2
Determinants of the Research Value of Biospecimens
Pathology is the study of the structural and functional changes brought about by disease or injury. Pathologists analyze biospecimens for such changes and thereby attempt to discern their causes. The present committee’s statement of task poses a number of questions regarding the future use of the Joint Pathology Center (JPC) repository’s biospecimens collection in clinical care, education, and research activities. This chapter lays the groundwork for addressing those questions by providing information on the means of preserving biospecimens, on methods for analyzing and assessing their research value, and on how the details of preservation, storage, documentation, and the applications for which they are intended may affect prospects for their use. It focuses on scientific and technical considerations; legal and ethical issues are addressed in Chapter 3.
COLLECTION AND PRESERVATION OF BIOSPECIMENS
Three types of biologic material may be collected during pathologic investigations: tissues and cells removed during surgery or obtained specifically for diagnosis via biopsy; cytologic material, including that from fine-needle aspiration biopsy, brushings, or swabs; and whole blood. The main objective in diagnostic pathology and pathology laboratories is to provide accurate diagnosis of a disease and additional pathologic information needed to define a prognosis and determine appropriate therapeutic strategies. Clinical data—including information about the patient and her or his medical history, physical examination, and diagnostic imaging, such as X-rays, CT scans, and the like—are also collected to inform evaluations.
This section briefly addresses how specimens are handled after collection, focusing on the types of samples found in the JPC repository. Figure 2-1 shows the relationship between the various materials collected and their forms of preservation for pathologic analysis.
The pathology workflow for tissues comprises collection or excision from the patient; visual examination of the macroscopic specimen (called a gross specimen); initial stabilization; transfer to a laboratory; selection of material from the gross specimen for further analysis; fixation; further visual examination; histopathologic, biochemical, or molecular analyses; and storage.
After collection or excision and any initial diagnostic evaluation, specimens are typically either frozen or chemically stabilized for transport. The essential processing steps for laboratory preparation of samples that are not maintained in a frozen state are summarized below.
Fixation. Fixative solutions stabilize tissue structure and biochemical constituents by coagulating (cross-linking, denaturing, and precipitating) proteins and thereby prevent cellular hydrolytic enzymes, which are released when cells die, from degrading tissue components and rendering tissues inadequate for microscopy. Fixation also immobilizes fats and carbohydrates, reduces or eliminates enzymatic and immunologic reactivity,
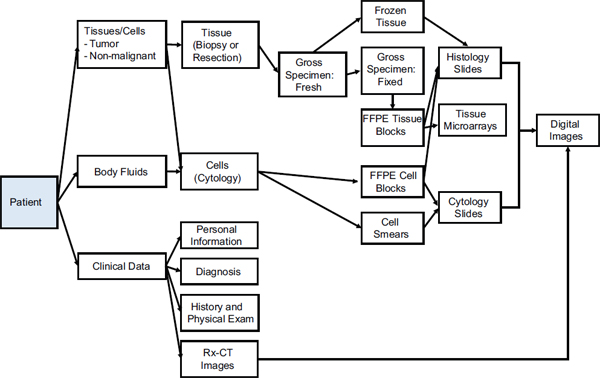
FIGURE 2-1 Relationship between pathologic materials collected and form in which they are preserved for analysis.
and kills microorganisms that are present in tissues. The fixative routinely used in pathology is 10 percent neutral buffered formalin, a buffered aqueous solution of formaldehyde. Fixation yields “wet tissue” that is either stored in an air-tight container or processed further as delineated below.
Embedding. Specimen water (about 70 percent of tissue mass) is replaced with paraffin wax, and the specimen is surrounded by paraffin in a mold to provide support during sectioning and to aid in preservation. Formalin-fixed paraffin-embedded (FFPE) tissue is one of the predominant forms in which pathologic specimens are stored.
Sectioning. Sections are cut on a microtome, which has a blade similar to a single-edge razor blade, that is advanced through a block of paraffin-embedded tissue. The shavings—about 5 µm thick, about twice the thickness of a human hair—are placed in water, and the floating shavings are picked up on 1 × 3-in. glass slides. A given tissue block can be recut many times, although specific slices may differ in the amounts and types of tissues (for example, primary tumor vs. normal tissue) present.
Staining. Tissue components can be distinguished with selective absorption of dyes to facilitate viewing under a microscope. The stain routinely used in histology is hematoxylin and eosin. There are special methods for highlighting components (such as microorganisms) that do not stain well with the customary preparations. Pathologists commonly use tissue sections prepared in this manner in their analyses.
Researchers use tissue microarrays (TMAs) constructed from diagnostic blocks of FFPE tissue to permit simultaneous evaluation of expression of specific pathologic features—proteins by immunohistochemistry, for example—in hundreds of individual tissue samples from different patients on a single slide (Rimm et al., 2011; Voduc et al., 2008). TMAs are assembled by using a needle to core an FFPE tissue block and extract a 0.5- to 2.0-mm piece that is placed into a predrilled master paraffin block that may contain up to 400 cores. Sections from the resulting block may be cut with a microtome, placed on a slide, stained, and analyzed. In cancer research, TMAs are used to analyze the frequency of a molecular alteration in different tumor subtypes, as detected by immunohistochemical and molecular techniques, to enable evaluation of potential diagnostic and prognostic markers by correlating staining patterns with light microscopy and clinical information, which may also contain outcome measures (Camp et al., 2008; Kapur, 2011). Advantages of using TMAs include minimal tissue use, lower reagent costs, faster results, and the ability to define a set of cases that
have related diseases and clinical annotations and to use many such cases in direct parallel analysis on the same slide.
Advances in technology also permit high-fidelity capture of the information contained on slides and in diagnostic images in digital form. Digitization removes the opportunity for further biologic analysis but facilitates storage, sharing, and, if desired, deidentification of specimens.
Educational, clinical consultation, and research uses impose different demands on the physical state of a specimen and the documentation that accompanies it. This section presents a brief summary of the considerations that influence the assessment of fitness for those uses.
Educational Uses
Educational uses have the lowest bar for molecular quality and physical integrity of material and therefore can, in principle, permit the greatest variety of samples. However, it is uncommon to keep fresh, frozen, or even preserved samples in a manner that would allow ready distribution beyond the location at which they were generated. Indeed, unfixed tissue poses a risk of infection and should be handled only by persons who are trained in handling potentially infectious agents. The risk associated with fixed tissues is much lower, but such agents as prions are not inactivated by normal fixation methods. Fixed specimens can be encased in plastic to facilitate handling, eliminate the risk of contagion, and enhance the educational value, but this is not commonly done. For those reasons, in most contexts, gross specimens are likely to have their most effective and widespread educational use in image form.
Microscope slides are extremely useful for education, are easy to ship and return, and carry a very low risk of contagion. They have a long but finite shelf-life. However, there is little need for samples of extremely rare entities, except in advanced residency and fellowship training, and slides of common entities are abundantly available. The current trend is toward archiving of teaching slides digitally and viewing them with virtual microscopy. That avoids the problems of image or slide degradation and of slide distribution. Digital microscopy allows a teacher and a student to collaborate as though they are at the same microscope even though they may be separated by large distances.
Accompanying clinical information often improves the pedagogic value of specimens, but it is not always required.
Clinical Consultation Uses
Clinical consultation was a major function of the Armed Forces Institute of Pathology, and the JPC continues to serve this function for the Military Health System, the Department of Defense, and other federal agencies. In the past, it was common to limit the materials for consultation to stained microscopic sections. Advances in diagnostic procedures now require the ability to extend the histopathologic description with immunohistochemical and molecular probes. That may require that unstained slides be included in the materials for consultation and, less often, that the tissue block or fresh-frozen tissue be available. Consultation also requires that detailed clinical information be provided to the consulting physician. Large medical centers can often carry out the more advanced procedures on their own and need only send appropriately stained slides to the consultant, whereas smaller centers might require that the consultant carry out the procedures. In general, fresher, unfixed tissues give better results in assays than do samples that have had longer intervals at room temperature or long periods in fixatives. When consultation is limited to examination of slides, digital (or “virtual”) microscopy via scanning of glass slides (Pantanowitz et al., 2011) allows rapid, interactive consultation without the need to transport and retrieve slides.
Research Uses
Research comprises a broad array of activities and a correspondingly broad array of requirements for pathologic specimens. Case reports and historical studies might need slides alone and be limited only by the condition of the slides, but more extensive studies of the mechanism of disease require the freshest materials possible with cryopreservation, snap freezing in liquid nitrogen, or relatively brief times in fixatives to obtain the best results. That means not that older materials or materials that have not been obtained under those conditions are without value—DNA sequences have been obtained even from Neanderthal bones—but simply that they make studies technically more difficult and limited in scope and preclude some studies as the signal-to-noise ratio tilts toward the noise.
In most cases, the more complete the accompanying clinical information is, the more valuable the sample. Nonetheless, many otherwise undocumented samples that have been well characterized histopathologically can be of value in genetic, microRNA (miRNA), and other functional studies. Studying diseases that have a high incidence, such as breast cancer, need not depend on suboptimal tissues or special collections inasmuch as specimens are readily available, whereas studying rare diseases may require greater
compromises with respect to sample quality and rely critically on special collections and repositories.
Limitations on the use of pathologic samples in research are addressed in greater detail later in this chapter.
TECHNOLOGIES USED TO MANAGE SPECIMEN ACQUISITION AND MANAGEMENT
Over the last two decades, technology development has created both challenges to and opportunities for human biospecimen resources. The sensitivity, specificity, multiplexing capability, and speed of operation of technologies for molecular analysis of all classes of biomolecules in human specimens have undergone transformative improvements, and further refinements are developed at ever-increasing rates. However, this greatly augmented analytic power has raised the bar for the quality of the biospecimens that serve as the source of analytes for the technology platforms, which are increasingly used in research and clinical care. Biospecimens acquired in the setting of standard clinical practice that formerly served as adequate sources of research material despite the varied, undocumented, and uncontrolled sources of preanalytic variation1 to which they were exposed are no longer adequate, let alone optimal, for new molecular-assessment platforms. Technology development specifically directed to the challenges related to biorepository operation in this environment has been essential in addressing the gap between the demand for and supply of high-quality human specimens for molecular research.
Biorepositories, such as the JPC, that comprise clinically derived samples collected in diverse settings and referred for pathologic consultation on disease, typically face greater complexities in ensuring that their collections meet quality standards for sensitive analytic platforms than do biobanks that were established for the express purpose of collecting specimens for research. Clinical consultation biorepositories have difficulty in controlling, recording, or assessing the sources of preanalytic variation that may compromise the molecular quality of their collections. Thus, technologic solutions for controlling processing and environmental variation and for assessing the molecular quality of processed or stored specimens have been essential for the continued evolution and usefulness of clinical consultation biorepositories for biomedical research.
___________________
1Preanalytic variation refers to any of the many biospecimen acquisition, handling, or processing procedures and environmental characteristics (such as temperature and humidity) to which a specimen may be exposed before analysis takes place. Preanalytic variation may alter the molecular quality or composition of a biospecimen and render it unsuitable for a specific type of analysis.
Technologic solutions for the problems in specimen acquisition, preservation, and management include the following:
- Shipping technologies that maintain specific environments during transport of samples and thus help to maintain the molecular quality of the samples for analysis at remote sites.
- Specimen fixation and other stabilization technologies that preserve the quality of labile biomolecules, allow optimal molecular preservation and histopathologic quality, or allow in situ stabilization of the specimen to preclude preanalytic variation incurred during specimen acquisition (for example, through surgical resection and pathologic handling).
- Information technology solutions that allow annotation of specimen collections with clinical data about the individual from whom the specimen is derived; consent to specimen collection, transfer, storage, and use; authorization of use of protected health information; pathologic data on the specimen (such as gross description and accompanying diagnosis); collection, processing, transportation, and storage data; quality-control data; specimen analysis data; radiologic imaging data; inventory tracking; overall (system-wide) quality-management data; and molecular analysis data.
- Specimen or molecular storage technologies, such as ambient-temperature (“dry-state”) storage.
- Molecular qualityassessment technologies for RNA, DNA, and proteins.
- Digital imaging and image analysis technologies for precise structure-based data associated with each specimen.
Those categories of technologies are not all equally developed, but all continue to improve rapidly and decrease in cost. Nevertheless, the rate of obsolescence of many of the technologies and the increasing knowledge of the effects of specific preanalytic factors on molecular analysis data require continual reassessment of the adequacy and functionality of technologies that are in place in light of the repository’s mission.
TECHNOLOGIES USED TO ANALYZE SPECIMENS
Researchers have several tools at their disposal for deriving clinical and research information from specimens. This section—which is based on review articles by West (2010) and Beck and colleagues (2010)—identifies some of the technologies used in analysis and discusses how preservation technique influences the ability to perform various types of analysis.
Table 2-1 summarizes the results of some recent research on the influ-
TABLE 2-1 Recent Research on the Influence of Preservation Method on Specimen Analysis Outcomes
| Collection or Preservation Methods | Tissue Types | Attributes |
| Fresh-frozen (Snap-frozen) | Pancreatic cancer specimens | RNA integrity was determined with microcapillary electrophoresis, using RNA integrity number (RIN) algorithm and results of laser-capture microdissection (LCM). Various ex-vivo procurement times (up to 10 min, 11–30 min, 31–60 min, over 1 h); banked over three periods (2001–2004, 2004–2006, 2006–2008) |
| Fresh-frozen | Invasive breast cancer tissues | Manual method: subjective evaluation of electropherogram; ratio method: ratio between 28S and 18S peaks; RIN |
| Room temperature, iced, saline solution, RNA-stabilizing buffer, snap-frozen (after 0.5, 1, 3, 6, 16 h) | Normal tonsil Normal colon | Structural RNA integrity via microchip electrophoresis |
| Snap-frozen (unfixed and immersed in RNA-stabilizing buffer), thawed for 0, 5, and 45 min, 1, 3, 6, and 16 h | Tonsil | Microchip gel electrophoresis and gene expression level via PCR |
| Snap-frozen, formalin-fixed paraffin-embedded (FFPE) | RNA quality | |
| FFPE | Diffuse large B-cell lymphoma | Gene expression |
| Effects | Source |
|
• 42 percent of human pancreas cancer specimens banked under a dedicated protocol yielded RNA with a RIN of ≥7 • Brief warm ex-vivo ischemia times did not adversely affect RNA quality (percentage of tissue with total RNA with RIN of ≥7 for ≤10 min, 42 percent; 11–30 min, 58 percent; 31–60 min, 33 percent; >60 min, 42 percent) • Long-term storage of banked pancreas cancer biospecimens did not adversely affect RNA quality (total RNA with RIN of ≥7 banked in 2001–2004, 44 percent; 2004–2006, 38 percent; 2006–2008, 50 percent); RNA retrieved from pancreatic cancer samples with RIN of C7 subject to LCM yielded RNA suitable for further downstream applications • Fresh-frozen pancreas tissue banked according to a standardized research protocol yields high-quality RNA in about 50 percent of specimens and can be used for enrichment with LCM; quality of tissues in the biobank was not adversely affected by slight variations in warm-ischemia times or different storage periods |
Rudloff et al., 2010 |
|
• Comparison between RNA quality (RIN) and gene expression analysis shows dense clustering of high-quality samples but weak clustering of low-quality samples • Manual and RIN methods are superior to ratio method |
Strand et al., 2007 |
|
• RNA stable in both tissues under all conditions for up to 6–16 h • Expression levels essentially stable when samples kept on ice • Marked regulation of single genes observed during room-temperature storage in normal saline and RNA-stabilizing buffer • RNA from 54 of 47 samples had proper ribosomal peaks • Nonfixed specimens may be transported on ice for hours with minimal influence |
Micke et al., 2006 |
|
• Minimal RNA degradation after 30 min • Relevant changes in some gene-expression levels at 45 min • Repetitive thawing cycles had similar effects on RNA integrity • Incubation in RNA-stabilizing buffer prevents RNA degradation |
Botling et al., 2009 |
|
• Introduced heating into extraction protocol to improve quality; incubation at 70°C for 20 min was applied to disrupt cross-links in FFPE without compromising RNA integrity • TaqMan detection influenced by master mix, amplicon size, and use of preamplification step • Comparable results in frozen and FFPE tissue |
Li et al., 2007 |
|
• Provided PCR protocol for gene-expression analysis • 62 of 65 samples “successfully” analyzed |
Votavová et al., 2009 |
| Collection or Preservation Methods | Tissue Types | Attributes |
| FFPE | Parathyroid | Proteome quality |
| Fresh-frozen, FFPE | Colon adenoma | Proteome quality Liquid chromatography |
| Frozen, FFPE | Frozen/optimal cutting temperature (OCT)-embedded livers (rats) | Proteome quality Liquid chromatography |
| Effects | Source |
|
• 163 unique proteins identified via mass spectrometry • Similar results via sodium dodecyl sulfate (SDS)-out in gel-free method • Antigenicity not always preserved in Western blot • Despite some limitations due to extensive formalin-induced covalent cross-linking, results suggest that FFPE extracts may be an alternative source for large-cohort samples when frozen samples are unavailable |
Donadio et al., 2011 |
|
• “The major difference between frozen and FFPE proteomes was a decrease in the proportions of lysine C-terminal to arginine C-terminal peptides observed, but these differences had little effect on the proteins identified.” • “Analysis of archival colon adenoma FFPE specimens indicated equivalent numbers of MS/MS spectral counts and protein group identifications from specimens stored for 1, 3, 5, and 10 years.” • “Analysis of the combined frozen and FFPE data showed a 92 percent overlap in the protein groups identified. Comparison of gene ontology categories of identified proteins revealed no bias in protein identification based on subcellular localization.” • “Archival samples displayed a modest increase in methionine oxidation, from approximately 17 percent after one year of storage to approximately 25 percent after 10 years.” • “These data demonstrate the equivalence of proteome inventories obtained from FFPE and frozen tissue specimens and provide support for retrospective proteomic analysis of FFPE tissues for biomarker discovery.” |
Sprung et al., 2009 |
|
• “Comparable molecular mass representation was found in extracts from FFPE and OCT-frozen tissue sections, whereas protein yields were slightly less for the FFPE sample.” • “The numbers of shared proteins identified indicated that robust proteomic representation from FFPE tissue and LCM [laser capture microdissection] did not negatively affect the number of identified proteins from either OCT-frozen or FFPE samples.” • “Subcellular representation in FFPE samples was similar to OCT-frozen, with predominantly cytoplasmic proteins identified. Biologically relevant protein changes were detected in atorvastatintreated FFPE liver samples, and selected atorvastatin-related proteins identified by MS were confirmed by Western blot analysis. These findings demonstrate that formalin fixation, paraffin processing, and LCM do not negatively impact protein quality and quantity as determined by MS and that FFPE samples are amenable to global proteomic analysis.” |
Scicchitano et al., 2009 |
| Collection or Preservation Methods | Tissue Types | Attributes |
| FFPE | Heart tissue (mice) | Proteome quality |
| FFPE | Liver (mouse) | Proteome quality |
| FFPE (different temperatures [4, 20–25, 37°C] and storage times [0–12 months]) | Liver, kidney, heart, brain, lung, spleen (rat) | RNA quality |
| FFPE (different fixation periods) | RNA quality | |
ence of preservation method on specimen analysis outcomes. It is intended not as a comprehensive survey of the literature but as an illustration of work in this field.
Protein Expression
Immunohistochemistry
Most gene-expression profiling studies aim to address clinical questions with biologic insight. Conventional gene-expression profiling, in which thousands of measurements are made, is not yet an efficient clinical tool. It is expensive, is technically demanding, and requires arduous tissue-handling protocols for optimal results (for example, rapidly freezing fresh tissue and maintaining it frozen at well-controlled, very low temperatures). However, such approaches as multiplex polymerase chain reaction (PCR) are emerging as useful options (Parker et al., 2009). Such techniques offer important
| Effects | Source |
|
• “Incubation of tissue sections at high temperature with a novel extraction buffer . . . resulted in improved protein recovery.” • “This is an indication of the formation of protein-protein complexes by cross-linking, and of protein fragmentation due to prolonged sample storage.” |
Azimzadeh et al., 2010 |
|
• “It was found that incubation of tissue in a lysis buffer containing 6 M guanidine hydrochloride at high temperature led to the highest protein yield and the largest number of proteins identified. The peptides and proteins identified from formalin-fixed tissue were first comprehensively compared with those identified from frozen-fresh tissue. It was found that a majority of peptides identified from fixed tissue were unmodified and proteome coverage for the analysis of fixed tissue was not obviously compromised by the formalin fixation process.” |
Jiang et al., 2007 |
|
• RIN 7 for 1–3 days of storage at 4°C • RIN 5–6 for 1 year at 4°C • 20°C and above yielded poorer results (and poor RNA amplification) • RNA quality not adversely affected by long interaction with fixative • RT–PCR quality is affected by long interaction with fixative • Sample size influences quality: the thicker the sample, the longer it takes for fixative penetration and the lower the RNA quality; similarly for RT–PCR |
von Ahlfen et al., 2007 |
|
• Optimal fixation period 12–24 h, yielded best RNA. |
Chung et al., 2008 |
improvements in reproducibility and dynamic range, but more traditional immunohistochemistry for protein detection still has a great role in diagnostic pathology.
A strength of immunohistochemistry is the ability to perform single-cell identification and functional analysis in the context of an archival specimen. Immunohistochemistry is robust for use with a wide variety of materials obtained for pathologic analysis. Its utility is independent of the size of the specimen, working well with very small biopsies, and is often robust in non-ideal conditions. For example, the method can often identify signal in tissue that shows extensive necrosis or is contaminated with normal, inflammatory, or cancer tissue. Although it has been in clinical use for decades (Warnke et al., 1983), the field of immunohistochemistry is not stagnant. For example, a technique that uses multiple antibody stains on a single slide with different reporter dyes has been developed clinically in the last decade and now has a number of clinical applications (West, 2010). These multidimensional assays can be useful in identifying relationships between different cell types (such
as breast luminal and myoepithelial cells) or distinguishing a subgroup of a single cell type (such as proliferating lymphocytes).
Immunohistochemistry requires expert interpretation, though, and this can lead to variations in results from one laboratory to another. Another limitation of the technique is that it can be difficult to control for variation in tissue quality or stain quality. That can be a problem in dealing with clinical specimens because ischemic time, fixation time, and variations in tissue processing can decrease signal-to-noise ratio and may lead to false-negative or false-positive results. Morphology often supplies the necessary backup to a single protein biomarker but requires expert interpretation. Although poor tissue preservation is easily recognizable when underfixation is the problem, the more subtle tissue degradation due to overfixation—which preserves tissue architecture but destroys the macromolecules because of extensive cross-linking—may not be evident in morphology alone (Werner et al., 2000).
Immunofluorescence
Immunohistochemistry using immunofluorescence is an emerging technology for protein detection that is likely to have an important impact on both medical clinical work and research. Several reports of improved immunofluorescence in FFPE sections have been published (Bataille et al., 2006; Ferri et al., 1997; Mason et al., 2000; Niki et al., 2004), but the techniques have yet to become widely used because of the dearth of the needed specialized microscopic technology in most laboratories and the lack of operators who are comfortable with the techniques and their challenges. Difficult features in FFPE material, such as autofluorescence, can be reduced with the use of new reagents, such as Sudan Black B. New tools, such as confocal microscopes, can also largely reduce the problem of autofluorescence. Data obtained with these techniques are likely to be incorporated into the clinical workflow as image analysis migrates from the microscope to the computer.
Proteome Analysis
The emergence of proteomic methods has enabled researchers to interrogate expressed proteins from a number of different tissue types systematically. That allows exploratory studies as opposed to studies that depend on prior knowledge of the proteins that are being studied, as do immunohistochemical studies (Hanash et al., 2008). Proteomic methods generally require high-quality tissue because changes in protein composition can lead to difficulties with identification of specific peptides due to the features of mass spectrometry (Aebersold and Mann, 2003).
The study of proteins extracted from FFPE material is particularly challenging owing to intraprotein and interprotein cross-linking that results
from the interaction with formaldehyde, the active ingredient in formalin. This issue is difficult to bypass in that the practical goal of fixation is to preserve the morphologic features of the tissue for pathologic analysis. The preservation is achieved by strengthening the integrity of the tissue with fixation. The protein cross-linking properties of formaldehyde are well suited for this whereas other fixatives that are more gentle on protein quality fail to preserve tissue structure to the standards generally expected in patient-care settings. Recent progress has been made in using FFPE tissue for proteomics. A number of groups have used steps to reverse the covalent interactions caused by formaldehyde cross-linking, including incubation at high temperatures (Prieto et al., 2005; Shi et al., 2006). These methods have allowed novel protein identifications in FFPE material.
Gene-Expression Profiling
Since the method of gene-expression microarrays was developed in the mid-1990s, genomewide expression profiling has been used widely in research (Janssens and van Duijn, 2008; Kraft et al., 2009; van der Net et al., 2009; Xu et al., 2009). Experiments with gene-expression profiling have led to important advances in our understanding of a wide variety of human conditions, but research efforts with clinically derived specimens have been frustrated by lack of specimens amenable to the biochemistry of the technique. A major obstacle to the translation of gene-expression profiling to specimens gathered for clinical purposes has been the fact that such analyses are best performed on fresh-frozen tissue, and few specimens are stored as fresh-frozen. Indeed, essentially all clinical tumor specimens are stored as FFPE (Hewitt et al., 2008). That fixation and storage technique results in extensive RNA fragmentation and alteration of hybridization qualities (Hewitt et al., 2008). Several groups have attempted to use clinical specimens in FFPE for gene-expression profiling with microarrays (Coudry et al., 2007; Farragher et al., 2008; Frank et al., 2007; Linton et al., 2008; Penland et al., 2007; Scicchitano et al., 2009) with less than ideal results.
Multiplex Real-Time Polymerase Chain Reaction2
Multiplex PCR has recently been shown to be robust in archival material. It is able to provide quantitative information compared with the typically qualitative information supplied with immunohistochemistry. The clinical utility of multiplex PCR has been shown in connection with a number of diseases, such as breast cancer. A multiplex kit is already available for clinical use in the quantitative measurement of prognosis of
___________________
2PCR is a technique used to reproduce pieces of DNA or RNA for analysis.
primary breast cancers. In addition, multiplex real-time (RT) PCR has been demonstrated to outperform immunohistochemistry in the characterization of breast carcinomas into their molecular subtypes (Parker et al., 2009).
cDNA-Mediated Annealing, Selection, Extension, and Ligation
Multiplex RT–CR has been shown to be useful in analyzing the expression levels of tens of genes. Other molecular approaches allow the analysis of thousands of genes. One of these is complementary DNA-mediated annealing, selection, extension, and ligation (DASL) (Ravo et al., 2008), which uses a two-probe oligonucleotide pair to measure each complementary DNA (cDNA) target. DASL uses two small oligonucleotides placed 1−20 nucleotides apart on the annealed target cDNA. Extension and ligation steps fuse these two oligonucleotides and the ligation event indicates identification of the presence of the target RNA. The technique provides high sensitivity and accuracy for measuring RNA expression in archival material. One major drawback is that the probe pairs must be synthesized, so few gene probe pairs (typically fewer than 10,000) are available for target interrogation (West, 2010).
RNA-Seq for Archival Material: 3SEQ and SAGE-Seq
RNAseq refers to methods that use high-throughput “next-generation” sequencing to produce reads of transcripts from each gene in a sample. A typical RNA-seq experiment generates tens of millions of sequencing reads. An expression level of each gene can be generated by counting how many reads have mapped to any given transcript.
Two novel RNA-seq protocols—3′-end sequencing for expression quantification (3SEQ) and serial analysis of gene expression-seq (SAGE-seq)—have been designed to work with mRNA extracted from FFPE tissue (Beck et al., 2010; Wu et al., 2010). As noted above, RNA purified from archival tissue does not work well with microarray methods. However, the small RNA fragments purified from archival tissue, typically 100−500 nucleotides, are well suited for small libraries for short-read (50−100 nt) sequencing.
Degraded mRNA from FFPE material can be purified with polyA selection, and reads can then be obtained from the ends (either single or paired) of the fragments. The reads are short but contain more than enough unique sequence to be confidently mapped to the reference genome. Transcript counts can thus be obtained in that the reads usually are in the three-prime untranslated region (3′-UTR) abutting the polyA tail, which leads to a convenient aggregation of signal that increases the power of the later statistical analysis. An important advantage over microarray profiling, which predefines the transcripts that will be measured by choices made in
the spotting of sequences in the array construction, is that the 3SEQ and SAGE-Seq methods generate reads from all polyA RNA molecules, including those from unannotated genes and transcripts and alternative 3′-UTRs.
RNA in situ Hybridization
An alternative method for evaluating gene expression in paraffin-embedded tissue is to use RNA in situ hybridization (ISH). The ISH technique is particularly useful for the examination of newly found genes of interest. Rather than waiting 4−6 months for a conventional antipeptide antiserum that has a success rate of 5–10 percent, ISH probes can be generated in less than a month. A number of methods are being developed. The ISH technique for archival material used most widely was developed by St. Croix and colleagues (2000) and Iacobuzio-Donahue and colleagues (2003). It uses RNA probes that are 400−600 nucleotides long. The signal is amplified with a tyramide-based system, and this is followed by development with traditional markers, either chromogenic or fluorescent substrates. The advantage of RNA ISH probes over antisera or antibodies is that one can include so-called sense strands or mis-sense probes as controls (Clarke and Shimono, 2011). Moreover, specificity of the probes is highly likely inasmuch as the probes are designed to hybridize with the 3′-UTR of each gene, an area that is highly specific for individual genes.
Despite the capabilities of ISH, immunohistochemistry appears to be more useful with very old specimens; RNA in specimens more than 20 years old appears to be less well preserved for detection with ISH. Although antigens for immunohistochemistry diminish over time, they are more resistant to deterioration than is mRNA. This issue becomes important for clinical studies that use cases with long followup.
miRNA Studies
In contrast with DNA and mRNA, the quality of miRNA does not change substantially with fixation. The key feature is the length of the miRNA molecules. These molecules are typically quite short, averaging 22 nucleotides, and so are less susceptible to fragmentation by formaldehyde (Li et al., 2007; Liu et al., 2009). The isolation of mRNA compared with miRNA can be complicated by substantial RNA degradation.
DNA Studies
Many of the studies performed with RNA can be applied equally to DNA, but high-throughput sequencing is an emerging technique specifically for use with archival DNA. The presence of mutations in multiple
cancer-related genes is being applied to DNA extracted from FFPE tumor-tissue specimens with PCR-based approaches. Recent advances in high-throughput next-generation sequencing methods permit the analysis of mutations in 200–400 genes from small amounts of FFPE-extracted DNA. DNA, in general, is much more stable that RNA. However, all nucleic acids are susceptible to depurination in highly acidic environments, and the preservation of DNA in archival tissues depends heavily on the quality of the fixative used. Depurination can lead to strand cleavage, which would create problems for many of the modern molecular techniques. Thus, if unbuffered formalin was used originally to fix the archival material—as might be the case for older specimens in the JPC repository—it could lead to substantial degradation of the DNA (Akalu and Reichardt, 1999; Bonin et al., 2010). There are few published data on the quality of reads generated from high-throughput sequencing of archival DNA.
Elemental and Chemical Studies
Tissues are routinely analyzed for trace minerals in clinical pathology laboratories for diagnostic purposes. Trace minerals are preserved and can easily be localized and measured in FFPE tissues. For example, hepatic iron concentrations are essentially the same whether assessed in fresh tissues or in paraffin-embedded tissues (Torbenson, 2011). However, the alterations in tissue quality, such as a change in weight due to the replacement of water with less-dense paraffin, can necessitate correction factors (Bischoff et al., 2008). The routine fixation process, with numerous alcohol washes to dehydrate the tissue, results in the removal of many lipophilic small molecules. Elemental concentrations were studied in the past largely with atomic absorption spectroscopy or energy-dispersive X-ray analysis, but new methods are available now (Becker and Jakubowski, 2009; Harrington et al., 2010; Mizuhira et al., 2000).
LIMITATIONS IN THE USE OF PATHOLOGIC SAMPLES IN RESEARCH
Several factors influence whether pathologic samples obtained for clinical consultation purposes will be fit for use in research. This section identifies the major issues that limit the research use of such specimens and addresses how the details of collection, preservation, storage, and documentation can affect fitness for research use.
The specimen preservation–stabilization process might be inappropriate for or incompatible with the technologic or scientific demands of the research analysis.
The suitability of any specific specimen for analysis depends on the analyte that is to be assessed and the technique to be used for the analysis—that is, whether it is “fit for purpose.” As previously noted, the most common stabilization method used in pathologic practice is 10 percent neutral buffered-formalin fixation followed by dehydration with graded alcohols, xylene exchange, impregnation by liquid paraffin, and permanent preservation in paraffin. While the paraffin is maintained in liquid state, the specimen is spatially oriented in a small, thin rectangular container and then rapidly cooled until the paraffin is solidified. Adequacy of fixation of a pathologic specimen is determined largely according to the dimensions of the tissue (when it is submerged in formalin in the gross state and when it is sampled for single aliquots to be individually processed into tissue blocks, formalin penetrates about 1 cm into tissue) and the total time in fixative. In general, inadequate preservation due to underfixation is far more deleterious to samples than is prolonged fixation. Both fixation in a molecular cross-linking fixative, such as formalin, and paraffin embedding with exposure to high temperatures may compromise the molecular quality of samples.
Other related issues concern variations in concentration, pH, or buffering of formalin or the use of fixatives other than 10 percent neutral buffered formalin. The optimal formaldehyde concentration in a fixative and the optimal pH of the solution may depend on the biomolecule of interest, especially in the case of immunohistochemical analysis. Practice is not standardized, and these measures are not always recorded in pathology reports and thus represent important unknowns in research. In the past, the use of such fixatives as Bouins solution (which contained picric and glacial acetic acid with formaldehyde) or special hematopathologic fixatives that contained metals, such as B5 zinc or B5 mercuric chloride, was common, and the fixative type was sometimes not recorded as a part of the pathologic record. Many of those fixatives are incompatible with modern molecular-analysis platforms. When specimens for research are accrued from multiple institutions that use different stabilization and preservation practices, data from molecular analyses may be neither reliable nor comparable or institution-specific batch effects may be seen.
Important variation in pathologic processing also occurs before stabilization. For example, the length of time that elapses between specimen removal from a patient and specimen fixation (“time to fixation” or “cold ischemia time”) and the conditions to which the specimen is exposed during this period, such as desiccation or various room temperatures (as opposed to refrigeration), may seriously alter both the molecular quality and the molecular content of the specimen and thus create artifacts that may be misinterpreted as reflecting disease biology. In some cases, the variations may even compromise histologic quality. Highly labile biomolecules may
be lost altogether as time to fixation increases. Those factors are neither controlled nor recorded in common clinical practice. Rapid stabilization of tissue with cryopreservation of various kinds (for example, immersion in liquid nitrogen, freezing in the vapor phase of liquid nitrogen before immersion, immersion in dry ice–isopentane slush, embedding in a freezing medium based on polyethylene glycol–sucrose,3 and placement in a −80°C freezer or a cryostat) may be used in exceptional circumstances but are not the standard of care for all tissues. Thus, if frozen (unfixed) tissue is needed for research, this requirement often cannot be met pathologic tissue. If viable cells or tissues are required for research, pathologic samples cannot fulfill this requirement unless viable aliquots are preserved appropriately at the moment of acquisition of the specimen and not used as a part of a retrospective pathologic sample set.
Sample aliquot content might be unknown to the investigator, and too little of the lesion of research interest might remain in the residual tissue after clinical workup.
Each paraffin tissue block processed for a case is thinly sliced (at 5-µm intervals) with a microtome to produce sections that are placed on glass microscope slides and stained for histopathologic analysis under a light microscope. As previously noted, the standard histopathologic stain is hematoxylin and eosin, but a wide variety of stains may be used on adjacent tissue sections to reveal special features as a part of the pathologic workup. Unstained sections also are often cut and set aside for or used for immunohistochemical analyses of various types required for pathologic diagnosis and characterization. In many cases, small amounts of tissue may remain in a block after complete workup, and this can result in inadequate residual amounts for research. Inadequacy of residual tissue for research is particularly problematic if the original specimen from the patient is very small or the lesion of interest is small, focal, or both and thus not of sufficient quality (or, indeed, at present all) in the residual tissue in a block. The lack of quality control of pathologic tissue blocks made available for research, to verify the nature and content of the remaining tissue, can detract from their usefulness for research. It can also skew data from investigational studies if the residual blocks are assumed to be representative of the overall diagnosis but are depleted of diagnostic tissue.
The original pathologic diagnosis might be unconfirmed or incorrect.
Unconfirmed or incorrect diagnosis of pathologic material is uncommon (Lind et al., 1995; Ramsey and Gallagher, 1992; Renshaw et al., 2003a,b; Safrin and Bark, 1993; Wakely et al., 1998), but in cases of rare diseases or
___________________
3Also known as optimal cutting temperature (OCT) compound.
diseases that have a documented, notoriously high degree of interobserver variation in diagnostic interpretation, it can be an important issue for both patient management and research. In such cases, it is considered standard of care to seek a second, specialty pathologic opinion, such as the consultative opinions produced for specimens submitted to the JPC repository and its predecessors. Verification of diagnosis by obtaining a new pathologic analysis in every case used for research is recommended.
However, it may also be problematic for research if the diagnosis that accompanies the case is outdated and no longer appropriately classifies the disease. Outdated diagnostic terminology (correct diagnosis but arcane language) or outdated diagnostic criteria for identification (classification according to a schema that is no longer in use) may cause considerable difficulty in mapping a historical case to a current diagnostic category accurately. Over the span of decades that the JPC repository has existed, pathologic classification of disease has evolved substantially. In some disease categories, such a hematopathologic malignancies, entire disease classifications have changed repeatedly because knowledge of pathogenesis has grown. Modern diagnosis of lymphoma and leukemia may require delineation of specific molecular features that were never tested for in older cases. Depending on the specific preanalytic variation associated with a historical case, it may not even be possible to test accurately for the molecular features required for diagnostic classification.
The more common problem in all disease categories is the lack of standardization in diagnostic terminology that was widespread in pathology for many years. That has been exemplified both by the use of a given diagnostic term for different disease entities and by the use of multiple diagnostic terms for a given disease entity (Cooper, 2006). A researcher using a historical case may not have the requisite expertise to interpret the existing diagnostic terminology accurately and map it to the current diagnostic terminology standard correctly. Failure to reclassify cases correctly according to current diagnostic standards and current diagnostic terminology for any of the above reasons may skew research data.
Preanalytic variations related to preoperative or intraoperative factors may create molecular artifacts.
Many drugs used in preoerative and intraoperative periods and such surgical events as devascularization or arterial ligation with cessation of blood supply during resection (called warm ischemia time) may cause changes in the molecular profiles of resected tumor and normal tissues and preclude use of specimens for research. Shifts in molecular profiles due to iatrogenic interventions may not be recognized as artifacts and may be mistakenly interpreted as disease signatures. Some drugs used in perioperative and interoperative periods have powerful molecular effects and are, in fact,
used specifically for these effects (Juhl, 2010), although little research has been published on this topic. In addition, the type of surgical procedure, the individual surgeon performing the procedure, and the use of robotic instruments are variables that affect the duration of the resection and the resulting tissue ischemia in the resection specimen. Although some information related to the number and types of drugs given before and during an operation may be available in clinical records, such as the anesthesia report, it is uncommon to have these data available to a biorepository. The committee does not know whether or how many specimens in the JPC repository are annotated with data related to perioperative variables that may be pertinent to molecular research on the molecular profiles of tissue samples.
Perhaps the most common preanalytic variable with an important effect on results of analytic tests, such as immunohistochemical staining, is the time that elapses after surgical removal of tissue until it is stabilized with fixation or freezing (cold ischemia time). The process of structural and biochemical tissue degradation occurring during this time is termed autolysis. The amount of time can be substantial, ranging from minutes to hours, and is rarely recorded in the pathology report. It may cause gain or loss of signal on molecular analysis, depending on the molecular entity in question. An analysis done without knowledge of the duration of the time to fixation and without an intrinsic control within the same specimen that can serve as a reference (for example, surrounding normal tissue that is known to express or not to express the molecule constitutively) may be misleading or even completely incorrect (Spruessel et al., 2004).
Storage conditions or duration may compromise specimen quality.
Oxidation occurs in both blocks and cut sections, but it is much greater on cut surfaces exposed to room air. Thus, tissue blocks stored at temperatures below the melting point of paraffin yield cut sections that show little deterioration of immunohistochemical signal compared with freshly embedded controls for as little as 2 years or as long as 25 years (Engel and Moore, 2011). RNAases are active even in FFPE tissues. For many antigens, immunoreactivity has been shown to deteriorate more rapidly if specimens are stored as cut sections rather than whole blocks; both the time line and the magnitude of the effect are antigen-dependent.
On a crude level, paraffin blocks that have been stored under conditions that do not include climate control may lead to melting of paraffin blocks in warm weather or destruction from other causes. Gross specimens, albeit formalin-fixed, may undergo dehydration, fungal contamination, and putrefying deterioration when stored under conditions that expose them to environmental extremes. All those issues have affected specimens held by the JPC at some point over the course of the repository’s history (Asterand, 2008).
Case-matched normal control samples from a tumor patient might be unavailable.
A normal specimen is needed as a source of a reference genome in tumor patients in for correct identification of tumor-specific mutations. Depending on the disease, the diagnostic or therapeutic procedure that produced the specimen, or the tissue remaining in a case after diagnostic analysis, it may not be possible to meet this requirement.
The clinical data associated with specimens may be inadequate, inaccurate, or nonstandardized.
Even if biospecimens are of sufficient quantity and quality for a particular molecular analysis, their value for translational research may be severely limited by the amount, type, and quality of clinical data that are available for an individual case and by the consistency of the data on the many cases that may be required for a study. Clinical data provide the essential functional or biologic behavioral correlations that define the medical relevance of the molecular data. The lack of relevant clinical data elements limits the value of the molecular analysis data for prediction and the conclusions that can be drawn.
The type and amount of clinical data provided by the physician requesting consultation was not prescribed in a standardized fashion by the JPC repository. Furthermore, it did not require that any clinical data submitted adhere to a standardized format.4 Thus, clinical data associated with cases varies widely in quality, quantity, and consistency, which may create important limitations in the utility of the JPC biospecimens for research studies.
In some cases, it may be possible to acquire missing clinical data from the medical record in the institution in which the referred case originated. For some institutions, such as those within the Department of Veterans Affairs health system or the Department of Defense and many private institutions, the acquisition of additional data elements for a case may be technically facilitated by using an electronic medical record (EMR). Nevertheless, additional technical hurdles—such as the difficulty in combining data from an EMR with the JPC’s information technology and the unreliability or unavailability of identifiers that would allow data from disparate sources to be combined—may limit the ease with which the data can be transferred or may preclude electronic transfer altogether. Acquiring or transferring additional data by nonelectronic means is labor-intensive, is expensive, and adds a risk of introducing errors. However, given the number of hospital mergers and closings in recent years, it may not be easy to trace a patient
___________________
4The Contributor’s Consultation Request Form used in recent years does require that some standardized information be submitted. The most recent version of the form is reproduced in Appendix B.
record through the original submitting institution. The ability to acquire additional data may also be constrained by ethical and legal restrictions, such as privacy laws, as noted briefly below.
Consent from the source might be inadequate for use, permission to recontact might be lacking, or recontact might be prohibitively expensive.
Although the acquisition of additional clinical data for a case may be technically possible, there may be circumstances where it would be prohibited or restricted by ethical or legal issues related to recontact or privacy laws, including the federal Health Insurance Portability and Accountability Act of 1996 (PL 104-191, 110 Stat. 1936, and accompanying regulations). In other situations—especially where a number of years have passed since the data or specimen were collected—it may require extensive research to locate the source individual. Those issues may render specific cases unusable for some types of research. Chapter 3 addresses consent and other ethical, legal, and regulatory aspects of use of the JPC materials for consultative, educational, and research purposes.
Aebersold R, Mann M. 2003. Mass spectrometry-based proteomics. Nature 422(6928):198-207.
Akalu A, Reichardt JK. 1999. A reliable PCR amplification method for microdissected tumor cells obtained from paraffin-embedded tissue. Genetic Analysis 15:229-233.
Asterand. 2008. Assessment of the Department of Defense’s tissue repository located at the Armed Forces of Pathology in Washington DC. Detroit, MI: Asterand, Inc.
Azimzadeh O, Barjaktarovic Z, Aubele M, Calzada-Wack J, Sarioglu H, Atkinson MJ, Tapio S. 2010. Formalin-fixed paraffin-embedded (FFPE) proteome analysis using gel-free and gel-based proteomics. Journal of Proteome Research 9(9):4710-4720.
Bataille F, Troppmann S, Klebl F, Rogler G, Stoelcker B, Hofstadter F, Bosserhoff AK, Rummele P. 2006. Multiparameter immunofluorescence on paraffin-embedded tissue sections. Applied Immunohistochemistry & Molecular Morphology 14(2):225-228.
Beck AH, Weng Z, Witten DM, Zhu S, Foley JW, Lacroute P, Smith CL, Tibshirani R, van de Rijn M, Sidow A, West RB. 2010. 3-end sequencing for expression quantification (3SEQ) from archival tumor samples. PLoS One 5(1):e8768.
Becker JS, Jakubowski N. 2009. The synergy of elemental and biomolecular mass spectrometry: New analytical strategies in life sciences. Chemical Societies Review 38(7):1969-1983.
Bischoff K, Lamm C, Erb HN, Hillebrandt JR. 2008. The effects of formalin fixation and tissue embedding of bovine liver on copper, iron, and zinc analysis. Journal of Veterinary Diagnostic Investigation 20(2):220-224.
Bonin S, Hlubek F, Benhattar J, Denkert C, Dietel M, Fernandez PL, Hofler G, Kothmaier H, Kruslin B, Mazzanti CM, Perren A, Popper H, Scarpa A, Soares P, Stanta G, Groenen PJ. 2010. Multicentre validation study of nucleic acids extraction from FFPE tissues. Virchows Archiv 457:309-317.
Botling J, Edlund K, Segersten U, Tahmasebpoor S, Engström M, Sundström M, Malmström PU, Micke P. 2009. Impact of thawing on RNA integrity and gene expression analysis in fresh frozen tissue. Diagnostic Molecular Pathology 18(1):44-52.
Camp RL, Neumeister V, Rimm DL. 2008. A decade of tissue microarrays: progress in the discovery and validation of cancer biomarkers. Journal of Clinical Oncology 26:5630-5637.
Chung JY, Braunschweig T, Williams R, Guerrero N, Hoffmann KM, Kwon M, Song YK, Libutti SK, Hewitt SM. 2008. Factors in tissue handling and processing that impact RNA obtained from formalin-fixed, paraffin-embedded tissue. Journal of Histochemistry and Cytochemistry 56(11):1033-1042.
Clarke M, Shimono Y. 2011. Methods and compositions relating to carcinoma stem cells. US Patent application number: 20110021607. Publication date: 01/27/2011. http://www.faqs.org/patents/app/20110021607 (accessed August 6, 2012).
Cooper K. 2006. Errors and error rates in surgical pathology: an Association of Directors of Anatomic and Surgical Pathology survey. Archives of Pathology & Laboratory Medicine 130(5):607-609.
Coudry RA, Meireles SI, Stoyanova R, Cooper HS, Carpino A, Wang X, Engstrom PF, Clapper ML. 2007. Successful application of microarray technology to microdissected formalin-fixed, paraffin-embedded tissue. Journal of Molecular Diagnostics 9:70-79.
Donadio E, Giusti L, Cetani F, Da Valle Y, Ciregia F, Giannaccini G, Pardi E, Saponaro F, Torregrossa L, Basolo F, Marcocci C, Lucacchini A. 2011. Evaluation of formalin-fixed paraffin-embedded tissues in the proteomic analysis of parathyroid glands. Proteome Science 9(1):29.
Engel KB, Moore HM. 2011. Effects of pre-analytical variables on the detection of proteins by immunohistochemistry in formalin-fixed paraffin-embedded tissues. Archives of Pathology & Laboratory Medicine 135(5):537-543.
Farragher SM, Tanney A, Kennedy RD, Paul Harkin D. 2008. RNA expression analysis from formalin-fixed paraffin embedded tissues. Histochemistry and Cell Biology 130:435-445.
Ferri GL, Gaudio RM, Castello IF, Berger P, Giro G. 1997. Quadruple immunofluorescence: a direct visualization method. Journal of Histochemistry and Cytochemistry 45(2):155-158.
Frank M, Doring C, Metzler D, Eckerle S, Hansmann ML. 2007. Global gene expression profiling of formalin-fixed paraffin-embedded tumor samples: A comparison to snap-frozen material using oligonucleotide microarrays. Virchows Archive 450:699-711.
Hanash SM, Pitteri SJ, Faca VM. 2008. Mining the plasma proteome for cancer biomarkers. Nature 452(7187):571-579.
Harrington CF, Vidler DS, Jenkins RO. 2010. Analysis of organometal(loid) compounds in environmental and biological samples. Metal Ions in Life Sciences 7:33-69.
Hewitt SM, Lewis FA, Cao Y, Conrad RC, Cronin M, Danenberg KD, Goralski TJ, Langmore JP, Raja RG, Williams PM, Palma JF, Warrington JA. 2008. Tissue handling and specimen preparation in surgical pathology: issues concerning the recovery of nucleic acids from formalin-fixed, paraffin-embedded tissue. Archives of Pathology and Laboratory Medicine 132:1929-1935.
Iacobuzio-Donahue CA, Maitra A, Olsen M, Lowe AW, van Heek NT, Rosty C, Walter K, Sato N, Parker A, Ashfaq R, Jaffee E, Ryu B, Jones J, Eshleman JR, Yeo CJ, Cameron JL, Kern SE, Hruban RH, Brown PO, Goggins M. 2003. Exploration of global gene expression patterns in pancreatic adenocarcinoma using cDNA microarrays. American Journal of Pathology 162(4):1151-1162.
Janssens AC, van Duijn CM. 2008. Genome-based prediction of common diseases: advances and prospects. Human Molecular Genetics 17(R2):R166-R173.
Jiang X, Jiang X, Feng S, Tian R, Ye M, Zou H. 2007. Development of efficient protein extraction methods for shotgun proteome analysis of formalin-fixed tissues. Journal of Proteome Research 6(3):1038-1047.
Juhl H. 2010. Preanalytical aspects: A neglected issue. Scandinavian Journal of Clinical and Laboratory Investigation Supplementum 242:63-65.
Kapur P. 2011. Tailoring treatment of rectal adenocarcinoma: immunohistochemistry for predictive biomarkers. Anticancer Drugs 22:362-370.
Kraft P, Wacholder S, Cornelis MC, Hu FB, Hayes RB, Thomas G, Hoover R, Hunter DJ, Chanock S. 2009. Beyond odds ratios—communicating disease risk based on genetic profiles. Nature Reviews Genetics 10(4):264-269.
Li J, Smyth P, Flavin R, Cahill S, Denning K, Aherne S, Guenther SM, O’Leary JJ, Sheils O. 2007. Comparison of miRNA expression patterns using total RNA extracted from matched samples of formalin-fixed paraffin-embedded (FFPE) cells and snap frozen cells. BMC Biotechnology 7:36.
Lind A, Bewtra C, Healy JC, Sims KL. 1995. Prospective peer review in surgical pathology. American Journal of Clinical Pathology 104(5):560-566.
Linton KM, Hey Y, Saunders E, Jeziorska M, Denton J, Wilson CL, Swindell R, Dibben S, Miller CJ, Pepper SD, Radford JA, Freemont AJ. 2008. Acquisition of biologically relevant gene expression data by Affymetrix microarray analysis of archival formalin-fixed paraffin-embedded tumours. British Journal of Cancer 98:1403-1414.
Liu A, Tetzlaff MT, Vanbelle P, Elder D, Feldman M, Tobias JW, Sepulveda AR, Xu X. 2009. MicroRNA expression profiling outperforms mRNA expression profiling in formalin-fixed paraffin-embedded tissues. International Journal of Clinical and Experimental Pathology 2:519-527.
Mason DY, Micklem K, Jones M. 2000. Double immunofluorescence labelling of routinely processed paraffin sections. Journal of Pathology 191(4):452-461.
Micke P, Ohshima M, Tahmasebpoor S, Ren ZP, Ostman A, Pontén F, Botling J. 2006. Biobanking of fresh frozen tissue: RNA is stable in nonfixed surgical specimens. Laboratory Investigation 86(2):202-211.
Mizuhira V, Hasegawa H, Notoya M. 2000. Fixation and imaging of biological elements: heavy metals, diffusible substances, ions, peptides, and lipids. Progress in Histochemistry and Cytochemistry 35(2):67-183.
Niki H, Hosokawa S, Nagaike K, Tagawa T. 2004. A new immunofluorostaining method using red fluorescence of PerCP on formalin-fixed paraffin-embedded tissues. Journal of Immunological Methods 293(1-2):143-151.
Pantanowitz L, Valenstein PN, Evans AJ, Kaplan KJ, Pfeifer JD, Wilbur DC, Collins LC, Colgan TJ, 2011. Review of the current state of whole slide imaging in pathology. Journal of Pathology Informatics 2:36.
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, Marron JS, Nobel AB, Mardis E, Nielsen TO, Ellis MJ, Perou CM, Bernard PS. 2009. Supervised risk predictor of breast cancer based on intrinsic subtypes. Journal of Clinical Oncology 27:1160-1167.
Penland SK, Keku TO, Torrice C, He X, Krishnamurthy J, Hoadley KA, Woosley JT, Thomas NE, Perou CM, Sandler RS, Sharpless NE. 2007. RNA expression analysis of formalin-fixed paraffin-embedded tumors. Laboratory Investigation 87:383-391.
Prieto DA, Hood BL, Darfler MM, Guiel TG, Lucas DA, Conrads TP, Veenstra TD, Krizman DB. 2005. Liquid tissue: Proteomic profiling of formalin-fixed tissues. BioTechniques Suppl:32-35.
Ramsey AD, Gallagher PJ. 1992. Local audit of surgical pathology. 18 months experience of peer review-based quality assessment in an English teaching hospital. American Journal of Surgical Pathology 16(5):476-482.
Ravo M, Mutarelli M, Ferraro L, Grober OM, Paris O, Tarallo R, Vigilante A, Cimino D, De Bortoli M, Nola E, Cicatiello L, Weisz A. 2008. Quantitative expression profiling of highly degraded RNA from formalin-fixed, paraffin-embedded breast tumor biopsies by oligonucleotide microarrays. Laboratory Investigation 88(4):430-440.
Renshaw A, Norberto-Cartagena N, Granter SR, Gould EW. 2003a. Agreement and error rates using blinded review to evaluate surgical pathology of biopsy material. American Journal of Clinical Pathology 119(6):797-800.
Renshaw A, Young M, Jiroutek MR. 2003b. How many cases need to be reviewed to compare performance in surgical pathology? American Journal of Clinical Pathology 119(3):388-391.
Rimm DL, Nielsen TO, Jewell SD, Rohrer DC, Broadwater G, Waldman F, Mitchell KA, Singh B, Tsongalis GJ, Frankel WL, Magliocco AM, Lara JF, Hsi ED, Bleiweiss IJ, Badve SS, Chen B, Ravdin PM, Schilsky RL, Thor A, Berry DA; Cancer and Leukemia Group B Pathology Committee. 2011. Cancer and Leukemia Group B Pathology Committee Guidelines for Tissue Microarray Construction representing multicenter prospective clinical trial tissues. Journal of Clinical Oncology 16:2282-2290.
Rudloff U, Bhanot U, Gerald W, Klimstra DS, Jarnagin WR, Brennan MF, Allen PJ. 2010. Biobanking of human pancreas cancer tissue: impact of ex-vivo procurement times on RNA quality. Annals of Surgical Oncology 17(8):2229-2236.
Safrin R, Bark C. 1993. Surgical pathology signout. Routine review of every case by a second pathologist. American Journal of Surgical Pathology 17(11):1190-1192.
Scicchitano MS, Dalmas DA, Boyce RW, Thomas HC, Frazier KS. 2009. Protein extraction of formalin-fixed, paraffin-embedded tissue enables robust proteomic profiles by mass spectrometry. Journal of Histochemistry and Cytochemistry 57(9):849-860.
Shi SR, Liu C, Balgley BM, Lee C, Taylor CR. 2006. Protein extraction from formalin-fixed, paraffin-embedded tissue sections: quality evaluation by mass spectrometry. Journal of Histochemistry and Cytochemistry 54:739-743.
Spruessel A, Steimann G, Jung M, Lee SA, Carr T, Fentz AK, Spangenberg J, Zornig C, Juhl HH, David KA. 2004. Tissue ischemia time affects gene and protein expression patterns within minutes following surgical tumor excision. BioTechniques 36(6):1030-1037.
Sprung RW Jr, Brock JW, Tanksley JP, Li M, Washington MK, Slebos RJ, Liebler DC. 2009. Equivalence of protein inventories obtained from formalin-fixed paraffin-embedded and frozen tissue in multidimensional liquid chromatography-tandem mass spectrometry shotgun proteomic analysis. Molecular & Cellular Proteomics 8(8):1988-1998.
St. Croix B, Rago C, Velculescu V, Traverso G, Romans KE, Montgomery E, Lal A, Riggins GJ, Lengauer C, Vogelstein B, and Kinzler KW. 2000. Genes expressed in human tumor endothelium. Science 289(5482):1197-1202.
Strand C, Enell J, Hedenfalk I, Fernö M. 2007. RNA quality in frozen breast cancer samples and the influence on gene expression analysis—a comparison of three evaluation methods using microcapillary electrophoresis traces. BMC Molecular Biology 8:38.
Torbenson M. 2011. Iron in the liver: A review for surgical pathologists. Advances in Anatomic Pathology 18(4):306-317.
van der Net JB, Janssens AC, Sijbrands EJ, Steyerberg EW. 2009. Value of genetic profiling for the prediction of coronary heart disease. American Heart Journal 158:105-110.
Voduc D, Kenney C, Nielson TO. 2008. Tissue microarrays in clinical oncology. Seminars in Radiation Oncology 18:89-97.
von Ahlfen S, Missel A, Bendrat K, Schlumpberger M. 2007. Determinants of RNA quality from FFPE samples. PLoS One 2(12):e1261.
Votavová H, Forsterová K, Stritesky J, Velenska Z, Trněný M. 2009. Optimized protocol for gene expression analysis in formalin-fixed, paraffin-embedded tissue using real-time quantitative polymerase chain reaction. Diagnostic Molecular Pathology 18(3):176-182.
Wakely S, Baxendine-Jones J, Gallagher PJ, Mullee M, Pickering R. 1998. Aberrant diagnoses by individual surgical pathologists. American Journal of Surgical Pathology 22(1):77-82.
Warnke RA, Gatter KC, Falini B, Hildreth P, Woolston RE, Pulford K, Cordell JL, Cohen B, De Wolf-Peeters C, Mason DY. 1983. Diagnosis of human lymphoma with monoclonal antileukocyte antibodies. New England Journal of Medicine 309:1275-1281.
Werner M, Chott A, Fabiano A, Battifora H. 2000. Effect of formalin tissue fixation and processing on immunohistochemistry. The American Journal of Surgical Pathology 24(7):1016-1019.
West RB. 2010. Expression profiling in soft tissue sarcomas with emphasis on synovial sarcoma, gastrointestinal stromal tumor, and leiomyosarcoma. Advances in Anatomic Pathology 17(5):366-373.
Wu ZJ, Meyer CA, Choudhury S, Shipitsin M, Maruyama R, Bessarabova M, Nikolskaya T, Sukumar S, Schwartzman A, Liu JS, Polyak K, Liu XS. 2010. Gene expression profiling of human breast tissue samples using SAGE-Seq. Genome Research 20(12):1730-1739.
Xu J, Sun J, Kader AK, Linström S, Wiklund F, Hsu FC, Johansson JE, Zheng SL, Thomas G, Hayes RB, Kraft P, Hunter DJ, Chanock SJ, Isaacs WB, Grönberg H. 2009. Estimation of absolute risk for prostate cancer using genetic markers and family history. The Prostate 69(14):1565-1572.




























