. L. Wohlfiel, and N. L. Wengenack. 2009. Identification of Cryptococcus gattii by use of L-canavanine glycine bromothymol blue medium and DNA sequencing. Journal of Clinical Microbiology 47(11):3669–3672.
Kwon-Chung, K. J., and J. E. Bennett. 1984a. Epidemiologic differences between the two varieties of Cryptococcus neoformans. American Journal of Epidemiology 120(1):123–130.
———. 1984b. High prevalence of Cryptococcus neoformans var. gattii in tropical and subtropical regions. Zentralblatt Fuer Bakteriologie,Microbiologie, und Hygiene (Reihe A) 257(2): 213–218.
Kwon-Chung, K. J., I. Polacheck, and J. E. Bennett. 1982. Improved diagnostic medium for separation of Cryptococcus neoformans var. neoformans (serotypes A and D) and Cryptococcus neoformans var. gattii (serotypes B and C). Journal of Clinical Microbiology 15(3):535–537.
Kwong-Chung, K., T. Boekhout, J. W. Fell, and M. Diaz. 2002. Proposal to conserve the name Cryptococcus gattii against C. hondurianus and C. bacillisporus (Basidiomycota, Hymenomycetes, Tremellomycetidae). Taxon 51:804–806.
Lalloo, D., D. Fisher, S. Naraqi, I. Laurenson, P. Temu, A. Sinha, A. Saweri, and B. Mavo. 1994. Cryptococcal meningitis (C. neoformans var. gattii) leading to blindness in previously healthy Melanesian adults in Papua New Guinea. Quarterly Journal of Medicine 87(6):343–349.
Lane, M., J. McBride, and J. Archer. 2004. Steroid responsive late deterioration in Cryptococcus neoformans variety gattii meningitis. Neurology 63(4):713–714.
Laurenson, I., S. Naraqi, N. Howcroft, I. Burrows, and S. Saulei. 1993. Cryptococcal meningitis in Papua New Guinea: Ecology and the role of eucalypts. Medical Journal of Australia 158 (3):213.
Laurenson, I. F., A. J. Trevett, D. G. Lalloo, N. Nwokolo, S. Naraqi, J. Black, N. Tefurani, A. Saweri, B. Mavo, J. Igo, and D. A. Warrell. 1996. Meningitis caused by Cryptococcus neoformans var. gattii and var. neoformans in Papua New Guinea. Transactions of the Royal Society of Tropical Medicine and Hygiene 90(1):57–60.
Laurenson, I. F., D. G. Lalloo, S. Naraqi, R. A. Seaton, A. J. Trevett, A. Matuka, and I. H. Kevau. 1997. Cryptococcus neoformans in Papua New Guinea: A common pathogen but an elusive source. Journal of Medical and Veterinary Mycology 35(6):437–440.
Li, S. S., and C. H. Mody. 2010. Cryptococcus. Proceedings of the American Thoracic Society 7(3):186–196.
Litvintseva, A. P., R. Thakur, L. B. Reller, and T. G. Mitchell. 2005. Prevalence of clinical isolates of Cryptococcus gattii serotype C among patients with AIDS in sub-Saharan Africa. Journal of Infectious Disease 192(5):888–892.
Lockhart, S. R., J. M. Schupp, J. D. Gillece, D. M. Engelthaler, and S. A. Balajee. 2010. Next gen sequencing helps unravel the molecular epidemiology of the emerging fungal pathogen Cryptococcus gattii. Presentation M-617. Paper presented at the Interscience Conference on Antimicrobial Agents and Chemotherapy, Boston, MA, September 12–15, 2010.
Loperena-Alvarez, Y., P. Ren, X. Li, D. J. Bopp, A. Ruiz, V. Chaturvedi, and C. Rios-Velazquez. 2010. Genotypic characterization of environmental isolates of Cryptococcus gattii from Puerto Rico. Mycopathologia 170(4):279–285.
Lopez-Martinez, R., J. L. Soto-Hernandez, L. Ostrosky-Zeichner, L. R. Castanon-Olivares, V. Angeles-Morales, and J. Sotelo. 1996. Cryptococcus neoformans var. gattii among patients with cryptococcal meningitis in Mexico. First observations. Mycopathologia 134(2):61–64.
Lui, G., N. Lee, M. Ip, K. W. Choi, Y. K. Tso, E. Lam, S. Chau, R. Lai, and C. S. Cockram. 2006. Cryptococcosis in apparently immunocompetent patients. QJM 99(3):143–151.
MacDougall, L., S. E. Kidd, E. Galanis, S. Mak, M. J. Leslie, P. R. Cieslak, J. W. Kronstad, M. G. Morshed, and K. H. Bartlett. 2007. Spread of Cryptococcus gattii in British Columbia, Canada, and detection in the Pacific Northwest, USA. Emerging Infectious Diseases 13(1):42–50.
Mitchell, D. H., T. C. Sorrell, A. M. Allworth, C. H. Heath, A. R. McGregor, K. Papanaoum, M. J. Richards, and T. Gottlieb. 1995. Cryptococcal disease of the CNS in immunocompetent hosts: Influence of cryptococcal variety on clinical manifestations and outcome. Clinical Infectious Diseases 20(3):611–616.
Mitchell, T. G., and J. R. Perfect. 1995. Cryptococcosis in the Era of AIDS—100 years after the discovery of Cryptococcus neoformans. Clinical Microbiology Reviews 8(4):515–48.
Morgan, J., K. M. McCarthy, S. Gould, K. Fan, B. Arthington-Skaggs, N. Iqbal, K. Stamey, R. A. Hajjeh, and M. E. Brandt. 2006. Cryptococcus gattii infection: Characteristics and epidemiology of cases identified in a South African province with high HIV seroprevalence, 2002–2004. Clinical Infectious Diseases 43(8):1077–1080.
Ngamskulrungroj, P., C. Serena, F. Gilgado, R. Malik, and W. Meyer. 2010. Global VGIIa isolates are of comparable virulence to the major fatal Cryptococcus gattii Vancouver Island outbreak genotype. Clinical Microbiology and Infection 17(2):251–258.
Okamoto, K., S. Hatakeyama, S. Itoyama, Y. Nukui, Y. Yoshino, T. Kitazawa, H. Yotsuyanagi, R. Ikeda, T. Sugita, and K. Koike. 2010. Cryptococcus gattii genotype VGIIa infection in man, Japan, 2007. Emerging Infectious Diseases 16(7):1155–1157.
Olivares, L. R., K. M. Martinez, R. M. Cruz, M. A. Rivera, W. Meyer, R. A. Espinosa, R. L. Martinez, and G. M. Santos. 2009. Genotyping of Mexican Cryptococcus neoformans and C. gattii isolates by PCR-fingerprinting. Medical Mycology 20:1–9.
Park, A. 2010. The “killer fungus”: Should we be scared? TIME, April 23, 2010.
Park, B. J., K. A. Wannemuehler, B. J. Marston, N. Govender, P. G. Pappas, and T. M. Chiller. 2009. Estimation of the current global burden of cryptococcal meningitis among persons living with HIV/AIDS. AIDS 23(4):525–530.
Perfect, J. R., and A. Casadevall. 2011. The history of cryptococcus and cryptococcosis. In Cryptococcus: From human pathogen to model yeast, pp 17–26, edited by J. Heitman, T. R. Kozel, K. J. Kwon-Chung, J. R. Perfect, and A. Casadevall. Washington, DC: ASM Press.
Perfect, J. R., W. E. Dismukes, F. Dromer, D. L. Goldman, J. R. Graybill, R. J. Hamill, T. S. Harrison, R. A. Larsen, O. Lortholary, M. H. Nguyen, P. G. Pappas, W. G. Powderly, N. Singh, J. D. Sobel, and T. C. Sorrell. 2010. Clinical practice guidelines for the management of cryptococcal disease: 2010 update by the Infectious Diseases Society of America. Clinical Infectious Diseases 50(3):291–322.
Randhawa, H. S., A. Y. Mussa, and Z. U. Khan. 2001. Decaying wood in tree trunk hollows as a natural substrate for Cryptococcus neoformans and other yeast-like fungi of clinical interest. Mycopathologia 151(2):63–99.
Saha, D. C., I. Xess, and N. Jain. 2008. Evaluation of conventional & serological methods for rapid diagnosis of cryptococcosis. Indian Journal of Medical Research 127(5):483–488.
Santos, W. R., W. Meyer, B. Wanke, S. P. Costa, L. Trilles, J. L. Nascimento, R. Medeiros, B. P. Morales, C. Bezerra Cde, R. C. Macedo, S. O. Ferreira, G. G. Barbosa, M. A. Perez, M. M. Nishikawa, and S. Lazera Mdos. 2008. Primary endemic Cryptococcosis gattii by molecular type VGII in the state of Para, Brazil. Memórias do Instituto Oswaldo Cruz 103(8):813–818.
Seaton, R. A., A. J. Hamilton, R. J. Hay, and D. A. Warrell. 1996a. Exposure to Cryptococcus neoformans var. gattii—a seroepidemiological study. Transactions of the Royal Society of Tropical Medicine and Hygiene 90(5):508–512.
Seaton, R. A., S. Naraqi, J. P. Wembri, and D. A. Warrell. 1996b. Predictors of outcome in Cryptococcus neoformans var. gattii meningitis. QJM 89(6):423–428.
Seaton, R. A., N. Verma, S. Naraqi, J. P. Wembri, and D. A. Warrell. 1997. Visual loss in immunocompetent patients with Cryptococcus neoformans var. gattii meningitis. Transactions of the Royal Society of Tropical Medicine and Hygiene 91(1):44–49.
Slobodniuk, R., and S. Naraqi. 1980. Cryptococcal meningitis in the central province of Papua New Guinea. Papua New Guinea Medical Journal 23(3):111–116.
Speed, B., and D. Dunt. 1995. Clinical and host differences between infections with the two varieties of Cryptococcus neoformans. Clinical Infectious Diseases 21(1):28–34; discussion 35–36.
Springer, D. J., and V. Chaturvedi. 2010. Projecting global occurrence of Cryptococcus gattii. Emerging Infectious Diseases 16(1):14–20.
Steele, K. T., R. Thakur, R. Nthobatsang, A. P. Steenhoff, and G. P. Bisson. 2010. In-hospital mortality of HIV-infected cryptococcal meningitis patients with C. gattii and C. neoformans infection in Gaborone, Botswana. Medical Mycology 8(8):1112–1115
Upton, A., J. A. Fraser, S. E. Kidd, C. Bretz, K. H. Bartlett, J. Heitman, and K. A. Marr. 2007. First contemporary case of human infection with Cryptococcus gattii in Puget Sound: Evidence for spread of the Vancouver Island outbreak. Journal of Clinical Microbiology 45(9):3086–3088.
Villanueva, E., M. Mendoza, E. Torres, M. B. de Albornoz, M. E. Cavazza, and G. Urbina. 1989. [Serotyping of 27 Cryptococcus neoformans strains isolated in Venezuela]. Acta Cientifica Venezolana 40(2):151–154.
Warkentien, T., and N. F. Crum-Cianflone. 2010. An update on Cryptococcus among HIV-infected patients. International Journal of STD and AIDS 21(10):679–684.
Woods, M. L., II, R. MacGinley, D. P. Eisen, and A. M. Allworth. 1998. HIV combination therapy: Partial immune restitution unmasking latent cryptococcal infection. AIDS 12(12):1491–1494.
Joseph Heitman,32,33Edmond J. Byrnes III32,34and John R. Perfect32,33
Abstract
How microbial pathogens emerge to cause outbreaks and become established as agents of disease in humans involves genetic exchange, zoonotic transmission, and perturbations of ecosystems and habitats. The threat of emerging infectious diseases is particularly poignant for eukaryotic pathogens, the fungi, and parasites, given that these microbes are more difficult to treat and have complex genomes and lifecycles. A sobering recent development has been the emergence and reemergence of several fungal pathogens in both humans and other animals, including Geomyces destructans in bats, Batrachochytrium dendrobatidis in amphibians, Nosema ceranae in bees (colony collapse disorder), and Cryptococcus gattii in humans and other animals in the Pacific Northwest. Here we review issues surrounding the C. gattii outbreak that began on Vancouver Island in 1999 and has expanded into the United States in Washington, Oregon, and California and has the potential to expand further. The focus will be on the emergence of C. gattii in the United States, including the appearance of a novel, highly virulent genotype and the potential role of sexual reproduction in the emergence of novel pathogens and their dispersal via airborne spores.
Introduction
The early history of cryptococcosis was documented in single or small series of cases. From an initial case of tibial osteomyelitis with the encapsulated Cryptococcus neoformans yeast in 1895 until a seminal monograph on this disease by Littman and Zimmer in 1956, the entire repertoire of reports in the medical literature numbered less than 300 cases (Littman and Zimmer, 1956). This was a humble beginning for this cosmopolitan, encapsulated basidiomycete that has now emerged into an outbreak mode in the new millennium. In the first half-century of its known existence, many of the clinical features of cryptococcosis were well described, including its propensity to invade the central nervous
_____________________
32 Department of Molecular Genetics and Microbiology, Duke University Medical Center.
33 Division of Infectious Diseases, Department of Medicine, Duke University Medical Center.
34 Current address: Division of Infectious Diseases, Johns Hopkins University School of Medicine, Baltimore, MD, USA 21287-0005.
system. The occurrence of outbreaks of mastitis (persistent inflammation of the udders) in dairy herds in which hundreds of animals were infected brought the realization that cryptococcal infection outbreaks can occur in mammals (Pounden et al., 1952; Simon et al., 1953). These outbreaks in animals and the link to the environment again were emphasized with reports of goats developing Cryptococcus gattii infection in Spain temporally linked to the importation of Eucalyptus trees (Baro et al., 1998). These two outbreaks in animals vividly demonstrated that cryptococcal infections could change from sporadic to outbreak as the endemic status changes. However, the first prescient report that recognized the future emergence of human cryptococcosis was from Kaufman and Blumer at the Centers for Disease Control and Prevention (CDC) when they called cryptococcosis “an awakening giant” mycoses (Kaufman and Blumer, 1978). As clinical mycologists in a major reference laboratory, they observed increasing numbers of cases as the immunosuppressed population dramatically increased due to the use of immunosuppressive therapies in modern medicine. In 1983 to 1984, reports of opportunistic cryptococcal infections coinciding with the early natural history of HIV infection provided insights into the emerging association of HIV infection and cryptococcosis (Lerner and Tapper, 1984; Vieira et al., 1983). In a landmark epidemiologic work in 2009, Park and colleagues from the CDC estimated that, in association with the AIDS pandemic, there were approximately 1 million cases of cryptococcosis per year worldwide, with at least 600,000 deaths per year in the past 5 years due to cryptococcosis (Park et al., 2009). While this massive outbreak continues, on another front, an outbreak of C. gattii infections on Vancouver Island has been observed over the past decade and has now migrated down into the Northwest United States, infecting humans and other mammals (Byrnes and Heitman, 2009; Byrnes et al., 2009, 2010; DeBess et al., 2010; Fraser et al., 2005; Kidd et al., 2004; MacDougall et al., 2007; Upton et al., 2007).35 Cryptococcosis started as a medical curiosity in a few patients, but because of medical interventions with immunosuppression, an immunosuppressive viral pandemic (HIV/AIDS), and a change in local climates, this yeast has become more prevalent in clinical medicine. Its present impact supports its title as a major emerging fungal disease or, to be more direct, “the giant is fully awake.”
Vancouver Island C. gattii Outbreak and Expansion into the United States
Our focus in this chapter will be the C. gattii outbreak that began on Vancouver Island in 1999 and has now expanded into the Canadian mainland in British Columbia and into the United States. A considerable body of knowledge is available with respect to the life and virulence cycles for this pathogenic yeast (Heitman, 2011). C. neoformans and C. gattii are closely aligned species. C. neoformans is prevalent in the environment globally, and C. gattii has been
_____________________
35 See also the contributed manuscript by Harris in Appendix A (pages 207–225).
thought to be geographically restricted to tropical and subtropical regions until its recent emergence in the relatively temperate climate on Vancouver Island. We are exposed to both organisms from the environment; C. neoformans is typically associated with pigeon guano and less commonly with trees, whereas C. gattii is commonly isolated from trees and soil, and also present in the air on Vancouver Island. We are exposed by inhaling spores and desiccated yeast cells, both of which can cause an initial pulmonary infection. This can be cleared, recede into a dormant latent form, cause fulminant pneumonia, or even make its way to the central nervous system via the bloodstream to infect both the covering of the brain (meninges) and the brain itself (meningoencephalitis). For reviews of the virulence and lifecycles, see Idnurm et al. (2005) and Kronstad et al. (2011).
Of particular importance are recent studies from the CDC’s Park and colleagues (2009) that reveal that Cryptococcus has reached pandemic proportions. More than a million cases occur globally each year. This is largely in the context of the AIDS pandemic and results in more than 600,000 attributable mortalities, approximately a third of all AIDS-associated deaths. This is thought to be largely attributable to C. neoformans, but there is also likely to be a more substantial burden of C. gattii infection occurring globally than is currently appreciated given that few clinical microbiology laboratories routinely assign Cryptococcus species status. It is in this global context that we consider the expanding and ongoing outbreak of C. gattii in the Pacific Northwest (Figure A10-1).
Cryptococcus is a species complex, and it is important to know which pathogen you are dealing with in the context of infection. For example, this is particularly important with E. coli, in which various strains are causing infections such as EPEC, EHEC, UPEC, and VTEC (enteropathogenic E. coli, enterohaemorrhagic E. coli, uropathogenic E. coli, and verotoxin-producing E. coli). This is also true with fungal pathogens. Two species are currently recognized: C. neoformans and C. gattii (Figure A10-2). The two can be readily distinguished in clinical microbiology labs on L-canavanine glycine bromothymol blue (CGB) agar, exploiting the ability of C. gattii to cause a pH change on this media, resulting in a blue chromogenic reaction (Figures A10-2A and B). We now appreciate that the group of isolates currently recognized as C. gattii spans four cryptic species groups. Currently, these are recognized as the VGI, VGII, VGIII, and VGIV molecular types (Figure A10-2C) based on molecular phylogenetic analyses that show each as a distinct, well-defined group (Bovers et al., 2008; Fraser et al., 2005; Ngamskulrungroj et al., 2009a). We also know from the whole-genome analysis for the representative VGI (WM276) and VGII (R265) isolates, which has just been completed at the Broad Institute and the University of British Columbia in Vancouver by Jim Kronstad and colleagues, that at a whole-genome level the different molecular types are not exchanging genetic information (D’Souza et al., 2011). Similarly, the molecular types are four genetically isolated cryptic species based on robust analysis of “molecular barcodes” throughout their genomes us-
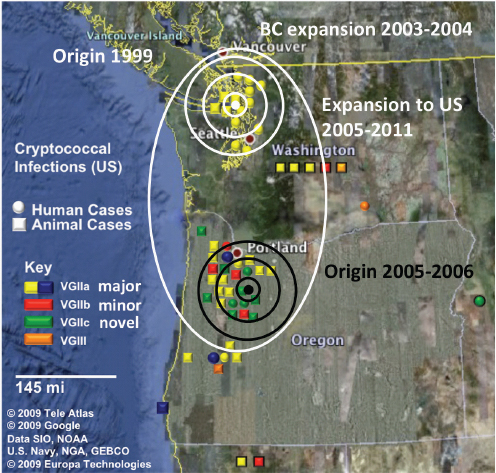
Figure A10-1 The C. gattii outbreak expanded into, and emerged within, the United States (n = 56). BC = British Columbia.
ing multilocus sequence typing (MLST) (Bovers et al., 2008; Fraser et al., 2005). This distinction of cryptic species is important because the VGII cryptic species is causing the outbreak in the Pacific Northwest, whereas the VGI lineage is more prevalent and more commonly causing disease in Australia (see Heitman, 2011).
The outbreak of C. gattii in the Pacific Northwest began in 1999 and prior to this, no C. gattii infections were reported as causing disease in this region of the world. The first cases came to the attention of astute clinicians and veterinarians on the southeastern shores in Naniamo and Parksville on Vancouver Island (Duncan et al., 2005; Hoang et al., 2004; Stephen et al., 2002). Over the past decade, there have been approximately 260 cases with an approximately 10 percent attributable mortality rate, and many infections in animals. Interested read-
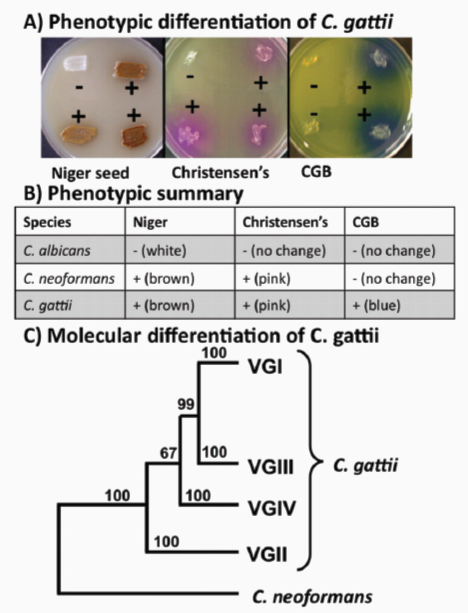
Figure A10-2 Cryptococcus pathogenic species complex. Panel A shows growth on niger seed media that detects production of the pigment melanin (+) or its absence (–), production of urease (+) as a pink color on Christensen’s agar, and growth and a blue color change (+) on CGB agar. Panel B shows a summary of the phenotypic data shown in panel A. Panel C is a phylogenetic tree of the relationships between the molecular types in the species complex. C. neoformans: opportunistic, AIDS, global, pigeon guano. C. gattii: primary, non-AIDS, tropical, arboreal. CGB = L-canavanine-glycine-bromothymol blue.
SOURCE: Panel A modified from Byrnes et al., PLoS Pathogens, 2010 (figure S2). Panel B courtesy of Heitman. Panel C modified from Chapter 22, Figure 2 of Carter et al. (2011).
are referred to a reprint36 included in this volume from Karen Bartlett, who played a critical role in identifying the environmental source of the organism on Vancouver Island (Bartlett et al., 2008), and to the manuscript from Julie Harris from the CDC Cryptococcus working group.37 Our charge in this article was to consider two aspects of the outbreak: first, its expansion into the United States, and second, the possible roles of sexual reproduction in the origin of the outbreak isolates and the ongoing production of airborne infectious spores.
The first cases associated with the Vancouver Island outbreak in patients from mainland British Columbia with no travel history to the island appeared in 2003–2004 (Bartlett et al., 2008). Environmental sampling studies provided evidence that C. gattii expanded across the water to a broader niche, including the Canadian mainland (MacDougall et al., 2007). The southern tip of Vancouver Island is very close to the U.S. border, and therefore a key question was if and when the outbreak might spread into the United States. The San Juan Archipelago is a part of Washington state, located as near as 5 km from the gulf islands off the coast of Vancouver Island. The first C. gattii index case in the United States was an elderly patient with leukemia on Orcas Island, Washington, who presented with a pulmonary C. gattii nodule in January 2006 (Upton et al., 2007). Since then, the outbreak has expanded into the United States from Canada, and we summarize here the evidence and key findings.
From 2006 to 2010, approximately 70 cases in patients have been reported, and the most complete records are attributable to the efforts of the CDC Cryptococcus working group. The cases in humans and animals are shown in Figure A10-1, and the icons represent cases for which we have isolates in the laboratory that have been molecularly analyzed. Two types of isolates are circulating on Vancouver Island: the VGIIa/major genotype, which causes approximately 95 percent of the infections, and the VGIIb/minor genotype, which represents the remaining 5 percent of isolates (Fraser et al., 2005). Both of these genotypes have been found in the environment on Vancouver Island and cause infection in both patients and animals, and both have now emerged within the United States. Of particular interest is a completely novel genotype that has emerged in Oregon: VGIIc, or the novel genotype (Byrnes et al., 2009, 2010). VGIIc has not yet been found in Washington state, Vancouver Island, or anywhere else in the world; thus, it is a completely new emergence from 2005 to 2010 in Oregon.
We can identify genotypes by applying MLST of genetic barcodes throughout the genome. In this technique, coding and non-coding regions of genes are PCR amplified, sequenced, and assigned a unique allele number. The alleles are then color coded and organized in a tabular format in which each line represents a different isolate associated with the outbreak or a global isolate in the strain collection. This allows one to appreciate that there has been what appears to be a large clonal expansion of the VGIIa/major genotype in the region that dominates
_____________________
36 See contributed manuscript by Bartlett in Appendix A (pages 101–116).
37 See contributed manuscript by Harris in Appendix A (pages 207–225).
the outbreak on Vancouver Island and its expansion into Puget Sound and beyond in Washington state. There are fewer isolates of the VGIIb/minor genotype, but these have been found on Vancouver Island, in Oregon, and most recently in Washington state. The VGIIc/novel genotype has thus far been found only in Oregon and has several, unique alleles not found to date in any other isolates examined from global sources. The CDC has identified one isolate from a patient in Idaho that appears to be closely related to the VGIIc/novel genotype, but it may be the result of travel exposure (DeBess et al., 2010).
Where did these VGII outbreak genotypes originate? The VGIIa/major genotype is indistinguishable across 30 MLST loci and several variable number of tandem repeat loci from the NIH444 strain, which was isolated in the early 1970s from a sputum sample from a patient in Seattle (Fraser et al., 2005). This isolate is the type strain for C. gattii and is molecularly indistinguishable from the VGIIa/major outbreak strain at all loci examined thus far. Therefore, the major outbreak strain appears to have been in this geographic region for at least 40 years and possibly even longer. The VGIIb/minor genotype isolates are indistinguishable from isolates from a fully recombining sexual population in Australia (Campbell et al., 2005a,b; Fraser et al., 2005). The VGIIc/novel genotype has thus far only been identified in isolates collected in Oregon, and we have not observed it in a large collection of more than 200 C. gattii isolates collected globally. It appears as if the VGIIc/novel genotype either emerged locally in Oregon or, alternatively, it may be present in an undersampled environmental niche that remains to be discovered. Further evidence that this new genotype in Oregon is novel involves haplotype network mapping. In this approach, the MLST alleles are compared and we apply a computer algorithm to predict the ancestral allele. The alleles are then organized into a diagram rooted with the ancestral allele, and alleles derived from it by mutations and genetic drift are indicated with lines and circles. In this analysis alleles that are arisen from the ancestral allele long ago lie closest to the ancestral allele, whereas alleles that arose recently lie distal. The MLST alleles that are private to the VGIIc genotype are distal in these haplotype networks, suggesting that they arose recently. The alleles that the VGIIc alleles are derived from come from isolates that originated from either Australia or South America (Byrnes et al., 2010). This analysis then suggests that those alleles might represent possible sites of origin of at least some of the genetic material in the Oregon VGIIc/novel outbreak isolate.
We have conducted mammalian virulence studies in a mouse inhalation model, both at the Wadsworth Center in Albany with our collaborators Ping Ren, Sudha Chaturvedi, and Vishnu Chaturvedi and also at Duke University (Byrnes et al., 2010; Fraser et al., 2005). The VGIIa/major genotype and the VGIIc/novel genotype isolates are both highly virulent in the mouse model used, whereas the minor VGIIb genotype is considerably attenuated by direct comparison (Byrnes et al., 2010). Isolates collected globally that share many but not all markers with the VGIIa/major genotype are considerably less virulent than VGIIa (Byrnes
et al., 2010). This includes, for example, isolates from both California and South America. Of note, the NIH444 type strain (a VGIIa/major genotype isolate) is also somewhat less virulent than contemporary outbreak VGIIa/major genotype strains, which may reflect either attenuation as a result of long-term storage and passage; variation in virulence even among the VGIIa/major isolates; or unknown genetic differences between NIH444 and the VGIIa/major outbreak genotypes in regions of the genome not yet analyzed. Therefore, it appears as if virulence has increased with the emergence of the VGIIa/major outbreak genotype and the VGIIc/novel genotype from their original source strains.
Several investigators have been addressing why outbreak isolates are more virulent than other genotypes. There have been two prescient studies. Jim Kronstad and colleagues at the University of British Columbia in Vancouver have shown that there is less protective inflammation in the lungs of mice infected with the outbreak isolates (Cheng et al., 2009). This may then lead to increased virulence if the host fails to mount a sufficient, protective immune response. In addition, the work from Robin May’s lab at the University of Birmingham has described an increased intracellular proliferation rate of the VGIIa and VGIIc outbreak isolates in macrophages and linked mitochondrial function to the robustness of their interaction with host immune cells (Byrnes et al., 2010; Ma et al., 2009). Given that Cryptococcus is a pathogen able to grow either outside or inside of host immune cells, an enhanced proliferation rate in the context of the macrophage intracellular milieu may lead to enhanced virulence. These studies are excellent starting points to use to begin to dissect the virulence mechanisms at the interface with host immune cells in the lung.
To summarize, this outbreak was originally restricted geographically to Vancouver Island and then spread to the mainland of British Columbia. Starting in 2006 (and maybe earlier in one case in Oregon in 2005), it emerged within the United States. Thus, there clearly appears to have been a geographic expansion from Vancouver Island across Puget Sound to reach Washington, mostly involving the VGIIa/major genotype. However, there is more diversity in strains from Oregon, involving both the VGIIa/major and VGIIb/minor genotypes observed on Vancouver Island, but also including the novel VGIIc genotype that has not been identified on Vancouver Island. In essence, this looks like an outbreak within an outbreak, which may be of independent origins. It is as though two pebbles have been dropped into a pond at different times, one earlier than the other, and they have generated concentric waves that are now expanding outward and intersecting. There is considerable concern that this outbreak will continue to expand geographically given that there are cases and isolates now from Idaho and California (Personal communication, Shawn Lockhart, CDC Mycotic Diseases Branch, May 2011). There is also a well-documented risk to travelers to the endemic region, who then return to locations around the world with an infection of the VGIIa/major isolate and present with a very unusual fungal infection that is not commonly described in those regions and that might confuse clinicians
(Chambers et al., 2008; Georgi et al., 2009; Hagen et al., 2010; Lindberg et al., 2007).
The Role of Sexual Reproduction in Pathogen Emergence and Outbreaks
The second charge of this article was to consider the role of sexual reproduction in the emergence of pathogens and how it might contribute to adapting to a novel niche. We might start with just a very general question of, why sex? For those investigators who focus on bacteria or other prokaryotes, their mechanisms for genetic transfer typically involve horizontal gene transfer. But for the eukaryotic pathogens, including fungi, parasites, and the oomycete plant pathogens such as Phytophthora infestans (the cause of potato blight), their genetic exchange mechanisms involve sexual reproduction. The general theme that emerges for all of these groups of eukaryotic pathogens is that sexual reproduction plays a critical role in their diversity and, in many cases, also in their infection cycles (Heitman, 2006, 2010).
What benefits are conferred by sexual reproduction? This process enables the exchange of genetic information and generates diversity, and it can also purge the genome of deleterious mutations. It also allows these organisms to stay one step ahead of their transposable elements, which might otherwise accumulate to litter the genome.
We know a great deal about the sexual reproduction of Cryptococcus because of the pioneering work of June Kwon-Chung at the National Institutes of Health some 30 years ago (Kwon-Chung, 1975, 1976a,b). There are two mating types, called a and α (Hull and Heitman, 2002; McClelland et al., 2004). A panoply of signals regulates the interactions of cells of opposite mating type, allowing fusion and completion of their sexual cycle, thereby generating infectious spores. We know from both historic and recent experimental studies that these spores are indeed infectious propagules (Botts and Hull, 2010; Botts et al., 2009; Giles et al., 2009; Sukroongreung et al., 1998; Velagapudi et al., 2009; Zimmer et al., 1984).
Two of the many signals that regulate the C. gattii sexual cycle are interactions with plants and extreme desiccation (Xue et al., 2007). As noted by Karen Bartlett, there is more Cryptococcus found in the air in Vancouver Island in July when it is very hot and dry (Bartlett, 2010). Indeed, these are also the ideal conditions for stimulating the sexual cycle, at least under laboratory conditions. Thus, Cryptococcus may mate more in the hot and dry conditions of July than in other times of the year.
Sex in Cryptococcus produces spores, which are infectious. There is also a link of the mating type to virulence that has focused interest on its sexual cycle (Kwon-Chung et al., 1992; Nielsen et al., 2003, 2005a,b; Okagaki et al., 2010). One of the most curious features of this species is that most of the Cryptococcus population is just one mating type (α), even though the sexual cycle that was originally defined in the laboratory requires both opposite mating types (Hull
and Heitman, 2002). This fact raised a central conundrum for the entire field of cryptococcal pathogenesis: If there is a well-maintained a-α opposite-sex sexual cycle, but the a mating type is extremely rare, if not completely absent in many populations, how then can a sexual cycle be an important step of the infectious cycle for this fungus? It might be that it is not important, such that these unisexual populations were just clonally reproducing mitotically as yeasts. However, we discovered that C. neoformans can undergo an unusual sexual cycle involving only one of the two mating types (Lin et al., 2005).
The opposite-sex cycle that we have known about for 30 years is depicted in the lower panel of Figure A10-3. Then in 2005, an alternative sexual cycle was reported called unisexual reproduction or same-sex mating, and this is depicted in the upper panel of Figure A10-3 (Lin et al., 2005; Wang and Lin, 2011). This alternative sexual cycle only involves α cells. They can fuse with another α cell from the population, or they can, in extreme cases, fuse with themselves, undergo meiosis, and produce spores.
You might wonder, what would be the point of undergoing sexual reproduction with yourself? There is no genetic diversity to admix in this circumstance. We always think about sexual reproduction as involving two parents with very
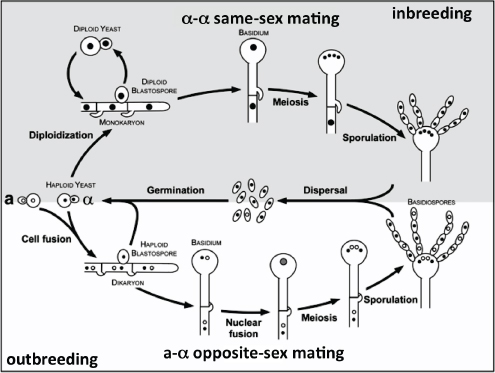
Figure A10-3 Cryptococcus neoformans can reproduce unisexually and bisexually.
different genetic compositions. It turns out that mating with yourself can also lead to the generation of genetic diversity. Sex itself can serve as something of a mutagen to generate genotypic and phenotypic diversity. This strategy turns completely on its head what we traditionally think of as the primary role of sexual reproduction. An analogy might be the appearance of mismatch repair mutator mutations in bacterial pathogens, which arise to result in the generation of genome-wide mutations and are then swept from the population as a consequence of their concomitant deleterious effects.
As an example, in preliminary studies we isolated 100 progeny produced by this same-sex mating cycle and looked for phenotypic diversity. We have found within this set of just 100 isolates clear examples of azole resistance, temperature-sensitive growth, hyperfilamentous growth, and increased melanin production. By comparison, analysis of 100 progeny produced by mitotic clonal growth yielded no such examples. Many of the phenotypically distinct isolates turn out to be aneuploid in one way or another. This is based on comparative genome hybridization analysis showing that one of these morphologically distinct isolates now has an extra copy of chromosome 10. The presence of an extra copy of a chromosome is termed aneuploidy. We often associate aneuploidy with very deleterious consequences. We do not have to go further than Down syndrome as an example to think about what might be bad about aneuploidy. On the other hand, in the microbial kingdom, aneuploidy can be a rich source of phenotypic diversity. There are well-documented studies in Candida albicans that an a special type of chromosome formed by duplicated arms of a chromosome (termed an isochromosome) can form and drive fluconazole resistance (Selmecki et al., 2006, 2008, 2009). Similarly, we also know that azole heteroresistance in Cryptococcus is driven by aneuploidy for chromosome 1 (Hu et al., 2008; Sionov et al., 2009, 2010). A variety of other experimental evolution studies have brought to the forefront the idea that aneuploidy might be a driving force for genotypic and phenotypic plasticity (Pavelka et al., 2010; Rancati et al., 2008; Torres et al., 2007, 2010). So this may be one mechanism by which a same-sex mating cycle could engender phenotypic and genotypic diversity in a population without needing a partner that is genetically divergent.
Another way to view same-sex mating is that it is a mechanism for selfing. Selfing/inbreeding is common in fungi and the paradigm is mating type switching in S. cerevisiae, which allows mother–daughter cell mating. Same-sex mating is another route to self, but not via mating type switching. It is a question at some level of outcrossing vs. inbreeding. Same-sex mating superimposes much more limited genetic diversity on a well-adapted fungal genome that has run the gauntlet of natural selection. As such, unisexual reproduction may confer one of the benefits of sex (generation of diversity), but at the same time avoid one of its costs (breaking apart well-adapted genomic/genetic configurations). For organisms that are well adapted to a niche, this more limited genetic diversity may better answer subtle changes in selection pressures.
We have alluded to the fact that there are restricted geographic areas where fungal sex is extant. Why is that? Why would you have these little areas and pockets in Africa and that is where they have sex, and in the rest of the world they don’t have sex except for the unusual unisexual cycle? At least for Cryptococcus, what it looks like is, where they have opposite-sex mating seems to be where there are restricted populations where both mating types still exist. We know the most about this from C. neoformans, where there is an extant population that is on a very specific indigenous tree in South Africa, in Botswana (Litvintseva et al., 2011). But everywhere else in the world, you find just the alpha mating type (Hull and Heitman, 2002). So there is a niche in nature where opposite-sex mating is occurring. Until 5 years ago everyone presumed everywhere else it was asexual and mitotic. But what we are coming to appreciate is that there is just a different version of the sexual cycle occurring in these other populations where there is just one mating type.
This is interesting as another classic example is Phytophthora infestans. In Mexico and in Peru and other areas of South America, that is where the sexual cycle occurs, new versions arise, and then they are often exported on potato crops. That was the source of the Irish potato famine. But why there is a sexual cycle in such a restricted place is not really clear. Toxoplasma gondii might be another interesting pathogen to consider. It only has its sexual cycle in cats and other felids, not in other animals of any sort (Heitman, 2006). So what is it about the gastrointestinal tract of a cat that is so different from a mouse or a human that allows this parasite to undergo its sexual cycle there? The parasite sexual cycles are even more bizarre than some of the fungal ones, and restricted to extreme niches (Heitman, 2006, 2010).
What about the specific example of C. gattii sexual reproduction? June Kwon-Chung elucidated the C. gattii sexual cycle more than 30 years ago (Kwon-Chung, 1976a,b). Spores were produced in her classic studies via an a-α opposite-sex mating cycle that she defined. We were approached by several of the Australian groups working on the VGI genotypes in the late 1990s, including Wieland Meyer and Dee Carter, and we spent more than 5 years recapitulating this sexual cycle under laboratory conditions. This required a great deal of heroic effort on the part of several individuals in various laboratories, in large part because fecundity can be reduced with long-term passage and storage and because the VGI molecular type that is predominant in Australia is more recalcitrant in mating assays compared to the VGII and VGIII C. gattii molecular types. We were finally able to recapitulate a sexual cycle for C. gattii involving the formation of dikaryotic hyphae with special cells linking the hyphal cells (fused clamp cells) culminating in the production of basidia and basidiospores, all of the morphological features of the sexual cycle, including meiotic recombination (Campbell et al., 2005a,b; Fraser et al., 2003). Based on this advance, we were able to show that the vast majority of the outbreak isolates from Vancouver Island are fertile in laboratory crosses (Byrnes et al., 2010; Campbell et al., 2005a;
Fraser et al., 2003). These are a-α matings that were conducted, but we stress the point that every single isolate that is associated with the outbreak is of the α mating type. No one has found any a isolates occurring anywhere on Vancouver Island or in Washington or Oregon.
How might mating occur on Vancouver Island and in the Pacific Northwest in this unisexual population? In previous studies we observed that an association of Cryptococcus with plants can stimulate the fungal sexual cycle (Xue et al., 2007). Part of the idea in testing for this phenomenon in the first place was that plants, and more specifically trees, are the environmental niche for C. gattii (Heitman et al., 2011). For the plant pathogenic fungus Ustilago maydis, which infects corn, the infectious form is the filamentous dikaryotic hyphae produced by the sexual cycle, which is stimulated by interaction with the plant. In fact, U. maydis has to be in its sexual form to infect the plant host. We found something very similar with Cryptococcus. We were unable to infect a plant efficiently with the haploid yeast, but the filamentous dikaryotic state will infect plants and stimulate a plant defense response (Xue et al., 2007). In turn, if we put seedlings distant from a fungal mating mixture on a plate, the plants secrete small molecules, such as inositol, that stimulate completion of the sexual cycle and production of spores (Xue et al., 2007, 2010). We observed this with seedlings of Arabidopsis or Eucalyptus and are now exploring this with Douglas fir because of the link to this indigenous tree species in the Pacific Northwest outbreak. This line of investigation suggests that this may be one niche in nature where the sexual cycle is occurring to produce spores as small airborne infectious propagules.
Another critical line of investigation involves population genetic studies conducted by Dee Carter from the University of Sydney, Australia. She has focused on the sexual cycle that may be occurring in Eucalyptus tree hollows. She discovered that in populations with both mating types, they can engage in opposite-sex mating, but in other tree hollows where only α isolates are found, they are also sexually recombining (Saul et al., 2008). So it seems as if the organism has two extant sexual cycles, one opposite sex and one same sex, and whom you mate with depends on who your neighbors are in the population. All of her studies have focused on the VGI cryptic species, which has very limited genetic diversity and an extremely high level of apparent inbreeding (Campbell and Carter, 2006; Campbell et al., 2005a,b; Carter et al., 2007, 2011). All of the isolates on Vancouver Island associated with the clonal outbreak are of a different cryptic species, VGII. However, based on Dee Carter’s studies of VGI sexual reproduction, the presumption is that there may be both forms of sexual reproduction occurring also for the VGII isolates in nature, associated with the environmental niche in trees or plants for the VGII lineage and involving both opposite and same-sex mating depending on where a isolates are found in nature.
We have advanced the population genetic analysis in two ways with our global isolates to look for measures of sexual reproduction and recombination occurring in the population. One of these is a simple test called an allele com-
patibility test. The basic premise is that if you cross two strains that differ at two markers, you obtain four types of progeny produced by sexual processes by reassorting two unlinked loci (i.e., AB by ab yields AB, ab, Ab, and aB) (Carter et al., 2011), which we all know from studying Punnett squares. So the multilocus sequence loci can be analyzed as two unlinked alleles in the population. If you find all four possible combinations, this is indicative of sexual reproduction and recombination occurring in the population. By looking at the multilocus sequence markers, we find that these measures of recombination are rampant throughout the VGII global population (Byrnes et al., 2010). In addition, if we just examine the sequences of the multilocus sequence alleles themselves, which represent approximately 1 kilobase of sequence each, we can find examples of hybrid recombinant alleles that implicate isolates as potential parents or potential offspring (Byrnes et al., 2010). A lot of mitotic recombination must be occurring to see evidence of recombination within just a random 1-kilobase sequence in a 20-megabase genome. The potential parents that are identified by this type of analysis originate from South America, Africa, and Australia (Byrnes et al., 2010). These findings suggest that sexual reproduction is occurring globally in multiple environments and locations.
We went on to examine these global isolates for fertility in the lab and documented, by scanning with electron microscopy, the production of meiotic spores, which are the products of sex that may be infectious propagules found in the air on Vancouver Island. The majority of the α strains we analyzed are fertile. We have found a very limited number of a strains in the global population. For example, we found 6 out of 200 (~3 percent) are this minor mating type. Five out of six of these a strains are fertile in the laboratory, and thus they may be undergoing opposite-sex mating in nature. Because five of these six a isolates are from South America, it may be a site in which opposite-sex mating is still extant in the population (Escandon et al., 2007; Fraser et al., 2005; Ngamskulrungroj et al., 2008). We know that C. neoformans has also retained extant opposite-sex mating, but that it is geographically restricted to Botswana and South Africa and occurring in the VNB C. neoformans lineage (Litvintseva et al., 2003, 2007).
When we return to consider the VGIIb minor genotype and its relationship to global isolates, this provides critical insight. For instance, we looked at 30 MLST alleles, the VGIIb outbreak isolates are completely indistinguishable from isolates from a fertile, unisexual, sexually recombining population in Australia that was identified by Dee Carter and colleagues (Campbell et al., 2005a,b). It suggests that this population may be the ancestral source for the VGIIb/minor outbreak genotype isolates now circulating in the Pacific Northwest. While one might posit just the opposite (transfer from Vancouver Island to Australia), the diversity of the population in Australia supports this as the ancestral rather than the derived population. Thus, the most parsimonious model is that the isolates from the outbreak originated from Australia (Figure A10-4).
Based on a broad comparison of the isolates on Vancouver Island, the VGIIa/
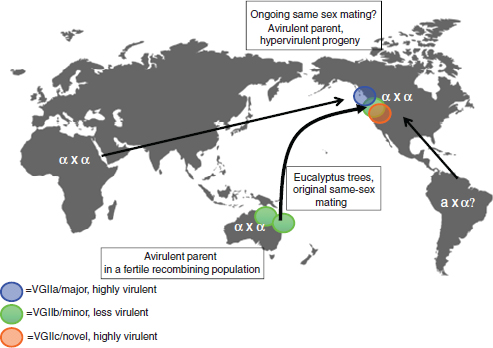
FIGURE A10-4 Sexual reproduction and the origin of an outbreak.
major and the VGIIb/minor genotypes share half of their genetic markers and differ at the other half. So in a very simple model, they might be progeny from a genetic cross (i.e., siblings), or one might be a parent and the other an offspring. In this simple conceptual framework, one could imagine an isolate undergoing mating with an unknown isolate in nature to produce the outbreak isolates. However, the VGIIa/major and the VGIIb/minor genotypes on Vancouver Island are all of the same mating type, α. When we look in detail at the mating type locus, we can see that they are molecularly distinct from each other (Fraser et al., 2005) and are not identical by descent. So, in essence, they have two different ancestries for this region of their genome that dictates their mating type. A more parsimonious model then is that genetic crosses that led to the production of these isolates would have involved two parents with different α mating alleles in a same-sex mating cycle. It is also possible that both types of crosses have occurred (opposite-sex and same-sex mating) and that there has been more than one cross involved here in the genesis of these isolates that are responsible for the outbreak and also for the global isolates to which they are related.
Conclusions and Perspective
To summarize, three molecular genotypes are found circulating in the Pacific Northwest. The most prominent is the VGIIa/major genotype (Figure A10-1 in yellow), and we can date its origin to at least the early 1970s to an isolate from a patient in Seattle. The VGIIb/minor genotype is shown in red and it is molecularly indistinguishable from isolates from a fertile recombining population in Australia. In green is a completely novel VGIIc genotype that has emerged in Oregon. Two of these genotypes (VGIIa, VGIIc) are highly virulent in both macrophage and mouse models (Byrnes et al., 2010).
From where do these VGII isolates originate? We find VGII isolates globally: in Africa, Australia, and South America. But based on available evidence to date, the most parsimonious model is that at least one of these genotypes (VGIIb/minor) came from a population in Australia, given that isolates from the two are indistinguishable (Figure A10-4). In terms of the crosses hypothesized to have been involved in the origin of the outbreak, all of the populations that have been identified are all α. There are no a mating types of this lineage that have been found on Vancouver Island, in the Pacific Northwest, or in Australia. The only places where VGII mating type a isolates have been found so far are South America and Greece. So it may be that in Australia and the Pacific Northwest that same-sex mating drives diversity in the population, but opposite-sex mating is a possibility in South America. In the context of the outbreak, we also envision that same-sex mating is contributing to the production of infectious spores, which are the aerosolized small particles detected with Anderson air samplers. It is entirely possible that those propagules are desiccated yeast cells, spores, or a mixture of both. Experiments to test if these are actually spores in nature are being planned, and if spores are present in the air on Vancouver Island, it seems likely that they are being produced by same-sex mating occurring in the environment, given that the population is α unisexual.
Disease caused by C. gattii vs. C. neoformans has been the subject of several reviews, and despite the impression that C. gattii frequently causes infection in apparently immunocompetent hosts and commonly presents with focal cryptococcomas in the lung or brain (Chen et al., 2000; Mitchell et al., 1995; Speed and Dunt, 1995), it has been difficult to clinically distinguish between C. neoformans and C. gattii disease. In fact, the presentation of C. gattii infections in Vancouver and the Pacific Northwest outbreak appear similar to C. neoformans infections, and disease is not limited to the immunocompetent host. C. gattii infections such as those caused by the VGIII molecular type/cryptic species can be even found in HIV-infected individuals in endemic areas (Byrnes et al., 2011; Chaturvedi et al., 2005). Another aspect that has been reported to distinguish these two species is their response to therapy. It has been suggested that C. gattii infections require longer therapy (West et al., 2008). However, in vitro susceptibility testing has produced inconsistent results; some investigators have found similar minimum inhibitory concentrations (MICs) whereas others have found higher MICs to azoles for
C. gattii compared to C. neoformans (Thompson et al., 2009; Torres-Rodriguez et al., 2008). It is always difficult to be precise in determining the response to therapy when in a normal host; a component of immune reconstitution inflammatory syndrome may influence clinical judgment of treatment failure. A recent study primarily in HIV-infected patients demonstrated that there was no difference in the outcome of C. gattii vs. C. neoformans infections in a single medical center (Steele et al., 2010). This is interesting and may be due to small numbers, but early results comparing C. gattii infections in the Vancouver Island vs. the Pacific Northwest outbreaks suggested a higher mortality in the United States (Kluger et al., 2006). After review of present data on outcomes in the literature, in vitro drug susceptibility studies, and clinical experience, the 2010 Infectious Diseases Society of America Cryptococcal Guidelines support the treatment of C. gattii infections in a manner similar to those of C. neoformans (Perfect et al., 2010). However, C. gattii infections may produce more cryptococcomas in the brain or lung, and chronic conditions such as hydrocephalus and a slow response to antifungal therapy may be prominent features of disease with this species.
Despite the lack of precise differences in C. gattii vs. C. neoformans disease clinically, which would require immediate identification of the species, there are differences in epidemiology regarding the range of infections. It is also suggested on a pathobiological basis that C. gattii may present more with acute disease rather than reactivation disease. This hypothesis is based on (1) antibody studies in children where C. neoformans, but not C. gattii, antibodies are frequently detected, and (2) the higher percentage of disease that is observed in immunocompetent patients with C. gattii (Goldman et al., 2001). It has also been shown that C. gattii may induce a different host immune response compared to C. neoformans (Cheng et al., 2009). Furthermore, the Vancouver Island outbreak represents an ideal arena to check for genetic susceptibility to C. gattii infection because a vast number of individuals were exposed to high numbers of infectious propagules, but only a minority of those exposed was identified with clinical disease. However, it is clear through both animal studies and transplant cases that C. gattii infections can reactivate (Dromer et al., 1992; Kluger et al., 2006). Finally, although major virulence factors (melanin, capsule, high-temperature growth) are similar, some of the networks such as the trehalose pathway connecting these virulence factors are different in these two species (Ngamskulrungroj et al., 2009b; Petzold et al., 2006). The possibility that differences in both host and microbe genetics may influence the course and management of infections will drive studies on this unique example of a fungal outbreak in humans as we continue its analysis.
Acknowledgments
We thank Wenjun Li and Yonathan Lewit for their contributions to the analysis of the outbreak; Dee Carter, Vishnu Chaturvedi, Jim Kronstad, Kieren Marr,
and Robin May for collaboration; Rory Duncan, Chris Lambros, and Dennis Dixon from the National Institute of Allergy and Infectious Diseases (NIAID) and Victoria McGovern from the Burroughs Wellcome Fund for their support; and Stephen Johnston for his prescient questions. We also explicitly thank all of the Forum members and the Institute of Medicine for shining a very bright light on the fungal kingdom, and for the invitation to participate. Our research is supported by NIH/NIAID R37 grant AI39115.
References
Baro, T., J. M. Torres-Rodriguez, M. H. De Mendoza, Y. Morera, and C. Alia. 1998. First identification of autochthonous Cryptococcus neoformans var. gattii isloated from goats with predominantly severe pulmonary disease in Spain. Journal of Clinical Microbiology 36(2):458–461.
Bartlett, K. 2010. Knowing where to look—environmental sources of cryptococcal disease in human and animal residents in the Pacific Northwest. Presentation given at the December 14–15, 2010 public workshop, “Fungal Diseases: An Emerging Challenge to Human, Animal, and Plant Health.” Forum on Microbial Threats, Institute of Medicine, Washington, DC.
Bartlett, K. H., S. E. Kidd, and J. W. Kronstad. 2008. The emergence of Cryptococcus gattii in British Columbia and the Pacific Northwest. Current Infectious Disease Reports 10(1):58–65.
Botts, M. R., and C. M. Hull. 2010. Dueling in the lung: How Cryptococcus spores race the host for survival. Current Opinion in Microbiology 13(4):437–442.
Botts, M. R., S. S. Giles, M. A. Gates, T. R. Kozel, and C. M. Hull. 2009. Isolation and characterization of Cryptococcus neoformans spores reveal a critical role for capsule biosynthesis genes in spore biogenesis. Eukaryotic Cell 8:595–605.
Bovers, M., F. Hagen, E. E. Kuramae, and T. Boekhout. 2008. Six monophyletic lineages identified within Cryptococcus neoformans and Cryptococcus gattii by multi-locus sequence typing. Fungal Genetics and Biology 45(4):400–421.
Byrnes, E. J., and J. Heitman. 2009. Cryptococcus gattii outbreak expands into the Northwestern United States with fatal consequences. F1000 Biology Reports 1:62.
Byrnes, E. J., III, R. J. Bildfell, S. A. Frank, T. G. Mitchell, K. A. Marr, and J. Heitman. 2009. Molecular evidence that the range of the Vancouver Island outbreak of Cryptococcus gattii infection has expanded into the Pacific Northwest in the United States. Journal of Infectious Diseases 199(7):1081–1086.
Byrnes, E. J., III, W. Li, Y. Lewit, H. Ma, K. Voelz, P. Ren, D. A. Carter, V. Chaturvedi, R. J. Bildfell, R. C. May, and J. Heitman. 2010. Emergence and pathogenicity of highly virulent Cryptococcus gattii genotypes in the northwest United States. PLoS Pathogens 6(4):e1000850.
Byrnes, E. J., W. Li, P. Ren, Y. Lewit, K. Voelz, et al. 2011. A diverse population of Cryptococcus gattii molecular type VGIII in Southern California HIV/AIDS patients. PLoS Pathogens. accepted in principle pending revision.
Campbell, L. T., and D. A. Carter. 2006. Looking for sex in the fungal pathogens Cryptococcus neoformans and Cryptococcus gattii. FEMS Yeast Research 6(4):588–598.
Campbell, L. T., B. J. Currie, M. Krockenberger, R. Malik, W. Meyer, J. Heitman, and D. Carter. 2005a. Clonality and recombination in genetically differentiated subgroups of Cryptococcus gattii. Eukaryotic Cell 4(8):1403–1409.
Campbell, L. T., J. A. Fraser, C. B. Nichols, F. S. Dietrich, D. Carter, and J. Heitman. 2005b. Clinical and environmental isolates of Cryptococcus gattii from Australia that retain sexual fecundity. Eukaryotic Cell 4(8):1410–1419.
Carter, D., N. Saul, L. Campbell, T. Bui, and M. Krockenberger. 2007. Sex in natural populations of Cryptococcus gattii. In Sex in fungi: Molecular determination and evolutionary implications, edited by J. Heitman, J. Kronstad, J. Taylor, and L. Casselton. Washington, DC: ASM Press. Pp. 477–488.
Carter, D., L. Campbell, N. Saul, and M. Krockenberger. 2011. Sexual reproduction of Cryptococcus gattii: A population genetics perspective. In Cryptococcus: From human pathogen to model yeast, edited by J. Heitman, T. R. Kozel, J. K. Kwon-Chung, J. R. Perfect, and A. Casadevall. Washington, DC: ASM Press. Pp. 299–311.
Chambers, C., L. MacDougall, M. Li, and E. Galanis. 2008. Tourism and specific risk areas for Cryptococcus gattii, Vancouver Island, Canada. Emerging Infectious Diseases 14(11):1781–1783.
Chaturvedi, S., M. Dyavaiah, R. A. Larsen, and V. Chaturvedi. 2005. Cryptococcus gattii in AIDS patients, southern California. Emerging Infectious Diseases 11(11):1686–1692.
Chen, S., T. Sorrell, G. Nimmo, B. Speed, B. Currie, D. Ellis, D. Marriott, T. Pfeiffer, D. Parr, and K. Byth. 2000. Epidemiology and host- and variety-dependent characteristics of infection due to Cryptococcus neoformans in Australia and New Zealand. Australasian Cryptococcal Study Group. Clinical Infectious Diseases 31(2):499–508.
Cheng, P. Y., A. Sham, and J. W. Kronstad. 2009. Cryptococcus gattii isolates from the British Columbia cryptococcosis outbreak induce less protective inflammation in a murine model of infection than Cryptococcus neoformans. Infection and Immunity 77(10):4284–4294.
DeBess, E., et al. 2010. Emergence of Cryptococcus gattii—Pacific Northwest, 2004–2010. Morbidity and Mortality Weekly Report 59:865–868.
Dromer, F., O. Ronin, and B. Dupont. 1992. Isolation of Cryptococcus neoformans var. gattii from an Asian patient in France: Evidence for dormant infection in healthy subjects. Journal of Medical and Veterinary Mycology 30(5):395–397.
D’Souza, C. A., J. W. Kronstad, G. Taylor, R. Warren, M. Yuen, G. Hu, W. H. Jung, A. Sham, S. E. Kidd, K. Tangen, N. Lee, T. Zeilmaker, J. Sawkins, G. McVicker, S. Shah, S. Gnerre, A. Griggs, Q. Zeng, K. Bartlett, W. Li, X. Wang, J. Heitman, J. E. Stajich, J. A. Fraser, W. Meyer, D. Carter, J. Schein, M. Krzywinski, K. J. Kwon-Chung, A. Varma, J. Wang, R. Brunham, M. Fyfe, B. F. Ouellette, A. Siddiqui, M. Marra, S. Jones, R. Holt, B. W. Birren, J. E. Galagan, and C. A. Cuomo. 2011. Genome variation in Cryptococcus gattii, an emerging pathogen of immunocompetent hosts. mBio 2(1):1–11.
Duncan, C., C. Stephen, S. Lester, and K. H. Bartlett. 2005. Sub-clinical infection and asymptomatic carriage of Cryptococcus gattii in dogs and cats during an outbreak of cryptococcosis. Medical Mycology 43(6):511–516.
Escandon, P., P. Ngamskulrungroj, W. Meyer, and E. Castaneda. 2007. In vitro mating of Colombian isolates of the Cryptococcus neoformans species complex. Biomedica 27(2):308–314.
Fraser, J. A., R. L. Subaran, C. B. Nichols, and J. Heitman. 2003. Recapitulation of the sexual cycle of the primary fungal pathogen Cryptococcus neoformans variety gattii: Implications for an outbreak on Vancouver Island. Eukaryotic Cell 2:1036–1045.
Fraser, J. A., S. S. Giles, E. C. Wenink, S. G. Geunes-Boyer, J. R. Wright, S. Diezmann, A. Allen, J. E. Stajich, F. S. Dietrich, J. R. Perfect, and J. Heitman. 2005. Same-sex mating and the origin of the Vancouver Island Cryptococcus gattii outbreak. Nature 437(7063):1360–1364.
Georgi, A., M. Schneemann, K. Tintelnot, R. C. Calligaris-Maibach, S. Meyer, R. Weber, and P. P. Bosshard. 2009. Cryptococcus gattii meningoencephalitis in an immunocompetent person 13 months after exposure. Infection 37(4):370–373.
Giles, S. S., T. R. Dagenais, M. R. Botts, N. P. Keller, and C. M. Hull. 2009. Elucidating the pathogenesis of spores from the human fungal pathogen Cryptococcus neoformans. Infection and Immunity 77:3491–3500.
Goldman, D. L., H. Khine, J. Abadi, D. J. Lindenberg, La Pirofski, R. Niang, and A. Casadevall. 2001. Serologic evidence for Cryptococcus neoformans infection in early childhood. Pediatrics 107(5):E66.
Hagen, F., S. van Assen, G. J. Luijckx, T. Boekhout, and G. A. Kampinga. 2010. Activated dormant Cryptococcus gattii infection in a Dutch tourist who visited Vancouver Island (Canada): A molecular epidemiological approach. Medical Mycology 48(3):528–531.
Heitman, J. 2006. Sexual reproduction and the evolution of microbial pathogens. Current Biology 16(17):R711–R725.
———. 2010. Evolution of eukaryotic microbial pathogens via covert sexual reproduction. Cell Host and Microbe 8(1):86–99.
———. 2011. Cryptococcus from human pathogen to model yeast, edited by J. Heitman, T. R. Kozel, J. K. Kwon-Chung, J. R. Perfect, and A. Casadevall. Washington, DC: ASM Press. P. 646.
Hoang, L. M., J. A. Maguire, P. Doyle, M. Fyfe, and D. L. Roscoe. 2004. Cryptococcus neoformans infections at Vancouver Hospital and Health Sciences Centre (1997–2002): Epidemiology, microbiology and histopathology. Journal of Medical Microbiology 53(Pt 9):935–939e.
Hu, G., I. Liu, A. Sham, J. E. Stajich, F. S. Dietrich, and J. W. Kronstad. 2008. Comparative hybridization reveals extensive genome variations in the AIDS-associated pathogen Cryptococcus neoformans. Genome Biology 9:R41.
Hull, C. M., and J. Heitman. 2002. Genetics of Cryptococcus neoformans. Annual Review of Genetics 36:557–615.
Idnurm, A., Y. S. Bahn, K. Nielsen, X. Lin, J. A. Fraser, and J. Heitman. 2005. Deciphering the model pathogenic fungus Cryptococcus neoformans. Nature Reviews Microbiology 3(10):753–764.
Kaufman, L., and S. Blumer. 1978. Cryptococcosis: The awakening giant. Proceedings of the Fourth International Conference on the Mycoses: PAHO Scientific Publications No. 356. Pp. 176–184.
Kidd, S. E., F. Hagen, R. L. Tscharke, M. Huynh, K. H. Bartlett, M. Fyfe, L. Macdougall, T. Boekhout, K. J. Kwon-Chung, and W. Meyer. 2004. A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proceedings of the National Academy of Sciences, USA 101(49):17258–17263.
Kluger, E. K., H. K. Karaoglu, M. B. Krockenberger, P. K. Della Torre, W. Meyer, and R. Malik. 2006. Recrudescent cryptococcosis, caused by Cryptococcus gattii (molecular type VGII), over a 13-year period in a Birman cat. Medical Mycology 44(6):561–566.
Kronstad, J. W., R. Attarian, B. Cadieux, J. Choi, C. A. D’Souza, E. J. Griffiths, J. M. Geddes, G. Hu, W. H. Jung, M. Kretschmer, S. Saikia, and J. Wang. 2011. Expanding fungal pathogenesis: Cryptococcus breaks out of the opportunistic box. Nature Reviews Microbiology 9(3):193–203.
Kwon-Chung, K. J. 1975. A new genus, Filobasidiella, the perfect state of Cryptococcus neoformans. Mycologia 67:1197–1200.
———. 1976a. Morphogenesis of Filobasidiella neoformans, the sexual state of Cryptococcus neoformans. Mycologia 68 (4):821–833.
———. 1976b. A new species of Filobasidiella, the sexual state of Cryptococcus neoformans B and C serotypes. Mycologia 68(4):943–946.
Kwon-Chung, K. J., J. C. Edman, and B. L. Wickes. 1992. Genetic association of mating types and virulence in Cryptococcus neoformans. Infection and Immunity 60(2):602–605.
Lerner, C. W., and M. L. Tapper. 1984. Opportunistic infection complicating acquired immune deficiency syndrome: Clinical features of 25 cases. Medicine (Baltimore) 63(3):155–164.
Lin, X., C. M. Hull, and J. Heitman. 2005. Sexual reproduction between partners of the same mating type in Cryptococcus neoformans. Nature 434(7036):1017–1021.
Lindberg, J., F. Hagen, A. Laursen, J. Stenderup, and T. Boekhout. 2007. Cryptococcus gattii risk for tourists visiting Vancouver Island, Canada. Emerging Infectious Diseases 13(1):178–179.
Littman, M. L., and L. E. Zimmer. 1956. Cryptococcosis. New York: Grune & Stratton, Inc.
Litvintseva, A. P., R. E. Marra, K. Nielsen, J. Heitman, R. Vilgalys, and T. G. Mitchell. 2003. Evidence of sexual recombination among Cryptococcus neoformans serotype A isolates in sub-Saharan Africa. Eukaryotic Cell 2(6):1162–1168.
Litvintseva, A. P., X. Lin, I. Templeton, J. Heitman, and T. Mitchell. 2007. Many globally isolated AD hybrid strains of Cryptococcus neoformans originated in Africa. PLoS Pathogens 3(8):e114.
Litvintseva, A. P., I. Carbone, J. Rossouw, R. Thakur, N. P. Govender, T. G. Mitchell. 2011. Evidence that the human pathogenic fungus Cryptococcus neoformans var. grubii may have evolved in Africa. PLoS One 6(5):e19688.
Ma, H., F. Hagen, D. J. Stekel, S. A. Johnston, E. Sionov, R. Falk, I. Polacheck, T. Boekhout, and R. C. May. 2009. The fatal fungal outbreak on Vancouver Island is characterized by enhanced intracellular parasitism driven by mitochondrial regulation. Proceedings of the National Academy of Sciences, USA 106(31):12980–12985.
MacDougall, L., S. E. Kidd, E. Galanis, S. Mak, M. J. Leslie, P. R. Cieslak, J. W. Kronstad, M. G. Morshed, and K. H. Bartlett. 2007. Spread of Cryptococcus gattii in British Columbia, Canada, and detection in the Pacific Northwest, USA. Emerging Infectious Diseases 13(1):42–50.
McClelland, C. M., Y. C. Chang, A. Varma, and K. J. Kwon-Chung. 2004. Uniqueness of the mating system in Cryptococcus neoformans. Trends in Microbiology 12(5):208–212.
Mitchell, D. H., T. C. Sorrell, A. M. Allworth, C. H. Heath, A. R. McGregor, K. Papanaoum, M. J. Richards, and T. Gottlieb. 1995. Cryptococcal disease of the CNS in immunocompetent hosts: Influence of cryptococcal variety on clinical manifestations and outcome. Clinical Infectious Diseases 20(3):611–616.
Ngamskulrungroj, P., T. C. Sorrell, A. Chindamporn, A. Chaiprasert, N. Poonwan, and W. Meyer. 2008. Association between fertility and molecular sub-type of global isolates of Cryptococcus gattii molecular type VGII. Medical Mycology 46(7):665–673.
Ngamskulrungroj, P., F. Gilgado, J. Faganello, A. P. Litvintseva, A. L. Leal, K. M. Tsui, T. G. Mitchell, M. H. Vainstein, and W. Meyer. 2009a. Genetic diversity of the Cryptococcus species complex suggests that Cryptococcus gattii deserves to have varieties. PLoS ONE 4(6):e5862.
Ngamskulrungroj, P., U. Himmelreich, J. A. Breger, C. Wilson, M. Chayakulkeeree, M. B. Krockenberger, R. Malik, H. M. Daniel, D. Toffaletti, J. T. Djordjevic, E. Mylonakis, W. Meyer, and J. R. Perfect. 2009b. The trehalose synthesis pathway is an integral part of the virulence composite for Cryptococcus gattii. Infection and Immunity 77(10):4584–4596.
Nielsen, K., G. M. Cox, P. Wang, D. L. Toffaletti, J. R. Perfect, and J. Heitman. 2003. Sexual cycle of Cryptococcus neoformans var. grubii and virulence of congenic a and α isolates. Infection and Immunity 71(9):4831–4841.
Nielsen, K., G. M. Cox, A. P. Litvintseva, E. Mylonakis, S. D. Malliaris, D. K. Benjamin, Jr., S. S. Giles, T. G. Mitchell, A. Casadevall, J. R. Perfect, and J. Heitman. 2005a. Cryptococcus neoformans α strains preferentially disseminate to the central nervous system during coinfection. Infection and Immunity 73(8):4922–4933.
Nielsen, K., R. E. Marra, F. Hagen, T. Boekhout, T. G. Mitchell, G. M. Cox, and J. Heitman. 2005b. Interaction between genetic background and the mating-type locus in Cryptococcus neoformans virulence potential. Genetics 171(3):975–983.
Okagaki, L. H., A. K. Strain, J. N. Nielsen, C. Charlier, N. J. Baltes, F. Chretien, J. Heitman, F. Dromer, and K. Nielsen. 2010. Cryptococcal cell morphology affects host cell interactions and pathogenicity. PLoS Pathogens 6(6):e1000953.
Park, B. J., K. A. Wannemuehler, B. J. Marston, N. Govender, P. G. Pappas, and T. M. Chiller. 2009. Estimation of the current global burden of cryptococcal meningitis among persons living with HIV/AIDS. AIDS 23(4):525–530.
Pavelka, N., G. Rancati, J. Zhu, W. D. Bradford, A. Saraf, L. Florens, B. W. Sanderson, G. L. Hattem, and R. Li. 2010. Aneuploidy confers quantitative proteome changes and phenotypic variation in budding yeast. Nature 468(7321):321–325.
Perfect, J. R., W. E. Dismukes, F. Dromer, D. L. Goldman, J. R. Graybill, R. J. Hamill, T. S. Harrison, R. A. Larsen, O. Lortholary, M. H. Nguyen, P. G. Pappas, W. G. Powderly, N. Singh, J. D. Sobel, and T. C. Sorrell. 2010. Clinical practice guidelines for the management of cryptococcal disease: 2010 update by the Infectious Diseases Society of America. Clinical Infectious Diseases 50(3):291–322.
Petzold, E. W., U. Himmelreich, E. Mylonakis, T. Rude, D. Toffaletti, G. M. Cox, J. L. Miller, and J. R. Perfect. 2006. Characterization and regulation of the trehalose synthesis pathway and its importance in the pathogenicity of Cryptococcus neoformans. Infection and Immunity 74(10):5877–5887.
Pounden, W. D., J. M. Amberson, and R. F. Jaeger. 1952. A severe mastitis problem associated with Cryptococcus neoformans in a large dairy herd. American Journal of Veterinary Medicine 13(47):121–128.
Rancati, G., N. Pavelka, B. Fleharty, A. Noll, R. Trimble, K. Walton, A. Perera, K. StaehlingHampton, C. W. Seidel, and R. Li. 2008. Aneuploidy underlies rapid adaptive evolution of yeast cells deprived of a conserved cytokinesis motor. Cell 135(5):879–893.
Saul, N., M. Krockenberger, and D. Carter. 2008. Evidence of recombination in mixed-mating-type and alpha-only populations of Cryptococcus gattii sourced from single eucalyptus tree hollows. Eukaryotic Cell 7(4):727–734.
Selmecki, A., A. Forche, and J. Berman. 2006. Aneuploidy and isochromosome formation in drug-resistant Candida albicans. Science 313(5785):367–370.
Selmecki, A., M. Gerami-Nejad, C. Paulson, A. Forche, and J. Berman. 2008. An isochromosome confers drug resistance in vivo by amplification of two genes, ERG11 and TAC1. Molecular Microbiology 68(3):624–641.
Selmecki, A. M., K. Dulmage, L. E. Cowen, J. B. Anderson, and J. Berman. 2009. Acquisition of aneuploidy provides increased fitness during the evolution of antifungal drug resistance. PLoS Genetics 5(10):e1000705.
Simon, J., R. E. Nichols, and E. V. Morse. 1953. An outbreak of bovine cryptococcosis. Journal of the American Veterinary Medical Association 122(910):31–35.
Sionov, E., Y. C. Chang, H. M. Garraffo, and K. J. Kwon-Chung. 2009. Heteroresistance to fluconazole in Cryptococcus neoformans is intrinsic and associated with virulence. Antimicrobial Agents and Chemotherapy 53(7):2804–2815.
Sionov, E., H. Lee, Y. C. Chang, and K. J. Kwon-Chung. 2010. Cryptococcus neoformans overcomes stress of azole drugs by formation of disomy in specific multiple chromosomes. PLoS Pathogens 6(4):e1000848.
Speed, B., and D. Dunt. 1995. Clinical and host differences between infections with the two varieties of Cryptococcus neoformans. Clinical Infectious Diseases 21:28–34.
Steele, K. T., R. Thakur, R. Nthobatsang, A. P. Steenhoff, and G. P. Bisson. 2010. In-hospital mortality of HIV-infected cryptococcal meningitis patients with C. gattii and C. neoformans infection in Gaborone, Botswana. Medical Mycology 48(8):1112–1115.
Stephen, C., S. Lester, W. Black, M. Fyfe, and S. Raverty. 2002. Multispecies outbreak of cryptococcosis on southern Vancouver Island, British Columbia. Canadian Veterinary Journal 43(10):792–794.
Sukroongreung, S., K. Kitiniyom, C. Nilakul, and S. Tantimavanich. 1998. Pathogenicity of basidiospores of Filobasidiella neoformans var. neoformans. Medical Mycology 36:419–424.
Thompson, G. R., III, N. P. Wiederhold, A. W. Fothergill, A. C. Vallor, B. L. Wickes, and T. F. Patterson. 2009. Antifungal susceptibilities among different serotypes of Cryptococcus gattii and Cryptococcus neoformans. Antimicrobial Agents and Chemotherapy 53(1):309–311.
Torres, E. M., T. Sokolsky, C. M. Tucker, L. Y. Chan, M. Boselli, M. J. Dunham, and A. Amon. 2007. Effects of aneuploidy on cellular physiology and cell division in haploid yeast. Science 317(5840):916–924.
Torres, E. M., N. Dephoure, A. Panneerselvam, C. M. Tucker, C. A. Whittaker, S. P. Gygi, M. J. Dunham, and A. Amon. 2010. Identification of aneuploidy-tolerating mutations. Cell 143(1):71–83.
Torres-Rodriguez, J. M., E. Alvarado-Ramirez, F. Murciano, and M. Sellart. 2008. MICs and minimum fungicidal concentrations of posaconazole, voriconazole and fluconazole for Cryptococcus neoformans and Cryptococcus gattii. Journal of Antimicrobial Chemotherapy 62(1):205–206.
Upton, A., J. A. Fraser, S. E. Kidd, C. Bretz, K. H. Bartlett, J. Heitman, and K. A. Marr. 2007. First contemporary case of human infection with Cryptococcus gattii in Puget Sound: Evidence for spread of the Vancouver Island outbreak. Journal of Clinical Microbiology 45(9):3086–3088.
Velagapudi, R., Y. P. Hsueh, S. Geunes-Boyer, J. R. Wright, and J. Heitman. 2009. Spores as infectious propagules of Cryptococcus neoformans. Infection and Immunity 77(10):4345–4355.
Vieira, J., E. Frank, T. J. Spira, and S. H. Landesman. 1983. Acquired immune deficiency in Haitians: Opportunistic infections in previously healthy Haitian immigrants. New England Journal of Medicine 308(3):125–129.
Wang, L., and X. Lin. 2011. Mechanisms of unisexual mating in Cryptococcus neoformans. Fungal Genetics and Biology 48(7):651–660
West, S. K., E. J. Byrnes, S. Mostad, R. Thompson, R. Barnes, et al. 2008. Emergence of Cryptococcus gattii in the Pacific Northwest United States. Paper presented at the 48th meeting of ICAAC/IDSA.
Xue, C., Y. Tada, X. Dong, and J. Heitman. 2007. The human fungal pathogen Cryptococcus can complete its sexual cycle during a pathogenic association with plants. Cell Host and Microbe 1(4):263–273.
Xue, C., T. Liu, L. Chen, W. Li, I. Liu, J. W. Kronstad, A. Seyfang, and J. Heitman. 2010. Role of an expanded inositol transporter repertoire in Cryptococcus neoformans sexual reproduction and virulence. mBio 1(1):e00084–10.
Zimmer, B. L., H. O. Hempel, and N. L. Goodman. 1984. Pathogenicity of the basidiospores of Filobasidiella neoformans. Mycopathologia 85(3):149–153.
YEAST INFECTIONS—HUMAN GENETICS ON THE RISE38
Steven M. Holland39and Donald C. Vinh39
We lead inextricably mycotic lives: yeasts leaven our bread, ferment our wine and beer, and inhabit our skins, mouths, and gastrointestinal tracts; however, not all is harmony. Hippocrates reported aphthous ulcers consistent with thrush in patients with severe debilitation, but it was not until the 1840s that in the newly emerging field of clinical experimental medicine that thrush—as well as other mycotic conditions, including ringworm—was recognized as being caused by transmissible fungi. In 1923 Christine Marie Berkhout named what we now call Candida albicans, for the white robe, toga candida, worn by Roman senators and senatorial candidates (Emmons et al., 1977). Fungal infections in general, and candida infections in particular, are important markers of innate or acquired immune dysfunction. However, despite the estimated 1.5 million species of fungi in the world, precious few cause human disease, and those that do (in the
_____________________
38 Originally published as Holland, Steven M; Vinh, Donald C.2009. Yeast Genetics on the Rise. New England Journal of Medicine. Article DOI: 10.1056/NEJMe0907186. Copyright © 2009 Massachusetts Medical Society. Available at: http://www.nejm.org/doi/full/10.1056/NEJMe0907186.
39 Steven M. Holland, M.D., and Donald C. Vinh, M.D. From the Laboratory of Clinical Infectious Diseases, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD.
absence of iatrogenic factors) usually cause inapparent or mucocutaneous infection. Therefore, genetic factors in both the pathogen and the host must be key to an understanding of who gets disease and why.
Despite the ubiquity and importance of fungi, little is known about specific human genetic predispositions to them. There are three distinct categories of fungi: filamentous molds (e.g., aspergillus, mucor, and trichophyton), dimorphic fungi (e.g., the endemic mycoses, histoplasma, blastomyces, coccidioides, and sporothrix, a non-endemic pathogen), and yeasts (e.g., candida, cryptococcus, and trichosporon). In the absence of iatrogenic factors, visceral infections of filamentous mold occur almost exclusively in patients with chronic granulomatous disease, whose capacity to generate phagocyte superoxide is defective. Severe invasive aspergillosis develops in these patients (who have normal lungs), as do, on occasion, infections with invasive candida; interestingly, mucocutaneous candidiasis is not encountered in chronic granulomatous disease. In contrast, development of most other cases of pulmonary aspergillosis that are unrelated to immunosuppression requires previous airway damage, such as bronchiectasis or bullous disease. Dimorphic fungi cause inapparent or relatively mild disease in the vast number of people who are otherwise normal. Severe pneumonia and disseminated disease occur infrequently, as in cases of advanced human immunodeficiency virus (HIV) and in patients with defects of the interferon-γ–interleukin-12 axis, indicating that intracellular killing plays a crucial role in the body’s defense against these infections. Mucocutaneous candidiasis, vaginal candidiasis, thrush, and onychomycosis occur in a variety of disparate conditions, ranging from diabetes mellitus to the autoimmune polyendocrinopathy–candidiasis–ectodermal dystrophy (APECED) syndrome, to HIV, to Job’s (hyper-IgE) syndrome. Our recent identification of mutations in signal transducer and activator of the transcription 3 gene (STAT3) as the cause of Job’s syndrome unexpectedly provides a link to the important articles by Ferwerda et al. (2009) and Glocker et al. (2009) in this issue of the Journal.
Whereas mucocutaneous candidiasis has long been recognized as a consequence of profound lymphocyte dysfunction, or lymphopenia, it occurs in Job’s syndrome in patients with a relatively normal number of functioning lymphocytes. Mutations in the gene encoding transcription factor STAT3 impair numerous pathways, including the generation of interleukin-17–committed T lymphocytes (Th17 cells). Th17 cells fill a gap in previous knowledge about lymphocyte-mediated killing: If Th1 (interferon-γ–producing) lymphocytes are controlling intracellular infection and Th2 (interleukin-4–producing) lymphocytes are directing antibody-mediated protection against extracellular infection, which cells protect the exposed mucosa and epithelium? Among the products of Th17 cells is interleukin-22, which synergizes with interleukin-17 in the epithelial synthesis of cationic antimicrobial peptides, such as defensins. STAT3 is also central to the induction and signaling of itself, interleukin-17, and interleukin-22 (Fig. A11-1) (Minegishi et al., 2009).
We encounter most microbes first at epithelial surfaces through cell-
Figure A11-1 (facing page). Mechanisms of Fungal Sensing and Control. The β-glucan on the budding yeast forms of candida bind to dectin-1 on epithelial cells and phagocytes, leading to activation of the CARD9 signaling complex (along with Bcl-10–MALT1) (Panel A). This in turn produces cytokines that help drive CD4+ T lymphocytes toward the Th17 phenotype, a STAT3-dependent process. Th17 lymphocytes elaborate interleukin-17 (augmenting neutrophil production and recruitment) and interleukin-22, which are synergic in the STAT3-dependent production of antimicrobial peptides by epithelial cells. The generation of superoxide by the phagocytic NADPH oxidase system is crucial for the killing of filamentous molds such as aspergillus (Panel B). Interactions between macrophages and lymphocytes are required for the control of intracellular infections such as the dimorphic fungi. In response to fungal infection, macrophages release interleukin-12, which acts on its cognate receptor on T lymphocytes and natural killer (NK) lymphocytes. Interleukin-12–stimulated lymphocytes release interferon-γ, which acts on macrophages through STAT1 to kill intracellular fungi. Interferon-γ also augments tumor necrosis factor α (TNF-α), which acts in part through the nuclear factor κB (NF-κB) to kill fungi and drive inflammatory responses. Molecules with identified mutations relevant to fungal susceptibility are shown in red.
receptors. These receptors couple the binding of specific microbial components to signal transduction pathways, which results in local and systemic immune responses, including activation of the central activator of inflammation, nuclear factor κB. Specific immunodeficiencies involving receptor molecules and their signaling have been characterized for herpes simplex virus encephalitis, Neisseria meningitidis, and Streptococcus pneumoniae (Beutler, 2009). As expected, the closer the defect is to the initial contact with the pathogen, the more likely it is that the susceptibility will be microbiologically narrow. In contrast, defects that occur at the convergence of several pathways tend to be broader and more severe. Dectin-1 is the cell-surface receptor for β-glucan, a major component of the budding yeast-cell wall, and its signaling travels through a series of molecules, including caspase-recruitment–domain (CARD) protein 9, leading to activation of nuclear factor κB.
Ferwerda et al. identified a Dutch family with impaired in vitro responses to β-glucan. The spectrum of disease was limited to nails and mucosa. Interestingly, the ages at clinical disease presentation were 10 to 12 years for the homozygous daughters but 40 and 55 years for the heterozygous mother and father, respectively, suggesting both hormonal and gene-dose effects. Generation of interleukin-6 and interleukin-17 was impaired only in response to the dectin-1 ligand, β-glucan. The specific stop codon the authors identified in dectin-1 is remarkably common in some parts of Africa and Europe (allele frequency, 3 to 7%), suggesting that unrecognized forces maintain it in populations. Glocker et al. identified an extended Iranian family with predominantly mucocutaneous but also fatal candidiasis of the central nervous system caused by mutations in the critical dectin-1 signal transduction molecule, CARD9, impairing both dectin-1 signaling and Th17 production. CARD9 also funnels a variety of other cell-
surface and intra-cellular signals, including the p38 mitogen-activated protein kinase (MAPK) and Jun N-terminal kinase (JNK) pathways, possibly accounting for its greater clinical severity as compared with isolated dectin-1 deficiency. The CARD9 mutation appears to be rare, and its rarity is commensurate with its severity.
Important caveats concerning both reports are that the key manifestations are mucocutaneous, whereas the functional studies were performed on leukocytes. It remains to be proven whether the key mechanisms in these cases of severe candidiasis consist of impaired dectin-1 signaling at the epithelial level or impaired leukocyte activation of epithelium, mediated through cytokines such as interleukin-17.
The long and largely happy coexistence of humans and fungi has necessitated the existence of ways to detect and control fungi, keeping them in their place. Apparently, these pathways are different for candida, aspergillus, and mucor. Although we do not yet understand the entire conversation between leukocytes and epithelium, with these two reports we have now overheard some of the key words that will enable us to listen more thoughtfully.
No potential conflict of interest relevant to this article was reported.
References
Beutler BA. TLRs and innate immunity. Blood 2009;113:1399–407.
Emmons CW, Binford CH, Utz JP, Kwon-Chung KJ. Medical mycology. 3rd ed. Philadelphia: Lea & Febiger, 1977.
Ferwerda B, Ferwerda G, Plantinga TS, et al. Human dectin-1 deficiency and mucocutaneous fungal infections. N Engl J Med 2009;361:1760–7.
Glocker E-O, Hennigs A, Nabavi M, et al. A homozygous CARD9 mutation in a family with susceptibility to fungal infections. N Engl J Med 2009;361:1727–35.
Minegishi Y, Saito M, Nagasawa M, et al. Molecular explanation for the contradiction between systemic Th17 defect and localized bacterial infection in hyper-IgE syndrome. J Exp Med 2009;206:1291–301.
THE INCREASED RISK OF GLOBAL WHEAT RUST PANDEMICS: PUTTING YELLOW RUST INTO PERSPECTIVE
Mogens Støvring Hovmøller40
Infectious diseases of humans are receiving great attention due to their direct and immediate impact on human mortality (King et al., 2006; WHO, 2008),
______________
40 Aarhus University, Faculty of Agricultural Sciences, Department of Integrated Pest Management, Flakkebjerg, 4200 Slagelse, Denmark. Mogens.hovmoller@agrsci.dk.
whereas the diseases of crop plants are subject to much less publicity although they are threatening crop productivity, food security, and thereby the livelihood of billions of people around the world. Currently, wheat rust fungi are among the top 10 constraints for food production in many parts of the world, such as sub-Saharan Africa, East Asia, and South Asia (FAO, 2010; Waddington et al., 2010), mainly due to their epidemic potential and transboundary spread by wind (Brown and Hovmøller, 2002).
According to U.S. Department of Agriculture statistics, global wheat production was generally lower in 2010 compared to the previous year (USDA, 2011). The reduced productivity was caused by several factors, such as unfavorable weather conditions, resulting in large-scale flooding, droughts, and bushfires in different parts of the world, as well as severe yellow rust epidemics (USDA, 2010).
Rust diseases have been recognized as being harmful to wheat since ancient Greece. Theophrastus of Eressus (371–286 BC), the founder of botany and a pupil of Aristoteles, noted in his book ΠΕΡΙ ΦΥΤΩΝ ΑΙΤΙΩΝ (The Causes of Plants) that some plants of barley were more susceptible to the rusts than others, the most susceptible denoted “Achilles Barley,” and he recognized the importance of sowing in due time to reduce rust diseases on different crop plants such as wheat, barley, peas, and beans (Theophrastus, 1990). Wheat rust may be caused by three plant pathogenic fungi, Puccinia graminis (Leonard and Szabo, 2005), Puccinia triticina (Bolton et al., 2008), and Puccinia striiformis (Hovmøller et al., 2011). The corresponding diseases are termed black (stem) rust, brown (leaf) rust, and yellow (stripe) rust, respectively (synonymous terms in brackets) (Figure A12-1).
The three wheat rust fungi are biotrophic and heteroaecious, meaning they generally require a primary wheat host for asexual reproduction and an alternate host to complete sexual reproduction. Berberis spp. have been known to serve as an alternate host for stem rust for more than two centuries. For instance, Schøler (1818), who was a school teacher in Denmark, demonstrated a link between stem rust on barberry and some cereals via repeated infection experiments. His results started a so-called “barberry quarrel,” where the usefulness of barberry eradication programs was discussed. Later, de Bary (1865, 1866) demonstrated the complete life cycle of the stem rust fungus. However, the problems related to barberry adjacent to cereal fields were observed by farmers in Europe much earlier, with the first local laws against barberry being implemented in France in the 17th century and in the Americas in the 18th century (Roelfs, 1982). Surprisingly, the discovery of barberry as an alternate host for yellow rust infecting wheat and Poa grass, respectively, was not made until very recently (Jin et al., 2010). The alternate host of P. triticina depend on the primary host: Isolates from cultivated wheat and wild emmer have Thalictrum speciosissimum (in the Ranunculaceae) as alternate host, whereas several species in the Boraginaceae, such as Anchusa aggregata, Anchusa italica, Echium glomeratum, and Lycopsis arvensis, may

Figure A12-1 Typical macroscopic symptoms of rust infections on adult wheat plants. Puccinia striiformis Westend (yellow/stripe rust) [left photo], Puccinia triticina (brown/leaf rust) [center photo], and Puccinia graminis tritici (black/stem rust) [right photo].
serve as alternate host for P. triticina from wild wheat and rye (Anikster et al., 1997). Although this paper deals with all three wheat rusts, the primary focus is yellow rust, which has spread at unprecedented scales in recent years and caused severe epidemics even in areas where the disease was previously non-significant or absent (Hovmøller et al., 2010).
Yellow Rust Epidemiology
Wheat rusts are highly epidemic on susceptible cultivars in a rust-favorable environment. In the past, yellow rust was considered most harmful in cool and wet climates, whereas leaf rust and stem rust epidemics were favored by warmer temperatures. They are all characterized by passive spreading of airborne urediniospores carried by the wind, potentially across hundreds of kilometers (Hovmøller et al., 2002; Kolmer, 2005; Leonard and Szabo, 2005; Zadoks, 1961). Since 2000, epidemics of yellow rust in particular have accelerated in many areas (Hovmøller et al., 2010). In the United States, annual losses due to wheat yellow
rust exceeded 1 million metric tons over several years, and in each of the years 2003 and 2010 they mounted to 2.4 million tons (Long, 2000–2010), despite increased and widespread use of agrochemicals already in 2003 (Chen, 2005). In China, yellow rust is considered the most damaging disease on wheat, which is grown on more than 20 million hectares (Wan et al., 2004). In three yellow rust epidemic years, annual losses varied between 1.8 and 6 million tons; in 2002, losses of 1.3 million tons were recorded and 1.9 million tons were saved by treating at least 6 million hectares with fungicides (Wan et al., 2004). Yellow rust epidemics reached record levels in northern Africa in 2009 (Ezzahiri et al., 2009) and also in central and western Asia (Mboup et al., 2009), where more than 90 percent of important wheat varieties were susceptible to the disease (Sharma et al., 2009). Areas particularly affected by yellow rust epidemics in 2009 and 2010 are illustrated in Figure A12-2.
New genetic variability of the pathogen, resulting from natural population dynamic forces such as mutation, recombination, and migration, may be followed by host-induced selection, which is a major driver of changes in gene and genotypic frequencies of biotrophic plant pathogens (Hovmøller et al., 1997).
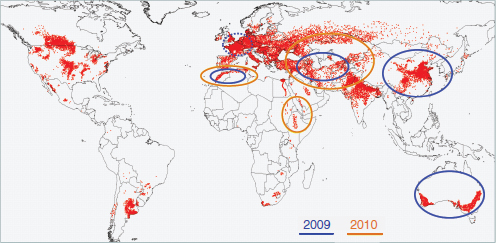
Figure A12-2 Map indicating the distribution of global wheat production and regions of recent yellow rust epidemics. Each red dot is equivalent to 20,000 tons of wheat grain production. The blue and orange circles indicate some of the areas in which severe yellow rust epidemics on wheat were reported in 2009 and 2010. Yellow rust severity varied greatly from field to field within the highlighted areas, depending on susceptibility of actual host cultivar, local environmental conditions, pathogen race distribution, and disease management practices. The epidemics in Northwest Europe in 2009 (dotted circle) were restricted to relatively few cultivars of wheat and triticale.
SOURCE: Adapted from Trethowan et al. (2002). Dr. Dave Hodson, Food and Agriculture Organization of the United Nations, Rome, is acknowledged for kind permission to reprint the map.
In agricultural systems, where crop cultivars with similar or identical resistance specificities are cultivated across large areas, the role of selection is greatly enhanced (Bayles et al., 2000; Hovmøller et al., 1997; McDonald and Linde, 2002). Therefore, crop varieties that are resistant when first deployed in agriculture may become susceptible after some years in cultivation. This phenomenon, which is often referred to as the boom-and-bust cycle, has been reported in numerous cases for the wheat rusts (e.g., Bayles et al., 2000; Chen, 2005; Enjalbert et al., 2005; Hovmøller, 2001; Wellings, 2007). The boom-and-bust cycle is closely related to resistance genes in the host and two categories of pathogen traits, virulence and aggressiveness.
Virulence and Aggressiveness
The yellow rust fungus is specialized on the cereal host at both species and cultivar levels (Hovmøller et al., 2011). The former refers to variation in the infection and reproduction capacity of the pathogen on different host genera (e.g., wheat, barley, rye, tricicale), whereas the second level of specialization occurs within the host genus, that is, at the cultivar level. The latter is typically based on the interaction between host resistance gene products and pathogen avirulence gene products in a gene-for-gene relationship (Flor, 1956). Mutation in the avirulence gene may result in virulence, which can be defined as the qualitative ability of the pathogen to cause disease by compromising a matching resistance gene in the host. In case the host cultivar possesses more than a single resistance gene, the virulence phenotype, representing a specific combination of different virulences, becomes critical in determining the fate of host susceptibility (e.g., Hovmøller and Henriksen, 2008; Johnson, 1992; Line and Qayoum, 1992). The virulence phenotype can be resolved from the outcome of a race analysis, which is carried out by inoculating a rust isolate onto a set of host differential lines carrying known resistance genes, for details (see, e.g., Hovmøller and Justesen, 2007a).
The study of race dynamics in P. striiformis populations has often been an important part of early warning and yellow rust strategies in Europe (Hovmøller and Henriksen, 2008; Johnson, 1992), North America (Chen and Penman, 2005; Line and Qayoum, 1992), China (Wan et al., 2007), and Australia (Wellings, 2007). Since 1967, where a national race survey was established in the United Kingdom (Johnson, 1992), similar programs were established in Germany (Flath and Bartels, 2002), France (De Vallavieille-Pope et al., 1990), and Denmark (Hovmøller, 2001). In recent years, European data on P. striiformis virulence dynamics have been made publicly available via the Eurowheat database at www.eurowheat.org, which also describes options for disease control, such as cultural practices and fungicide efficacies (Jørgensen et al., 2010). Systematic race surveys were initiated in the United States in the late 1960s after a series of severe yellow rust epidemics on wheat (Line and Qayoum, 1992). Systematic race surveys have also been implemented in China (Chen et al., 2009; Wan et al., 2004),
India (Prashar et al., 2007), Australia (Wellings, 2007; Wellings and McIntosh, 1990), and South Africa (Boshoff et al., 2002).
In addition to the virulence phenotype, pathogen aggressiveness, which is a quantitative measure of the ability of a virulent isolate to cause disease on a susceptible host plant, is a key determinant for the rate of pathogen spread and evolution. Some of the recent epidemics since 2000 have been ascribed to the emergence of yellow rust strains with increased aggressiveness and tolerance to warm temperatures (Hovmøller et al., 2008). In this context, strain was defined as a group of P. striiformis isolates sharing both molecular marker phenotype and virulence phenotype. For instance, Milus et al. (2009) demonstrated a significantly increased aggressiveness at both high and low temperatures, compared with isolates of pre-2000 races from North America and Europe (Milus et al., 2009). At the low-temperature regime, gradually changing from 10°C (night) to 18°C (day), the aggressive strains produced more than 70 percent more spores per infected leaf area compared to reference isolates from before 2000. At the high-temperature regime (12°C at night to 28°C during the day), isolates of the aggressive strains produced approximately 150 percent more spores per infected leaf area. Milus and colleagues (2009) concluded that the aggressive strain had most likely enhanced the yellow rust epidemics in North America and contributed to the spread of yellow rust into areas that were previously considered too warm for yellow rust epidemics. The aggressive strain detected in North America was indistinguishable from a strain detected as exotic incursion in Australia in 2002 (Hovmøller et al., 2008; Wellings et al., 2003). In Europe, a nearly identical strain was first detected in 2000, whereas first appearance of this strain in the Red Sea area and in western and central Asia is unknown (Hovmøller et al., 2008). According to time and area, these observations may be comparable to the famous panglobal spread of the Irish potato famine fungus Phytophthora infestans (Goodwin et al., 1994).
The distinction between virulence and aggressiveness may be subject to controversy because virulence toward a resistance gene with minor effects may be manifest by increased disease progress and spore production. These are the same variables used to measure aggressiveness. However, in case of virulence, there should be evidence for a significant host–pathogen interaction, whereas in the case of aggressiveness, the host–pathogen interaction is ideally non-significant. This implies that for a range of host cultivars with varying levels of rust susceptibility, they should rank the same for aggressive and non-aggressive isolates.
The Need for Coordinated Global Action
The number of foreign incursions of wheat rust races has increased significantly in recent years. Park et al. (2010) reported nine foreign incursions of wheat stem rust and wheat leaf rust into Australia between 1925 and 2005. Of these seven had occurred since 1969. Hovmøller et al. (2011) documented at least six
exotic incursions of yellow rust at continental scales since 1979, often resulting in the spread of yellow rust epidemics to new regions or continents. Scherm and Coakley (2003) noted that the rate of exotic pathogen invasion in the United States had increased from about five instances per decade from 1940 to 1970 to more than three times that number during the 1990s. This increasing rate of incursions of the yellow rust fungus may partly be ascribed to the emergence of new, highly aggressive rust strains combined with increasing human travel and commerce. However, regardless of the reasons for the increasing spread, the situation escalates, emphasizing the relevance of pathogen surveys covering larger areas and a need for global coordination. The use of crop cultivars with similar or identical resistance specificities across large wheat-growing areas supports this conclusion.
The accelerating wheat yellow rust epidemics requires coordinated multinational action to complement the ongoing initiatives of the Borlaug Global Rust Initiative (www.globalrust.org) to fight the multivirulent strain of wheat stem rust, known as Ug99 (Northoff, 2007, 2008). The speed by which threatening new strains can be diagnosed and reported widely is essential, and the precision by which traits such as virulence and aggressiveness are diagnosed is another bottleneck (Hovmøller et al., 2011). Precise diagnosis of both traits require pure and correct identification of the seed stocks of standard differential wheat varieties used for the race assays, genetically pure pathogen isolates, and well-defined experimental conditions for temperature, humidity, light, and plant nutrition, as well as trained staff who can assess and interpret the virulence phenotype and aggressiveness. The acknowledgment of these challenges led to the establishment of a Global Rust Reference Center (GRRC) in 2008 targeting yellow rust, supported by the International Center for Agricultural Research in the Dry Areas (ICARDA), the International Maize and Wheat Improvement Center (CIMMYT), and Aarhus University (Denmark) (Hovmøller et al., 2010). GRRC is accessible for receiving yellow rust samples on a year-round basis, which is a major advance complementing the capacities of existing regional and national rust diagnostic laboratories. At the Borlaug Global Rust Initiative (BGRI) Coordination Meeting in Syria in September 2009 (Dold, 2009), a proposal was made to extend the activities of GRRC to all wheat rust fungi, that is, yellow rust, brown rust, and stem rust.
The long-term aims of a Global Rust Reference Center would be the tracking of new wheat rust incursions and assessment of ongoing pathogen evolution at global scales, supplying data to online early-warning systems for farmers, breeders, and policy makers, like the current “Rust Spore” monitoring system run by the Food and Agriculture Organization of the United Nations (FAO, 2010). Another main activity would be to assist in training of students and junior scientists in rust pathology, and maintenance and extension of a unique world wheat rust collection to facilitate resistance breeding efforts. The activities should complement ongoing regional and national survey activities and research efforts
by BGRI partners, national counterparts, and advanced research institutions, resulting in stronger global efforts to mitigate the consequences of the inherent pathogen dynamics. At present, the activities at GRRC are limited to race analysis of relatively few pathogen samples per year in addition to training activities. To become sustainable in the long term, GRRC must contain a considerable portfolio of activities to reduce vulnerability due to potential changes in staff, management, and political awareness.
The identification of sources of resistance from cultivated and wild crop relatives to wheat is probably the most urgent task to reduce the threats posed by the wheat rusts, including yellow rust. Plants have an immunity system with several layers. Race-specific resistance provides protection only during one or few steps of the infection process and is therefore considered vulnerable to pathogen evolution whereas partial resistance, based on several or many sources of resistance, is considered more durable. Nevertheless, breeding for increased partial and non-host resistance protecting wheat against a broad spectrum of pathogen species and races can be done by large-scale field screening of thousands of breeding lines by applying targeted pathogen species and strains.
Prospects for New Research
The serious impacts and characteristics of wheat yellow rust epidemics should remind society about the vulnerability of global food production and the need to understand the interactions between agricultural crops, their pathogens, and the environment. Until recently, investigations of the basic biological and molecular mechanisms of yellow rust evolution have been hampered by the biotrophic lifestyle of yellow rust, precluding the use of molecular tools used for fungi that can be cultured on artificial media, and by the lack of an experimental system for genetic studies. These limitations may soon be overcome by the significant advances of molecular technologies, new insights into pathogen biology, and the cloning of host resistance genes, which were achieved in recent years. Molecular markers are probably the best established tools for resolving recent as well as past dispersal and evolutionary events in P. striiformis (Hovmøller and Justesen, 2007b; Hovmøller et al., 2008; Mboup et al., 2009). Next generation sequencing is currently the most promising tool for the identification of genetic changes related to virulence in fungal pathogens (Stergiopoulos and de Wit, 2009). A Danish–British genome sequencing initiative was started in 2009 (Walter et al., 2009), followed by a more comprehensive U.S.-led initiative (Broad Institute, 2009) aiming to resolve the whole genome sequence of P. striiformis. Such knowledge will most certainly speed up the discovery of genes involved in pathogenicity. The recent discovery of barberry as sexual host for P. striiformis (Jin et al., 2010), which has solved a century-old mystery about the yellow rust lifecycle, represents another major breakthrough that can serve as the basis for the development of an experimental system allowing classical genetic studies.
Functional characterization of isolated rust genes is still a great challenge because of the biotrophic lifestyle, which is hampering genetic transformation of P. striiformis. The identification of sources of resistance in wheat, especially for resource-poor areas in Africa and Asia, is probably the most urgent task to reduce the threats posed by wheat rust pathogens. However, this will require a long-term commitment and combined efforts of breeders, pathologists, and biologists to keep pace with the wheat rusts, which can evolve rapidly to compromise genetic resistance of the wheat host (Hovmøller and Justesen, 2007b; Wellings and McIntosh, 1990).
Concluding Remarks
The fight against infectious crop diseases has become a collective responsibility and requires a collective investment with a global, long-term political and scientific commitment. Such a global network will need to establish the physical and human resources to support the progress of wheat rust management technologies, that is, pathogen monitoring and resistance breeding, training programs for farmers and pathologists, and scientific progress. Hence a major priority of a global network should be to link researchers and plant breeders (1) with each other and with appropriate partners in resource-poor countries that are affected most by sudden wheat rust epidemics, and (2) with political authorities to allow for rapid actions. The establishment of the BGRI in 2006, inspired by Dr. Norman Borlaug, the pioneer of the Green Revolution and the front leader of cereal rust resistance breeding, represents a major milestone in this respect. The real challenge is to ensure sustained activities beyond ongoing, short-term projects. In the long term, surveillance and prevention measures of wheat rust epidemics will only be successful if performed on a coordinated, global scale. Borlaug used to say: “Rust never sleeps”—and events of recent years have shown how right he was.
References
Bayles, R. A., K. Flath, M. S. Hovmøller, and C. de Vallavieille-Pope. 2000. Breakdown of the Yr17 resistance to yellow rust of wheat in northern Europe. Agronomie 20(7):805–811.
Bolton, M. D., J. A. Kolmer, and D. F. Garvin. 2008. Wheat leaf rust caused by Puccinia triticina. Molecular Plant Pathology 9(5):563–575.
Boshoff, W. H. P., Z. A. Pretorius, and B. D. van Niekerk. 2002. Establishment, distribution, and pathogenicity of Puccinia striiformis f. sp tritici in South Africa. Plant Disease 86(5):485–492.
Broad Institute. 2009. Genome sequencing of wheat stripe rust and comparative genomics of Puccinia spp. U.S. Department of Agriculture. http://www.reeis.usda.gov/web/crisprojectpages/219263.html. (accessed March 28, 2011).
Brown, J. K. M., and M. S. Hovmøller. 2002. Aerial dispersal of pathogens on the global and continental scales and its impact on plant disease. Science 297(5581):537–541.
Chen, W. Q., L. R. Wu, T. G. Liu, S. C. Xu, S. L. Jin, Y. L. Peng, and B. T. Wang. 2009. Race dynamics, diversity, and virulence evolution in Puccinia striiformis f. sp tritici, the causal agent of wheat stripe rust in China from 2003 to 2007. Plant Disease 93(11):1093–1101.
Chen, X. M. 2005. Epidemiology and control of stripe rust Puccinia striiformis f. sp tritici on wheat. Canadian Journal of Plant Pathology-Revue Canadienne De Phytopathologie 27 (3):314–337.
Chen, X., and L. Penman. 2005. Stripe rust epidemic and races of Puccinia striiformis in the United States in 2004. Phytopathology 95(6):S19.
de Bary, A. 1865. Neue Untersuchungen über die Uredineen, insbesondere die Entwicklung der Puccinia. Monatsbericht der Koeniglich-Preussischen Akademie der Wissenschaften zu Berlin. 15–49.
———. 1866. Neue Untersuchungen über Uredineen, Zweite Mittheilung. Monatsberichten der Akademie der Wissenschaften zu Berlin. 205–215.
De Vallavieille-Pope, C., H. Picardformery, S. Radulovic, and R. Johnson. 1990. Specific resistance factors to yellow rust in seedlings of some French wheat varieties and races of Puccinia striiformis Westend in France. Agronomie 10(2):103–113.
Dold, M. 2009. Top wheat experts call for scaling up efforts to combat Ug99 and other wheat rusts, 11 September 2009. http://www.eurekalert.org/pub_releases/2009-09/bc-twe091009.php (accessed March 28, 2011).
Enjalbert, J., X. Duan, M. Leconte, M. S. Hovmøller, and C. De Vallavieille-Pope. 2005. Genetic evidence of local adaptation of wheat yellow rust (Puccinia striiformis f. sp tritici) within France. Molecular Ecology 14(7):2065–2073.
Ezzahiri, B., A. Yahyaoui, and M. S. Hovmøller. 2009. An analysis of the 2009 epidemic of yellow rust on wheat in Morocco. Paper presented at the Fourth Regional Yellow Rust Conference for Central and West Asia and North Africa, Antalya, Turkey, October 11–12, 2009.
FAO (Food and Agriculture Organization of the United Nations). 2010. WHEAT RUST—Threat to farmers and global food security. http://www.fao.org/agriculture/crops/core-themes/theme/pests/wrdgp/en/ (accessed March 28, 2011).
Flath, K., and G. Bartels. 2002. Virulence situation in Austrian and German populations of wheat yellow rust. Arbeitstagung 2001 der Vereinigung der Pflanzenzuchter und Saatgutkaufleute Osterreichs gehalten vom 20. bis 22. November 2001 in Gumpenstein, Irdning, Austria. 51–56. Verlag und Druck der Bundesanstalt ffn alpenllnstalt Landwirtschaft Gumpenstein.
Flor, H. H. 1956. The complementary genic systems in flax and flax rust. In Advances in Genetics, edited by M. Demerec. New York: Academic Press.
Goodwin, S. B., B. A. Cohen, and W. E. Fry. 1994. Panglobal distribution of a single clonal lineage of the Irish potato famine fungus. Proceedings of the National Academy of Sciences, USA 91(24):11591–11595.
Hovmøller, M. S. 2001. Disease severity and pathotype dynamics of Puccinia striiformis f. sp tritici in Denmark. Plant Pathology 50(2):181–189.
Hovmøller, M. S., and K. E. Henriksen. 2008. Application of pathogen surveys, disease nurseries and varietal resistance characteristics in an IPM approach for the control of wheat yellow rust. European Journal of Plant Pathology 121(3):377–385.
Hovmøller, M. S., and A. F. Justesen. 2007a. Appearance of atypical Puccinia striiformis f. sp tritici phenotypes in north-western Europe. Australian Journal of Agricultural Research 58 (6):518–524.
———. 2007b. Rates of evolution of avirulence phenotypes and DNA markers in a northwest European population of Puccinia striiformis f. sp tritici. Molecular Ecology 16:4637–4647.
Hovmøller, M. S., H. Ostergard, and L. Munk. 1997. Modelling virulence dynamics of airborne plant pathogens in relation to selection by host resistance in agricultural crops. In The gene-for-gene relationship in plant–parasite interactions, edited by I. R. Crute, E. B. Holub, and J. J. Burdan. Oxfordshire: CAB International.
Hovmøller, M. S., A. F. Justesen, and J. K. M. Brown. 2002. Clonality and long-distance migration of Puccinia striiformis f. sp tritici in North-West Europe. Plant Pathology 51(1):24–32.
Hovmøller, M. S., A. H. Yahyaoui, E. A. Milus, and A. F. Justesen. 2008. Rapid global spread of two aggressive strains of a wheat rust fungus. Molecular Ecology 17(17):3818–3826.
Hovmøller, M. S., S. Walter, and A. F. Justesen. 2010. Escalating threat of wheat rusts. Science 329 (5990):369–369.
Hovmøller, M. S., C. K. Sørensen, S. Walter, and A. F. Justesen. 2011. Diversity of Puccinia striiformis on cereals and grasses. Annual Review of Phytopathology 49. doi: 10.1146/annurev-phyto-072910-095230.
Jin, Y., L. J. Szabo, and M. Carson. 2010. Century-old mystery of Puccinia striiformis life history solved with the identification of Berberis as an alternate host. Phytopathology 100(5):432–435.
Johnson, R. 1992. Reflections of a plant pathologist on breeding for disease resistance, with emphasis on yellow rust and eyespot of wheat. Plant Pathology 41(3):239–254.
Jørgensen, L. N., M. S. Hovmøller, J. G. Hansen, P. Lassen, B. Clark, R. Bayles, B. Rodemann, M. Jahn, K. Flath, T. Goral, J. Czembor, P. du Cheyron, C. Maumene, C. de Pope, and G. C. Nielsen. 2010. EuroWheat.org—A support to integrated disease management in wheat. Outlooks on Pest Management 21(4):173–175.
King, D. A., C. Peckham, J. K. Waage, J. Brownlie, and M. E. J. Woolhouse. 2006. Infectious diseases: Preparing for the future. Science 313(5792):1392–1393.
Kolmer, J. A. 2005. Tracking wheat rust on a continental scale. Current Opinion in Plant Biology 8(4):441–449.
Leonard, K. J., and L. J. Szabo. 2005. Stem rust of small grains and grasses caused by Puccinia graminis. Molecular Plant Pathology 6(2):99–111.
Line, R. F., and A. Qayoum. 1992. Virulence, aggressiveness, evolution, and distribution of races of Puccinia striiformis (the cause of stripe rust of wheat) in North America, 1968–87. USDA Technical Bulletin 1788.
Long, D. L. 2000–2010. Small grain losses due to rust. http://www.ars.usda.gov/Main/docs.htm?docid=10123 (accessed June 17, 2011).
Mboup, M., M. Leconte, A. Gautier, A. M. Wan, W. Chen, C. de Vallavieille-Pope, and J. Enjalbert. 2009. Evidence of genetic recombination in wheat yellow rust populations of a Chinese over-summering area. Fungal Genetics and Biology 46(4):299–307.
McDonald, B. A., and C. Linde. 2002. Pathogen population genetics, evolutionary potential, and durable resistance. Annual Review of Phytopathology 40:349–379.
Milus, E. A., K. Kristensen, and M. S. Hovmøller. 2009. Evidence for increased aggressiveness in a recent widespread strain of Puccinia striiformis f. sp. tritici causing stripe rust of wheat. Phytopathology 99(1):89–94.
Northoff, E. 2007. Wheat killer spreads from East Africa to Yemen: New partnership formed to monitor and prevent spread of dangerous fungus. Food and Agriculture Organization of the United Nations (FAO). http://www.fao.org/newsroom/en/news/2007/1000537/index.html (accessed March 28, 2011).
———. 2008. Wheat killer detected in Iran: Dangerous fungus on the move from East Africa to the Middle East. Food and Agriculture Organization of the United Nations (FAO). http://www.fao.org/newsroom/en/news/2008/1000805/index.html (accessed March 28, 2011).
Park, R., T. Fetch, D. Hodson, Y. Jin, K. Nazari, M. Prashar, and Z. Pretorius. 2010. International surveillance of wheat rust pathogens—progress and challenges. http://www.globalrust.org/db/attachments/about/19/1/BGRI%20oral%20papers%202010.pdf (accessed March 28, 2011).
Prashar, M., S. C. Bhardwaj, S. K. Jain, and D. Datta. 2007. Pathotypic evolution in Puccinia striiformis in India during 1995–2004. Australian Journal of Agricultural Research 58(6):602–604.
Roelfs, A. P. 1982. Effects of barberry eradication on stem rust in the United States. Plant Disease 66(2):177–181.
Scherm, H., and S. M. Coakley. 2003. Plant pathogens in a changing world. Australasian Plant Pathology 32(2):157–165.
Schøler, N. P. 1818. En afhandling om Berberissens skadelige Virkning på Sæden. Landoec. Tid. 8, 289–336 (in Danish).
Sharma R. C., A. Amanov, Z. Khalikulov, C. Martius, Z. Ziyaev, S. Alikulov. 2009. Wheat yellow rust epidemic in Uzbekistan in 2009. Proceedings of the 4th Regional Yellow Rust Conference for Central and West Asia and North Africa, Antalya, Turkey, October 10–12.
Stergiopoulos, I., and P. J. G. M. de Wit. 2009. Fungal effector proteins. Annual Review of Phytopathology 47(1):233–263.
Theophrastus, E. 1990. De Causis Plantarum, books III IV, edited and translated by B. Einarson and G. K. K. Link. Cambridge (Massachusetts), London (England): Harvard University Press.
Trethowan, R. M., D. Hodson, H.-J. Braun, W. H. Pfeiffer, and M. van Ginkel. 2002. Wheat breeding environments. In Impacts of international wheat breeding research in the developing world, 1988–2002, edited by M. A. Lantican, H. J. Dubin, and M. L. Morris. Mexico, D.F.: CIMMYT.
USDA (U.S. Department of Agriculture). 2010. Middle East: Yellow rust epidemic affects regional wheat crops. http://www.pecad.fas.usda.gov/highlights/2010/06/Middle%20East/ (accessed March 28, 2011).
———. 2011. 2010/2011 wheat production. http://www.pecad.fas.usda.gov/ogamaps/default.cfm?cmdty=Wheat&attribute=Production (accessed March 28, 2011).
Waddington, S., X. Li, J. Dixon, G. Hyman, and M. de Vicente. 2010. Getting the focus right: Production constraints for six major food crops in Asian and African farming systems. Food Security 2(1):27–48.
Walter, S., E. Kemen, J. K. M. Brown, J. D. G. Jones, M. S. Hovmøller, and A. F. Justesen. 2009. Omics approaches to understand the nature of virulence in Puccinia striiformis f.sp. tritici. Paper presented at the 12th International Cereal Rusts and Powdery Mildews Conference, Antalya, Turkey, October 13–16, 2009.
Wan, A., Z. Zhao, X. M. Chen, Z. He, S. Jin, Q. Jia, G. Yao, J. Yang, B. Wang, and G. Li. 2004. Wheat stripe rust epidemic and virulence of Puccinia striiformis f. sp. tritici in China in 2002. Plant Diseases 88:896–904.
Wan, A. M., X. M. Chen, and Z. H. He. 2007. Wheat stripe rust in China. Australian Journal of Agricultural Research 58(6):605–619.
Wellings, C. R. 2007. Puccinia striiformis in Australia: A review of the incursion, evolution, and adaptation of stripe rust in the period 1979–2006. Australian Journal of Agricultural Research 58(6):567–575.
Wellings, C. R., and R. A. McIntosh. 1990. Puccinia striiformis f.sp. tritici in Australasia: Pathogenic changes during the first 10 years. Plant Pathology 39(2):316–325.
Wellings, C. R., D. G. Wright, F. Keiper, and R. Loughman. 2003. First detection of wheat stripe rust in western Australia: Evidence for a foreign incursion. Australasian Plant Pathology 32(2):321–322.
WHO (World Health Organization). 2008. The World Health Report 2008—primary health care (now more than ever), edited by T. Evans and W. Van Lerberghe. Geneva, Switzerland: WHO.
Zadoks, J. C. 1961. Yellow rust on wheat studies in epidemiology and physiologic specialization. European Journal of Plant Pathology 67(3):69–256.
FUNGAL PATHOGENESIS IN PLANTS AND ANIMALS: SIMILARITIES AND DIFFERENCES
Barbara Howlett41
Introduction
Although the vast majority of fungal species do not cause disease, ones that affect plants are responsible for significant economic losses (Skamnioti and Gurr, 2009). In the past couple of decades, fungal diseases of humans have become an increasing threat, especially for people who are immunologically compromised (Sexton and Howlett, 2006). Consequently there has been a rapid explosion in knowledge about fungal pathogenesis of animals and this has been taken up by people studying plant pathogenic fungi. Pathogenesis involves the interaction of two partners with input from the environment, a concept described as the “disease triangle” in plant pathology. The “damage-response” concept developed for animal pathogens emphasizes that the outcome of an interaction is determined by the amount of damage incurred on the host (Casadevall and Pirofski, 2003). These concepts are useful reminders of the complexity of the interaction and the interdependence of host and pathogen.
Tools to Study Fungal Pathogenesis
The large number of fungi with sequenced genomes and recent advances in genetic manipulation of fungi are leading to an improved understanding of mechanisms associated with disease. Comparative genomics is being used to identify candidate genes involved in disease. Amplification of particular gene families within a genome is consistent with lifestyles: For instance, many plant pathogens have large gene families that encode enzymes to degrade the cuticle and cell wall, which coats plant cells, while animal pathogens (e.g., Coccidioides immitis, the cause of valley fever) have gene families encoding enzymes that degrade proteins of the skin, which is usually the initial barrier to invasion (Sharpton et al., 2009). Transcriptomics can show which genes are turned on; microarrays have been used extensively and many datasets are publicly available (Cairns et al., 2010). Proteomic analysis of culture filtrates provides information about secreted proteins (secrotome) that are accessible to the host and thus may play a role in the interaction; this approach has been applied to identify such proteins of Leptosphaeria maculans, the blackleg pathogen of canola (Vincent et al., 2009).
______________
41 The University of Melbourne.
The generation of fungal mutants via tagged random insertional mutagenesis allows identification of novel genes involved in disease, while targeted gene knockout or gene silencing allows functional analysis of candidate pathogenicity genes. Analyses of human pathogenic fungi generally rely on cell lines and animal models with different immunosuppression regimes. By contrast, plant pathogens can be studied directly on their hosts. Many more fungal species infect plants compared to animals and thus more plant fungal systems than animal fungal systems are studied, but with a shallower focus. Plants obviously can be manipulated with fewer ethical issues than those associated with animal experimentation. Findings from model systems can often be applied to other host–pathogen interactions. The best studied plant fungal system for ascomycetes is rice and Magnaporthe oryzae (cause of rice blast) (Ebbole, 2007) and for basidiomycetes is maize and Ustilago maydis (cause of corn smut) (Brefort et al., 2009). Model animal fungal systems include immune-suppressed mice and Aspergillus fumigatus, Candida albicans, or Cryptococcus spp; recently amoeboid and non-vertebrate animal model systems, particularly the insect Galleria mellonella, have been used (Mylonakis et al., 2007). General similarities and differences between fungal pathogens of plants and animals are described in Table A13-1 and are discussed in more detail later in this chapter.
The Invasion and Disease Process
The disease process can be considered as a series of consecutive steps beginning with recognition between host and fungus, penetration of the host, colonization and avoidance of host defenses, development of disease symptoms, and finally fungal reproduction and dispersal. These steps are developmentally regulated, and fungal mutants can be generated that are arrested at particular stages (for review see Sexton and Howlett, 2006). The mutated genes are often referred to as pathogenicity genes and are broadly categorized as to where they appear to act in the disease pathway. This is an arbitrary classification for convenience. Clearly there are pathogenicity genes with pleiotropic effects at many steps in the pathway, as some processes (e.g., recognition, pH regulation, oxidative burst, signaling) occur several times. Pathogenicity genes of fungi that attack plants have been reviewed recently (Van de Wouw and Howlett, 2011). Interestingly the steps related to symbiotic relationships between plant and fungi are similar to those involved in pathogenesis; thus the classification of plant–fungus interactions as pathogenic or symbiotic is indistinct.
Fungal inoculum of plants is usually spores (sexual or asexual) that then germinate on the surface of the plant. Animal pathogenic fungi can exist in forms such as yeasts, conidia, or hyphal fragments. Some such fungi undergo dimorphic switching. In some cases, the yeast form is more pathogenic than the hyphal phase, while in others the converse is true. A key class of hydrophobic proteins, hydrophobins coat the fungal cell wall, which is composed generally of
Table A13-1 General Similarities and Differences Between Fungal Pathogens of Plants and Animals
| Feature | Plants | Animals |
| Number of fungal pathogens | Many | Few |
| Importance | Cause 25–30% crop losses | Usually affect immunocompromised animals |
| Lifestyle and environmental niche | Often biotroph; necrotroph; also saprophytes | Often soil saprophytes; not obligates |
| Experimental systems | Many; often shallow focus | Few; different host cell and immunosuppression regimes of non-humans |
| Host specificity | Often host species or cultivar specificity | Sometimes species specific, not genotype specific |
| Mode of entry into hosts | Via stomata, or breaching surface barriers via enzymes or pressure created by infection structures | Via inhalation, ingestion, or wounds in skin |
| Dispersal | Between hosts | Not usually between hosts |
| Reproduction | Sexual and asexual | Usually asexual (ascomycetes) |
| Overall pathogen requirements | Often complex set of physical and molecular barriers to overcome | Inocula small enough to enter host, survive at 37°C and avoid immune responses |
chitin, a polymer of β 1,4 N-acetyl glucosamine, β 1,3 glucans, as well as other polysaccharides and proteins.
Plants have more complex physical barriers to invasion than animals do. Epidermal cells on the plant surface have a cuticle comprising epoxy fatty acids, and a cell wall that is composed of a matrix of proteins and interacting carbohydrates, including β 1,3 and β 1,4 glucans such as cellulose (Carpita and Gibeaut, 1993). By contrast the animal surface, the stratum corneum, is much thinner and the cells are highly keratinized (Proksch et al., 2008). Fungi enter animals by inhalation, ingestion, or wounds. Basically, for a fungus to be an animal pathogen, its major features include being small enough to enter tissue, able to survive at 37°C, and able to evade host immune responses (Sexton and Howlett, 2006). Indeed, the temperature optimum for many fungi is significantly lower than 37°C, which has been proposed as an explanation for why relatively few fungi are important pathogens of mammals (Robert and Casadevall, 2009). The mammalian characteristics of maintenance and close regulation of body temperature are proposed to have a selective advantage in conferring resistance to many fungal pathogens (Bergman and Casadevall, 2010).
Fungi enter plants via the stomatal apertures, where air exchange occurs; by digesting the cuticle and cell wall with hydrolytic enzymes; or by developing infection structures, appressoria, which accumulate high concentrations of glycerol and puncture the surface due to high turgor pressure (Van de Wouw and Howlett, 2011). As mentioned above, fungi enter animals through the skin or via inhalation or wounds; an exception is Histoplasma capsulatus, which enters the host via receptor-mediated endocytosis (Woods, 2003).
The first basal or innate layer of defense is very similar in plants and animals. It involves binding of pathogen associated molecular patterns (PAMPs) to pattern recognition receptors on the host membrane surface. This triggers signaling pathways that induce a range of defense responses, including production of reactive oxygen species and sometimes programmed cell death. This response is termed “pathogen triggered immunity.” PAMPs common to plant and animal pathogenic fungi include wall carbohydrate fragments, such as chitin oligosaccharides or β 1,3 glucans. As well as conservation among PAMPs, there is also conservation in structural domains of Pattern Recognition Receptors (for review see Zipfel, 2009).
In plants often the responses triggered by innate basal immunity are not strong enough to stop pathogen invasion. Consequently a second round of recognition occurs involving effector-triggered immunity (Jones and Dangl, 2006). Often there is a “gene for gene” interaction between avirulence (effector) genes in the pathogen and resistance genes in the plant, such that the pathogen is unable to attack host genotypes with the corresponding resistance genes. Thus there is usually a high degree of host specificity with plant diseases, with only particular varieties (genotypes) of a single species being susceptible. In contrast to plant diseases, most animal diseases do not display host genotype specificity, although some fungi only cause disease in certain animal species. Apart from innate basal immunity, the immune system of mammals is very different from that of plants. The responses that come into play if the pathogen is not stopped by the innate basal immunity response include the innate complement system, circulating cells such as phagocytes that can internalize and destroy pathogen cells, and adaptive antibody-mediated defenses (Speth et al., 2004).
Fungal plant pathogens have a range of lifestyles and nutritional requirements. Many have a saprophytic phase, and some are obligate, surviving only on their hosts. Some (biotrophs) require hosts to be living, while others (necrotrophs) kill plant tissues. Some fungi are both biotrophs and necrotrophs at different stages during growth in planta. Some fungi such as mycorrhizae and endophytes colonize plants in a symbiotic relationship deriving carbon from photosynthesis by the plant and often conferring drought tolerance on the plant. Fungi that infect animals often have a saprophytic lifestage in the soil, and few if any are obligate.
After invasion, fungi then need to colonize the host, derive nutrition, and avoid or subvert host defense responses. In many plant–pathogen interactions, defense responses include the hypersensitive response, whereby an oxidative burst
by the plant generates reactive oxygen species associated with the programmed cell death of host cells. This can arrest pathogen growth, particularly that of biotrophs. Necrotrophic fungi can subvert this defense process to derive nutrition from the dead host tissue. Toxins produced by necrotrophic fungi also kill plant tissue. However, they are generally not often important disease determinants. This is also often the case for secondary metabolite toxins produced by fungi that attack animals. Their role may be to protect fungi against predators such as insects, nematodes, and amoebae during their saprophytic growth phase in the soil (Kempken and Rohlfs, 2010).
To complete its lifecycle, a fungus must reproduce and exit the host to find another host. Many animal pathogenic ascomycetes do not have a sexual phase. However, the sexual cycle of basidiomycetes such as Cryptococcus spp. is extremely important in virulence, as discussed by Heitman.42 Both animal- and plant-pathogenic fungi reproduce mitotically within the host. Although plant-to-plant transmission of fungal disease is very common, direct transmission of fungal pathogens between mammalian hosts is unusual. In plant pathogens, conidia (vegetative spore) production usually occurs after infection has been established and lesions have developed. Conidia are spread from plant to plant by wind or in water droplets. However, many pathogens of mammals are transmitted by inhalation as hyphae or as arthroconidia, in dust and wind (e.g., C. immitis) or soil. Increasingly, biofilms, whereby a community of microorganisms attaches to a solid surface, mediate dispersal of fungi in hospital environments (Ramage et al., 2009).
A Fungal Pathogen That Infects Plants and Animals
Given some of the similarities in strategies that fungi use to cause disease in plants and animals, it is perhaps surprising that few organisms have been reported to infect both plants and animals. A single isolate of Fusarium oxysporum f. sp. lycopersici can infect both plants and animals. The animal experimental system involves injection of the tail vein of immunodepressed mice and the plant system involves incubation on intact tomato roots (Ortoneda et al., 2004). Mutants in several genes of F. oxysporum f. sp. lycopersici have been tested for their virulence in the animal and plant models. A mitogen activated protein kinase gene (fmk1) and an Rho1 GTPase, which are both involved in signaling, are not required for virulence in mice, but are essential for virulence in tomatoes. In contrast, the transcription factor PacC, which mediates the environmental pH signal, and the photosensor WC-1, which interacts with a light-sensitive transcription factor, WC-2, are necessary for full virulence in mice, but are not essential for virulence in tomatoes (Martínez-Rocha et al., 2008; Ortoneda et al., 2004; Ruiz-Roldan et al., 2008). These findings highlight redundancy in some signaling pathways
______________
42 See contributed manuscript by Heitman in Appendix A (pages 226–248).
Table A13-2 Fungicides Used to Control Plant and Animal Diseases
| Fungicide Group | Target | Plants | Animals |
| Azoles | Ergosterol biosynthesis in fungal membrane | Widely used | Widely used |
| Echinocandins | β 1,3 glucan synthesis | Not used | New drug |
| Polyenes: Amphotericin B | Ergosterol biosynthesis | Not used | Yes |
| Nikkomycin Z | Chitin synthesis CHS I | Not used | Trialed against C. immitis |
| Strobulurins | Mitochondrial cytochrome bc complex | Used but resistance can develop: G143A mutations in cytochrome bc | Not used; toxic |
| Bion | Salicylic acid analog: Mimics systemic acquired resistance in plants | Used in horticulture; expensive | Not effective as dependent on plant defense responses |
and the complexity of these interactions. It will be interesting to compare global transcriptional analyses of tomato and mouse tissue infected with this fungus to further identify networks and signaling components regulated in response to interaction with two extremely different host types.
Control of Fungal Diseases
Animal diseases are controlled mainly by a well-functioning immune system of a potential host. There are no approved vaccines against fungi, and fungicides often are not efficacious (Ostrosky-Zeichner et al., 2010). Plant diseases are controlled by a combination of approaches. These include management of inocula; for instance, removing infected stubble (crop trash) at the end of the growing season; using fungicides; and introgressing (breeding) resistance genes, often from related species, into varieties. For diseases of both types of hosts, a range of fungicides is employed; choices depend on toxicity and cost/benefit ratio (Table A13-2). Azoles target ergosterol biosynthesis in the fungal membrane and are the most common group of fungicides used for diseases on both types of hosts. Echinocandins, which target synthesis of β 1,3 glucans in the fungal cell wall, are now being used to combat fungal diseases of humans, but would not be effective against fungal plant pathogens given that plant cell walls also have β 1,3 glucans. Nikkomycin Z, which targets chitin synthesis, is being used to control C. immitis infections (Ostrosky-Zeichner et al., 2010),43 while strobilurins,
______________
43 See article by Galgiani in Appendix A (pages 196–207).
which target mitochondrial cytochrome bc complex, are deployed against plant diseases, but resistance to them can develop and these molecules have a degree of toxicity toward animals (Bartlett et al., 2002). Bion, which is a salicyclic acid analogue, can induce systemic acquired resistance in some plants (Beckers and Conrath, 2007). This molecule is used more in horticulture than broad-acre grain crops due to its cost.
Deploying resistance genes is an important strategy in controlling plant disease. When corresponding avirulence genes are mutated or deleted, the fungus is not recognized by the plant, and infection ensues. Thus resistance is overcome, often resulting in severe yield losses. Some fungi can more readily overcome resistance than others; such fungi are deemed to have “high evolutionary potential” and they usually outcross prolifically, producing large numbers of wind-borne sexual spores as inocula (McDonald and Linde, 2002). These properties enable such fungi to adapt to selection pressures imposed by extensive sowing of crop varieties with resistance conferred by single genes. Thus the frequency of virulent isolates will increase and resistance in the plant can break down. An example of such a breakdown of resistance occurred in 2003 when resistance in canola to blackleg disease caused by L. maculans broke down (Sprague et al., 2006). This resulted in up to 90 percent yield losses in regions of Australia, costing farmers up to $20 million (B. J. Howlett, unpublished). The resistance had been conferred by a canola gene named Rlm1. The corresponding avirulence effector gene, AvrLm1, has been cloned and is located in a very “plastic” part of the L. maculans genome (Gout et al., 2006).
Recently the genome sequence of this fungus has been acquired (Rouxel et al., 2011). The fungus has a unique structure. More than one third comprises repetitive elements composed of degenerated transposons. Gene-rich regions with high GC content alternate with gene-poor regions with high AT content with sharp boundaries between them. Disease-related genes such as effectors are interspersed in the gene-poor regions among repetitive DNA. This organization contributes to the readily changeable nature of the fungal genome and enables effector genes to be readily gained, lost, or mutated (Rouxel et al., 2011; Van de Wouw et al., 2010). Markers are now available for several other avirulence effector genes (Fudal et al., 2007; Parlange et al., 2007), and thus the frequency of virulent isolates within fungal populations in the field that are virulent toward a particular resistance gene can be monitored. This information allows farmers to rotate the disease resistance genes in the canola varieties that they sow from year to year. This strategy, which can be applied to other plant diseases, can help prevent outbreaks and maximizes the duration of effectiveness of resistance genes.
Closing Remarks
The development of fungal diseases of plants and animals has many parallels. A fungus must overcome many hurdles to successfully invade a plant and cause
disease. There is a continuum from disease to damage to symbiosis with plant–fungal interactions. The most important properties of a human pathogen are to be able to survive at 37°C and to avoid the immune responses. The use of fungicides to control disease is made on the basis of specificity and after cost/benefit analyses. Such decision making about a broad acreage crop is very different from that about a human patient. An important strategy to control plant disease is to incorporate resistance genes. However, virulence toward this resistance can be selected for, due to the high evolutionary potential of some fungi. Thus resistance genes have to be managed (rotated) by farmers to maximize longevity of resistance.
References
Bartlett, D. W., J. M. Clough, J. R. Godwin, A. A. Hall, M. Hamer, and B. Parr-Dobrzanski. 2002. The strobilurin fungicides. Pest Management Science 58:649–652.
Beckers, G. J., and U. Conrath. 2007. Priming for stress resistance: From the lab to the field. Current Opinion in Plant Biology 10:425–431.
Bergman, A., and A. Casadevall. 2010. Mammalian endothermy optimally restricts fungi and metabolic costs. MBIO 1(5):e00212–10.
Brefort, T., G. Doehlemann, A. Mendoza-Mendoza, S. Reissmann, A. Djamei, and R. Kahmann. 2009. Ustilago maydis as a pathogen. Annual Review of Phytopathology 47:423–445.
Cairns, T., F. Minuzzi, and E. Bignell. 2010. The host-infecting fungal transcriptome. FEMS Microbiology Letters 307:1–11.
Carpita, N. C., and D. M. Gibeaut 1993. Structural models of primary-cell walls in flowering plants—consistency of molecular structures with the physical properties of the wall during growth. Plant Journal 3:1–30.
Casadevall, A., and L.A. Pirofski. 2003. The damage–response framework of microbial pathogenesis. Nature Review Microbiology 1:17–24.
Ebbole, D. J. 2007. Magnaporthe as a model for understanding host–pathogen interactions. Annual Review of Phytopathology 45:437–456.
Fudal, I., S. Ross, L. Gout, F. Blaise, M. L. Kuhn, M. R. Eckert, L. Cattolico, S. Bernard-Samain, M. H. Balesdent, and T. Rouxel. 2007. Heterochromatin-like regions as ecological niches for avirulence genes in the Leptosphaeria maculans genome: Map-based cloning of AvrLm6. Molecular Plant Microbe Interactions 20:459–470.
Gout, L., I. Fudal, M. L. Kuhn, F. Blaise, M. Eckert, L. Cattolico, M.-H. Balesdent, and T. Rouxel. 2006. Lost in the middle of nowhere: The AvrLm1 avirulence gene of the Dothideomycete Leptosphaeria maculans. Molecular Microbiology 60:67–80.
Jones, J. D. G., and J. Dangl. 2006. The plant immune system. Nature 444:323–329.
Kempken, F., and M. Rohlfs. 2010. Fungal secondary metabolite biosynthesis—a chemical defence strategy against antagonistic animals? Fungal Ecology 3:107–114.
Martínez-Rocha, A. L., M. I. G. Roncero, A. López-Ramirez, M. Mariné, J. Guarro, G. Martínez-Cadena, and A. Di Pietro. 2008. Rho1 has distinct functions in morphogenesis, cell wall biosynthesis and virulence of Fusarium oxysporum. Cellular Microbiology 10:1339–1351.
McDonald, B. A., and C. Linde. 2002. Pathogen population genetics, evolutionary potential, and durable resistance. Annual Review of Phytopathology 40:349–379.
Mylonakis, E., A. Casadevall, and F. M. Ausubel. 2007. Exploiting amoeboid and non-vertebrate animal model systems to study the virulence of human pathogenic fungi. PLoS Pathogens 3:859–865.
Ortoneda, M., J. Guarro, M. P. Madrid, Z. Caracuel, M. I. Roncero, E. Mayayo, and A. Di Pietro. 2004. Fusarium oxysporum as a multihost model for the genetic dissection of fungal virulence in plants and mammals. Infection and Immunity 72:1760–1766.
Ostrosky-Zeichner, L., A. Casadevall, J. N. Galgiani, F. C. Odds, and J. H. Rex. 2010. An insight into the antifungal pipeline: Selected new molecules and beyond. Nature Reviews Drug Discovery 9:719–727.
Parlange, F., G. Daverdin, I. Fudal, M. L. Kuhn, M.-H. Balesdent, F. Blaise, B. Grezes-Besset, and T. Rouxel. 2007. Leptosphaeria maculans avirulence gene AvrLm4-7 confers a dual recognition specificity by the Rlm4 and Rlm7 resistance genes of oilseed rape, and circumvents Rlm4mediated recognition through a single amino acid change. Molecular Microbiology 71:851–863.
Proksch, E., J. M. Brandner, and J. M., Jensen. 2008. The skin: An indispensable barrier. Experimental Dermatology 17:1063–1072.
Ramage, G., E. Mowat, B. Jones, C. Williams, and J. Lopez-Ribot. 2009. Our current understanding of fungal biofilms. Critical Reviews in Microbiology 35:340–355.
Robert, V. A., and A. Casadevall. 2009. Vertebrate endothermy restricts most fungi as potential pathogens. Journal of Infectious Diseases 200:1623–1626.
Rouxel, T., J. Grandaubert, J. K. Hane, C. Hoede, A. P. van de Wouw, A. Couloux, V. Dominguez, V. Anthouard, P. Bally, P. Bourras, A. J. Cozijnsen, L. M. Ciuffetti, A. Degrave, A. Dilmaghani, L. Duret, I. Fudal, S. B. Goodwin, L. Gout, N. Glaser, J. Linglin, G. H. J. Kema, N. Lapalu, C. B. Lawrence, K. M. May, M. Meyer, B. Ollivier, J. Poulain, C. L. Schoch, A. Simon, J. W. Spatafora, A. Stachowiak, B. G. Turgeon, B. M. Tyler, D. M. Vincent, J. Weissenbach, J. Amselem, H. Quesneville, R. P. Oliver, P. Wincker, M.-H. Balesdent, and B. J. Howlett. 2011. Effector diversification within compartments of the Leptosphaeria maculans genome affected by repeat induced point mutations. Nature Communications 2:202.
Ruiz-Roldan, M. C., V. Garre, J. Guarro, M. Marine, and M. I. G. Roncero. 2008. Role of the white collar 1 photoreceptor in carotenogenesis, UV resistance, hydrophobicity and virulence of Fusarium oxysporum. Eukaryotic Cell 7:1227–1230.
Sexton, A. C., and B. J. Howlett. 2006. Parallels in fungal pathogenesis on plant and animal hosts. Eukaryotic Cell 5:1941–1949.
Sharpton, T. J., J. E. Stajich, S. D. Rounsley, M. J. Gardner, J. R. Wortman, V. S. Jordar, R. Maiti, C. D. Kodira, D. E. Neafsey, Q. Zeng, C. Y. Hung, C. McMahan, A. Muszewska, M. Grynberg, M. A. Mandel, E. M. Kellner, B. M. Barker, J. N. Galgiani, M. J. Orbach, T. N. Kirkland, G. T. Cole, M. R. Henn, B. W. Birren, and J. W. Taylor. 2009. Comparative genomic analyses of the human fungal pathogens Coccidioides and their relatives. Genome Research 19:1722–1731.
Skamnioti, P., and S. J. Gurr. 2009. Against the grain: Safeguarding rice from rice blast disease. Trends in Biotechnology 27:141–150.
Speth, C., G. Rambach, C. Lass-Florl, M. P. Dierich, and R. Wurzner. 2004. The role of complement in invasive fungal infections. Mycoses 47:93–103.
Sprague, S. J., S. J. Marcroft, H. L. Hayden, and B. J. Howlett. 2006. Major gene resistance to blackleg in Brassica napus overcome within three years of commercial production in southeastern Australia. Plant Disease 90:190–198.
Van de Wouw, A. P., and B. J. Howlett. 2011. Fungal pathogenicity genes in the age of “omics.” Molecular Plant Pathology 12:507–514.
Van de Wouw, A. P., A. J. Cozijnsen, J. K. Hane, P. C. Brunner, B. A. McDonald, R. P. Oliver, and B. J. Howlett. 2010. Evolution of linked avirulence effectors in Leptosphaeria maculans is affected by genomic environment and exposure to resistance genes in host plants. PLoS Pathogens 6:e1001180.
Vincent, D., M. H. Balesdent, J. Gibon, S. Claverol, D. Lapaillerie, A. M. Lomenech, F. Blaise, T. Rouxel, F. Martin, M. Bonneu, J. Amselem, V. Dominguez, B. J. Howlett, P. Wincker, J. Joets, M. H. Lebrun, and C. Plomion. 2009. Hunting down fungal secretomes using liquid-phase IEF prior to high resolution 2-DE. Electrophoresis 30:4118–4136.
Woods, J. 2003. Knocking on the right door and making a comfortable home: Histoplasma capsulatum intracellular pathogenesis. Current Opinion in Microbiology 6:327–331.
Zipfel, C. 2009. Early molecular events in PAMP-triggered immunity. Current Opinion in Plant Biology 12:414–420.
CLIMATE, GLOBALIZATION, AND TRADE: IMPACTS ON DISPERSAL AND INVASION OF FUNGAL PLANT PATHOGENS
Michael Jeger,44Marco Pautasso,44and James Stack45
Climate change is likely to become a major issue in plant pathology over the coming years. In this overview, we provide recent evidence for this statement. Moreover, we point out the importance for future plant disease management of climate change interactions with other global change drivers, such as increased long-distance trade. Recent advances in aerobiology, together with new molecular tools used in landscape and geographical genetics, can help in addressing the challenges posed by an increasing number of emerging plant diseases worldwide. There is increasing evidence that climate change will be a key issue in how plant pathogens will affect food security and ecosystem health.
Less knowledge is available on the potential impacts of climate change on biological control of exotic fungal plant pathogens. Network theory is a promising tool to improve biosecurity in the face of the increased volumes of traded plants coupled with climate warming. Although there are now many reviews of the literature on the topic of climate change and plant diseases, there is a need to keep up with the rapid development of the subject.
Introduction
The emergence of fungal plant pathogens can occur as the result of at least three processes: (1) the evolution of new genotypes within a pathogen population that is endemic to a habitat or location (e.g., through genetic recombination); (2) the introduction of an exotic pathogen genotype into a new, receptive habitat or location; and/or (3) the natural selection of new pathogen genotypes from an endemic population as a result of changes in host genotype, host species, or external pressures (e.g., environmental conditions, fungicide applications, changes in plant cultural practices). Emergence is a function of introduction (via dispersal or evolution), establishment (i.e., adaptation to the habitat or location), and spread
______________
44 Division of Biology, Imperial College London, Silwood Park, Ascot, SL5 7PY, U.K.
45 Department of Plant Pathology, Kansas State University, 4024 Throckmorton Plant Sciences Center, Manhattan, KS 66506-5502.
(i.e., dispersal via natural or human-mediated mechanisms). Weather variables, including wind patterns and storms, and the global trade in plants and plant products facilitate the dispersal of plant pathogens and thus provide opportunities for the emergence of fungal plant pathogens. Further, weather is a primary regulator of invasion into new habitats and locations by plant pathogens. Consequently, climate change may provide unprecedented opportunities for the emergence of fungal plant pathogens through the dispersal of fungal pathogens to new locations and/or through habitat modifications for both pathogens and plants.
Climate, globalization, and trade are major drivers of the dispersal and invasion of fungal plant pathogens, affecting agricultural and horticultural crops, plantation and forest trees, and plants in natural/semi-natural environments. Patterns of weather, notably large-scale air movements, have been implicated in long-range dispersal of several economically important pathogens between and within continents. In some cases, continental dispersal of plant pathogens is tracking and catching up with previous movements of crop plants, in turn accompanying human population migrations that occurred over centuries, sometimes millennia. The rate of movement of humans, crop plants, and plant pathogens has intensified markedly in recent decades with the increased globalization of the world economy. Expansion of trade pathways and technological innovations make possible the production, harvesting, storage, marketing, shipping, and air freight of food crops, ornamental plants, and plant products to an extent that previously was not possible (Figure A14-1).
From a broad perspective, climate and global change are drivers of large-scale ecological perturbations that facilitate novel “biomixing” and “ecological fitting” (Agosta et al., 2010). These phenomena can lead to rapid host switching, the emergence of hybrid pathogens, and invasion of new infectious diseases and pests (Brasier, 2008; Palm, 1999). Changes in plant phenology occur in a variety of ways, depending on species and geography. Such changes impact on plant interactions with fungi in general (Gange et al., 2011; Kauserud et al., 2010) and with fungal pathogens in particular (Grulke, 2011; Marçais et al., 2009). The consequence is that multiple species at many sites need to be studied in order to understand and predict regional change and impact (Ibanez et al., 2010). An alternative strategy has been to focus on multiple drivers of global change on a single species (Baeten et al., 2010; Matesanz et al., 2009; Paajanen et al., 2011). Global environmental change arises from CO2 enrichment, increased nitrogen deposition, climate shifts, biotic interactions, and land use change (Tylianakis et al., 2008). These factors have pervasive effects on antagonistic and mutualistic interactions between plants and fungi and in some cases increase the severity of pathogen infection while weakening mutualisms. Interactions must be expected between tree genetic diversity, variation in phenology, resistance to defoliators and fungal pathogens, increased CO2, and ozone concentration affecting tree growth and mortality; some of these interactions have been reviewed by Pautasso (2009).
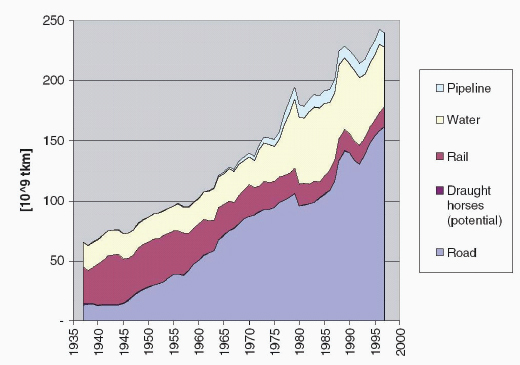
Figure A14-1 The increase in goods (109 tons × km) moved in the United Kingdom from the 1930s to the 1990s.
SOURCE: Schulz (2004).
In this overview we consider the impact of climate, globalization, and trade on dispersal and invasion of fungal plant pathogens in the broader ecological context described above. We make a distinction between dispersal that is mediated by natural means, mostly atmospheric, and that which is mediated by human intervention of one form or another. This is partly a matter of convenience: we recognize that at some scales of dispersal both means can be important. With respect to pathogen emergence, dispersal is ineffective without establishment, spread, and persistence—the elements of invasion.
Natural Dispersal of Fungal Plant Pathogens
Fungal spores that escape the boundary layer of crop canopies can be transported over long distances subject to their biophysical characteristics and meteorological conditions. The study of such transport belongs firmly in the domain of aerobiology, a discipline that owes much of its development to the effect of aero-allergens on human health (Dallafior and Sesartic, 2010). An account of an early pioneer of aerobiology, Philip Gregory, and the link with fungi of agricultural and human health concern, can be found in Lacey et al. (1997). Weather has
significant effects on the incidence of aero-allergens, including the abundance and biodiversity of spores of the fungal plant pathogen Alternaria alternata (Magyar et al., 2009). Under conditions predicted by climate change, changes in planting practices and modified crop management may be required to keep allergen concentrations under control (Beggs, 2010).
Even if climate change is becoming important in allergen aerobiology (D’Amato and Cecchi, 2008), vegetation normally will be the main source of fungal spores in the atmosphere. Thirty-two genera of fungi arising from vegetation sources were found across both cultivated and urban areas in three regions in Egypt, with a clear association with weather conditions and many implications for the spread of human and plant diseases (Awad, 2005). Fungal concentration in the atmosphere may not be the best indicator of health risk, which may be more associated with the predominant aero-allergen present (Awad, 2005). The effects of meteorological factors on atmospheric dispersal, in biophysical terms relevant for plant pathogens (including viruses and bacteria), was provided in a recent review (Jones and Harrison, 2004).
In some cases, the dispersal of fungal plant pathogens has been modeled explicitly using biophysical principles. A recent example has been the dispersal of Phakopsora pachyrhizi, causing soybean rust (Andrade et al., 2009). The American continent was free of P. pachyrhizi until 2001. After its introduction in Paraguay, the pathogen rapidly became established throughout Bolivia, Brazil, and Argentina, possibly due to a combination of large-scale cultivation of the plant host, international movement of infected material, and long-distance natural dispersal. Urediniospores of this fungus have been shown to remain viable long enough to be able to travel hundreds of kilometers (Savage et al., 2010). In 2004, Asian soybean rust was reported in the continental United States (Goellner et al., 2010). The fungus now overwinters in warm southern U.S. locations. Spore escape was modeled and combined with a standard large-scale transport model to forecast spore deposition over U.S. soybean production areas. Canopy turbulence and canopy porosity were found to be key determinants of spore escape.
Some effort has been made to test the validity of transport models (Skelsey et al., 2009). Spijkerboer et al. (2002) evaluated the Gaussian plume model (GPM) for predicting and describing spore dispersal over a potato crop. The main purpose was prediction of Phytophthora infestans, but for experimental purposes they used a commonly used fern spore. They concluded that the GPM was not applicable in risk assessments, unless combined with site-specific information at the source, such as spore escape in relation to wind speed.
Other more empirical approaches can also be used for specific purposes. High-speed imaging showed that, by synchronizing the ejection of thousands of spores, ascomycete fungi such as the pathogen Sclerotinia sclerotiorum form an air flow that carries spores around intervening obstacles to atmospheric currents and new infection sites (Roper et al., 2010). Mundt et al. (2009) used simple empirical relationships based on the inverse power law to describe the spread of
plant diseases such as wheat stripe rust, wheat stem rust, potato late blight, and Southern corn leaf blight. Much of the earlier literature on this approach is cited by the authors. They found that the estimated power law parameter varied little over five orders of magnitude on a distance scale. Evidence was found to support the hypothesis that disease advances through accelerating, rather than constant, waves. Integration of (unmanned) aerial measurements of spore concentration at various distances from the source with simulation of spore flight trajectories is important to develop reliable decision support systems to predict risk of disease spread, as shown for Phytophthora infestans in potato fields in Virginia (Aylor et al., 2011; Techy et al., 2010).
A feature of recent work on dispersal has been the integration with population genetics aspects of pathogen diversity (Hovmøller et al., 2008; Montarry et al., 2010). This can operate at the level of host specialization and biotrophy and the ways in which wind dispersal acts as a survival mechanism—what has been termed “Oases in the desert” (Brown et al., 2002). Equally, the strongly stochastic nature of long-distance dispersal can lead to founder effects in the pathogen population (Brown and Hovmøller, 2002). In such cases, pathogen genotypes that successfully establish in new regions and/or on new cultivars may be different from those at the source—the so-called founder effect. Such effects were found in the migration of the fungus causing black Sigatoka of bananas, Mycosphaerella fijiensis, from Southeast Asia to Africa and Latin America (Brown and Hovmøller, 2002; Rivas et al., 2004). In both of the new regions founder effects were present in the new invasions. Unresolved for this pathogen is the relative importance of air-dispersed ascospores (probably limited) and the movement of infected plant material (largely unrecorded). The fungus Corynospora cassiicola has a wide geographical range in the tropics and subtropics and many plant hosts. Common fungal lineages were widely distributed geographically, indicating long-distance dispersal of clonal lineages, but also previously unrecognized genetic diversity involving some degree of host specialization on some hosts (Dixon et al., 2009). The advent of modern molecular tools in epidemiology provides a step change in both the tracking of dispersal of novel fungal genotypes and in risk assessment of emerging fungal diseases in plants and in animals (Gladieux et al., 2011; Moslonka-Lefebvre et al., 2011).
Climate and Plant Diseases
Weather and climate generally have major impacts on diseases caused by fungal plant pathogens. This topic has had extensive historical coverage that it is not possible to cover in this overview. What is more significant and of immediate concern is how climate change will impact the distribution and severity of diseases caused by known pathogens and the emergence of new invasive pathogens. This topic has also had its fair share of literature reviews: recently, e.g., in the context of structural change in the international horticultural industry (Dehnen-
Schmutz et al., 2010); the evolution of the phytosanitary regulatory framework (MacLeod et al., 2010); forest health and adaptive management (Parks and Bernier, 2010); the impacts on plant health and carbon sequestration in Australia (Singh et al., 2010); cool season grain legume crops and their diseases (Thomas, 2010); urban trees and their pathogens (Tubby and Webber, 2010); diseases in tropical plantation crops (Ghini et al., 2011); the disease triangle and changes in plant phenology (Grulke, 2011); rice diseases and pests (Haq et al., 2011); diseases of food crops (Luck et al., 2011); the geographical distribution of plant pathogens (Shaw and Osborne, 2011); and plant pathogens in Sweden (Roos et al., 2011).
Here, we refer to climate change to include trends in air composition as well as trends in global warming. Air composition in terms of SO2 concentration has been shown to be associated with the relative prevalence of two important fungal pathogens of wheat (Fitt et al., 2011), a finding made possible by the application of molecular analytical tools to archived plant material from the long-term Broad-balk experiments at Rothamsted Research in the United Kingdom. Elevated atmospheric CO2 and ozone concentrations decreased the incidence of downy mildew disease in soybean, but increased the severity of disease caused by Fusarium virguliforme (Eastburn et al., 2009). Changes in precipitation and temperature resulted in increased disease severity for both diseases, and there were indirect effects due to treatment effects on canopy structure and leaf age. Similar kinds of results were obtained in studies of four pepper diseases (Shin and Yun, 2010). Elevated CO2 and temperature treatments increased the rate of progress for two bacterial diseases, but not for a stramenopile disease (Phytophthora capsici) or a fungal disease (Colletotrichum acutatum).
The relationship between climate change, plant diseases, and food security (Chakraborty and Newton, 2011) considers international cooperation and integrated solutions, including disease management issues, to be essential to meet the food demands of the growing world population. Plant breeding for climate-related traits such as drought avoidance or tolerance (Khan et al., 2010) must also take into account disease resistance. The arable sector will be critical in this respect, with mitigation and adaptation strategies with respect to plant disease control likely to become a key area (Fitt et al., 2010). Because of the variation in crop growth and pathogen environmental requirements, geographical divides in crop yield and productivity may become more pronounced (Butterworth et al., 2010). Climate change will also impact on the incidence of mycotoxins in food (Russell et al., 2009), a much-neglected topic in relation to food security and human and animal health.
Forest health is one area where the impact of climate change and biological invasions is providing a clear signal (Kliejunas et al., 2008; Sturrock et al., 2011; Woods et al., 2010). In turn, emerging forest diseases under climate change can lead to a positive feedback by reducing the carbon stocks of affected forests (Peltzer et al., 2010), as seen in British Columbia with the developing Dothis-
troma needle blight epidemic following the devastating mountain pine beetle outbreaks. The resilience of northern boreal forests to rapid climate change can be questioned (Chapin et al., 2010), as can forest establishment under conditions of permafrost thaw where fungal pathogens may affect seedling survival (Camill et al., 2010). In forest tree nurseries in Finland, rust, powdery mildew, and other fungal leaf diseases are already causing more problems because of climate warming (Lilja et al., 2010). A more optimistic outlook due to improved adaptive management practices is presented with respect to white pine blister rust (Hunt et al., 2010), where white pines have broad ecological ranges and are less likely to be maladapted thus succumbing to the disease, and hence may be more resilient in the long term. Similar issues will need to be faced with regard to urban trees (Tubby and Webber, 2010) and Mediterranean forests (Attorre et al., 2011; La Porta et al., 2008), where the impact of non-native insect pests and fungal pathogens introduced through trade pathways (see later sections) is already being observed.
The above discussion concerns general issues regarding climate change and plant diseases. There have been many accounts of global/climate change impacts on diseases caused by specific fungal plant pathogens in recent years. A selection of these is cited with summary comments in Table A14-1.
Climate and Beneficial/Biocontrol Fungi
Compared with studies on plant pathogenic fungi, there have been relatively few studies on the effects of climate change on beneficial plant-associated fungi. There has been some consideration of mycorrhizal associations and endophytic fungi, but little on tritrophic mycoparasitic interactions (Singh et al., 2009). In a review of the results of 135 studies, Compant et al. (2010) found that elevated CO2 had a positive influence on the incidence of arbuscular and ectomycorrhizal fungi, whereas the effects on endophytic fungi were more variable. Effects of temperature were idiosyncratic, with positive, neutral, and negative effects equally common. Plant growth-promoting fungi (as with bacteria) positively affected plants subject to a degree of drought stress. Considerable research has been done with tritrophic interactions among arthropod pests and their natural enemies (Thomson et al., 2009), including fungal entomopathogens; similar research is lacking for biocontrol of plant pathogens (Ghini et al., 2008). The expectation must be that prediction will be difficult because of the indirect and direct effects on biocontrol fungi, unless there is a good understanding of tritrophic interactions (Thomson et al., 2009). Biological control of weeds using fungi is simply a case of a plant pathogenic fungus being used for a beneficial purpose. Cirsium arvense is a troublesome weed with world-wide distribution. Climate change will exacerbate invasion and persistence of the weed and make biological control using a variety of pathogenic fungi more difficult (Tiley, 2010).
| Pathogen | Host | Reference | Comments |
| Valsa | Alder | Worrall | Previous oscillation period in damage of ~21 years |
| melandiscus | et al. (2010) | likely to dampen with warmer summers and with no period of recovery | |
| Erisiphe necatrix | Grapevine | Pugliese et al. (2010) | Increase in CO2 concentration did not affect incidence possibly due to increased photosynthesis with higher CO2 and temperature |
| Phytophthora citricola | Beech | Fleishmann et al. (2010) | Elevated CO2 and low N supply enhanced susceptibility, but host compensation followed |
| Cercospora spp. | Red bud and Sweet bud | McElrone et al. (2010) | Incidence/severity under elevated CO2 greater with above average rainfall and temperature (one species), but mitigated by higher photosynthetic efficiency |
| Leptosphaeria maculans and L. biglobosa | Oilseed rape | Stonard et al. (2010) | Geographic variation in species may be exacerbated by climate change with more damaging species expanding in range |
| Didymella rabiei | Chickpea | Frenkel et al. (2010) | Isolates infecting crop and wild Cicer show different adaptation to high temperature, with the potential for hybrids infecting both hosts over a broader temperature range |
| Phakopsora pachyrhizi | Soybean | Del Ponte and Esker (2008) | Integrating epidemiological and meteorological factors suggest no restricted overwintering areas in Brazil |
| Seiridium cardinale | Mediterranean cypress | Zocca et al. (2008) | Pathogen is associated with insect vectors, which are able to reach the range margin and thus the continuous threat of arrival in the expanding range |
| Cronartium ribicola | White pine | Frank et al. (2008) | Pathogen arrived in New Mexico in ~1970 with upper flow synoptic classification indicating early June 1969 most favourable |
| Fusarium spp. | Wheat | Xu et al. (2008) | Environmental conditions differentially affect infection and colonization process and the comparative abundance of six toxigenic species in the head blight disease |
| Fusarium spp. | Maize | Horvath (2003) | Damage by beetles provides conducive growing conditions for the toxigenic fungi, with feeding behavior changing under drought conditions |
Human-Mediated Dispersal of Fungal Plant Pathogens
Thus far we have considered natural dispersal and invasion of fungal plant pathogens, where the main drivers are weather variables, or more generally climate. Although pathogens have always accompanied humans and their crops over the centuries, most notably with the historical migrations of human populations (Guillemaud et al., 2011; Money, 2007), there have been instances where crops moved to new areas have escaped, at least temporarily, the pathogens endemic in the original distributional range of the crops (Mitchell and Power, 2003). Equally, there are cases where crops grown in a new environment have been exposed to novel pathogens. These movements have taken place over centuries with periods of time for adaptation (assisted through plant breeding) or for the pathogen to reencounter the host in a new environment. With the globally connected world that now exists, these time scales are much shorter—decades or, in some cases, much less. The consequence has been the emergence of new fungal plant pathogen species or levels of subspecific variants that were previously unrecorded. For example, since the publication of Erwin and Ribeiro’s 1996 Handbook on the genus, 39 new species of Phytophthora and two species of hybrids have been formally described (Ersek and Ribeiro, 2010). It is unlikely that this increase is due solely to the advent of modern molecular taxonomic techniques. More likely it is due to the ability of this genus to adapt to new hosts in new environments, through encounters made possible by new pathways.
Less spectacular has been the historical accumulation of non-indigenous forest pests in the United States, where some 450 insect and pathogen species have colonized since European settlement (Aukema et al., 2010). Some 16 pathogenic species have caused substantial damage to trees. This finding is more in keeping with analyses over 10 taxonomic groups of alien species (including fungi), which suggested a historical legacy going back at least a century (Essl et al., 2011). This result would imply that Britain is more at risk for invasion of new exotic species than other European countries (as has happened, for instance, for Phytophthora ramorum), given its historical links to its Commonwealth (Figure A14-2). However, even where that is the case, the corollary is that the impact of current global activity will be even more manifest in the decades to come (Crooks, 2005).
Dispersal Through Trade Pathways
Even in a country with an efficient and visible plant quarantine service, such as Australia, there are problems in defining what is present and what is absent (Hyde et al., 2010). These authors make a case for a reinventory of Australia’s plant pathogens and consider five fungal groups in which what were thought to be species were in fact species complexes. Without this level of discrimination concerning what is in the country, it will not be possible to operate an effective quarantine and plant protection service. The often tortuous relationship between plant quarantine and trade barriers can be a problematic political issue (MacLeod
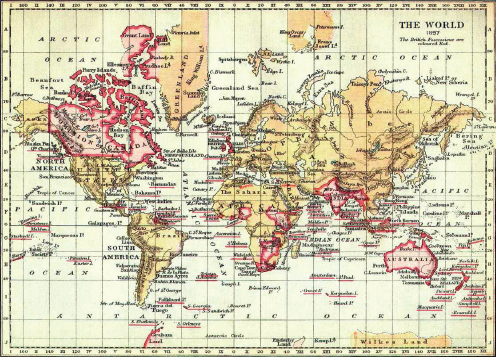
Figure A14-2 The world in 1897, with British possessions marked in red.
SOURCE: Cambridge University Library available at Wikimedia Commons.
et al., 2010; Perrings et al., 2010). Emerging plant diseases can be seen as negative externalities deriving from the international trade in plants (Lansink, 2011). The removal of trade barriers, however, can be beneficial and in some cases desirable. For example, seed trade legislation was designed primarily to protect trade and return royalties to contemporary plant breeders. Increasingly, the importance of exploiting the genetic diversity present in cereal land races has been recognized, but to exploit their use fully, changes in legislation will be required (Newton et al., 2010). Also the genetic diversity of target plant pathogens should be used in building comprehensive collections to allow efficient, reliable, and specific diagnostic and detection tools in the national and international trade (Barba et al., 2010).
The Sanitary and Phytosanitary Agreement of the World Trade Organisation specifies that countries cannot regulate against unknown pests, yet many invasive alien forest insects and pathogens were new to science when first recorded in a new environment (Britton et al., 2010). To counter this, effective surveillance systems are required to facilitate early detection; these are lacking in many nations. Britton et al. (2010) propose a global network of sentinel plantings based in historical gardens and arboreta to enable early detection and rapid response
to such incursions. To support early detection, diagnostic capability and capacity are needed to provide rapid and accurate identification of emerging or introduced pathogens. Progress has been made in establishing diagnostic capability in rural areas of some developing nations and improving diagnostic capability in some developed nations (Boa, 2007; Miller et al., 2009; Stack, 2010).
Mitigating the nursery stock pathway (for forest trees and ornamentals) for undescribed pathogens will be extremely difficult. In general, analysis has been lacking on how the structure of trade pathways affects the spread of plant pathogens in the nursery trade. Pautasso et al. (2010c) analyzed the hierarchical structure in terms of the proportions of producers, wholesalers, and retailers involved in the trade. Despite the many uncertainties associated with commercially sensitive trade information, it was concluded that disease management options should concentrate on the middle tier of the nursery hierarchy, particularly in the absence of hubs (superconnected nodes). If hubs are present, then control is better targeted at them because (ceteris paribus) the higher the correlation between links in and out of nodes, the lower the epidemic threshold (Moslonka-Lefebvre et al., 2009). In addition, the number of outgoing links from the starting node of simulated epidemics in small-size directed networks (a realistic assumption for plant trade among countries and within regions) is strongly correlated with epidemic final size at equilibrium (Pautasso et al., 2010b). These analyses were motivated by the outbreaks of the emerging plant pathogen Phytophthora ramorum but have broader generality for systems where susceptibility and infection are not too incompatible. Rather, the states form two poles of a continuum (a realistic assumption for the plant trade, where premises and plant shipments can have a varying proportion of infected plants).
In 2009, the United States imported 1.2 billion live plants, 2.6 million pounds of seed for planting, and countless shipments of horticultural plants that pass through inspection and enter the retail distribution system in less time than the latent period for most diseases. The tremendous volume of trade of plants and plant products crossing borders and ecological areas and the speed of that trade precludes inspection and interception as a primary strategy for preventing the emergence of plant pathogens (Stack, 2010; Stack et al., 2010). On average, 1–2 percent of U.S. imports are inspected. Even with effective surveillance and diagnostic systems, the emergence of fungal plant pathogens will continue. Existing natural dispersal pathways at the site of introduction, in addition to local and regional trade channels, provide opportunities for spread from the sites of introduction.
The first comprehensive inventory of alien fungi and oomycetes recorded in France since 1800 was recently reported (Desprez-Loustau et al., 2010), with some 65 percent being plant pathogens. Using this dataset, the factors influencing invasion success were investigated, with an emphasis on forest tree pathogens. There was an influence of climatic factors, but human population size and its relationship with imports of plants in trade was a major explanatory variable.
In some cases, it is not simply the occurrence of a plant health problem in trade that is the issue, but that of human or animal health, notably with regard to mycotoxins. For example, the occurrence of tracheomycosis associated with Gibberella xylarioides on Robusta coffee in East Africa (Hindorf, 1998) has added to the long-known mycotoxin problem in raw coffee, limiting imports, especially to the European Union. Similarly, the increasing trade volumes of fresh produce due to consumer appreciation of the health benefits brought by consumption of fresh fruit and vegetables are associated with a greater occurrence of outbreaks of foodborne human pathogens, such as Salmonellae, Escherichia coli, noroviruses, and hepatitis A (Berger et al., 2010).
Dispersal of Plant Pathogens as a Cause and Consequence of Warfare and Social Unrest
Perhaps it is no coincidence that sometimes new pathogen problems emerge during periods of warfare and social upheaval. The Bengal famine in the mid 1940s due to Cochliobolus miyabeanus was associated with severe weather, but it also occurred at the end of World War II, when many military, political, and social issues constrained the ability to deal with the problem: enough food was there, but it was not distributed to those who needed it (Padmanabhan, 1973). In some cases, famines due to fungal pathogens can lead to, rather than be caused by, periods of social change, as shown by the Irish migrations following the famine caused by Phytophthora infestans in the middle of the 19th century (Fry and Goodwin, 1997). In central Italy, the invasion of stone pine stands by North American isolates of Heterobasidion annosum was linked to the movement of U.S. troops during World War II (Gonthier et al., 2007). Since that time, there has been a rate of advance of 1.3 km/year along invasion corridors, with the North American taxon dominant, but also some hybridization with the European taxon. The two pathogens are active in the Circeo National Park, where they cause extensive (up to 30 m in radius) gaps in Pinus pinea plantations (Scirè et al., 2011). Subsequent studies showed that invasion success was due to the reproductive potential of the invader not being reduced during the dry seasons, when compared with the resident (Garbelotto et al., 2010). Unusually, this example of a fungal plant pathogen has been commented on (“Trees become casualties of war”) in a respected medical journal (Dixon, 2005). Although there have been cases of plant pathogen outbreaks during wars and revolutions, today there is a trend away from looking at climate change with a disaster-focused perspective to an emphasis on long-term livelihood security and adaptation (Conway and Schipper, 2011). For example, the shift from today’s widespread reliance on chemical fungicides to the adoption of alternative and less toxic products such as inorganic salts may happen following democratic regulations and in agreement with consumers’ perceptions, rather than as a result of, for example, mass protests or social unrest (Deliopoulos et al., 2010).
The intentional introduction of plant pathogens as a strategy of warfare has long been considered (Madden and Wheelis, 2003). Concern regarding the creation and deployment of novel pathogen genotypes with high virulence and/ or expanded host range could certainly result in the emergence of novel fungal plant pathogens. Although there are clear examples of state-sponsored programs from the past, there is considerable speculation about the relative importance of intentional introductions in the face of introductions resulting from trade and natural events (Stack et al., 2010; Young et al., 2008).
Dispersal Associated with the Introduction of Novel Crops and Plant Species
Farmers world-wide tend to maximize the potential of the crops they grow, in ways that are appropriate to their socioeconomic circumstances, but often only sensible in the short term or not considering side effects such as enhanced opportunities for plant pathogens. For example, modern agriculture relies heavily on increasing the size of fields to enable a more efficient mechanization and economies of scale. But this leads to more uniform landscapes, with potential repercussions on epidemic thresholds of plant pathogens (Moslonka-Lefebvre et al., 2011). Not just in natural ecosystems (Scherrer and Körner, 2011), but probably also in agriculture, small-scale topographic variability will be important for plant species to be able to cope with climate change. In some cases, there may be opportunities to grow a novel crop because the changing climate makes production possible in areas not previously suitable. This may lead to hosts for plant pathogens that were previously unavailable.
In other cases, the cultivation of novel crops arises from other, often political inducements: for example, the requirement that each country in the European Union must derive a certain percentage of its energy use from renewable sources. Hence, there has been a diversion of food crop (cereal) production to bioenergy use, and the planting of non-food crops such as willow, and other short-rotation coppiced trees, and grasses such as Miscanthus for such use. As pointed out by Stewart and Cromey (2010), new disease threats, often from fungal plant pathogens, are likely to emerge among the existing ones. This is likely to occur because the crop is new or because the cultivation method (e.g., high-density monoculture) is different from previous practice. In conservation biology, there is a hefty discussion on the issue of assisted migration (managed relocation), which is thought to be going to be necessary for the many species not able to track the predicted rapid variations in climate, although it will result in artificial shifts in species distributions and ecosystems (Loss et al., 2011). In such debates, little attention has been paid to the likelihood that assisted migration will result in unwanted introductions of, for example, exotic plant pathogens.
Invasion Biology and Biodiversity
Dispersal and invasion of pathogenic fungi are significant not only for crop and other managed plant populations such as commercial forests and grasslands, but also for native plant communities (Alexander, 2010; Mordecai, 2011). In such plant communities, invasive alien plants have long been considered a threat to native biodiversity. Conversely, we now admit that native fungal diseases are important components of healthy forests (Ostry and Laflamme, 2009). However, invasive plant pathogens (either introduced with alien plant invaders or through transfer from plants in trade) can pose a threat to ecosystems (Busby and Canham, 2011; Evans and Finkral, 2010; Holzmüller et al., 2010; Loo, 2009; Newcombe and Dugan, 2010; Orwig, 2002).
Phytophthora ramorum is one such example. On the West Coast of the United States, the pathogen affects a range of forest tree and shrub species, from bay laurel to redwoods and tanoaks. In the United Kingdom (U.K.), the main host in woodlands that supports sporulation of Phytophthora ramorum is Rhododendrum ponticum, itself an invasive plant against which eradication efforts have been targeted, whereas the main native tree species that suffers mortality is beech. The heathland environment and its biodiversity, notably Vaccinium and heather species, are also at risk (Harwood et al., 2009). More worryingly, the pathogen has now been discovered affecting large areas of commercially planted Japanese larch (Larix kaempferi), introduced into the United Kingdom in the 19th century. Japanese larch also supports sporulation of Phytophthora ramorum. Consequently, this poses a threat not only to commercial production, but also to neighboring native trees not previously thought to be susceptible, or never tested (Brasier and Webber, 2010). As has happened in California, infection, tree mortality, and disease spread affect, and in turn are impacted by, forest composition and other aspects such as landscape connectivity, proximity to infected plant nurseries, and woodland history (Davis et al., 2010; Xu et al., 2009).
This example supports the hypothesis mentioned in the Introduction that emerging fungal diseases often result from host shift speciation within a particular ecological context (Giraud et al., 2010). The impact on native biodiversity can usefully be seen from this viewpoint. In return, reduced biodiversity frequently increases disease transmission, including in plants, despite the observation that areas of high endemic biodiversity may harbor a greater source pool of pathogens (Keesing et al., 2010). In general, conserving diverse plant communities reduces the incidence of disease (Pautasso et al., 2005; Turnbull et al., 2010). Of course there are two-way interactions between wild and crop plants, as noted earlier in the case of chickpea and native Cicer. Macrophomina phaseolina affects a broad range of native plant communities and crops in Kansas. Saleh et al. (2010) found that isolates from crops grown in the vicinity of plants in tallgrass prairie could interchange with the more diverse wild isolates with potential dangers arising from novel recombinations.
Irrespective of how an initial introduction occurred—through natural or
human-mediated means—invasion into endemic communities depends very much on ecological circumstances. For example, the origins of fungi in New Zealand are diverse, a few ancient with many more recent arrivals. Some more recent arrivals have evolved into local endemic species, whereas others are maintained through regular transoceanic arrivals (Johnston, 2010). Since 1980 the number of fungal species (saprobic, mycorrhizal, and pathogenic) has doubled. The kinds of fungi present in New Zealand are constantly changing as a consequence of historical changes in plant and animal biota as well as those exotic fungal species that have become naturalized. The fungal biodiversity of New Zealand may appear today as a rather special case in its exotic nature, but could well become the norm for many other countries if current trends in global trade and carbon emissions will carry on unrelented.
The rates of spread and ecological impact of an introduced plant pathogen through a natural plant community depend on the phylogenetic structure of the plant community (Gilbert and Webb, 2007). Although many of the tropical fungal pathogens investigated by Gilbert and Webb (2007) are polyphagous, the likelihood of any one infecting two species decreases with phylogenetic distance. However, using arbitrary cut-offs, such as plant genus or family, may underestimate the risks of local spread. It would be interesting to integrate this phylogenetic approach with the framework provided by the geographic mosaic theory of coevolution, which was tested for mining moths and leaf rust (Melampsora) of Populus tremula along a latitudinal gradient in Sweden (Albrectsen et al., 2010). Climate change can be expected to lead to twists in geographic clines of parasite damage, as well as to shifts in the phylogenetic signal of plant host susceptibility, but it is likely that phylogenetic and spatial structure will still be present even in novel communities and climates.
Much of this overview has been concerned with dispersal and invasion of fungal plant pathogens into crop plants or forest trees (Liebhold et al., 1995). Grassland constitutes a large proportion of the Earth’s terrestrial ecosystems, with varying levels of human exploitation (Ellis et al., 2010). Grassland ecosystems have been less studied than crops or forests in the context of climate change and plant health. Reductions in grassland plant richness, a measure of plant biodiversity, appear to increase vulnerability to the spread of fungal plant diseases (Knops et al., 1999), with aspects of global change, especially elevated CO2 and nitrogen addition, increasing the pathogen load, suggesting this is an important mechanism by which global change affects grassland ecosystems.
Concluding Comments
In cropland, forests, and grasslands, experimentation is needed at the landscape level to investigate plant health adaptation approaches to climate change (Holdenrieder et al., 2004). Similar experimentation, if accompanied by subsequent data analysis and dissemination, is also likely to be beneficial for pathway
control and diagnostic systems (Albers et al., 2010; Norton and Taylor, 2010). Monitoring of plant pathosystems and associated organisms under changing conditions is key to refine models, so as to successfully predict further changes (Luedeling et al., 2011). Similarly, review of the burgeoning literature on the subjects of climate change, land use, and plant health is essential and should be funded on a long-term basis (Jeger and Pautasso, 2008; Pautasso et al., 2010a). Analyses of past and current trends and simulations of future plant health scenarios are likely to be more influential and effective if accompanied by stakeholder involvement and interactions with policy experts (Dwyer, 2011; Mills et al., 2011). Climate change and other global change drivers are not the only factors involved in plant disease epidemiology, so it is still important to consider the important role of, for example, improvements in agronomic and silvicultural practices in reducing current disease potential, whether or not the climate is likely to become more conducive to a certain chronology of diseases in the coming decades (Savary et al., 2011). Nonetheless, it is time to start adapting to the challenges to plant health posed by the future climate change scenarios (Juroszek and von Tiedemann, 2011; Legrève and Duveiller, 2010; Olesen et al., 2011; Planinšek et al., 2011).
The emergence of fungal plant pathogens, facilitated by climate change and globalization, will challenge our ability to meet future food demands for a growing population and to achieve sustainable environments that provide the ecosystem services on which societies depend. Policies informed by science and an infrastructure that supports plant health are prerequisites to that future.
Acknowledgments
Many thanks to C. Brasier, B. D. L. Fitt, M. Garbelotto, K. Garrett, O. Holdenrieder, M. W. Shaw, and J. Webber for insights and discussions. This overview was partly funded by the Rural Economy and Land Use Programme (RELU), U.K.
References
Agosta, S. J., N. Janz, and D. R. Brooks. 2010. How specialists can be generalists: Resolving the “parasite” paradox and implications for emerging infectious disease. Zoologia 27:151–162.
Albers, H. J., C. Fischer, and J. N. Sanchirico. 2010. Invasive species management in a spatially heterogeneous world: Effects of uniform policies. Resource and Energy Economics 32:483–499.
Albrectsen, B. R., J. Witzell, K. M. Robinson, S. Wulff, V. M. C. Luquez, R. Ågren, and S. Jansson. 2010. Large scale geographic clines of parasite damage to Populus tremula. L. Ecography 33:483–493.
Alexander, H. M. 2010. Disease in natural plant populations, communities, and ecosystems: Insights into ecological and evolutionary processes. Plant Disease 94:492–503.
Andrade, D., Z. T. Pan, W. Dannevik, and J. Zidek. 2009. Modeling soybean rust spore escape from infected canopies: Model description and preliminary results. Journal of Applied Meteorology and Climatology 48:789–803.
Attorre, F., M. Alfò, M. De Sanctis, F. Francescani, R. Valenti, M. Vitale, and F. Bruno. 2011. Evaluating the effects of climate change on tree species abundance and distribution in the Italian peninsula. Applied Vegetation Science 14:242–255.
Aukema, J. E., D. G. McCullough, B. Von Holle, A. M. Liebhold, K. Britton, and S. J. Frankel. 2010. Historical accumulation of nonindigenous forest pests in the continental United States. Bioscience 60:886–897.
Awad, A. H. A. 2005. Vegetation: A source of air fungal bio-contaminant. Aerobiologia 21:53–61.
Aylor, D. E., D. G. Schmale, E. J. Shields, M. Newcomb, and C. J. Nappo. 2011. Tracking the potato late blight pathogen in the atmosphere using unmanned aerial vehicles and Lagrangian modeling. Agriculture and Forest Meteorology 151:251–260.
Baeten, L., P. De Frenne, K. Verheyen, B. J. Graae, and M. Henry. 2010. Forest herbs in the face of global change: A single-species-multiple-threats approach for Anemone nemorosa. Plant Ecology and Evolution 143:19–30.
Barba, M., I. Van den Bergh, A. Belisario, and F. Beed. 2010. The need for culture collections to support plant pathogen diagnostic networks. Research in Microbiology 161:472–479.
Beggs, P. J. 2010. Adaptation to impacts of climate change on aeroallergens and allergic respiratory diseases. International Journal of Environmental Research and Public Health 7:3006–3021.
Berger, C. N., S. V. Sodha, R. K. Shaw, P. M. Griffin, D. Pink, P. Hand, and G. Frankel. 2010. Fresh fruit and vegetables as vehicles for the transmission of human pathogens. Environmental Microbiology 12:2385–2397.
Boa, E. 2007. Plant healthcare for poor farmers: An introduction to the work of the Global Plant Clinic. APSnet http://www.apsnet.org/publications/apsnetfeatures/Pages/PoorFarmers.aspx (accessed June 13, 2011).
Brasier, C. M. 2008. The biosecurity threat to the U.K. and global environment from international trade in plants. Plant Pathology 57:792–808.
Brasier, C., and J. Webber. 2010. Sudden larch death. Nature 466:824–825.
Britton, K. O., P. White, A. Kramer, and G. Hudler. 2010. A new approach to stopping the spread of invasive insects and pathogens: Early detection and rapid response via a global network of sentinel plantings. New Zealand Journal of Forestry Science 40:109–114.
Brown, J. K. M., and M. S. Hovmøller. 2002. Aerial dispersal of pathogens at the global and continental scale and its impact on plant disease. Science 297:537–541.
Brown, J. K. M., M. S. Hovmøller, R. A. Wyand, and D.-Z. Yu. 2002. Oases in the desert: Dispersal and host specialization of biotrophic fungal pathogens of plants. In Dispersal, edited by J. M. Bullock, R. E. Kenwood, and R. S. Hails. Oxford, England: Blackwell Science. Pp. 395–409.
Busby, P. E., and C. D. Canham. 2011. An exotic insect and pathogen disease complex reduces aboveground tree biomass in temperate forests of eastern North America. Canadian Journal of Forest Research 41:401–411.
Butterworth, M. H., M. A. Semenov, A. Barnes, D. Moran, J. S. West, and B. D. L. Fitt. 2010. North–South divide: Contrasting impacts of climate change on crop yields in Scotland and England. Journal of the Royal Society Interface 7:123–130.
Camill, P., L. Chihara, B. Adams, C. Andreass, A. Barry, S. Kalim, J. Limmer, M. Mandell, and G. Rafert. 2010. Early life history transitions and recruitment of Picea mariana in thawed boreal permafrost peatlands. Ecology 91:448–459.
Chakraborty, S., and A. C. Newton. 2011. Climate change, plant diseases and food security: An overview. Plant Pathology 60:2–14.
Chapin, F. S., A. D. McGuire, R. W. Ruess, T. N. Hollingsworth, M. C. Mack, J. F. Johnstone, E. S. Kasische, E. S. Euskirchen, J. B. Jones, M. T. Jorensen, K. Kielland, G. P. Kofinas, M. R. Turetsky, J. Yarie, A. H. Lloyd, and D. L. Taylor. 2010. Resilience of Alaska’s boreal forest to climatic change. Canadian Journal of Forest Research 40:1360–1370.
Compant, S., M. G. A. van der Heijden, and A. Sessitsch. 2010. Climate change effects in beneficial plant–microorganism interactions. FEMS Microbiology Ecology 73:197–214.
Conway, D., and E. L. F. Schipper. 2011. Adaptation to climate change in Africa: Challenges and opportunities identified. Global Environmental Change 21:227–237.
Crooks, J. A. 2005. Lag times and exotic species: The ecology and management of biological invasions in slow-motion. Ecoscience 12:316–329.
Dallafior, T. N., and A. Sesartic. 2010. Global fungal spore emissions, review and synthesis of literature data. Biogeosciences Discussions 7:8445–8475.
D’Amato, G., and L. Cecchi. 2008. Effects of climate change on environmental factors in respiratory allergic diseases. Clinical & Experimental Allergy 38:1264–1274.
Davis, F. W., M. Borchert, R. K. Meentenmeyer, A. Flint, and D. M. Rizzo. 2010. Pre-impact forest composition and ongoing tree mortality associated with sudden oak death in the Big Sur region, California. Forest Ecology and Management 259:2342–2354.
Dehnen-Schmutz, K., O. Holdenrieder, M. J. Jeger, and M. Pautasso. 2010. Structural change in the international horticultural industry: Some implications for plant health. Scientia Horticulturae 125:1–15.
Del Ponte, E. M., and P. D. Esker. 2008. Meteorological factors and Asian soybean rust epidemics—a systems approach and implications for risk assessment. Scientia Agricola 65:88–97.
Deliopoulos, T., P. S. Kettlewell, and M. C. Hare. 2010. Fungal disease suppression by inorganic salts: A review. Crop Protection 29:1059–1075.
Desprez-Loustau, M.-L., R. Courtecuisse, C. Robin, C. Husson, P.-A. Moreau, D. Blancard, M.-A. Selosse, B. Lung-Escarmant, D. Piou, and I. Sache. 2010. Species diversity and drivers of spread of alien fungi (sensu lato) in Europe with a particular emphasis on France. Biological Invasions 12:157–172.
Dixon, B. 2005. Trees become casualties of war. The Lancet 5:469.
Dixon, L. J., R. L. Schlub, K. Pernezny, and L. E. Datnof. 2009. Host specialization of Corynespora cassiicola. Phytopathology 99:1015–1027.
Dwyer, J. 2011. U.K. land use futures: Policy influence and challenges for the coming decades. Land Use Policy 28(4):674–683. doi:10.1016/j.landusepol.2010.12.002.
Eastburn, D. M., M. M. Degennaro, E. H. Delucial, O. Demody, and A. J. McElrone. 2009. Elevated atmospheric carbon dioxide and ozone alter soybean diseases at SoyFACE. Global Change Biology 16:320–330.
Ellis, E. C., K. K. Goldewijk, S. Siebert, D. Lightman, and N. Ramankutty. 2010. Anthropogenic transformation of the biomes, 1700 to 2000. Global Ecology & Biogeography 19:586–606.
Ersek, T., and O. K. Ribeiro. 2010. An annotated list of new Phytophthora species described post 1996. Global Change Biology Acta Phytopathologica et Entomologica Hungarica 45:251–266.
Essl, F., S. Dullinger, W. Rabitsch, P. E. Hulme, K. Hulber, V. Jarosik, I. Kleinbauer, F. Krausmann, I. Kuhn, W. Nentwig, M. Vila, P. Genovesi, F. Gherardi, M.-L. Desprez-Loustau, A. Roques, and P. Pysek. 2011. Socioeconomic legacy yields an invasion debt. Proceedings of the National Academy of Sciences, USA 108:203–207.
Evans, A. M., and A. J. Finkral. 2010. A new look at spread rates of exotic diseases in North American forests. Forest Science 56:453–459.
Fitt, B. D. L., N. Evans, P. Gladders, D. J. Hughes, J. W. Madwick, M. J. Jeger, J. A. Townsend, J. A. Turner, and J. S. West. 2010. Climate change and arable crop disease control; mitigation and adaptation. Proceedings of the 16th International Reinhardsbrunn Symposium, April 25–29. In press.
Fitt, B. D. L., B. A. Fraaije, P. Chandramohan, and M. W. Shaw. 2011. Impacts of changing air composition on severity of arable crop disease epidemics. Plant Pathology 60:44–53.
Fleischmann, F., S. Raidl, and W. F. Osswald. 2010. Changes in susceptibility of beech (Fagus sylvatica) seedlings towards Phytophthora citricola under the influence of elevated atmospheric CO2 and nitrogen fertilization. Environmental Pollution 158:1051–1060.
Frank, K. L., B. W. Geils, L. S. Kalkstein, and H. W. Thistle. 2008. Synoptic climatology of the long-distance dispersal of white pine blister rust II. Combination of surface and upper-level conditions. International Journal of Biometeorology 57:653–666.
Frenkel, O., T. L. Peever, M. I. Chilvers, H. Ozkilinc, C. Can, S. Abbo, D. Shtienberg, and A. Sherman. 2010. Ecological genetic divergence of the fungal pathogen Didymella rabiei on sympatric wild and domesticated Cicer spp. (Chickpea). Applied and Environmental Microbiology 76:30–39.
Fry, W. E., and S. B. Goodwin. 1997. Resurgence of the Irish potato famine fungus. Bioscience 47:363–370.
Gange, A. C., E. G. Gange, A. B. Mohammad, and L. Boddy. 2011. Host shifts in fungi caused by climate change? Fungal Ecology 4:184–190.
Garbelotto, M., R. Linzer, G. Nicolotti, and P. Gonthier. 2010. Comparing the influences of ecological and evolutionary factors on the successful invasion of a fungal forest pathogen. Biological Invasions 12:943–957.
Ghini, R., E. Hamada, and W. Bettiol. 2008. Climate change and plant diseases. Scientia Agricola 65:98–107.
Ghini, R., W. Bettiol, and E. Hamada. 2011. Diseases in tropical and plantation crops as affected by climate changes: Current knowledge and perspectives. Plant Pathology 60:122–132.
Gilbert, G. S., and C. O. Webb. 2007. Phylogenetic signal in plant pathogen—host range. Proceedings of the National Academy of Sciences, USA 104:4979–4983.
Giraud, T., P. Gladieux, and S. Gavrilets. 2010. Linking the emergence of fungal plant diseases with ecological speciation. Trends in Ecology & Evolution 25:387–395.
Gladieux, P., E. J. Byrnes, G. Aguiletta, M. C. Fisher, J. Heitman, and T. Giraud. 2011. Epidemiology and evolution of fungal pathogens in plants and animals. In Genetics and evolution of infectious diseases, edited by M. Tibayrenc. Elsevier Inc. Pp. 59–132.
Goellner, K., M. Loehrer, C. Langenbach, U. Conrath, E. Koch, and U. Schaffrath. 2010. Phakopsora pachyrhizi, the causal agent of Asian soybean rust. Molecular Plant Pathology 11:169–177.
Gonthier, P., G. Nicolotti, R. Linzer, F. Guglielmo, and M. Garbelotto. 2007. Invasion of European pine stands by a North American forest pathogen and its hybridization with a native interfertile taxon. Molecular Ecology 16:1389–1400.
Grulke, N. E. 2011. The nexus of host and pathogen phenology: Understanding the disease triangle with climate change. New Phytologist 189:8–11.
Guillemaud, T., M. Ciosi, E. Lombaert, and A. Estoup. 2011. Biological invasions in agricultural settings: Insights from evolutionary biology and population genetics. Comptes Rendus Biologies 334:237–246.
Haq, M., M. A. T. Mia, M. F. Rabbi, and M. A. Ali. 2011. Incidence and severity of rice diseases and insect pests in relation to climate change. In Climate change and food security in South Asia, edited by R. Lal, M. V. K. Sivakumar, S. M. A. Faiz, A. H. M. M. Rahman, and K. R. Islam. Berlin, Germany: Springer. Pp. 445–457.
Harwood, T. D., X. Xu, M. Pautasso, M. J. Jeger, and M. W. Shaw. 2009. Epidemiological risk assessment using linked network and grid based modelling: Phytophthora ramorum and P. kernoviae in the U.K. Ecological Modelling 220(23):3353–3361.
Hindorf, H. 1998. Current diseases of Coffea arabica and C. canephora in East Africa causing crop losses. Proceedings of the International Symposium on Crop Protection 50:861–865.
Holdenrieder, O., M. Pautasso, P. J. Weisberg, and D. Lonsdale. 2004. Tree diseases and landscape processes: The challenge of landscape pathology. Trends in Ecology & Evolution 19:446–452.
Holzmüller, E. J., S. Jose, and M. A. Jenkins. 2010. Ecological consequences of an exotic fungal disease in eastern U.S. hardwood forests. Forest Ecology and Management 259:1347–1353.
Horvath, Z. 2003. Damage in corn production and in hybrid multiplication caused by species of the Anthicidae (Coleoptera) family. Cereal Research Communications 31:421–427.
Hovmøller, M. S., A. H. Yahyaoui, E. A. Milus, and A. F. Justesen. 2008. Rapid global spread of two aggressive strains of a wheat rust fungus. Molecular Ecology 17:3818–3826.
Hunt, R. S., B. W. Geils, and K. E. Hummer. 2010. White pines, Ribes and blister rust: Integration and action. Forest Pathology 40:402–417.
Hyde, K. D., P. Chomnunti, P. W. Crous, J. Z. Groenewald, U. Damm, T. W. Ko Ko, R. G. Shivas, B. A. Summerell, and Y. P. Tan. 2010. A case for re-inventory of Australia’s plant pathogens. Persoonia 25:50–60.
Ibanez, I., R. B. Primack, A. J. Miller-Rushing, E. Ellwood, H. Higuchi, S. D. Lee, H. Kobori, and J. A. Silander. 2010. Forecasting phenology under global warming. Philosophical Transactions B 365:3247–3260.
Jeger, M. J., and M. Pautasso. 2008. Plant disease and global change—the importance of long-term data sets. New Phytologist 177:8–11.
Johnston, P. R. 2010. Causes and consequences of changes to New Zealand’s fungal biota. New Zealand Journal of Ecology 34:175–184.
Jones, A. M., and R. M. Harrison. 2004. The effects of meteorological factors on atmospheric bioaerosol concentrations—a review. Science of the Total Environment 326:151–180.
Juroszek, P., and A. von Tiedemann. 2011. Potential strategies and future requirements for plant disease management under a changing climate. Plant Pathology 60:100–112.
Kauserud, H., E. Heegaard, M. A. Semenov, L. Boddy, R. Halvorsen, L. C. Stige, T. H. Sparks, A. C. Gange, and N. C. Stenseth. 2010. Climate change and spring-fruiting fungi. Proceedings of the Royal Society B 277:1169–1177.
Keesing, F., L. K. Belden, P. Daszak, A. Dobson, C. D. Harvell, R. D. Holt, P. Hudson, A. Jolles, K. E. Jones, C. E. Mitchell, S. S. Myers, T. Bogich, and R. S. Ostfeld. 2010. Impacts of biodiversity on the emergence and transmission of infectious diseases. Nature 468:647–652.
Khan, H. R., J. G. Paull, K. H. M. Siddique, and F. L. Stoddard. 2010. Faba bean breeding for drought-affected environments: A physiological and agronomic perspective. Field Crops Research 115:279–286.
Kliejunas, J. T., B. W. Geils, M. J. Glaeser, E. M. Goheen, P. Hennon, M.-S. Kim, H. Kope, J. Stone, R. Sturrock, and S. J. Frankel. 2008. Climate and forest diseases of Western North America: A literature review. PSW-GTR: USDA Forest Service.
Knops, J. M. H., D. Tilman, N. M. Haddad, S. Naeem, C. E. Mitchell, J. Haarstad, M. E. Ritchie, K. M. Howe, P. B. Reich, E. Siemann, and J. Groth. 1999. Effects of plant species richness on invasion dynamics, disease outbreaks, insect abundances and diversity. Ecology Letters 2:286–293.
La Porta, N., P. Capretti, I. M. Thomsen, R. Kasanen, A. M. Hietala, and K. Von Weissenberg. 2008. Forest pathogens with higher damage potential due to climate change in Europe. Canadian Journal of Plant Pathology 30:177–195.
Lacey, J., M. E. Lacey, and B. D. L. Fitt. 1997. Philip Herries Gregory—1907–1986: Pioneer aerobiologist, versatile mycologist. Annual Review of Phytopathology 35:1–14.
Lansink, A. O. 2011. Public and private roles in plant health management. Food Policy 366:166–170.
Legrève, A., and E. Duveiller. 2010. Preventing potential disease and pest epidemics under a changing climate. In Climate change and crop production, edited by M. P. Reynolds. Wallingford, UK: CAB International. Pp. 50–70.
Liebhold, A. M., W. L. MacDonald, L. Bergdahl, and V. C. Mastro. 1995. Invasion by exotic forest pests: A threat to forest ecosystems. Forest Science Monographs 30:1–49.
Lilja, A., M. Poteri, R.-L. Petaisto, R. Rikala, T. Kurkelo, and R. Kasanen. 2010. Fungal diseases in forest nurseries in Finland. Silva Fennica 44:525–545.
Loo, J. A. 2009. Ecological impacts of non-indigenous invasive fungi as forest pathogens. Biological Invasions 11:81–96.
Loss, S. R., L. A. Terwilliger, and A. C. Peterson. 2011. Assisted colonization: Integrating conservation strategies in the face of climate change. Biological Conservation 144:92–100.
Luck, J., M. Spackman, A. Freeman, P. Trebicki, W. Griffiths, K. Finlay, and S. Chakraborty. 2011. Climate change and diseases of food crops. Plant Pathology 60:113–121.
Luedeling, E., K. P. Steinmann, M. Zhang, P. H. Brown, J. Grant, and E. H. Girvetz. 2011. Climate change effects on walnut pests in California. Global Change Biology 17:228–238.
MacKay, D. J. C. 2008. Sustainable energy—without the hot air. Cambridge: UIT.
MacLeod, A., M. Pautasso, M. J. Jeger, and R. Haines-Young. 2010. Evolution of the international regulation of plant pests and challenges for future plant health. Food Security 2:49–70.
Madden, L. V., and M. Wheelis. 2003. The threat of plant pathogens as weapons against U.S. crops. Annual Review of Phytopathology 41:155–176.
Magyar, D., G. Frenguelli, E. Bricchi, E. Tedeschini, P. Csontos, D.-W. Li, and J. Bobvos. 2009. The biodiversity of air spora in an Italian vineyard. Aerobiologia 25:99–109.
Marçais, B., M. Kavkova, and M.-L. Desprez-Loustau. 2009. Phenotypic variation in the phenology of ascospore production between European populations of oak powdery mildew. Annals of Forest Science 66:814.
Matesanz, S., A. Escudero, and F. Valladares. 2009. Impact of three global change drivers on a Mediterranean shrub. Ecology 90:2609–2621.
McElrone, A. J., J. G. Hamilton, A. J. Krafnick, M. Aldea, R. G. Knepp, and E. H. DeLucia. 2010. Combined effects of elevated CO2 and natural climatic variation on leaf spot diseases of redbud and sweetgum trees. Environmental Pollution 158:108–114.
Miller, S. A., F. D. Beed, and C. Lapierre Harmon. 2009. Plant disease diagnostic capabilities and networks. Annual Review of Phytopathology 47:15–38.
Mills, P., K. Dehnen-Schmutz, B. Ilbery, M. Jeger, G. Jones, R. Little, A. MacLeod, S. Parker, M. Pautasso, S. Pietravalle, and D. Maye. 2011. Integrating natural and social science perspectives on plant disease risk, management and policy formulation. Philosophical Transactions B 366:2035–2044.
Mitchell, C. E., and A. G. Power. 2003. Release of invasive plants from fungal and viral pathogens. Nature 421:625–627.
Money, N. P. 2007. The triumph of fungi: A rotten history. Oxford: Oxford University Press.
Montarry, J., D. Andrivon, I. Glais, R. Corbiere, G. Mialdea, and F. Delmotte. 2010. Microsatellite markers reveal two admixed genetic groups and an ongoing displacement within the French population of the invasive plant pathogen Phytophthora infestans. Molecular Ecology 19: 1965–1977.
Mordecai, E. 2011. Pathogen impacts on plant communities: Unifying theory, concepts, and empirical work. Ecological Monographs. In press. doi:10.1890/10–2241.1
Moslonka-Lefebvre, M., M. Pautasso, and M. J. Jeger. 2009. Disease spread in small-size directed networks: Epidemic threshold, correlation between links to and from nodes, and clustering. Journal of Theoretical Biology 260:402–411.
Moslonka-Lefebvre, M., A. Finley, I. Dorigatti, K. Dehnen-Schmutz, T. Harwood, M. J. Jeger, X. M. Xu, O. Holdenrieder, and M. Pautasso. 2011. Networks in plant epidemiology: From genes to landscapes, countries and continents. Phytopathology 101:392–403.
Mundt, C. C., K. E. Sackett, L. D. Wallace, C. Cowger, and J. P. Dudley. 2009. Long-distance dispersal and accelerating waves of disease: Empirical relationships. The American Naturalist 73:456–466.
Newcombe, G., and F. M. Dugan. 2010. Fungal pathogens of plants in the homogocene. In Molecular identification of fungi, edited by Y. Gherbawy and K. Voigt. Berlin, Germany: Springer. Pp. 3–34.
Newton, A. C., T. Akar, J. P. Baresel, P. J. Bebbeli, E. Bettencourt, K. V. Bladenopoulos, J. H. Czembor, D. A. Fasoula, A. Katsiotis, K. Koutis, M. Koutsika-Sotiriou, G. Kovacs, H. Larsson, M. A. A. P. de Carvalho, D. Rubiales, J. Russell, T. M. M. Dos Santos, and M. C. V. Patto. 2010. Cereal landraces for sustainable agriculture. Agronomy for Sustainable Development 30:237–269.
Norton, G., and M. Taylor. 2010. What pest is that? Recent developments in digital pest diagnostics. Outlooks on Pest Management 21:236–238.
Olesen, J. E., M. Trnka, K. C. Kersebaum, A. O. Skjelvag, B. Seguin, P. Peltonen-Sainio, F. Rossi, J. Kozyra, and F. Micale. 2011. Impacts and adaptation of European crop production systems to climate change. European Journal of Agronomy 34:96–112.
Orwig, D. A. 2002. Ecosystems to regional impacts of introduced pests and pathogens: Historical context, questions and issues. Journal of Biogeography 29:1471–1474.
Ostry, M. E., and G. Laflamme. 2009. Fungi and diseases—natural components of healthy forests. Botany 87:22–25.
Paajanen, R., R. Julkunen-Tiitto, L. Nybakken, M. Petrelius, R. Tegelberg, J. Pusenius, M. Rousi, and S. Kellomäki. 2011. Dark-leaved willow (Salix myrsinifolia) is resistant to three-factor (elevated CO2, temperature and UV-B-radiation) climate change. New Phytologist 190:161–168.
Padmanabhan, S. Y. 1973. The great Bengal famine. Annual Review of Phytopathology 11:11–24.
Palm, M. E. 1999. Mycology and world trade: A view from the front line. Mycologia 91:1–12.
Parks, C. G., and P. Bernier. 2010. Adaptation of forests and forest management to changing climate with emphasis on forest health: A review of science, policies and practices. Forest Ecology and Management 259:657–659.
Pautasso, M. 2009. Geographical genetics and the conservation of forest trees. Perspectives in Plant Ecology, Evolution and Systematics 11:157–189.
Pautasso, M., O. Holdenrieder, and J. Stenlid. 2005. Susceptibility to fungal pathogens of forests differing in tree diversity. In Forest diversity and function: Temperate and boreal systems, edited by M. Scherer-Lorenzen, C. Koerner, and E. D. Schulze. Berlin, Germany: Springer. Pp. 263–289.
Pautasso, M., K. Dehnen-Schmutz, O. Holdenrieder, S. Pietravalle, N. Salama, M. J. Jeger, E. Lange, and S. Hehl-Lange. 2010a. Plant health and global change—some implications for landscape management. Biological Reviews 85:729–755.
Pautasso, M., M. Moslonka-Lefebvre, and M. J. Jeger. 2010b. The number of links to and from the starting node as a predictor of epidemic size in small-size directed networks. Ecological Complexity 7:424–432.
Pautasso, M., X.-M. Xu, M. J. Jeger, T. D. Harwood, M. Moslonka-Lefebvre, and L. Pellis. 2010c. Disease spread in small-size directed trade networks: The role of hierarchical categories. Journal of Applied Ecology 47:1300–1309.
Peltzer, D. A., R. B. Allen, G. M. Lovett, D. Whitehead, and D. A. Wardle. 2010. Effects of biological invasions on forest carbon sequestration. Global Change Biology 16:732–746.
Perrings, C., S. Burgiel, M. Lonsdale, H. Mooney, and M. Williamson. 2010. International cooperation in the solution to trade-related invasive species risks. Annals of the New York Academy of Sciences 1195:198–212.
Planinšek, Š., L. Finér, A. del Campo, J. Alcazar, C. Vega-García, D. Dimitrov, and J. Capuliak. 2011. Adjustment of forest management strategies to changing climate. Ecological Studies 212:313–329.
Pugliese, M., M. L. Gullino, and A. Garibaldi. 2010. Effects of elevated CO2 and temperature on interactions of grapevine and powdery mildew: First results under phytotron conditions. Journal for Plant Diseases and Plant Protection 117:9–14.
Rivas, G. G., M. F. Zapater, C. Abadie, and J. Carlier. 2004. Founder effects and stochastic dispersal at the continental scale of the fungal pathogen of bananas Mycosphaerella fijiensis. Molecular Ecology 13:471–482.
Roos, J., R. Hopkins, A. Kvarnheden, and C. Dixelius. 2011. The impact of global warming on plant diseases and insect vectors in Sweden. European Journal of Plant Pathology 129:9–19.
Roper, M., A. Seminara, M. M. Band, A. Cobb, H. R. Dillard, and A. Pringle. 2010. Dispersal of fungal spores on a cooperatively generated wind. Proceedings of the National Academy of Sciences, USA 107:17474–17479.
Russell, R., M. Paterson, and N. Lima. 2009. How will climate change affect mycotoxins in food? Food Research International 43:1902–1914.
Saleh, A. A., H. U. Ahmed, T. C. Todd, S. E. Travers, K. A. Zeller, J. F. Leslie, and K. A. Garrett. 2010. Relatedness of Macrophomina phaseolina isolates from tallgrass prairie, maize, soybean and sorghum. Molecular Ecology 19:79–91.
Savage, D., M. J. Barbetti, W. J. MacLeod, M. U. Salam, and M. Renton. 2010. Timing of propagule release significantly alters the deposition area of resulting aerial dispersal. Diversity and Distributions 16:288–299.
Savary, S., A. Mila, L. Willocquet, P. Esker, O. Carisse, and N. McRoberts. 2011. Risk factors for crop health under global change and agricultural shifts: A framework of analyses using rice in tropical and subtropical Asia as a model. Phytopathology 101:696–709.
Scherrer, D., and C. Körner. 2011. Topographically controlled thermal-habitat differentiation buffers alpine plant diversity against climate warming. Journal of Biogeography 38:406–416.
Schulz, N. B. 2004. The transport system and society’s metabolism in the U.K. Population and Environment 26:133–155.
Scirè, M., E. Motta, and L. D’Amico. 2011. Behaviour of Heterobasidion annosum and H. irregulare isolates from central Italy in inoculated Pinus pinea seedlings. Mycological Progress 10:85–91.
Shaw, M. W., and T. M. Osborne. 2011. Geographic distribution of plant pathogens in response to climate change. Plant Pathology 60:31–43.
Shin, J.-W., and S.-C. Yun. 2010. Elevated CO2 and temperature effects on the incidence of four major chilli pepper diseases. Plant Pathology Journal 26:178–184.
Singh, D. P., D. Backhouse, and P. Kristiansen 2009. Interactions of temperature and water potential in displacement of Fusarium pseudograminearum from cereal residues by fungal antagonists. Biological Control 48:188–195.
Singh, S., S. Davey, and M. Cole. 2010. Implications of climate change for forests, vegetation and carbon in Australia. New Zealand Journal of Forestry Science 40:141–152.
Skelsey, P., W. A. H. Rossing, G. J. T. Kessel, and W. van der Werf. 2009. Scenario approach for assessing the utility of dispersal information in decision support for aerially spread plant pathogens, applied to Phytophthora infestans. Phytopathology 99:887–895.
Spijkerboer, H. P., J. E. Benier, D. Jaspers, H. J. Schouten, J. Goudriaan, R. Rabbinge, and W. van der Werf. 2002. Ability of the Gaussian plume model to predict and describe spore dispersal over a potato crop. Ecological Modelling 155:1–18.
Stack, J. P. 2010. Diagnostic networks for plant biosecurity. In Knowledge and technology transfer for plant pathology, edited by N. Hardwick and M. L. Gullino. Dordrecht, Netherlands: Springer. Pp. 59–74.
Stack, J. P., F. Suffert, and M. L. Gullino. 2010. Bioterrorism: A threat to plant biosecurity? In The role of plant pathology in food safety and food security, edited by R. N. Strange and M. L. Gullino. Dordrecht, Netherlands: Springer. Pp. 115–123.
Stewart, A., and M. Cromey. 2010. Identifying disease threats and management practices for bioenergy crops. Current Opinion in Environmental Sustainability 3:75–80.
Stonard, J. F., A. O. Latunde-Dada, Y.-J. Huang, J. S. West, N. Evans, and B. D. L. Fitt. 2010. Geographic variation in severity of phoma stem canker and Leptosphaeria maculans/L. biglobosa populations on U.K. winter oilseed rape (Brassica napus). European Journal of Plant Pathology 126:97–109.
Sturrock, R. N., S. J. Frankel, A. V. Brown, P. E. Hennon, J. T. Kliejunas, K. E. Lewis, J. J. Worrall, and A. J. Woods. 2011. Climate change and forest diseases. Plant Pathology 60:133–149.
Techy, L., D. G. Schmale, and C. A. Woolsey. 2010. Coordinated aerobiological sampling of a plant pathogen in the lower atmosphere using two autonomous unmanned aerial vehicles. Journal of Field Robotics 27:335–343.
Thomas, K. 2010. Climate change and management of cool season grain legume crops. In Impact of climate change on diseases of cool season grain legume crops, edited by S. S. Yadav, D. L. McNeil, R. Redden, and S.A. Patil. Berlin, Germany: Springer. Pp. 99–113.
Thomson, L. J., S. MacFadyen, and A. A. Hoffmann. 2009. Predicting the effects of climate change on natural enemies of agricultural pests. Biological Control 52:296–396.
Tiley, G. E. D. 2010. Biological flora of the British Isles: Cirsium arvense (L.) Scop. Journal of Ecology 98:938–983.
Tubby, K. V., and J. F. Webber. 2010. Pests and diseases threatening urban trees under a changing climate. Forestry 81:451–459.
Turnbull, L. A., J. M. Levine, A. J. F. Fergus, and J. S. Petermann. 2010. Species diversity reduces invasion success in pathogen-regulated communities. Oikos 119:1040–1046.
Tylianakis, J. M., R. K. Didham, J. Bascompte, and D. A. Wardle. 2008. Global change and species interactions in terrestrial ecosystems. Ecology Letters 11:1351–1363.
Woods, A. J., D. Heppner, H. H. Kope, J. Burleigh, and L. Maclauchlan. 2010. Forest health and climate change: A British Columbia perspective. The Forestry Chronicle 86:412–422.
Worrall, J. J., G. C. Adams, and S. C. Tharp. 2010. Summer heat and an epidemic of cytospora canker of Alnus. Canadian Journal of Plant Pathology 32:376–386.
Xu, X. M., P. Nicholson, M. A. Thomsett, D. Simpson, B. M. Cooke, F. M. Doohan, J. Brennan, S. Monaghan, A. Moretti, G. Mule, L. Hornok, E. Beki, J. Tatnell, A. Ritieni, and S. G. Edwards. 2008. Relationship between the fungal complex causing Fusarium head blight of wheat and environmental conditions. Phytopathology 98:69–78.
Xu, X. M., T. D. Harwood, M. Pautasso, and M. J. Jeger. 2009. Spatio-temporal analysis of an invasive plant pathogen (Phytophthora ramorum) in England and Wales. Ecography 32:504–516.
Young, J. M., C. Allen, T. Coutinho, T. Denny, J. Elphinstone, M. Fegan, M. Gillings, T. R. Gottwald, J. H. Graham, N. S. Iacobellis, J. D. Janse, M. A. Jacques, M. M. Lopez, C. E. Morris, N. Parkinson, P. Prior, O. Pruvost, J. Rodrigues Neto, M. Scortichini, Y. Takikawa Y, and C. D. Upper. 2008. Plant–pathogenic bacteria as biological weapons—real threats? Phytopathology 98:1060–1065
Zocca, A., C. Zanini, A. Aimi, G. Frigimelica, N. La Porta, and A. Battisti. 2008. Spread of plant pathogens and insect vectors at the northern range margin of cypress in Italy. Acta Oecologica 33:307–313.
EMERGING FUNGAL DISEASES OF WILD ANIMAL SPECIES
Luis R. Padilla46
Summary
Several fungal diseases of non-domestic animal species have been described as agents of epizootic proportions in wild animals in recent years. The emergence of these diseases has reshaped the understanding of the role of fungi as contagious diseases having an impact on wild animal populations. The recent description of fungal epizootics caused by Batrachochytrium dendrobatidis (Bd) in amphibians worldwide and Geomyces destructans in North American bats has called attention to the factors driving the emergence of these diseases. Two other fungal diseases of wild animals, lobomycosis in dolphins and penicilliosis in wild bamboo rats, have been recognized for their zoonotic potential and highlight the need for comprehensive, multidisciplinary approaches to understanding the ecology and epidemiology of these diseases in wild animal populations. Continued efforts aimed at mitigating the effects of fungal epizootics on wildlife populations and on
______________
46 Department of Animal Health, Smithsonian Conservation Biology Institute.
public health will only be successful through the identification of factors driving the emergence of these diseases across different taxa, and embracing the concept that wildlife may serve as sentinels of ecosystem health.
Introduction
Anthropogenic activities have been the likely driving factors behind the emergence of some diseases in wildlife (Daszak et al., 2000). In some cases, animal species whose status may have been threatened by other factors may now be faced with extinction as the emergent disease spreads through a diminished population. The recent spread of two major fungal epizootic agents, namely Batrachochytrium dendrobatidis and Geomyces destructans in amphibians and bats, respectively, could result in the largest changes to vertebrate populations in recorded history (Frick et al., 2010; Skerrat et al., 2007). The discovery of these fungal agents as major causes of wildlife epizootics in the past decade has revolutionized the way that biologists approach the detection and diagnosis of fungal diseases, challenging the prior misconception that fungal infections only occurred “sporadically or in small outbreaks” and were more important to captive wildlife, where captivity was thought to “increase susceptibility to these diseases” (Burek, 2001).
As novel pathogens have been discovered, or known ones recognized in novel hosts, in novel geographic areas, or with increased incidence, wildlife conservationists have been faced with the emergence of fungal diseases that threaten the status of wild animal populations. The factors driving disease in free-ranging wild animal species can no longer be easily differentiated from those affecting captive wildlife. The different factors have been blurred into a continuum of factors through modern globalization, inadvertent movement of disease, disease vectors, or animals themselves, through the trade of animals across geographical barriers and the mixing of potential hosts of disease that may not have been exposed to each other in their native habitats. Many of these anthropogenic actions have expanded the geographic range of some diseases or removed the natural barriers that had prevented their spread, exposing naïve hosts to pathogens to which they were not previously exposed. Concurrently, many of the environmental factors thought to predispose their captive counterparts to infectious diseases have been identified and minimized through modern captive animal science aimed at reducing stress levels, providing better environmental conditions, and through better quarantine, improving disease screening and recognition procedures.
Institutions dedicated to the captive care of animals for conservation purposes often encounter infectious diseases that would otherwise go unreported in those species. These provide a useful baseline of understanding the host–pathogen relationship systems. Captive animal populations can serve as viable models for recognizing or refining the understanding of disease mechanisms in individual hosts, which may not be possible in free-ranging animals. For in-
stance, in the 1990s, investigations by pathologists at the Smithsonian’s National Zoo of an apparently novel fungus affecting captive frogs led to the isolation of Batrachochytrium dendrobatidis (Pessier et al., 1999) and fulfillment of Koch’s postulates (Nichols et al., 2001) in what would subsequently be recognized as the most important infectious disease affecting wild amphibian populations on a global scale.
The widespread recognition of other fungal pathogens and their prevalence in wildlife populations has also changed the understanding of their role as agents of contagious potential and concern to human public health. Concern to human health from fungal diseases carried or propagated by wild animals has brought attention to the role of wildlife and interactions with humans in changing environments as indicators of global health and ecosystem stability. Direct zoonotic potential or a shared susceptibility to disease where wildlife have a high prevalence of infection is worthy of direct public concern, but fungal pathogens where the infection potential is limited to wild animals still carry an inherent cost to the health of an ecosystem and carry indirect impacts to human society that may be difficult to quantify.
This review article summarizes some of the fungal diseases currently known to affect wild animal species that could be encountered by veterinarians, conservationists, or animal professionals. This review is not intended as a comprehensive reference on the subject. Instead, it is meant to be a common point for calling attention to the impact of the recent emergence of specific fungal pathogens in wildlife species within the context of wildlife as sentinel species for recognizing changes to the health of an ecosystem.
Emergent Fungal Pathogens of Wild Animal Species
Batrachochytrium dendrobatidis
Batrachochytrium dendrobatidis is a member of a basal group of fungi, the Chytridiomycota, and the only member known to affect vertebrates. Bd infects amphibian species, with frogs most often reported. This emerging pathogen was discovered in the late 1990s (Berger et al., 1998; Longcore et al., 1999; Pessier et al., 1999) and has been recognized as a significant pathogen partially implicated in the global decline of amphibian populations (Schloegel et al., 2006; Skerrat et al., 2007). It has been retrospectively identified in North American amphibians as early as 1961 (Ouellet et al., 2005). Bd may be the most significant fungal infectious disease agent of vertebrate species, based on the global distribution, wide species host range, pathogenic potential, and ability to cause large-scale mortality.
Molecular genetic evidence from isolates collected from different locations worldwide suggests that recent Bd outbreaks were caused by a pathogen recently disseminated (Morehouse et al., 2003), and anthropogenic spread is suspected
in at least some introductions (Weldon et al., 2004), but the origin is still not conclusively known. Virulence and pathogenic potential varies among isolates, and some may show little or no pathogenicity to their hosts (Berger et al., 2005; Goka et al., 2009), but these host-adapted strains may still serve as reservoirs of strains that are pathogenic to other species. The naturally occurring bacterial flora of the host may produce antifungal proteins that decrease the ability of the fungi to colonize amphibian skin, and these bacteria may improve host survival (Harris et al., 2006). Mortality in affected amphibians is likely associated with physiologic abnormalities (electrolyte loss and osmotic imbalances) caused by damage to the permeable amphibian skin.
Clinical signs in affected frogs can be subtle and not detectable before sudden death (Pessier et al., 1999). Affected frogs show excessive shedding of skin, usually on the legs, feet, and ventrum (Nichols et al., 2001). Molecular techniques have been established as a cost-effective and rapid method to detect the organism in amphibian samples (Boyle et al., 2004; Kriger et al., 2006). Histopathology, showing intralesional organisms in keratinized layers of skin, and cytology of shed skin or imprints showing chytrid organisms have been used for diagnosing infections (Nichols et al., 2001). Bd has no known zoonotic potential.
Chrysosporium Anamorph of Nanniziopsis vriesii (CANV)
Nanniziopsis vriesii is an ascomycetous fungus recognized as a pathogen from skin lesions of reptiles, specifically lizards, snakes, and crocodilians. When cultured, Nanniziopsis vriesii produces an anamorph form that is described in the genus Chrysosporium and is undistinguishable from reptilian isolates (Paré and Sigler, 2006). Mycotic infections may be morphologically misidentified as other species of Chrysosporium, Geotrichum, or even Trichophyton. Historical reports in the literature of reptilian mycoses attributed to unspeciated members of these genera should be interpreted with caution, as CANV infections may have been previously underdiagnosed. CANV deserves special attention as an emerging fungal pathogen of reptiles because clinical disease has been recognized in a multitude of species with increasing frequency (Abarca et al., 2008; Bertelsen et al., 2005; Bowman et al., 2007; Nichols et al., 1999; Paré and Sigler, 2006; Paré et al., 2006; Thomas et al., 2002), but most reports involve captive or farmed reptiles. Unlike opportunistic fungal pathogens, which are often ubiquitous in the host’s environment, the CANV is extremely rare on healthy reptile integument (Paré et al., 2003), suggesting that this organism is a primary fungal pathogen in reptiles.
Preliminary molecular genetic evidence has suggested that the CANV represents a species complex with distinct host affinities (Paré and Sigler, 2006). Until further molecular work is completed, the magnitude of the CANV as a pathogen of free-ranging and captive reptile populations cannot be properly described,
but its potential as an emerging primary fungal pathogen of reptiles cannot be ignored.
Clinical signs vary by species, but when seen in inland bearded dragons (Pogona vitticeps), it causes a yellow skin discoloration dubbed “yellow skin disease” or “yellow fungus disease.” The disease can have a protracted, chronic course involving deep tissues (including muscles and bones) and cause disfigurement in addition to the necrotic skin lesions seen. The corresponding lesions can be diagnosed as CANV through histopathology of lesions and fungal cultures. Histopathology typically shows severe granulomatous, necrotic dermatomycosis with the characteristic intralesional fungal organisms. The CANV forms solitary, single-celled conidia that can be confused with the microconidia of dermatophytes (e.g., Trichophyton), and arthroconidia caused by fragmented hyphae, which may resemble the arthroconidia of Geotrichum or other Chrysosporium (Paré and Sigler, 2006).
A single case report of a mycotic brain abscess attributed to the Chrysosporium anamorph of Nanniziopsis vriesii in an HIV-seropositive Nigerian man has suggested that this reptile pathogen can be a zoonotic agent, although the route or source of exposure or infection in this patient could not be identified (Steininger et al., 2005).
Geomyces destructans
Geomyces destructans is a newly described psychrophilic fungus that affects North American bats during hibernation (Gargas et al., 2009), causing the characteristic fungal white growth after which the syndrome was originally named.
White-nose syndrome (WNS) could more accurately be dubbed bat geomycosis (per Chaturvedi and Chaturvedi, 2011), but the denomination of WNS has been used since its recognition (Blehert et al., 2009). Its impact on wild bat populations has earned it the distinction of being the second most significant vertebrate pathogen in recorded history (after Bd), if only because Bd has been documented in more species, with a wider global impact.
Bat geomycosis is a true emerging disease of epizootic proportion, appearing in upstate New York in 2006 (Blehert et al., 2009), but quickly spreading south and westward and devastating bat populations (Blehert et al., 2009; Frick et al., 2010). Host susceptibility to infection seems to vary with species. The fungus has also been identified in Europe (Puechmaille et al., 2010; Wibbelt et al., 2010), but to date, the severe, widespread mortality seen in North American bats has not been documented. Among North American bat species, either pathologic lesions or G. destructans DNA has been detected in the gray bat (Myotis grisescens), the Indiana bat (M. sodalis), the little brown bat (Myotis lucifugus), the northern long-eared bat (M. septentrionalis), the eastern small-footed bat (M. leibii), the southeastern bat (M. austroriparius), the cave bat (M. velifer), the tricolored bat
(Perimyotis subflavus), and the big brown bat (Eptesicus fuscus) (Foley et al., 2011).
Geomyces destructans infections in bats result in premature arousal from hibernation and abnormal behavior. Although the mechanisms by which death occurs are still under investigation, the inability to forage in the winter after premature arousal (due to lack of prey availability) and direct damage to the wing membranes, resulting in irreversible physiologic and homeostatic deficits from which bats cannot recover (Cryan et al., 2010), are significant factors.
Limiting human access to caves of concern has been a management tool implemented to limit the spread of the Geomyces fungus. One of the first theories to explain the rapid appearance and spread of this pathogen in North American bat populations suggested that the fungus may have been endemically established in European bat populations and that a recent anthropogenic introduction into North America may have led to disease in naïve populations, but this theory has not been tested. Histopathology of rostral muzzle and wing membranes is deemed important to confirm infections and establish true prevalence of this disease (Meteyer et al., 2009), although advanced diagnostic tools are likely being developed and refined.
As bats are significant providers of ecological services, such as insect control and pollination, the extinction or even reduction of bat populations is likely to have economic impacts on society beyond the immediate loss of biodiversity.
Penicillium marneffei
Penicillium marneffei is the only fungus of this genus known to be a primary pathogen of mammals. Penicilliosis marneffei is a systemic fungal disease of wild rodents and humans recognized in northeast India and Southeast Asia (Thailand, the Guangxi region of China, Vietnam, Taiwan, and Hong Kong). Penicillium marneffei was first identified from hepatic lesions in a bamboo rat (Rhizomys sinensis) from Dalat, South Vietnam (Caponi et al., 1956), and was subsequently identified as a human pathogen following an accidental exposure by a researcher (Vanittanakom et al., 2006). It is an emerging human disease, a primary pathogen to bamboo rats, and a threat to public health. Penicilliosis marneffei is third only to tuberculosis and cryptococcosis as the most common opportunistic infections in patients with AIDS in northern Thailand (Vanittanakom et al., 2006), and the source of more than 100 cases per year in the Guanxi region of China (Cao et al., 2011).
Affected rodents show ascites and enlargement of the liver, spleen, and lymph nodes. The fungal pathogens can be identified in multiple organs and ascitic fluid, but is most commonly cultured from the lung of affected rodents (Ajello et al., 1995).
Wild bamboo rats may be good sentinels of this sapronotic disease, and are significant in the ecology of this disease, although their exact role is not com-
pletely understood. Penicillium marneffei has been isolated from the internal organs of four species of bamboo rats (Rhizomys sinensis, Rhizomys pruinosus, Rhizomys sumatrensis, and Cannomys badius) and from the soil associated with their burrows (Vanittanakom et al., 2006). A high prevalence of infection and lesions among wild bamboo rats of the genera Rhizomys and Cannomys suggested that these wild animals may serve as enzootic reservoirs, but the prevalence varies across regions, and a wildlife reservoir has never been conclusively identified (Cao et al., 2011; Deng et al., 1986; Gugnani et al., 2004; Vanittanakom et al., 2006). When sympatric rodents have been sampled, only bamboo rats appear to harbor the organism (Gugnani et al., 2004), suggesting distinct host differences.
Possibly, bamboo rats are exposed from either an unidentified wildlife vector or an environmental source (Vanittanakom et al., 2006), and are not the wildlife hosts of this disease, although they can transmit the disease to humans. The role of wild bamboo rats as amplifiers or dispersers of infectious stages has not been eliminated. The initial case of human infection in 1959 in a researcher who became infected after injection from a needle used in laboratory inoculations (Vanittanakom et al., 2006) underscores the zoonotic risk posed by wildlife.
Although most human cases involve immunosuppressed individuals, infections in humans with normal immunity—and the lack of evidence for immunosuppression in affected bamboo rats—suggests that P. marneffei could be a primary mammalian fungal pathogen (Duong, 1996). A complete understanding of the role of wildlife in this disease and its dynamics in rodent populations is essential to better managing the threat to human health in endemic regions and mitigating the possible anthropogenic spread of this disease through travel, movement of commercial products (some of which may be environmental reservoirs), and the primary trade of wild animals. Penicilliosis marneffei remains one of the most enigmatic emerging fungal diseases of wildlife, and one whose primary zoonotic potential is not fully understood.
Lacazia loboi
Lobomycosis is a zoonotic disease of dolphins caused by Lacazia loboi, a cutaneous fungus in the order Onygenales that has never been cultured (Herr et al., 2001). The disease affects humans and dolphins in tropical and transitional tropical climates. Lobomycosis is endemic in certain human populations in Central and South America, and is likely endemic in regional dolphin populations (Murdoch et al., 2008), although the overall prevalence and many factors of its ecology are still unclear. The disease has been confirmed in two dolphin species, Guiana dolphins (Sotalia guianensis) (Van Bressem et al., 2009) and Atlantic bottlenose dolphins (Tursiops truncatus) (Caldwell et al., 1975), and has been suspected in Indo-Pacific bottlenose dolphins (T. aduncus) (Kiszka et al., 2009).
Transmission between dolphins is likely by direct contact, and affected dolphins show chronic white to pink verrucous, raised lesions that may coalesce
into large plaques or nodules predominantly on dorsal and pectoral fins, the head, fluke, and caudal peduncle (Murdoch et al., 2010). Lesions may be associated with sites of prior trauma, and many affected dolphins have shown impaired adaptive immunity, suggesting that the disease may represent an opportunistic infection in an immunocompromised host (Reif et al., 2008).
Some have suggested that the disease in dolphins is being more frequently reported (Van Bressem et al., 2009) and the reported geographic range may be expanding (Rotstein et al., 2009), making it a true emerging fungal disease of wildlife. In the Indian River lagoon of Florida, measured prevalence was 30 percent in Atlantic bottlenose dolphins (Tursiops truncatus) from the southern part of the lagoon in a 2006 survey (Reif et al., 2006). It was not detected in the northern portion (near Charleston, South Carolina), suggesting that localized geographical factors or environmental stressors may play a significant role in the incidence and distribution of the disease. A subsequent study (Murdoch et al., 2008) suggested that the disease is endemic and not an emerging disease in the Indian River dolphin population. A survey of Guiana dolphins (Van Bressem et al., 2009) in the Paranaguá estuary in Brazil suggested an increasing detection of a lobomycosis-like disease (missing histological confirmation) and suggested that the change was an indication of the health of the marine environment.
Coccidioides immitis and Coccidioides posadasii
Coccidioidomycosis is a systemic fungal disease known to affect a large number of domestic and non-domestic animal species, both in captivity and in the wild. It is a primary pathogen and can affect otherwise healthy individuals, although clinical disease is often seen in animals with recognized limited immune function or underlying disease. Coccidioides immitis is considered endemic to the Western Hemisphere, and is commonly recognized in the southwestern United States in the Sonoran life zone—areas characterized by alkalinic soils and an arid climate. Coccidioidomycosis is also reported in Mexico and Central and South America, although C. posadasii is often implicated in infections outside of the Southwestern United States. Anthropogenic disturbance of soil (e.g., construction), dust storms, and cycles of drought may precede outbreaks, but wind-blown spores have been hypothesized to play a role in cases of affected marine mammals far from endemic areas. Fungal spores may be concentrated around rodent burrows, but wild animals are not likely reservoirs of this disease and contagion is unlikely to occur between animals. Human exposure may occur during necropsies of affected carcasses by aerosolization of spores, but wild animals are likely insignificant in the epidemiology of the disease in wild animal populations.
Coccidioides has affected some captive endangered species disproportionately more than other species, and can be a threat to species with limited immune function. Specifically, the Przewalski’s horse (Equus przewalskii) has been disproportionately affected in areas where the disease may be endemic (Terio et al.,
2003). In one area of southern California, coccidioidomycosis was associated with high incidence of infection and was the leading cause of death in a population of this endangered species, posing a threat to the captive propagation of this species. The disease showed a predilection toward younger male horses, and it has been suggested that the immune system of some of these horses may not respond appropriately to infection with coccidioides (Terio et al., 2003). Limited immune function can be a limitation of propagation of endangered species with a limited genetic pool, and infectious diseases can have disproportionate effects on these populations, even when the epizootic potential is limited.
Actions for Managing and Mitigating the Effects of Fungal Diseases on Wild Animals
A basic understanding of the role of fungal diseases in populations is the most important, fundamental need to manage the spread of these diseases. Public expectation for action in the face of spreading epizootics is high, and basic knowledge gaps must be addressed to develop and implement effective management plans. Although the goal is to correct the product of detrimental anthropogenic influences on the occurrence or emergence of diseases, intentions and action plans should be tempered by the recognition that naturally occurring diseases are and have been a driving evolutionary and ecological force responsible for shaping and adapting community structures in wild populations. Adaptive management strategies should be scientifically sound.
In the absence of proven action plans for the control of specific fungal pathogens, general action plans for the adaptive management of fungal epizootics can be modeled following the template considered for the control of WNS in bats (Foley et al., 2011). These strategies can generally include: continuing disease and population surveillance; limiting further anthropogenic spread through biosecurity and quarantine measures; providing individual animal treatments to increase survival; creating protected populations or rescue populations with the goal of preserving genetic diversity; increasing population resistance to the disease; modifying the environment or environmental sources of exposure; selective culling of populations to limit transmission; and disseminating factual information to address both public perception and foster scientific collaboration and research.
Prospective monitoring and surveillance of disease and populations allows wildlife managers to establish objective criteria that can be used to document their impact following the emergence of a pathogen. In the case of a novel pathogen, such as the Geomyces destructans in North American bats, historical, basic data on the population status of native bat populations has allowed the documentation of changes to the population after the emergence of this disease. By contrast, the lack of historical specimens collected at the time of initial decline of amphibian populations, whose decline has been attributed to chytridiomycosis, has raised unanswerable questions about the cofactors that may have precipitated the
emergence of this fungal pathogen or its true impact on populations (McCallum, 2005). Active surveillance should rely on standardized case definitions, sampling methodologies, and data analysis and centralized sharing of results. Molecular techniques, some of which were in their infancy a mere 10 years ago, have allowed the differentiation of pathogenic strains for different fungal organisms and their relationship to the host and their immune defenses, and a better understanding of the ecology of fungal diseases. For instance, the advancement of molecular techniques as an adjunct to field surveillance techniques has proven valuable in understanding the dynamics of amphibian chytridiomycosis. The characterization and dissemination of reports of fungal diseases in novel hosts or new presentations in known hosts is an implied responsibility of veterinarians and animal biologists working with wildlife species, both in captive or free-ranging situations.
In the face of spreading epizootics, movement and entry restrictions or quarantine regulations are often imposed to halt or slow down the spread of an infectious disease. These restrictions can be as simple as prohibiting (human) visitor access to caves where bat geomycosis has been documented, or as complex as trade regulations and sanctions against the commercial movement of species. Instituting proper quarantine procedures when entering endemic areas and educating visitors and researchers about their own potential as vectors of disease can be effective methods to avoid irresponsible spread of pathogens. Responsible disinfection protocols can be established by scientists repeatedly monitoring affected populations.
Individual animal treatments may not be feasible or effective in halting the large-scale spread of epizootics, but are valuable if the animals treated are being used for captive propagation programs or if treatment is only needed for extremely small populations. Treatment of infected individuals may include antifungal agents, although the efficacy and safety of these for individual fungal pathogens has not always been established, and a specific discussion of antifungal agents and their efficacy in the treatment of specific pathogens is beyond the scope of this manuscript. The biggest limitations in delivering treatments to free-ranging wild animals is lack of access to treat individual animals, and lack of knowledge on the proportion of a population that would need to be effectively treated to reduce the continued spread of a fungal disease. Large-scale aerosolization (“fogging”) treatments have been suggested for specific cave environments affected by bat geomycosis, but the environmental effects of “untargeted” environmental treatments are likely to disrupt microbial communities within delicate cave ecosystems, and the efficacy for the treatment is unknown at this time.
The captive breeding of endangered species vulnerable to emerging infectious disease pandemics has been advocated as a temporary “rescue” measure until the threat of infectious disease can be mitigated. Captive breeding is particularly appealing as a putative solution when the infectious disease has had an anthropogenic component, or when human activities have compounded a threat to vulnerable populations. In principle, captive stock can be subsequently used to
supplement wildlife populations or reintroduce them, after the infectious disease threat is mitigated, to areas where the species underwent local extinction. Indeed, many species have been successfully bred in captivity for purposes of reintroduction after the initial threat of an infectious disease has threatened their survival.
A successful captive breeding and propagation program undertaken by the U.S. Fish and Wildlife Service (FWS) in the 1980s was aimed at the recovery of the black-footed ferret (Mustela nigripes). Although the species was considered extinct in 1979, the rediscovery of a small population in Wyoming in 1981 launched a recovery program for the species. When this extremely small population was further threatened by an outbreak of canine distemper virus after a sylvatic plague outbreak (Forrest et al., 1988; Williams et al., 1988), the FWS started a rescue effort to propagate the species in captivity from those wild founders. This program has now resulted in the reintroduction of the species into the wild at multiple sites, and has resulted in adequate early success in a relatively short time frame for the recovery of an entire species. Between 1987 and 2010, more than 6,500 ferret kits were produced, and over 2,300 animals have been released into the wild through 19 projects across 8 states, Canada, and Mexico (FWS, 2011).
Over the years, this program has benefited from advances in veterinary medicine to mitigate the threats posed by infectious diseases in the wild and in captivity; advances in captive husbandry and nutrition; establishment of biosecurity measures; application of modern principles of genetic management; and the use of assisted reproductive techniques to enhance genetic diversity. Similar multidisciplinary programs could be applicable to supplement endangered populations at risk of extinction from a fungal disease.
The unprecedented emergence of amphibian chitridiomycosis and the global scale of this threat have triggered the launch of concerted recovery efforts for amphibians under the Amphibian Ark (AARK). The AARK is a joint initiative by international partners (the World Association of Zoos and Aquariums, the International Union for Conservation of Nature [IUCN] Conservation Breeding Specialist Group, and the IUCN Amphibian Specialist Group) dedicated to “ensuring the global survival of amphibians, focusing on those that cannot currently be safeguarded in nature” (AARK, 2011). The AARK prioritizes and rescues amphibian species at risk of extinction while developing in-country capacity for the continued research and propagation of those species. Although aimed at the overall decline of amphibian populations, the prominent role of the chytrid fungus in the decline has prompted the AARK to develop and refine biosecurity procedures and management techniques to control and mitigate the effects of this disease. This active network of professionals is a model for the recovery of species through ex situ propagation and investigation, with the goal of returning and recovering threatened populations of amphibians to their natural habitats.
However, captive propagation may not always be the most viable solution in the face of a wildlife epizootic, and should never be seen as a substitute to directly addressing the anthropogenic factors driving the emergence of diseases threaten-
ing wildlife populations. The financial cost of captive propagation programs is variable and species-dependent, but inevitably significant. In addition, the life history and biological needs of some species may make them unsuitable candidates for captive recovery programs. Captive breeding should be seen as a short-term strategy for the recovery of endangered or threatened species, and should only be considered as a last resort, as in the previous examples, with defined goals and a target time line. Captive breeding should only be undertaken in conjunction with comprehensive efforts to create in situ capacity for disease and population monitoring, continued basic disease research, and efforts to contain or mitigate the spread of disease in the wild. The limitations of captive breeding programs have been previously discussed (Lynch and O’Hely, 2001; Snyder et al., 1996) and should always be weighed as part of the risk assessment on the viability of captive breeding strategies. Limitations include problems with establishing self-sufficient captive populations; species-specific and possibly limited success in reintroductions; relatively high costs; possible domestication and the selection and propagation of traits that may not be ideal for wild species survival; and possible exposure and establishment of unrecognized diseases in captive stock that could be carried to their wild counterparts.
Techniques that increase the population resistance to infection could be applicable to the management of fungal epizootics, specifically through vaccinations or bioaugmentation of biological defenses. The acquisition of immunity against fungal pathogens is not well known for all pathogens, and the complexity of pathogen–host relationships further exacerbates our gaps in knowledge. Although not widely used, fungal vaccines have been developed, even if their efficacy has varied widely. Different strategies in vaccine development, from targeting fungal cell wall characteristics (Cassone and Torosantucci, 2006) to developing recombinant vaccines that induce immunity to specific fungal proteins, could be employed. However, these techniques take time to develop and effectively implement on a large scale, and are potentially costly.
A novel technique being used to convey immunity to amphibians at risk of chytridiomycosis involves the use of skin bacteria that produce antifungal compounds. The addition of Janthobacterium lividum to the skin of mountain yellow-legged frogs (Rana muscosa) has been demonstrated to convey protection against challenge with B. dermatitis (Harris et al., 2009). Surveillance of wild populations has suggested that the presence of symbiotic skin bacteria capable of producing anti-Bd compounds is beneficial to populations in the wild (Lam et al., 2010). The bioaugmentation of anti-Bd bacterial flora has the potential to be a significant advance in increasing the survival of amphibians in areas where the fungus has become endemic. The recognition of similar symbionts in the skin of other species could help direct action plans toward increasing resistance to fungi in natural populations.
Environmental manipulations could be effective in captive efforts, or during short-term recovery of individual infected animals, but are not necessarily
feasible to apply in wild environments. Manipulating the pH of soil or water, and changing the temperature, humidity, or other microclimatic conditions, are possible ways to discourage the growth of certain life stages of fungal organisms. However, basic biological research is necessary before these manipulations could even be considered or applied in the wild. The detrimental effects of these manipulations to the environment would need to be understood and minimized, particularly when applied at scales large enough to make a difference to the health of a population of wild animals.
The mass culling of populations known to be affected by an emerging infectious disease has been advocated by domestic animal regulatory agencies and has been an effective tool in the eradication of diseases of agricultural interest and zoonotic concern. The efficacy of culling wildlife populations has not been established and efforts have been met with mixed results. Success would depend on a multitude of factors, including the pathogen, transmission dynamics, population biology of hosts and the reservoir species, prevalence and incidence of the disease, and culling techniques. Wildlife culling has been effective at meeting the immediate management goals in some wildlife health scenarios, but the overall impacts to ecosystems are often unknown, typically complex, and not easily predicted. For instance, long-term efforts to cull wild European badgers (Meles meles) by the British government to control tuberculosis (TB) in cattle have been successful at reducing incidence of TB in cattle in the immediate area where badgers are culled, but actually increased the incidence in adjoining areas (Donnelly et al., 2007). A disease model to examine the efficacy of culling bats in hibernacula as a way to slow the spread of WNS suggested that this strategy would be ineffective, in part due to high contact rates among colonial bat species (Hallam and McCracken, 2011). Culling is not likely to be a large-scale effective strategy for managing fungal infectious diseases.
Conclusion
Several emerging fungal diseases have recently been shown to affect wild animal species. Some of these diseases carry a zoonotic risk potential, but even those that do not directly affect human health are likely to carry a societal cost in terms of ecosystem health. Wild animal species can be sentinels of emerging diseases and are early indicators of overall ecosystem health. Advances in veterinary medicine have aided in the recognition of fungal pathogens, and new techniques are being developed to mitigate their impact on wild populations. However, the inherent responsibility falls to veterinarians, physicians, conservation biologists, and public health professionals to properly document and disseminate the findings of fungal diseases in novel hosts, geographic locations, or areas of increased frequency. Continued multidisciplinary surveillance of fungal diseases is essential to understand the impact of these emerging pathogens on wild animal populations, humans, and ecosystems.
References
AARK (Amphibian Ark). 2011. Amphibian Ark: About us. http://www.amphibianark.org/about-us/ (accessed February 27, 2011).
Abarca, M. L., J. Martorell, G. Castellá, A. Ramis, and F. J. Cabañes. 2008. Cutaneous hyalohyphomycosis caused by a Chrysosporium species related to Nannizziopsis vriesii in two green iguanas (Iguana iguana). Medical Mycology 46:349–354.
Ajello, L., A. A. Padhye, S. Sukroongreung, C. H. Nilaku, and S. Tantimavanic. 1995. Occurrence of Penicillium marneffei infections among wild bamboo rats in Thailand. Mycopathologia 131:1–8.
Berger, L., R. Speare, P. Daszak, D. E. Green, A. A. Cunningham, C. L. Goggin, R. Slocombe, M. A. Raga, A. D. Hyatt, K. R. McDonald, H. B. Hines, K. R. Lips, G. Marantelli, and H. Parke. 1998. Chytridiomycosis causes amphibian mortality associated with population declines in the rain forests of Australia and Central America. Proceedings of the National Academy of Sciences, USA 95:9031–9036.
Berger, L., G. Marantelli, L. L. Skerratt, and R. Speare. 2005. Virulence of the amphibian chytrid fungus Batrachochytrium dendrobatidis varies with the strain. Diseases of Aquatic Organisms 68:47–50.
Bertelsen, M. F., G. J. Crawshaw, L. Sigler, and D. A. Smith. 2005. Fatal cutaneous mycosis in tentacled snakes (Erpeton tentaculatum) caused by the Chrysosporium anamorph of Nannizziopsis vriesii. Journal of Zoo and Wildlife Medicine 36:82–87.
Blehert, D. S., A. C. Hicks, M. Behr, C. U. Meteyer, B. M. Berlowski-Zier, E. L. Buckles, J. T. H. Coleman, S. R. Darling, A. Gargas, R. Niver, J. C. Okoniewski, R. J. Rudd, and W. B. Stone. 2009. Bat white nose syndrome: An emerging fungal pathogen? Science 232:227.
Bowman, M. R., J. A. Paré, L. Sigler, J. P. Naeser, K. K. Sladky, C. S. Hanley, P. Helmer, L. A. Phillips, A. Brower, and R. Porter. 2007. Deep fungal dermatitis in three inland bearded dragons (Pogona vitticeps) caused by the Chrysosporium anamorph of Nannizziopsis vriesii. Medical Mycology 45:371–376.
Boyle, D. G., D. B. Boyle, V. Olsen, J. A. T. Morgan, and A. D. Hyatt. 2004. Rapid quantitative detection of chytridiomycosis (Batrachochytrium dendrobatidis) in amphibian samples using real time Taqman PCR assay. Diseases of Aquatic Organisms 60:141–148.
Burek, K. 2001. Mycotic diseases. In Infectious diseases of wild mammals, 3rd ed., edited by E. S. Williams and I. K. Barker. Ames, IA: Iowa State University Press. Pp. 514–531.
Caldwell, D. K., M. C. Caldwell, J. C. Woodard, L. Ajello, W. Kaplan, and H. M. McClure. 1975. Lobomycosis as a disease of the Atlantic bottle-nosed dolphin (Tursiops truncatus, Montagu, 1821). American Journal of Tropical Medicine and Hygiene 24:105–114.
Cao, C., L. Liang, W. Wang, H. Luo, S. Huang, D. Liu, J. Xu, D. A. Henk, and M. C. Fisher. 2011. Common reservoirs for Penicillium marneffei infection in humans and rodents, China. Emerging Infectious Diseases 17:209–214.
Caponi, M., G. Segretain, and P. Sureau. 1956. Penicilliosis de Rhizomis sinensis. Bulletin de la Société de Pathologie Exotique 49:418–421.
Cassone, A., and A. Torosantucci. 2006. Opportunistic fungi and fungal infections: The challenge of a single, general antifungal vaccine. Expert Review of Vaccines 5:859–867.
Chaturvedi, V., and S. Chaturvedi. 2011. What is in a name? A proposal to use geomycosis instead of white nose syndrome (WNS) to describe bat infection caused by Geomyces destructans. Mycopathologia 171:231–233.
Cryan, P. M., C. Uphoff, C. U. Meteyer, J. G. Boyles, and D. S. Blehert. 2010. Wing pathology of white-nose syndrome in bats suggests life-threatening disruption of physiology. BMC Biology 8:135–142.
Daszak, P., A. A. Cunningham, and A. D. Hyatt. 2000. Emerging infectious diseases of wildlife—threats to biodiversity and human health. Science 287:443–449.
Deng, Z. L., M. Yun, and L. Ajello. 1986. Human Penicilliosis marneffei and its relation to the bamboo rat (Rhizomys pruinosus). Journal of Medical and Veterinary Mycology 24:383–389.
Donnelly, C. A., G. Wei, W. T. Johnston, D. R. Cox, R. Woodroffe, F. J. Bourne, C. L. Cheeseman, R. S. Clifton-Hadley, G. Gettinby, P. Gilks, H. E. Jenkins, A. M. LeFevre, J. P. McInerney, and W. I. Morrison. 2007. Impacts of widespread badger culling on cattle tuberculosis: Concluding analyses from a large-scale field trial. International Journal of Infectious Diseases 11:300–308.
Duong, T. A. 1996. Infection due to Penicillium marneffei, an emerging pathogen: Review of 155 reported cases. Clinical Infectious Diseases 23:125–130.
Foley, J., D. Clifford, K. Castle, P. Cryan, and R. S. Ostfeld. 2011. Investigating and managing the rapid emergence of white-nose syndrome, a novel, fatal, infectious disease of hibernating bats. Conservation Biology 25(2):223–231.
Forrest, S. C., D. E. Biggins, L. Richardson, T. W. Clark, T. M. Campbell, K. A. Fagerstone, and E. T. Thorne. 1988. Population attributes for the black-footed ferret (Mustela nigripes) at Meeteetse, Wyoming, 1981–1985. Journal of Mammalogy 69:261–273.
Frick, W. F., J. F. Pollock, A. C. Hicks, K. E. Langwig, D. S. Reynolds, G. G. Turner, C. M. Butchkoski, and T. H. Kunz. 2010. An emerging disease causes regional population collapse of a common North American bat species. Science 329:679–682.
FWS (U.S. Fish and Wildlife Service). 2011. U.S. Fish and Wildlife Service endangered species reports: Prairie mountain region: Black footed ferret. http://www.fws.gov/mountain-prairie/species/mammals/blackfootedferret/ (accessed February 25, 2011).
Gargas, A., M. T. Trest, M. Christensen. T. J. Volk, and D. S. Blehert. 2009. Geomyces destructans sp. nov. associated with bat white-nose syndrome. Mycotaxon 108:147–154.
Goka, K., J. Yokoyama, Y. Une, T. Kuroki, K. Suzuki, M. Nakahara, A. Kobayashi, S. Inaba, T. Mizutani, and A. D. Hyatt. 2009. Amphibian chytridiomycosis in Japan: Distribution, haplotypes and possible route of entry into Japan. Molecular Ecology 18:4757–4774.
Gugnani, H., M. C. Fisher, A. Paliwal-Johsi, N. Vanittanakom, I. Singh, and P. S. Yadav. 2004. Role of Cannomys badius as a natural animal host of Penicillium marneffei in India. Journal of Clinical Microbiology 42:5070–5075.
Hallam, T. G., and G. F. McCracken. 2011. Management of the panzootic white-nose syndrome through culling of bats. Conservation Biology 25:189–194.
Harris, R. N., T. Y. James, A. Lauer, M. A. Simon, and A. Patel. 2006. Amphibian pathogen Batrachochytrium dendrobatidis is inhibited by the cutaneous bacteria of amphibian species. EcoHealth 3:53–56.
Harris, R. N., R. M. Brucker, J. B. Walke, M. H. Becker, C. R. Schwantes, D. C. Flaherty, B. A. Lam, D. C. Woodhams, C. J. Briggs, V. T. Vredenburg, K. P. C. Minbiole, T. Y. James, A. Lauer, M. A. Simon, and A. Pate. 2009. Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. The ISME Journal 3:818–824.
Herr, R. A., E. J. Tarcha, P. R. Taborda, J. W. Taylor, L. Ajello, and L. Mendoza. 2001. Phylogenetic analysis of Lacazia loboi places this previously uncharacterized pathogen with the dimorphic Onygenales. Journal of Clinical Microbiology 39:309–314.
Kiszka, J., M. F. Van Bressem, and C. Pusineri. 2009. Lobomycosis-like disease and other skin conditions in Indo-Pacific bottlenose dolphins Tursiops aduncus from the Indian Ocean. Diseases of Aquatic Organisms 84:151–157.
Kriger, K. M., J. M. Hero, and K. J. Ashton. 2006. Cost efficiency in the detection of chytridiomycosis using PCR assay. Diseases of Aquatic Organisms 71:149–154.
Lam, B. A., J. B. Walke, V. T. Vredenburg, and R. N. Harris. 2010. Proportion of individuals with anti-Batrachochytrium dendrobatidis skin bacteria is associated with population persistence in the frog Rana muscosa. Biological Conservation 143:529–531.
Longcore, J. E., A. P. Pessier, and D. K. Nichols. 1999. Batrachochytrium dendrobatidis gen. et sp. nov.: A chytrid pathogenic to amphibians. Mycologia 91:219–227.
Lynch, M., and M. O’Hely. 2001. Captive breeding and the genetic fitness of natural populations. Conservation Genetics 2:363–378.
McCallum, H. 2005. Inconclusiveness of chytridiomycosis as the agent in widespread frog declines. Conservation Biology 19:1421–1430.
Meteyer, C. U., E. L. Buckles, D. S. Blehert, A. C. Hicks, D. E. Green, V. Sheam-Bochsler, N. J. Thomas, A. Gargas, and M. J. Behr. 2009. Histopathologic criteria to confirm white-nose syndrome in bats. Journal of Veterinary Diagnostics and Investigation 21:411–414.
Morehouse, E. A., T. Y. James, A. R. D. Ganley, R. Vilgalys, L. Berger, P. J. Murphy, and J. E. Longcore. 2003. Multilocus sequence typing suggests the chytrid pathogen of amphibians is a recently emerged clone. Molecular Ecology 12:395–403.
Murdoch, M. E., J. S. Reif, M. Mazzoli, S. D. McCulloch, P. A. Fair, and G. D. Bossart. 2008. Lobomycosis in bottlenose dolphins (Tursiops truncatus) from the Indian River Lagoon, Florida: Estimation of prevalence, temporal trends, and spatial distribution. EcoHealth 5:289–297.
Murdoch, M. E., M. Mazzoli, S. McCulloch, S. Bechdel, G. O’Corry-Crowe, G. D. Bossart, and J. S. Reif. 2010. Lacaziosis in bottlenose dolphins Tursiops truncatus along the coastal Atlantic Ocean, Florida USA. Diseases of Aquatic Organisms 92:69–73.
Nichols, D. K., R. S. Weyant, E. W. Lamirande, L. Sigler, and R. T. Mason. 1999. Fatal mycotic dermatitis in captive brown tree snakes (Boiga irregularis). Journal of Zoo and Wildlife Medicine 30:111–118.
Nichols, D. K., E. W. Lamirande, A. P. Pessier, and J. E. Longcore. 2001. Experimental transmission of cutaneous chytridiomycosis in dendrobatid frogs. Journal of Wildlife Diseases 37:1–11.
Ouellet, M., I. Mikaelian, B. D. Pauli, J. Rodrigue, and D. M. Green. 2005. Historical evidence of widespread chytrid infection in North American amphibian populations. Conservation Biology 19:1431–1440.
Paré, J. A., and L. Sigler. 2006. Fungal diseases. In Reptile medicine and surgery, 2nd ed., edited by D. R. Mader. St. Louis, MO: Elsevier Inc. Pp. 217–226.
Paré, J. A., L. Sigler, K. L. Rypien, and C. F. C. Gibas. 2003. Cutaneous mycobiota of captive reptiles with notes on the scarcity of the Chrysosporium anamorph of Nanniziopsis vriesii. Journal of Herpetological Medicine and Surgery 13:10–15.
Paré, J. A., K. A. Coyle, L. Sigler, A. K. Maas III, and R. L. Mitchell. 2006. Pathogenicity of the Chrysosporium anamorph of Nannizziopsis vriesii for veiled chameleons (Chamaeleo calyptratus). Medical Mycology 44:25–31.
Pessier, A. P., D. K. Nichols, J. E. Longcore, and M. S. Fuller. 1999. Cutaneous chytridiomycosis in poison dart frogs (Dendrobates spp.) and White’s tree frogs (Litoria caerulea). Journal of Veterinary Diagnostics and Investigation 11:194–199.
Puechmaille, S. J., P. Verdeyroux, H. Fuller, A. Gouilh, M. Bekaert, and E. C. Teeling. 2010. Whitenose syndrome fungus (Geomyces destructans) in bat, France. Emerging Infectious Diseases 16:290–293.
Reif, J. S., M. S. Mazzoli, S. D. McCulloch, R. A. Varela, J. D. Goldstein, P. A. Fair, and G. D. Bossart. 2006. Lobomycosis in Atlantic bottlenose dolphins from the Indian River Lagoon, Florida. Journal American Veterinary Medical Association 228:104–108.
Reif, J. S., M. M. Peden-Adams, T. A. Romano, C. D. Rice, P. A. Fair, and G. D. Bossart. 2008. Immune dysfunction in Atlantic bottlenose dolphins (Tursiops truncatus) with lacaziosis. Medical Mycology 4:1–11.
Rotstein, D. S., L. G. Burdett, W. McLellan, L. Schwacke, T. Rowles, K. A. Terio, S. Schultz, and A. Pabst. 2009. Lobomycosis in offshore bottlenose dolphins (Tursiops truncatus), North Carolina. Emerging Infectious Diseases 15:588–590.
Schloegel, L. M., J. M. Hero, L. Berger, R. Speare, K. McDonald, and P. Daszak. 2006. The decline of the sharp-snouted day frog (Taudactylus acutirostris): The first documented case of extinction by infection in a free-ranging wildlife species? EcoHealth 3:35–40.
Skerratt, L. F., L. Berger, R. Speare, S. Cashins, K. R. McDonald, A. D. Phillott, H. B. Hines, and N. Kenyon. 2007. Spread of chytridiomycosis has caused the rapid global decline and extinction of frogs. EcoHealth 4:125–134.
Snyder, N. F. R., S. R. Derrickson, S. R. Bessinger, J. W. Wiley, T. B. Smith, W. D. Toone, and B. Miller. 1996. Limitations of captive breeding in endangered species recovery. Conservation Biology 10:338–348.
Steininger, C., J. van Lunzen, K. Tintelnot, I. Sobottka, H. Rohde, M. A. Horstkotte, and H. J. Stellbrink. 2005. Mycotic brain abscess caused by opportunistic reptile pathogen. Emerging Infectious Diseases 11:349–350.
Terio, K. A., I. H. Stalis, J. L. Allen, J. L. Stott, and M. B. Worley. 2003. Coccidioidomycosis in Przewalski’s horses (Equus przewalskii). Journal of Zoo and Wildlife Medicine 34:339–345.
Thomas, A. D., L. Sigler, S. Peucker, J. H. Norton, and A. Nielan. 2002. Chrysosporium anamorph of Nanniziopsis vriesii associated with fatal cutaneous mycoses in the salt-water crocodile (Crocodylus porosus). Medical Mycology 40:143–151.
Van Bressem, M. F., M. O. Santos, and J. E. Oshima. 2009. Skin diseases in Guiana dolphins (Sotalia guianensis) from the Paranaguá estuary, Brazil: A possible indicator of a compromised marine environment. Marine Environmental Research 67:63–68.
Vanittanakom, N., C. R. Cooper, M. C. Fisher, and T. Sirisanthana. 2006. Penicillium marneffei infection and recent advances in the epidemiology and molecular biology aspects. Clinical Microbiology Reviews 19:95–110.
Weldon, C., L. H. du Preez, A. D. Hyatt, R. Muller, and R. Speare. 2004. Origin of the amphibian chytrid fungus. Emerging Infectious Diseases 10:2100–2105.
Wibbelt, G., A. Kurth, D. Hellmann, M. Weishaar, A. Barlow, M. Veith, J. Prüger, T. Görföl, L. Grosche, F. Bontadina, U. Zöphel, H.-P. Seidl, P. M. Cryan, and D. S. Blehert. 2010. Whitenose syndrome fungus (Geomyces destructans) in bats, Europe. Emerging Infectious Diseases 16:1237–1242.
Williams, E. S., E. T. Thorne, M. J. Appel, and D. W. Belitsky. 1988. Canine distemper in black-footed ferrets (Mustela nigripes) from Wyoming. Journal of Wildlife Diseases 24:385–398.
THE EMERGENCE OF PHYTOPHTHORA RAMORUM IN NORTH AMERICA AND EUROPE
David M. Rizzo,47Ross K. Meentemeyer,48and Matteo Garbelotto49
Introduction
The emergence of fungal and fungal-like plant pathogens in agricultural and natural ecosystems can be triggered by a number of key changes in the host–pathogen–environment interaction (Anderson et al., 2004; Desprez-Loustau and Rizzo, 2011; Desprez-Loustau et al., 2007). The evolution of pathogens owing to selection pressure due to fungicides, deployment of resistant plant varieties, or changes in the environment (global to local) can allow previously known pathogens to increase in incidence locally or across wide geographic areas (Anderson et al., 2004). Movement of fungi from one part of the world to another may also lead to disease emergence and large-scale epidemics of plant pathogens (Desprez-Loustau et al., 2007; Rizzo, 2005). Because of the well-known impacts of exotic plant pathogens, much effort has been made at regional, national, and
______________
47 Department of Plant Pathology, University of California-Davis.
48 Department of Geography and Earth Science, University of North Carolina-Charlotte.
49 Department of Environmental Science, Policy and Management, University of California-Berkeley.
international levels to restrict movement of plant pathogens in order to protect agricultural crops. However, some of the greatest impacts of exotic plant pathogens have occurred in natural ecosystems. Well-known, high-impact fungal diseases that are considered to be caused by exotic pathogens include chestnut blight (caused by Cryphonectria parasitica), white pine blister rust (caused by Cronartium ribicola), Dutch elm disease (caused by Ophiostoma ulmi and O. novo-ulmi), and jarrah dieback (caused by Phytophthora cinnamomi) (Aukema et al., 2010; Desprez-Loustau et al., 2007; Rizzo, 2005).
Recently, Phytophthora ramorum has emerged as a presumed exotic causal agent of the forest disease “sudden oak death” that has had important impacts on coastal oak forests in California and Oregon (Rizzo and Garbelotto, 2003) and more recently in woodlands in the United Kingdom (Brasier and Webber, 2010). There are several reviews available that cover disease symptoms, biology, ecology, management, and history of P. ramorum (e.g., Davidson et al., 2003; Garbelotto and Rizzo, 2005; Grünwald et al., 2008; Kliejunas, 2010; Rizzo and Garbelotto, 2003; Rizzo et al., 2005). Information on the pathogen and the diseases it causes is updated regularly on the website of the California Oak Mortality Task Force (http//:www.suddenoakdeath.org). In this brief review, we give an overview of some of the factors responsible for the emergence of P. ramorum followed by current work on management of the pathogen.
The Pathogen
Phytophthora ramorum was unknown before it was observed to cause diseases of a number of host species in the mid-1990s in Europe and California. The pathogen was officially named P. ramorum in 2001 (Werres et al., 2001). Population studies indicate that P. ramorum in both North America and Europe has a genetic structure consistent with that expected of an introduced species and that it reproduces exclusively clonally (Ivors et al., 2004, 2006). P. ramorum has two mating types (A1, A2); to date, all North American isolates of P. ramorum have been found to be mating type A2, while all isolates in Europe (with rare exceptions) are A1 (Grünwald et al., 2008). Sexual reproduction outside of the laboratory has not been documented (Grünwald et al., 2008), although laboratory attempts at crossing the P. ramorum mating types have produced viable progeny with similar virulence to the parent types (Boutet et al., 2010). Three genetically distinct lineages (NA1, NA2, and EU1) have been identified within P. ramorum (Grünwald et al., 2008; Ivors et al., 2006). Only NA1 is present in California and Oregon forests, all three lineages are found in North American nursery populations, and only EU1 has been found in European nurseries and woodlands (Goss et al., 2009, 2011; Grünwald et al., 2008; Ivors et al., 2006).
The putative exotic nature of P. ramorum in North America is also supported by the findings that it is present in nurseries (Yakabe et al., 2009) yet is absent in historical herbarium collections (Monahan et al., 2008). However, the geographic origin of P. ramorum remains unknown. From a phylogenetic perspective, its
nearest relative is P. lateralis, another presumed exotic pathogen in California and Oregon that is the causal agent of Port-Orford cedar root disease (Rizzo et al., 2002). The geographic origin of P. lateralis was also unknown until it was recently found in the mountains of Taiwan (Brasier et al., 2010). This suggests a potential origin for P. ramorum in Asia (Brasier et al., 2010), but there is no direct evidence at this time.
P. ramorum is a generalist plant pathogen and has been found to infect more than 125 plant species including ferns, gymnosperms, monocots, and dicots (Grünwald et al., 2008; Kliejunas, 2010). The U.S. Department of Agriculture Animal and Plant Health Inspection Service Plant Protection and Quarantine maintains an updated list of regulated and associated hosts (http://www.aphis.usda.gov/). The diseases that P. ramorum causes on this wide range of hosts are expressed in two ways: canker infections that may cause tree mortality and nonlethal foliar and twig infections (known as ramorum blight) (Rizzo et al., 2005). Canker symptoms have been primarily described from oaks and other members of Fagaceae (known as sudden oak death). Recently, canker symptoms have been reported on trunks of Japanese larch in plantations in the United Kingdom (Brasier and Webber, 2010). No host-specific population genetic structure has been detected in P. ramorum. While there is variation in virulence within P. ramorum populations, isolates of the pathogen from one host show similar virulence on unrelated hosts (Hüberli and Garbelotto, 2011).
Transmission of P. ramorum in forests is primarily driven by spore production from foliar infections rather than from lethal cankers on the main stem of trees (Davidson et al., 2005, 2008, 2011). P. ramorum spreads naturally via spores over both short and long distances. Spread is primarily by rain splash, with the majority of propagules appearing to travel less than 10 m (Davidson et al., 2005; Mascheretti et al., 2008). Genetic information has been used to infer inoculum dilution curves and spread potential of the pathogen (Mascheretti et al., 2008, 2009). The curve describing such a relationship is bimodal and reminiscent of dispersal curves for relatively large particles, with a steep gradient indicating limited dispersal ability (most dispersal within 10 m, and decreasing to a minimum at about 500 m) and a secondary peak at 1–3 km, indicating medium-distance dispersal (Mascheretti et al., 2008). These values match observations based on the onset of new infestation foci in Oregon (Hansen et al., 2008). Long-distance spread of the pathogen in both North America and Europe appears to be primarily associated with movement of nursery plants (Goss et al., 2009; Ivors et al., 2006; Prospero et al., 2009).
Emergence of P. ramorum in North America
A new disease, described as sudden oak death (SOD), was first associated with mortality of tanoak (Notholithocarpus densiflorus) and coast live oak (Quercus agrifolia) in the San Francisco Bay area during the mid-1990s (Garbelotto et al., 2001). A Phytophthora species was identified and confirmed as the causal
agent of the disease in 2000 (Rizzo et al., 2002). In the United States, P. ramorum’s current geographic range in native forests extends from the Big Sur area in central California to southern Mendocino County, with two disjunct populations in Humboldt County, California, and one small population in Curry County, Oregon (Figure A16-1). P. ramorum has not become established in forests outside of this area. Potentially millions of tanoak and oak trees have been lost to the disease over the past 10 years (Meentemeyer et al., 2008c, 2011).
The pathogen has been associated with the horticultural industry in both the United States and Canada and has been consistently found in a number of nurseries. While dozens of plant species have been found infected in nurseries, the majority of infections have been associated with the genera Rhododendron, Camellia, Pieris, Kalmia, and Viburnum (Osterbauer et al., 2004; Parke et al., 2010a). Although P. ramorum has not been found in forests outside of California and Oregon, the pathogen has become established in streams in several states, including Washington, Alabama, Mississippi, Georgia, and South Carolina (Oak et al., 2010). These streams have been associated with infestations in nearby nurseries; in several rare instances vegetation along the banks has become infected by P. ramorum, but there is no evidence of terrestrial colonization of forests at this time outside of California and Oregon (Oak et al., 2010).
The emergence of P. ramorum in California and Oregon forests can be linked to several key factors: movement of ornamental plants (both into and within California) (Ivors et al., 2006; Prospero et al., 2009), the susceptibility and distribution of host plants in coastal forests (Anacker et al., 2008; Dodd et al., 2004, 2008; Hayden et al., 2011; Hüberli and Garbelotto, 2011; Meentemeyer et al., 2008a,b,c; Rizzo et al., 2005), and a suitable climate (Meentemeyer et al., 2004, 2011). P. ramorum appears to have originally spread from two focal points in California. Two P. ramorum populations (Marin and Santa Cruz) were identified using population genetic analysis as the two oldest sources of the pathogen in the state (Mascheretti et al., 2008, 2009). This genetic analysis corroborated anecdotal evidence from field observations and nursery records. By reconstructing the epidemic from this point, other locations were determined to be of intermediate age. For example, the Big Sur population was shown to be derived from the Santa Cruz population originally associated with nursery stock (Mascheretti et al., 2008). This analysis suggests at least eight separate introductions of P. ramorum within California from the two focal populations. Most of these introductions were likely human related via movement of nursery plants, but some introductions were likely caused by natural spread of the pathogen from forests (Cushman and Meentemeyer, 2008; Mascheretti et al., 2008, 2009).
P. ramorum can be found in a number of forest types along California and Oregon coasts; these range from drier oak mixed evergreen forests dominated by coast live oak to wetter forest types dominated by coast redwood or Douglas fir (Rizzo and Garbelotto, 2003). In these conifer-dominated forests, tanoak is the
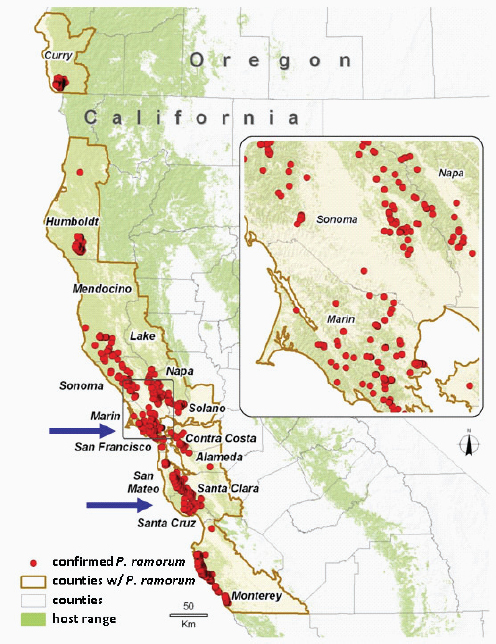
Figure A16-1 Current distribution of Phytophthora ramorum in California and Oregon forests. Red circles represent areas where P. ramorum and sudden oak death have been confirmed. The blue arrows indicate the general locations (Marin and Santa Cruz counties) where P. ramorum was initially introduced into California.
SOURCE: Figure courtesy of Ross K. Meentemeyer, University of North Carolina-Charlotte.
host most affected by sudden oak death. As forest types vary across a topographical landscape, there are associated changes in microclimate and, consequently, a likelihood of variation in the host–pathogen interaction and pathogen transmission (Condeso and Meentemeyer, 2007; Meentemeyer et al., 2008a, 2011). These changes in microclimate may be due to variation in edaphic factors such as aspect, soil strata, and hydrology that underlie the growth of a particular vegetation type. Differences in the ensuing physical structure of the vegetation itself also affect light availability and, consequently, temperature and moisture. These are crucial factors that determine survival and sporulation of most pathogens, including Phytophthora species. On the level of fine-scale host-pathogen interactions, these environmental variables may affect the timing and production of inoculum, the length of the infectious period, the degree of host susceptibility or pathogen virulence, pathogen survival through adverse conditions (dormancy), the timing and incidence of new infections, and the overall magnitude and pattern of disease epidemics.
Although P. ramorum infects more than 25 host species in these woodlands, nonlethal foliar lesions on bay laurel (Umbellularia californica) are the most important host tissue for sporulation by P. ramorum in the oak woodlands of California (Davidson et al., 2005, 2008, 2011). Levels of inoculum in through-fall rain are up to 20 times higher under bay laurel as opposed to other forest trees. In addition, at the landscape level, infection on bay laurel is known to precede infection on oak and tanoak trees, and the presence of this host is associated with higher levels of oak mortality (Cobb et al., 2010; Maloney et al., 2005). Consequently, infections on bay laurel leaves drive the spread of P. ramorum and the onset of lethal infections on oak and tanoak. Sporulation does occur on tanoak leaves and this species does appear capable of driving epidemics in the absence of bay laurel (Davidson et al., 2008). This situation is occurring in Oregon forest, where bay laurel is a relatively minor component of the forests (Hansen et al., 2008).
Various modeling efforts have been made to predict the spread and establishment of P. ramorum in California, Oregon, and other locations within North America (e.g., Kelly et al., 2007; Meentemeyer et al., 2004, 2008a,b, 2011; Vaclavik et al., 2010; Venette and Cohen, 2006). For example, a recent epidemiological model was used in combination with geographical modeling to predict the spread of P. ramorum through host populations in wildland forests, subject to fluctuating weather conditions (Meentemeyer et al., 2011). Application of the model to Californian landscapes over a 40-year period (1990–2030), since the approximate time of pathogen introduction, suggests that in the absence of extensive control, a 10-fold increase in disease spread will occur between 2010 and 2030 with most infection concentrated along the northern coast of California between San Francisco and Oregon (Meentemeyer et al., 2011). Long-range dispersal of inoculum to susceptible host communities in the Sierra Nevada foothills and coastal southern California leads to little secondary infection due to lower host
availability and less suitable weather conditions. However, a shift to wetter and milder conditions in future years would double the amount of disease spread in California through 2030 (Meentemeyer et al., 2011). In other areas of North America, the forests of the southern Appalachian Mountains are considered to be at the highest risk (Kelly et al., 2007; Rizzo et al., 2005; Venette and Cohen, 2006). The combination of susceptible oaks (e.g., Q. rubra), potential sporulating hosts found in the understory (Rhododendron, Kalmia), and a moist, relatively mild climate has the potential to support the emergence of P. ramorum if it is introduced into these areas (Spaulding and Rieske, 2011).
Emergence of P. ramorum in Europe
Around the same time that SOD was noted in California, a new Phytophthora species was observed to infect rhododendrons in nurseries and gardens in Germany and the Netherlands (Werres et al., 2001). A connection was made between the European and California Phytophthora species in December 2000 (Rizzo et al., 2002). Although initially described from Germany and the Netherlands, P. ramorum has now been identified in nurseries or gardens of most countries in Western Europe (Kliejunas, 2010). To date, P. ramorum has not caused the extensive damage in native European woodlands that has been seen in California forests; stem cankers caused by P. ramorum have only been found in the United Kingdom and the Netherlands. The pathogen is most widespread in the United Kingdom (including Ireland) in mixed-species woodlands, planted woodland gardens, heritage gardens, and national plant collections (Fichtner et al., 2011). Surveys for P. ramorum in Great Britain were initiated in 2001 and the pathogen has since been found at hundreds of sites, both in nursery systems and in woodlands (Kliejunas, 2010). P. ramorum causes extensive foliar necrosis and shoot dieback on Rhododendron ponticum (considered an invasive plant species) and bleeding cankers on European beech (Fagus sylvatica) as well as other tree species. Early surveys of disease incidence found that the majority of infected trees are located within 2 m of infected R. ponticum, suggesting a role of R. ponticum in disease transmission.
P. ramorum has recently emerged as a serious pathogen in Japanese larch (Larix kaempferi) plantations in the United Kingdom (Brasier and Webber, 2010). Japanese larch is an important timber tree in the United Kingdom and is grown in large plantations. While other conifers had been reported as hosts (e.g., redwood, Douglas fir), this was the first observation of extensive damage caused by P. ramorum to conifers. The key finding was that larch can serve both as a canker host, leading to death of large trees (as on oaks), as well as a foliar host that supports sporulation of P. ramorum (Brasier and Webber, 2010). The identification of larch as both a foliar and a canker host was unexpected and points to many gaps in the knowledge about the host range and biology of P. ramorum.
Disease Management
Management of P. ramorum–associated diseases in forests, woodlands, and urban areas has taken a multiscale approach ranging from individual trees to landscapes to international quarantines (Alexander and Lee, 2010; Frankel, 2008; Rizzo et al., 2005). Disease prevention and mitigation at the individual plant level or urban–wildland interface in California has been focused on chemical control or other programs designed to maintain health of plants. Some fungicides have been developed that act as protectants (e.g., phosphonates) against infection, but few chemicals have been developed that work once the plant is infected (Garbelotto and Schmidt, 2009; Garbelotto et al., 2009). Removal of inoculum-producing plants, such as bay laurel or rhododendron, has also been important at smaller scales to protect high-value oaks (Swiecki and Bernhardt, 2008). Education and involvement of local communities has been critical at the urban–wildland interface to the implementation of management programs (Alexander and Lee, 2010).
At larger landscape scales in wildland forest communities, management strategies for P. ramorum have included prevention, eradication, treatment, and restoration (Rizzo et al., 2005; Valachovic et al., 2008, 2010). Eradication has been attempted in some cases, most notably with tanoak forests in Oregon (Kanaskie et al., 2010) and larch plantations in the United Kingdom (Brasier and Webber, 2010), but has met with mixed success. Important successes have been balanced by continuing tree mortality in many areas (Kanaskie et al., 2010). Difficulties have been encountered in detecting the pathogen at an early enough stage for eradication to be completely effective at a landscape scale. Cryptic infection (i.e., with minimal or no symptoms) of foliage during the initial invasion of a site by P. ramorum has allowed the pathogen to stay one step ahead of detection efforts in many cases. The development of management strategies, beyond eradication, for forest lands following invasion by P. ramorum is still in the early stages (Rizzo et al., 2005; Valachovic et al., 2008, 2010). Decision making requires the ability to fit disease management into the context of other management goals (e.g., fire prevention, wildlife) within the broader forest landscape (Rizzo et al., 2005). Examples of approaches that are being tested include forest stand thinning to remove inoculum-producing hosts and use of prescribed fire (Valachovic et al., 2008, 2010).
The broadest scale for disease management, regional to international, is driven by regulations and management practices designed to prevent further spread of Phytophthora (Brasier, 2008; Frankel, 2008; Rizzo et al., 2005). In recent years, broadening of national and international quarantines designed to prevent pathogen movement has led to an increased effort to manage all Phytophthora diseases in nursery settings (Osterbauer et al., 2004; Parke et al., 2010a,b). While dozens of plant species have been found infected in nurseries, the majority of infections have been associated with the genera Rhododendron, Camellia, Pieris, Kalmia, and Viburnum (Osterbauer et al., 2004; Parke et al., 2010a). These plant species have become the focal point for development of best management practices and pathogen detection strategies (Parke et al., 2010a). The need for
pathogen detection in nursery plants as part of quarantine inspections has resulted in the development of a number of PCR-based molecular tests (e.g., Hayden et al., 2004; Vettraino et al., 2010).
One consequence of increased detection ability and monitoring for P. ramorum has been the discovery of many additional species of Phytophthora in nurseries and wildlands (e.g., Brasier, 2008; Hansen et al., 2003; Yakabe et al., 2009). Since 2000, the number of described species of Phytophthora has nearly doubled (Brasier, 2009). The ecology and potential long-term impacts of these many “new” species is unknown at this time. Limited knowledge of the baseline biodiversity of Phytophthora (and other fungal plant pathogens) in many areas makes it very difficult to determine if an organism is a newly arrived “exotic” species. Therefore, it becomes very difficult to predict if any of these recently described species will have the large ecological impacts of P. ramorum.
References
Alexander, J., and C. A. Lee. 2010. Lessons learned from a decade of sudden oak death in California: Evaluating local management. Environmental Management 46:315–328.
Anacker, B. L., N. E. Rank, D. Hüberli, M. Garbelotto, S. Gordon, T. Y. Harnik, R. Whitkus, and R. K. Meentemeyer. 2008. Susceptibility to Phytophthora ramorum in a key infectious host: Landscape variation in host genotype, host phenotype, and environmental factors. New Phytologist 177:756–766.
Anderson, P. K., A. A. Cunningham, N. G. Patel, F. J. Morales, P. R. Epstein, and P. Daszak. 2004. Emerging infectious diseases of plants: Pathogen pollution, climate change and agrotechnology drivers. Trends in Ecology & Evolution 19:535–544.
Aukema, J. A., S. McCullough, B. Von Holle, A. M. Liebhold, K. Britton, and S. J. Frankel. 2010. Historical accumulation of nonindigenous forest pests in the continental United States. BioScience 60:886–897.
Brasier, C. M. 2008. The biosecurity threat to the UK and global environment from international trade in plants. Plant Pathology 57:792–808.
———. 2009. Phytophthora biodiversity: How many Phytophthora species are there? In Proceedings of the fourth meeting of the International Union of Forest Research Organizations (IUFRO) Working Party S07.02.09: Phytophthoras in forests and natural ecosystems, edited by E. M. Goheen and S. J. Frankel. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-221. Pp. 101–116.
Brasier, C. M., and J. Webber. 2010. Sudden larch death. Nature 466:824–825.
Brasier, C. M., A. M. Vettraino, T. T. Chang, and A. Vannini. 2010. Phytophthora lateralis discovered in an old growth Chamaecyparis forest in Taiwan. Plant Pathology 59:595–603.
Boutet, X., A. Vercauteren, K. Heungens, F. Laurent, and A. Chandelier. 2010. Oospores progenies from Phytophthora ramorum. Fungal Biology 114:369–378.
Cobb, R. C., R. K. Meentemeyer, and D. M. Rizzo. 2010. Apparent competition in canopy trees determined by transmission rather than susceptibility. Ecology 91:327–333.
Condeso, T. E., and R. K. Meentemeyer. 2007. Effects of landscape heterogeneity on the emerging forest disease sudden oak death. Journal of Ecology 95:364–375.
Cushman, J. H., and R. K. Meentemeyer. 2008. Multi-scale patterns of human activity and the incidence of an exotic forest pathogen. Journal of Ecology 96:766–776.
Davidson, J. M., S. Werres, M. Garbelotto, E. M. Hansen, and D. M. Rizzo. 2003. Sudden oak death and associated diseases caused by Phytophthora ramorum. Online. Plant Health Progress, doi:10.1094/PHP-2003-0707-01-DG.
Davidson, J. M., A. C. Wickland, H. A. Patterson, K. Falk, and D. M. Rizzo. 2005. Transmission of Phytophthora ramorum in mixed-evergreen forests of California. Phytopathology 95:587–597.
Davidson, J. M., H. A. Patterson, and D. M. Rizzo. 2008. Sources of inoculum for Phytophthora ramorum in a redwood forest. Phytopathology 98:860–866.
Davidson, J. M., H. A. Patterson, A. C. Wickland, E. J. Fichtner, and D. M. Rizzo. 2011. Forest type influences transmission of Phytophthora ramorum in California oak woodlands. Phytopathology 101:492–501.
Desprez-Loustau, M. L., and D. M. Rizzo. 2011. Fungi. In Encyclopedia of invasive introduced species, edited by D. Simberloff and M. Rejmanek. Berkeley, CA: University of California Press. Pp. 258–263.
Desprez-Loustau, M. L., C. Robin, M. Buée, R. Courtecuisse, J. Garbaye, F. Suffert, I. Sache, and D. M. Rizzo. 2007. The fungal dimension of biological invasions. Trends in Ecology and Evolution 22:472–480
Dodd, R. S., D. Hüberli, V. Douhovnikoff, T. Y. Harnik, Z. Afzal-Rafii, and M. Garbelotto. 2004. Is variation in susceptibility to Phytophthora ramorum correlated with population genetic structure in coast live oak (Quercus agrifolia Née)? New Phytologist 165:203–214.
Dodd, R. S., D. Hüberli, W. Mayer, T. Y. Harnik, Z. Afzal-Rafii, and M. Garbelotto. 2008. Evidence for the role of synchronicity between host phenology and pathogen activity in the distribution of sudden oak death canker disease. New Phytologist 179:505–514.
Fichtner, E. J., D. M. Rizzo, S. Kirk, and J. Webber. 2011. Root infections may challenge management of invasive Phytophthora species in UK woodlands. Plant Disease 95:13–18.
Frankel, S. J. 2008. Sudden oak death and Phytophthora ramorum in the USA: A management challenge. Australasian Plant Pathology 37:19–25.
Garbelotto, M., and D. M. Rizzo. 2005. A California-based chronological review (1995–2004) of research on Phytophthora ramorum, the causal agent of sudden oak death. Phytopathologia Mediterranea 44:127–143.
Garbelotto, M., and D. J. Schmidt. 2009. Phosphonate controls sudden oak death pathogen for up to 2 years. California Agriculture 63:10–17.
Garbelotto, M., P. Svihra, and D. M. Rizzo. 2001. Sudden oak death syndrome fells three oak species. California Agriculture 55 (1):9–19.
Garbelotto, M., T. Y. Harnik, and D. J. Schmidt. 2009. Efficacy of phosphonic acid, metalaxyl-M and copper hydroxide against Phytophthora ramorum in vitro and in planta. Plant Pathology 58:111–119.
Goss, E. M., M. Larsen, G. A. Chastagner, D. R. Givens, and N. J. Grünwald. 2009. Population genetic analysis infers migration pathways of Phytophthora ramorum in US nurseries. PLoS Pathogens 5:e1000583.
Goss, E. M., M. Larsen, A. Vercauteren, S. Werres, K. Heungens, and N. J. Grünwald. 2011. Phytophthora ramorum in Canada: Evidence for migration within North America and from Europe. Phytopathology 101:166–171.
Grünwald, N. J., E. M. Gross, and C. Press, M. 2008. Phytophthora ramorum: A pathogen with a remarkably wide host range causing sudden oak death on oaks and ramorum blight on woody ornamentals. Molecular Plant Pathology 9(5):1–11.
Hansen, E. M., P. M. Reeser, J. M. Davidson, M. Garbelotto, K. Ivors, L. I. Douhan, and D. M. Rizzo. 2003. Phytophthora nemorosa sp. nov.: An aerial Phytophthora found in forests of California and Oregon. Mycotaxon 88:129–138.
Hansen, E. M., A. Kanaskie, S. Prospero, M. McWilliams, E. M. Goheen, N. Osterbauer, P. M. Reeser, and W. Sutton. 2008. Epidemiology of Phytophthora ramorum in Oregon tanoak forests. Canadian Journal of Forest Research 38:1133–1143.
Hayden, K. J., D. M. Rizzo, J. Tse, and M. Garbelotto. 2004. Development of a PCR-based diagnostic test for Phytophthora ramorum. Phytopathology 94:1075–1083.
Hayden, K. J., A. Nettel, R. S. Dodd, and M. Garbelotto. 2011. Will all the trees fall? Variable resistance to an introduced forest disease in a highly susceptible host. Forest Ecology and Management 261(11):1781–1791.
Hüberli, D. and M. Garbelotto. 2011 (in press). Phytophthora ramorum is a generalist plant pathogen with differences in virulence between isolates from infectious and dead-end hosts. Forest Pathology.
Ivors, K., K. J. Hayden, P. J. M. Bonants, D. M. Rizzo, and M. Garbelotto. 2004. AFLP and phylogenetic analyses of North American and European populations of Phytophthora ramorum. Mycological Research 108:378–392.
Ivors, K., M. Garbelotto, I. D. E. Vries, C. Ruyter-Spira, B. T. E. Hekkert, N. Rosenzweig, and P. J. M. Bonants. 2006. Microsatellite markers identify three lineages of Phytophthora ramorum in US nurseries, yet single lineages in US forest and European nursery populations. Molecular Ecology 15:1493–1505.
Kanaskie, A., E. M. Hansen, E. M. Goheen, N. Osterbauer, M. McWilliams, J. Laine, M. Thompson, S. Savona, H. Timeus, B. Woosley, W Sutton, P. Reeser, R. Schultz, and D. Hilburn. 2010. Detection and eradication of Phytophthora ramorum from Oregon forests, 2001–2008. In Proceedings of the sudden oak death fourth science symposium, edited by S. J. Frankel, J. T. Kliejunas, and K. M. Palmieri. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-229. Pp. 3–5.
Kelly, M., Q. Guo, D. Liu, and D. Shaari. 2007. Modeling the risk of a new invasive forest disease in the United States: An evaluation of five environmental niche models. Computers, Environment and Urban Systems 31:689–710.
Kliejunas, J. T. 2010. Sudden oak death and Phytophthora ramorum: A summary of the literature. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. 181 pp.
Maloney, P. E., S. C. Lynch, S. F. Kane, C. E. Jensen, and D. M. Rizzo. 2005. Establishment of an emerging generalist pathogen in redwood forest communities. Journal of Ecology 93:899–905.
Mascheretti, S., P. J. P. Croucher, A.Vettraino, S. Prospero, and M. Garbelotto. 2008. Reconstruction of the sudden oak death epidemic in California through microsatellite analysis of the pathogen Phytophthora ramorum. Molecular Ecology 17:2755–2768.
Mascheretti, S., P. J. P. Croucher, M. Kozanitas, L. Baker, and M. Garbelotto. 2009. Genetic epidemiology of the sudden oak death pathogen Phytophthora ramorum in California. Molecular Ecology 18:4577–4590.
Meentemeyer, R. K., D. M. Rizzo, W. Mark, and E. Lotz. 2004. Mapping the risk of establishment and spread of sudden oak death in California. Forest Ecology and Management 200:195–214.
Meentemeyer, R. K., B. L. Anacker, W. Mark, and D. M Rizzo. 2008a. Early detection of emerging forest disease using dispersal estimation and ecological niche modeling. Ecological Applications 18:377–390.
Meentemeyer, R. K., N. E. Rank, B. L. Anacker, D. M. Rizzo, and J. H. Cushman. 2008b. Influence of land-cover change on the spread of an invasive forest pathogen. Ecological Applications 18:159–171.
Meentemeyer, R. K., N. E. Rank, D. A. Shoemaker, C. B. Oneal, A. C. Wickland, K. Frangioso, and D. M. Rizzo. 2008c. Impact of sudden oak death on tree mortality in the Big Sur ecoregion of California. Biological Invasions 10:1243–1255.
Meentemeyer, R. K., N. J. Cunniffe, A. R. Cook, R. D. Hunter, D. M. Rizzo, and C. A. Gilligan. 2011. Application of stochastic epidemiological models to realistic landscapes: Spread of the sudden oak death pathogen in California (1990–2030). Ecosphere 2:Article 17.
Monahan, W. B., J. Tse, W. D. Koenig, and M. Garbelotto. 2008. Preserved specimens suggest non-native origins of three species of Phytophthora in California. Mycological Research 112: 757–758.
Oak, S. W., J. Hwang, S. N. Jeffers, and B. M. Tkacz. 2010. Phytophthora ramorum in USA streams from the national early detection survey of forests. In Proceedings of the sudden oak death fourth science symposium, edited by S. J. Frankel, J. T. Kliejunas, and K. M. Palmieri. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-229. Pp. 353–354.
Osterbauer, N. K., J. A. Griesbach, and J. Hedberg. 2004. Surveying for and eradicating Phytophthora ramorum in agricultural commodities. Plant Health Progress, doi:10.1094/PHP-2004-0309-02-RS.
Parke, J. L., N. J. Grünwald, C. Lewis, and V. Fieland. 2010a. A systems approach for detecting sources of Phytophthora contamination in nurseries. In Proceedings of the sudden oak death fourth science symposium, edited by S. J. Frankel, J. T. Kliejunas, and K. M. Palmieri. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-229. Pp. 67–68.
Parke, J. L., J. Pscheidt, R. Regan, J. Hedberg, and N. J. Grünwald. 2010b. The Phytophthora online course: Training for nursery growers. In Proceedings of the sudden oak death fourth science symposium, edited by S. J. Frankel, J. T. Kliejunas, and K. M. Palmieri. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-229. Pp. 355.
Prospero, S., N. J. Grunwald, L. M. Winton, and E. M. Hansen. 2009. Migration patterns of the emerging plant pathogen Phytophthora ramorum on the west coast of the United States of America. Phytopathology 99:739–749.
Rizzo, D. M. 2005. Exotic species and fungi: Interactions with fungal, plant and animal communities, In The fungal community (3rd ed.), edited by J. Dighton, P. Oudemans, and J. White. Boca Raton, FL: CRC Press. Pp. 857–877.
Rizzo, D. M., and M. Garbelotto. 2003. Sudden oak death: Endangering California and Oregon forest ecosystems. Frontiers in Ecology and the Environment 1:197–204.
Rizzo, D. M., M. Garbelotto, J. M. Davidson, G. W. Slaughter, and S. Koike. 2002. Phytophthora ramorum as the cause of extensive mortality of Quercus spp. and Lithocarpus densiflorus in California. Plant Disease 86:205–214.
Rizzo, D. M., M. Garbelotto, and E. M. Hansen. 2005. Phytophthora ramorum: Integrative research and management of an emerging pathogen in California and Oregon forests. Annual Review of Phytopathology 43:309–335.
Spaulding, H. L., and L. K. Rieske. 2011. A glimpse at future forests: Predicting the effects of Phytophthora ramorum on oak forests of southern Appalachia. Biological Invasions 6:1367–1375.
Swiecki, T. J., and E. Bernhardt. 2008. Increasing distance from California bay reduces the risk and severity of Phytophthora ramorum canker in coast live oak. In Proceedings of the third sudden oak death fourth science symposium, edited by S. J. Frankel, J. T. Kliejunas, and K. M. Palmieri. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-214. Pp. 181–194.
Vaclavik, T., A. Kanaskie, E. M. Hansen, J. L. Ohmann, and R. K. Meentemeyer. 2010. Predicting potential and actual distribution of sudden oak death in Oregon: Prioritizing landscape contexts for early detection and eradication of disease outbreaks. Forest Ecology and Management 260:1026–1035.
Valachovic, Y., C. Lee, J. Marshall, and H. Scanlon. 2008. Wildland management of Phytophthora ramorum in northern California forests. In Proceedings of the third sudden oak death fourth science symposium, edited by S. J. Frankel, J. T. Kliejunas, and K. M. Palmieri. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-214. Pp. 305–312.
———. 2010. Forest treatment strategies for Phytophthora ramorum. In Proceedings of the sudden oak death fourth science symposium, edited by S. J. Frankel, J. T. Kliejunas, and K. M. Palmieri. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. Gen. Tech. Rep. PSW-GTR-229. Pp. 239–248.
Venette, R. C., and S. D. Cohen. 2006. Potential climatic suitability for establishment of Phytophthora ramorum within the contiguous United States. Forest Ecology and Management 231:18–26.
Vettraino, A. M., S. Sukno, A. Vannini, and M. Garbelotto. 2010. Diagnostic sensitivity and specificity of different methods used by two laboratories for the detection of Phytophthora ramorum on multiple natural hosts. Plant Pathology 59:289–300.
Werres, S., R. Marwitz, W. A. Man in’t Veld, A. W. A. M. De Cock, P. J. M. Bonants, M. De Weerdt, K. Themann, E. Ilieva, and R. P. Baayen. 2001. Phytophthora ramorum sp. nov., a new pathogen on Rhododendron and Viburnum. Mycological Research 105:1155–1165.
Yakabe, L. E., C. L. Blomquist, S. L. Thomas, and J. D. MacDonald. 2009. Identification and frequency of Phytophthora species associated with foliar diseases in California ornamental nurseries. Plant Disease 93:883–890.
CLIMATE CHANGE, EXTREME WEATHER EVENTS, AND FUNGAL DISEASE EMERGENCE AND SPREAD
Compton J. Tucker,50Karina Yager,50,51Assaf Anyamba,50,52and Kenneth J. Linthicum53
Abstract
Empirical evidence from multiple sources shows the Earth has been warming since the late 19th century. More recently, evidence for this warming trend is strongly supported by satellite data since the late 1970s from the cryosphere, atmosphere, oceans, and land. Those data confirm increasing temperature trends and their consequences (e.g., reduced Arctic sea ice, rising sea level, ice sheet mass loss, etc.). At the same time, satellite observations of the Sun show remarkably stable solar cycles since the late 1970s, when direct observations of the Sun’s total solar irradiance began. Numerical simulation models, driven in part by assimilated satellite data, suggest that future warming trends will lead to not only a warmer planet, but also a wetter and drier climate, depending on location, in a fashion consistent with large-scale atmospheric processes. Continued global warming poses new opportunities for the emergence and spread of fungal disease, as climate systems change at regional and global scales, and as animal and plant species move into new niches.
Our contribution to this proceedings is organized as follows: First, we review empirical evidence for a warming Earth. Second, we show the Sun is not responsible for the observed warming. Third, we review numerical simulation
______________
50 Laboratory for Hydrospheric and Biospheric Science, NASA/Goddard Space Flight Center, Greenbelt, MD 20771.
51 Oak Ridge Associated Universities (ORAU).
52 GESTAR–Universities Space Research Association (USRA), Columbia, MD 21044.
53 USDA/ARS Center for Medical, Agricultural, and Veterinary Entomology, Gainesville, FL.
modeling results that project these trends into the future, describing the projected abiotic environment of our planet in the next 40 to 50 years. Fourth, we illustrate how Rift Valley fever outbreaks have been linked to climate, enabling a better understanding of the dynamics of these diseases, and how this has led to the development of an operational predictive outbreak model for this disease in Africa. Fifth, we project how this experience may be applicable to predicting outbreaks of fungal pathogens in a warming world. Last, we describe an example of changing species ranges due to climate change, resulting from recent warming in the Andes and associated glacier melt that has enabled amphibians to colonize higher elevation lakes, only to be followed shortly by the emergence of fungal disease in the new habitats.
Introduction: Observational Evidence for Global Warming
Among many non-scientists, there appears to be controversy over climate change and its causes. This is paradoxical because there is little debate over human-caused climate change within academic communities, except for some peripheral issues, as noted by Lockwood (2009) and others. Let us look to observational evidence from multiple and different sources to better understand climate change and its implications in this persisting pseudo-controversy (Figure A17-1).
A number of methods are used to collect consistent and long-term datasets to monitor Earth’s properties. Observations of temperatures by thermometers at the surface and by satellite microwave “sounding” of atmospheric temperature profiles are important components of our understanding of global temperature. Since 2003, we have been able to measure ocean temperature profiles using the ARGO global array of 3,000 free-drifting robotic probes that measure the temperature and salinity of the upper 2,000 meters of the ocean. This provides continuous monitoring of ocean temperature, salinity, and currents, with all data made publicly available within hours after collection (Lyman et al., 2010; Wells et al., 2009). The ARGO data are fundamental to climate studies because the oceans absorb ~90 percent of the heat from global warming.
We can measure the extent of glaciers and their variation over time, frequently drawing on historical paintings, photographs, maps, and satellite image archives to determine if they are getting smaller or larger. We also measure the extent of sea ice weekly and monthly using passive microwave radiometers. More recently, we have been able to measure ice mass variations for the Greenland and Antarctic ice sheets, using gravity data from the joint U.S.–German Gravity Recovery and Climate Experiment (GRACE) satellite mission (Swenson and Wahr, 2002).
We have measured sea level globally since 1993 using radar altimeters on-board satellites. Sea level is an unequivocal proxy for global warming: As the Earth warms, sea level rises; as it cools, sea level falls. Lastly, since 1979 we have been able to measure the Sun’s energy output using satellites above the Earth’s
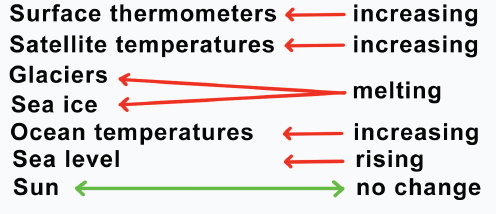

surface. The convergence of observational evidence from all of these sources makes a compelling case that the Earth is warming, and this warming is not due to the Sun.
Convergence of Observations Showing Global Warming
Surface Thermometers
Four global surface temperature datasets are available from the following locations: (1) National Aeronautics and Space Administration (NASA)/Goddard Institute of Space Studies (Hansen et al., 2010), (2) National Oceanic and Atmospheric Administration’s National Climatic Data Center (Smith et al., 2008), (3) the University of East Anglia’s Climate Research Unit (Rayner et al.. 2006), and (4) the Japanese Meteorological Agency.54 These four datasets all use the same input data and differ only in interpolation techniques between sparse observations, how the polar regions are treated, and the reference period for which means are calculated. Not surprisingly, they are very similar (Figure A17-2).
Atmospheric Temperature Profiles
Since late 1978, polar-orbiting satellites have provided global air temperature profiles with altitude or “soundings” using passive microwave radiometers operating between 23 and 89 GHz frequencies. These measurements started with
______________
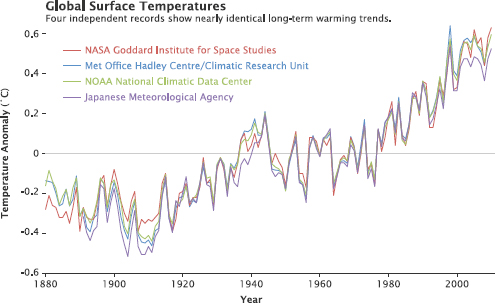
Figure A17-2 A comparison of the existing four global surface temperature datasets that are used in climate analyses. These datasets are based on the same input data and differ by interpolation among stations, treatment of missing data, and the length of the record. The data in this figure have been adjusted to a common baseline period.
SOURCE: Figure courtesy of Robert Simmon, NASA Earth Observatory, provided by Compton J. Tucker NASA Goddard Space Flight Center.
the microwave sounding-unit instruments and were followed by the advanced microwave sounding-unit instruments that began operation in 1988. As is not uncommon in science, early work using atmospheric temperature soundings produced a range of temperature trends. Recent work on atmospheric temperatures has shown no reasonable evidence of disagreement between these measurements and surface observations (Thorne et al., 2011).
Arctic Sea Ice
Satellite observation of sea ice is accomplished with a very high degree of accuracy. This results from spectral emissivity differences between open water (~0.4–0.5) and sea ice (~0.9–1.0) at a wavelength of 1.5 cm (Kwok, 2002). Furthermore, passive microwave radiometers are translucent to clouds, are weather insensitive, and operate from a polar orbit that provides near-daily observations of Arctic sea ice. According to the National Snow and Ice Data Center, all months (i.e., all Januaries, all Februaries, all Marches, etc.) from the late 1970s to 2010–2011 show declining Arctic sea ice with time.55
______________
Sea Level
Sea level is of direct interest to climate science because it varies directly with global mean temperature over short time scales. Temperature affects sea level through two mechanisms: (1) sea level rises through the thermal expansion of water as it warms, or it falls through thermal contraction of water as it cools; and (2) warmer global temperatures melt ice stored on land in glaciers and ice sheets, and the resulting ice loss raises sea level, while cooler global temperatures result in more water being stored on land in glaciers and ice sheets and sea level falls. Thus sea-level variations are an excellent, unambiguous indicator of planetary cooling or warming. For example, at the last glacial maximum, occurring before 20,000 years ago, sea level was >100 m lower than it is now (Clark et al., 2009). This huge quantity of water was stored on land in the form of glaciers and ice sheets (Lambeck et al., 2010).
Although tide gauges provide centennial-scale sea-level records from nearly 10 locations around the world, these few locations are insufficient for a global study of sea level. Researchers have also measured vertical accretion rates in salt marshes as a sea-level proxy, using radiocarbon, pollen, foraminifera, and other markers (reviewed in Mitchum et al., 2010). Since 1993, however, radar altimeters have measured sea level globally and directly with a high precision (Figure A17-3).
We have briefly reviewed global surface thermometer data, atmospheric temperature profile data from satellites, variations in Arctic sea ice, and sea-level data. All of these data unambiguously show the effects of increasing global temperatures.
Earth’s Climate and the Sun
Since the late 1970s, the study of the Sun with instruments on satellites has progressed with continuous observations being collected. Lockwood and Frohlich (2007, 2008) have shown that the three mechanisms where the sun can influence the Earth’s temperature (total solar irradiance, changes in the spectral distribution of solar irradiance, and the solar wind–magnetic field–cosmic ray–cloud hypothesis) have all been opposite to the observed increases in temperatures (Figure A17-4). The Sun’s output is currently at record low values since the satellite era began in the late 1970s (Lockwood, 2009). Thus the Sun is not to blame for the observed global warming since the late 1970s to the present.
Projections of Warming Trends on Weather and Climate
Numerical simulation models of the Earth’s weather and climate are called general circulation models because they simulate the circulation of the atmosphere. They are representations of the ocean, land, sea ice, and atmosphere where the Earth is a series of grid cells driven by energy, moisture, and pres-
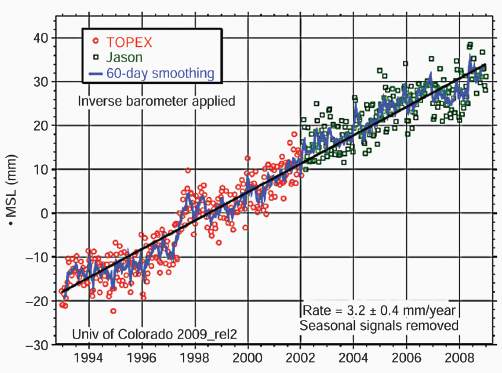
Figure A17-3 Sea-level rise based on radar altimeters from TOPEX and Jason, with seasonal variations removed.
SOURCE: Mitchum et al. (2010).
sure. Each grid cell interacts with adjacent cells horizontally and vertically to simulate climate (Figure A17-5). Model interactions are governed by systems of differential equations and incorporate climate-forcing factors such as land cover change, volcanic aerosols, and increasing greenhouse gas concentrations. Weather and climate models have been shown to be realistic at reproducing the global temperature and precipitation patterns of the 20th century and are widely used in weather and climate research (Delworth et al., 2006).
Climate model simulations, incorporating increasing greenhouse gas concentrations in the atmosphere, have been used to extrapolate precipitation patterns into the 21st century as surface temperatures increase. Several of these climate model simulation predictions can be described as “the wet getting wetter and the dry getting drier” (Held and Soden, 2006). The displacement of arid and semiarid zones northward results from an expansion from the Hadley circulation cell under global warming (Figure A17-6) (Lu et al., 2007). These changes in climate have direct impacts on vegetation and biodiversity across the globe, including species range shifts, changing phenology, new invasive species, and new disease outbreaks (Parmesan and Yohe, 2003; Walther et al., 2002).
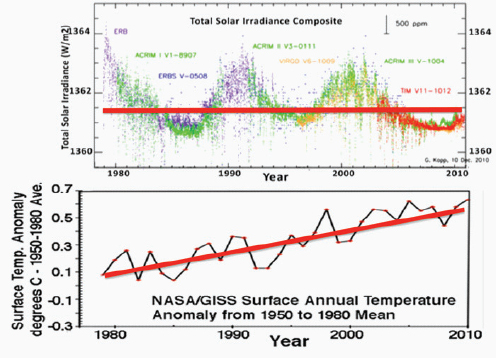
Figure A17-4 A comparison between the total solar irradiance (top) and the NASA/GISS surface temperature data (bottom), both from 1979 to 2010. This shows the sun is not responsible for the 1979 to 2010 increased surface temperatures.
Linkages Between Vector and Non-Vector Diseases and Climate
A variety of infectious diseases have been linked to variations in climate (reviewed in Patz et al., 2005). These include diarrheal diseases (Checkley et al., 1997), cholera (Colwell, 1996; Pascual et al., 2000), salmonella (Kovats et al., 2004), viral pneumonia (Ebi et al., 2001), hantavirus (Glass et al., 2002), influenza (Viboud et al., 2004), flea-associated plague (Parmenter et al., 1999), Culicoides biting midge-associated bluetongue (Baylis et al., 1999; Purse et al., 2005), African horse sickness (Baylis et al., 1999), mosquito-associated Murray Valley encephalitis (Nicholls, 1986), Ross River virus (Woodruff et al., 2002), dengue (Hopp and Foley, 2003; Linthicum et al., 2008), chikungunya (Chretien et al., 2007), malaria (Bouma and Dye, 1997; Bouma et al., 1996), and Rift Valley fever (Linthicum et al., 1999). Of these, we use the example of Rift Valley fever outbreaks to show the use of climate data to understand in what regions and at what times these disease outbreaks will occur.
The link between epizootics of Rift Valley fever and rainfall was first documented by Davies et al. (1985). Through an analysis of time-series rainfall data
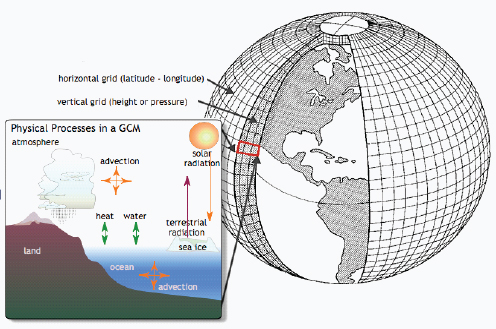
Figure A17-5 Representation of a general circulation model illustrating the grid cell nature of the model on the right, while on the left, many of the different important components of these models are shown.
SOURCE: Figure courtesy of the Center for Multiscale Modeling of Atmospheric Processes (CMMAP), Colorado State University, http://www.cmmap.org/learn/modeling/whatIs2.html.
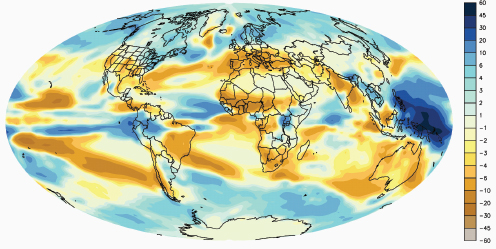
Figure A17-6 Change in precipitation between the 1971–2000 average and the 2091–2100 average in inches of liquid water/year (Held and Soden, 2006).
SOURCE: Geophysical Fluid Dynamics Laboratory, National Oceanic and Atmospheric Administration.
records from numerous stations in Kenya between 1950 and 1982, it was determined that periods with extended positive surplus rainfall corresponded to periods when Rift Valley fever epizootics occurred. Widespread, frequent, and persistent rainfall was shown to be a prominent feature of all epizootic periods. Heavy rainfall raises the level of the water table in certain areas, flooding grassland depressions that are the habitat of the immature stages of certain ground-pool–breeding mosquitoes of the genus Aedes. These findings have been collaborated by findings in southern Africa (Swanepoel, 1976) and West Africa (Bicout and Sabatier, 2004). Rift Valley fever virus is thought to be initially transmitted transovarially in these species. Under prolonged flooded conditions, large numbers of Culex species mosquitoes emerge and are an amplification vector for Rift Valley fever. Following the development of these conditions, Rift Valley fever first occurs in animals and subsequently in humans.
Linthicum et al. (1999) established that outbreaks of Rift Valley fever are closely coupled with above normal rainfall that is associated with the occurrence of the warm phase of El Niño/Southern Oscillation (ENSO) (Cane, 1983; Nicholson, 1986; Ropelewski and Halpert, 1987) and warm events in the equatorial western Indian Ocean (Anyamba et al., 2002; Birkett et al., 1999; Saji et al., 1999). Such warm ocean events precede by 2 to 3 months above normal and extended rainfall over East Africa, and are further enhanced when both the sea surface anomalies in the western Indian Ocean and equatorial central-eastern Pacific are synchronized. More than 90 percent of Rift Valley fever outbreak events since 1950 have occurred during warm ENSO events (Linthicum et al., 1999) (Figure A17-7). The interepizootic period is dominated by La Niña events (the cold phase of ENSO), which results in drought in East Africa and wetter than normal conditions in southern Africa (Anyamba et al., 2002; Nicholson and Entekhabi, 1986). Recent evidence shows that Rift Valley fever outbreaks in southern Africa are coupled with La Niña patterns (Anyamba et al., 2010). Interannual variability, in part driven by ENSO events with differential impacts on rainfall anomaly patterns in Eastern and Southern Africa, largely influences the temporal outbreak patterns of Rift Valley fever.
Our work on Rift Valley fever prediction thus uses climate data to inform us when and where regionally we should expect outbreaks. Subsequent detailed daily satellite observations identify where outbreaks will occur with a high degree of geographical specificity (~60 percent).
Prediction of Rift Valley Fever Outbreaks
Developed by Anyamba et al. (2002), prediction of Rift Valley fever outbreaks includes several components: (1) mapping of potential epizootic/epidemic regions through the combined use of satellite data, climate data, and historical reports; (2) closely following sea surface temperature anomalies with reference to phase and amplitude in the NINO 3.4 tropical Pacific and equatorial western
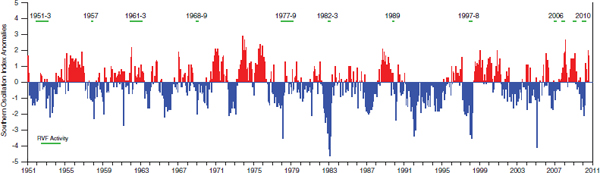
FIGURE A17-7 Rift Valley fever major outbreak events plotted against time and the Southern Oscillation Index, a measure of the phase of El Niño/Southern Oscillation events. Most Rift Valley fever outbreak events have occurred during the warm phase of ENSO (negative Southern Oscillation Index shown in blue).
SOURCE: Linthicum et al. (1999), and updated.
Indian Ocean areas; (3) monitoring patterns of outgoing long-wave radiation anomalies to infer and detect large-scale changes and shifts in the major atmospheric centers of tropical convection as a result of ENSO; and (4) monitoring patterns of normalized difference vegetation index anomalies over Africa as a proxy for excessive rainfall.
The first successful prediction using this system was made in 2006 (Anyamba et al., 2006, 2009) and provided a lead time of 3 to 4 months (Figure A17-8) to respond, although response and mitigation activities only started a month before the first reported outbreak. The predictions were subsequently confirmed by entomological and epidemiological field investigations of virus activity in the areas mapped to be at risk in Kenya, Somalia, and Tanzania with a geographic accuracy of 60 percent (Anyamba et al., 2009). Following the outbreak in East Africa, this system provided further predictions of outbreaks in Sudan in late 2007 and January 2008, 2009, and 2010 in southern Africa. These predictions and outbreak assessments are described in detail in Anyamba et al. (2010).
How Tools and Previous Approaches Could Have Relevance to Anticipating Conditions for Fungal Disease Emergence
We (Linthicum, Anyamba, and Tucker) have been studying the use of satellite data to predict Rift Valley fever outbreaks since the mid-1980s. Our study of Rift Valley fever occurrence led us to the antecedent role of high sea-surface temperatures in the tropical Pacific and western Indian Oceans that results in higher than average rainfall in East Africa, which triggers Rift Valley fever outbreaks. Alerted when the antecedent sea-surface temperature conditions are present, we then step up our near-real-time satellite data surveillance in East Africa that provides very specific location information for control measures to be put in place.
We propose a similar approach for anticipating conditions for fungal disease emergence: Use satellite data to map the abiotic conditions associated with fungal disease outbreaks; evaluate historical fungal outbreaks with respect to antecedent abiotic conditions; and use this knowledge to predict where and when future fungal outbreaks would occur.
A recent example how our prediction model could be applied elsewhere was the role that the very heavy 2011 summer rains have played in increased transmission of Murray Valley encephalitis virus, Ross River virus, and Kunjin virus in Australia (ProMed, 2011b). This same climate anomaly also produced widespread moist soil conditions and increased the likelihood of fungal diseases of cereal crops such as Puccinia graminis f. sp. tritici producing wheat stem rust (Figure A17-9), resulting in the release of warnings for the occurrence of fungal diseases in cereal crops in eastern and southern Australia (ProMed, 2011a).
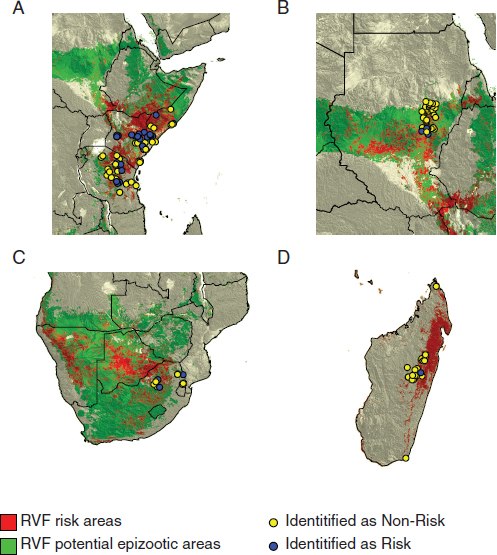
Figure A17-8 Summary Rift Valley fever (RVF) risk maps for (A) Eastern Africa: September 2006–May 2007; (B) Sudan: May 2007–December 2007; (C) Southern Africa: September 2007–May 2008; and (D) Madagascar: September 2007–May 2008. Areas shown in green represent Rift Valley fever potential epizootic areas. Areas shown in red represent pixels that were mapped by the prediction system to be at risk for RVF activity during the respective time periods. Blue dots indicate human cases identified to be with the Rift Valley fever risk areas, while yellow dots represent human cases in areas not mapped to be at risk.
SOURCE: Adapted from Anyamba et al. (2010).

Figure A17-9 Stem rust symptoms on wheat.
SOURCE: Agricultural research service, U.S. Department of Agriculture (http://www.ars.usda.gov/images/docs/9910_10104/stemrust_inset.jpg).
Tropical Glacier Recession, Amphibian Migration, and Subsequent Fungal Migration
Our group at NASA/Goddard Space Flight Center has documented New World tropical glacier variation from the mid-1980s to the present, including glaciers in the Cordillera Vilcanota in Peru. This heavily glaciated range, with multiple peaks over 6,000 m, is a key watershed for regional river systems, including the Amazon. Rapid environmental changes are documented in the region, including record tropospheric warming of 0.3°C per decade between 1974 and 1998 (Vuille et al., 2003), rise in freezing level (Diaz and Graham, 1996), and deglaciation (Bradley et al., 2006; Thompson et al., 2003). According to our current estimates, between the mid-1980s and mid-2000s, there has been approximately 30 percent glacial loss in this particular mountain range of southern Peru (Slayback and Tucker, in preparation). We have found warmer temperatures were largely responsible for tropical glacier recession in these areas (Figure A17-10). No variations in cloud cover or precipitation have been found, indicating global warming was the primary driver of glacial change.
Interdisciplinary research in the Cordillera Vilcanota, around Lake Sibinacocha, has been conducted for several years to investigate the impacts of climate change on local ecosystems (Seimon et al., 2009). This work includes research
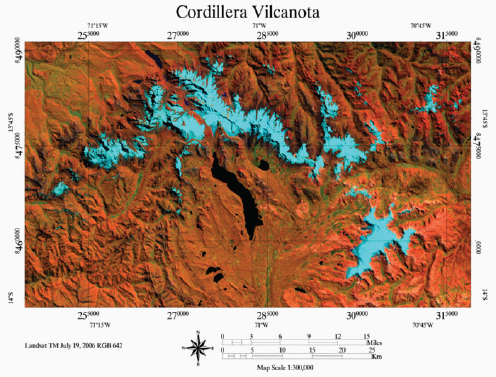
Figure A17-10 False-color Landsat satellite data (RGB 642) showing glaciers as the blue colors. The green colors represent green vegetation and the red colors represent areas of rock, sand, and soil.
SOURCE: Figure provided by Karina Yager, NASA-Goddard Space Flight Center.
on glaciers, vegetation, colonization of microbes in newly deglaciated soils, fossil plants, agropastoralism, species migration, and amphibian studies (Halloy et al., 2006; Nemergut et al., 2007; Seimon et al., 2007; Yager et al., 2010). Glacier retreat at higher elevations in the watershed has been rapid, resulting in the creation of new corridors and newly habitable areas for species migration and the upward range extension of numerous species, including plants, animals, amphibians, and pathogens.
Herpetologists on the team have documented the higher elevation colonization of three species of anurans—Telmatobius marmoratus, Rhinella spinulosa, and Pleurodema marmorata—that have expanded their ranges and moved to unprecedented elevations for amphibians (5,200–5,400 m) into new lakes and ponds created by recent deglaciation (Seimon et al., 2007). In the case of P. marmorata, climatic warming has resulted in an approximate 200 m vertical increase in its range, corresponding to the amount of glacier retreat since 1880 (Seimon et al., 2007).
These amphibian species are opportunistic in their adaptation to the warming climate by migrating to and spawning in ever-higher terrain. However, new climate conditions are also proving advantageous for the spread of epidemic disease, and in particular Chytridiomycosis. This pathogenic chytrid (Bd) produces aquatic zoospores on amphibian skin, and under certain conditions becomes a highly lethal infection (Seimon et al., 2005). Chytrid fungus has been linked to amphibian population declines and even species extinction across the globe (Daszak et al., 1999; Pounds et al., 2006).
New challenges are being presented for the long-term survival of amphibian species in this watershed with climate change. Since 2003, a year after Bd was first detected in this region, all three species have been decreasing in number, and T. marmoratus has not been documented in the Sibinacocha watershed since 2005 (Seimon et al., 2007; Yager et al., 2010). The current research indicates that recent warming, and intense solar heating of glacier ponds during the day, may be contributing to the ability of chytrid to expand and thrive at unprecedented altitudes and terrain (Seimon et al., 2007). In addition, as the glaciers continue to melt, ponds that were once inhabited by amphibians are experiencing a reduction in meltwater or are drying up altogether, leading to loss of habitat and contributing to subsequent population declines. Amphibians are some of the most sensitive species to environmental changes, and are becoming more susceptible to life-threatening disease and possible extinction under current climate patterns.
Conclusions
The Earth’s climate is warming and our Sun is not responsible. Weather and climate simulation models project even warmer temperatures by the middle of this century, with some areas getting wetter and others drier. These changing patterns of temperature and precipitation will alter endemic areas for various plant and animal diseases, including fungal pathogens. We reviewed how knowledge of climatic linkages is being used to predict the outbreak regions of Rift Valley fever in Africa, complemented by detailed satellite observations to identify specific locales where control measures should be undertaken. We advocate a similar approach to identify areas where fungal diseases may emerge: understand the biology of specific fungal pathogens; use satellite data to establish temperature and precipitation climatology in the areas of interest; associate this information with documented fungal outbreaks; and use this knowledge in conjunction with satellite data to predict the impacts of a changing and variable climate on fungal pathogens.
References
Anyamba, A., K. J. Linthicum, R. Mahoney, C. J. Tucker, and P. W. Kelley. 2002. Mapping potential risk of Rift Valley fever outbreaks in African savannas using vegetation index time series data. Photogrammatic Engineering & Remote Sensing 68(2):137–145.
Anyamba, A., J. Chretien, J. Small, C. J. Tucker, and K. J. Linthicum. 2006. Developing global climate anomalies suggest potential disease risks for 2006–2007. International Journal of Health Geographics 5:60. http://www.ij-healthgeographics.com/content/5/1/60 (accessed June 14, 2011).
Anyamba, A., J.-P. Chretien, J. Small, C. J. Tucker, P. Formenty, J. H. Richardson, S. C. Britch, D. C. Schnabel, R. L. Erickson, and K. J. Linthicum. 2009. Prediction of the Rift Valley fever outbreak in the Horn of Africa 2006–2007. Proceedings of the National Academy of Sciences, USA 106(3):955–959.
Anyamba, A., K. J. Linthicum, J. Small, S. C. Britch, E. Pak, S. de La Rocque, P. Formenty, A. W. Hightower, R. F. Breiman, J.-P. Chretien, C. J. Tucker, D. Schnabel, R. Sang, K. Haagsma, M. Latham, H. B. Lewandowski, S. O. Magdi, M. A. Mohamed, P.M. Nguku, J.-M. Reynes, and R. Swanepoel. 2010. Prediction, assessment of the Rift Valley fever activity in East and Southern Africa 2006–2008 and possible vector control strategies. American Journal of Tropical Medicine and Hygiene 81(5):43–51.
Baylis, M., P. Mellor, and R. Meiswinkel. 1999. Horse sickness and ENSO in South Africa. Nature 397(6720):574.
Bicout, D. J., and P. Sabatier. 2004. Mapping Rift Valley fever vectors and prevalence using rainfall variations. Vector-Borne and Zoonotic Diseases 4(1):33–42.
Birkett, C., R. Murtugudde, and T. Allan. 1999. Indian Ocean climate event brings floods to East Africa’s lakes and the Sudd Marsh. Geophysical Research Letters 26(8):1031–1034.
Bouma, M., and C. Dye. 1997. Cycles of malaria associated with El Niño in Venezuela. Journal of the American Medical Association 278(21):1772–1774.
Bouma, M., C. Dye, and J. van der Kay. 1996. Falciparum malaria and climate change in the Northwest Frontier Province of Pakistan. American Journal of Tropical Medicine and Hygiene 55(2):131–137.
Bradley, R. S., M. Vuille, H. F. Diaz, and W. Vergara. 2006. Threats to water supplies in the tropical Andes. Science 312(5781):1755–1756.
Cane, M. A. 1983. Oceanographic events during El Niño. Science 222(4629):77–90.
Checkley, W., L. D. Epstein, R. H. Gilman, D. Figuerora, R. I. Cama, and J. A. Patz. 2000. Effects of El Niño and ambient temperature on hospital admissions for diarrheal diseases in Peruvian children. Lancet 355(9202):442–450.
Chretien, J.-P., A. Anyamba, S. A. Bedno, F. Breiman, R. Sang, K. Sergon, A. M. Powers, C. O. Onyango, J. Small, C. J. Tucker, and K. J. Linthicum. 2007. Drought-associated chikungunya emergence along coastal East Africa. American Journal of Tropical Medicine and Hygiene 76(3):405–407.
Clark, P. U., A. S. Dyke, J. D. Shakun, A. E. Carlson, J. Clark, B. Wohlfarth, J. X. Mitrovica, S. W. Hostetler, and A. M. McCabe. 2009. The last glacial maximum. Science 325(5941):710–714.
Colwell, R. R. 1996. Global climate and infectious disease: The cholera paradigm. Science 274(5295): 2025–2031.
Daszak, P., L. Berger, A. A. Cunningham, A. D. Hyatt, D. E. Green, and R. Speare. 1999. Emerging infectious diseases and amphibian population declines. Emerging Infectious Diseases 5(6): 735–748.
Davies, F. G., K. J. Linthicum, and A. D. James. 1985. Rainfall and epizootic Rift Valley fever. Bulletin of the World Health Organization 63(5):941–943.
Delworth, T. L., A. J. Broccoli, A. Rosati, R. J. Stouffer, V. Balaji, J. A. Beesley, W. F. Cooke, K. W. Dixon, J. Dunne, K. A. Dunne, J. W. Durachta, K. L. Findell, P. Ginoux, A. Gnanadesikan, C. T. Gordon, S. M. Griffies, R. Gudgel, M. J. Harrison, I. M. Held, R. S. Hemler, L. W. Horowitz, S. A. Klein, T. R. Knutson, P. J. Kushner, A. R. Langenhorst, H. C. Lee, S. J. Lin, J. Lu, S. L. Malyshev, P. C. D. Milly, V. Ramaswamy, J. Russell, M. D. Schwarzkopf, E. Shevliakova, J. J. Sirutis, M. J. Spelman, W. F. Stern, M. Winton, A. T. Wittenberg, B. Wyman, F. Zeng, and R. Zhang. 2006. GFDL’s CM2 global coupled climate models—Part 1: Formulation and simulation characteristics. Journal of Climate 19(5):643–674.
Diaz, H. F., and N. E. Graham. 1996. Recent changes in tropical freezing heights and the role of sea surface temperature. Nature 383(6596):152–155.
Ebi, K. L., K. A. Exuzides, E. Lau, M. Kelsh, and A. Barnston. 2001. Association of normal weather periods and El Nino events with hospitalization for viral pneumonia in females: California, 1983–1998. American Journal of Public Health 91(8):1200–1208.
Glass, G. E., T. L. Yates, J. B. Fine, T. M. Shields, J. B. Kendall, A. G. Hope, C. A. Parmenter, C. J. Peters, T. G. Ksiazek, C.-S. Li, J. A. Patz, and J. N. Mills. 2002. Satellite imagery characterizes local animal reservoir populations of Sin Nombre virus in the southwestern United States. Proceedings of the National Academy of Sciences, USA 99(26):16817–16822.
Halloy, S., A. Seimon, and K. Yager. 2006. Multidimensional (climate, biodiversity, socioeconomics, agriculture) context of changes in land use in the Vilcanota watershed, Peru. In Land use change and mountain biodiversity, edited by E. Spehn, M. Liberman-Cruz, and C. Körner. Ft. Lauderdale, FL: CRC Press. Pp. 323–337.
Hansen, J., R. Ruedy, M. Sato, and K. Lo. 2010. Global surface temperature change. Reviews of Geophysics 48(RG4004, 29):1–29.
Held, I. M., and B. J. Soden. 2006. Robust responses of the hydrological cycle to global warming. Journal of Climate 19(21):5686–5699.
Hopp, M. J., and J. A. Foley. 2003. Worldwide fluctuations in dengue fever cases related to climate variability. Climatic Research 25(1):85–94.
Kovats, R. S., S. J. Edwards, S. Hajat, B. G. Armstrong, K. L. Ebi, B. Menne, and the Collaborating Group. 2004. The effect of temperature on food poisoning: A time-series analysis of salmonellosis in ten European countries. Epidemiological Infections 132(3):443–453.
Kwok, R. 2002. Sea ice concentration estimates from satellite passive microwave radiometry and openings from SAR ice motion. Geophysical Research Letters 29(9):25-1–25-4.
Lambeck, K., C. D. Woodroffe, F. Antonioli, M. Anzidei, W. R. Gehrels, J. Laborel, and A. J. Wright. 2010. Paleoenvironmental records, geophysical modeling, and reconstructions of sea-level trends and variability on centennial and longer timescales. In Understanding sea level rise and variability, edited by P. L. Woodworth, J. A. Church, T. Aarup, and W. S. Wilson. New York: Wiley-Blackwell. Pp. 61–121.
Linthicum, K. J., A. Anyamba, C. J. Tucker, P. W. Kelley, M. F. Myers, and C. J. Peters. 1999. Climate and satellite indicators to forecast Rift Valley fever epidemics in Kenya. Science 285(5426):397–400.
Linthicum, K. J., S. C. Britch, A. Anyamba, J. Small, C. J. Tucker, J.-P. Chretien, and R. Sithipraasasna. 2008. Ecology of disease: The intersection of human and animal health. In Vector-borne diseases—Understanding the environmental, human health, and ecological considerations, Institute of Medicine of The National Academies. Washington, DC: National Academy Press. Pp. 78–88.
Lockwood, M. 2009. Solar change and climate: An update in the light of the current exceptional solar minimum. Proceedings of the Royal Society, Series A 466(2114):303–329.
Lockwood, M., and C. Frohlich. 2007. Recent oppositely-directed trends in solar climate forcings and the global mean surface air temperature. Proceedings of the Royal Society, Series A 463(2086):2447–2460.
———. 2008. Recent oppositely-directed trends in solar climate forcings and the global mean surface air temperature. Different reconstructions of the total solar irradiance variation and dependence on response time scale. Proceedings of the Royal Society, Series A. 464(2094):1367–1385.
Lu, J., G. A. Vecchi, and T. Reichler. 2007. Expansion of the Hadley cell under global warming. Geophysical Research Letters 34(L06805):1–5.
Lyman, J. M., S. A. Good, V. V. Gouretski, M. Ishii, G. C. Johnson, M. D. Palmer, S. M. Smith, and J. K. Willis. 2010. Robust warming of the global upper ocean. Nature 465(7296):334–337.
Mitchum, G. T., R. S. Nerem, M. A. Merrifield, and W. R. Gehrels. 2010. Modern sea-level-change estimates. In Understanding sea level rise and variability, edited by P. L. Woodworth, J. A. Church, T. Aarup, and W. S. Wilson. New York: Wiley-Blackwell. Pp. 122–142.
Nemergut, D. R., S. P. Anderson, C. C. Cleveland, A. P. Martin, A. E. Miller, A. Seimon, and S. K. Schmidt. 2007. Microbial community succession in an unvegetated, recently deglaciated soil. Microbial Ecology 53(1):110–122.
Nicholls, N. 1986. A method for predicting Murray Valley encephalitis in southeast Australia using the Southern Oscillation. Australian Journal of Experimental Biology and Medical Science 64:587–594.
Nicholson, S. E. 1986. The spatial coherence of African rainfall anomalies: Inter-hemispheric teleconnections. Journal of Climate and Applied Meteorology 25(10):1365–1381.
Nicholson, S. E., and D. Entekhabi. 1986. The quasi-periodic behavior of rainfall variability in Africa and its relationship to the Southern Oscillation. Archive Fur Meteorologie Geophysik und Bioklimatologie. Series A 34(3–4):311–348.
Parmenter, R. R., E. P. Yadav, C. A. Parmenter, P. Ettestad, and K. L. Gage. 1999. Incidence of plague associated with increased winter–spring precipitation in New Mexico. American Journal of Tropical Medicine and Hygiene 61(5):814–821.
Parmesan, C., and G. Yohe. 2003. A globally coherent fingerprint of climate change impacts across natural systems. Nature 421(6918):37–42.
Pascual, M., X. Rodo, S. Ellner, R. R. Colwell, and M. Boum. 2000. Cholera dynamics and El Niño-Southern Oscillation. Science 289(5485):1766–1769.
Patz, J. A., D. Campbell-Lendrum, T. Holloway, and J. A. Foley. 2005. Impact of regional climate change on human health. Nature 438(7066):310–317.
Pounds, J. A., M. R. Bustamante, L. A. Coloma, J. A. Consuegra, M. P. L. Fogden, P. N. Foster, E. La Marca, K. L. Masters, A. Merino-Viteri, R. Puschendorf, S. R. Ron, G. A. SanchezAzofeifa, C. J. Still, and B. E. Young. 2006. Widespread amphibian extinctions from epidemic disease driven by global warming. Nature 439(7073):161–167.
ProMed. 2011a. Fungal diseases, cereals—Australia: (Eastern, South) Alert. 18 Feb. 2011. Archive number: 20110218.0531.
———. 2011b. Murray Valley encephalitis/Ross River—Australia (03): (Western Australia). 14 April 2011. Archive number: 20110414.1173.
Purse, B. V., P. S. Mellor, D. J. Rogers, A. R. Samuel, P. P. Mertens, and M. Baylis. 2005. Climate change and the recent emergence of bluetongue in Europe. Nature Review of Microbiology 3:171–181.
Rayner, N. A., P. Brohan, D. E. Parker, C. K. Folland, J. J. Kennedy, M. Vanicek, T. Ansell, and S. F. B. Tett. 2006. Improved analyses of changes and uncertainties in marine temperature measured in situ since the mid-nineteenth century: The HadSST2 dataset. Journal of Climate 19(3):446–469.
Ropelewski, C. F., and M. S. Halpert. 1987. Global and regional scale precipitation patterns associated with El Niño/Southern Oscillation. Monthly Weather Review 115(8):1606–1626.
Saji, N. H., B. N. Goswami, P. N. Vinayachandran, and T. Yamagata. 1999. A dipole mode in the tropical Indian Ocean. Nature 401(6751):360–363.
Seimon, T. A., G. Hoernig, P. Sowell, S. Halloy, and A. Seimon. 2005. Identification of chytridiomycosis in Telmatobius marmoratus at 4450 m in the Cordillera Vilcanota of southern Peru. In Studies on the Andean frogs of the genera Telmatobius and Batrachophrynus, Monografıas de Herpetología, Vol. 7, edited by E. O. Lavilla and I. De la Riva. Valencia, Spain: Asociación Herpetológica Española, Monografías de Herpetología. Pp. 275–283.
Seimon, T. A., A. Seimon, P. Daszak, S. Halloy, L. Schloegel, C. Aguilar, P. Sowell, A. D. Hyatt, B. Konecky, and J. E. Simmons. 2007. Upward range extension of Andean anurans and chytridiomycosis to extreme elevations in response to tropical deglaciation. Global Change Biology 13(1):288–299.
Seimon, A., K. Yager, T. Seimon, S. Schmidt, A. Grau, S. Beck, C. García, A. Tupayachi, P. Sowell, J. Touval, and S. Halloy. 2009. Changes in biodiversity patterns in the high Andes: Understanding the consequences and seeking adaptation to global change. Mountain Forum Bulletin 9(2):25–27.
Slayback, D., and C. J. Tucker. In preparation.
Smith, T. M., R. W. Reynolds, T. C. Peterson, and J. Lawrimore. 2008. Improvements to NOAA’s historical merged land–ocean surface temperature analysis (1880–2006). Journal of Climate 21(10):2283–2293.
Swanepoel, R. 1976. Studies on the epidemiology of Rift Valley fever. Journal of the South African Veterinary Association 47:93–94.
Swenson, S., and J. Wahr. 2002. Methods of inferring regional surface-mass anomalies from Gravity Recovery and Climate Experiment (GRACE) measurements of time-variable gravity. Journal of Geophysical Research 107(B9):2193–2205.
Thompson, L. G., E. Mosley-Thompson, M. E. Davis, P. N. Lin, K. Henderson, and T. A. Mashiotta. 2003. Tropical glacier and ice core evidence of climate change on annual to millennial time scales. Climatic Change 59(1–2):137–155.
Thorne, P. W., J. R. Lanzante, T. C. Peterson, D. J. Seidel, and K. P. Shine. 2011. Tropospheric temperature trends: History of an ongoing controversy. Wiley Interdisciplinary Reviews: Climate Change 2(1):66–88.
Viboud, C., K. Pakdaman, P.-Y. Boelle, M. L. Wilson, M. F. Myers, A.-J. Valleron, and A. Flahault. 2004. Association of influenza epidemics with global climate variability. European Journal of Epidemiology 19(11):1055–1059.
Vuille, M., R. S. Bradley, M. Werner, and F. Keimig. 2003. 20th century climate change in the tropical Andes: Observations and model results. Climate Change 59(1–2):75–99.
Walther, G.-R., E. Post, P. Convey, A. Menzel, C. Parmesan, T. J. C. Beebee, J. M. Fromentin, O. Hoegh-Guldberg, and F. Bairlein. 2002. Ecological responses to recent climate change. Nature 416(6879):389–395.
Wells, N. C., S. A. Josey, and R. E. Hadfield. 2009. Towards closure of regional heat budgets in the North Atlantic using Argo floats and surface flux datasets. Ocean Science 5(2):59–72.
Woodruff, R. E., C. Guest, M. G. Garner, N. Becker, J. Lindesay, T. Carvan, and K. Ebi. 2002. Predicting Ross River virus epidemics from regional weather data. Epidemiology 13(4):384–393.
Yager, K., S. Halloy, T. Seimon, A. Seimon, A. Tupayachi Herrera, S. Schmidt, P. Sowell, J. Farfan, and L. N. Thompson. 2010. The Ecology and Environmental History of the GLORIA site in the Cordillera Vilcanota, Peru. Proceedings Paper from the 6th International GLORIA (Global Observation Research Initiative in Alpine Environments) Meeting: Perth, Scotland.
Vance T. Vredenburg,56Cheryl J. Briggs,57and Reid Harris58
Abstract
Amphibian biodiversity is currently facing a severe crisis having recently experienced declines in 42 percent of all species, and as many as 32 percent are
______________
56 Department of Biology, San Francisco State University, San Francisco, CA.
57 Department of Ecology, Evolution and Marine Biology, University of California Santa Barbara, Santa Barbara, CA.
58 Department of Biology, James Madison University, Harrisonburg, VA.
threatened with imminent extinction. The most alarming extinctions and declines have occurred enigmatically in protected, apparently pristine habitats. An emerging infectious disease, chytridiomycosis, is directly linked to the recent extinction or serious decline of hundreds of amphibian species and is increasingly proposed as a primary threat to amphibians. Chytridiomycosis, caused by the fungal pathogen Batrachochytrium dendrobatidis (Bd), infects the skin of amphibians and has been described as causing the greatest loss of vertebrate biodiversity due to disease in recorded history. The severity of the current amphibian biodiversity crisis suggests that Bd is a fundamentally new challenge to amphibians from previous global and environmental changes. While many amphibian species are susceptible to this fungal pathogen, others are silent carriers exhibiting no signs of disease. How amphibian hosts survive with Bd infection is still unknown; however, host immunity, differences in pathogen virulence, and environmental differences that may limit growth and reproduction of the pathogen have all been proposed as possible mechanisms that could lead to Bd-infected host survival. Here we examine amphibian declines in one of the best-documented systems, the Sierra Nevada of California, and review the role that the amphibian skin microbiome may play in host–pathogen dynamics of the chytridiomycosis-amphibian host system.
Introduction
The amphibians are long-term survivors (existing on earth for more than 350 million years) that endured four previous mass extinctions (e.g., 95 percent of all living species were lost in the Permian-Triassic extinction) (Wake and Vredenburg, 2008). Through these extinctions, not only did all three orders of amphibians escape extinction, but most families and genera survived (Wake and Vredenburg, 2008). Today, the amphibians, presently including more than 6,800 species (AmphibiaWeb, 2011), are the most threatened group of vertebrates with over 40 percent of species in decline and over 30 percent threatened with extinction (Stuart et al., 2004). There are many potential causes for the declines, but an emerging infectious disease, chytridiomycosis (Longcore et al., 1999), caused by the infectious fungal pathogen Batrachochytrium dendrobatidis (Bd) is the most alarming (Daszak et al., 2000). This fungal pathogen is associated with the recent decline or extinction of more than 200 species of amphibians (Fisher et al., 2009; Skerratt et al., 2007). Epizootic waves of chytridiomycosis have been identified in Australia, Central and South America, and the western United States (Fisher et al., 2009). In each case the epizootic caused mass mortality and collapse of amphibian faunas (Berger et al., 1998, Lips et al., 2006, Vredenburg et al., 2010a). Once Bd invades naive amphibian host populations, they can collapse very quickly. In Panamanian sites, 50 percent of species were extirpated 4 to 6 months after Bd invaded (Lips et al., 2006). The surviving species declined in population size by 80 percent (Lips et al., 2006). In California, a moving epizootic wave of Bd is causing the collapse of entire metapopulations of ranid frogs
in some of the most protected and pristine areas in the United States (Vredenburg et al., 2010a). In Australia, most of the damage was done before the pathogen was identified (Laurance et al., 1996), but post hoc studies estimate that Bd was responsible for the decline and disappearance of a majority of amphibian species along the diverse eastern tropical montane areas (Fisher et al., 2009).
The Decline of the Mountain Yellow-legged Frog in the Sierra Nevada of California
The dramatic decline of California’s mountain yellow-legged frog (a species complex consisting of Rana muscosa and Rana sierrae; Vredenburg et al., 2007) is emblematic of global amphibian declines (Stuart et al., 2004). Historically, lakes in California’s Sierra Nevada were often inhabited by hundreds of frogs and thousands of tadpoles (Grinnell and Storer, 1924) of this diurnal and highly aquatic taxon (Vredenburg et al., 2005). By 1997, despite the fact that the majority of their habitat is fully protected, R. sierrae and R. muscosa had disappeared from 93 percent and 96 percent, respectively, of their historic range (Vredenburg et al., 2007; Figure A18-1). As a consequence of this decline the two species of mountain yellow-legged frog have gone from being the most common vertebrates in the Sierra Nevada (Grinnell and Storer, 1924) to species classified as “critically endangered” (Stuart et al., 2004). These two species are not the only ones in trouble in the protected landscape of the Sierra Nevada. In fact, five of the seven species of amphibians that occur in the high-elevation areas of the Sierra Nevada (>2,000 m above sea level) are threatened in the range (R. muscosa, R. sierrae, Bufo canorus, B. boreas, and Ambystoma macrodactylum), while only two species (Hydromantes platycephalus and Pseudacris regilla) are thought to have stable populations (Stebbins and Cohen, 1995).
The declines of amphibians that occur in high-elevation protected habitats around the world have been described as enigmatic (Stuart et al., 2004). Decline hypotheses in these areas include the negative effects of increasing UV radiation (Blaustein et al., 1994), pesticide drift (Davidson, 2004), introduced species (Vredenburg, 2004), climate change (Pounds et al., 1999), and disease (Longcore et al., 1999). In the Sierra Nevada, most studies have focused on the decline of the two species of yellow-legged frog. To date, studies have not directly addressed climate change and have shown no evidence for UV radiation effects (Vredenburg et al., 2010b) or pesticide drift (Bradford et al., 2011), but they have shown large negative effects from introduced species (Knapp and Matthews, 2000) and disease (Vredenburg et al., 2010a). Like many montane areas worldwide, the vast majority of the high-elevation portion of the Sierra Nevada, consisting of more than 15,000 glacial lakes, was historically fishless (Knapp and Matthews, 2000). Despite the fact that most of the habitat is protected from habitat destruction, humans changed the landscape dramatically by planting non-native fishes. By 1997 most of the lakes and streams in the area contained self-reproducing
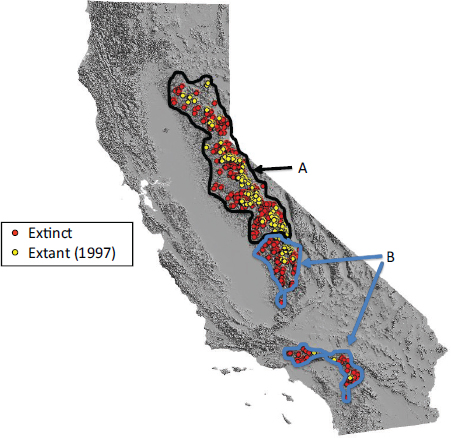
Figure A18-1 Decline of (A) Sierra Nevada mountain yellow-legged frog, Rana sierrae, and (B) southern mountain yellow-legged frog, Rana muscosa, in California, USA. Yellow points indicate extant populations in 1997 and red points indicate extinct populations compared to historically documented sites.
SOURCE: Adapted from Vredenburg et al. (2007).
non-native trout, which have contributed significantly to the decline of the two species of yellow-legged frog (Knapp and Matthews, 2000; Vredenburg, 2004). In 2004 a whole-lake experimental study determined that predation on tadpoles by introduced trout was causing frog population extirpations (Vredenburg, 2004). The same study also demonstrated that threatened frog populations would recover quickly when habitat was restored by removing the non-native fish (Vredenburg, 2004). Additional research showed that habitat restoration (to the fishless condition) worked across the mountain range for both yellow-legged frog species (Knapp et al., 2007). Unfortunately, further research ultimately revealed that amphibian declines in the Sierra Nevada, perhaps like other areas around the world, are complex in nature and can be caused by multiple factors that may act
synergistically. Although it was not understood until 2010, an infectious fungal disease (chytridiomycosis) was also causing population collapse of the yellow-legged frog (Vredenburg et al., 2010a), but it just happened that fish-removal studies occurred ahead of the epizootic wave.
Differential Outcomes for Frog Host Populations After Pathogen Invasion
Chytridiomycosis is caused by the waterborne fungal pathogen Batrachochytrium dendrobatidis, whose only known hosts are larval and adult amphibians. This pathogen was first described in the late 1990s (Berger et al., 1998; Longcore et al., 1999) and is now known to inhabit six continents (Skerratt et al., 2007). The infective stage is a free-living flagellated zoospore that encysts in the skin of an amphibian and develops into a zoosporangium. Zoosporangia produce zoospores via asexual reproduction (sexual reproduction may also occur; Morgan et al., 2007) that are released into the water through a discharge tube (Berger et al., 2005). Tadpoles of most species are minimally affected by chytridiomycosis and the effects on frogs are highly variable, with some species succumbing to the disease within weeks and others experiencing few negative effects (Fisher et al., 2009; Skerratt et al., 2007). Chytridiomycosis likely causes frog mortality by severely disrupting epidermal functions and leading to osmotic imbalance (Voyles et al., 2009).
Despite a growing literature on chytridiomycosis (Fisher et al., 2009; Rosenblum et al., 2010), we have little predictive power in assessing why Bd presence in some areas causes epizootic die-offs while in other areas amphibian hosts survive Bd infections in an apparently enzootic state for years with little or no effect on host population survival (Briggs et al., 2010). Our studies of yellow-legged frogs in California offer one of the most complete insights into the host–pathogen dynamics of chytridiomycosis (Briggs et al., 2010; Vredenburg et al., 2010a). Along with an important study from Central America (Lips et al., 2006), we show that Bd epizootics occur when Bd invades completely naive host populations. We describe the dynamics of Bd and its spread into three naive metapopulations of yellow-legged frogs, each consisting of collections of connected subpopulations (n = 80) within widely separated drainage basins in Sequoia and Kings Canyon National Park (SEKI). Our study was the first to capture the spread of Bd on a small scale (more than hundreds of kilometers) and provided insights into how the pathogen spreads between host populations. The Bd epizootic moved linearly across the landscape at approximately 700 m per year. In one basin (Sixty Lake Basin, Figure A18-2, panels F–J), Bd took 4 years to invade all of the subpopulations (Figure A18-2) and we proposed that Bd was probably invading new host populations through movement of frog hosts as opposed to movement by water (movement was upstream) or by birds or other flying organisms that might carry Bd. (Bd always infected the nearest uninfected host population.) With the data we collected from 1996 to 2008, we demonstrate that Bd invaded naive host
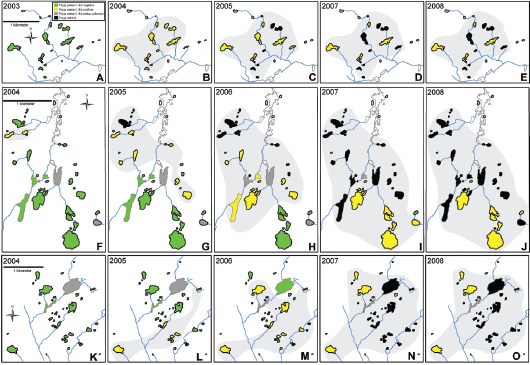
Figure A18-2 Maps of the three study metapopulations showing the spread of Bd and frog population status (adults only) during a 4-year period following the initial detection of Bd. Depicted are (A–E) Milestone Basin, (F–J) Sixty Lake Basin, and (K–O) Barrett Lakes Basin. Lake color (green, yellow, and black) shows the Bd infection and frog population status, and the light gray shaded region surrounds the area in which frog populations were Bd-positive in each year. Lakes shown with a thick black outline are fishless, and a thin gray outline indicates that non-native fish were present. The infection status of frog populations depicted in panels A and K is based on mouthpart surveys of 459 tadpoles. The infection status of frog populations in panels B–J and L–O is based on 4,591 skin swabs analyzed using a real-time PCR assay.
SOURCE: Vredenburg et al. (2010a).
populations of frogs and that infection prevalence quickly reached 100 percent (Figure A18-3, panel B), but mass mortality and host population collapse did not occur until the average Bd infection load (or intensity of infection on individuals) for a population reached more than 104 zoospore equivalents (Figure A18-3, panels A and C; Vredenburg et al., 2010a). This condition was later named the “Vredenburg 10,000 Zoospore Rule” (Kinney et al., 2011) and was shown to also predict mortality in neotropical salamanders (Cheng et al., 2011).
In the Sierra Nevada, infection by Bd did not always lead to epizootic events and host population collapse. About 100 km north of our SEKI study area in and around Yosemite National Park, we describe the Bd–frog dynamics under stable enzootic conditions (Briggs et al., 2010), which contrast sharply with conditions we describe above in the same species of yellow-legged frog in SEKI (Vredenburg et al., 2010a). We discovered that in the Yosemite area, frog host population dynamics are strikingly different and that host populations survive, at least for a decade, despite infection by Bd (Briggs et al., 2010). We determined that while infection prevalence (proportion of infected hosts in a population) could be high (more than 60 percent), the infection intensity of hosts remained low (Briggs et al., 2010), well below the Vredenburg 10,000 Zoospore Rule (Kinney et al., 2011). In fact, several other studies have found that host amphibians do not die when Bd infection intensities remain low (Cheng et al., 2011; Kinney et al., 2011; Kriger et al., 2007; Retallick et al., 2004). Why the dynamics of the host–pathogen interaction is so different in different sites and among different species remains a mystery, but we believe the key to host survival is the low infection intensities on individual host frogs (Briggs et al., 2010; Kinney et al., 2011; Vredenburg et al., 2010a) and salamanders (Cheng et al., 2011).
Mutualistic Bacteria Play a Role in Amphibian Resistance to Fungal Disease
There are several factors that may prevent fungal infection intensities from reaching a lethal threshold in amphibian hosts, such as exposure to a thermal environment that limits Bd growth (Piotrowski et al., 2001), immune response by the host (Ramsey et al., 2010; Rollins-Smith et al., 2000), or density-dependent host-pathogen dynamics (Briggs et al., 2010). Another possibility is that a host’s microbiome may disrupt fungal invasion and growth; in particular, species of mutualistic bacteria may be able to produce compounds that deter fungal colonization and growth. The idea that microbial symbionts may protect amphibian hosts from fungal pathogens was discovered, in a sense, by accident. Until relatively recently, it was assumed that amphibians had few microbes living on their skin since it is generally accepted that amphibians wear their defenses, like armor, on the outsides of their bodies (Duellman and Trueb, 1986). Amphibians are distinct among vertebrates in that they contain specialized skin secretion glands (granular glands) that produce toxins, presumably to deter predators (Duellman
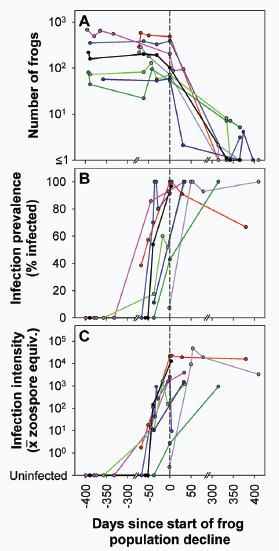
Figure A18-3 Frog–Bd dynamics in eight intensively sampled populations in Milestone and Sixty Lake basins before and after detection of Bd: (A) frog counts (adults + subadults) from visual encounter surveys; (B) infection prevalence, defined as the fraction of skin swabs collected from each population on each date positive for Bd; and (C) infection intensity, defined as the average zoospore equivalents on swabs collected from each population on each date. Data are from frog populations that were sampled more than once per year, that experienced more than 80 percent declines by the end of 2006, and for which the decline in the number of frogs was greater than 10. This last criterion excluded populations that were very small before Bd arrival. Populations are aligned along the x axis such that “0” represents the date on which each frog population began to decline. This was calculated for each population by determining the date at which the number of postmetamorphic frogs dropped below 20 percent of the average population count before that point.
SOURCE: Vredenburg et al. (2010a).
and Trueb, 1986). Many species are well known for this characteristic, such as the poison dart frogs of the neotropics (genus Dendrobates) that produce neurotoxins strong enough to incapacitate monkeys hit by blow-darts from hunters who rub the darts on the skin of the frogs, or the toxic newts (genus Taricha) that produce the same chemical defense compounds, known as tetrodotoxins, found in deadly puffer fish.
Amphibians, although generally tied to water, have colonized many diverse habitats, including many with no standing or running water (Duellman, 1999). In fact, many diverse species have developed reproductive modes that free them in some capacity or completely from water (e.g., direct developers). The four-toed salamander (Hemidactylium scutatum), although not a direct developer, is of interest for this chapter because the female will lay her eggs outside water along steep banks or edges of ponds and she will brood them or guard them until they hatch as larvae. Originally it was presumed that the female was guarding her eggs against egg predators, but behavioral experiments showed that unattended eggs were not eaten by egg predators; in fact, the eggs were unpalatable to many predators (Hess and Harris, 2000), and died instead from fungal infections. Egg-brooding females protect their eggs by inoculating them with colonies of beneficial bacteria (Harris et al., 2006). A laboratory study later showed that the bacterium, Janthinobacteria lividum, cultured from female four-toed salamanders inhibited the growth of Bd (Harris et al., 2006).
Could the information from egg-brooding salamanders in the eastern United States provide insight into why some populations of yellow-legged frogs in California survived Bd infection while others did not? In 2005 we collected bacterial cultures from susceptible and nonsusceptible populations of Rana muscosa and R. sierrae. We found that individual frogs from nonsusceptible populations were more likely to contain culturable populations of the symbiotic bacterium Janthinobacteria lividum, the same species of bacteria discovered on the four-toed salamander (Woodhams et al., 2007). A laboratory study to test whether inoculations of J. lividum could protect susceptible R. muscosa from Bd infections was then initiated. Adult R. muscosa were collected from the wild in SEKI in an area ahead of the Bd epizootic wave and were transported to the Harris Laboratory at James Madison University. The laboratory experiment consisted of three treatments of frogs: one group received J. lividum only, another group received Bd only, and a third was first inoculated with J. lividum and was then infected with Bd. As expected, all of the frogs in the Bd-only treatment died from chytridiomycosis within several weeks, but the frogs inoculated with J. lividum before being infected survived, as did the frogs that only received inoculations of J. lividum, providing evidence that bacterial symbionts could alter the outcome of Bd infections in susceptible host populations (Harris et al., 2009).
Could Bioaugmentation of the Skin Microbiome on Host Frogs Make Bd Epizootics Less Deadly?
The successful laboratory experiment with yellow-legged frogs showed that inoculations with Janthinobacterium lividum can effectively inhibit Bd growth and colonization (Harris et al., 2009). The production of violacein, an antifungal compound (Brucker et al., 2008), appears to be the mechanism used by J. lividum to combat the colonization of Bd (Harris et al., 2009). Our field cultures from admittedly limited Rana muscosa and R. sierrae populations indicated that a small proportion of individual frog hosts even in susceptible populations did naturally include J. lividum in their skin microbiome (<10 percent compared to >80 percent from susceptible and nonsusceptible populations of hosts, respectively; Lam et al., 2010). Nevertheless, because the Bd epizootic was extirpating hundreds of yellow-legged frog populations, we decided to initiate a field bioaugmentation experiment of J. lividum on frogs at a Bd epizootic site in the Sierra Nevada. We planned a replicated whole-lake experiment in the spring of 2010 at a site called Dusy Basin in the northeastermost section of Kings Canyon National Park. This site represented some of the very last susceptible populations of R. sierrae that had yet experienced the Bd epizootic by 2009. By the time we arrived after the spring snowmelt in 2010, there was only one population in Dusy Basin of R. sierrae left containing living adults; all other frogs in the basin were dead, presumably killed by Bd.
In July 2010 we captured 40 adult R. muscosa at a single lake in Dusy Basin and collected bacteria culture samples. Of the 40 frogs sampled, only one successfully resulted in a bacterial culture of J. lividum. We used this single culture to grow large volumes of J. lividum and returned 2 weeks later to Dusy Basin to inoculate as many frogs as we could find with live cultures of J. lividum. We captured 100 adult frogs at the single remaining population in the basin and treated 80 with bacterial baths, leaving 20 as controls. All frogs were marked individually with microchips (PIT tags) and were treated equally in terms of handling time. Control frogs were soaked in pond water from the lake while treatment frogs were soaked in provosoli solution containing live populations of J. lividum in broth. Each animal was weighed, measured, swabbed (for Bd PCR assay; Boyle et al., 2004; Hyatt et al., 2007), and released. We then revisited the site five times in the remaining days of summer, capturing as many frogs as we could find. Each frog was checked for a microchip, weighed, swabbed, and released. By the end of summer, 6 control frogs had been recaptured and 62 bacterially treated frogs had been recaptured. The highest Bd infection intensities were recovered on the control frogs, several control frogs contained infections with >10,000 zoospore equivalents, a lethal infection for the species (Vredenburg et al., 2010a). Due to the high elevation of the site (~3,352 m) and the extremely large amount of precipitation during the winter of 2010–2011, we are not able to return to the site until mid-July 2011. At this time we have no way of knowing what the fate of the population may be. If the microbiome bioaugmentation technique is successful,
it holds great hope for the conservation of susceptible populations of amphibians globally. If the technique does not work, then we must continue to try to understand the factors that may keep Bd infection intensities on individual host amphibians low, because we currently have no way of controlling the spread of this highly virulent pathogen.
Acknowledgments
We thank Harold Werner, David Graber, and Danny Boiano of the National Park Service for their support of our project and to SEKI for providing the proper research permits. We especially thank Lilia Torres, without whom we would not have been able to culture Janthinobacterium lividum, and David Daversa, who collected most of the field data. We thank Anand Varma for taking wonderful photographs; Celeste Dodge, Cory Singer, Sam McNally, and Lauren Gillespie for all the hard work; and Roland Knapp and his field crew for logistics, field work, and planning. We also thank all of the Forum members and the Institute of Medicine for the invitation to participate and LeighAnne Olsen for her patience and kindness to the lead author. Our research is supported by National Science Foundation Grant EF-0723563 as part of the joint National Science Foundation–National Institutes of Health Ecology of Infectious Disease program and by startup funds provided to V. Vredenburg by the College of Science and Engineering at San Francisco State University.
References
AmphibiaWeb: Information on amphibian bioloby and conservation [web application]. Berkeley, California: AmphibiaWeb. Available: http://amphibiaweb.org/ (accessed 2011).
Berger, L., R. Speare, P. Daszak, D. E. Green, A. A. Cunningham, C. L. Goggin, R. Slocombe, M. A. Ragan, A. D. Hyatt, K. R. McDonald, H. B. Hines, K. R. Lips, G. Marantelli, and H. Parkes. 1998. Chytridiomycosis causes amphibian mortality associated with population declines in the rain forests of Australia and Central America. Proceedings of the National Academy of Sciences, USA 95(15):9031–9036.
Berger, L., A. D. Hyatt, R. Speare, and J. E. Longcore. 2005. Life cycle stages of the amphibian chytrid Batrachochytrium dendrobatidis. Diseases of Aquatic Organisms 68(1):51–63.
Blaustein, A. R., P. D. Hoffman, D. G. Hokit, J. M. Kiesecker, S. C. Walls, and J. B. Hays. 1994. UV repair and resistance to solar UV-B in amphibian eggs: A link to population declines? Proceedings of the National Academy of Sciences, USA 91(5):1791–1795.
Boyle, D. G., D. B. Boyle, V. Olsen, J. A. T. Morgan, and A. D. Hyatt. 2004. Rapid quantitative detection of chytridiomycosis (Batrachochytrium dendrobatidis) in amphibian samples using real-time TaqMan PCR assay. Diseases of Aquatic Organisms 60(2):141–148.
Bradford, D. F., R. A. Knapp, D. W. Sparling, M. S. Nash, K. A. Stanley, N. G. Tallent-Halsell, L. L. McConnell, and S. M. Simonich. 2011. Pesticide distributions and population declines of California, USA, alpine frogs, Rana muscosa and Rana sierrae. Environmental Toxicology and Chemistry 30(3):682–691.
Briggs, C. J., R. A. Knapp, and V. T. Vredenburg. 2010. Enzootic and epizootic dynamics of the chytrid fungal pathogen of amphibians. Proceedings of the National Academy of Sciences, USA 107(21):9695–9700.
Brucker, R. M., R. N. Harris, C. R. Schwantes, T. N. Gallaher, D. C. Flaherty, B. A. Lam, and K. P. C. Minbiole. 2008. Amphibian chemical defense: Antifungal metabolites of the microsymbiont Janthinobacterium lividum on the salamander Plethodon cinereus. Journal of Chemical Ecology 34(11):1422–1429.
Cheng, T. L., S. M. Rovito, D. B. Wake, and V. T. Vredenburg. 2011. Coincident mass extirpation of neotropical amphibians with the emergence of the infectious fungal pathogen Batrachochytrium dendrobatidis. Proceedings of the National Academy of Sciences, USA 108(23):9502–9507.
Daszak, P., A. A. Cunningham, and A. D. Hyatt. 2000. Emerging infectious diseases of wildlife—threats to biodiversity and human health. Science 287(5452):443–449.
Davidson, C. 2004. Declining downwind: Amphibian population declines in California and historical pesticide use. Ecological Applications 14(6):1892–1902.
Duellman, W. E. 1999. Patterns of distribution of amphibians. Baltimore: The Johns Hopkins University Press.
Duellman, W. E., and L. Trueb. 1986. Biology of amphibians. New York: McGraw-Hill.
Fisher, M. C., T. W. J. Garner, and S. F. Walker. 2009. Global emergence of Batrachochytrium dendrobatidis and amphibian chytridiomycosis in space, time, and host. Annual Review of Microbiology 63:291–310.
Grinnell, J., and T. Storer. 1924. Animal life in the Yosemite. Berkeley: University of California Press.
Harris, R. N., T.Y. James, A. Lauer, M. A. Simon, and A. Patel. 2006. The amphibian pathogen Batrachochytrium dendrobatidis is inhibited by the cutaneous bacteria of amphibian species. EcoHealth 3:53–56.
Harris, R. N., R. M. Brucker, J. B. Walke, M. H. Becker, C. R. Schwantes, D. C. Flaherty, B. A. Lam, D. C. Woodhams, C. J. Briggs, V. T. Vredenburg, and K. P. C. Minbiole. 2009. Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. ISME Journal 3(7):818–824.
Hess, Z. J., and R. N. Harris. 2000. Eggs of Hemidactylium scutatum (caudata: Plethodontidae) are unpalatable to insect predators. Copeia 2000:597–600.
Hyatt, A. D., D. G. Boyle, V. Olsen, D. B. Boyle, L. Berger, D. Obendorf, A. Dalton, K. Kriger, M. Hero, H. Hines, R. Phillott, R. Campbell, G. Marantelli, F. Gleason, and A. Colling. 2007. Diagnostic assays and sampling protocols for the detection of Batrachochytrium dendrobatidis. Diseases of Aquatic Organisms 73(3):175–192.
Kinney, V. C., J. L. Heemeyer, A. P. Pessier, and M. J. Lannoo. 2011. Seasonal pattern of Batrachochytrium dendrobatidis infection and mortality in Lithobates areolatus: Affirmation of Vredenburg’s “10,000 zoospore rule”. PLoS ONE 6(3):e16708.
Knapp, R. A., and K. R. Matthews. 2000. Non-native fish introductions and the decline of the mountain yellow-legged frog from within protected areas. Conservation Biology 14(2):428–438.
Knapp, R. A., D. M. Boiano, and V. T. Vredenburg. 2007. Removal of nonnative fish results in population expansion of a declining amphibian (mountain yellow-legged frog, Rana muscosa). Biological Conservation 135(1):11–20.
Kriger, K. M., F. Pereoglou, and J.-M. Hero. 2007. Latitudinal variation in the prevalence and intensity of chytrid (Batrachochytrium dendrobatidis) infection in eastern Australia. Conservation Biology 21(5):1280–1290.
Lam, B. A., J. B. Walke, V. T. Vredenburg, and R. N. Harris. 2010. Proportion of individuals with anti-Batrachochytrium dendrobatidis skin bacteria is associated with population persistence in the frog Rana muscosa. Biological Conservation 143(2):529–531.
Laurance, W. F., K. R. McDonald, and R. Speare. 1996. Epidemic disease and the catastrophic decline of Australian rain forest frogs. Conservation Biology 10(2):406–413.
Lips, K. R., F. Brem, R. Brenes, J. D. Reeve, R. A. Alford, J. Voyles, C. Carey, L. Livo, A. P. Pessier, and J. P. Collins. 2006. Emerging infectious disease and the loss of biodiversity in a neotropical amphibian community. Proceedings of the National Academy of Sciences, USA 103:3165–3170.
Longcore, J. E., A. P. Pessier, and D. K. Nichols. 1999. Batrachocytrium dendrobatidis gen. et sp. nov., a chytrid pathogenic to amphibians. Mycologia 91:219–227.
Morgan, J. A. T., V. T. Vredenburg, L. J. Rachowicz, R. A. Knapp, M. J. Stice, T. Tunstall, R. E. Bingham, J. M. Parker, J. E. Longcore, C. Moritz, C. J. Briggs, and J. W. Taylor. 2007. Population genetics of the frog-killing fungus Batrachochytrium dendrobatidis. Proceedings of the National Academy of Sciences, USA 104(34):13845–13850.
Piotrowski, J. S., S. L. Annis, and J. E. Longcore. 2001. Physiology, zoospore behavior, and enzyme production of Batrachochytrium dendrobatidis, a chytrid pathogenic to amphibians. Phytopathology 91(6 Suppl):S121.
Pounds, J. A., M. P. L. Fogden, and J. H. Campbell. 1999. Biological response to climate change on a tropical mountain. Nature 398(6728):611–615.
Ramsey, J. P., L. K. Reinert, L. K. Harper, D. C. Woodhams, and L. A. Rollins-Smith. 2010. Immune defenses against Batrachochytrium dendrobatidis, a fungus linked to global amphibian declines, in the South African clawed frog, Xenopus laevis. Infection and Immunity 78(9):3981–3992.
Retallick, R. W. R., H. McCallum, and R. Speare. 2004. Endemic infection of the amphibian chytrid fungus in a frog community post-decline. PLoS Biology 2(11):1965–1971.
Rollins-Smith, L. A., J. E. Longcore, S. K. Taylor, J. C. Shamblin, J. M. Krepp, and C. Carey. 2000. Antimicrobial peptides of frog skin: A possible defense against pathogens associated with global amphibian declines. Developmental and Comparative Immunology 24(Suppl 1):S20.
Rosenblum, E. B., J. Voyles, T. J. Poorten, and J. E. Stajich. 2010. The deadly chytrid fungus: A story of an emerging pathogen. PLoS Pathogens 6:e10.
Skerratt, L. F., L. Berger, R. Speare, S. Cashins, K. R. McDonald, A. D. Phillott, H. B. Hines, and N. Kenyon. 2007. Spread of chytridiomycosis has caused the rapid global decline and extinction of frogs. EcoHealth 4(2):125–134.
Stebbins, R. C., and N. W. Cohen. 1995. A natural history of amphibians. Princeton, NJ: Princeton University Press.
Stuart, S., J. S. Chanson, N. A. Cox, B. E. Young, A. S. L. Rodrigues, D. L. Fishman, and R. W. Waller. 2004. Status and trends of amphibian declines and extinctions worldwide. Science 306:1783–1786.
Voyles, J., S. Young, L. Berger, C. Campbell, W. F. Voyles, A. Dinudom, D. Cook, R. Webb, R. A. Alford, L. F. Skerratt, and R. Speare. 2009. Pathogenesis of chytridiomycosis, a cause of catastrophic amphibian declines. Science 326:582–585.
Vredenburg, V. T. 2004. Reversing introduced species effects: Experimental removal of introduced fish leads to rapid recovery of a declining frog. Proceedings of the National Academy of Sciences, USA 101(20):7646–7650.
Vredenburg, V. T., G. Fellers, and C. Davidson. 2005. The mountain yellow-legged frog Rana muscosa (camp 1917). In Status and conservation of U.S. amphibians, edited by M. Lanoo. Berkeley: University of California Press. Pp. 563–566.
Vredenburg, V. T., R. Bingham, R. Knapp, J. A. T. Morgan, C. Moritz, and D. Wake. 2007. Concordant molecular and phenotypic data delineate new taxonomy and conservation priorities for the endangered mountain yellow-legged frog. Journal of Zoology (London) 271(4):361–374.
Vredenburg, V. T., R. A. Knapp, T. S. Tunstall, and C. J. Briggs. 2010a. Dynamics of an emerging disease drive large-scale amphibian population extinctions. Proceedings of the National Academy of Sciences, USA 107(21):9689–9694.
Vredenburg, V. T., J. M. Romansic, L. M. Chan, and T. Tunstall. 2010b. Does UV-B radiation affect embryos of three high elevation amphibian species in California? Copeia 2010(3):502–512.
Wake, D. B., and V. T. Vredenburg. 2008. Are we in the midst of the sixth mass extinction? A view from the world of amphibians. Proceedings of the National Academy of Sciences, USA 105:11466–11473.
Woodhams, D. C., V. T. Vredenburg, M. A. Simon, D. Billheimer, B. Shakhtour, Y. Shyr, C. J. Briggs, L. A. Rollins-Smith, and R. N. Harris. 2007. Symbiotic bacteria contribute to innate immune defenses of the threatened mountain yellow-legged frog, Rana muscosa. Biological Conservation 138:390–398.
THE EFFECT OF TRADE-MEDIATED SPREAD OF AMPHIBIAN CHYTRID ON AMPHIBIAN CONSERVATION
Ché Weldon59and Matthew C. Fisher60
Introduction
Accelerating losses of amphibian biodiversity were brought to the forefront of conservation concerns in 1989 at the First World Congress of Herpetology, and the National Research Council Workshop the following year (Blaustein and Wake, 1995). Although obvious causes of amphibian declines were known (e.g., habitat destruction and alteration, introduction of exotic species, pollution), the causes of some declines and disappearances were undetermined, such as enigmatic declines in tropical Australia, Meso-America, and western United States. Detailed retrospective studies of declines in California national parks (Drost and Fellars, 1996) and Monteverde Cloud Forest Preserve in Costa Rica (Pounds et al., 1997) gave conclusive evidence of community-wide declines. Amphibian species have disappeared or declined at such an alarming rate over the past few decades that the phenomenon is now considered to be an amphibian extinction crisis (Mendelson et al., 2006; Wake and Vredenburg, 2008).
In 2004, the Global Amphibian Assessment (GAA) conducted by the International Union for Conservation of Nature (IUCN) revealed that 32.5 percent of the world’s ca. 6,000 amphibian species are currently threatened with extinction, and more than 120 have already disappeared (Stuart et al., 2004). Furthermore, half of 435 rapidly declining species were threatened by enigmatic processes (e.g., disease, climate change) and disease was detected in a significant proportion of families (15/36 families) with threatened species (Stuart et al., 2004). While long suspected, little direct evidence for the emergence of infection existed until a new pathogen to science, the fungal chytrid Batrachochytrium dendrobatidis (Bd), was found in dead and dying frogs collected in the mid-1990s from declining populations in Australia and Panama (Berger et al., 1998; Longcore et al., 1999).
______________
59 Unit for Environmental Research: Zoology, Private Bag X6001, North-West University, Potchefstroom 2520, South Africa.
60 Department of Infectious Disease Epidemiology, Faculty of Medicine, Imperial College London, London W2 1PG, U.K.
Many amphibian declines considered at the time to be the cause of introduced species or anthropogenic activities are now known to have been partly due to chytridiomycosis (Green et al., 2002; Muths et al., 2003). More recently, Bd has been identified as a rapidly emerging amphibian pathogen, capable of acting as both the proximate and ultimate cause of amphibian extinction (Fisher et al., 2009a; Schloegel et al., 2006).
In this article, the contribution of the amphibian–chytridiomycosis spread pathway to amphibian declines is investigated, and conventional as well as novel approaches to amphibian conservation with regard to disease threat are evaluated. The essay argues for greater collaboration among science, conservation, and policy if the chytridiomycosis panzootic is to be warded off more efficiently, and summarizes possible strategies to meet that goal.
The Chytridiomycosis–Amphibian Spread Pathway
Through trade and travel, humans introduce species both intentionally and inadvertently. There is a lucrative global trade in amphibians for purposes ranging from experimental animals to cuisine delicacies to pets. The GAA reported that there are 278 amphibian species in pet trade, with the main centers for export being the wet tropics (Stuart et al., 2008). The commercial collection of wild amphibians has often been unsustainable, resulting in reductions in even highly protected amphibian populations such as the Japanese Giant salamander and the Mountain Chicken frog of the islands of Montserrat and Dominica (Anita et al., 2007; Wang et al., 2004).
The conservation threat of traded species affects local and endemic species when introduced exotic and non-native species become invasive and establish feral populations. In addition, diseases and parasites are often carried along unintentionally with their hosts, allowing them to cross geographic boundaries that historically contained these agents at their source of origin (Cunningham et al., 2003). Not many pathogens become established with invasive species, but those that do have the potential to seriously threaten native wildlife (Lyles and Dobson, 1993). Amphibians are known for their ability to become invasive, as demonstrated by the Invasive Species Specialist Group (ISSG) list of 100 of the world’s most invasive alien species, which includes three amphibians (ISSG, 2008). Also listed is Bd, for which invasive amphibians are now known to be the primary vectors of spread (Garner et al., 2006). In Australia, for example, Bd is found among others in Cane toads, a recently introduced invasive alien species (Fisher and Garner, 2007). This stands as a testimony to the complexity of biological invasion issues where not only is Bd an invasive alien species, but its spread is facilitated by an invasive alien toad. Thus, aspects of the disease impacts on amphibian populations around the world result from the unintentional introduction of a pathogen as an invasive alien species. This leads to a paradox in conservation management, namely that in order to contain the pathogen threatening the organ-
isms targeted for conservation (e.g., amphibians), the very group conservationists are trying to protect need to be mitigated to prevent the pathogen from spreading.
The disease pathway is not always a single invasive host–pathogen system, but may involve multiple host species that are either directly or indirectly related to the source population. Several invasive frog species play a role in the exportation of Bd from South Africa and its ongoing spread and impact on amphibian populations around the world. Trade in African clawed frogs (Xenopus laevis) since the 1930s was identified as one of the first human-mediated pathways for spreading Bd internationally (Weldon et al., 2004). Feral populations that became established in countries where the species had been moved (Weldon et al., 2007) allowed for potential transmission of Bd to naïve amphibian species. Such a link was hypothesized to exist between X. laevis and the American bullfrog Rana catesbeiana, an invasive species and major vector of the disease (Daszak et al., 2004; Weldon et al., 2004). Various other strong links have been confirmed between the international amphibian trade for use in the food, pet, and laboratory industry, and the global dispersal of Bd (Fisher and Garner, 2007; Garner et al., 2006; Schloegel et al., 2009). A second species traded from South Africa, Xenopus gilli, was identified as the source of a Bd infection transmitted to captive Alytes muletensis in Spain (Walker et al., 2008). These infected toads were later reintroduced into their native habitat on the island of Mallorca, thus introducing the pathogen into this disease-naïve ecosystem. What makes this case even more extraordinary is that Bd was transmitted from one critically endangered species to another, dismissing the assumption that the more common, widely distributed species are mostly responsible for disease spread. This case study demonstrates the caution with which reintroduction programs should be approached when there is a potential for disease organisms to be vectored alongside their hosts; it is now clear that invasive frogs and toads have spontaneously exposed many native species of amphibians around the world to Bd, often with catastrophic results.
Conventional Amphibian Conservation Strategy
Conventional approaches to amphibian conservation focus on mitigating factors that have been threatening amphibian populations for more than a century. Primary threats that have traditionally received conservation attention include habitat loss and fragmentation, overexploitation, and species invasion (Stuart et al., 2008). Because disease is a relative newcomer on the scene of threats to amphibian biodiversity, it has traditionally received little conservation attention and proven mitigation measures are generally underdeveloped.
A common approach to conserving species is to make habitat or site the conservation priority as some form of protected area. The species that require protection are then viewed as important features that are present at that site and that qualify it for a particular protected-area designation status (Tucker, 2005). However, no amphibian species is protected from becoming infected with Bd by
the site conservation designations approach because the distribution of disease agents is not restricted by political boundaries. This observation is underscored by the fact that many of the population declines at the start of the chytridiomycosis panzootic occurred at pristine and protected areas.
A close relationship exists between emerging infectious disease and alien invasive species, as demonstrated by the pathways of spread for the chytridiomycosis-amphibian system (Fisher et al., 2009a). So, how can we manage the spread of such trade-associated pathogens? Invasive species can either be managed through the control of introduced species (physical, chemical, or biological control) or through the prevention of introduction (mainly through cross-boundary policies and laws). However, the strategies used in the control or prevention of invasive species do not necessarily account for the spread of infectious disease. Physical control is more effective for relatively contained invasion of conspicuous species and therefore not appropriate for pathogen control. Chemical (antifungal) or biocontrol would be more suitable for a widespread invasion such as Bd, but non-target mortality of amphibian hosts or other fungal species resulting from in situ application remains a high risk. Ultimately, the effective control of introduced species requires rapid response by designated authorities when an invasion is first detected, given that a country’s border-level biosecurity has failed. However, unless detection is rapid, the costs of removing an invasive host species is high, and there is no guarantee that cross-transmission will not have occurred to endemic species (e.g., as was found with the transmission of Bd from North American bullfrogs to European Bufo bufo) (Garner et al., 2006). Thus, it is essential that invasion pathways are not only identified, but effectively regulated if invasion is to be prevented. Because the spread of Bd is associated with the amphibian trade, all intentional introductions of amphibians should be regarded as potential intentional introductions of the pathogen. In general, control of intentional introductions fall under the jurisdiction of governmental agencies responsible for the introductions. Unintentional introductions of Bd (e.g., infected frogs that escape from captivity) occur at a smaller spatial and temporal scale and are more challenging to manage. Regardless of scale, effective prevention of invasion requires a certain amount of legislation and policy development.
Another shortfall of conventional conservation is that the laws governing the conservation of biodiversity do not sufficiently deal with wildlife disease issues. International laws governing management and the conservation of biodiversity consist of agreements by cooperating countries through treaties, conventions, and voluntary participation. For instance, the Convention on Biological Diversity (CBD) consists of members who pledge to pursue a comprehensive approach to conserving biological diversity. The CBD states that all participating governments should attempt to prevent the introduction of, eradicate, or control those alien species that threaten ecosystems, habitats, or species. The conservation challenge, however, is for the individual countries that are either the source or recipient of invasive species to act on these general statements of good intent. A good example
is the Lacy Act of 1900 that identifies a priority list of invasive species requiring conservation management. The federal agencies responsible for implementing the law are the U.S. Fish and Wildlife Service (FWS) and the National Oceanic and Atmospheric Association, but the FWS does not provide specific authority to prohibit the importation of a pathogen.
International trade in species whose existence may be threatened as a result of commerce and other trafficking is regulated by the Convention on International Trade in Endangered Species of Wild Fauna and Flora (CITIES). However, comprehensive data covering all amphibians is hard to find because there is no global database or monitoring system for the trade in non-CITIES species (Schlaepfer et al., 2005). Consequently, control over the international spread of Bd is not regulated under this process when the vectors are non-threatened species such as X. laevis and R. catesbeiana. The World Organisation for Animal Health (OIE) provides guidelines that can be used by member countries to protect themselves from the introduction of diseases and pathogens. Only two amphibian pathogens (Bd and Ranavirus) are currently listed as notifiable to the OIE, thus requiring regulation of the amphibian trade aimed at the prevention of disease spread. Because non-members are under no obligation to adhere to the guidelines set by the OIE and because recommendations rely on voluntary participation and lack any legally binding mechanisms, amphibians traded by both member and non-member countries are often not accompanied by veterinary certification. This shortfall in international legislation may cause ambiguity with respect to biosecurity associated with trade, ultimately contributing to the unabated global spread of pathogens.
Conservation Strategy for Amphibian Disease Threat
The contemporary approach to biological conservation is to begin with an assessment of threatened taxa, which can be used by conservationists as a source of reference for identifying species that should be prioritized for conservation research. When species are threatened by a pathogen with no apparent host specificity, however (as is the case for Bd), species evaluation should take on the form of a risk assessment based on biological and environmental characteristics that predispose species to disease susceptibility. Prediction models are best applied in a precautionary context, when information is used proactively to prevent a catastrophe to biodiversity (e.g., introduction of a novel pathogen). Preventing the importation of non-indigenous species in the first place is an important tool to invasive species management, but a strategy is also needed to effectively contain harmful non-indigenous species once they have become established (Schlaepfer et al., 2005).
Predict Disease Susceptibility and Emergence
The GAA indicated disease as the third most significant threat to Anura globally, but only the seventh for salamanders, with no caecilians known to be threatened by disease (Stuart et al., 2008). When finer analyses are performed (family down to species level), the data become more appropriate as a conservation management tool. Various studies at global and regional scales have produced predictive Bd emergence maps and species priority lists for conservation based on multiple characteristics associated with susceptibility to Bd (www.bd-maps.net; Bielby et al., 2008; Rödder et al., 2009; Ron, 2005) (Figure A19-1). Some of the more common denominators collectively used in Bd predictive modeling include climate, altitude, conservation status, and reproductive mode of amphibian species. These integrated models can be used to predict which specific groups in any newly infected region would have a high probability of being impacted by the disease. The question of chytridiomycosis emergence is complicated by evidence that different strains of Bd can exhibit varying degrees of virulence (Berger et al., 2005; Fisher et al., 2009b). Therefore, the cross-regional/continental introduction of infected amphibians could introduce novel strains of Bd that are more virulent than those already present in a region. Consequently, there is an urgent need to abate the introduction of Bd into regions already affected by chytridiomycosis.
Population Management Strategies
Despite a growing wildlife disease phenomenon as human encroachment into natural areas intensifies, making ecological systems more susceptible to infection and its consequences, surprisingly little research has been done on the management of wildlife diseases (Daszak et al., 2001; Fenichel and Horan, 2007). Mendelson et al. (2006) stated that the only hope to save amphibian species from extinction is management through a combination of coordinated in situ actions with ex situ husbandry programs (e.g., survival-assurance and research colonies) on an unprecedented scale. The mandate of the IUCN is that all critically endangered and extinct in the wild taxa should be subjected to ex situ management to ensure recovery of wild populations (IUCN, 2004).
Assurance-survival populations A management strategy that results from using predictive modeling is to collect endangered species for assurance-survival populations by moving ahead of the planned spread of disease. Thus, by collecting wild animals before the arrival of disease and breeding them in captivity disease-mediated decline is prevented. Assurance-survival populations have particular application when species face threats that cannot be easily or swiftly mitigated in the wild (Pavajeau et al., 2008). The 2005 IUCN Amphibian Conservation Action Plan (ACAP) white papers state that assurance-survival populations are mandatory for amphibian species that will not persist in the wild long enough to recover naturally once environments are restored (e.g., disease is controlled)
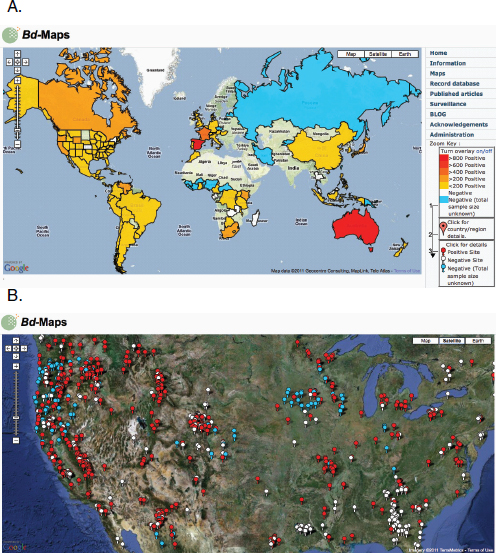
Figure A19-1 Maps indicating (A) the global prevalence and (B) regional U.S. prevalence of Batrachochytrium dendrobatidis as mapped by www.bd-maps.net.
SOURCE: The Global Bd-Mapping Project, www.bd-maps.net.
(IUCN, 2005). This strategy was successfully harnessed in Panama ahead of an encroaching Bd epidemic wave (Lips et al., 2006). Threatened, regional endemics and other specifically selected amphibian species were rescued from El Copé and El Valle and placed in the El Valle Amphibian Conservation Center, an in-country ex situ facility with the objective of long-term maintenance of species until pertinent threats could be mitigated and the species could be reintroduced (Gagliardo et al., 2008). A group of concerned conservation organizations, including the IUCN/SSC (Species Survival Commission) Conservation Breeding Specialist
Group, the World Association of Zoos and Aquariums, and the IUCN/SSC Amphibian Specialist Group initiated the Amphibian Ark (AArk), an organization that provides all levels of support to ex situ actions around the world (Pavajeau et al., 2008). Ex situ conservation centers, in addition to maintaining captive assurance populations, provide a great opportunity for much-needed research on the management of wildlife diseases.
Some concerns that have to be addressed when initiating captive assurance populations is the need of substantial resources for the construction of biosecure facilities, training of keepers, and support of amphibian husbandry requirements (Gagliardo et al., 2008; Pavajeau et al., 2008). In addition, for animals salvaged from areas with recognized disease-mediated die-offs, it is necessary to control the population-limiting infection to help ensure the success of the captive colony and to prevent introduction of disease to other collection animals (Pessier, 2008).
Disease eradication Fortunately, a variety of commonly used disinfectants, including 1 percent sodium hypochlorite (chlorine bleach) and quaternary ammonium compounds, as well as heat (60°C for 5 min) and desiccation, are effective against Bd (Johnson et al., 2003; Webb et al., 2007). Infected and diseased animals from zoological collections have been treated successfully with various antifungal medicines, especially itraconazole baths (Nichols et al., 2000; Parker et al., 2002; Pessier and Mendelson, 2010). Before any chemical treatment can be widely applied, its efficacy must be tested because no treatment for chytridiomycosis has proven to be consistently effective across different amphibian species and life stages (Young et al., 2007). The use of elevated environmental temperatures (37°C for 16 hours) may also clear Bd infection in heat-tolerant species (Woodhams et al., 2003).
However, treatment of Bd in wild amphibian populations presents its own challenges, and we are still far from being able to eliminate Bd from the environment or stop its dispersal. Although Bd can be killed by a range of antifungal medications, it is both impractical and ecologically dangerous to attempt such treatments in the wild due to unknown effects on ecosystems fungal components. However, mathematical models predict that mass mortalities can be reduced if infection loads in affected populations are reduced (Briggs et al., 2010). Furthermore, it is possible to eradicate a disease if the aggregate wildlife population is harvested at a specific level: below a threshold population density and below which disease prevalence declines and above which a disease becomes epidemic if a threshold exists (McCallum et al., 2001). Some of these alternative approaches have been attempted by Bosch et al. (2010) at Peñalara Natural Park in Madrid with the Common midwife toad (Alytes obstetricans), the Mallorcan midwife toad (A. muletensis), and the Betic midwife toad (A. dickhilleni).
Methods employed on wild-caught tadpoles included (1) thermal treatment (21°C) in captivity followed by release after metamorphosis, (2) chemical treatment with itraconazole baths in captivity and subsequent release of tadpoles into
ponds that had been desiccated during treatment, and (3) itraconazole baths in situ in artificial pools combined with habitat improvement, which included desiccation before releasing the tadpoles. The different methods resulted in varying success, although generally, released tadpoles became reinfected and metamorphs had a higher survival rate than what was determined for previous seasons. It seems that Bd eradication is difficult to achieve and that attempts at keeping infection levels controlled in the hope of achieving natural selection against the disease may be the only current approach for in situ mitigation (Bosch et al., 2010). In tandem with antifungal mitigations, the use of probiotic amphibian-associated bacteria holds promise for managing levels of chytrid infection in nature. Specifically, Janthinobacterium lividum has shown to clear infections in captive populations and is currently being widely trialed in a variety of field settings (Harris et al., 2009).
According to Schlaepfer et al. (2005), management practices that rely on continuous intervention are not sustainable indefinitely due to human and monetary resource limitations. Instead, Schlaepfer et al. (2005) suggest a novel approach where native species can respond to changes in their selective regime via evolution or learning that allows coexistence of native and introduced species in cases where the eradication of invasive species is not successful. Although this approach is suggested for invasive species that threaten local species through predation or competition, its application for a pathogen like Bd—capable of rapid population extirpations and even extinction through disease—remains to be explored. Some evidence exists that amphibian populations that have experienced chytridiomycosis-associated decline may partially recover and persist with endemic Bd infection (Retallick et al., 2004; Woodhams et al., 2007). These observations raise hope that some species can acquire resistance to chytridiomycosis in the wild, lessening the reliance on the development of survival-assurance colonies for species threatened by this disease (Mendelson et al., 2006; Young et al., 2007).
Reintroduction Ultimately, the goal of population management that begins with the capture of wild amphibians is to combine ex situ treatment with the selection of resistant individuals that could form the basis of lineages for future reintroduction. Before amphibians can be released into the wild, a number of precautions related to infectious disease need to be considered. Collections that comprise several species increase the likelihood of infection with a variety of potentially novel infectious agents (Cunningham, 1996), a situation that warrants prerelease screening for chytridiomycosis and other infectious disease, both known and unknown. Pessier (2008) demonstrated how the spectrum of infectious agents affecting amphibians is more varied than realized previously and argued for enhancement of the understanding of disease in conservation programs, which includes permanent quarantine housing of species combined with comprehensive disease surveillance and treatment. However, it is possible that hosts raised in
captivity, who are continually treated to eliminate infections, may suffer a disadvantage when released into the wild because of increased susceptibility to infection caused by relaxed selection for resistance, especially in traits that are costly to maintain (Smith et al., 2009). Given this possibility, species management programs, especially those that include captive breeding and reintroductions, may need to focus on maintaining the levels of immunity or variation in resistance that are present in natural populations (Altizer and Pedersen, 2008).
Summary
The diversity and apparent increase in wildlife diseases, including chytridiomycosis, have raised concerns that pathogens may pose a substantial threat to biodiversity. The amphibian extinction crisis has been held to represent the greatest species conservation challenge in the history of humanity (Pavajeau et al., 2008). In part, this crisis has been accelerated by the spread of Bd through the global amphibian trade. The ability of some amphibians to become alien invasive species while functioning as carriers of chytridiomycosis complicates conservation management of the matter. Response to amphibian threats usually involves introducing or rising protection measures of the species and/or their habitats. However, dealing with chytridiomycosis in wildlife populations requires novel strategies that conventional approaches to and policy for amphibian conservation do not adequately provide. It requires greater collaboration among wildlife ecologists, veterinarians, and conservation organizations to provide a more comprehensive mitigation strategy that may significantly reduce the risk of chytridiomycosis-mediated population declines.
The global conservation community responded to this need in the ACAP, of which the AArk plays an integral part, to select species that would otherwise go extinct for captive assurance populations until they can be secured in the wild (Pavajeau et al., 2008). Such facilities are ideally positioned to combine the management of endangered species through assurance colonies with opportunities for conservation research that may include the development of new approaches to mitigate wildlife diseases. Future efforts to better manage and protect the planet’s amphibian diversity will depend on innovation from both the sciences and the world of policies and regulation. A wide spectrum of international and domestic regulation regimes already play a significant role in shaping the conservation status of amphibian biodiversity, but considerate input from the scientific community will make implementation more effective.
References
Altizer, S., and A. B. Pedersen. 2008. Host–pathogen evolution, biodiversity and disease risks for natural populations. In Conservation biology: Evolution in action, edited by S. Carroll and C. Fox. Oxford, UK: Oxford University Press. Pp. 259–278.
Anita, M., R. S. Thorpe, E. Hypolite, and A. James. 2007. A report on the status of the herpetofauna of the Commonwealth of Dominica, West Indies. Applied Herpetology 4(2):177–194.
Berger, L., R. Speare, P. Daszak, D. E. Green, A. A. Cunningham, C. L. Goggin, R. Slocombe, M. A. Ragan, A. D. Hyatt, K. R. McDonald, H. B. Hines, K. R. Lips, G. Marantelli, and H. Parkes. 1998. Chytridiomycosis causes amphibian mortality associated with population declines in the rain forests of Australia and Central America. Proceedings of the National Academy of Sciences, USA 95:9031–9036.
Berger, L., G. Marantelli, L. F. Skerratt, and R. Speare. 2005. Virulence of the amphibian chytrid fungus Batrachochytrium dendrobatidis varies with the strain. Diseases of Aquatic Organisms 68:47–50.
Bielby, J., N. Cooper, A. A. Cunningham, T. W. J. Garner, and A. Purvis. 2008. Predicting susceptibility to future declines in the world’s frogs. Conservation Letters 1:82–90.
Blaustein, A. R., and D. B. Wake. 1995. The puzzle of declining amphibian populations. Scientific American 272:52–57.
Bosch, J., S. Fernández-Beaskoetxea, and B. Martín-Beyer. 2010. Time for chytridiomycosis mitigation in Spain. Aliens: The Invasive Species Bulletin 30:54–58.
Briggs, C. J., R. A. Knapp, and V. T. Vredenburg. 2010. Enzootic and epizootic dynamics of the chytrid fungal pathogen of amphibians. Proceedings of the National Academy of Sciences, USA 107:9695–9700.
Cunningham, A. A. 1996. Disease risks of wildlife translocations. Conservation Biology 10:349–353.
Cunningham, A. A., P. Daszak, and J. P. Rodriguez. 2003. Pathogen pollution: Defining a parasitological threat to biodiversity conservation. Journal of Parasitology 89(Suppl):S78–S83.
Daszak, P., A. A. Cunningham, and A. D. Hyatt. 2001. Anthropogenic environmental change and the emergence of infectious disease in wildlife. Acta Tropica 78:103–116.
Daszak, P., A. Strieby, A. A. Cunningham, J. E. Longcore, C. C. Brown, and D. Porter. 2004. Experimental evidence that the bullfrog (Rana catesbeiana) is a potential carrier of chytridiomycosis, an emerging fungal disease of amphibians. Herpetological Journal 14:201–207.
Drost, C. A., and G. M. Fellars. 1996. Collapse of a regional frog fauna in the Yosemite area of the California Sierra Nevada, USA. Conservation Biology 10(2):414–425.
Fenichel, E. P. and R. D. Horan. 2007. Jointly-determined ecological thresholds and economic tradeoffs in wildlife disease management. Natural Resource Modeling 20(4):511–547.
Fisher, M. C., and T. W. J. Garner. 2007. The relationship between the emergence of Batrachochytrium dendrobatidis, the international trade in amphibians and introduced amphibian species. Fungal Biology Reviews 21:2–9.
Fisher, M. C., T. W. J. Garner, and S. F. Walker. 2009a. The global emergence of Batrachochytrium dendrobatidis and amphibian chytridiomycosis in space, time and host. Annual Review of Microbiology 63:291–310.
Fisher, M. C., J. Bosch, Z. Yin, D. A. Stead, J. Walker, L. Selway, A. J. Brown, L. A. Walker, N. A. Gow, J. E. Stajich, and T. W. J. Garner. 2009b. Proteomic and phenotypic profiling of the amphibian pathogen Batrachochytrium dendrobatidis show that genotype is linked to virulence. Molecular Ecology 18:415–426.
Gagliardo, R., P. Crump, E. Griffith, J. Mendelson, H. Ross, and K. Zippel. 2008. The principles of rapid response for amphibian conservation, using the programmes in Panama as an example. International Zoo Yearbook 42:125–135.
Garner, T. W. J., M. W. Perkins, P. Govindarajulu, D. Seglie, S. Walker, A. Cunningham, and M. C. Fisher. 2006. The emerging amphibian pathogen, Batrachochytrium dendrobatidis, globally infects introduced populations of the North American bullfrog, Rana catesbeiana. Biology Letters 2:455–459.
Green, D. E., K. A. Converse, and A. Schrader. 2002. Epizootiology of sixty-four amphibian morbidity and mortality events in the USA, 1996–2001. Annals of the New York Academy of Sciences 969:323–339.
Harris, R. N., R. M. Brucker, J. B. Walke, M. H. Becker, C. R. Schwantes, D. C. Flaherty, B. A. Lam, D. C. Woodhams, C. J. Briggs, V. T. Vredenburg, and K. P. C. Minbiole. 2009. Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. International Society for Microbial Ecology 3:818–824.
ISSG (Invasive Species Specialist Group). 2008. IUCN ISSG—About Us. http://www.issg.org/about. htm (accessed March 25, 2011).
IUCN (International Union for Conservation of Nature). 2004. IUCN technical guidelines on the management of ex-situ populations for conservation. Gland: IUCN. http://www.iucn.org/dbtwwpd/edocs/Rep-2002-017.pdf (accessed March 25, 2011).
———. 2005. Amphibian conservation summit declaration. IUCN. http://intranet.iucn.org/webfiles/doc/SSC/SSCwebsite/GAA/ACAP_Summit_Declaration.pdf (accessed March 25, 2011).
Johnson, M. L., L. Berger, L. Phillips and R. Speare. 2003. Fungicidal effects of chemical disinfectants, UV light, desiccation and heat on the amphibian chytrid Batrachochytrium dendrobatidis. Diseases of Aquatic Organisms 57:255–260.
Lips, K. R., F. Brem, R. Brenes, J. D. Reeve, R. A. Alford, J. Voyles, C. Carey, L. Livo, A. P. Pessier, and J. P. Collins. 2006. Emerging infectious disease and the loss of biodiversity in a neotropical amphibian community. Proceedings of the National Academy of Sciences, USA 103(9):3165–3170.
Longcore, J. E., A. P. Pessier, and D. K. Nichols. 1999. Batrachochytrium dendrobatidis gen. et sp. nov., a chytrid pathogenic to amphibians. Mycologia 91(2):219–227.
Lyles, A. M., and A. P. Dobson. 1993. Infectious disease and intensive management: Population dynamics, threatened hosts, and their parasites. Journal of Zoo and Wildlife Medicine 24:315–326.
McCallum, H., N. Barlow, and J. Hone. 2001. How should pathogen transmission be modeled? Trends in Ecology and Evolution 16(6):295–300.
Mendelson, J. R., III, K. R. Lips, R. W. Gagliardo, G. B. Rabb, J. P. Collins, J. E. Diffendorfer, P. Daszak, R. Ibanez, K. C. Zippel, D. P. Lawson, K. M. Wright, S. N. Stuart, C. Gascon, H. R. Da Silva, P. A. Burrowes, R. L. Joglar, E. Lamarca, S. Lotters, L. H. Du Preez, C. Weldon, A. Hyatt, J. V. Rodriguez-Mahecha, S. Hunt, H. Robertson, B. Lock, C. J. Raxworthy, D. R. Frost, R. C. Lacy, R. A. Alford, J. A. Campbell, G. Parra-Olea, F. Bolanos, J. J. C. Domingo, T. Halliday, J. B. Murphy, M. H. Wake, L. A. Coloma, S. L. Kuzmin, M. S. Price, K. M. Howell, M. Lau, R. Pethiyagoda, M. Boone, M. Lannoo, A. R. Blaustein, A. Dobson, R. A. Griffiths, M. L. Crump, D. B. Wake, and E. D. Brodie. 2006. Confronting amphibian declines and extinctions. Science 313:48.
Muths, E., P. S. Corn, A. P. Pessier, and D. E. Green. 2003. Evidence for disease-related amphibian decline in Colorado. Biological Conservation 110:357–365.
Nichols, D. K., E. W. Lamirande, and A. P. Pessier. 2000. Experimental transmission and treatment of cutaneous chytridiomycosis in poison dart frogs (Dendrobates azureus and Dendrobates tinctorius). In Joint proceedings American Association of Zoo Veterinarians and the International Association for Aquatic Animal Medicine, edited by C. K. Baer. Yulee, FL: American Association of Zoo Veterinarians. Pp. 42–44.
Parker, J. M., I. Mikaelian, N. Hahn, and H. E. Diggs. 2002. Clinical diagnosis and treatment of epidermal chytridiomycosis in African clawed frogs (Xenopus tropicalis). Comparative Medicine 52:265–268.
Pavajeau, L., K. C. Zipel, R. Gibson, and K. Johnson. 2008. Amphibian Ark and the 2008 Year of the Frog Campaign. International Zoo Yearbook 42:24–29.
Pessier, A. P. 2008. Management of disease as a threat to amphibian conservation. International Zoo Yearbook 42:30–39.
Pessier, A. P., and J. R. Mendelson (eds.). 2010. A manual for control of infectious diseases in amphibian survival assurance colonies and reintroduction programs. Apple Valley, MN: IUCN/SSC Conservation Breeding Specialist Group.
Pounds, J. A., M. P. L. Fogden, J. M. Savage, and G. C. Gorman. 1997. Tests of null models for amphibian declines on a tropical mountain. Conservation Biology 11:1307–1322.
Retallick, R. W. R., H. McCallum, and R. Speare. 2004. Endemic infection of the amphibian chytrid fungus in a frog community post-decline. PLoS Biology 2:1965–1971.
Rödder, D., J. Kielgast, J. Bielby, S. Schmidtlein, J. Bosch, T. W. J. Garner, M. Veith, S. Walker, M. C. Fisher, and S. Lötters. 2009. Global amphibian extinction risk assessment for the panzootic chytrid fungus. Diversity 1:52–66.
Ron, S. R. 2005. Predicting the distribution of the amphibian pathogen Batrachochytrium dendrobatidis in the New World. Biotropica 37(2):209–222.
Schlaepfer, M. A., P. W. Sherman, B. Blossey, and M. C. Runge. 2005. Introduced species as evolutionary traps. Ecology Letters 8:241–246.
Schloegel, L. M., J. M. Hero, L. Berger, R. Speare, K. McDonald, and P. Daszak. 2006. The decline of the sharp-snouted day frog (Taudactylus acutirostris): The first documented case of extinction by infection in a free-ranging wildlife species? EcoHealth 3:35–40.
Schloegel, L. M., A. Picco, A. M. Kilpatrick, A. Hyatt, and P. Daszak. 2009. Magnitude of the U.S. trade in amphibians and presence of Batrachochytrium dendrobatidis and ranavirus infection in imported North American bullfrogs (Rana catesbeiana). Biological Conservation 142:1420–1426.
Smith, K. F., K. Acevedo-Whitehouse, and A. B. Pedersen. 2009. The role of infectious diseases in biological conservation. Animal Conservation 12:1–12.
Stuart, S., J. S. Chanson, N. A. Cox, B. E. Young, S. L. Rodrigues, D. L. Fischman, and R. W. Waller. 2004. Status and trends of amphibian declines and extinctions worldwide. Science 306:1783–1786.
Stuart, S. N., M. Hoffmann, J. S. Chanson, N. A. Cox, R. J. Berridge, P. Ramani, and B. E. Young (eds.). 2008. Threatened amphibians of the world. International Union for Conservation of Nature. Gland, Switzerland, and Conservation International, Arlington, VA. Barcelona, Spain; Lynx Editions.
Tucker, G. 2005. Biodiversity evaluation methods. In Handbook of biodiversity methods, edited by D. Hill, M. Fasham, G. Tucker, M. Shewry, and P. Shaw. Cambridge, UK: Cambridge University Press. Pp. 65–101.
Wake, D. B., and V. T. Vredenburg. 2008. Are we in the midst of the sixth mass extinction? A view from the world of amphibians. Proceedings of the National Academy of Sciences, USA 105(Suppl):11466–11473.
Walker, S. F., J. Bosch, T. Y. James, A. P. Litvintseva, J. A. O. Valls, S. Piña, G. García, G. A. Rosa, A. A. Cunningham, S. Hole, R. Griffiths, and M. C. Fisher. 2008. Invasive pathogens threaten species recovery programs. Current Biology 18:R853–R854.
Wang, X., K. Zhang, Z. Wang, Y. Ding, W. Wu, and S. Huang. 2004 The decline of the Chinese giant salamander Andrias davidianus and implications for its conservation. Oryx 38:197–202.
Webb, R., D. Mendez, L. Berger, and R. Speare. 2007. Additional disinfectants effective against the amphibian chytrid fungus Batrachochytrium dendrobatidis. Diseases of Aquatic Organisms 74:13–16.
Weldon, C., L. H. Du Preez, A. D. Hyatt, R. Muller, and R. Speare. 2004. Origin of the amphibian chytrid fungus. Emerging Infectious Diseases 10(12):2000–2005.
Weldon, C., A. L. De Villiers, and L. H. Du Preez. 2007. Quantification of the African clawed frog trade from South Africa, with implications for biodiversity conservation. African Journal of Herpetology 56:77–83.
Woodhams, D. C., R. A. Alford, and G. Marantelli. 2003. Emerging disease of amphibians cured by elevated body temperature. Diseases of Aquatic Organisms 55:65–67.
Woodhams, D. C., V. T. Vrendenberg, M. Simon, D. Billheimer, B. Shakhtour, Y. Shyr, C. Briggs, L. Rollins-Smith, and R. Harris. 2007. Symbiotic bacteria contribute to innate immune defenses of the threatened mountain yellow-legged frog, Rana muscosa. Biological Conservation 138:390–398.
Young, S., L. Berger, and R. Speare. 2007. Amphibian chytridiomycosis: Strategies for captive management and conservation. International Zoo Yearbook 41:85–95.
WHITE-NOSE SYNDROME FUNGUS (GEOMYCES DESTRUCTANS) IN BATS, EUROPE61
Gudrun Wibbelt, Andreas Kurth, David Hellmann, Manfred Weishaar,
Alex Barlow, Michael Veith, Julia Prüger, Tamás Görföl,
Lena Grosche, Fabio Bontadina, Ulrich Zöphel, Hans-Peter Seidl,
Paul M. Cryan, and David S. Blehert62
White-nose syndrome is an emerging disease in North America that has caused substantial declines in hibernating bats. A recently identified fungus (Geomyces destructans) causes skin lesions that are characteristic of this disease. Typical signs of this infection were not observed in bats in North America before white-nose syndrome was detected. However, unconfirmed reports from Europe indicated white fungal growth on hibernating bats without associated deaths. To investigate these differences, hibernating bats were sampled in Germany, Switzerland, and Hungary to determine whether G. destructans is present in Europe. Microscopic observations, fungal culture, and genetic analyses of 43 samples from 23 bats indicated that 21 bats of 5 species in 3 countries were colonized by G. destructans. We hypothesize that G. destructans is present throughout Europe and that bats in Europe may be more immunologically or behaviorally resistant to G. destructans than their congeners in North America because they potentially coevolved with the fungus.
White-nose syndrome (WNS) is a recently emerged wildlife disease in North America, which in 4 years has resulted in unprecedented deaths of hibernating bats in the northeastern United States (Blehert et al., 2009; Reichard and Kunz, 2009; Turner and Reeder, 2009), and is a widespread epizootic disease among bats. Although we have searched the literature describing observations of hibernating bats, we have been unable to find any similar historical accounts of white fungus growing on live hibernating bats in North America before the recent emergence of WNS.
______________
61 Emerging Infectious Diseases * www.cdc.gov/eid * Vol. 16, No. 8, August 2010.
62 Author affiliations: Leibniz Institute for Zoo and Wildlife Research, Berlin, Germany (G. Wibbelt); Robert Koch Institute, Berlin (A. Kurth); University of Oldenburg, Oldenburg, Germany (D. Hellmann); Bat Conservation Working Group, Gusterath, Germany (M. Weishaar); Veterinary Laboratory Agency, Somerset, UK (A. Barlow); Trier University, Trier, Germany (M. Veith); Coordination Agency for Bat Protection in Thuringia, Erfurt, Germany (J. Prüger); Nature Conservation Foundation of Tolna County, Szekszárd, Hungary (T. Görföl); Echolot GbR, Münster, Germany (L. Grosche); SWILD–Urban Ecology and Wildlife Research, Zurich, Switzerland (F. Bontadina); Saxonian State Office for Environment, Agriculture and Geology, Dresden-Pillnitz, Germany (U. Zöphel); Technical University Munich, Munich, Germany (H.-P. Seidl); US Geological Survey, Fort Collins, Colorado, USA (P. M. Cryan); and US Geological Survey, Madison, Wisconsin, USA (D. S. Blehert).
DOI: 10.3201/eid1608.100002
In North America, WNS is known to affect 6 species of bats that use hibernation as their winter survival strategy: the big brown bat (Eptesicus fuscus), the eastern small-footed bat (Myotis leibii), the little brown bat (M. lucifugus), the northern long-eared bat (M. septentrionalis), the tricolored bat (Perimyotis subflavus), and the Indiana bat (M. sodalis) (Blehert et al., 2009; Courtin et al., 2010; Turner and Reeder, 2009). Since its detection in February 2006 in a popular tourist cave near Albany, New York, USA, WNS has spread >1,300 km into Connecticut, Massachusetts, New Hampshire, New Jersey, Pennsylvania, Tennessee, Vermont, Virginia, and West Virginia in the United States and the provinces of Ontario and Quebec in Canada (Blehert et al., 2009, Turner and Reeder, 2009; United States Geological Survey, 2010) in a pattern suggesting the spread of an infectious agent.
A recently discovered psychrophilic (cold-loving) fungus, Geomyces destructans (Gargas et al., 2009), has consistently been isolated from bats that meet the pathologic criteria for WNS, including colonization of skin by fungal hyphae causing characteristic epidermal erosions and ulcers that can progress to invasion of underlying connective tissue (Meteyer, et al.,2009; Reichard and Kunz, 2009). G. destructans is identified by its distinctive asymmetrically curved conidia and has a unique taxonomic position among other Geomyces spp. described to date (Gargas et al., 2009) Its closest genetic relative is G. pannorum, a ubiquitous psychrophilic, keratinolytic fungus that has been isolated from a variety of sources and geographic regions, including soil and the fur of wild mammals in France (Chabasse et al., 1987), floors of trains and ferryboats in Italy (Mercantini et al., 1989), boreal forests in Canada (de Bellis et al., 2007), and environmental samples from Arctic regions (Kochkina et al., 2007; Mercantini et al., 1989). G. pannorum var. pannorum has been reported as an unusual dermatophyte infecting fingernails and superficial skin of humans who have a history of close contact with soil and dust (Gianni et al., 2003, Zelenková, 2006). However, G. destructans differs from other common soil fungi of North America in its ability to invade the living tissues of hibernating bats.
After WNS was described in North America (Blehert et al., 2009), reports dating back to the early 1980s (Feldmann, 1984) described repeated observations of white fungal growth on muzzles of hibernating bats in Germany. However, these bats lacked the characteristics of WNS such as associated deaths. Moreover, fungus was not identified. In response to WNS in North America, researchers in Europe initiated a surveillance effort during the winter of 2008–09 for WNS-like fungal infections among hibernating populations of bats in Europe. G. destructans in Europe was previously reported in 1 hibernating bat that was sampled in France during March 2009 (Puechmaille et al., 2010).
In this report, we describe results of a more extensive effort by scientists from 4 countries in Europe (Germany, United Kingdom, Hungary, and Switzerland) to obtain and analyze samples from hibernating bats with white patches on their faces or wing membranes. Our objectives were to identify the fungus
colonizing such affected hibernating bats in Europe and to clarify its geographic distribution over a broad area of Europe.
Materials and Methods
During ongoing annual population surveys of caves and mines conducted by national nongovernmental organizations, hibernating bats with obvious fungal growth on their bodies (Figure A20-1, panel A) were opportunistically sampled in Germany, Switzerland, and Hungary; samples were also obtained from 2 dead bats from the same hibernaculum in the United Kingdom. Approximately 366 hibernacula were visited during mid-February–mid-April 2009: 336 in Germany, 20 in Hungary, and 10 in Switzerland. Two to 214 hibernating animals were observed at each site, with the exception of 2 sites in Germany, which harbored 2,000–7,000 animals at each site.
Samples were collected from live bats by using 2 methods. Touch imprints were obtained by holding adhesive tape against affected areas of skin or fur, or fur clippings were obtained from affected areas of bat muzzles. All species of bats in Europe are strictly protected under the Flora, Fauna, Habitat Guidelines of the European Union (92/43/ EEC) (http://ec.europa.eu/environment/nature/legislation/ habitatsdirective/index_en.htm) and The Agreement on the Conservation of Populations of European Bats (www.eurobats.org). We did not have permission to invasively sample or kill individual animals for histologic analysis to confirm skin infection by G. destructans (Meteyer et al., 2009). Samples were shipped to the Leibniz Institute of Zoo and Wildlife Research (IZW), Berlin, Germany, for further investigations.
Twenty adhesive tape samples were first screened by using light microscopy, and 21 hair samples were examined by using scanning electron microscopy for conidia characteristic of G. destructans (Figure A20-1, panel B). Two of the submitted samples (2 greater horseshoe bats from the United Kingdom) consisted of entire bat carcasses. Although the carcasses were examined externally for fungal growth on muzzle skin and hair, specimens were too decomposed to conduct internal pathologic examinations. Tape or hair samples from all bats were further investigated by using direct PCR amplification of fungal rRNA gene internal transcribed spacer (ITS) region DNA (ITS1, 5.8S, and ITS2). Total nucleic acids were extracted from culture, tape, or hair samples by using PrepMan Ultra Reagent (Applied Biosystems, Darmstadt, Germany) following the manufacturer’s instructions.
The rRNA gene ITS region DNA was amplified by using PCR with primers ITS4 and ITS5 (White et al., 1990) and GoTaq DNA polymerase (Promega, Madison, WI, USA). Cycling parameters were an initial 2-min denaturation at 98°C; followed by 30 cycles of denaturation at 98°C for 10 s, annealing at 50°C for 30 s, and extension at 72°C for 1 min; and a final extension at 72°C for 7 min. For fungal isolates, rRNA gene small subunit (SSU) DNA was amplified by us-
ing PCR with primers nu-SSU-0021–5′ (White et al., 1990) and nu-SSU-1750–3′ (Gargas and Taylor, 1992) as above, except the extension time was increased to 2 min. Sequencing primers were PCR primers; nu-SSU-0402–5′ (Gargas and Taylor, 1992), nu-SSU-1150–5′ (White et al., 1990), nu-SSU-0497–3′ (Gargas and Taylor, 1992), and nu-SSU-1184–3′ (Gargas et al., 1995) were added for SSU. PCR products were sent to the Robert Koch Institute, Berlin, Germany, for direct sequencing.
Culture analyses of samples were performed at Munich University Hospital and IZW. After examining tape impressions by using light microscopy, we identified small areas with fungal conidia characteristic of G. destructans and excised them with a sterile scalpel blade. Half of the excised material was used for PCR; the remaining sample and samples of individual hairs with microscopic indication of G. destructans were immediately placed onto Sabouraud dextrose agar plates containing gentamicin and chloramphenicol and incubated at 4°C and 8°C. G. destructans isolates obtained during this study are maintained at IZW.
Results
We obtained and analyzed samples from live bats with obvious fungal growth on their bodies found between mid-February and the end of March 2009 at 11 sites (8 in Germany, 1 in Hungary, and 2 in Switzerland). Samples were also obtained from an additional bat in Germany in February 2008 and from 2 dead bats from a site in the United Kingdom in March 2009 (Tables A20-1, A20-2) All 12 hibernacula sampled contained 1–5 animals that exhibited obvious fungal growth. Forty-three samples were obtained from these 12 hibernacula and represented 23 adult bats of 6 species: 1 Brandt bat (M. brandtii), 3 pond bats (M. dasycneme), 1 Daubenton bat (M. daubentonii), 1 lesser mouse-eared bat (M. oxygnathus), 15 greater mouse-eared bats (M. myotis), and 2 greater horseshoe bats (Rhinolophus ferrumequinum).
After direct PCR amplification and DNA sequence analysis of fungal rRNA gene ITS regions, genetic signatures 100% identical with those from G. destructans type isolate NWHC 20631–21 (GenBank accession no. EU884921) were identified from 21 of 23 bats examined: 15/15 from Germany, 2/2 from Hungary, and 4/4 from Switzerland. Both bats from the United Kingdom were colonized by Penicillium sp. (Tables A20-1, A20-2). Fungi with conidia morphologically identical to those of G. destructans (Figure A20-1, panel B) as described by Gargas et al. (2009) were isolated in axenic cultures from 8 of 23 bats examined: 3/15 from Germany, 1/2 from Hungary, and 4/4 from Switzerland) (Tables A20-1, A20-2; Figure A20-2).
Consistent with published descriptions for G. destructans (Gargas et al., 2009), fungal colonies grew slowly and within 14 days attained diameters of 1.0 mm at 4°C and 4.0–5.0 mm at 8°C; no growth occurred at 25°C. The sensitivity of our method for isolating G. destructans from bat hair was comparable to
Table A20-1 Bats Tested for Geomyces destructans by Using Microscopy, Fungal Culture, or PCR Analysis, by Country, Europe*
| No. positive/no. tested | ||||||
| Species (common name) | Germany Switzerland | Hungary | United Kingdom | |||
| Myotis myotis (greater mouse-eared bat) | 10/10 4/4 | 1/1 | - | |||
| M. dasycneme (pond bat) | 3/3 - | - | - | |||
| M. daubentonii (Daubenton bat) | 1/1 - | - | - | |||
| M. brandtii (Brandt bat) | 1/1 - | - | - | |||
| M. oxygnathus (lesser mouse-eared bat) | - - | 1/1 | - | |||
|
Rhinolophus ferrumequinum (greater horseshoe bat) |
- - | - | 0/2 | |||
*-, species not obtained in this country.
published diagnostic sensitivity for culturing G. destructans from bat skin (Lorch et al., 2010). Subsequent PCR/DNA sequencing analyses of the 8 isolates indicated that they all had rRNA gene ITS and SSU region DNA sequences identical to those of G. destructans type isolate NWHC 20631–21 (GenBank accession nos. EU884921 for ITS and FJ231098 for SSU).
Unlike other bats sampled in this study, the 2 greater horseshoe bats from the United Kingdom were found dead, and their nostrils were colonized by Penicillium sp. These bats did not fulfill the pathologic criteria for WNS (Meteyer, et al., 2009) because fungal hyphae did not invade the epidermis but remained within the superficial layer of the epidermal stratum corneum. A more complete description of the postmortem analysis of the greater horseshoe bats has been reported (Barlow et al., 2009). G. destructans was not isolated in culture, and its genetic signature was not identified by PCR and DNA sequencing of samples collected from greater horseshoe bats.
Discussion
Laboratory analyses demonstrated that 5 species of the genus Myotis in Europe harbored G. destructans; male and female bats were equally affected. Despite laboratory confirmation that bats obtained in Germany, Switzerland, and Hungary were colonized by G. destructans, deaths were not observed at collection sites. Puechmaille et al. (2010) reported a similar observation with a greater mouse-eared bat in France. Additionally, a lesser mouse-eared bat from Hungary with visible fungal infection during hibernation, from which G. destructans was isolated, was recaptured 5 months later (August 2009) and showed no external signs of fungal infection. On February 19, 2010, the same bat was again observed in the same hibernaculum without any visible sign of fungal growth. However, 7 other bats within that group of 55 animals displayed obvious fungal growth but were not sampled for this study.
Table A20-2 Fungal Culture and PCR Results for 23 Bats with Evidence of Fungal Colonization Tested by Light or Electron Microscopy, Europe*
| GenBank accession no. | ||||||||
| Country/location no.† | Sample source | Species | Collection date | No./hibernacula | PCR result | Culture result | ITS | SSU rRNA |
| Germany/4 | Hair 2 | Myotis dasycneme | 2008 Feb 25 | 10 | + | + | GU350437 | GU350442 |
| Germany/8 | Hair 7 | M. myotis | 2009 Mar 3 | 214 | + | + | GU350436 | GU350441 |
| Germany/7 | Tape 8 | M. myotis | 2009 Mar 7 | 57 | + | + | GU999986 | GU999983 |
| Hungary/9 | Hair 16 | M. myotis | 2009 Mar 29 | 64 | + | + | GU350434 | GU350439 |
| Switzerland/10 | Tape 10 | M. myotis | 2009 Apr 5 | 25 | + | + | GU350433 | GU350438 |
| Switzerland/10 | Tape 11 | M. myotis | 2009 Apr 5 | 25 | + | + | GU999984 | GU999981 |
| Switzerland/11 | Tape 12 | M. myotis | 2009 Apr 5 | 25 | + | + | GU999985 | GU999982 |
| Switzerland/10 | Tape 20 | M. myotis | 2009 Apr 11 | 25 | + | + | GU350435 | GU350440 |
| Germany/1 | Hair 1 | M. myotis | 2009 Feb 21 | 65 | + | – | HM222616 | – |
| Germany/6 | Hair 20 | M. myotis | 2009 Mar 13 | 100 | + | – | HM222617 | – |
| Germany/2 | Tape 1 | M. myotis | 2009 Feb 26 | ≈2,000 | + | – | HM222618 | – |
| Germany/2 | Tape 2 | M. myotis | 2009 Feb 26 | ≈2,000 | + | – | HM222619 | – |
| Germany/8 | Tape 5 | M. myotis | 2009 Mar 3 | 214 | + | – | HM222620 | – |
| Germany/8 | Tape 6 | M. myotis | 2009 Mar 3 | 214 | + | – | HM222621 | – |
| Germany/7 | Tape 9 | M. myotis | 2009 Mar 7 | 57 | + | – | HM222622 | – |
| Germany/6 | Tape 16 | M. myotis | 2009 Mar 13 | 100 | + | – | HM222623 | – |
| Germany/8 | Hair 6 | M. brandtii | 2009 Mar 3 | 214 | + | – | HM222624 | – |
| Germany/5 | Hair 3 | M. dasycneme | 2009 Feb 28 | 29 | + | – | HM222625 | – |
| Germany/6 | Tape 17 | M. dasycneme | 2009 Mar 13 | 100 | + | – | HM222626 | – |
| Germany/3 | Hair 17 | M. daubentonii | 2009 Mar 5 | ≈7,000 | + | – | HM222627 | – |
| Hungary/9 | Tape 13 | M. oxygnathus | 2009 Mar 29 | 64 | + | – | HM222628 | – |
| United Kingdom/12 | Hair 10 | Rhinolophus ferrumequinum | 2009 Mar 11 | 558 | –‡ | –‡ | HM222629 | – |
| United Kingdom/12 | Hair 11 | R. ferrumequinum | 2009 Mar 11 | 558 | –‡ | –‡ | HM222630 | – |
*ITS, internal transcribed spacer; SSU, small subunit; tape, touch imprint with adhesive tape.
†Location number corresponds to a hibernation site in Figure A20-2.
‡Although samples were negative for G. destructans, they were positive for Penicillium sp. by PCR and culture.
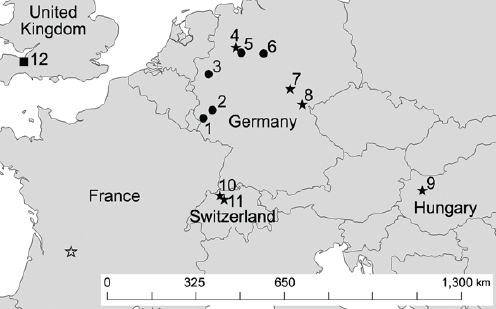
Figure A20-2 Locations in Europe of bats positive for Geomyces destructans by PCR alone (circles) or by PCR and culture (solid stars) and bats negative for G. destructans but positive for other fungi (square). Numbers for locations correspond to those in Table A20-2. Sites 7, 8, and 9 had additional bats that were positive for G. destructans only by PCR. Location of a bat positive for G. destructans in France (Puechmaille et al., 2010) is indicated by an open star. Some sites had >1 bat species with evidence of colonization by G. destructans.
In contrast, decreases in hibernating bat colonies infected by G. destructans in North America are often >90% (Reichard and Kunz, 2009; Turner and Reeder, 2009), and mortality rates similar in magnitude would be difficult to miss among closely monitored winter populations of bats in Europe. Biologists in Germany and Switzerland have conducted annual censuses of bat hibernacula since the 1930s and 1950s, respectively. In Hungary, the largest hibernacula have been annually monitored since1990. Similar death rates to those caused by WNS in hibernating bats in North America have never been documented in countries in Europe in which G. destructans has now been identified.
Although distribution of G. destructans in bats across Europe has not been exhaustively characterized, opportunistic sampling conducted as part of this study during the winter of 2008–09 demonstrated that the fungus was present on bats in 3 countries (Figure A20-2). The 2 most distant points from which bats colonized with G. destructans have been identified were separated by >1,300 km. Despite the observed distribution of G. destructans in Europe (Figure A20-2), the 5 bat species from which G. destructans was detected migrate average distances <100 km between their summer and winter roosting sites (Hutterer et al., 2005),
indicating that the fungus is most likely spread as local bat populations emerge from hibernacula, disperse, and interact with populations within their dispersal range. Identification of bats colonized by G. destructans from such distant sites, in addition to the relatively homogenous distribution of the fungus among sites in Germany, suggests that G. destructans may be widespread in Europe.
Regardless of widespread occurrence of G. destructans among bat species in Europe (Figure A20-2), deaths of bats in Europe caused by WNS, similar to those caused by WNS in North America, have not been observed. Although no bat species migrates between Europe and North America or is present on both continents (Dietz et al., 2009; Nowak, 1999), many species of the genus Myotis are infected by G. destructans on each continent. Although the mechanism(s) by which hibernating bats died because of infection with G. destructans in North America is not yet understood, bat species in Europe may exhibit greater resistance or respond differently to infection by this fungus than their counterparts in North America.
Before the emergence of WNS in North America, large aggregations of hibernating bats ranging from 1,000 to 50,000 animals were common in caves and mines of affected regions, and many hibernation sites in regions of North America still unaffected by WNS contained tens of thousands of bats during winter (some contain hundreds of thousands) (Barbour and Davis, 1969). In contrast, aggregations of bats hibernating in caves and mines in Europe rarely exceed 1,000 animals. However, larger hibernating groups have been observed at a few natural sites, such as a cave in northern Germany with 13,000–18,000 bats (Petermann and Boye, 2006) and human-made structures (e.g., Daubenton bats in bunkers and catacombs) (Dietz et al., 2009). If host density plays a role in G. destructans transmission or deaths of bats, such as through increased disturbance of clustered bats, the bats in Europe may experience lower mortality rates because they form smaller hibernation groups composed of small clusters or individual bats. Apparent continental differences in susceptibility of hibernating bats to deaths associated with skin infection by G. destructans may indicate either circumstantial or evolved resistance in bats in Europe.
G. destructans has been detected in North America only in states and provinces where WNS has also been observed and in contiguous states. Recent emergence and spread of G. destructans with associated deaths of bats throughout hibernacula in the northeastern United States (Turner and Reeder, 2009) may suggest ecologic release of an exotic pathogen into an uninfected ecosystem. Although this suggestion remains a hypothesis and how G. destructans may have been introduced to the United States is not known, initial documentation of WNS in a popular tourist cave near Albany, New York (Blehert et al., 2009), suggests that a human vector could have been involved.
There are many examples of unintended introductions of fungal pathogens, particularly of those affecting plants and ectothermic animals with tissue temperatures permissive to fungal infection (Casadevall, 2005; Desprez-Loustau et al.,
2007; Robert and Casadevall, 2009). One case with striking similarities is the panzootic chytrid fungus (Batrachochytrium dendrobatidis), which has caused global decreases among amphibian species (Fisher et al., 2009). As with skin infection by B. dendrobatidis in amphibians, which can alter body electrolyte levels and lead to cardiac arrest (Voyles et al., 2009), skin infection by G. destructans in hibernating bats may also kill by causing irreversible homeostatic imbalance because wing membranes play major roles in water balance, circulation, and thermoregulation of hibernating bats during winter (Davis, 1970; Makanya and Mortola, 2007).
Bat species in Europe may be immunologically or behaviorally resistant to G. destructans because of having coevolved with the fungus. Additionally, microbial flora of bat skin or other abiotic surfaces in bat hibernacula in Europe may have also coevolved to incorporate G. destructans as a nonpathogenic component of the microbial community. Conversely, possible recent introduction of G. destructans into the United States, with subsequent infection of bat species in North America and ecosystems not infected with the fungus, provides a potential explanation for the devastating effects of WNS in North America. Although bats are reservoirs of various pathogens (Calisher et al., 2006; Wibbelt et al., 2009), research into the immune function of bats, particularly during hibernation, is just beginning.
In conclusion, nondetrimental colonization of bat species in Europe by G. destructans may be relatively common (Figure A20-2), and historical reports (Feldmann, 1984) suggest that such colonization of hibernating bats in Europe has occurred for several decades. In contrast to recent mass deaths associated with G. destructans skin infection, which is the hallmark of WNS in North America, bats in Europe appear to coexist with G. destructans. Studies to investigate mechanisms of pathogenesis, microbial ecology, and phylogeography of G. destructans will be essential for developing a comprehensive understanding of WNS. In particular, testing the hypotheses that bats in Europe are more resistant to fungal skin infection by G. destructans, that G. destructans was introduced from Europe to North America, and that environmental circumstances limit the pathogenicity of G. destructans in Europe seem warranted. Divergent manifestations of infection by G. destructans in bats in Europe and North America provide a unique opportunity to address these research objectives with the ultimate goals of better understanding WNS and developing sound strategies to manage the devastating effects of this emerging wildlife disease in North America.
Acknowledgments
We thank N. Jahn, D. Viertel, A. Kus, and C. Kohl for excellent technical assistance; A. Beck, T. Filip, M. Franz, S. Gloor, R. Güttinger, A. Kiefer, V. Korn, G. Mäscher, B. Máté, C.D. Morawitz, C. Morris, P. Paulovics, M. Piskorski, F. Putzmann, W. Schorcht, and C. Tress for help surveying sites and retrieving sam-
ples; A. Gargas, K. Schuler, and 2 anonymous reviewers for providing thoughtful comments on earlier drafts of the manuscript; and M. Riccucci for help with the literature search for previous reports of fungi in bats in Europe.
This study was supported by Bat Conservation Switzerland.
Dr Wibbelt is a senior veterinary pathologist at the Leibniz Institute for Zoo and Wildlife Research, Berlin, Germany. Her research interests include infectious diseases in wild animals, particularly bats.
References
Barbour RW, Davis WH. Bats of America. Lexington (KY): The University Press of Kentucky; 1969.
Barlow A, Ford S, Green R, Morris C, Reaney S. Investigations into suspected white-nose syndrome in two bats in Somerset. Vet Rec. 2009;165:481–2.
Blehert DS, Hicks AC, Behr M, Meteyer CU, Berlowski-Zier BM, Buckles EL, et al. Bat white-nose syndrome: an emerging fungal pathogen? Science. 2009;323:227. DOI: 10.1126/science.1163874
Calisher CH, Childs JE, Field HE, Holmes KV, Schountz T. Bats: an important reservoir host of emerging viruses. Clin Microbiol Rev. 2006;19:531–45. PubMed DOI: 10.1128/CMR.00017-06
Casadevall A. Fungal virulence, vertebrate endothermy, and dinosaur extinction: is there a connection? Fungal Genet Biol. 2005;42:98–106. DOI: 10.1016/j.fgb.2004.11.008
Chabasse D, Guiguen C. Couatarmanac’h A, Launay H, Reecht V, de Bièvre C. Keratinophilic fungal flora isolated from small wild mammals and rabbit-warren in France. Discussion on the fungal species found [in French]. Ann Parasitol Hum Comp. 1987;62:357–68.
Courtin F, Stone W, Risatti G, Gilbert K, Van Kruiningen H. Pathologic findings and liver elements in hibernating bats with white-nose syndrome. Vet Pathol. 2010;47:214–9. DOI: 10.1177/0300985809358614
Davis WH. Hibernation: ecology and physiological ecology. In: Wimsatt WA, editor. Biology of bats. Vol. 1. New York: Academic Press; 1970. p. 265–300.
de Bellis T, Kernaghan G, Widden P. Plant community influences on soil microfungal assemblages in boreal mixed-wood forests. Mycologia. 2007;99:356–67. DOI: 10.3852/mycologia.99.3.356
Desprez-Loustau ML, Robin C, Buée M, Courtecuisse R, Garbaye J, Suffert F, et al. The fungal dimension of biological invasions. Trends Ecol Evol. 2007;22:472–80. DOI: 10.1016/j. tree.2007.04.005
Dietz C, von Helversen O, Nill D. Bats of Britain, Europe and North-west Africa. London: A and C Black Publishers; 2009.
Feldmann R. Teichfledermaus–Myotis dasycneme (Boie, 1825). Die Säugetiere Westfalens. Münster: Westfälisches Museum für Naturkunde; 1984. pp. 107–11.
Fisher MC, Garner TW, Walker SF. Global emergence of Batra-chochytrium dendrobatidis and amphibian chytridiomycosis in space, time, and host. Annu Rev Microbiol. 2009;63:291–310. DOI: 10.1146/annurev.micro.091208.073435
Gargas A, dePriest P, Taylor J. Positions of multiple insertions in SSU rDNA of lichen forming fungi. Mol Biol Evol. 1995;12:208–18.
Gargas A, Taylor J. Polymerase chain reaction (PCR) primers for amplifying and sequencing 18S rDNA from lichenized fungi. Mycologia. 1992;84:589–92. DOI: 10.2307/3760327
Gargas A, Trest MT, Christensen M, Volk TJ, Blehert DS. Geomyces destructans sp. nov. associated with bat white-nose syndrome. Mycotaxon. 2009;108:147–54.
Gianni C, Caretta G, Romano C. Skin infection due to Geomyces pannorum var. pannorum. Mycoses. 2003;46:430–2. DOI: 10.1046/ j.1439-0507.2003.00897.x
Hutterer R, Ivanova T, Meyer-Cords C, Rodrigues L. Bat migrations in Europe: a review of banding data and literature. Bonn (Germany): German Agency for Nature Conservation; 2005.
Kochkina GA, Ivanushkina NE, Akimov VN, Gilichinskii DA, Ozerskaia SM. Halo- and psychrotolerant Geomyces fungi from arctic cryopegs and marine deposits [in Russian]. Mikrobiologiia. 2007;76:39–44.
Lorch JM, Gargas A, Meteyer CU, Berlowski-Zier BM, Green DE, Shearn-Bochsler V, et al. Rapid polymerase chain reaction diagnosis of white-nose syndrome in bats. J Vet Diagn Invest. 2010;22: 224–30.
Makanya AN, Mortola JP. The structural design of the bat wing web and its possible role in gas exchange. J Anat. 2007;211:687–697. PubMed DOI: 10.1111/j.1469-7580.2007.00817.x
Mercantini R, Marsella R, Cervellati M. Keratinophilic fungi isolated from Antarctic soil. Mycopathologia. 1989;106:47–52. DOI: 10.1007/BF00436926
Mercantini R, Marsella R, Prignano G, Moretto D, Marmo W, Leonetto F, et al. Isolation of keratinophilic fungi from the dust of ferry boats and trains in Italy. Mycoses. 1989;32:590–4.
Meteyer CU, Buckles EL, Blehert DS, Hicks AC, Green DE, Shearn-Bochsler V, et al. Histopathologic criteria to confirm white-nose syndrome in bats. J Vet Diagn Invest. 2009;21:411–4.
Nowak R. Walker’s mammals of the world. Baltimore: The Johns Hopkins University Press; 1999.
Petermann R, Boye P. National report on bat conservation in the Federal Republic of Germany 2003–2006. Bonn (Germany): Eurobats; 2006.
Puechmaille SJ, Verdeyroux P, Fuller H, Ar Gouilh M, Bekaert M, Teeling EC. White-nose syndrome fungus (Geomyces destructans) in bat, France. Emerg Infect Dis. 2010;16:290–3.
Reichard JD, Kunz TH. White-nose syndrome inflicts lasting injuries to the wings of little brown myotis (Myotis lucifugus). Acta Chiropterologica. 2009;11:457–64. DOI: 10.3161/150811009X485684
Robert VA, Casadevall A. Vertebrate endothermy restricts most fungi as potential pathogens. J Infect Dis. 2009;200:1623–6. DOI: 10.1086/644642
Turner GR, Reeder DM. Update of white-nose syndrome in bats, September 2009. Bat Research News. 2009;50:47–53.
United States Geological Survey. Update on white-nose syndrome: Tennessee finding. USGS wildlife health bulletin. Reston (VA): The Survey; 2010 [cited 2010 May 19]. http://www.nwhc.usgs.gov/ disease_information/white-nose_syndrome/
Voyles J, Young S, Berger L, Campbell C, Voyles WF, Dinudom A, et al. Pathogenesis of chytridiomycosis, a cause of catastrophic amphibian declines. Science. 2009;326:582–5. DOI: 10.1126/science.1176765
White T, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TH, editors. PCR protocols: a guide to methods and applications. San Diego (CA): Academic Press; 1990. p. 315–22.
Wibbelt G, Speck S, Field H. Methods for assessing diseases in bats. In: Kunz T, Parsons S, editors. Ecological and behavioral methods for the study of bats. Baltimore: Johns Hopkins University Press; 2009. p. 775–94.
Zelenková H. Geomyces pannorum as a possible causative agent of dermatomycosis and onychomycosis in two patients. Acta Dermatovenerol Croat. 2006;14:21–5
PAN-EUROPEAN DISTRIBUTION OF WHITE-NOSE SYNDROME FUNGUS (GEOMYCES DESTRUCTANS) NOT ASSOCIATED WITH MASS MORTALITY*
Sébastien J. Puechmaille,63,64,65,66Gudrun Wibbelt,66,67 Vanessa Korn,68
Hubert Fuller,63Frédéric Forget,69Kristin Mühldorfer,67Andreas Kurth,70
Wieslaw Bogdanowicz,71Christophe Borel,72Thijs Bosch,73Thomas Cherezy,74
Mikhail Drebet,75Támás Görföl,76Anne-Jifke Haarsma,77Frank Herhaus,78
Guénael Hallart,79Matthias Hammer,80Christian Jungmann,81Yann Le Bris,82
Lauri Lutsar,83Matti Masing,84Bart Mulkens,85Karsten Passior,86
Martin Starrach,87Andrzej Wojtaszewski,88Ulrich Zöphel,89and
Emma C. Teeling63,64
______________
63 School of Biology and Environmental Science, University College Dublin, Dublin, Ireland
64 Conway Institute of Biomolecular and Biomedical Research, Dublin, Ireland
65 Email: s.puechmaille@gmail.com
66 These authors contributed equally to this work.
67 Leibniz Institute for Zoo and Wildlife Research, Berlin, Germany
68 Office for Faunistic and Landscape Ecology, Schöneberg, Germany
69 Plecotus Working Group,Association Natagora, Brussels, Belgium
70 Robert Koch Institute, Berlin, Germany
71 Museum and Institute of Zoology, Polish Academy of Sciences, Warszawa, Poland
72 Commission de Protection des Eaux, du Patrimoine, de l’Environnement, du Sous-sol et des Chiroptères—Lorraine, Velaine-en-Haye, France
73 Dutch Bat Workers Group,Nijmegen, The Netherlands
74 Coordination Mammalogique du Nord de la France, Béthune, France
75 Podilski Tovtry National Nature Park, Kamenets-Podilsky, Ukraine
76 Nature Conservation Foundation of Tolna County, Szekszárd, Hungary
77 Centre for Ecosystem Studies, Alterra and Wageningen University, Wageningen, The Netherlands
78 Biology Station Oberberg, Nümbrecht, Germany
79 Société d’Etude et de Protection de la Nature en Thiérache, Le Chaudron, Origny-en-Thiérache, France
80 Department of Biology, Center for Bat Conservation in Northern Bavaria, Erlangen University, Erlangen, Germany
81 Nature and Biodiversity Conservation Union Rhineland-Palatine, Birkenfeld, Germany
82 Bretagne Vivante SEPNB, Roussimel, Glénac, France
83 Estonian Fund for Nature, Tartu, Estonia
84 Sicista Development Centre, Tartu, Estonia
85 Bat Working Group, Natuurpunt VZW, Belgium
86 Nature and Biodiversity Conservation Union Southern Lower-Saxony,Nordstemmen, Germany
87 Biotope Mapping Cooperation, Herford, Germany
88 Institute of Natural Fibres and Medicinal Plants, Poznan, Poland
89 Saxonian State Office for Environment Agriculture and Geology, Dresden-Pillnitz, Germany
*Originally printed as: Sébastien Peuchmaille, Gudrun Wibbelt, Vaness Korn, Hubert Fuller, Frédéric Forget, et al. (2011) Pan-European Distribution of White-Nose Syndrome Fungus (Geomyces destructans) Not Associated with Mass Mortality. PLoS ONE. 6(4): e19167. doi:10.1371/journal. pone.0019167.
Abstract
Background:
The dramatic mass mortalities amongst hibernating bats in Northeastern America caused by ‘‘white nose syndrome’’(WNS) continue to threaten populations of different bat species. The cold-loving fungus, Geomyces destructans, is the most likely causative agent leading to extensive destruction of the skin, particularly the wing membranes. Recent investigations in Europe confirmed the presence of the fungus G. destructans without associated mass mortality in hibernating bats in six countries but its distribution remains poorly known.
Methodology/Principal Findings:
We collected data on the presence of bats with white fungal growth in 12 countries in Europe between 2003 and 2010 and conducted morphological and genetic analysis to confirm the identity of the fungus as Geomyces destructans. Our results demonstrate the presence of the fungus in eight countries spanning over 2000 km from West to East and provide compelling photographic evidence for its presence in another four countries including Romania, and Turkey. Furthermore, matching prevalence data of a hibernaculum monitored over two consecutive years with data from across Europe show that the temporal occurrence of the fungus, which first becomes visible around February, peaks in March but can still be seen in some torpid bats in May or June, is strikingly similar throughout Europe. Finally, we isolated and cultured G. destructans from a cave wall adjacent to a bat with fungal growth.
Conclusions/Significance:
G. destructans is widely found over large areas of the European continent without associated mass mortalities in bats, suggesting that the fungus is native to Europe. The characterisation of the temporal variation in G. destructans growth on bats provides reference data for studying the spatio-temporal dynamic of the fungus. Finally, the presence of G. destructans spores on cave walls suggests that hibernacula could act as passive vectors and/or reservoirs for G. destructans and therefore, might play an important role in the transmission process.
Introduction
White nose-syndrome (WNS) is a devastating disease causing mass mortalities in hibernating bats in North-America. In May 2009, it was estimated that over one million bats had died from the disease which was first documented in February 2006 at Howe’s Cave, West of Albany, New York (Anonymous, 2009). A visually conspicuous white fungus grows on the face, ears, or wings of
stricken bats with hyphae penetrating deep into the connective tissue of glabrous skin and snout (Meteyer et al., 2009) and causing severe damage (Reichard and Kunz, 2009). The fungus associated with WNS is a newly described, psychrophilic (cold-loving) species (Geomyces destructans) (Gargas et al., 2009), closely related to other psychrophilic species of Geomyces (Rice and Currah, 2006; Puechmaille et al., 2010). Although it is not yet conclusively proven whether G. destructans is the causative agent of the disease or if other co-factors are necessary for disease to occur, the fungus is always found on bats at WNS sites where hibernating bats experience mass mortalities (Blehert et al., 2009). To date, bacteriological, virological, parasitological and pathological evaluations as well as studies of toxic contaminants have not identified the consistent presence of any other agents/cause of death. The lack of evidence for the involvement of other agents or compounds reinforces the suspicion that G. destructans is the causative agent of WNS mortality (Meteyer et al., 2009; Blehert et al., 2009; Kannan et al., 2010; Courtin et al., 2010).
Geomyces destructans has been found in nine species of bats in North-America, from the provinces of Ontario and Quebec in Canada south and west to the states of Tennessee and Oklahoma in the USA (Anonymous, 2010). Three recent studies investigating samples collected in 2008–2010 have shown that G. destructans was also present in six European countries (France, Germany, Switzerland, Czech Republic, Slovakia & Hungary) (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010). Nevertheless, the geographic coverage of these studies was limited and the extent of the distribution of G. destructans in Europe remains poorly known. In this paper, we combine previously published data on the distribution of G. destructans in Europe (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010) with new data from twelve countries covering 2,400 km from West to East (France to Turkey) and 1,900 km from North to South (Estonia to Turkey) to demonstrate the widespread presence of G. destructans on multiple species of hibernating bats in Europe without associated mass mortality.
Results
Review of data on Geomyces destructans in European bats, 2008–2010
Although photographs of bats with fungal growth similar to G. destructans were published in Germany in the 1980’s (Feldmann, 1825), and also taken in the 1990’s in the Czech Republic (Martínková et al., 2010), there have been no confirmed records of G. destructans in Europe prior to 2008 (Puechmaille et al., 2010; Wibbelt et al., 2010). In 2010, G. destructans has been confirmed by morphological and genetic analyses from samples collected during the winters 2007/2008, 2008/2009 and 2009/2010 in six European countries (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010). In France, Hungary,
Switzerland and Slovakia, the fungus has been confirmed from 1–2 location(s) per country, whereas it has been confirmed at 8 sites in Germany and 23 sites in the Czech Republic (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010). All confirmed detections of G. destructans in Europe have been made by isolating and/or genetically identifying the fungus from hairs, swabs or touch imprints from bats (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010). In Europe, eight species of Myotis have been observed being colonised by G. destructans: M. Myotis, M. blythii (referred to as M. oxygnathus in [Wibbelt et al., 2010]), M. mystacinus, M. daubentonii, M. dasycneme, M. nattereri, M. bechsteinii and M. brandtii. Species from other bat families were present in the caves with infected individuals (e.g., Miniopteridae: Miniopterus schreibersii; Rhinolophidae: Rhinolophus hipposideros and R. ferrumequinum),but no G. destructans has been confirmed from these species. Previous extensive surveys of cave fungi in Europe (i.e., Nováková, 2009; Mosca and Campanino, 1962; Bastian et al., 2009) or fungi associated with insects hibernating in underground sites (Kubátová and Dvořák, 2005) never reported G. destructans in their inventory, although some other species of Geomyces were recovered (Nováková, 2009; Mosca and Campanino, 1962; Bastian et al., 2009).
New data on G. destructans in Europe 2003–2010
During winter hibernation counts, a total of 107 bats from 56 sites in twelve European countries were reported to have visible white fungal growth (Tables A21-1, A21-2, A21-3 and Figure A21-1). This represents the first records from nine countries (Austria, Belgium, Denmark, Estonia, The Netherlands, Poland, Romania, Turkey and Ukraine). One hundred and five bats were alive and two of them were found dead in hibernacula. These 107 bats belonged to eight different species of Myotis: M. Myotis (59), M. dasycneme (26), M. mystacinus (9), M. daubentonii (4), M. Myotis/blythii (3), M. blythii (3), M. nattereri (1), M. escalerai/sp. A (1) and M. brandtii (1). Of these, molecular and morphological identifications of the colonising fungus were carried out in 23 cases (Table A21-1), while only photographic evidence was obtained for a further 50 cases (Table A21-2 and Figure A21-2). The remaining 34 cases were based on reports of visual observations of a white fungal growth on bat snouts and/or ears (Table A21-3), which was very similar to pictures presented in Figure A21-2. All 84 bats reported in Tables A21-2 and A21-3 are considered as Gd-suspects (bats showing fungal growth that is thought to be G. destructans).
Geomyces destructans identification
Out of a total of 107 bats with fungal growth, 22 were sampled, 16 with touch imprints and 6 with cotton swabs. The 22 bats sampled (20 alive and 2 dead) belonged to the species of Myotis from which G. destructans had been pre-
| Country | Lat | Lon | Date | Host species | Culture | PCR | GenBank No. |
| France* | 49.9 | 4.1 | 04/03/2010 | Myotis mystacinus | GuH-04032010 | -‡ | n/a |
| France* | 50.6 | 2.5 | 22/02/2010 | Myotis nattereri | ThC-22022010 | -‡ | n/a |
| France† | 47.7 | –2.1 | 04/03/2010 | Myotis myotis | Mmyo-FR-1 | + | JF502415 |
| Belgium | 49.8 | 5.3 | 03/04/2010 | Myotis myotis | Mmyo-BE-1 | + | JF502414 |
| Belgium† | 50.8 | 5.6 | 18/03/2010 | Myotis mystacinus | Mmys-BE-1 | + | JF502407 |
| Belgium† | 50.8 | 5.6 | 18/03/2010 | Myotis mystacinus | n/a | + | n/a |
| Netherlands† | 52.0 | 5.8 | 09/03/2010 | Myotis daubentonii | Mdau-NL-1 | + | JF502411 |
| Netherlands | 52.1 | 4.3 | 27/02/2010 | Myotis dasycneme | Mdas-NL-1 | + | JF502410 |
| Germany† | 49.7 | 7.4 | 10/03/2010 | Myotis myotis | Mmyo-DE-12 | + | JF502401 |
| Germany | 49.8 | 9.6 | 22/03/2010 | Myotis myotis | Mmyo-DE-14 | + | JF502403 |
| Germany | 50.7 | 13.7 | 20/03/2010 | Myotis myotis | Mmyo-DE-13 | + | JF502402 |
| Germany | 50.9 | 7.5 | 18/04/2009 | Myotis myotis | Mmyo-DE-10 | + | JF502399 |
| Germany | 51.2 | 8.1 | 21/03/2010 | Myotis mystacinus | Mmys-DE-2 | + | JF502409 |
| Germany | 51.2 | 8.1 | 21/03/2010 | Myotis mystacinus | Mmys-DE-3 | + | n/a |
| Germany | 51.2 | 8.1 | 07/03/2010 | Myotis myotis | Mmyo-DE-11 | + | JF502400 |
| Germany | 51.2 | 8.1 | 07/03/2010 | Myotis myotis | Mmyo-DE-16 | + | n/a |
| Germany† | 52.3 | 9.5 | 23/03/2010 | Myotis myotis | Mmyo-DE-15 | + | JF502404 |
| Germany† | 52.3 | 9.4 | 23/03/2010 | Myotis mystacinus | Mmys-DE-1 | + | JF502408 |
| Hungary | 47.1 | 17.6 | 24/03/2010 | Myotis myotis | Mmyo-HU-2 | + | JF502405 |
| Hungary | 47.1 | 17.6 | 24/03/2010 | Myotis myotis | Mmyo-HU-3 | + | n/a |
| Poland | 50.8 | 16.7 | 07/03/2010 | Myotis myotis | Mmyo-PL-1 | + | JF502413 |
| Estonia#,† | 59.3 | 24.6 | 01/06/2010 | Myotis brandtii | EsT-01062010 | + | JF502412 |
| Ukraine | 48.7 | 26.6 | 17/03/2010 | Myotis myotis | Mmyo-UA-1 | + | JF502406 |
*Dead bat.
#Environmental sample (individual observed 23/05/2010; see text for further explanations).
†Photograph of the bat shown in Figure A21-2.
‡Samples were negative for G. destructans but amplified another fungus. doi:10.1371/journal.pone.0019167.t001
viously isolated (see list above). In some cases, we were not able to discriminate between M. Myotis and M. blythii (referred to as M. Myotis/blythii) as well as between the newly recognised M. escalerai (Ibañez et al., 2006; Cabrera, 1904) and Myotis sp. A (Garcia-Mudarra et al., 2009), a yet undescribed cryptic species from the M. nattereri species complex (Ibañez et al., 2006; Mayer et al., 2007). Additionally, swab samples were collected from the tunnel wall of an Estonian hibernaculum. On the 23rd May 2010, a M. brandtii was observed in this hibernaculum with white fungal growth on its snout (Figure A21-2A) but no sample was collected at the time. When the site was revisited for sample collection on the 1st of June 2010, the bat had left the site so samples were collected by swabbing the wall of the tunnel where the bat was seen nine days before. Four cotton swabs were used to sample different areas a few centimetres around the location where
Table A21-2 Suspected Photographic Records of Geomyces destructans on Hibernating Bats in Europe.
| Country | Lat. | Lon. | Date | Host species | No. Individual |
| France | 44.8 | 1.6 | 25/04/2008 | Myotis myotis | 1 |
| France† | 42.6 | 2.2 | 26/06/2010 | Myotis escalerai/sp.A | 1 |
| France | 47.7 | 22.1 | 04/03/2010 | Myotis myotis | 1 |
| France† | 45.0 | 2.0 | 13/02/2010 | Myotis myotis | 2 |
| France | 47.3 | 6.2 | 04/03/2010 | Myotis myotis | 3 |
| France | 50.4 | 3.5 | 01/03/2008 | Myotis mystacinus | 1 |
| France | 47.2 | 1.4 | 24/02/2010 | Myotis myotis | 2 |
| Belgium | 50.8 | 5.6 | 09/02/2008 | Myotis dasycneme | 1 |
| Belgium | 50.8 | 5.6 | 20/03/2008 | Myotis daubentonii | 1 |
| Belgium | 50.8 | 5.6 | 17/01/2010 | Myotis dasycneme | 1 |
| Belgium† | 50.3 | 5.9 | 07/03/2010 | Myotis myotis | 1 |
| Belgium | 50.8 | 5.7 | 13/03/2010 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 26/03/2008 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 18/02/2008 | Myotis dasycneme | 1 |
| Netherlands | 52.0 | 5.7 | 04/03/2010 | Myotis mystacinus | 1 |
| Denmark† | 56.4 | 9.1 | 14/03/2010 | Myotis dasycneme | 2 |
| Germany | 51.8 | 10.8 | 02/02/2008 | Myotis myotis | 1 |
| Germany | 51.6 | 10.5 | 07/02/2010 | Myotis myotis | 1 |
| Germany | 51.7 | 10.3 | 20/03/2010 | Myotis myotis | 1 |
| Germany† | 52.3 | 9.5 | 23/03/2010 | Myotis mystacinus | 1 |
| Germany | 52.3 | 9.5 | 23/03/2010 | Myotis dasycneme | 1 |
| Germany | 52.1 | 8.2 | 21/03/2007 | Myotis daubenonii | 1 |
| Germany | 52.1 | 8.2 | 14/03/2007 | Myotis dasycneme | 1 |
| Germany | 52.2 | 8.0 | 04/02/2008 | Myotis myotis | 1 |
| Austria† | 46.8 | 16.0 | 07/02/2007 | Myotis myotis | 1 |
| Hungary | 47.1 | 17.6 | 24/02/2007 | Myotis myotis | 1 |
| Hungary | 47.1 | 17.6 | 23/02/2009 | Myotis myotis | 1 |
| Hungary | 46.2 | 18.1 | 03/03/2009 | Myotis myotis/blythii | 1 |
| Hungary | 48.5 | 20.5 | 18/02/2010 | Myotis blythii | 1 |
| Hungary | 47.1 | 17.6 | 19/02/2010 | Myotis blythii | 1 |
| Hungary† | 47.1 | 17.6 | 19/02/2010 | Myotis myotis | 2 |
| Poland† | 50.8 | 16.7 | 07/03/2010 | Myotis myotis | 1 |
| Ukraine† | 48.8 | 26.6 | 13/02/2010 | Myotis myotis | 1 |
| Ukraine | 48.8 | 26.6 | 17/03/2010 | Myotis myotis | 8 |
| Romania† | 46.8 | 22.6 | 29/03/2008 | Myotis blythii | 1 |
| Romania | 45.4 | 25.2 | 14/03/2009 | Myotis myotis/blythii | 1 |
| Turkey† | 41.9 | 27.9 | 22/03/2009 | Myotis myotis/blythii | 1 |
†Photograph of the bat shown in Figure A21-2. doi:10.1371/journal.pone.0019167.t002
the bat was observed. The four swabs were then streaked onto four Sabouraud’s agar plates each and monitored regularly to physically remove any fungal growth that was not similar to G. destructans. Although the amount of fungi varied per swab sample, G. destructans was recovered from all four swabs, henceforth considered as a single sample, bringing the total of samples analysed to 23. No mass mortality was reported at any of the sites investigated.
Table A21-3 Suspected Visual Records of Geomyces destructans on Hibernating Bats in Europe
| Country | Lat. | Lon. | Date | Host species | No. Individual |
| France | 49.1 | 6.6 | 06/04/2009 | Myotis myotis | 1 |
| France | 48.5 | 6.9 | 28/02/2009 | Myotis myotis | 1 |
| France | 48.3 | 7.1 | 29/03/2009 | Myotis myotis | 1 |
| France | 48.3 | 5.7 | 16/03/2008 | Myotis myotis | 1 |
| France | 47.9 | 6.8 | 03/03/2010 | Myotis myotis | 2 |
| France | 49.5 | 5.2 | 04/03/2010 | Myotis myotis | 1 |
| France | 48.9 | 0.3 | 06/02/2010 | Myotis myotis | 1 |
| France | 47.2 | 5.7 | 20/02/2010 | Myotis myotis | 3 |
| Netherlands | 52.1 | 4.3 | 10/03/2005 | Myotis dasycneme | 2 |
| Netherlands | 52.1 | 4.3 | 24/06/2006 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 07/03/2007 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 15/03/2008 | Myotis dasycneme | 3 |
| Netherlands | 52.1 | 4.3 | 30/03/2008 | Myotis dasycneme | 2 |
| Netherlands | 52.1 | 4.3 | 05/04/2008 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 12/04/2008 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 13/02/2004 | Myotis dasycneme | 2 |
| Netherlands | 52.1 | 4.3 | 05/04/2003 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 26/03/2008 | Myotis dasycneme | 1 |
| Netherlands | 52.1 | 4.3 | 10/03/2005 | Myotis dasycneme | 1 |
| Germany | 50.9 | 13.3 | 23/03/2010 | Myotis daubentonii | 1 |
| Germany | 49.9 | 7.4 | 14/03/2010 | Myotis Myotis | 1 |
| Ukraine | 48.8 | 26.6 | 17/03/2010 | Myotis Myotis | 4 |
| Romania | 47.0 | 22.4 | 08/04/2008 | Myotis Myotis | 1 |
doi:10.1371/journal.pone.0019167.t003
Out of 23 samples investigated in the laboratory, 14 of the 16 touch imprint samples presented characteristic conidia when observed under a microscope and two of them were doubtful; none of the cotton swabs were inspected under a microscope prior to culture. Cultures from 22 of these samples were successful. The two dead bats investigated did not reveal the presence of G. destructans but other fungal species such as Mucor sp. and Cladosporium sp. were identified (data not shown).
DNA was isolated from the 22 cultures of which 20 showed morphological similarity to G. destructans (e.g., curved conidia) and from one touch imprint with unsuccessful culture attempts. Amplification and sequencing of the internal transcribed spacer ITS) region (ITS1, 5.8S, and ITS2) was preferred over the small subunit (SSU) rDNA as it was shown to be more informative and was comparable to both European (Puechmaille et al., 2010; Wibbelt et al., 2010) and North American G. destructans (Blehert et al., 2009, Chaturvedi et al., 2010). All sequences obtained were identical and showed 100% similarity with previously published ITS sequences of G. destructans available on GenBank (retrieved on
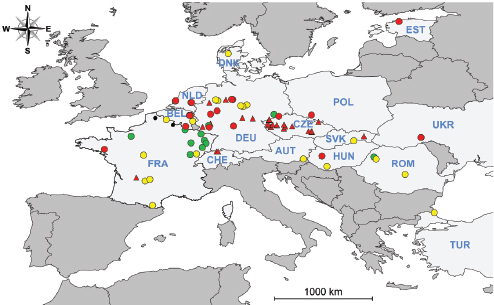
October 13th) (Puechmaille et al., 2010; Blehert et al., 2009, Wibbelt et al., 2010; Chaturvedi et al., 2010).
Seasonal distribution of G. destructans
The monitoring of one site over two consecutive winters showed an absence of Gd-suspect bats from September until the end of January (Figure A21-S1). The first Gd-suspect bats were reported in February each year (16/02/2007 and 07/02/2008) and their numbers peaked in March (Figure A21-3). In April, the total number of bats and the number of Gd-suspect bats decreased as bats left the hibernacula. However, as the number of Gd-suspect bats decreased more slowly than the total numbers of bats, the highest prevalence was observed in April (Figure A21-S1). Prevalence varied between years for the same period of the year and reached values in the range of 18–25% in 2007 (14th–28th March)
Figure A21-2 (facing page). Photographic evidence showing bats with confirmed or suspected growth of G. destructans. Photographs of cases confirmed by genetic analysis, from (A) Estonia (M. brandtii, May 23rd 2010, © L. Lutsar), (B) Poland (M. myotis, March 7th 2010, © A. Wojtaszewski), (C) Germany (M. myotis, March 10th 2010, © C. Jungmann), (D) France (M. myotis, March 4th 2010, © Y. Le Bris), (E) Netherlands (M. daubentonii, March 9th 2010, © T. Bosch), (F) Germany (M. myotis, March 23rd 2010, © K. Passior) (G) Belgium (M. mystacinus, March 18th 2010, © B. Mulkens), (H) Germany (M. mystacinus, March 23rd 2010, © K. Passior) or bats with white-fungal growth suspected as G. destructans from (I) Denmark (M. dasycneme, March 14th 2010, © B. Ohlendorf), (J) Austria (M. myotis, February 2nd 2007, © O. Gebhardt), (K) Hungary (M. myotis, February 19th 2010, © T. Görföl), (L) Belgium (M. myotis, March 7th 2010, © F. Forget), (M) France (M. myotis, February 13th 2010, © J. Vittier), (N) Ukraine (M. myotis, February 13th 2010, © A.-T. Bashta), (O) France (M. escalerai/sp. A, June 25th 2010, © F. Blanc), (P) Turkey (M. myotis/blythii, March 22nd 2009, © M. Doker), and (Q) Romania (M. blythii, March 29th 2008, © B. Szilárd). doi:10.1371/journal.pone.0019167.g002
or 28–55% in 2008 (13th–28th March) when the numbers of Gd-suspect bats are at the highest. The distributions of reported cases were similar between the two years, although more cases were reported in April in the winter 2007/2008 (Figure A21-3). In April 2008, the monitoring of three marked bats with white fungal growth clearly showed that after a bat had changed its position within the hibernaculum or when it was leaving the hibernaculum, the visible white fungal growth disappeared (Figure A21-4), most likely as a result of self-grooming.
The temporal distribution of reported cases of live Gd-suspect bats from throughout Europe (this study, n = 105) was combined with information available from previously reported cases of G. destructans (Puechmaille et al., 2009; Wibbelt et al., 2010) (n = 22) to investigate the seasonal variation across multiple sites in Europe. The temporal range of reported cases of Gd-suspect live bats and bats confirmed with G. destructans (n = 127) was not evenly distributed throughout the winter/ spring, with about 2/3rd of the cases reported in March (81/127; Figure A21-3). The number of reported cases more than doubled between February (30 cases) and March (81 cases). The earliest case was reported on January 17th from Belgium and the three latest cases were observed on May 23rd in Estonia, in June 24th in the Netherlands and June 25th in France (Tables A21-1, A21-2, A21-3, Figures A21-2A and A21-2).
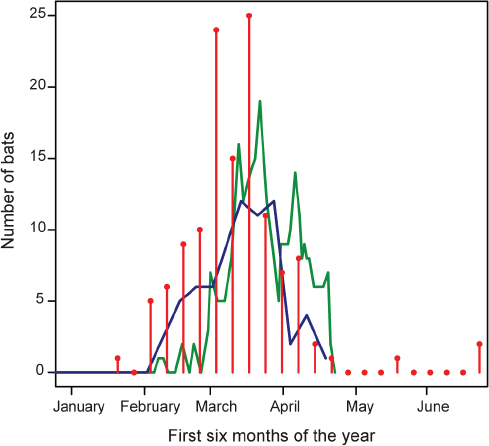
Figure A21-3 Seasonal changes of the number of live bats reported with white fungal growth in Europe. The number of bats with visible white fungal growth at an hibernaculum in Germany was monitored during the winter 2006/2007 (blue line) and the winter 2007/2008 (green line). The vertical red lines represent the number of Gd-suspect bats (or confirmed) observed across twelve European countries (n = 127) from 2003 until 2010. In the X-axis, the thick marks represent the start of each month. doi:10.1371/journal. pone.0019167.g003
Discussion
Presence of G. destructans in Europe
G. destructans was first identified in Europe in 2008–2009 (Puechmaille et al., 2009; Wibbelt et al., 2010) but increasing photographic evidence suggest that the fungus was present in Europe well before this date (this study, [Martínková et al., 2010; Feldmann 1984]). Most previous studies investigating fungi in European caves, including bat guano (Nováková, 2009; Mosca and Campanino, 1962; Groth et al., 1999; Nováková and Kolařik, 2010) reported Geomyces species, but none had curved conidia so far typical of G. destructans. In the Czech Republic,
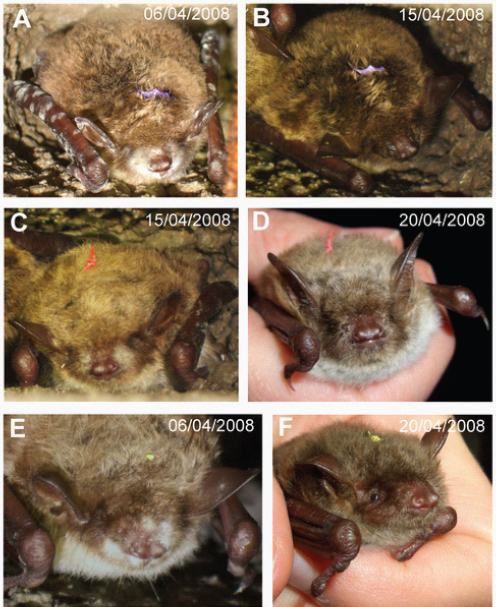
FIGURE A21-4 Indirect evidence of bats grooming off G. destructans during hibernation. Photographic evidence showing three different M. dasycneme individuals (A–B, C–D and E–F) observed at two different dates, first with visible fungal growth (A, C, E) and later without visible fungal growth (B, D, F). The bat in A–B changed its position within the hibernaculum whereas the other two (C–D and E–F) were captured when leaving the hibernaculum (© V. Korn). doi:10.1371/journal.pone.0019167.g004
Kubátová & Dvořák (2005) investigated fungi associated with insects hibernating in underground sites but did not find Geomyces species. To our knowledge, only one study in Europe has investigated fungi present in bats’ skin and hair samples where, based on our current knowledge, G. destructans is most likely to be found. During the winter 1999/2001, Larcher et al. (2003) collected 25 samples of hair and skin swabs from six species, including three Myotis Myotis, but did not find any Geomyces species. It is important to note that most fungal cultures have been carried out at temperatures above 24–25°C, temperatures at which G. destructans does not grow (Gargas et al., 2009; Chaturvedi et al., 2010), which could explain why although present, this fungal species had never been reported in Europe before the study of Puechmaille et al. (2010).
Combining previously published data from France, Germany, Switzerland, Hungary, The Czech Republic and Slovakia (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010), additional data collected from France, Germany and Hungary (this study), and new data from Belgium, The Nether-lands, Poland, Estonia and Ukraine (this study), we demonstrate here that G. destructans is widespread in Europe. We consider the photographic evidence of bats with white fungus matching the characteristic growth pattern (e.g., Figure A21-2; pictures from Romania and Turkey) to most likely represent G. destructans, because so far all tested live European bats with such white fungal growth on their nose, similar to Figure A21-2, have been confirmed to carry that species of fungus. These findings further support the fact that G. destructans is widespread across Europe. However, to confirm the presence of G. destructans in Europe prior to 2008, historical collections of bat specimens (or cave soil samples), especially specimens collected during the hibernation period, should be screened for the fungus.
As depicted in Figure A21-1, most cases of bats with G. destructans (confirmed and suspected) have been found from North-eastern France through Belgium, The Netherlands, Germany and the Czech Republic. However, it is not clear whether this pattern reflects an actual higher occurrence and/or prevalence of the fungus in these regions or if it is at least partly due to sampling bias, whereby the fungus is more likely to be detected in regions with a higher number of underground sites visited every winter or in regions were the fungus is specifically sought. In our opinion, it is most likely that this large-scale pattern is due to a sampling bias. For example, the largest number of sites with G. destructans in any European country was reported from the Czech Republic (76 localities with suspected or confirmed G. destructans) were most sites have been searched for signs of the fungus (>800 hibernacula) (Martínková et al., 2010).
G. destructans growth on bats
The clear seasonal peak in the number of observations of bats with white fungal growth indicates an increasing prevalence or detectability of G. destructans as winter passes. This suggests that bats might acquire G. destructans late
during the hibernation period or that the fungus is carried by bats at the onset of hibernation but needs time to develop the visible white fungal growth due to the phenology of the fungus. Therefore, the absence of visible white fungal growth on bats when observed with the naked eye may not directly reflect the absence of G. destructans, but rather just the absence of visible fungal colonies. Further complicating matters, our ability to detect G. destructans growth on bats can substantially differ with proximity to the bats (i.e., low ceiling versus high ceiling) or the location of the bat (ceiling versus crevices).
Our results confirm the suggestion of Martínková et al. (2010) by showing that during the hibernation period, bats can remove the fungus from their snout, ears and wings to a point where the fungus is no longer visible to the naked eye, although some spores might still be present on their skin. During hibernation, bats arouse every two weeks on average (Brack and Twente, 1985; Twente et al., 1985) and if bats consistently groom off the fungus on these occasions, our ability to visually detect the fungus, if present, will be considerably reduced. We also showed that towards the end of the hibernation period, bats were emerging from the hibernaculum without visible signs of the fungus despite showing visible white fungal growth from two weeks to five days before leaving the hibernaculum. It would be important to investigate whether bats carry spores out of hibernacula and as a result could possibly contaminate maternity roosts and maternity mates as suggested by Hallam and McCracken (2011).
Factors affecting G. destructans prevalence
Although it is not possible to clearly identify the mechanism responsible for the sudden increase in the prevalence of G. destructans in late February and March, these data suggest that shorter winter periods should be associated with lower prevalence. This prediction seems to hold as in the Mediterranean region, where hibernation periods are shorter (Rodrigues, 2003), no bats with visually conspicuous fungal growth have yet been reported during winter cave surveys. The case reported from Southern France (June 25th 2010, Figure A21-2) was found in the Pyrenees mountains at ca. 1700 m a.s.l. and hence, is not considered typical of the Mediterranean climate. It is nevertheless too early to conclude on this association between G. destructans prevalence and the hibernation duration, as other factors would need to be considered such as for example, the higher temperature observed in hibernacula in the Mediterranean region compared to other regions in Europe (Rodrigues, 2003). Higher temperatures in hibernacula have been associated with more frequent arousals in Rhinolophus ferrumequinum (Ransome, 1971; Arlettaz et al., 2000; Park et al., 2000). Considering that this association holds for other species, as a consequence of more frequent arousals, bats are expected to groom more often and therefore, reduce the probability of a visible fungal growth to develop. More surveys and strategic sampling efforts are needed to uncover whether the length of the hibernation period and/or climatic
conditions have a direct or indirect effect on the growth rates, prevalence, and detectability of G. destructans on bats.
It is crucial that the change in prevalence or detectability over the hibernation period is considered when comparing prevalence across sites and/or years. Our results from monitoring one site throughout the hibernation period over two consecutive years as well as reported cases from multiple sites in multiple years show that bats with fungal growth are first seen in January, the number of cases slowly increases into February and peaks in March, then in April when bats emerge from hibernation it drops again. Our results are in agreement with recent results from the Czech Republic where in the winter 2009/2010, the number of sites with bats with white fungal growth increased from 4.1% in January/February (33/800 sites; regular bat monitoring) to 77.5% in late February/March (76/98 sites; additional inspections) (Martínková et al., 2010) The Czech study reported that this increase in G. destructans prevalence was ‘‘suggestive of an epizootic spread of the fungus’’ (Martínková et al., 2010); we propose an alternative explanation whereby the increase in prevalence of G. destructans in late winter (March) might regularly (yearly) occur in Europe but has gone unnoticed. Nearly all hibernation counts in previous years were carried out between December and mid-February when prevalence/detectability of G. destructans is low, but not in March (Battersby, 2010) when the prevalence/detectability of G. destructans is at its highest (Figure A21-3). Although the total numbers of bats in the hibernacula decreased through April as bats left for the maternity colonies, our results show that there is a high probability of fungal growth developing on the remaining individuals. This further supports our hypothesis proposed above and links the duration of the hibernation period with the prevalence of G. destructans. By increasing the sample size, some cases might be reported earlier in the hibernation season or later through the summer, but we expect that the general pattern observed will not change. Despite these difficulties in assessing the occurrence of the fungus on bats, our data are consistent with other studies (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010), and also demonstrate that the most commonly encountered bat species with G. destructans in Europe is the largest species of Myotis on the continent, Myotis myotis. In countries/regions (i.e., the Netherlands, Northwest Germany) where M. dasycneme is more commonly encountered in hibernacula, G. destructans prevalence can reach high levels in that species. It is interesting to note that neither Pipistrellus pipistrellus nor Miniopterus schreibersii have been observed with G. destructans (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010), although these two species are known to hibernate in aggregations of tens of thousands of individuals, especially the latter (Furman and Özgül, 2004; Nagy and Postwana, 2011; Benda et al., 2003; Serra-Cobo et al., 1998). Although rare, hibernacula of a few thousands and up to about 34,000 individuals are also known for species of Myotis in Europe (Furman and Özgül, 2004; Nagy and Postwana, 2011; Kokurewicz, 2009; Arthur and Lemaire, 2009; Sachanowicz et al., 2006; Dietz et al., 2009).
G. destructans outside of the hibernation period
We observed three individual bats with white fungal growth around their nose (one confirmed as G. destructans) from May and June, when they were still torpid in cold underground sites. This represents the first mention of individuals with G. destructans colonisation outside of the hibernation period and raises questions about the role of these individuals in the persistence of the fungus in bat populations. Pipistrellus pipistrellus During the summer period, while females aggregate in colonies to raise their young, it remains largely unknown where males are roosting (e.g., Senior et al., 2005). Furthermore, during the swarming season in late summer/autumn, large numbers of individuals aggregate in caves, mines or tunnels and come in close contact with each other (chasing, mating) (Senior et al., 2005; Parsons and Jones, 2003; Parsons et al., 2003a; Parsons et al., 2003b; Rivers et al., 2006; Rivers et al., 2005), which could represent an opportunity for G. destructans to be transmitted between individuals.
We isolated G. destructans from the environment surrounding hibernating bats. The presence of viable spores of G. destructans on the surfaces of hibernation sites has huge implications for the understanding of disease transmission mechanisms and disease modelling (Hallam and McCracken, 2011) It seems likely that cave walls could serve as a passive vector and/or reservoir for G. destructans spores. It is not yet known how long these spores can remain viable but fungal spores generally remain viable for extended periods. Bats entering these sites in autumn (for swarming and/or hibernation) could become contaminated with spores of G. destructans left from bats infected during the previous winter. In North-America, Lindner et al. (2010) successfully amplified ITS sequences identical to G. destructans DNA from soil samples collected during the winter 2008–2009 at three bat hibernacula and stressed the importance of considering the environment as a reservoir for G. destructans and in the dynamics of WNS transmission. Our results confirm this and further suggest that more work is needed to understand the persistence of G. destructans on hibernacula walls (reservoir or passive vector) where they are in physical contact with bats.
Insights into the origin of G. destructans and WNS
The wide distribution of G. destructans in Europe and the absence of associated mortality supports the hypothesis that G. destructans has co-evolved with European bats and only recently arrived in North America where it is causing unprecedented mass mortalities (Puechmaille et al., 2010; Blehert et al., 2009; Martínková et al., 2010; Wibbelt et al., 2010). Alternatively, G. destructans could have been present on both continents and a virulent strain could have evolved in North-America. Until the relationships between G. destructans populations across continents are clarified, precautions should be taken to minimise the chances of transcontinental movement of viable G. destructans (Puechmaille et al., 2011).
During the two years monitoring at one site in Germany where G. destructans prevalence reached high levels in March-April, not a single dead bat was found. This is in agreement with previous studies (Puechmaille et al., 2010; Martínková et al., 2010; Wibbelt et al., 2010) reporting that the presence of G. destructans in bats from Europe is not associated with mass mortality. This sharply contrasts with mass mortalities reported in North America where hundreds or thousands of dead bats are found in hibernacula towards the end of the hibernation period. Recent pathological investigations of bats dying from WNS in North America led Cryan et al. (2010) to propose that mortality was caused by important disruptions of wing-dependant physiological functions due to infection by G. destructans. In North America, the fungus deeply invades wings tissues (Meteyer et al., 2009) and causes damages that are thought to alter homeostasis and water balance, resulting in more frequent arousals than bats can afford with their fat reserves, leading to death by starvation (Cryan et al., 2010). The pathology associated with G. destructans colonisation in Europe is not yet known. We believe that the first step in understanding mortality differences between bats from Europe and North America rely on understanding pathological differences incurred by the fungus on the bats’ wings. As a result, we urge the necessity to carry out pathological investigation of live bats from Europe colonised by G. destructans. Despite the absence of mortality associated with the presence of G. destructans in Europe, it would be necessary to investigate whether chronic infections with the fungus are compromising the health of individuals, especially in M. Myotis and M. dasycneme, which show high prevalence of the fungus towards the end of the hibernation period.
Phylogeographic studies of European bat species have shown that in the last 100,000 years, some species colonised Europe from Western Asia (Flanders et al., 2009), including Myotis blythii (Berthier et al., 2006; Currat et al., 2008) which has been found with G. destructans (Wibbelt et al., 2010). Assuming that G. destructans can be transported over long distances by bats, we speculate that the distribution of G. destructans is probably not limited to Europe and possibly extends eastwards into Russia, Western and Central Asia. Further surveys are necessary to clarify the global distribution of G. destructans.
Conclusions
We have shown here that G. destructans, the most likely causative agent of WNS in North America, is widespread in Europe, but is not associated with mass mortality. The prevalence of visible fungal growth on bats increases in February/March before sharply decreasing when bats emerge from hibernation. We also isolated viable G. destructans from the walls of an underground site suggesting that the hibernacula could act as passive vectors and/ or reservoirs for G. destructans and therefore, might play an important role in the transmission process. Further research is needed to clarify the global prevalence of G. destructans and
identify variables (e.g., temperature, humidity and hibernation length) explaining regional differences. Finally, further research is needed in different parts of the globe, especially temperate region of the Northern and Southern hemispheres, to precisely determine the global distribution of G. destructans.
Materials and Methods
Sample collection
During ongoing population censuses carried out at hibernacula in different countries across Europe and during additional hibernacula surveys carried out for the purpose of this study, information on bats with visible white fungal growth on snouts and/or ears was recorded. Whenever possible, sterile dry cotton swabs (Puechmaille et al., 2010) or adhesive tape touch imprints (Wibbelt et al., 2010) were used to collect fungal material from the bats. In Estonia, samples were collected from the wall of the tunnel where a bat with characteristic white fungus was observed nine days prior to the sampling. Where no sample collection was possible, a photograph was taken of the bat (photographic record). In cases where neither sample collection nor photographic evidence was obtained, the record was classified as visual observation. Live hibernating bats with powdery, white fungal growth on their noses were considered suspects of infection by G. destructans (Gd-suspects) but not suspected of having WNS. There is presently no data supporting the occurrence of WNS in Europe and the co-occurrence of the fungus with lesions characteristic of WNS (Meteyer et al., 2009) has not (yet) been reported in Europe (Wibbelt et al., 2010; Barlow et al., 2005). Although, prevalence of G. destructans can reach high levels in some European species (i.e., Myotis myotis, M. dasycneme) in late winter (especially in March), it can be expected that by chance alone some bats dying from causes unrelated to the presence of G. destructans will also be carrying the fungus. Unless the criteria for the diagnosis of WNS are met (confirmation by histo-pathology and PCR) (Meteyer et al., 2009) WNS should not be assumed as a cause of mortality in dead bats found in hibernacula of Europe. Various species of fungi have been identified on dead bats (Wibbelt et al., 2010; Voyron et al., 2011), most of them likely being saprophytes that colonise bat carcasses post-mortem.
Fungal cultures
In the laboratory, samples were treated as in Puechemaille et al. (2010) for swabs and following Wibbelt et al. (2010) for touch imprints. Briefly, swabs were streak-plated onto plates of Sabouraud’s agar, supplemented with 0.1% mycological peptone. For touch imprints, small areas with fungal conidia characteristic of G. destructans were identified by light microscopy and the tape was disinfected and excised before being transferred for culture to Sabouraud’s agar.
The plates were sealed with parafilm and incubated inverted in the dark at 10°C. A fungal growth developed within 14 days, from which single spore cultures were established.
Molecular identification
Each culture was sequenced for one molecular marker, the rRNA gene internal transcribed spacer (ITS, ca. 930 bp.) region (ITS1, 5.8S, and ITS2) to further confirm species identity. The DNA extraction, PCR amplification and DNA sequencing followed protocols described in Puechmaille et al. (2010). Briefly, DNA was extracted using the Qiagen Blood and Tissue kit following the manufacturer’s instructions with slight modifications (after step 3, we added an incubation time of 10 minutes at 70°C). PCR reactions were carried out in 25 mL containing 1 mL of DNA extract (at 10–75 ng/mL), 1.5 mmol/L MgCl2, 0.4 mmol/L each primer (Forward: ITS4, 5′-TCCTCCGCTTATTGATATGC – 3′; Reverse: ITS5, 5′-GGAAGTAAAAGTCGTAACAAGG – 3′; (White et al., 1990), 0.2 mmol/L dNTP, 1x PCR buffer and 1 U Platinum Taq DNA Polymerase High Fidelity (Invitrogen). PCR cycling conditions were: initial step 15′ at 95°C, then 10 cycles of 30″ at 95°C, 1′45″ at 60°C (reduce of 2°C every 2 cycles), 1′ at 72°C, following by 30 cycles of 30″ at 95°C, 1′45″ at 50°C and 1′ at 72°C. PCR products were purified and sequenced by Macrogen Inc. (Seoul, Korea) in both directions using the PCR primers. Complementary sequences were assembled and edited for accuracy using CodonCode Aligner 3.0.3 (www.codoncode.com/aligner/download.htm).
Monitoring of visible fungal growth on bats
One site situated in Northwest Germany (Latitude: 52.1; Longitude: 8.2) near the city of Osnabrück was monitored over two consecutive winters, 2006/2007 (5th September until 19th May) and 2007/2008 (28th August until 23rd April). The monitoring consisted of counting the total number of bats at the site as well as the number of bats with visible white fungal growth similar to the pictures presented in Figures A21-2 and A21-4. The counts were done by the same person (V. Korn) every 4 days on average during the first year and every 2.5 days on average during the second year. The procedures complied with guidelines of the American Society of Mammalogists and were carried out under permit number FBD7.2 60 from the Administration of the County of Osnabrück, Department of Environment.
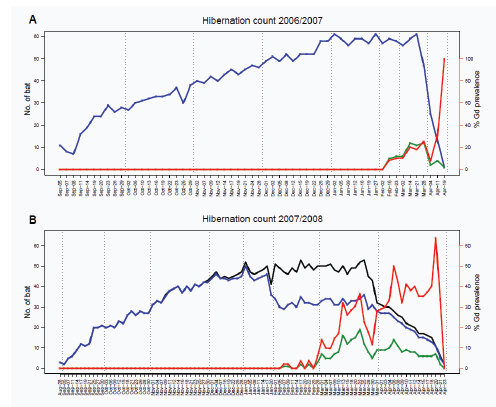
FIGURE A21-S1 Monitoring of bats at an hibernaculum in Germany during (A) the winter 2006/2007 (September 5th 2006 until April 19th 2007) and (B), the winter 2007/2008 (August 28th 2007 until April 23rd 2008). The blue line represents the total number of bats counted whereas the green line represents the number of bats with visible white fungal growth (Gd-suspects). Dotted vertical lines separate counts from each month. Note that the number of counts per month was not equal between months. In (B), the black line represents the total number of bats counted whereas the blue line represents the total number of bats bar one portion of the hibernaculum where bats grouped densely (ca. 20 individuals) and did not allow a reliable identification of the number of bats with white fungal growth. The green line represents the number of bats with visible white fungal growth (Gd-suspects) counted at the hibernaculum without considering individuals densely grouping at one place in the hibernaculum. The group of about 20 individuals formed while the hibernaculum was partially flooded, likely as a result of bats changing position to avoid drowning. Note that the right Y-axis scale is different between (A) and (B).
Acknowledgments
We would like to thank Dóczy Annamária, Andriy-Taras Bashta, Frédéric Blanc, Sándor Boldogh, Gaby Bollen, Thomas Chatton, Emrah Coraman, Jére Csaba, Simon Dutilleul, Mehmet Doker, Oliver Gebhardt, Lena Godlevska, René Janssen, Daniel Lefèvre, Barti Levente, Vadim Martyniuk, Gerhard Mascher, Mykola Matveev, Bernd Ohlendorf, Rian Pulles, Tony Rock, Wolfgang Rackow, Sébastien Roué, Bücs Szilárd, Abigel Szodoray-Parádi, Farkas Szodoray-Parádi and Julien Vittier for providing us with their field observations. The comments of Paul Cryan, Paul Racey, Natalia Martínková and an anonymous reviewer helped to improve a previous version of the manuscript.
Author Contributions
Conceived and designed the experiments: SJP GW VK ECT. Performed the experiments: SJP GW HF VK KM AK. Analyzed the data: SJP GW. Contributed reagents/materials/analysis tools: SJP GW VK HF KM AK FF WB CB TB TC MD TG AJH FH GH MH CJ YLB LL MM BM KP MS AW UZ ECT. Wrote the paper: SJP GW.
References
A national plan for assisting states, federal agencies, and tribes in managing white-nose syndrome in bats. 2010 Draft v. 10.21.2010.16 p.
Arlettaz R, Ruchet C, Aeschimann J, Brun E, Genoud M, et al. (2000) Physiological traits affecting the distribution and wintering strategy of the bat Tadarida teniotis. Ecology 81:1004–1014.
Arthur L, Lemaire M (2009) Les Chauves-souris de France, Belgique, Luxembourg et Suisse: Mèze: Biotope, Paris: Muséum national d’Histoire naturelle. 576 p.
Barlow A, Ford S, Green R, Morris C, Reaney S (2009) Investigation into suspected white-nose syndrome in two bat species in Somerset. Vet Rec 165:481–482.
Bastian F, Alabouette C, Saiz-Jimenez C (2009) The impact of arthropods on fungal community structure in Lascaux Cave. J Appl Microbiol 106:1456–1462.
Battersby J (2010) Guidelines for surveillance and monitoring of European bats. Bonn, Germany. 95 p.
Benda P, Ivanova T, Horáček I, Hanák V, Cerveny J, et al. (2003) Bats (Mammalia: Chiroptera) of the Eastern Mediterranean. Part 3. Review of bat distribution in Bulgaria. Acta Soc Zool Bohem 67:245–357.
Berthier P, Excoffier L, Ruedi M (2006) Recurrent replacement of mtDNA and cryptic hybridization between two sibling bat species Myotis Myotis and Myotis blythii. Proc R Soc B 273:3101–3109.
Blehert DS, Hicks AC, Behr M, Meteyer CU, Berlowski-Zier BM, et al. (2009) Bat white-nose syndrome: an emerging fungal pathogen? Science 323: 227.
Brack V, Twente JW (1985) The duration of the period of hibernation of three species of vespertilionid bats. I. Field studies. Can J Zoolog 63:2952–2954.
Cabrera A (1904) Ensayo monogra´fico sobre los quirópteros de España. Mem Soc Españ Hist Nat 2:249–292.
Chaturvedi V, Springer DJ, Behr MJ, Ramani R, Li X, et al. (2010) Morphological and molecular characterizations of psychrophilic fungus Geomyces destructans from New York bats with White Nose Syndrome (WNS). PLoS ONE 5: e10783.
Courtin F, Stone WB, Risatti G, Gilbert K, Van Kruiningen HJ (2010) Pathologic findings and liver elements in hibernating bats with white-nose syndrome. Vet Pathol 47:214–219.
Cryan P, Meteyer CU, Boyles JG, Blehert DS (2010) Wing pathology of whitenose syndrome in bats suggests life-threatening disruption of physiology. BMC Biol 8: 135.
Currat M, Ruedi M, Petit RJ, Excoffier L (2008) The hidden side of invasions: massive introgression by local genes. Evolution 62:1908–1920.
Dietz C, Von Helversen O, Dietmar N (2009) Bats of Britain, Europe & Northwest Africa. London: A & C Black Publishers Ltd. 400 p.
Flanders J, Jones G, Benda P, Dietz C, Zhang S, et al. (2009) Phylogeography of the greater horseshoe bat, Rhinolophus ferrumequinum: contrasting results from mitochondrial and microsatellite data. Mol Ecol 18:306–318.
Furman A, Özgül A (2004) The distribution of cave-dwelling bats and conservation status of underground habitats in Northwestern Turkey. 120:243–248.
Garcia-Mudarra JL, Ibañez C, Juste J (2009) The straits of Gibraltar: barrier or bridge to IberoMoroccan bat diversity? Biol J Linn Soc 96:434–450.
Gargas A, Trest MT, Christiensen M, Volk TJ, Blehert DS (2009) Geomyces destructans sp. nov. associated with bat white-nose syndrome. Mycotaxon 108:147–154.
Geomyces destructans Widespread in Europe White nose syndrome science strategy meeting II, 2009. Consensus Statement. Austin, Texas.
Groth I, Vetermann R, Scuetze B, Schumann P, Saiz-Jimenez C (1999) Actinomycetes in karstic caves of northern Spain (Altamira and Tito Bustillo). J Microbiol Meth 36:115–122.
Hallam TG, McCracken GF (2011) Management of the panzootic White-Nose Syndrome through culling of bats. Conserv Biol 25:189–194.
Ibañez C, Garcia-Mudarra JL, Ruedi M, Stadelmann B, Juste J (2006) The Iberian contribution to cryptic diversity in European bats. Acta Chiropt 8:277–297.
Kannan K, Hun Yun S, Rudd RJ, Behr M (2010) High concentrations of persistent organic pollutants including PCBs, DDT, PBDEs and PFOS in little brown bats with white-nose syndrome in New York, USA. Chemosphere 80:613–618.
Kokurewicz T (2009) Management Plan for the Natura 2000 site ‘‘Nietoperek’’ (Western Poland). International Conference Military Heritage, Utrecht, The Netherlands.
KubátováA, Dvořák L (2005) Entomopathogenic fungi associated with insect hibernating in underground shelters. Czech Mycol 57:221–237.
Larcher G, Bouchara JP, Pailley P, Montfort D, Beguin H, et al. (2003) Fungal biota associated with bats in Western France. J Mycol Méd 13:29–34.
Lindner DL, Gargas A, Lorch JM, Banik MT, Glaser J, et al. (2010) DNA-based detection of the fungal pathogen Geomyces destructans in soils from bat hibernacula. Mycologia. In press.
Martínková N, Bačkor P, Bartonička T, Blažkova´ P, Červený J, et al. (2010) Increasing incidence of Geomyces destructans fungus in bats from the Czech Republic and Slovakia. PLoS ONE 5:e13853.
Mayer F, Dietz C, Kiefer A (2007) Molecular species identification boosts bat diversity. Front Zool 4:4.
Meteyer CU, Buckles EL, Blehert DS, Hicks AC, Green DE, et al. (2009) Histopathologic criteria to confirm white-nose syndrome in bats. J Vet Diagn Invest 21:411–414.
Mosca AML, Campanino F (1962) Analisi micologiche del terreno di grotte piemontesi. Allionia. pp. 27–43.
Nagy ZL, Postwana T (2011) Seasonal and geographical distribution of cavedwelling bats in Romania: implications for conservation. Anim Conserv 14:74–86.
Nováková A (2009) Microscopic fungi isolated from the Domica Cave system (Slovak Karst National Park, Slovakia). A review. Int J Speleol 38:71–82.
Nováková A, Kolařik M (2010) Chrysosporium speluncarum, a new species resembling Ajellomyces capsulatus, obtained from bat guano in caves of temperate Europe. Mycol Progress 9:253–260.
Park KJ, Jones G, Ransome RD (2000) Torpor, arousal and activity of hibernating Greater Horseshoe bats (Rhinolophus ferrumequinum). Funct Ecol 14:580–588.
Parsons KN, Jones G (2003) Dispersion and habitat use by Myotis daubentonii and Myotis nattereri during the swarming season: implications for conservation. Anim Conserv 6:283–290.
Parsons KN, Jones G, Davidson-Watts I, Greenaway F (2003a) Swarming of bats at underground sites in Britain-implications for conservation. Biol Conserv 111:63–70. Geomyces destructans Widespread in Europe PLoS ONE | www.plosone.org 10 April 2011 | Volume 6 | Issue 4 | e19167 Parsons KN, Jones G, Greenaway F (2003b) Swarming activity of temperate zone microchiropteran bats: effect of season, time of night and weather conditions. J Zool 261:257–264.
Puechmaille SJ, Verdeyroux P, Fuller H, Ar Gouilh M, Bekaert M, et al. (2010) White-nose syndrome fungus (Geomyces destructans) in bat, France. Emerging Infectious Diseases 16:290–293.
Puechmaille SJ, Fuller H, Teeling EC (2011) Effect of sample preservation methods on the viability of Geomyces destructans, the fungus associated with whitenose syndrome in bats. Acta Chiropt. In press.
Ransome RD (1971) The effect of ambient temperature on the arousal frequency of the hibernating greater horseshoe bat, Rhinolophus ferrumequinum, in relation to site selection and the hibernation state. J Zool 164:353–371.
Reichard JD, Kunz TH (2009) White-nose syndrome inflicts lasting injuries to the wings of little brown Myotis (Myotis lucifugus). Acta Chiropt 11:457–464.
Rice AV, Currah RS (2006) Two new species of Pseudogymnoascus with Geomyces anamorphs and their phylogenetic relationship with Gymnostellatospora. Mycologia 98:307–318.
Rivers NM, Butlin RK, Altringham JD (2005) Genetic population structure of Natterer’s bats explained by mating at swarming sites and philopatry. Mol Ecol 14:4299–4312.
Rivers NM, Butlin RK, Altringham JD (2006) Autumn swarming behaviour of Natterer’s bats in the UK: population size, catchment area and dispersal. Biol Conserv 127:215–226.
Rodrigues L, Zahn A, Rainho A, Palmeirim JM (2003) Contrasting the roosting behaviour and phenology of an insectivorous bat (Myotis Myotis) in its southern and northern distribution ranges. Mammalia 67:321–335.
Sachanowicz K, Sachanowicz M, Piksa K (2006) Distribution patterns, species richness and status of bats in Poland. Vespertilio 9–10:151–173.
Senior P, Butlin RK, Altringham JD (2005) Sex and segregation in temperate bats. Proc R Soc Lond B 272:2467–2473.
Serra-Cobo J, Sanz-Trullen V, Martinez-Rica JP (1998) Migratory movements of Miniopterus schreibersii in the north-east of Spain. Acta Theriol 43:271–283.
Twente JW, Twente J, Brack V (1985) The duration of the period of hibernation of three species of vespertilionid bats. II. Laboratory studies. Can J Zoolog 63:2955–2961.
Voyron S, Lazzari A, Riccucci M, Calvini M, Varese GC (2011) First mycological investigations on Italian bats. Hystrix. In press.
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, eds. PCR protocols: a guide to methods and applications: Academic Press, Inc., New York. pp. 315–322.
Wibbelt G, Kurth A, Hellmann D, Weishaar M, Barlow A, et al. (2010) White-Nose Syndrome fungus (Geomyces destructans) in bats, Europe. Emerging Infectious Diseases 16:1237–1242.