Informatics and Personalized Medicine
POINTS HIGHLIGHTED IN THE KEYNOTE ADDRESS BY LEROY HOOD
• The digital information of the genome and the environmental information that modifies the digital genomic information connect to produce the phenotype through biological networks.
• A systems view of disease postulates that disease is the result of perturbation of one or more biological networks, leading to altered expression of information.
• User-, domain-, and data-driven analytic informatics tools will allow researchers to decipher the billions of data points collected for an individual and use them to define and understand individual wellness and disease.
• Together, systems biology and advanced informatics tools applied to disease and wellness provide the foundation for a personalized approach to medicine, allowing health care to be more predictive, preventive, personalized, and participatory.
Presenting the keynote address of the workshop, Leroy Hood, president and co-founder of the Institute for Systems Biology, shared his perspective on how the tremendous volumes of data currently becoming available can be utilized to advance medicine. An essential problem in biology in the 21st
century is complexity, Hood said. He described his vision for the convergence of systems biology and the “digital revolution” to transform medicine into an enterprise that he has termed “proactive P4 medicine,” in which P4 refers to predictive, preventive, personalized, and participatory.
Hood said that the digital revolution has provided three key elements that will play a central role in medicine going forward: (1) big datasets, (2) social and business networks that evolve from the knowledge gained from big datasets, and (3) digital personalized devices that will lead to the generation of a “quantized self.”
Hood predicted that in 10 years, each individual will be surrounded by a virtual data cloud of billions of data points from many different types of networks (e.g., genome, proteome, transcriptome, epigenome, phenome, single cells, transactional, telehealth, social media). Big datasets have a lot of noise, and we must be able to analyze the individual types of data and put them into higher meta-level structures that will increase the signal and reduce the noise. Then we will be able to integrate these data to make predictive and actionable models to guide and inform health and medicine, he said.
AN INTEGRATIVE SYSTEMS APPROACH TO BIOLOGY, MEDICINE, AND COMPLEXITY
Biological complexity comes from the random and chaotic process of Darwinian evolution. Evolution arises from random mutations, is driven by environmental challenges, and selects solutions building on past successes, Hood said. To understand how a complex system achieves its end goal, one has to define all of the elements present, their interconnectivity, and their dynamics. This is the essence of systems biology.
Hood described his personal involvement in four paradigm changes that led him to the conceptualization of P4 medicine. First, bringing engineering to biology catalyzed high-throughput biology (e.g., instruments for automated sequencing and synthesizing genes and proteins). High--throughput biology, he noted, was the beginning of large datasets in biomedical research. Second, automated sequencing led to the human genome project, which democratized genes (i.e., made them available to all biologists) and created a complete gene “parts list” that was essential for systems biology. Making automated sequencing a reality required a chemist, an engineer, a computer scientist, and a biologist, and Hood realized that the futures of biology and technology were intertwined. For the third
paradigm change, Hood founded the first cross-disciplinary university biology department to couple technology and analytic tool development to leading-edge biology research. Then, in 2000, he launched the first institute for the study of systems biology.
Simply stated, systems biology is a holistic and integrative approach to studying biological complexity, where frontier biology drives new technologies, which in turn catalyze novel domain-driven and data-driven computational tools; systems medicine is the application of the strategies, technologies, and computational tools of systems biology to disease and wellness; and P4 medicine is the clinical application of systems medicine to patients.
An integrated systems approach to disease is essential for dealing with complexity, Hood said, and he elaborated on the five pillars of that philosophy:
1. Viewing biology and medicine as informational sciences is one key to deciphering complexity.
2. Systems biology infrastructure involves a cross-disciplinary culture, democratization of data-generation and data-analysis tools, and the power of model organisms to decipher complexity.
3. Holistic systems experimental approaches enable deep insights into disease mechanisms and new approaches to diagnosis and therapy.
4. Emerging technologies enable large-scale data acquisition and permit exploration of new dimensions of patient data space.
5. Transforming analytic tools allow researchers to decipher the billions of data points for the individual, detailing wellness and disease.
Biology and Medicine as Informational Sciences
Human phenotypes are specified by two types of biological information: the digital information of the genome and the environmental information that modifies this digital information. These interact to generate the phenotype through biological networks that capture, transmit, process, and pass the information on to simple and complex molecular machines that execute the biological functions of life. Both the digital information and the environmental information are needed to understand how a system works. Hood pointed out that all levels of the biological information hierarchy—from DNA to RNA, proteins, interactions and networks, cells, organs, individuals, populations, and ecologies—are modified by environmental signals. If one wanted to understand the system at the level of the
cell, for example, all of the preceding levels of information in the hierarchy would have to be captured and integrated in a way that can elucidate the contributions of the environmental signals in each of those steps.
Each technology will require a domain-driven software pipeline for acquisition, validation, storage, mining, integration, visualization, and modeling of data. Validation1 is key, Hood stressed, and there currently are very few examples of good validation. In addition, different laboratories may not generate data that are interoperable, even if they use the same systems.
Systems Biology Infrastructure
Leading-edge biology research drives the development of technology and computation, facilitating the exploration of data in new dimensions, as well as a cycle of biological information (Figure 3-1). What is required for this cycle to be successful, Hood said, is a cross-disciplinary research environment, including biology, chemistry, computer science, engineering, mathematics, physics, and perhaps other disciplines, working together and communicating effectively in teams. A systems biology approach also facilitates democratization of the tools for data generation and analysis, so that any individual scientist can use them, regardless of the size of the project. This does not mean that every scientist has to learn how to do everything, Hood said, but there should be an environment that creates opportunities.
Holistic Systems Experimental Approaches
The systems approach to biology is holistic, seeking to understand how the individual components assemble and collectively create a functional whole. Hood used the analogy of a radio as a collection of individual transistors assembled onto circuit boards that together convert radio waves into sound waves. Human beings are essentially a collection of biological circuits and networks that process information into health or disease outcomes.
In an experimental systems biology approach, one creates a model from
![]()
1 Validation refers to processes to ensure that the data are accurate and reproducible. Validation may also entail assessing an assay’s measurement performance characteristics to determine the range of conditions under which the assay will give reproducible and accurate data (analytical validation), as well as assessing a test’s ability to accurately and reliably predict the clinically defined disorder or phenotype of interest (clinical/biological validation).
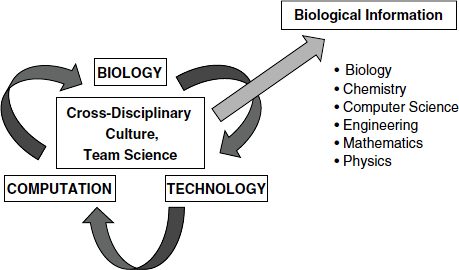
FIGURE 3-1 Systems biology infrastructure.
SOURCE: Hood presentation (February 27, 2012); Hood and Flores, 2012. Reprinted with permission from New Biotechnology.
extant data, formulates a hypothesis, and iteratively tests the model through experimental perturbations of the system. This can be both hypothesis driven and hypothesis generating and can produce large datasets. Experimental systems biology involves global analysis of all components (e.g., DNA, RNA, protein), evaluation of the dynamics of systems (both temporal and spatial), and integration of the different datasets from the system. Hood noted that large datasets are subject to two types of noise: technical noise resulting from how the data are acquired, managed, and analyzed and biological noise that arises as a natural consequence. Subtractive analysis can be used to help minimize biological noise. Ultimately, the goal is to convert data into knowledge.
A Systems View of Disease
A systems view of disease postulates that disease arises when one or more biological networks become perturbed, thereby altering the information they express. Systems medicine, Hood said, is really a network of networks. There are genetic networks at the level of the DNA, molecular networks at the level of proteins and other small molecules, cellular networks, organ networks, and social networks (Figure 3-2). We need to be able, from the various -omics data, to generate networks at each of these
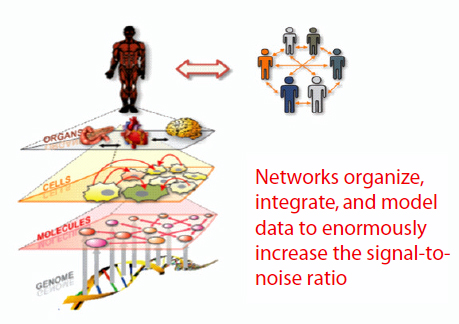
FIGURE 3-2 Systems medicine: A network of networks.
SOURCE: Hood presentation (February 27, 2012); Hood and Flores, 2012. Reprinted with permission from New Biotechnology.
levels and to integrate those networks to begin to understand the system. From a research perspective, networks are powerful ways of organizing, integrating, and modeling data, significantly increasing the signal-to-noise ratio. A systems approach to disease also allows the researcher to follow the disease from inception to end.
As an example, Hood described a systems approach to studying neuro-degeneration in a mouse model. Briefly, inbred mice were injected with infectious prion particles and followed for changes in the transcriptome of their brains relative to the brains of normal littermates. Surprisingly, Hood and colleagues observed that 7,400 genes, or one-third of the mouse genome, were differentially expressed. The researchers then constructed eight combinations of inbred mouse strains and prion strains, carefully designed to exhibit different biologies that could be subtracted away. For example, one construct was a double knockout for the prion gene. When infectious prion particles were injected, the mice did not show signs of disease, but their brains underwent changes. Those changes that were not related to disease could be subtracted away. Subtractive transcriptome analysis based on this and the seven other mouse-prion strain combinations
resulted in 300 differentially expressed genes. Subsequent serial histopathology identified four major disease-perturbed networks that played a major role in this prion disease process, and two-thirds of the genes mapped into these four networks. The remaining 100 genes defined six new networks that were previously unknown participants in prion disease. This is the power of global analyses, Hood said. There was also a sequential disease perturbation of all 10 of these networks, and the combined dynamics of these 10 networks can explain nearly all of the pathophysiology of prion disease (Hwang et al., 2009).
This study provided many new insights into potential biomarkers and diagnostics, Hood said. For example, he and his colleagues were able to demonstrate presymptomatic diagnosis of prion disease. They also established a fingerprint of the normal levels of 100 brain-specific proteins in human blood. The levels of proteins whose cognate networks become perturbed in disease will likely be altered in the blood, resulting in a unique fingerprint for each disease. Hood predicted that organ-specific blood proteins also will have utility for early disease detection, disease stratification, disease progression, following the progress of therapy, and assessing recurrences. Systems-driven blood-based diagnostics will be the key to P4 medicine, he said.
Emerging technologies allow for large-scale data acquisition and analysis. Hood described four technology-driven projects with potential commercial applications on which the Institute for Systems Biology is currently working:
1. Complete genome sequencing of families—integrating genomics and genetics to find disease genes
2. The Human Proteome Project—selected reaction monitoring (SRM) mass spectrometry assays for all human proteins
3. Clinical assays for patients—allowing new dimensions of data space to be explored
4. The Second Human Genome Project—mining all complete human genomes and associated phenotypic or clinical data for the predictive medicine of the future
Family Genome Sequencing
The family genome sequencing project began with a family of two unaffected parents and two children each with genetic disease (Roach et al., 2010). By sequencing the genomes of a family of four and applying the principles of Mendelian genetics, one can identify 70 percent of the sequencing errors in the family, identify rare variants, determine the chromosomal haplotypes, determine the intergenerational mutation rate, and identify candidate genes for simple Mendelian diseases.
Hood predicted that within a decade, the human genome will be a part of every patient’s medical record, and in 5 years, sequencing a human genome could cost as little as $100. He also predicted that any societal and scientific objections to routine genome sequencing will fade as actionable gene variants are identified. Every year, for example, patients could be checked for newly discovered actionable gene variants and provided with information that might be relevant to optimizing their health.
Hood added that the Institute for Systems Biology has developed a variety of software packages to manage the data from the family genomics project.
The Human Proteome Project
The Institute for Systems Biology pioneered four major advances that led to the consideration of a human proteome project, Hood explained. The first advance, the Trans Proteomic Pipeline, is a suite of software programs that validate mass spectrometry data. The software was developed using a bottom-up approach, driven by domain expertise and data. After validation, data are entered into the second new tool, the Peptide Atlas. The third new approach is targeted proteomics, which is the ability to use triple-quadrupole mass spectrometry to analyze 100 to 200 proteins in one hour. Finally, Hood and colleagues created targeted proteomic assays for most of the known 20,333 human proteins, and the results are being cataloged in the SRMAtlas database.
Moving forward, Hood suggested that the key clinical technology is not likely to be mass spectrometry but microfluidic protein chips. The Institute for Systems Biology, in collaboration with Caltech, currently has a prototype chip containing 50 ELISAs (enzyme-linked immunosorbent assays) that can be completed in about 5 minutes using 300 nanoliters of plasma. In the future, Hood would like to be able to assay 2,500 organ-specific blood
proteins (50 from each of 50 organs) from millions of patients and follow them longitudinally to monitor wellness (as opposed to disease assessment) of each of those major organs.
Clinical Assays
Another focus of the Institute for Systems Biology is the development of individual patient information–based clinical assays. There are genomic and proteomic assays in development as well as single-cell analysis and induced pluripotent stem cells (iPS cells) for biology research and diagnostics development.
Single-cell analysis will impact the way we think about cancer, Hood said. For example, he observed quantitative transcriptome clustering of single cells from the human glioblastoma cell line U87. This is in contrast to a whole-tumor sequencing approach, where the signals of the individual cells are averaged and noise is enhanced. The reasons for this cellular heterogeneity are as yet unknown.
One ongoing project that Hood described involves differentiating iPS cells from healthy and diseased individuals into neurons, exposing them to environmental signals, and then using global and single-cell -omics analysis to try to understand the relative contributions of the digital genome and the environmental signals. Another example of the use of iPS cells is the stratification of complex genetic diseases such as Alzheimer’s disease. The project involves creating iPS cells for each individual in the family of an affected individual; differentiating the cells into neurons; conducting single-cell analyses to identify and sort quantized cell states; exposing the sorted neuron populations to environmental probes such as drugs, ligands, or small interfering RNA (siRNA); and analyzing the transcriptome, select proteomes, microRNAome, etc. The intent is to stratify different subtypes of disease-perturbed networks and their response to environmental signals. The substratified populations of Alzheimer’s patient cells could be provided to pharmaceutical companies to test the more than 100 drugs under investigation for Alzheimer’s.
Domain-Driven, Transforming Analytic Tools
The last pillar of the integrative systems approach to disease that Hood described is the development of bottom-up, domain-driven analytic tools that will allow researchers to decipher the billions of data points collected
for an individual and use them to define and understand individual wellness and disease. Hood mentioned The Cancer Genome Atlas (TCGA) as one example of genomic data integration and analysis. There are also efforts toward vertical integration, that is, integration of networks.
Hood advocated an open source platform for the development of software, suggesting that the advantages outweigh the challenges. In addition, he said software development should be driven by users, domain expertise, and data. Because of the complexity of biology, development needs to be bottom-up.
Hood highlighted several challenges to software development, including integrating biological expertise with statistical and computational expertise. Other challenges are integrating individual software packages or modules into coherent platforms for comprehensive modeling and determining the level of granularity of biological information that is needed.
APPLICATIONS OF SYSTEMS MEDICINE: THE P4 APPROACH
Together, systems biology and advanced informatics tools applied to disease and wellness provide the foundation for a personalized approach to medicine, allowing health care to be more predictive, preventive, personalized, and participatory. Hood outlined his perspectives and predictions about how a P4 approach to medicine could look in the future (Box 3-1).
Information Technology for Health Care
Information technology infrastructure is key to the advancement of P4 medicine, Hood said. There will be a need for sufficient infrastructure development and maintenance cycles; storage solutions for the vast amount of genomics, proteomics, and other data; and analytic tools that can access stored data from desktop platforms. Hood reiterated that systems should be open source, extensible, and interoperable, and the development of solutions should be domain expertise driven and data driven (i.e., bottom-up).
Among the technology and informatics challenges Hood listed were the lack of standards for electronic medical information; how to handle conventional medical records and histories as well as molecular, cellular, and phenotypic data; how to identify the actionable gene variants in individual genome sequences; and how to handle the comparative and subtractive analyses of billions of genomes and associated phenotypic data. Such challenges have also been noted by the President’s Council of Advisors on
BOX 3-1
P4 Medicine: Perspectives from Leroy Hood on What the Future Could Hold
Predictive
• In 10 years, most individuals will have had their genome sequenced.
• Genomic information will be used to design probabilistic health history, a lifelong strategy that will optimize individual wellness and manage the potential for disease.
• Individuals will have regular, multiparameter blood measurements done, assaying perhaps up to 2,500 organ-specific blood proteins at once, to predict any transition from health into disease and facilitate a timely response.
Preventive
• Taking a systems medicine approach, therapeutic and preventive drugs and vaccines will be developed to modify disease-perturbed networks so that they operate in a more normal fashion.
• Maintaining wellness (rather than treatment of disease) will increasingly be the focus of medicine.
Personalized
• Individuals are genetically unique, differing by 6 million nucleotides from one another.
• Each patient will serve as his or her own control for analysis of the vast, longitudinal datasets that will be generated.
Participatory
• The patient will become the center of the P4 health care network, and patient-driven social networks for disease and wellness will be a driving force for change.
• Society should be able to access patient data after de--identification and make them available to biologists for pioneering the predictive medicine approaches of the future.
• Patients, physicians, and other members of the health care community will have to be educated about P4 medicine so they can participate fully.
SOURCE: Hood presentation (February 27, 2012).
Science and Technology (PCAST) in its recent report on the use of health information technology to improve health care (PCAST, 2010).
The digital revolution will generate enormous amounts of useful personal data including, for example, imaging data, longitudinal data, and social network data, existing together in a dynamic “network of networks,” Hood concluded. There will have to be ways to integrate all of these data into predictive models and actionable opportunities, he said.
Hood, L., and M. Flores. 2012. A personal view on systems medicine and the emergence of proactive P4 medicine: Predictive, preventive, personalized and participatory. New Biotechnology March 18. [Epub ahead of print].
Hwang, D., I. Y. Lee, H. Yoo, N. Gehlenborg, J. H. Cho, B. Petritis, D. Baxter, R. Pitstick, R. Young, D. Spicer, N. D. Price, J. G. Hohmann, S. J. Dearmond, G. A. Carlson, and L. E. Hood. 2009. A systems approach to prion disease. Molecular Systems Biology 5:252.
PCAST (President’s Council of Advisors on Science and Technology). 2010. Report to the President: Realizing the full potential of health information technology to improve healthcare for all Americans: The path forward. http://www.whitehouse.gov/sites/default/files/microsites/ostp/pcast-health-it-report.pdf (accessed July 10, 2012).
Roach, J. C., G. Glusman, A. F. Smit, C. D. Huff, R. Hubley, P. T. Shannon, L. Rowen, K. P. Pant, N. Goodman, M. Bamshad, J. Shendure, R. Drmanac, L. B. Jorde, L. Hood, and D. J. Galas. 2010. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328(5978):636-639.












