STUDYING ZOONOTIC DISEASES IN THE NATURAL HOST
John W. Lowenthal,10Michelle L. Baker,10Cameron R. Stewart,10Christopher Cowled,10Celine Deffrasnes,10Lin-Fa Wang,10,11and Andrew G. D. Bean10
Introduction
Several factors, including the recent growth and geographic expansion of human populations and the intensification of agriculture combined with habitat disruption caused by climate change and deforestation, has meant that now, more than ever, there is a greater risk of emerging infectious diseases (EIDs) being transmitted to humans from wild and domesticated animals (Jones et al., 2008; Taylor et al., 2001). Moreover, increased global travel means there is a greater likelihood that EIDs will rapidly spread. Over the past three decades the incidence of EIDs has risen in humans, with around 70 percent being zoonotic in nature, and the majority being caused by viruses (Jones et al., 2008; Taylor et al., 2001) (Figure A6-1). Containment of these EID outbreaks has often been difficult owing to their unpredictability and the absence of effective control measures, such as vaccines and antiviral therapeutics. In addition, there is a lack of essential knowledge of the host immune responses induced by zoonotic viruses, particularly those that provide protection.
The Impact of EIDs
The World Health Organization has warned that the source of the next human pandemic is likely to be zoonotic and that wildlife is a prime culprit (see http://www.who.int/zoonoses/diseases/en).
While the current list of known EIDs is a major concern, it is the unknown EIDs out there, with a potential for efficient human-to-human transmission, that may pose the biggest threat. Over the past decade there have been a number of epidemics, raising the concern that they are precursors to a pandemic.
_______________
10 CSIRO Biosecurity Flagship, Australian Animal Health Laboratory, Geelong, Victoria, 3220, Australia.
11 Program in Emerging Infectious Diseases, Duke-NUS Graduate Medical School, Singapore 169857, Singapore.
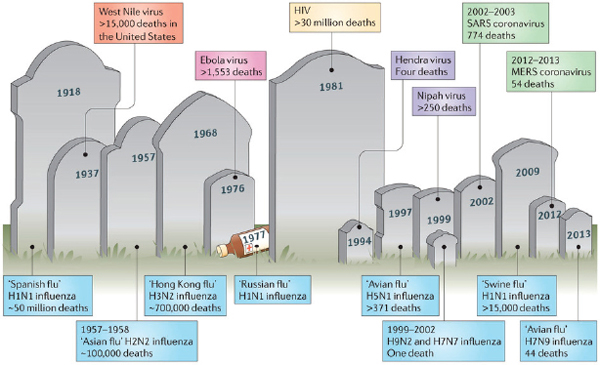
FIGURE A6-1 Emergence of zoonoses. Over the past century, humanity has witnessed the emergence of numerous zoonotic infections that have resulted in varying degrees of human fatalities. Influenza viruses originating from birds account for an important portion of these deaths and recently many new zoonotic viruses originating in bats, such as Hendra virus, Nipah virus, and severe acute respiratory syndrome coronavirus (SARS-CoV), have caused outbreaks with high mortality rates.
NOTE: Human immunodeficiency virus (HIV), West Nile virus (WNV), Novel coronavirus (MERS-CoV). H7N9 avian influenza as of August 11, 2013; MERS-CoV as of September 20, 2013.
SOURCE: Bean et al., 2013.
- The SARS epidemic in 2003–2004 claimed more than 800 lives and cost more than $80 billion to the global economy. It was shown to have involved virus transmission from bats to civet cats to humans.12
- In 2012 a novel coronavirus emerged in the Middle East (MERS-CoV) (Zaki et al., 2012) with a 30 percent mortality rate for the more than 500 cases so far confirmed, raising the concerns for a SARS-like pandemic.13
- Highly pathogenic H5N1 avian influenza virus has decimated poultry production in Asia and claimed more than 350 lives since 2003 with regular outbreaks continuing.14
- A new strain of virus (H7N9), which has never previously been seen in humans, appeared in April 2013. While this current strain of the avian influenza is not in a form that is able to transmit from human to human, there is still the possibility that it could mutate and trigger a serious pandemic (Yu et al., 2013).15
- Hendra virus in Australia, Nipah virus in Malaysia and Bangladesh, and hemorrhagic fever viruses (Ebola and Marburg) have over the past two decades emerged from bats via intermediate hosts such as horses and pigs to infect and kill humans.
A One Health Approach
Numerous emerging disease concerns are closely connected to the ever-increasing interactions between humans and wildlife. A number of drivers are associated with the emergence of disease from wildlife and spread to and among humans (Patz et al., 2004):
- The escalated need for food production to meet present and future demand has led to the intrusion of agriculture into previously untouched areas of the native environment (Pulliam et al., 2012).
- The impact of climate change has resulted in disturbances in ecosystems and a redistribution of disease reservoirs and vectors.
- Increased globalization and travel has significantly increased the chance, extent, and spread at which disease transmission occurs.
With this in mind, there has been a growing initiative to more closely address this animal–human–ecosystem interface. The term One Health describes a collaborative effort from multiple disciplines to support a holistic approach in the development of health strategies for people, animals, and the environment.
_______________
12 See http://www.who.int/csr/don/archive/disease/severe_acute_respiratory_syndrome/en/index.html.
13 See http://www.who.int/csr/disease/coronavirus_infections/archive_updates/en/index.html.
14 See http://www.who.int/influenza/human_animal_interface/H5N1_cumulative_table_archives/en.
15 See http://www.who.int/influenza/human_animal_interface/influenza_h7n9/en/index.html.
One Health unifies clinical and veterinary health and directly links this with environmental health research. The development of a framework directed at strengthening alliances between these sectors has facilitated the development and application of effective and sustainable community health strategies. There is a growing view that a One Health approach will be critically important for our preparedness for the next zoonotic pandemic.
Coevolution of Hosts and Pathogens
In many cases, natural host reservoirs seem to coexist with human pathogens, including zoonotic viruses, in the absence of disease, demonstrating the importance of this coevolutionary relationship. Bats are one example of a group of mammals that has a long coevolutionary history with the viruses they harbor. Although infection with viruses such as SARS, Hendra, and Ebola appears to result in little pathology in bats, infection of other susceptible hosts often causes severe disease and has fatal consequences. The evolution of unique immune mechanisms for the control of viral replication may be a mechanism for the ability of bats to coexist with viruses. Furthermore, it has been suggested that the severe pathology and disease that often occurs as a result of the spillover of viruses into other vertebrate hosts may also result from the disturbance of this finely tuned interaction of viral proteins with their targets in host cells (Wang et al., 2011). Understanding how bats and other natural reservoirs, such as birds, coexist with human pathogens could potentially lead to the discovery of mechanisms that control viral replication, and these could eventually be applied to help protect other susceptible species.
Animal Models for Zoonotic Pathogens
Traditional Animal Models
Mouse models have been fundamental to our understanding of immune responses to infection and disease outcomes. Indeed, mice have become the traditional “workhorse” because of their ease of handling, fast generation time, and the ready availability of mouse-specific reagents (Legrand et al., 2006). However, for a better understanding of EIDs, the laboratory mouse may not be the most appropriate model. There are often many differences in the symptoms of disease between the natural transmission and human hosts. Frequently, zoonotic infections appear as asymptomatic and nonlethal in the natural reservoir host, yet induce severe and potentially lethal disease in humans or other spillover hosts. Nevertheless, there are numerous factors that are likely to contribute to these differences including anatomical, physiological, metabolic, and behavioral traits as well as how the immune systems of these hosts interact with the same disease agent.
Nontraditional Laboratory Animal Models
There are many examples in the literature where nontraditional animal models have been highly informative for our understanding of host responses to pathogens (Hein and Griebel, 2003). For several decades the chicken has been used to study immunology, sexual development and developmental biology of the limbs, and the nervous system and brain (Le Douarin and Dieterlen-Lievre, 2013). Indeed, with the exception of mice and humans, arguably the most thoroughly characterized immune system is that of the chicken. Other animals have also provided a wealth of knowledge concerning aspects of immunity and human disease. For example, bats are now being used to study several emerging viruses such as Hendra, and ferrets are widely accepted as an excellent model for influenza infection—they are naturally susceptible to infection with human influenza viruses, and the disease pathology they develop resembles that of humans infected with influenza (Belser et al., 2011).
Natural Reservoir Hosts and Spillover Events
Why is it important to use both the natural animal reservoir and spillover host to study the host responses to zoonotic pathogens? In particular, understanding the differences between the immune systems of domesticated and wild animal hosts and comparing them to that of humans is crucial for unravelling the complex disease mechanisms involved in zoonotic infections (Figure A6-2). Furthermore, by studying the pathogen in its natural host we may be able to devise efficient control measures in that host, thereby disrupting their transmission to humans. This has important implications for predicting, preventing, and controlling spillover events and for the development of novel therapeutics and diagnostics. Table A6-1 lists selected zoonotic viruses and their reservoir hosts, susceptible hosts, and transmission hosts.
One example is the case of highly pathogenic avian influenza (HPAI). Waterfowl are natural hosts for avian influenza viruses and after infection with HPAI develop what appears to be a limited inflammatory response of the respiratory system, usually with little or no mortality. By contrast, in chickens and in humans HPAI viruses can induce a rapid and strong inflammatory response and potential hypercytokinemia, often referred to as a cytokine storm (Clark, 2007), and the infection may become systemic and induce severe disease symptoms (Karpala et al., 2011; Lee et al., 2007; Tisoncik et al., 2012). Chickens are acutely susceptible to infection with H5N1 strains of HPAI, which typically cause death within 18–36 hours. Studying waterfowl, such as ducks, and comparing their immune responses to influenza virus with those of the chicken may provide invaluable insight into the “aberrant” immune reactions that occurs in influenza spillover hosts, such as chickens, pigs, and humans (Figure A6-3).
Another interesting example is Hendra virus, which does not cause disease in fruit bats, the natural reservoir for this infection, but induces severe disease
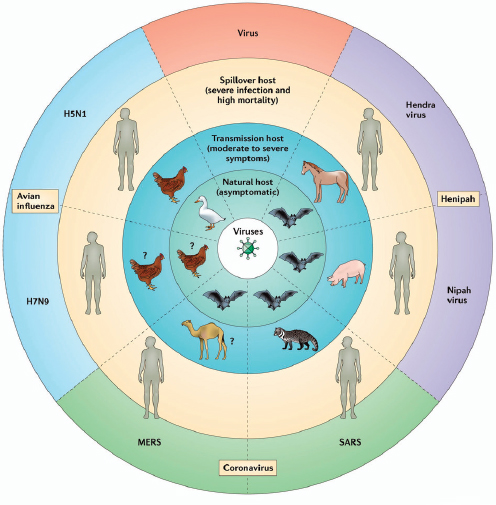
FIGURE A6-2 The outcome of disease severity is influenced by the host–pathogen interaction. Many zoonotic agents cause little or no signs of disease in their natural host such as wild birds and bats, while transmission hosts may present symptoms ranging from moderate (such as pigs for AI) to severe (such as horses for HeV) signs. The terminal or spillover host, such as humans in the case of H5N1 and HeV infections, can present with very severe symptoms and high mortality rates. For some of the most recently emerging EIDs such as H7N9 and MERS-CoV, natural and transmission hosts have not been identified.
SOURCE: Bean et al., 2013.
in horses and humans (reviewed in Mahalingam et al., 2012). The study of disease pathogenesis and immune responses to Hendra virus in horses has led to the development of a horse vaccine that will help reduce the rates of Hendra virus transmission from horses to humans (Mahalingam et al., 2012). Nevertheless, studying zoonotic viruses in wildlife is complex. For example, the fact that
| Disease (virus) | Known Reservoir Hosts | Other Susceptible Hosts | Transmitted to Humans by | |
| Influenzaa | ||||
| Avian (H5N1, H7N9, H7N7, H9N2, H3N2, and others) | Waterfowl | Bats | Chickens | |
| Wild birds | Cats | |||
| Swine (H1N1, H3N2) | Dogs | |||
| Pigs | Ferrets | Pigs | ||
| Foxes | ||||
| Pigs | ||||
| Horses | ||||
| Poultry (chicken, duck, turkey) | ||||
| Marine mammals | ||||
| SARS (SARS coronavirus)b | Bats | Civet cats | Civet cats | |
| Dengue fever (Dengue virus)c | Primates | Unknown | Mosquitoes | |
| Hendra (Hendra virus)d | Bats | Horses | Horses | |
| Ferrets | ||||
| Rabies (Rabies virus and other lyssaviruses)e | Bats | Cats | Bats | |
| Cattle | Dogs | |||
| Coyotes | ||||
| Dogs | ||||
| Foxes | ||||
| Horses | ||||
| Mongooses | ||||
| Primates | ||||
| Raccoons | ||||
| Sheep | ||||
| Skunks | ||||
| Wolves | ||||
| Ebola viral hemorrhagic fever (Ebola virus)f | Bats | Primates | Primates | |
| Japanese encephalitis (Japanese encephalitis virus)g | Pigs | Horses | Bats | |
| Mosquitoes | ||||
| Wild birds | ||||
| West Nile virus encephalitis (West Nile virus)h | Domestic and wild birds | Bats | Mosquitoes | |
| Camels | Birds | |||
| Horses | ||||
| Marine mammals | ||||
| Reptiles | ||||
| > 30 vertebrate species | ||||
a Centers for Disease Control and Prevention, 2013; World Health Organization, 2013a; World Organisation for Animal Health, 2013; Reperant et al., 2012; Swenson et al., 2010; Tong et al., 2012.
b Centers for Disease Control and Prevention, 2013; Shi and Hu, 2008.
c Centers for Disease Control and Prevention, 2013; Carver et al., 2009.
d Centers for Disease Control and Prevention, 2013; Clayton et al., 2013.
e Centers for Disease Control and Prevention, 2013; World Organisation for Animal Health, 2013; Hatz et al., 2012; Rupprecht et al., 2011.
f Centers for Disease Control and Prevention, 2013; World Health Organization, 2013a.
g Centers for Disease Control and Prevention, 2013.
h Centers for Disease Control and Prevention, 2013; World Organisation for Animal Health, 2013.
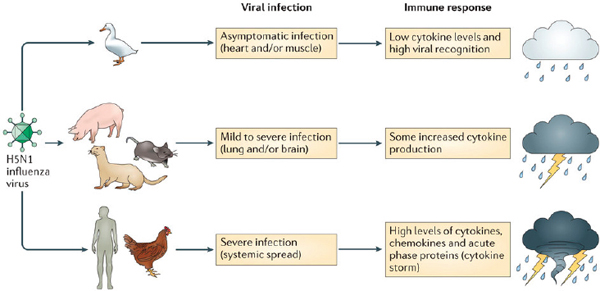
FIGURE A6-3 The host immune response to an infection influences the disease outcome. Infection with H5N1 can cause very different disease outcomes in different reservoir and spillover hosts. Waterfowl such as wild ducks are the natural virus host and develop a limited inflammatory response associated with low levels of cytokine expression. Intermediate hosts such as mice, pigs, and ferrets, often used as laboratory models, display mild to severe symptoms (depending on the H5N1 virus strain used) associated with elevated levels of proinflammatory cytokines. By contrast, spillover hosts such as chickens and humans display a rapid and strong inflammatory response, often referred to as hypercytokinemia (or cytokine storm), and the infection becomes systemic, causing severe disease symptoms and high mortality rates.
SOURCE: Bean et al., 2013.
infectious agents, such as henipaviruses and lyssaviruses, that cause lethal diseases in humans can coexist peacefully with bats raises many questions. It is still unclear how the bat immune system keeps these pathogens in check and enables host survival while allowing enough viral replication to facilitate transmission of the virus to spillover hosts. Other outstanding questions are how host–pathogen interactions are influenced by the genetics of the population, environmental factors, changes in demographics, food supply, co-infections, and interactions with other species as well as particular physiological, anatomical, and metabolic features of the host.
Despite the benefits of using nontraditional animal models, working with these systems presents a number of challenges. These may include a limited access to suitable subjects of the desired species, particularly in the case of fauna that may need to be wild caught and may be members of a species of high conservation value; lack of knowledge or experience in handling and husbandry methods suitable for high levels of biocontainment that will also meet contemporary ethics and welfare requirements; a paucity of reagents such as species-specific antibodies required to conduct traditional immunological experiments; and limited or no genome sequence information for the purpose of developing molecular biology tools. For species not traditionally used in the laboratory, new policies and procedures for housing, husbandry, restraint, and sampling may need to be developed on a case-by-case basis. In such instances, staff within the veterinary departments of zoological parks, zoologists, and veterinarians who specialize in exotic or unusual pets are valuable resources.
Comparative Immunology Studies
A major goal of studying immunology in natural hosts is to explain how infection with the same pathogen can have such vastly different outcomes in different species. Investigation of immunity in natural hosts is therefore an area that may yield significant discoveries and illuminate the nature of successful immune responses to agents that are typically associated with adverse disease outcomes in humans and other species. There is an extensive range of technologies that have been used to understand the pathogenesis and immune responses in reservoir and transmission species. These include molecular genetic tools such as DNA and RNA sequencing, transcriptomics, RNAi, miRNA, and genome-editing meganucleases. In addition, posttranslational analysis tools such proteomics, kinomics, and other protein modifications provide additional information.
Comparative genomics is a powerful approach for identifying genetic determinants underlying phenotypical differences between species. The recent advances in high-throughput sequencing techniques have facilitated whole-genome sequencing of a large number of species, including some that are reservoir hosts of important zoonotic viruses, and others that are susceptible to disease caused by those same viruses. Comparative analysis of these genomes can identify gene
candidates for disease-susceptibility or disease-resistance phenotypes. Quantitative transcriptomics by sequencing, for example RNA-sequencing (RNA-seq) and small RNA-seq, have enormous potential for identifying crucial differences between species in host responses to virus infection. These analyses can provide unprecedented detail and can be easily performed on species for which no species-specific reagents are available, as is the case for the natural hosts of many zoonotic viruses. Another application with strong potential in this field is genome-wide screening using RNAi (reviewed in Meliopoulos et al., 2012). As genome sequencing and RNAi technologies continually develop, it will soon be possible to compare host genes required for virus replication across virus-susceptible and virus-resistant species, such as chickens and ducks in the case of HPAI.
A major goal of sequencing bat genomes was to understand the genetic basis of virus–host interactions in a natural reservoir host (Zhang et al., 2013). More recently, analysis of the duck genome and virus-infected duck transcriptome has continued this trend. It has been observed that avians generally encode fewer cytokines than mammals, and appear to lack α- and θ-defensins. In contrast, ducks featured lineage-specific duplications of β-defensin and butyrophilin-like genes, suggesting a possible connection with the fundamental differences in disease outcome observed between chickens and ducks infected with HPAI (Huang et al., 2013). An area of potential interest will be to revisit genome sequences of animals with the hindsight that they are natural hosts for diseases of relevance to humans, for example the cat, dog, goat, armadillo, and camel genomes.
Clinical Implications of Wild Animal Studies
There are a number of key practical outcomes that research in wild and livestock animal species could achieve. For example, if we better understand influenza infection in pigs and birds, will we be better able to predict where the next pandemic might emerge or be able to develop vaccines or antivirals for the animals that prevent the crossover to humans? For example, if we understand how the bat and duck immune system responds to viruses, will this help us to develop new therapeutics and vaccines for preventing fatal infections caused by these viruses in humans? Can we engineer livestock that will be resilient to EIDs, such as influenza and Nipah viruses, thereby blocking the transmission cycle?
The ability of the segmented influenza genome to continually re-sort within different animal host species has a crucial impact on the epidemiology of influenza outbreaks. For example, the combination of viral gene segments from four virus strains circulating in three different species (human, swine, and poultry) led to the emergence of swine-origin influenza virus (S-OIV) in the human population (Itoh et al., 2009). The relevance of a lack of preexisting immunity became apparent as mortality and morbidity began to rise sharply in the 20- to 40-year-old age group (Chowell et al., 2009), while the older age group was associated with a lower infection risk (Fisman et al., 2009) that was due to a higher prevalence
of preexisting cross-antibodies that cross-reacted with the 2009 H1N1 virus. The impact of a novel influenza virus arising from wildlife species was again felt in February 2013, when an avian-origin influenza virus emerged in Zhejiang, China, causing more than 400 reported human infections and 100 deaths. This virus appears to have emerged from the mixing of influenza viruses from several avian sources, including ducks and wild birds (Chen et al., 2013). It was the first report of an H7N9 influenza virus infecting humans, and was met with little preexisting immunity. Furthermore, vaccines against H7N9 are predicted to be poorly immunogenic due to few T cell epitopes on the H7 molecule compared to other HA subtypes (De Groot et al., 2013). Despite this particular virus being highly pathogenic in humans, natural infections with H7N9 viruses in chickens, ducks, and other birds are asymptomatic and elicit an immune response that can be detected serologically (WHO, 2013b). This is in stark contrast to H5N1 and H7N7 where disease in humans was associated with a highly pathogenic phenotype in poultry (Perdue and Swayne, 2005). The immunologic component to this disease in birds suggests that further study of these emerging viruses in poultry and other bird species are required to understand factors influencing disease susceptibility and transmission.
Identification of key differences in immune pathways between susceptible and nonsusceptible hosts may offer clues to develop disease intervention strategies. As mentioned previously, for HPAI infection, ducks and chickens represent natural and spillover hosts, respectively. Chickens have lost expression of the innate immune sensor retinoic acid-inducible protein I (RIG-I), which may be an important clue in explaining why they suffer close to 100 percent mortality from HPAI. On the other hand, ducks (which have intact RIG-I expression) develop only mild symptoms in response to HPAI infection and usually survive. It has been shown that the transfection of duck RIG-I into chicken cells induced IFN-β promoter activity and limited the replication of low and highly pathogenic avian influenza strains, suggesting a key role of RIG-I in the ability of ducks to be resistant to influenza-mediated disease (Barber et al., 2010). While the consequences of the absence of RIG-I in chickens to other viral infection is not understood, it would nevertheless be of interest to generate transgenic chickens that express duck RIG-I and investigate whether these animals are less susceptible to disease following HPAI infection or indeed infection with other viruses.
Concluding Remarks
What can we learn from nature’s experiments? Studying the responses to zoonotic pathogens in the natural reservoir host and comparing it to the responses in spillover hosts will help identify key processes in disease susceptibility and transmission. With the current emphasis on a One Health approach, researchers are turning to an analysis of the immune response of natural host species for a greater understanding of emerging zoonotic diseases. Together with adoption of
new genomic information and genome editing capability we can identify new strategies to prevent and minimize the impact of EIDs and enhance our pandemic preparedness.
References
Barber, M. R., J. R. Aldridge, Jr., R. G. Webster, and K. E. Magor. 2010. Association of RIG-I with innate immunity of ducks to influenza. Proceedings of the National Academy of Sciences of the United States of America 107(13):5913-5918.
Bean, A. G. D., M. L. Baker, C. R. Stewart, C. Cowled, C. Deffrasnes, L.-F. Wang, and J. W. Lowenthal. 2013. Studying immunity to zoonotic diseases in the natural host—keeping it real. Nature Reviews: Immunology 13(12):851-861.
Belser, J. A., J. M. Katz, and T. M. Tumpey. 2011. The ferret as a model organism to study influenza A virus infection. Disease Models & Mechanisms 4(5):575-579.
Carver, S., A. Bestall, A. Jardine, and R. S. Ostfeld. 2009. Influence of hosts on the ecology of arboviral transmission: Potential mechanisms influencing dengue, Murray Valley encephalitis, and Ross River virus in Australia. Vector Borne and Zoonotic Diseases 9(1):51-64.
Chen, Y., W. Liang, S. Yang, N. Wu, H. Gao, J. Sheng, H. Yao, J. Wo, Q. Fang, D. Cui, Y. Li, X. Yao, Y. Zhang, H. Wu, S. Zheng, H. Diao, S. Xia, Y. Zhang, K. H. Chan, H. W. Tsoi, J. L. Teng, W. Song, P. Wang, S. Y. Lau, M. Zheng, J. F. Chan, K. K. To, H. Chen, L. Li, and K. Y. Yuen. 2013. Human infections with the emerging avian influenza A H7N9 virus from wet market poultry: Clinical analysis and characterisation of viral genome. Lancet 381(9881):1916-1925.
Chowell, G., S. M. Bertozzi, M. A. Colchero, H. Lopez-Gatell, C. Alpuche-Aranda, M. Hernandez, and M. A. Miller. 2009. Severe respiratory disease concurrent with the circulation of H1N1 influenza. New England Journal of Medicine 361(7):674-679.
Clark, I. A. 2007. The advent of the cytokine storm. Immunology and Cell Biology 85(4):271-273.
Clayton, B. A., L. F. Wang, and G. A. Marsh. 2013. Henipaviruses: An updated review focusing on the pteropid reservoir and features of transmission. Zoonoses and Public Health 60(1):69-83.
De Groot, A. S., M. Ardito, F. Terry, L. Levitz, T. M. Ross, L. Moise, and W. Martin. 2013. Low immunogenicity predicted for emerging avian-origin H7N9: Implication for influenza vaccine design. Human Vaccines & Immunotherapeutics 9(5):950-956.
Fisman, D. N., R. Savage, J. Gubbay, C. Achonu, H. Akwar, D. J. Farrell, N. S. Crowcroft, and P. Jackson. 2009. Older age and a reduced likelihood of 2009 H1N1 virus infection. New England Journal of Medicine 361(20):2000-2001.
Hatz, C. F., E. Kuenzli, and M. Funk. 2012. Rabies: Relevance, prevention, and management in travel medicine. Infectious Disease Clinics of North America 26(3):739-753.
Hein, W. R., and P. J. Griebel. 2003. A road less travelled: Large animal models in immunological research. Nature Reviews: Immunology 3(1):79-84.
Huang, Y., Y. Li, D. W. Burt, H. Chen, Y. Zhang, W. Qian, H. Kim, S. Gan, Y. Zhao, J. Li, K. Yi, H. Feng, P. Zhu, B. Li, Q. Liu, S. Fairley, K. E. Magor, Z. Du, X. Hu, L. Goodman, H. Tafer, A. Vignal, T. Lee, K. W. Kim, Z. Sheng, Y. An, S. Searle, J. Herrero, M. A. Groenen, R. P. Crooijmans, T. Faraut, Q. Cai, R. G. Webster, J. R. Aldridge, W. C. Warren, S. Bartschat, S. Kehr, M. Marz, P. F. Stadler, J. Smith, R. H. Kraus, Y. Zhao, L. Ren, J. Fei, M. Morisson, P. Kaiser, D. K. Griffin, M. Rao, F. Pitel, J. Wang, and N. Li. 2013. The duck genome and transcriptome provide insight into an avian influenza virus reservoir species. Nature Genetics 45(7):776-783.
Itoh, Y., K. Shinya, M. Kiso, T. Watanabe, Y. Sakoda, M. Hatta, Y. Muramoto, D. Tamura, Y. SakaiTagawa, T. Noda, S. Sakabe, M. Imai, Y. Hatta, S. Watanabe, C. Li, S. Yamada, K. Fujii, S. Murakami, H. Imai, S. Kakugawa, M. Ito, R. Takano, K. Iwatsuki-Horimoto, M. Shimojima, T. Horimoto, H. Goto, K. Takahashi, A. Makino, H. Ishigaki, M. Nakayama, M. Okamatsu, K. Takahashi, D. Warshauer, P. A. Shult, R. Saito, H. Suzuki, Y. Furuta, M. Yamashita, K. Mitamura, K. Nakano, M. Nakamura, R. Brockman-Schneider, H. Mitamura, M. Yamazaki, N. Sugaya, M. Suresh, M. Ozawa, G. Neumann, J. Gern, H. Kida, K. Ogasawara, and Y. Kawaoka. 2009. In vitro and in vivo characterization of new swine-origin H1N1 influenza viruses. Nature 460(7258):1021-1025.
Jones, K. E., N. G. Patel, M. A. Levy, A. Storeygard, D. Balk, J. L. Gittleman, and P. Daszak. 2008. Global trends in emerging infectious diseases. Nature 451(7181):990-993.
Karpala, A. J., J. Bingham, K. A. Schat, L. M. Chen, R. O. Donis, J. W. Lowenthal, and A. G. Bean. 2011. Highly pathogenic (H5N1) avian influenza induces an inflammatory T helper type 1 cytokine response in the chicken. Journal of Interferon & Cytokine Research 31(4):393-400.
Le Douarin, N. M., and F. Dieterlen-Lievre. 2013. How studies on the avian embryo have opened new avenues in the understanding of development: A view about the neural and hematopoietic systems. Development Growth & Differentiation 55(1):1-14.
Lee, N., C. K. Wong, P. K. Chan, S. W. Lun, G. Lui, B. Wong, D. S. Hui, C. W. Lam, C. S. Cockram, K. W. Choi, A. C. Yeung, J. W. Tang, and J. J. Sung. 2007. Hypercytokinemia and hyperactivation of phospho-p38 mitogen-activated protein kinase in severe human influenza A virus infection. Clinical Infectious Diseases 45(6):723-731.
Legrand, N., K. Weijer, and H. Spits. 2006. Experimental models to study development and function of the human immune system in vivo. Journal of Immunology 176(4):2053-2058.
Mahalingam, S., L. J. Herrero, E. G. Playford, K. Spann, B. Herring, M. S. Rolph, D. Middleton, B. McCall, H. Field, and L. F. Wang. 2012. Hendra virus: An emerging paramyxovirus in Australia. Lancet Infectious Diseases 12(10):799-807.
Meliopoulos, V. A., L. E. Andersen, K. F. Birrer, K. J. Simpson, J. W. Lowenthal, A. G. Bean, J. Stambas, C. R. Stewart, S. M. Tompkins, V. W. van Beusechem, I. Fraser, M. Mhlanga, S. Barichievy, Q. Smith, D. Leake, J. Karpilow, A. Buck, G. Jona, and R. A. Tripp. 2012. Host gene targets for novel influenza therapies elucidated by high-throughput RNA interference screens. FASEB Journal 26(4):1372-1386.
Patz, J. A., P. Daszak, G. M. Tabor, A. A. Aguirre, M. Pearl, J. Epstein, N. D. Wolfe, A. M. Kilpatrick, J. Foufopoulos, D. Molyneux, D. J. Bradley, Working Group on Land Use Change and Disease Emergence. 2004. Unhealthy landscapes: Policy recommendations on land use change and infectious disease emergence. Environmental Health Perspectives 112(10):1092-1098.
Perdue, M. L., and D. E. Swayne. 2005. Public health risk from avian influenza viruses. Avian Diseases 49(3):317-327.
Pulliam, J. R., J. H. Epstein, J. Dushoff, S. A. Rahman, M. Bunning, A. A. Jamaluddin, A. D. Hyatt, H. E. Field, A. P. Dobson, P. Daszak, and Henipavirus Ecology Research Group. 2012. Agricultural intensification, priming for persistence and the emergence of Nipah virus: A lethal bat-borne zoonosis. Journal of the Royal Society Interface 9(66):89-101.
Reperant, L. A., T. Kuiken, and A. D. Osterhaus. 2012. Influenza viruses: from birds to humans. Human Vaccines and Immunotherapeutics 8:7-16.
Rupprecht, C. E., A. Turmelle, and I. V. Kuzmin. 2012. A perspective on lyssavirus emergence and perpetuation. Current Opinion in Virology 1:662-670.
Shi, Z., and Z. Hu. 2008. A review of studies on animal reservoirs of the SARS coronavirus. Virus Research 133(1):74-87.
Swenson, S. L., L. G. Koster, M. Jenkins-Moore, M. L. Killian, E. E. DeBess, R. J. Baker, D. Mulrooney, R. Weiss, J. Galeota, and A. Bredthauer. 2010. Natural cases of 2009 pandemic H1N1 influenza A virus in pet ferrets. Journal of Veterinary Diagnostic Investigation 22(5):784-788.
Taylor, L. H., S. M. Latham, and M. E. Woolhouse. 2001. Risk factors for human disease emergence. Philosophical Transactions of the Royal Society of London B: Biological Sciences 356(1411): 983-989.
Tisoncik, J. R., M. J. Korth, C. P. Simmons, J. Farrar, T. R. Martin, and M. G. Katze. 2012. Into the eye of the cytokine storm. Microbiology and Molecular Biology Reviews 76(1):16-32.
Tong, S., Y. Li, P. Rivailler, C. Conrardy, D. A. Castillo, L. M. Chen, S. Recuenco, J. A. Ellison, C. T. Davis, I. A. York, A. S. Turmelle, D. Moran, S. Rogers, M. Shi, Y. Tao, M. R. Weil, K. Tang, L. A. Rowe, S. Sammons, X. Xu, M. Frace, K. A. Lindblade, N. J. Cox, L. J. Anderson, C. E. Rupprecht, and R. O. Donis. 2012. A distinct lineage of influenza A virus from bats. Proceedings of the National Academy of Sciences of the United States of America 109(11):4269-4274.
Wang, L. F., P. J. Walker, and L. L. Poon. 2011. Mass extinctions, biodiversity and mitochondrial function: Are bats “special” as reservoirs for emerging viruses? Current Opinion in Virology 1(6):649-657.
WHO (World Health Organization). 2013a. Zoonoses-Diseases. http://www.who.int/zoonoses/diseases/en (accessed 2013).
WHO. 2013b. Overview of the emergence and characteristics of the avian influenza A (H7N9) virus. http://www.who.int/influenza/human_animal_interface/influenza_h7n9/WHO_H7N9_review_31May13.pdf (accessed 2013).
World Organisation for Animal Health. 2013. Animal Disease Information Summaries. http://www.oie.int/en/for-the-media/animal-diseases/animal-disease-information-summaries (accessed 2013).
Yu, H., B. J. Cowling, L. Feng, E. H. Lau, Q. Liao, T. K. Tsang, Z. Peng, P. Wu, F. Liu, V. J. Fang, H. Zhang, M. Li, L. Zeng, Z. Xu, Z. Li, H. Luo, Q. Li, Z. Feng, B. Cao, W. Yang, J. T. Wu, Y. Wang, and G. M. Leung. 2013. Human infection with avian influenza A H7N9 virus: An assessment of clinical severity. Lancet 382(9887):138-145.
Zaki, A. M., S. van Boheemen, T. M. Bestebroer, A. D. Osterhaus, and R. A. Fouchier. 2012. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. New England Journal of Medicine 367(19):1814-1820.
Zhang, G., C. Cowled, Z. Shi, Z. Huang, K. A. Bishop-Lilly, X. Fang, J. W. Wynne, Z. Xiong, M. L. Baker, W. Zhao, M. Tachedjian, Y. Zhu, P. Zhou, X. Jiang, J. Ng, L. Yang, L. Wu, J. Xiao, Y. Feng, Y. Chen, X. Sun, Y. Zhang, G. A. Marsh, G. Crameri, C. C. Broder, K. G. Frey, L. F. Wang, and J. Wang. 2013. Comparative analysis of bat genomes provides insight into the evolution of flight and immunity. Science 339(6118):456-460.















