Evolutionary Transitions in
Bacterial Symbiosis
![]()
JOEL L. SACHS,*†RYAN G. SKOPHAMMER,*AND
JOHN U. REGUS*
Diverse bacterial lineages form beneficial infections with eukaryotic hosts. The origins, evolution, and breakdown of these mutualisms represent important evolutionary transitions. To examine these key events, we synthesize data from diverse interactions between bacteria and eukaryote hosts. Five evolutionary transitions are investigated, including the origins of bacterial associations with eukaryotes, the origins and subsequent stable maintenance of bacterial mutualism with hosts, the capture of beneficial symbionts via the evolution of strict vertical transmission within host lineages, and the evolutionary breakdown of bacterial mutualism. Each of these transitions has occurred many times in the history of bacterial–eukaryote symbiosis. We investigate these evolutionary events across the bacterial domain and also among a focal set of well-studied bacterial mutualist lineages. Subsequently, we generate a framework for examining evolutionary transitions in bacterial symbiosis and test hypotheses about the selective, ecological, and genomic forces that shape these events.
Ancestrally, bacteria and archaea persisted solely as free-living cells in terrestrial and aquatic habitats. Along with the evolution and diversification of animals and plants, the past 500 million years have also witnessed a massive radiation of bacteria. Bacterial
_____________
*Department of Biology, University of California, Riverside, CA 92521. †To whom correspondence should be addressed. E-mail: joels@ucr.edu.
lineages have evolved diverse mechanisms to gain entry and proliferate in the tissues and cells of multicellular eukaryotes (Merhej et al., 2009; Carvalho et al., 2010; Medina and Sachs, 2010; Toft and Andersson, 2010), and these symbionts vary in their effect on hosts from harmful to beneficial (Medina and Sachs, 2010; Toft and Andersson, 2010). Archaea have also evolved associations with hosts, but these interactions do not appear as diverse or ubiquitous. Bacterial symbioses (defined in the broad sense) include persistent, intimate associations between bacteria and other species and date back at least to the origins of eukaryotes (Sagan, 1967). Bacterial parasites range from infectious diseases that rapidly exploit hosts before infecting new individuals, to bacteria that are transmitted vertically from host parent to offspring and manipulate host reproduction to favor their own spread (Stouthamer et al., 1999). Parasitic bacteria have received intense focus from researchers over the last century because harmful infections represent a critical challenge to human health and economic interests. In contrast, except for a few early pioneers (Buchner, 1921), researchers have only recently focused on the biology of bacteria that enhance host fitness: bacterial mutualists (Sachs et al., 2011).
Bacterial mutualists are diverse (Williams et al., 2007, 2010; Merhej et al., 2009; Wu et al., 2009; Carvalho et al., 2010; Medina and Sachs, 2010; Philippot et al., 2010; Toft and Andersson, 2010) and exhibit a variety of lifestyles and coevolutionary relationships with eukaryote hosts (Sachs et al., 2011) (Table 2.1). First, beneficial bacteria vary in their degree of reliance on hosts for reproduction. Whereas some bacterial-derived organelles and endosymbionts cannot live independently of hosts, most bacterial mutualists retain extensive environmental phases and form infections that are facultative for the bacterium (Szathmáry and Smith, 1995; Nyholm and McFall-Ngai, 2004; Sachs et al., 2011). Second, bacterial mutualists inhabit diverse host tissues ranging from skin, mucosa, leaves, and roots to inter- and intracellular spaces. Some bacterial mutualists inhabit specialized structures in hosts (Becking, 1970; Savage, 1977; Sprent et al., 1987; Douglas, 1989; Bright and Sorgo, 2003; Currie et al., 2006; Nussbaumer et al., 2006; Visick and Ruby, 2006; Goettler et al., 2007; Vaishnava et al., 2008; Pflugfelder et al., 2009; Ran et al., 2010), whereas others range widely in host mucosa or other unstructured tissues (Hirose, 2000; Hirose et al., 2009; Kaltenpoth et al., 2009) (Table 2.1). Finally, bacterial mutualists provide a great variety of benefits to hosts, including nutrients (Becking, 1970; Sprent et al., 1987; Douglas, 1989; Hirose, 2000; Hooper et al., 2002; Ran et al., 2010), bioluminescence (Nyholm and McFall-Ngai, 2004), and antibiotic production (Currie et al., 1999; Kaltenpoth et al., 2005; Kost et al., 2007). Although bacterial mutualists by definition provide a net fitness benefit to hosts,
they can also bear features that exploit hosts (Frank, 1996a,b; Sachs and Wilcox, 2006; Simms et al., 2006; Weeks et al., 2007; Oliver et al., 2008; Heath et al., 2010; Sachs et al., 2010a, 2011). As we detail later, each of these variables (degree of reliance on hosts, type of host habitat, and type of benefit provided to host) can modulate evolutionary transitions in bacterial symbiosis and can explain how and why transitions occur.
Here, we investigate evolutionary transitions that have occurred in the history of bacterial mutualism. We focus on (i) the origins of host association in bacteria (transitions in which environmental bacteria evolve to form intimate and persistent associations with hosts irrespective of effects on host fitness), (ii) the origins of bacterial mutualism from other types of bacterial lifestyles, (iii) shifts to the stable maintenance of bacterial mutualism, (iv) the capture of bacterial mutualists (via the evolution of strict vertical transmission within host lineages), and (v) the evolutionary breakdown of bacterial mutualism. Each of these events has occurred multiple times in the evolution of bacteria. Only symbiont capture possibly constitutes a “major evolutionary transition,” defined as an integrating event in which partners lose the ability to replicate independently (Szathmáry and Smith, 1995). However, loss of independence often only occurs for the symbiont.
To study broad patterns and genetic drivers of transitions, we investigate phylogenomic data that span the bacterial domain (Williams et al., 2007, 2010; Merhej et al., 2009; Wu et al., 2009; Philippot et al., 2010; Toft and Andersson, 2010) (Fig. 2.1), and to study fine-scale patterns, we also analyze a focal set of bacterial mutualists (Table 2.1). Our domain-level data sources include a phylogeny with 350 bacterial taxa sampled from 20 phyla (Wu et al., 2009), coupled with phenotypic host-association data (Boussau et al., 2004; Merhej et al., 2009; Bright and Bulgheresi, 2010; Philippot et al., 2010; Toft and Andersson, 2010). The focal systems include beneficial symbionts chosen to represent host and bacterial diversity, breadth in symbiotic services, and variety in transmission modes. Our analysis of historical and selective scenarios that characterize transitions in bacterial symbiosis complements other work that has focused on genomic changes (Merhej et al., 2009; Carvalho et al., 2010; Medina and Sachs, 2010; Toft and Andersson, 2010). The phylogeny of Wu and colleagues (2009) and the review by Toft and Andersson (2010) are particularly germane to this study as they provide the domain-level dataset that we use to test hypotheses.
There are caveats to consider when inferring the evolutionary history of bacterial symbiosis at broad phylogenetic scales. First is the challenge of assigning host-association traits to bacterial species. Recent work suggests that fitness benefits provided by bacteria to hosts can be context dependent (Heath and Tiffin, 2007; Oliver et al., 2008; Heath et
TABLE 2.1. Fourteen Focal Bacterial-Host Mutualisms Analyzed
| Symbiont, Host | Benefits Provided by Bacteria to Host | Host Localization | Transmissibn Among Hosts |
| Rhizobia [e.g..Sawada et al.(2003)], legumes | Nitrogen fixation (Sprentet al., 1987) | Nodules (Sprentet al., 1987) | Horizontal transmission (Sprentet al., 1987) with free-living stages (Sachs et al., 2009) |
| Frankia spp., actinorhizal plants | Nitrogen fixation (Becking, 1970) | Nodules (Becking, 1970) | Horizontal transmission with free-living stages (Huss-Danell and Frej, 1986) |
| Pseudonocardia spp. (fungus-growing ants) | Antibiotics (Currie et al.,1999; Kost et al., 2007) | Crypt structures on exoskeleton (Currie et al.,2006) | Vertical transmission to offspring ant colonies (Currie et al., 1999) and horizontal transmission with environmental pool (Mueller et al., 2008, 2010) |
| Endoriftia persepfmie, tubeworm | All nutrients (Nussbaumer et al., 2006) | Lobules in host trophosome (Bright and Sorgo, 2003; Nussbaumer 2006; Pflugfelder et al., 2009) | Horizontal with free-living stages (Nussbaumer et al.,et al., 2006) |
| Burkholderia spp., stinkbugs | Unknown nutrients (Kikuchi et al.,2007) | Midgut crypts (Kikuchi et al., 2011) | Horizontal with free- living stages (Kikuchi et al., 2011) |
| Bacleroides thetaiotaomicron, humans | Nutrients OHooper et al., 2002) | Crypt structures in gut (Savage, 1977; Vaishnava et al., 2008) | Horizontal transmission (Savage, 1977) with free-living stages (Carson et al., 2005) |
| Vibrio fisdieri, bobtail squids | Bioluminescence (Kyholm and McFall-Ngai, 2004) | Deep crypts in light organ (Visick and Ruby, 2006) | Horizontal transmission with free-living stages (Visick and Ruby, 2006) |
| Host Association Origins | Mutualism Breakdown | Forces Stabilizing Bacterial Mutualism |
| Mutualist (Fig. 2.1) | Abandonment events (Sawada et al., 2003; Sachs et al., 2009, 2010a) | Partner choice (Kiers el al., 2003; Simms et al., 2006; Sachs et al., 2010b) |
| Mutualist (Fig. 2.1) (Normand et al., 1996) | No evidence | Unknown, host localization consistent with partner choice |
| Mutualisl (Mueller el al., 2010) | Abandonment events (Mueller et al., 2010) | Byproducts (see discussion), no evidence of partner choice (Kost et al., 2007) |
| Ambiguous (Williams etal., 2010) | No evidence | Unknown, host localization consistent with partner choice |
| Parasite (Fig. 2.1) | No evidence | Byproducts (Discussion), partner fidelity feedback (Wilkinson, 1999; Turnbaugh et al., 2009), partner choice (Vaishnava et al., 2008) |
| Parasite (Fig. 2.1) | Abandonment events (Kishiguchi and Nair, 2003) | Partner choice (Sachs et al., 2004,2010b) |
| Symbiont, Host | Benefits Provided by Bacteria to Host | Host Localization | Transmissibn Among Hosts |
| vrochioron spp.,didemnid ascidians | photosynthates Wares (Hirose, 2000) | Unstructured in cloacal cavity (Hirose, 2000; Ran et al., 2010) | Vertical transmission via physical transfer to larvae (Hirose, 2000). No known free-living state (Kojima and Hirose, 2010) |
| Coriobacterium glomerans, firebugs | Aid in digestion (Kaltenpoth et al., 2009) | Unstructured in guts (Kaltenpoth et al., 2009) | Vertical transmission (via egg inoculation. Little potential for horizontal transmission or free-living stages (Kaltenpoth et al., 2009) |
| Streptomyces philanihi, beewolves | Antibiotics (Kaltenpoth et al., 2005) | Lobed antennomere reservoirs in antennae (Goettler et al., 2007) | Vertical transmission via brood provisioning of bacteria (Kaltenpoth et aL, 2010). No known free-living state |
| "Mycetocyte" bacteria, diverse insects (Douglas, 1989) | Amino acids, vitamins (Douglas, 1989) | Unstructured in mycetocytes in diverse tissues (Douglas, 1989) | Vertical transmission via host transfer to oocytes, eggs, or larvae (Douglas, 1989) |
| Cyanobaclerium spp., water fern | Nitrogen fixation (Ran et al., 2010) | Cavities in leaves (Ran et al., 2010) | Vertical transmission via bacterial motility, no free-living stage (Ran etal., 2010) |
| Plastids, plants Mitochondria, eukaryotes | Photosynthates Metabolism | Unstructured, intracellular | Iransovanal, no tree-living stage |
Notes: Bacterial symbionts are indicated with genus and species when possible, and hosts are identified with common names. "Mutualist Benefits" specifies the types of resources or services that the bacterial symbionts provide to their hosts. "Host Localization" specifies the location that the bacteria inhabit during the majority of or key parts of their interactions with hosts and whether these localesare structured spatially. 'Transmission Among Hosts" specifies transmission mode, and presence of free-living stages are identified. "Host-Association Origins" specifies the inferred ancestral condition at the origin of host association in the described lineage(s). "Mutualism
| Host Association Origins | Mutualism Breakdown | Forces Stabilizing Bacterial Mutualism |
| Mutualist (MunchhoH et a., 2007) | No evidence or mutualism breakdown | Vertical transmission promotes partner fidelity feedback |
| Ambiguous (Kaltenpoth et al 2009) | No evidence of mutualism breakdown | Vertical transmission promotes partner fidelity feedback |
| Mutualist (Kaltenpoth et al., 2006) | No evidence of mutualism breakdown | Vertical transmission promotes partner fidelity feedback |
| Parasite (Fig. 2.1) | No evidence of mutualism breakdown | Vertical transmission promotes partner fidelity feedback |
| Mutualist (Svenning et al., 2005) | No evidence of mutualism breakdown | Vertical transmission promotes partner fidelity feedback |
| Mutualist (Turner et al., 1999) Parasite (Williams et al., 2007) | No evidence of mutualism breakdown | Vertical transmission promotes partner fidelity feedback |
Breakdown” specifies evidence of evolutionary transitions in bacterial lineages from mutualism to other lifestyles, with “abandonment” referring to transitions from mutualism to an environmental lifestyle. “Forces Stabilizing Bacterial Mutualism” specifies potential forces stabilizing cooperation in a bacterial mutualist lineage, divided into the three model classes [byproduct cooperation, partner choice, and partner fidelity feedback (Sachs et al., 2004)].
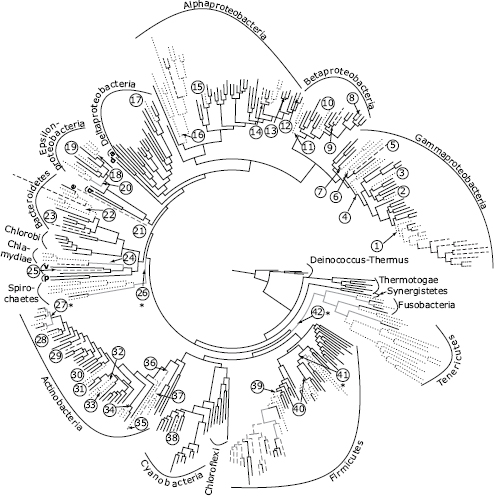
FIGURE 2.1 Inferred evolutionary history of bacterial host association. Ancestral states are inferred on a domain-level bacterial phylogeny modified from a previous study (Wu et al., 2009). The tree is a maximum likelihood reconstruction of a concatenated set of 31 single-copy genes from 350 bacterial species chosen to optimize phylogenetic sampling. Phyla and proteobacterial classes are labeled with their full names (e.g., Gammaproteobacteria; Firmicutes) or single-letter abbreviations (a, Acidobacteria; d, Defferribacteres; q, Aquificae; e, Elusimicrobia; v, Verrucomicrobia; p, Planctomycetes). Branch shades represent host-associated traits on the tips of the tree and inferred states on ancestral nodes (black, environmental; dashed gray, commensal; dashed black, mutualist; dotted, parasite). Host association traits were obtained from a prior review (Toft and Andersson, 2010). We inferred a minimum of 42 origins of host association (labeled 1–42). Origins at five nodes had equivocal parsimony reconstructions, noted with asterisks. Equivocal ancestral states are represented by gray branches. Additional origins are equally parsimonious at these nodes and provide an upper bound for global origins at 52.
al., 2010) and evolutionarily labile (Weeks et al., 2007; Sachs et al., 2010a, 2011), potentially blurring mutualist and parasite categories. Nonetheless, although striking exceptions exist (Weeks et al., 2007; Oliver et al., 2008), the majority of well-studied bacterial taxa can be unambiguously categorized into host-association categories (Moran and Wernegreen, 2000; Philippot et al., 2010; Toft and Andersson, 2010). Second is the challenge of accurately inferring past evolutionary events, which requires a robust and well-sampled phylogeny. The bacterial tree we use is well supported (Wu et al., 2009), but the sampling is sparse (relative to the domain of bacteria represented) and likely biased (only sequenced taxa are included). Finally, predictions about selective factors that drive transitions must be considered with caution, as phylogenetic comparisons often cannot distinguish evolution that predates the origins of host association from the consequences of these transitions. Our fine-scale analysis of the 14 focal symbioses serves as a complementary approach to help mitigate these challenges (Table 2.1).
ORIGINS OF HOST ASSOCIATION IN BACTERIAL LINEAGES
Origins of host association are transitions in which bacteria that live independently in the environment evolve to form intimate and persistent associations with hosts. To evolve host association, bacteria must be able to compete with other microbes on host surfaces, evade negative host responses, uptake novel resources on or inside the host, and ultimately gain transmission to new hosts. Considering these potential hurdles, one unanswered question is whether origins of host association are rare in bacterial lineages. Another question is whether certain bacteria taxa are more likely to evolve host association. In a phenotypic sense, this latter question addresses whether some bacteria bear preadaptations to host association.
Analyzing host association origins on a domain-level bacterial tree (Wu et al., 2009; Toft and Andersson, 2010) (Fig. 2.1 for taxon information), we inferred an environmental ancestral condition for the most recent common ancestor of bacteria and a minimum of 42 origins of host association across bacteria (Methods). An environmental ancestral condition is logical (as bacteria predate eukaryote hosts by at least 1 billion years) and is consistent with other analyses (Boussau et al., 2004). Origins of host association are diversely distributed across bacteria, emerging independently in at least 11 bacterial phyla. Proteobacteria, Actinobacteria, and Firmicutes each exhibit multiple origins of host association, whereas a few phyla such as Chlorobi, Chloroflexi, and Planctomycetes have never evolved host association (Madigan et al., 2009; Wu et al., 2009; Toft and Andersson, 2010).
Toft and Andersson (2010) predicted that bacterial preadaptations to host association might be ecological in nature, including access to mobile genes in soil and oceans and physical contact with diverse hosts, characteristics identified as common in Proteobacteria (Snel et al., 2002; Toft and Andersson, 2010). Although Proteobacteria exhibit 20 host-association origins, the evolutionary rate of host-association origins in this lineage (estimated as origins per adjusted branch length; Methods) is typical for eubacteria. Bacterial preadaptation to a host-associated lifestyle might also be genetically based, which is not mutually exclusive from ecological preadaptation. Several studies have begun to investigate genomic content changes correlated with transitions in host association, for instance by comparing phylogenetic relationships and genetic characteristics among bacterial mutualists, parasites, and related environmental species (Dale et al., 2001; Sawada et al., 2003; Horn et al., 2004; Frank et al., 2005; Ruby et al., 2005; Ma et al., 2006; Carvalho et al., 2010). The Rhizobiales represent an excellent case study, as these α-Proteobacteria include environmental bacteria, parasites, and mutualists (Sawada et al., 2003; Carvalho et al., 2010). Genomic comparisons of 19 species in this lineage uncovered a relatively small subset of loci unique to the host-associated species and revealed that these loci most often originated in host-associated lineages via horizontal transfer from other host-associated bacteria (Carvalho et al., 2010). Other lineages that encompass parasitic and mutualistic bacteria also show a similar pattern in which host-association loci exhibit evidence of horizontal gene transfer (Dale et al., 2001; Horn et al., 2004; Frank et al., 2005; Ruby et al., 2005; Ma et al., 2006). In summary, we found many origins of host association across bacteria and little evidence consistent with ecological or genomic predispositions to host association. The data suggest that transitions to host association might be constrained only by access to and compatibility with horizontally transferred loci that engender host-association traits (Toft and Andersson, 2010). Nonetheless, ecological constraints to host association cannot be ruled out; the bacterial taxa that have apparently never evolved host association might lack access to habitats with compatible hosts.
ORIGINS OF BACTERIAL MUTUALISM
Fundamental questions about the origins of bacterial mutualisms remain unresolved. Do bacterial mutualists evolve from parasitic ancestors or do they represent independent origins of host association (Ewald, 1987; Szathmáry and Smith, 1995; Corsaro et al., 1999; Moran and Wernegreen, 2000; Medina and Sachs, 2010)? If bacterial mutualists evolved from parasite ancestors, this predicts that transitions from
parasitism to mutualism have occurred, whereas if mutualists originate separately from parasites, this predicts that mutualists have evolved directly from environmental taxa. Two scenarios have been suggested to resolve this issue. Ewald (1987) introduced a detailed hypothesis for the origin of bacterial mutualism in which (i) an ancestral parasite infects hosts via both horizontal and vertical transmission, (ii) a mutation knocks out the parasite’s horizontal transmission pathway, and (iii) subsequent vertical transmission of the bacterium selects for reduced virulence and the enhancement of mutualistic traits [as vertical transmission can link reproductive interests of symbionts and hosts (Fine, 1975; Frank, 1996a,b; Sachs et al., 2004)]. This scenario is controversial because host-associated bacteria are thought to lack the genomic potential to easily switch from parasitism to mutualism (Moran and Wernegreen, 2000). The alternative hypothesis is that bacterial mutualists evolve directly from environmental bacteria, which is also problematic because it implies that free-living ancestors exhibited traits that could offer immediate benefits to hosts (Ewald, 1987).
We can empirically examine these alternative hypotheses by using the bacterial domain dataset (Wu et al., 2009; Toft and Andersson, 2010) and our focal systems (Table 2.1). At the domain level, many host-associated lineages are poorly sampled (Fig. 2.1), so this analysis must be considered preliminary. Bacteria on the domain-level tree include species classified as commensals, mutualists, and parasites (Toft and Andersson, 2010). Among the 42 host-association origins we reconstructed, 32 are inferred to have originated as parasites, 9 are inferred to have mutualist origins, and 1 origin is ambiguous (Fig. 2.1). Several mutualist taxa are nested in parasitic clades, consistent with three independent transitions from parasitism to mutualism (Fig. 2.1). It is unknown whether the evolution of vertical transmission drove these transitions because, in most lineages, the taxon sampling is poor and the order of events cannot be resolved. Among the nine mutualist lineages that evolved directly from environmental ancestors, six are nitrogen fixing. Consistent with Ewald’s (1987) hypothesis, nitrogen fixation is an ancient bacterial trait (Raymond et al., 2004) that can potentially offer hosts immediate benefits. However, as we observed earlier for the origins of host association, nitrogen fixation loci are also prone to horizontal transfer as parts of genome islands. This creates a scenario in which bacterial mutualists can evolve de novo from environmental ancestors via the gain of a core set of symbiosis loci (Sullivan et al., 1995; Sachs et al., 2010a).
Among the 14 focal taxa, we can infer the host-association origins of 12 (Table 2.1). Three of the lineages that likely represent transitions from parasitism to mutualism are vertically transmitted (Burkholderia spp., “Mycetocyte” bacteria, mitochondria), consistent with the hypoth-
esis that loss of horizontal transmission drove the origin of mutualism (Ewald, 1987). The history of the mitochondrion is somewhat ambiguous. Although some authors have suggested that mitochondria originated from a parasitic lineage of rickettsial bacteria (Moran and Wernegreen, 2000), no analysis of which we are aware has tested this hypothesis explicitly. In none of these cases can we resolve whether vertical transmission evolved before or after the transition from parasitism to mutualism. Seven of the symbioses are inferred to have originated as mutualists directly from environmental ancestors. As described earlier, these lineages carry traits that can offer immediate benefits to hosts, including antibiotic production, nitrogen fixation, and photosynthesis (Table 2.1). More detailed phylogenetic analysis is needed to resolve whether these cooperative traits predate the host association, as predicted by Ewald (1987). Finally, there are two symbioses that do not fit any of the aforementioned hypotheses. Both Bacteroides thetaiotaomicron and Vibrio fischeri are mutualists inferred to have evolved from parasites with no history of vertical transmission. For B. thetaiotaomicron (a dominant gut symbiont in humans), there is the possibility of pseudovertical transmission (Wilkinson, 1999; Turnbaugh et al., 2009). This is the hypothesis that hosts are more likely to transmit symbionts to kin, which approximates the effects of vertical transmission (Wilkinson, 1999). In summary, mutualist bacteria can evolve from environmental or parasitic ancestors. Bacterial phenotypes that offer immediate benefits to hosts are thought to promote origins of mutualism in environmental bacterial lineages, but well-studied cases implicate horizontal gene transfer (Sullivan et al., 1995; Sachs et al., 2010a) as an alternative. Vertical transmission is a predicted driver of transitions from parasitism to mutualism, but there is relatively little support for vertical transmission preceding the origin of mutualism (Weinert et al., 2009).
MAINTENANCE OF BACTERIAL MUTUALISM
In mutualist bacteria, it can be challenging to explain what prevents the spread of cheater mutants; symbionts that gain in fitness by exploiting hosts and giving little or nothing in return (Sachs et al., 2004). Three classes of models have been proposed for the maintenance of cooperation between species—byproduct cooperation, partner fidelity feedback, and partner choice (Axelrod and Hamilton, 1981; Bull and Rice, 1991; Sachs et al., 2004; Foster and Wenseleers, 2006)—and each of these models applies to bacterial mutualism. Byproduct cooperation occurs when the benefit provided by the symbiont to the host exists as an automatic consequence of a selfish trait, and thus byproduct cooperation carries no net cost for the symbiont (Brown, 1983; Connor, 1995). Partner
fidelity feedback exists when fitness benefits delivered from a symbiont to its host feed back as returned benefits to the symbiont, such that beneficial symbionts are rewarded and harmful symbionts experience reduced fitness (Bull and Rice, 1991; Simms and Taylor, 2002; Sachs et al., 2004). Fitness feedbacks are only expected when symbionts and hosts interact repeatedly over time, such as occurs with vertical transmission. Partner choice occurs when hosts preferentially reward beneficial symbionts and or sanction cheaters, thus producing a selective advantage for symbiont cooperation (Bull and Rice, 1991; Denison, 2000; Sachs et al., 2004). To what degree is byproduct cooperation, partner fidelity feedback, or partner choice responsible for the maintenance of cooperative symbioses? These models can work independently or in concert with each other (Sachs et al., 2004; Foster and Wenseleers, 2006); however, little empirical research has compared their prevalence.
Among our 14 focal symbioses, byproduct cooperation can mostly be ruled out, such as in Rhizobia, in which nitrogen fixation is costly and occurs only during the symbiosis (Sachs and Simms, 2008). In contrast, we are not aware of examples in which byproduct cooperation has been demonstrated. Such scenarios are certainly possible. For instance, Actinomycete bacteria produce antibiotics on fungus-farming ants that keep the ants’ fungal gardens pathogen-free (Table 2.1) (Currie et al., 1999). Antibiotic production is an anticompetitive function that benefits bacteria directly, whether on the surface of an ant or free in the soil, so it likely qualifies as a byproduct. Similarly, the symbiont B. thetaiotaomicron benefits humans by foraging and catabolizing compounds that the host cannot otherwise digest (Sonnenburg et al., 2005). The consumption of complex molecules and releasing of simpler compounds also must benefit Bacteroides directly. Byproduct cooperation is likely important for the origins of cooperative symbioses (Sachs et al., 2004), but when interactions have been established, hosts are expected to rapidly evolve traits to promote the infection and proliferation of beneficial symbionts (Connor, 1995; Foster and Wenseleers, 2006). For the B. thetaiotaomicron-human symbiosis, these host traits might include mechanisms to bias symbiont transmission to offspring [to maximize partner fidelity (Wilkinson, 1999; Turnbaugh et al., 2009)] or mechanisms to favor beneficial strains over more selfish ones [e.g., partner choice (Vaishnava et al., 2008)].
There is vigorous debate over the relative importance of partner fidelity feedback versus partner choice (Bull and Rice, 1991; Simms and Taylor, 2002; West et al., 2002a,b; Weyl et al., 2010; Archetti et al., 2011). Partner fidelity feedback is often equated with vertically transmitted symbioses, as vertical transmission tightly correlates symbiont and host reproductive interests (Sachs et al., 2004; Foster and Wenseleers,
2006). By this measure, partner fidelity is widespread across bacteria with multiple origins and diverse mechanisms of vertical transmission (Table 2.1). However, vertical transmission does not guarantee symbiont cooperation, as even rare opportunities for horizontal transfer or the potential to manipulate host reproduction can lead to parasitic bacterial phenotypes. For example, vertically transmitted parasites [such as some Wolbachia lineages (Weeks et al., 2007)] manipulate hosts to maximize their own transmission by biasing host sex ratio toward females (they are not transmitted to males) or by inducing cytoplasmic incompatibility (Stouthamer et al., 1999). On the contrary, most symbionts are horizontally transmitted (Nyholm and McFall-Ngai, 2004; Sachs et al., 2011). Under horizontal transmission, multiple symbiont genotypes often infect hosts, and, with rare exceptions (Sachs and Wilcox, 2006), partner fidelity is predicted to be weak (West et al., 2002a,b). Partner choice can efficiently select for symbiont cooperation under these conditions (Bull and Rice, 1991; Denison, 2000; West et al., 2002a,b; Foster and Wenseleers, 2006). Partner choice has been best demonstrated for legumes that form symbioses with nitrogen-fixing Rhizobia (Kiers et al., 2003; Simms et al., 2006; Sachs et al., 2010b) and squids that form symbioses with bioluminescent V. fischeri (Visick et al., 2000; Sachs et al., 2004). In both examples, hosts exhibit mechanisms to reward cooperative symbionts and punish cheaters. It can be difficult to experimentally distinguish partner-fidelity feedback from partner choice (Weyl et al., 2010). However, one approach is to assess if symbionts are spatially structured within the host. The degree to which hosts can spatially separate symbiont genotypes is a key prerequisite for partner choice mechanisms (Denison, 2000; West et al., 2002a,b; Sachs et al., 2004), but should have no bearing on partner fidelity feedback. Many hosts of horizontally transmitted bacteria have evolved specialized structures that can separate symbionts that vary in their fitness effects on the host and potentially aid in distinguishing beneficial strains from cheaters (Becking, 1970; Savage, 1977; Sprent et al., 1987; Douglas, 1989; Bright and Sorgo, 2003; Currie et al., 2006; Nussbaumer et al., 2006; Visick and Ruby, 2006; Goettler et al., 2007; Vaishnava et al., 2008; Pflugfelder et al., 2009; Ran et al., 2010) (Table 2.1 and Fig. 2.2). In most of these examples, there is no more than a correlation between symbiotic structure on hosts and the potential for partner choice. However, these data become powerful when coupled with phylogenetic and ecological information. Kikuchi and colleagues (2011) analyzed the presence and structure of midgut crypts among 124 species of stinkbugs that vary in diet as well as the presence of horizontally transmitted Burkholderia symbionts (Table 2.1 and Fig. 2.2). They found that (i) stinkbugs exhibit multiple Burkholderia genotype infections, a key prerequisite for partner choice;
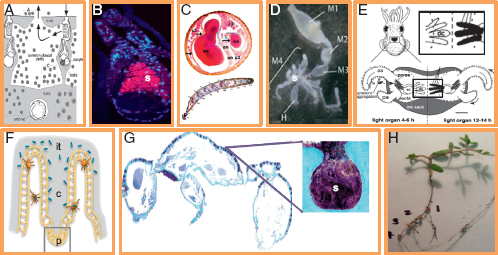
FIGURE 2.2 Symbiont housing structures and their potential to promote spatial structure. (A) Host Ascidian Diplosoma spp. and symbiont Prochloron spp. unstructured in host cloacal cavity [reprinted from Hirose et al. (2009)]. (B) Host hydrothermal tubeworm Riftia pachyptila with symbiont Endoriftia persephone (s) unstructured in host trophosome [reprinted from Nussbaumer et al. (2006)]. (C) Antenna of host beewolf Philanthus triangulum with symbiont Streptomyces (ws) housed in structured serial antennomere reservoirs (cross section above; longitudinal section below) [reprinted from Goettler et al. (2007)]. (D). Four-chambered midgut of host stinkbug Dimorphopterus pallipes with symbiont Burkholderia spp. (s) housed in structured crypts of fourth midgut section (m4) [reprinted from Kikuchi et al. (2011)]. (E) Juvenile squid host Euprymna scolopes during colonization by symbiont Vibrio fischeri, housed in structured deep crypts [dc; adapted from Visick and Ruby (2006)]. (F) Host mouse small intestine and symbiont Bacteroides thetaiotaomicron in structured crypts of Lieberkuhn (c) based with Paneth cells (p) [adapted from Vaishnava et al. (2008)]. (G) Dorsal cross section of host ant Cyphomyrmex longiscapus with Actinomyces symbionts (s) housed in structured crypts [reprinted from Currie et al. (2006)]. (H) Host legume Lotus strigosus with symbiont Bradyrhizobium japonicum structured in four numbered nodules (photo by J. L. Sachs).
(ii) the Burkholderia symbiosis has evolved in some, but not all, of the stinkbug species that exhibit midgut crypts; (iii) there is no evidence that the Burkholderia symbiosis has evolved in stinkbug species without such crypts; and (iv) crypts are not strictly correlated with different feeding habits of the bugs. These data suggest that crypts—which can potentially separate beneficial from harmful symbionts (Kikuchi et al., 2011)—are a key factor promoting stability in this bacterial mutualism. In summary, there is controversy over the relative importance of
partner-fidelity feedback and partner choice as the key selective forces that maintain bacterial mutualisms (Bull and Rice, 1991; Simms and Taylor, 2002; West et al., 2002a,b; Weyl et al., 2010; Archetti et al., 2011). However, spatial separation among symbiont genotypes is a predicted indicator of partner choice (Denison, 2000; West et al., 2002a,b; Sachs et al., 2004), and such structure is common.
SYMBIONT CAPTURE
Symbiont capture occurs when bacteria that can replicate in the environment evolve to be strictly vertically transmitted within hosts and lose independent life stages. The most basal form of transmission is horizontal and likely occurs when bacteria are acquired from environmental pools (Huss-Danell and Frej, 1986; Nussbaumer et al., 2006; Mueller et al., 2008, 2010; Sachs et al., 2009; Barke et al., 2010). In other cases of horizontal transmission, the symbiont taxa can be found in the environment (Nishiguchi and Nair, 2003; Carson et al., 2005), but most transmission likely occurs among hosts (Savage, 1977; Wilkinson, 1999; Turnbaugh et al., 2009; Wollenberg and Ruby, 2009) with little contribution from environmental pools. Vertical transmission modes range from direct symbiont transfer within host germ lines to host behavioral mechanisms that supplement offspring with symbionts (Bright and Bulgheresi, 2010) (Table 2.1). Moreover, some bacteria cannot be easily categorized into horizontal or vertical transmission modes. For instance, some bacterial lineages are transmitted vertically, but in rare events, get horizontally transmitted to novel hosts, likely through vectors or predation (Russell et al., 2003; Dale and Moran, 2006). In most cases, captured lineages of bacteria are mutualists (our focus here), but obligate intracellular parasites such as Wolbachia and Rickettsia can also exhibit strict vertical transmission.
Symbionts with strict vertical transmission exhibit reduced effective population size and are subject to the accumulation of deleterious mutations and gene loss (Moran, 2003; Toh et al., 2006), transfer of DNA to host genomes (Martin and Herrmann, 1998), and obligate reliance on the host for basic nutrient synthesis (Shigenobu et al., 2000). Captured symbionts also experience reduced access to novel genetic material via horizontal gene transfer (Dale and Moran, 2006; Toft and Andersson, 2010), which limits the potential for novel functions to evolve and for recombination to restore function to degraded genomes. Such genome degradation tends to worsen over time (Moran et al., 2009) and ultimately cause loss of functions that are required for life outside of the host (Merhej et al., 2009). Hence, vertical transmission is often an irreversible evolutionary endpoint.
An unexplored question about symbiont capture is whether host, symbiont, or joint mechanisms are responsible for these evolutionary transitions. Although the evolution of vertical transmission can be costly to symbionts, hosts experience benefits including transmitting mutualists to offspring, minimizing symbiont diversity, and reducing mixing among symbiont genotypes, all of which promote symbiont cooperation (Frank, 1996a,b; Sachs et al., 2004). Thus, symbiont capture should be correlated with the evolution of host mechanisms to control transmission (Frank, 1996a). In some cases, hosts have specialized structures with no obvious function other than to transfer bacteria to offspring. Female stinkbugs bear organs on their ovipositors (Kikuchi et al., 2009) that transfer symbionts to their eggs. The ascidian Diplosoma similis (Hirose, 2000; Hirose et al., 2009; Kojima and Hirose, 2010) exhibits a specialized “plant rake,” which it extends into its cloacal cavity during spawning and thus transfers bacterial symbionts to newly spawned larvae. In many cases, vertical transmission relies on specific host behaviors, such as when females smear symbionts onto eggs, egg cases, or cocoons of offspring (Douglas, 1989; Hirose, 2000; Kaltenpoth et al., 2005, 2010; Hirose et al., 2009; Kikuchi et al., 2009; Kojima and Hirose, 2010). However, bacterial mutualists can also promote their own vertical transmission. Among insect symbionts that inhabit mycetocyte structures within their hosts (Table 2.1), the bacteria sometimes migrate in the host from their mycetocyte structures to the host ovaries (Douglas, 1989). Wolbachia that infect Drosophila use the host microtubule cytoskeleton and transport system to maximize vertical transmission (Ferree et al., 2005). Moreover, the bacterial symbiont of the water fern Azolla filiculoides differentiates into a motile form and actively moves from adult plant leaves to infect the sporocarp of offspring plants (Ran et al., 2010). In all the examples in which the symbiont bears mechanisms to promote vertical transmission, there is no free-living existence and no potential for horizontal transfer (Table 2.1). Not surprisingly, when vertical transmission is the only mechanism to invade new hosts, symbiont traits are selected to enhance its efficiency. In summary, symbiont capture within host lineages involves a suite of deleterious effects that degrade symbiont genomes while providing benefits to hosts. As predicted by theory, the evolution of symbiont capture appears to be mostly driven by host mechanisms, but only a handful of bacterial–host interactions have been studied in detail (Bright and Bulgheresi, 2010).
BREAKDOWN OF SYMBIOSIS
There is debate about the evolutionary robustness of mutualisms, of which beneficial microbe–host interactions are a subset. Mutualist pop-
ulations have been predicted to be prone to extinction (Vandermeer and Boucher, 1978), the spread of cheater mutants (Axelrod and Hamilton, 1981; Bull and Rice, 1991), and reversions to free-living existence (Vandermeer and Boucher, 1978; Keeler, 1985; Holland et al., 2004), but other research predicts that mutualisms are robust to these challenges (Doebeli and Knowlton, 1998; Ferrière et al., 2007; Douglas, 2008). Evolutionary transitions that result in the loss of mutualistic traits (Sachs and Simms, 2006) can be divided into transitions from mutualism to parasitism and transitions from mutualism to free-living status (i.e., abandonment of mutualism). Ancient bacterial mutualisms (Sagan, 1967; Moran, 2003; Keeling, 2010; Ran et al., 2010) serve as empirical examples of long-term robustness, but it is unknown whether such stability is common.
To what degree does mutualism breakdown occur in bacteria? We can investigate the evolutionary stability of bacterial mutualism by using the domainwide phylogeny (Wu et al., 2009; Toft and Andersson, 2010) (Fig. 2.1) and our focal symbioses (Table 2.1). The domainwide data can be considered only preliminary because of the paucity of dense taxon sampling. We could only infer two evolutionary transitions from mutualism to other lifestyles: one transition from mutualism to parasitism and one abandonment of mutualism. Nonetheless, this is a surprising paucity of transitions considering that we inferred 72 evolutionary transitions on the tree (Figs. 2.1 and 2.3).
Among the 14 focal systems, there is evidence of mutualism breakdown in four, all of which involve transitions from mutualism to free-living status in symbionts with extensive free-living stages (Table 2.1). Two particularly dynamic examples of mutualism breakdown have been uncovered in symbionts of ants (Mueller et al., 2010) and stinkbugs (Kikuchi et al., 2007, 2011). In the case of the ants, the symbionts are antibiotic-producing Actinobacteria that live in cuticular crypts supported by specialized exocrine glands (Currie et al., 2006). Lineages of these Actinobacteria have likely undergone multiple transitions between host-associated and environmental status based on the intermixing of symbiotic and environmental genotypes on a population-level phylogeny (Mueller et al., 2010). Similarly, a phylogeny of the Burkholderia bug symbionts encompasses many environmental isolates, consistent with multiple transitions from symbiotic to environmental status (Kikuchi et al., 2011). Evidence for abandonment of symbiosis has also been found among rhizobial lineages, some of which are related to plant and mammal parasites as well as environmental bacterial species (Sawada et al., 2003), suggesting the potential for multiple transitions among mutualism, parasitism, and environmental lifestyles (Sachs and Wilcox, 2006) likely driven by horizontal transfer events of symbiosis loci (Young and
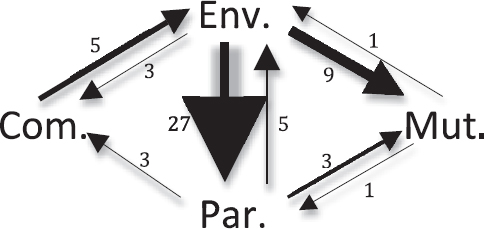
FIGURE 2.3 Path diagram of evolutionary transitions among bacterial host-association types. Transitions among four bacterial host-association types inferred in the tree by Wu and colleagues (2009) using lifestyle data from Toft and Andersson (2010). Thirteen transitions were undetermined on the tree as a result of ambiguity. There were zero transitions between mutualism and commen-salism and zero transitions from commensalism to parasitism. Arrow sizes are scaled to the number of transitions between host-association types. Note: Com., commensal; Env., environmental; Mut., mutualist; Par., parasite.
Haukka, 1996). More focused analyses have inferred multiple events of evolutionary abandonment of mutualism within Bradyrhizobium populations (Sachs et al., 2009, 2010a), but found no evidence of transitions from mutualism to parasitism (Sachs et al., 2010a). In Bradyrhizobium, the abandonment of mutualism appears to be driven by degradation or wholesale loss of symbiosis loci encoded on a genome island (Sachs et al., 2010a). Finally, there is evidence of abandonment of mutualism within lineages of beneficial V. fischeri, with at least three evolutionary transitions from mutualism to environmental status (Nishiguchi and Nair, 2003) (Table 2.1). In summary, among different lifestyles that bacteria can exhibit, mutualism with hosts appears to be evolutionary stable with few transitions to other lifestyles. We found transitions from mutualism to free-living status, but virtually no evidence of transitions from mutualism to parasitism.
DISCUSSION
The evolutionary history of bacterial mutualism is rich and ancient. The origin of host association appears to be a readily surmountable step for bacteria. The commonness and near universality of this transition suggests that it is selectively advantageous and might be rarely affected
by ecology. The evolution of bacterial mutualism is also common and phylogenetically diverse, and can occur via multiple routes. Bacterial mutualism most often appears to emerge from environmental ancestors. This can occur because the ancestral bacteria bear key traits (that can immediately benefit hosts) or by horizontal gene transfer of symbiosis loci (Sullivan et al., 1995; Sachs et al., 2010a), but neither mechanism is well understood. Bacterial mutualism can also arise from parasitic ancestors. It has been predicted that transitions from parasitism to mutualism are promoted by the evolution of vertical transmission (Ewald, 1987); however, more detailed work is needed to test this hypothesis. When bacterial mutualism has evolved, it can be stabilized via several selective mechanisms (Sachs et al., 2004). Partner choice, concomitant with the ability of hosts to spatially structure bacterial genotypes, is likely the dominant force maintaining bacterial mutualism.
Bacterial symbiosis first evolved with horizontal transmission, and several bacterial lineages have subsequently evolved strict vertical transmission. Some of the most ancient cases of bacterial mutualism exhibit vertical transmission, so this transition can promote the evolutionary stability of symbioses. We hypothesize that transitions from horizontal to obligate vertical transmission are host driven, as hosts (but not symbionts) most benefit from these transitions. Finally, evolutionary losses of bacterial mutualism are rare compared with other transitions in bacterial symbiosis. Evolutionary reversions from mutualism to environmental status occur in some bacterial lineages, potentially driven by the degradation or deletion of genes that encode symbiotic traits (Sachs et al., 2010a). In contrast, there is virtually no evidence in the phylogenetic record of transitions from mutualism to parasitism, thus refuting theory that predicts that mutualisms are vulnerable to fixation of cheater mutants (Axelrod and Hamilton, 1981; Bull and Rice, 1991; Sachs et al., 2004). The lack of transitions from mutualism to parasitism suggests that (i) bacterial mutualisms are evolutionarily robust or (ii) transitions from mutualism to parasitism are themselves unstable [and lead to extinctions or other stable states (Sachs and Simms, 2006)].
METHODS
We analyzed evolutionary transitions on a published 350-species bacterial phylogeny reconstructed by using a concatenated alignment of 31 proteins with maximum likelihood [PhyML (Guindon and Gascuel, 2003)] and an AMPHORA pipeline (Wu and Eisen, 2008; Wu et al., 2009) (Fig. 2.1, Table 2.1). Host-associated phenotypes were assigned based on a recent review (Toft and Andersson, 2010) that included host-association classifications of parasitic, mutualistic, commensal, or no interaction. We
divided classifications into two characters: (i) association (host-associated or environmental) and (ii) type of host interaction (parasitic, mutualistic, commensal). Ancestral states were inferred by using parsimony [Mesquite 2.74 (Maddison and Maddison, 2010)]. When two equally parsimonious ancestral state reconstructions were found, we noted the ambiguity and listed a minimum estimate of transitions (Fig. 2.1).
To compare the relative frequencies of host-association origins among different bacterial lineages, we estimated the rate of origins over evolutionary time for each phylum and the complete tree. Rates were calculated by dividing the total number of origins of host association in a lineage by an adjusted sum of the taxon’s branch length. The adjusted sum included only branches on which transitions from an environmental lifestyle to host association could occur (i.e., summed branch length of the taxon minus host-associated descendant branches of previously accounted origins and individual branches on which host association has been lost). The unit of branch length is the expected number of amino acid substitutions per site.
For focal symbiont taxa, we analyzed phylogenies containing the lineages of interest to assess whether host association originated from parasitic ancestors or free-living ancestors and to search for evidence of mutualism breakdown. Ancestral states for symbiotic lineage and evidence of mutualism breakdown were inferred by using parsimony on the available phylogenies (Normand et al., 1996; Turner et al., 1999; Nishiguchi and Nair, 2003; Sawada et al., 2003; Ruby et al., 2005; Svenning et al., 2005; Kaltenpoth et al., 2006, 2009; Kikuchi et al., 2007, 2011; Münchhoff et al., 2007; Williams et al., 2007, 2010; Sachs et al., 2009, 2010a; Mueller et al., 2010).
This page intentionally left blank.






















