ANIMAL RESERVOIRS OF MIDDLE EAST RESPIRATORY SYNDROME CORONAVIRUS
Jonathan H. Epstein1and Kevin J. Olival1
Introduction
Middle East respiratory syndrome coronavirus (MERS-CoV) is a newly recognized group C β-coronavirus within the family Coronaviridae that was first isolated from a Saudi patient suffering from severe respiratory disease in June 2012 (Zaki et al., 2012). A phylogenetic analysis of the complete viral genome indicated that it was related to severe acute respiratory syndrome (SARS) coronavirus, making it the second β-coronavirus to be identified in humans, although it was of a different lineage (van Boheemen et al., 2012). As of July 2014, MERS-CoV (formerly hCoV-EMC) has caused more than 837 laboratory-confirmed cases of human infection in 21 countries with an overall mortality rate of approximately 35 percent (WHO, 2014). The majority of cases have occurred in Saudi Arabia, and human-to-human transmission has resulted in several clusters of cases, some of which have included mild or asymptomatic infections (Milne-Price et al., 2014).
_______________
1 EcoHealth Alliance, New York, NY.
Epidemiological studies have identified hospital-based infections as responsible for several clusters of cases; however, there were many cases that did not report contact with other MERS-CoV patients, and the source of their infection was unknown. The genetic relationship to SARS and other bat-associated CoVs lead to early suspicion that this, too, was a bat coronavirus, yet it was unclear which species it was associated with and whether patients had had direct exposure to bats or whether another animal host may have been involved in human infections (van Boheemen et al., 2012). Here we review current evidence for the animal origins of MERS-CoV and the involvement of a domestic animal reservoir, dromedary camels, in human infection.
Early Evidence for a Bat Reservoir for MERS-CoV
Since the discovery of as SARS-like CoV in Rhinolophus bat species in China and Hong Kong (Lau et al., 2005; Li et al., 2005) in 2004, there has been a huge proliferation of coronavirus studies in bats worldwide. Subsequent studies have described other SARS-like CoVs in bats in Asia, Africa, and Europe (Rihtaric et al., 2010; Tong et al., 2009; Yuan et al., 2010), as well as a diversity of other coronaviruses in bats around the world (Anthony et al., 2013; August et al., 2012; Falcon et al., 2011; Osborne et al., 2011; Shirato et al., 2012; Tao et al., 2012). Phylogenetic analyses of coronaviruses from bats from both the Old and New World, humans, and other known mammalian and avian coronaviruses have shown a greater diversity of viral species compared to other taxonomic host groups, and support the hypothesis that major groups within the family Coronaviridae originated in bats (Drexler et al., 2014; Huynh et al., 2012).
Evidence of Host Range from Receptor Binding Studies
Characterizing receptor binding for a given coronavirus can provide insights into the range of potential host species and tissue types that the virus may infect (Graham and Baric, 2010). SARS coronavirus requires the angiotensin-converting enzyme 2 (ACE2) receptor to enter cells (Eickmann et al., 2003). In humans, ACE2 receptors are found in lung and small intestine epithelial tissue, which was where SARS-CoV replication primarily occurs resulting in severe lower respiratory tract and gastrointestinal tract infection (Hamming et al., 2004; Ksiazek et al., 2003; Nicholls et al., 2003). The SARS-like coronavirus strains first identified in horseshoe bats were closely related to SARS-CoV (88 to 92 percent nucleotide homology across the full genome), but they did not enter cells expressing human or civet ACE2 receptors, nor could SARS-CoV infect cells expressing bat ACE2, leaving doubt as to whether they were the direct progenitor of SARS-CoV (Ren et al., 2008). Recently, a SARS-like virus with 95 percent homology to SARS-CoV and that does use the human ACE2 receptor was isolated from Rhinolophus sinicus, providing the most convincing evidence to date that
bats are the natural reservoir for SARS-CoV and that direct zoonotic transmission from bats is possible (Ge et al., 2013). Early genetic characterization led to the recognition that MERS-CoV was related to SARS-CoV and to two other group C β-coronaviruses, HKU4 and HKU5, which generated the hypothesis that MERS-CoV also had a bat reservoir (Woo et al., 2012; Zaki et al., 2012). In vitro infection of various mammalian cell lines with MERS-CoV (then called hCoV-EMC) showed that the virus did not use the ACE2 receptor and that it was able to replicate in a variety of animal cell lines, suggesting the possibility of a broad mammalian host range including nonhuman primates, pigs, and multiple species of bats (Muller et al., 2012). Similarly, Eckerle et al. used cell lines from common Arabian livestock and other mammal species to show that MERS-CoV replicates efficiently in goat, camel, bat, human, and African green monkey cells, but less efficiently in cow, sheep, bank vole, and shrew cell lines (Eckerle et al., 2014).
Although BtCoV-HKU4 and BtCoV-HKU5 coronaviruses, which were identified in two species of vespertilionid bats (Tylonycterus pachypus and Pipistrellis abramus, respectively), were closely related to MERS-CoV, phylogenetic analysis revealed that these bats were unlikely its natural reservoir (Lau et al., 2013; Woo et al., 2012). An in vitro study using virus surface spike proteins from HKU4 and HKU5 demonstrated that HKU4 binds to the dipeptidyl peptidase 4 (DPP4) receptor, but HKU5 does not (Yang et al., 2014). Yang et al. further showed that HKU4 had a stronger affinity to bat cells over human cells; the opposite pattern was observed in MERS-CoV.
Shortly after the first human case of MERS-COV was identified in 2012, coronaviruses closely related to MERS were found in bat species in Mexico, Thailand, Europe, and Africa, illustrating the wide geographic and bat family range of viruses similar to MERS-CoV and adding support to the hypothesis that MERS-CoV had originated in bats; however, the potential host species in Saudi Arabia could not yet be deduced (Annan et al., 2013; Anthony et al., 2013; Ithete et al., 2013; Wacharapluesadee et al., 2013).
The Search for the Natural Reservoir in Saudi Arabia
The initial investigation of the natural reservoir for MERS-CoV began in October 2012 in Bisha, Saudi Arabia, the town where the index patient had lived (Figure A1-1). Despite the initial hypothesis that MERS-CoV had bat origins, there was no information available from the medical records of the first patient that described any contact with bats or other animals. Furthermore, there was little known about the bat fauna or distribution of other wildlife in this area. It was also unknown at the time whether MERS-CoV was capable of infecting other species, including domestic livestock. The investigation included a visit to the patient’s households and place of business to observe whether bats and domestic animals were present. Interviews with the patient’s surviving family members and an absence of evidence of bats in the households (he had multiple residences)
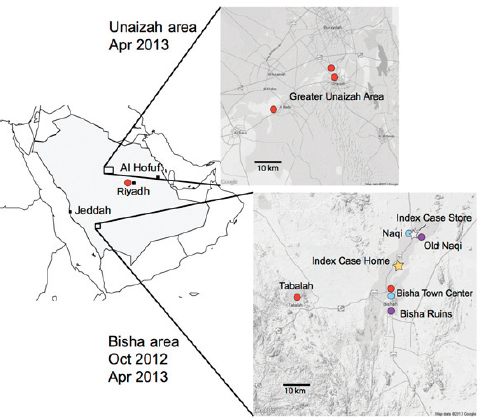
FIGURE A1-1 Map of the initial investigation of bats as a reservoir for MERS-CoV in Bisha and Unaizah.
SOURCE: Memish et al., 2013.
suggested that direct contact with bats was unlikely. The patient was a 60-year-old business man who owned four camels, kept as companion animals in a paddock next to his house, and a flock of sheep and goats kept at his business about 15 km north of Bisha. His business was a hardware shop with a large warehouse, and although bats were observed foraging in a palm grove behind the warehouse, there was no evidence of bats roosting inside the building, which again suggested direct contact with bats or their excreta inside his home or business was unlikely. Seven different species of bats representing four families (Rhinopoma hardwickii, Rhinopoma microphyllum, Taphozous perforatus, Pipistrellus kuhlii, Eptesicus bottae, Eidolon helvum, and Rosettus aegyptiacus) were captured and sampled either near his business or in Bisha on two separate field investigations (Figure A1-1). In April 2013, in addition to the Bisha area, additional bat specimens were collected from other geographic areas with human cases including Unaizah and Riyadh (Figure A1-1). Insectivorous bats were primarily found
roosting in abandoned buildings, one occasionally inhabited building, and captured during evening emergence from the roost, while frugivorous bats (Rousettus and Eidolon) were captured while foraging. Fecal samples, blood, and oropharyngeal swabs were collected from each bat, and fecal samples were also collected by laying out plastic sheets beneath roosts. Samples were screened at the Center for Infection and Immunity at Columbia University using pan coronavirus and MERS-CoV-specific polymerase chain reaction (PCR) assays (Memish et al., 2013). A short 190nt fragment of viral RNA that was 100 percent identical to MERS-CoV in the RdRp region was detected in a fecal sample from an Egyptian tomb bat (Taphozous perforatus) captured in Bisha (Memish et al., 2013). One of 29 tomb bats was positive, indicating a prevalence of 3.5 percent (95% CI 0–20%). Although similar coronaviruses had been identified in other bat species from other regions, this represented the first finding of RNA matching MERS-CoV in a bat in Saudi Arabia. Although this finding provided a valuable indication that Taphozous perforatus may be a reservoir for MERS-CoV, a broader survey is needed to confirm the finding and potentially identify other bat species that may be involved as reservoirs for MERS-CoV or related viruses on the Arabian Peninsula. For example, HKU10 CoV (an α-coronavirus) was found to naturally infect two bat species from distinct families (Pteropodidae and Hipposideridae) that had ecological overlap (Lau et al., 2012). It remains unknown how the index patient was infected with MERS-CoV, and possibilities include infection by another MERS case or zoonotic transmission. The finding of MERS-CoV in a bat did not rule out the potential involvement of other animal hosts as being involved in human infection, as transmission directly from bats to humans seemed unlikely given the limited opportunities for exposure to bat excreta.
Potential Livestock Hosts
The first two cases of MERS-CoV had no information regarding possible exposure to animals in their clinical history. As the numbers of cases increased, and information was reported to WHO and the international community, there continued to be little or no information about animal exposure. The four camels as well as the sheep that were owned by the index patient from Bisha all tested negative for MERS-CoV (Alagaili et al., 2014). The cellular receptor used by MERS-CoV was later identified as the DPP4 receptor, which is conserved across many mammalian species and found in a variety of tissue types including lung and kidney epithelium (Raj et al., 2013). As previously noted, in vitro studies of host range using cell lines suggested a breadth of potential hosts. Muller et al. and Eckerle et al. determined that MERS-CoV could infect bat cell lines derived from six species as well as pig, camel, sheep, nonhuman primate, and human cell lines (Eckerle et al., 2014; Muller et al., 2012).
By August 2013, human cases had been reported from several countries including Jordan, Qatar, and the United Arab Emirates, though most were from
Saudi Arabia, and most reported infections were the result of human-to-human transmission (Assiri et al., 2013a,b). However, human-to-human transmission was limited (R0 ~ 0.69), indicating MERS-CoV was not easily transmitted among people, and supporting the hypothesis that repeated spillover from an animal reservoir may be occurring (Breban et al., 2013). The first study to provide evidence of a domestic animal host found IgG antibodies specific to MERS-CoV in dromedary camel herds in Oman and the Canary Islands (Reusken et al., 2013b). One hundred percent of the camels tested in Oman (n = 50) and 14 percent (n = 105) of Spanish camels were positive for MERS-CoV antibodies. Several species of domestic animals in various countries including Oman, Egypt, Jordan, and Saudi Arabia were screened for antibodies against MERS-CoV, including sheep, goats, cattle, and buffalo, but all were negative (Alagaili et al., 2014; Hemida et al., 2013; Perera et al., 2013; Reusken et al., 2013a). Six of 126 sheep were positive for MERS-CoV reactive antibodies in Jordan; however, none of the six had neutralizing antibodies, and it was suspected that there was cross-reactivity with the antigen used for the initial screening assay (Reusken et al., 2013a). Anti-MERS-CoV antibodies have subsequently been found in camels in Egypt, Qatar, and the United Arab Emirates at high prevalence (Alexandersen et al., 2014; Chu et al., 2014; Meyer et al., 2014; Nowotny and Kolodziejek, 2014). A study of dromedary camels in Saudi Arabia examined sera from 2013 and dating back to 1993 found antibodies to MERS-CoV, indicating that it had been circulating in camels in Saudi Arabia for at least 20 years (Alagaili et al., 2014). The same group detected MERS-CoV RNA in nasal swabs from adult and juvenile camels, and isolates were obtained from nasal swabs from camels in Saudi Arabia in 2014 (Briese et al., 2014). Sequences from camels were 99 percent identical to human full-genome sequences, yet individual camels were found to be infected by multiple, closely related strains of MERS-CoV, or viral quasispecies (Briese et al., 2014). The observed genetic diversity of MERS-CoV in camels, which was greater than in humans, and the sequence homology between human and camel strains, suggested that multiple introductions from camels may be occurring and that there may be some bottleneck selection if only certain strains are being transmitted from camels to humans (Briese et al., 2014; Cotten et al., 2014). These findings did not specifically provide direct evidence for zoonotic transmission from camels; however, support for this hypothesis is provided by two other studies. In October 2013, two patients with laboratory confirmed MERS-CoV infection had a history of contact with camels on their farm. An investigation found MERS-CoV RNA in nasal swabs from 5 of 14 camels tested within a week of detection of the first human case. Sequences from the camels and two patients were closely related (Haagmans et al., 2014). In a second investigation, MERS-CoV was isolated from a patient in Jeddah, Saudi Arabia, who became ill after having had contact with nasal discharge while treating several of his camels that were ill. MERS-CoV was also isolated from one of his camels, and the sequence matched that of the isolate from the patient (Azhar et al., 2014). It is
unclear whether camels experience severe pathology or disease from MERS-CoV infection, although the infected camels in Jeddah were reported to have had nasal discharge (Azhar et al., 2014). MERS-CoV antibodies and viral RNA have been detected in camel milk in Qatar, and experimentally the virus has been shown to be stable in camel milk, though it is currently unknown whether consumption of raw milk has led to human infections (Reusken et al., 2014a; van Doremalen et al., 2014). Definitive proof of transmission from camels to humans, or of a potential mechanism for transmission, has not yet been characterized, although there is a preponderance of serological evidence now that MERS-CoV is circulating widely in camels both in the Middle East and in parts of Africa (Chu et al., 2014; Corman et al., 2014a; Perera et al., 2013; Reusken et al., 2014b).
Camel Trade as a Driver of MERS-CoV Emergence
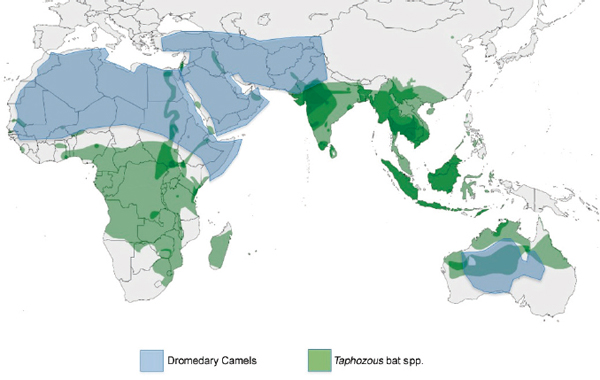
Antibodies to MERS-CoV have been identified in dromedary camel serum samples in both Saudi Arabia and Kenya dating as far back as 1992, indicating that MERS-CoV has been present in camel populations for more than 20 years in both Africa and Saudi Arabia (Alagaili et al., 2014; Corman et al., 2014a). The majority of the world’s camels (82.2 percent) produced between 1992 and 2013 have come from northern Africa, with Somalia and Sudan being the top two producers (Figure A1-2) (Food and Agriculture Organization of the United Nations Statistics Division, 2014). The Kingdom of Saudi Arabia imports most of its camels from Africa, and it is the top camel importer in the world, with an average of 60,900 camels imported per year between 1992 and 2011 (Food and Agriculture Organization of the United Nations Statistics Division, 2014). Given the apparent ubiquity of anti-MERS-CoV IgG found in camels in several African countries, including Ethiopia, Kenya, Tunisia, Egypt, and Nigeria, it is likely that MERS-CoV-infected camels have been imported into Saudi Arabia multiple times over the past two decades. It is also possible that MERS-CoV was introduced to Saudi Arabia from Africa via the camel trade, which presents an alternate hypothesis to one proposed by Memish et al. (2013) that spillover of MERS-CoV from its original bat host occurred in Saudi Arabia. Indeed, one of the MERS-CoV isolates obtained by Briese et al. in Saudi Arabia came from a juvenile (< 1 year old) camel of African origin (Briese et al., 2014). While multiple bat species could potentially be involved in the ecology and zoonotic spillover of MERS-CoV, if we look at just the distribution of Taphozous spp. bats we see that their species ranges overlap with dromedary camel distribution in parts of northern Africa, Arabia, and even Australia (Figure A1-3). In Saudi Arabia, bats were observed roosting in abandoned structures used to occasionally house camels and other livestock species (Epstein and Olival, pers. comm.), and thus the ecological potential for spillover exists although more studies are needed to better quantify this overlap. The wide distribution of anti-MERS-CoV antibodies in dromedary camels in Africa also suggests that zoonotic transmission has occurred there, and
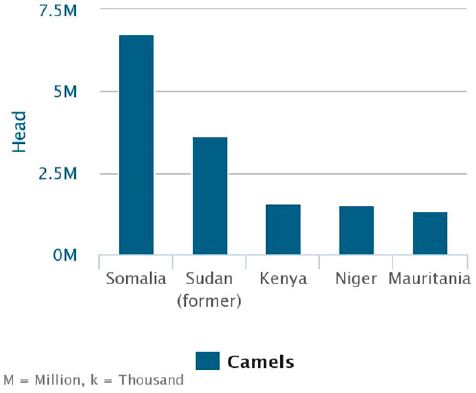
FIGURE A1-2 The top five camel-producing countries (1992–2013).
SOURCE: Data from FAO.
further studies are warranted to determine whether MERS-CoV is circulating in human populations in Africa.
Discussion and Future Research Needs
While there has been an abundance of data collected that suggests an apparent ubiquity of MERS-CoV infection in camels both in the Middle East and Africa, little is understood about potential mechanisms of zoonotic transmission or the frequency of zoonotic transmission. To date, no case-control study has identified high-risk exposures in human MERS cases, other than contact with another MERS case. Nor have there been sufficient epidemiological studies that might identify human MERS cases in Africa. Information about animal exposure has been absent or vague in the majority of reported cases from Saudi Arabia. The outbreak investigation in Qatar of two cases of MERS identifies a history of contact between the cases and sick camels, as well as confirmed MERS-CoV infection in the camels, which provides the best evidence to date that camels may be involved in MERS-CoV transmission to people. Phylogenetic analyses
of MERS-CoV and MERS-related β-coronaviruses in bats support the hypothesis that bats are the natural reservoir for MERS-CoV; however, broader studies are needed to identify which bat species (and it may that more than one is involved) may be responsible for camel and/or human infections either in the Middle East, or more likely, in Africa. There is anecdotal evidence for bat–livestock and bat–human contact in Saudi Arabia, but detailed ecological investigations are needed to quantify the level of contact and identify the specific interfaces that may increase risk of spillover of MERS-CoV and MERS-related CoVs in Arabia and northern Africa. MERS-related CoVs have been identified in bat species in Africa (Annan et al., 2013; Ithete et al., 2013), although more extensive surveys of bats in countries where camels are produced, as well as identification of specific bat–camel interfaces, would provide valuable data on the potential for spillover from bats to camels or people. It will also be important to rule out the involvement of other wildlife species in MERS circulation. MERS-related coronaviruses have been identified in European hedgehogs (Erinaceus europaeus) (Corman et al., 2014b). A related species of hedgehog, Paraechinus aethiopicus, is commonly found across the Middle East and parts of northern Africa; however, to date there have been no epizootiological studies in wildlife species other than bats.
Multidisciplinary, ecological, and virological studies of MERS coronavirus will help further elucidate possible wildlife origins; mechanisms for primary human infections; risk factors for human infection; and the mechanisms and frequency of spillover from bats to camels or other domestic livestock, all of which would help answer the question of whether MERS-CoV is a recently emerging virus in humans or one which has simply escaped detection until 2012. As with SARS-CoV in China or Nipah virus in Bangladesh, the risk of human infection may be reduced when the proximal source of infection (e.g., pteropid bats and date palm sap for Nipah virus) is identified and transmission is interrupted (Nahar et al., 2010). However, without knowing the wildlife reservoir, the risk of reintroduction into animal or human populations cannot be managed. For nearly 10 years following the discovery that SARS-like CoVs were carried by Rhinolophid bats, it was assumed that SARS required intermediary animal hosts such as civets to become transmissible to people. The recent discovery of a strain of SARS-CoV in the same bats that uses the ACE2 receptor provided evidence that direct bat-to-human transmission was possible, and underscored the importance of reducing bat–human exposure. Opportunities for direct contact with bats or their excreta appear to be limited in Saudi Arabia, but more studies are needed to quantify and characterize this. Camel trade is likely the major driver of primary human infection in Saudi Arabia and other Gulf countries, where the majority of camels are imported from Africa. Expanded human and camel surveillance; educational outreach and public health initiatives designed to reduce exposure to camel bodily fluids, particularly nasal secretions; and improved infection control practices in health care settings will be instrumental in reducing the incidence of MERS in human populations.
References
Alagaili, A. N., T. Briese, N. Mishra, V. Kapoor, S. C. Sameroff, P. D. Burbelo, E. de Wit, V. J. Munster, L. E. Hensley, I. S. Zalmout, A. Kapoor, J. H. Epstein, W. B. Karesh, P. Daszak, O. B. Mohammed, and W. I. Lipkin. 2014. Middle East respiratory syndrome coronavirus infection in dromedary camels in Saudi Arabia. mbio 5(2):e00884-14.
Alexandersen, S., G. P. Kobinger, G. Soule, and U. Wernery. 2014. Middle East respiratory syndrome coronavirus antibody reactors among camels in Dubai, United Arab Emirates, in 2005. Transboundary and Emerging Diseases 61(2):105-108.
Annan, A., H. J. Baldwin, V. M. Corman, S. M. Klose, M. Owusu, E. E. Nkrumah, E. K. Badu, P. Anti, O. Agbenyega, B. Meyer, S. Oppong, Y. A. Sarkodie, E. K. V. Kalko, P. H. C. Lina, E. V. Godlevska, C. Reusken, A. Seebens, F. Gloza-Rausch, P. Vallo, M. Tschapka, C. Drosten, and J. F. Drexler. 2013. Human betacoronavirus 2c EMC/2012-related viruses in bats, Ghana and Europe. Emerging Infectious Diseases 19(3):456-459.
Anthony, S., R. Ojeda-Flores, O. Rico-Chávez, I. Navarrete-Macias, C. Zambrana-Torrelio, M. K. Rostal, J. H. Epstein, T. Tipps, E. Liang, M. Sanchez-Leon, J. Sotomayor-Bonilla, A. A. Aguirre, R. Ávila, R. A. Medellín, T. Goldstein, G. Suzán, P. Daszak, and W. I. Lipkin. 2013. Coronaviruses in bats from Mexico. Journal of General Virology.
Assiri, A., J. A. Al-Tawfiq, A. A. Al-Rabeeah, F. A. Al-Rabiah, S. Al-Hajjar, A. Al-Barrak, H. Flemban, W. N. Al-Nassir, H. H. Balkhy, R. F. Al-Hakeem, H. Q. Makhdoom, A. I. Zumla, and Z. A. Memish. 2013a. Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: A descriptive study. Lancet Infectious Diseases 13(9):752-761.
Assiri, A., A. McGeer, T. M. Perl, C. S. Price, A. A. Al Rabeeah, D. A. T. Cummings, Z. N. Alabdullatif, M. Assad, A. Almulhim, H. Makhdoom, H. Madani, R. Alhakeem, J. A. Al-Tawfiq, M. Cotten, S. J. Watson, P. Kellam, A. I. Zumla, Z. A. Memish, and K. M.-C. I. Team. 2013b. Hospital outbreak of Middle East respiratory syndrome coronavirus. New England Journal of Medicine 369(5):407-416.
August, T. A., F. Mathews, and M. A. Nunn. 2012. Alphacoronavirus detected in bats in the United Kingdom. Vector-Borne and Zoonotic Diseases 12(6):530-533.
Azhar, E. I., S. A. El-Kafrawy, S. A. Farraj, A. M. Hassan, M. S. Al-Saeed, A. M. Hashem, and T. A. Madani. 2014. Evidence for camel-to-human transmission of MERS coronavirus. New England Journal of Medicine 370(26):2499-2505.
Breban, R., J. Riou, and A. Fontanet. 2013. Interhuman transmissibility of Middle East respiratory syndrome coronavirus: Estimation of pandemic risk. Lancet 382(9893):694-699.
Briese, T., N. Mishra, K. Jain, I. S. Zalmout, O. J. Jabado, W. B. Karesh, P. Daszak, O. B. Mohammed, A. N. Alagaili, and W. I. Lipkin. 2014. Middle East respiratory syndrome coronavirus quasispecies that include homologues of human isolates revealed through whole-genome analysis and virus cultured from dromedary camels in Saudi Arabia. mbio 5(3):e01146-14.
Chu, D. K. W., L. L. M. Poon, M. M. Gomaa, M. M. Shehata, R. A. P. M. Perera, D. Abu Zeid, A. S. El Rifay, L. Y. Siu, Y. Guan, R. J. Webby, M. A. Ali, M. Peiris, and G. Kayali. 2014. MERS coronaviruses in dromedary camels, Egypt. Emerging Infectious Diseases 20(6):1049-1053.
Corman, V. M., J. Jores, B. Meyer, M. Younan, A. Liljander, M. Y. Said, I. Gluecks, E. Lattwein, B.-J. Bosch, J. F. Drexler, S. Bornstein, C. Drosten, and M. A. Mueller. 2014a. Antibodies against MERS coronavirus in dromedary camels, Kenya, 1992-2013. Emerging Infectious Diseases 20(8):1319-1322.
Corman, V. M., R. Kallies, H. Philipps, G. Goepner, M. A. Mueller, I. Eckerle, S. Bruenink, C. Drosten, and J. F. Drexler. 2014b. Characterization of a novel betacoronavirus related to Middle East respiratory syndrome coronavirus in European hedgehogs. Journal of Virology 88(1): 717-724.
Cotten, M., S. J. Watson, A. I. Zumla, H. Q. Makhdoom, A. L. Palser, S. H. Ong, A. A. Al Rabeeah, R. F. Alhakeem, A. Assiri, J. A. Al-Tawfiq, A. Albarrak, M. Barry, A. Shibl, F. A. Alrabiah, S. Hajjar, H. H. Balkhy, H. Flemban, A. Rambaut, P. Kellam, and Z. A. Memish. 2014. Spread, circulation, and evolution of the Middle East respiratory syndrome coronavirus. mbio 5(1):e01062-13.
Drexler, J. F., V. M. Corman, and C. Drosten. 2014. Ecology, evolution and classification of bat coronaviruses in the aftermath of SARS. Antiviral Research 101:45-56.
Eckerle, I., V. M. Corman, M. A. Müller, M. Lenk, R. G. Ulrich, and C. Drosten. 2014. Replicative apacity of MERS coronavirus in livestock cell lines. Emerging Infectious Diseases 20(2):276-279.
Eickmann, M., S. Becker, H. D. Klenk, H. W. Doerr, K. Stadler, S. Censini, S. Guidotti, V. Masignani, M. Scarselli, M. Mora, C. Donati, J. H. Han, H. C. Song, S. Abrignani, A. Covacci, and R. Rappuoli. 2003. Phylogeny of the SARS coronavirus. Science 302(5650):1504-1505.
Falcon, A., S. Vazquez-Moron, I. Casas, C. Aznar, G. Ruiz, F. Pozo, P. Perez-Brena, J. Juste, C. Ibanez, I. Garin, J. Aihartza, and J. E. Echevarria. 2011. Detection of alpha and betacoronaviruses in multiple Iberian bat species. Archives of Virology 156(10):1883-1890.
Food and Agriculture Organization of the United Nations Statistics Division. 2014. FAOSTAT. FAO.
Ge, X. Y., J. L. Li, X. L. Yang, A. A. Chmura, G. J. Zhu, J. H. Epstein, J. K. Mazet, B. Hu, W. Zhang, C. Peng, Y. J. Zhang, C. M. Luo, B. Tan, N. Wang, Y. Zhu, G. Crameri, S. Y. Zhang, L. F. Wang, P. Daszak, and Z. L. Shi. 2013. Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature 503(7477):535-538.
Graham, R. L., and R. S. Baric. 2010. Recombination, reservoirs, and the modular spike: Mechanisms of coronavirus cross-species transmission. Journal of Virology 84(7):3134-3146.
Haagmans, B. L., S. H. S. Al Dhahiry, C. B. E. M. Reusken, V. S. Raj, M. Galiano, R. Myers, G.-J. Godeke, M. Jonges, E. Farag, A. Diab, H. Ghobashy, F. Alhajri, M. Al-Thani, S. A. Al-Marri, H. E. Al Romaihi, A. Al Khal, A. Bermingham, A. D. M. E. Osterhaus, M. M. AlHajri, and M. P. G. Koopmans. 2014. Middle East respiratory syndrome coronavirus in dromedary camels: An outbreak investigation. Lancet Infectious Diseases 14(2):140-145.
Hamming, I., W. Timens, M. L. C. Bulthuis, A. T. Lely, G. J. Navis, and H. van Goor. 2004. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. Journal of Pathology 203(2):631-637.
Hemida, M. G., R. A. Perera, P. Wang, M. A. Alhammadi, L. Y. Siu, M. Li, L. L. Poon, L. Saif, A. Alnaeem, and M. Peiris. 2013. Middle East respiratory syndrome (MERS) coronavirus seroprevalence in domestic livestock in Saudi Arabia, 2010 to 2013. Eurosurveillance 18(50):21-27.
Huynh, J., S. Li, B. Yount, A. Smith, L. Sturges, J. C. Olsen, J. Nagel, J. B. Johnson, S. Agnihothram, J. E. Gates, M. B. Frieman, R. S. Baric, and E. F. Donaldson. 2012. Evidence supporting a Zoonotic origin of human coronavirus strain NL63. Journal of Virology 86(23):12816-12825.
Ithete, N. L., S. Stoffberg, V. M. Corman, V. M. Cottontail, L. R. Richards, M. C. Schoeman, C. Drosten, J. F. Drexler, and W. Preiser. 2013. Close relative of human Middle East respiratory syndrome coronavirus in bat, South Africa. Emerging Infectious Diseases 19(10):1697-1699.
Ksiazek, T. G., D. Erdman, C. S. Goldsmith, S. R. Zaki, T. Peret, S. Emery, S. Tong, C. Urbani, J. A. Comer, W. Lim, P. E. Rollin, S. F. Dowell, A. E. Ling, C. D. Humphrey, W. J. Shieh, J. Guarner, C. D. Paddock, P. Rota, B. Fields, J. DeRisi, J. Y. Yang, N. Cox, J. M. Hughes, J. W. LeDuc, W. J. Bellini, and L. J. Anderson. 2003. A novel coronavirus associated with severe acute respiratory syndrome. New England Journal of Medicine 348(20):1953-1966.
Lau, S. K. P., P. C. Y. Woo, K. S. M. Li, Y. Huang, H. W. Tsoi, B. H. L. Wong, S. S. Y. Wong, S. Y. Leung, K. H. Chan, and K. Y. Yuen. 2005. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proceedings of the National Academy of Sciences of the United States of America 102(39):14040-14045.
Lau, S. K. P., K. S. M. Li, A. K. L. Tsang, C. T. Shek, M. Wang, G. K. Y. Choi, R. T. Guo, B. H. L. Wong, R. W. S. Poon, C. S. F. Lam, S. Y. H. Wang, R. Y. Y. Fan, K. H. Chan, B. J. Zheng, P. C. Y. Woo, and K. Y. Yuen. 2012. Recent transmission of a novel alphacoronavirus, bat coronavirus HKU10, from Leschenault’s rousettes to Pomona leaf-nosed bats: First evidence of interspecies transmission of coronavirus between bats of different suborders. Journal of Virology 86(21):11906-11918.
Lau, S. K. P., K. S. M. Li, A. K. L. Tsang, C. S. F. Lam, S. Ahmed, H. L. Chen, K. H. Chan, P. C. Y. Woo, and K. Y. Yuen. 2013. Genetic characterization of betacoronavirus lineage c viruses in bats reveals marked sequence divergence in the spike protein of pipistrellus bat coronavirus HKU5 in Japanese pipistrelle: Implications for the origin of the novel Middle East respiratory syndrome coronavirus. Journal of Virology 87(15):8638-8650.
Li, W. D., Z. L. Shi, M. Yu, W. Z. Ren, C. Smith, J. H. Epstein, H. Z. Wang, G. Crameri, Z. H. Hu, H. J. Zhang, J. H. Zhang, J. McEachern, H. Field, P. Daszak, B. T. Eaton, S. Y. Zhang, and L. F. Wang. 2005. Bats are natural reservoirs of SARS-like coronaviruses. Science 310(5748):676-679.
Memish, Z. A., N. Mishra, K. J. Olival, S. F. Fagbo, V. Kapoor, J. H. Epstein, R. AlHakeem, A. Durosinloun, M. A. Asmari, A. Islam, A. Kapoor, T. Briese, P. Daszak, A. A. A. Rabeeah, and W. I. Lipkin. 2013. Middle East respiratory syndrome coronavirus in bats, Saudi Arabia. Emerging Infectious Diseases 19(11):1819-1823.
Meyer, B., M. A. Mueller, V. M. Corman, C. B. E. M. Reusken, D. Ritz, G.-J. Godeke, E. Lattwein, S. Kallies, A. Siemens, J. van Beek, J. F. Drexler, D. Muth, B.-J. Bosch, U. Wernery, M. P. G. Koopmans, R. Wernery, and C. Drosten. 2014. Antibodies against MERS coronavirus in dromedaries, United Arab Emirates, 2003 and 2013. Emerging Infectious Diseases 20(4):552-559.
Milne-Price, S., K. L. Miazgowicz, and V. J. Munster. 2014. The emergence of the Middle East respiratory syndrome coronavirus. Pathogens and Disease 71(2):119-134.
Muller, M. A., V. S. Raj, D. Muth, B. Meyer, S. Kallies, S. L. Smits, R. Wollny, T. M. Bestebroer, S. Specht, T. Suliman, K. Zimmermann, T. Binger, I. Eckerle, M. Tschapka, A. M. Zaki, A. D. M. E. Osterhaus, R. A. M. Fouchier, B. L. Haagmans, and C. Drosten. 2012. Human coronavirus EMC does not require the SARS-coronavirus receptor and maintains broad replicative capability in mammalian cell lines. mbio 3(6):e00515-12.
Nahar, N., R. Sultana, E. S. Gurley, M. J. Hossain, and S. P. Luby. 2010. Date palm sap collection: Exploring opportunities to prevent Nipah transmission. EcoHealth 7(2):196-203.
Nicholls, J. M., L. L. M. Poon, K. C. Lee, W. F. Ng, S. T. Lai, C. Y. Leung, C. M. Chu, P. K. Hui, K. L. Mak, W. Lim, K. W. Yan, K. H. Chan, N. C. Tsang, Y. Guan, K. Y. Yuen, and J. S. M. Peiris. 2003. Lung pathology of fatal severe acute respiratory syndrome. Lancet 361(9371):1773-1778.
Nowotny, N., and J. Kolodziejek. 2014. Middle East respiratory syndrome coronavirus (MERS-CoV) in dromedary camels, Oman, 2013. European Communicable Disease Bulletin 19(16):pii-20781.
Osborne, C., P. M. Cryan, T. J. O’Shea, L. M. Oko, C. Ndaluka, C. H. Calisher, A. D. Berglund, M. L. Klavetter, R. A. Bowen, K. V. Holmes, and S. R. Dominguez. 2011. Alphacoronaviruses in New World bats: Prevalence, persistence, phylogeny, and potential for interaction with humans. PLoS ONE 6(5):e19156.
Perera, R. A., P. Wang, M. R. Gomaa, R. El-Shesheny, A. Kandeil, O. Bagato, L. Y. Siu, M. M. Shehata, A. S. Kayed, Y. Moatasim, M. Li, L. L. Poon, Y. Guan, R. J. Webby, M. A. Ali, J. S. Peiris, and G. Kayali. 2013. Seroepidemiology for MERS coronavirus using microneutralisation and pseudoparticle virus neutralisation assays reveal a high prevalence of antibody in dromedary camels in Egypt, June 2013. Eurosurveillance 18(36):8-14.
Raj, V. S., H. H. Mou, S. L. Smits, D. H. W. Dekkers, M. A. Muller, R. Dijkman, D. Muth, J. A. A. Demmers, A. Zaki, R. A. M. Fouchier, V. Thiel, C. Drosten, P. J. M. Rottier, A. Osterhaus, B. J. Bosch, and B. L. Haagmans. 2013. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 495(7440):251-254.
Ren, W., X. Qu, W. Li, Z. Han, M. Yu, P. Zhou, S.-Y. Zhang, L.-F. Wang, H. Deng, and Z. Shi. 2008. Difference in receptor usage between severe acute respiratory syndrome (SARS) coronavirus and SARS-like coronavirus of bat origin. Journal of Virology 82(4):1899-1907.
Reusken, C. B., M. Ababneh, V. S. Raj, B. Meyer, A. Eljarah, S. Abutarbush, G. J. Godeke, T. M. Bestebroer, I. Zutt, M. A. Mueller, B. J. Bosch, P. J. Rottier, A. D. Osterhaus, C. Drosten, B. L. Haagmans, and M. P. Koopmans. 2013a. Middle East respiratory syndrome coronavirus (MERS-CoV) serology in major livestock species in an affected region in Jordan, June to September 2013. Eurosurveillance 18(50):14-20.
Reusken, C. B., B. L. Haagmans, M. A. Mueller, C. Gutierrez, G.-J. Godeke, B. Meyer, D. Muth, V. S. Raj, L. Smits-De Vries, V. M. Corman, J.-F. Drexler, S. L. Smits, Y. E. El Tahir, R. De Sousa, J. van Beek, N. Nowotny, K. van Maanen, E. Hidalgo-Hermoso, B.-J. Bosch, P. Rottier, A. Osterhaus, C. Gortazar-Schmidt, C. Drosten, and M. P. G. Koopmans. 2013b. Middle East respiratory syndrome coronavirus neutralising serum antibodies in dromedary camels: A comparative serological study. Lancet Infectious Diseases 13(10):859-866.
Reusken, C. B., E. A. Farag, M. Jonges, G. J. Godeke, A. M. El-Sayed, S. D. Pas, V. S. Raj, K. A. Mohran, H. A. Moussa, H. Ghobashy, F. Alhajri, A. K. Ibrahim, B. J. Bosch, S. K. Pasha, H. E. Al-Romaihi, M. Al-Thani, S. A. Al-Marri, M. M. AlHajri, B. L. Haagmans, and M. P. Koopmans. 2014a. Middle East respiratory syndrome coronavirus (MERS-CoV) RNA and neutralising antibodies in milk collected according to local customs from dromedary camels, Qatar, April 2014. Eurosurveillance 19(23):8-12.
Reusken, C. B., L. Messadi, A. Feyisa, H. Ularamu, G.-J. Godeke, A. Danmarwa, F. Dawo, M. Jemli, S. Melaku, D. Shamaki, Y. Woma, Y. Wungak, E. Z. Gebremedhin, I. Zutt, B.-J. Bosch, B. L. Haagmans, and M. P. G. Koopmans. 2014b. Geographic distribution of MERS coronavirus among dromedary camels, Africa. Emerging Infectious Diseases 20(8):1370-1374.
Rihtaric, D., P. Hostnik, A. Steyer, J. Grom, and I. Toplak. 2010. Identification of SARS-like coronaviruses in horseshoe bats (Rhinolophus hipposideros) in Slovenia. Archives of Virology 155(4): 507-514.
Shirato, K., K. Maeda, S. Tsuda, K. Suzuki, S. Watanabe, H. Shimoda, N. Ueda, K. Iha, S. Taniguchi, S. Kyuwa, D. Endoh, S. Matsuyama, I. Kurane, M. Saijo, S. Morikawa, Y. Yoshikawa, H. Akashi, and T. Mizutani. 2012. Detection of bat coronaviruses from Miniopterus fuliginosus in Japan. Virus Genes 44(1):40-44.
Tao, Y., K. Tang, M. Shi, C. Conrardy, K. S. M. Li, S. K. P. Lau, L. J. Anderson, and S. X. Tong. 2012. Genomic characterization of seven distinct bat coronaviruses in Kenya. Virus Research 167(1):67-73.
Tong, S. X., C. Conrardy, S. Ruone, I. V. Kuzmin, X. L. Guo, Y. Tao, M. Niezgoda, L. Haynes, B. Agwanda, R. F. Breiman, L. J. Anderson, and C. E. Rupprecht. 2009. Detection of novel SARS-like and other coronaviruses in bats from Kenya. Emerging Infectious Diseases 15(3):482-485.
van Boheemen, S., M. de Graaf, C. Lauber, T. M. Bestebroer, V. S. Raj, A. M. Zaki, A. D. M. E. Osterhaus, B. L. Haagmans, A. E. Gorbalenya, E. J. Snijder, and R. A. M. Fouchier. 2012. Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. mbio 3(6):e00473-12.
van Doremalen, N., T. Bushmaker, W. B. Karesh, and V. J. Munster. 2014. Stability of Middle East respiratory syndrome coronavirus in milk. Emerging Infectious Diseases 20(7):1263-1264.
Wacharapluesadee, S., C. Sintunawa, T. Kaewpom, K. Khongnomnan, K. J. Olival, J. H. Epstein, A. Rodpan, P. Sangsri, N. Intarut, A. Chindamporn, K. Suksawa, and T. Hemachudha. 2013. Identification of group C betacoronavirus from bat guano fertilizer, Thailand. Emerging Infectious Diseases 19(8):1349-1351.
WHO (World Health Organization). 2014. Middle East respiratory syndrome coronavirus (MERS-CoV) - update. http://www.who.int/csr/don/2014_07_23_mers/en (accessed August 29, 2014).
Woo, P. C. Y., S. K. P. Lau, K. S. M. Li, A. K. L. Tsang, and K.-Y. Yuen. 2012. Genetic relatedness of the novel human group C betacoronavirus to Tylonycteris bat coronavirus HKU4 and Pipistrellus bat coronavirus HKU5. Emerging Microbes & Infections 1.