2
Envisioning a Rapid Learning System for Biomarker Tests for Molecularly Targeted Therapies
The charge to the Committee on Policy Issues in the Clinical Development and Use of Biomarkers for Molecularly Targeted Therapies, as noted in Chapter 1 of this report, was to examine the regulatory, reimbursement, and clinical practice policy issues that currently influence the adoption of biomarker tests for molecularly targeted therapies into routine clinical practice. Biomarker tests do not exist in a vacuum; rather, they are part of a complex health care system. The active, broad-based participation of, and collaboration among, multiple stakeholders—including patients, health care providers, payers, health care organizations, regulatory agencies, test developers and therapy manufacturers, professional organizations, and researchers—is required to transform the significant potential of biomarker tests for molecularly targeted therapies into the reality of improved patient care through precision medicine. In an effort to address the interrelated regulatory, reimbursement, and clinical practice challenges, the committee calls for the development of a rapid learning system approach that supports the most effective and appropriate use of biomarker tests and their associated molecularly targeted therapies, with continuous evidence development and ongoing assessment of their value.
This chapter provides an overview of the concept of a learning health care system, and describes a number of efforts to establish such systems. It lays the foundation for the committee’s vision of an integrated, systematic approach to accelerating the appropriate use of biomarker tests for molecularly targeted therapies to improve patient outcomes and enhance the cost-effective use of relatively expensive targeted therapies. This chap-
ter serves as a preview for the three chapters that follow. They discuss the three interrelated components (supportive policy environment, supporting data infrastructure, and processes to improve patient care) of the rapid learning system envisioned by the committee.
LEARNING HEALTH CARE SYSTEM
The notion of a learning organization was developed by organizational strategist Peter Senge, who advanced the concept in his book The Fifth Discipline (Senge, 1990). The idea was subsequently applied in the context of health care. A learning health care system is viewed as an approach to generating evidence about the quality, safety, and value of health care, using electronic health records (EHRs), large complex health care datasets known as “big data,”1 and learning networks to support and accelerate the practice of evidence-based medicine. Such a system is envisioned as a way to bridge knowledge gaps, promote the adoption of best practices, facilitate learning from the outcomes of patient care, and expedite translation of lessons learned into improvements in patient care (Etheredge, 2007).
The Institute of Medicine (IOM) led some of the recent foundational work on the learning health care system concept, starting with the initial workshop, The Learning Healthcare System (IOM, 2007), followed by numerous other workshops—convened first by the Roundtable on Evidence-Based Medicine, and later by the Roundtable on Value & Science-Driven Health Care;2 altogether the IOM produced a series of 11 workshop summaries on various facets of the learning health care system.3 In addition, the National Academies of Sciences, Engineering, and Medicine’s National Cancer Policy Forum explored a learning health care system for cancer (IOM, 2010). The concept of a learning health care system became the focus of increasing research efforts to refine and apply the concept (Abernethy et al., 2010; Etheredge, 2007, 2014; Ginsburg and Kuderer, 2012; Ginsburg et al., 2011; Schilsky et al., 2014; Sledge et al., 2013; Slutsky, 2007; Tunis et al., 2007; Wallace et al., 2014; Yu, 2015). In parallel, the National Research Council’s (NRC’s) report Toward Precision
___________________
1 “Big data has been described as the rapidly increasing size of available data, the speed with which those data are produced, and the ways in which data are represented. It also can refer not only to the data, but to the possibilities of discovering new knowledge by leveraging massive data collections in novel ways” (Krumholz, 2014, p. 1163).
2 For a complete list of the work of the Roundtable on Value & Science-Driven Health Care see https://www.nationalacademies.org/hmd/Activities/Quality/VSRT.aspx. The Roundtable’s name was changed to the Leadership Consortium for Value & Science-Driven Health Care. See http://nam.edu/programs/value-science-driven-health-care.
3 See http://www.nap.edu/catalog/13301 (accessed August 4, 2015).
Medicine: Building a Knowledge Network for Biomedical Research and a New Taxonomy of Disease called for a new national research and database system to revolutionize research, clinical care, and public health (NRC, 2011).
The IOM’s work culminated in a 2013 consensus report Best Care at Lower Cost: The Path to Continuously Learning Health Care in America, which concluded that “achieving a learning health care system—one in which science and informatics, patient-clinician partnerships, incentives, and culture are aligned to promote and enable continuous and real-time improvement in both the effectiveness and efficiency of care—is both necessary and possible for the nation” (IOM, 2013a, p. 17). The potential beneficial impact of implementing such a system was highlighted in the Health Affairs July 2014 issue, which focused on the role of “big data,” and an examination of a new rapid-learning agenda (Etheredge, 2014). The IOM continues work in this area, most recently with the publication of the workshop summary Genomics-Enabled Learning Health Care Systems (IOM, 2015). Additionally, the Leadership Consortium for Value & Science-Driven Health Care, now under the auspices of the National Academy of Medicine, continues to examine this multifaceted issue.4
The concept of a learning health care system has been articulated in various ways, including a rapid learning health care system (Abernethy et al., 2010; Etheredge, 2007, 2014; Ginsburg et al., 2011), a continuously learning health care system (IOM, 2013a), and a knowledge-generating health care system (IOM, 2015). Etheredge’s working definition of a rapid learning health care system is elegant in its simplicity: “a health system that learns as quickly as possible about the best treatments for each patient—and delivers it” (Etheredge, 2014, p. 1156). In a learning health care system, closed feedback loops between clinical practice and research enable one to inform the other as both work to improve the efficiency and effectiveness of the health care system (Ginsburg, 2014; IOM, 2015). Regardless of the terminology, learning health systems share the common goal of drawing on clinical data to “learn from every patient, and feed the knowledge of ‘what works best’ back to clinicians, public health professionals, patients and other stakeholders to create cycles of continuous improvement” (Friedman et al., 2015, p. 44).
The creation of such a system of continuous improvement would enable the health care system to “routinely study its own behavior” (Friedman et al., 2015, p. 44) and gain important insight on ways to address issues related to health care quality, cost, and safety, and to improve patient care. The collection, analysis, and shared use of clinical data form the
___________________
4 The Leadership Consortium for Value & Science-Driven Health Care manages five Innovation Collaboratives focusing on a range of issues. See http://nam.edu/programs/value-science-driven-health-care.
cornerstone of a learning health system. Indeed, through the analysis of data—from clinical trials, translational science, patient treatments and outcomes, and other data sources—the learning process occurs as data are shared, transformed into knowledge, and subsequently applied to patient care. As Yu succinctly points out: “Knowledge achieves clinical utility only when it becomes actionable to improve patient health” (Yu, 2015, p. 206). Data have the ability not only to improve clinical decision making for better patient outcomes, but also to transform clinical research, as large databases “enable observational studies on a scale and at a speed randomized controlled trials cannot approach” (Weil, 2014, p. 1110).
The successful development of a rapid learning system for biomarker tests for molecularly targeted therapies depends on the ability of health care organizations to collect and handle large quantities of data; this presents a particular challenge in the age of precision medicine and its complex biomarker tests and molecularly targeted therapies. Biomarker data and outcomes data are distinct, and both are necessary as are patient-level data (Abernethy, 2015). Thus, an effective rapid learning system requires appropriate data collection. In addition, all stakeholders, including health care providers, researchers, and payers, must be able to access, analyze, and use the data effectively, requiring a supportive health information technology infrastructure (IOM, 2013c).
Other data challenges include a lack of uniformity of data; data are often collected in free-text format rather than structured format, rendering analysis difficult (Kean et al., 2012). Moreover, sharing of data across health care organizations and settings may be difficult due to lack of standardized data definitions. Finally, data captured in a rapid learning system may be more biased or inaccurate than data collected in clinical trials (IOM, 2013c). An IOM workshop sponsored by the Patient-Centered Outcomes Research Institute (PCORI)5 focused on conducting analytical studies in a learning health care system, and examined analytic methods for improving the reliability and validity of results from such studies (IOM, 2013b).
These challenges notwithstanding, the rationale for a learning health care system is simple and clear: health care providers need better, more complete information and knowledge about what works best for individual patients, and they need it as fast as possible. Given the complexity of advanced technologies such as biomarker tests for molecularly targeted therapies, and the high cost of new therapies, accurate information on
___________________
5 Congress authorized the creation of PCORI as part of the Affordable Care Act of 2010. PCORI is a nonprofit, nongovernmental organization, whose mission is to fund comparative effectiveness research with the goal of helping patients, health care providers, payers, and policy makers make informed health care decisions.
what works and what does not work for each patient is critical to improving patient outcomes well as the cost-effectiveness of health care. The transition to a learning health care system would be advantageous for all health care stakeholders, with benefits shared by patients and health care providers, researchers, and payers and the health care system as a whole.
EXAMPLES OF LEARNING HEALTH CARE SYSTEMS
Many of the foundational elements necessary for the creation of a learning health care system are already in place, including widespread use of EHRs, registries (for many conditions, including cancers), a robust clinical trial infrastructure, and biorepositories linked to clinical data (IOM, 2013c). A number of organizations have adapted and applied concepts of the learning health system. These initiatives and other programs that generate data and foster collaboration in clinically relevant research serve as a rich base of experience from which to draw insights for the establishment of a rapid learning system for the most effective use of biomarker tests for molecularly targeted therapies. Selected initiatives are discussed below, and additional examples are found in Box 2-1.
Federal Agencies
The National Institutes of Health (NIH) is involved in a number of collaborative research initiatives that could serve as models for a learning health system. NIH launched the Electronic Medical Records and Genomics (eMERGE) Network through the National Human Genome Research Institute in 2007. The goal of this network is to “develop, disseminate, and apply approaches to research that combine DNA repositories with EHRs for large-scale, high-throughput genetic research.” The first phase of the initiative (2007-2011) used genome-wide association analysis to examine the relationship between genetic variation and at least two human traits. A key goal of the second phase (2011-2015) is to examine the best approaches to incorporate genetic variants into EHRs for use in clinical care.6
The Department of Veterans Affairs’ (VA’s) Office of Research & Development is funding the Million Veterans Program (MVP), a national, voluntary research program to study how genes affect health. The MVP program will build one of the world’s largest databases on genetic, military exposure, lifestyle, and health information.7 The VA also plans to use a learning health care system approach with veterans who are diagnosed with non-small-cell lung cancer. Test results from gene sequencing
___________________
6 See http://www.genome.gov/27540473 (accessed September 3, 2015).
7 See http://www.research.va.gov/mvp (accessed August 14, 2015).
panels will be used to direct therapy, and the information will be used for research purposes (IOM, 2015).
The learning health care system may be facilitated through collaboration among federal agencies. The Centers for Medicare & Medicaid Services (CMS), for example, could support a genetics-enabled learning health care system through its Innovation Center, which could test and advance best practices in genomics-enabled cancer care, using pay-for-performance approaches to improve quality (Etheredge, 2014; IOM, 2015).
Research Collaborations
Another example of a learning health system approach is the American Association for Cancer Research (AACR)–sponsored project called Genomics, Evidence, Neoplasia, Information, Exchange (GENIE). GENIE is a multi-phase, multi-year, data-sharing initiative to develop a regulatory-grade registry that captures and links clinical-grade cancer genomic data with clinical outcomes from thousands of patients sequenced at cancer centers in the United States, Canada, France, and the Netherlands. The registry will provide the statistical analysis to improve clinical decision making, which is a key unmet need particularly for rare cancers and rare variants in common cancers. The project also will aggregate, harmonize and share clinical-grade next-generation sequencing data from routine clinical practice. AACR will work closely with the Food and Drug Administration (FDA) to ensure data could be accepted as evidence necessary for regulatory approval.8
Another relevant initiative is the Global Alliance for Genomics and Health (GA4GH). This international coalition of 360 organizational members from 35 countries includes agencies, universities and biomedical research institutions, health care providers, information technology and life-sciences companies, research funders, and patient advocacy organizations. The goal of the alliance is to “establish a common framework of harmonized approaches to enable effective and responsible sharing of genomic and clinical data” (Lawler et al., 2015, p. 1133). The alliance’s first cancer-specific demonstration project, the BRCA Challenge, brings together leaders in research and clinical care to develop a catalog of breast cancer susceptibility gene (BRCA) variants according to their phenotypic effects. Another GA4GH effort, the Actionable Cancer Genome Initiative, focuses on harmonizing the data from different clinical sequencing efforts to implement a data-sharing approach to facilitate the use of datasets to guide patient care (Lawler, 2015).
PCORI has developed a national research network, the National Patient-Centered Clinical Research Network (PCORNet), designed to support comparative effectiveness research and clinical trials that take place in clinical care settings. PCORNet provides funding to health system–based networks whose members include hospitals, health information exchanges, and federally qualified health centers that collect electronic health information in the process of providing clinical care. PCORNet also funds patient-powered research networks focused on specific medi-
___________________
8 See http://www.aacr.org/Research/Research/Pages/aacr-project-genie.aspx (accessed January 15, 2016).
cal conditions, such as muscular dystrophy, Crohn’s disease, and arthritis, with the research controlled by patients.9
The first phase of PCORNet focused on creating infrastructure to support observational and interventional studies across multiple networks. Subsequent work will focus on conducting research using the integrated datasets (Curtis et al., 2014; Fleurence et al., 2014). One such example is ImproveCareNow, which has evolved into PEDSnet, a network of eight of the largest pediatric academic health centers in the United States. Together they provide care for more than 2 million children annually. With PCORI funding, PEDSnet aims to create a national pediatric distributed learning health system linked to three disease-specific networks, focused on complex congenital heart disease, childhood obesity, and pediatric inflammatory bowel disease (Forrest et al., 2014).
PEDSnet has formed partnerships with two national data systems and will link administrative data with clinical data from the member hospitals. Once the system is complete, it will represent the broadest pediatric big data project in the nation, and will be instrumental in ensuring effective large-scale observational research and clinical trials. Such collaboration and data-sharing efforts are critical because pediatric disorders are typically rare diseases; consequently no single health care institution has sufficiently large patient groups to produce broadly generalizable knowledge (Forrest et al., 2014).
Professional Societies
The American Society of Clinical Oncology’s (ASCO’s) Cancer Learning Intelligence Network for Quality (CancerLinQ) embraces the vision of a rapid learning system described in the IOM workshop summary report A Foundation for Evidence-Driven Practice: A Rapid Learning System for Cancer Care (IOM, 2010). ASCO also drew on the IOM’s consensus reports such as Best Care at Lower Cost (IOM, 2013a) and Delivering High-Quality Cancer Care (IOM, 2013c) for guidance in creating CancerLinQ (Schilsky et al., 2014).
Launched in 2015, CancerLinQ is a physician-led data informatics system that draws comprehensive data from EHRs and practice management systems from participating oncology practices. The system captures patient data from EHRs at the point of care, both process (what was done) as well as outcomes data. The system is designed to provide real-time clinical decision support to assist physicians in treatment planning, as well as point-of-care assessments to physicians regarding the quality of
___________________
9 See a list of patient-powered networks at http://www.pcornet.org/patient-powered-research-networks (accessed October 22, 2015).
their work. The data will be used to revise ASCO Clinical Practice Guidelines and clinical decision support tools that will provide physicians with the latest developments in a complex and rapidly changing environment (Schilsky et al., 2014; Sledge et al., 2013).
Other rapid learning system approaches focused on biomarker tests for choosing or optimizing therapy are beginning to demonstrate, on a small scale, their potential to improve patient care. For example, Vanderbilt University Medical Center continuously develops and refines clinical management algorithms to define biomarker testing protocols, in order to lower unnecessary testing or improve rates of testing when clinically indicated (Stead, 2015). Similarly, Geisinger Health System has deployed clinical decision support that queries EHRs for genetic test results indicating hypersensitivity to the HIV drug abacavir, and alerts clinicians to perform testing if results are unavailable (Williams, 2015). Both of these examples of integrating EHRs and clinical decision support are tied into feedback systems measuring outcomes, utilization, efficiency, cost, and other metrics, in order to improve health care delivery and patient care (Stead, 2015; Williams, 2015).
However, as knowledge of the implications of some biomarker tests (for example, those predicting risk of cardiac events) increases, it is becoming clear that institutions and clinical laboratories operating on their own may be unable to fully characterize those relationships (Van Driest et al., 2016). Forming broad networks with a shared commitment to continuous learning may offer a potential approach to this challenge.
The examples discussed above support the feasibility of, and demonstrate the need for, an interconnected system focused on rapid learning for biomarker tests for molecularly targeted therapies. A rapid learning system for biomarker tests for molecularly targeted therapies such as that envisioned by the committee could leverage the existing digital infrastructure and analytics capabilities and catalyze efforts to forge linkages between existing databases and learning systems. Ongoing focus would be on further strengthening cooperation and collaboration and creating solid partnerships among related systems to ensure that a critical learning opportunity is not wasted.
A RAPID LEARNING SYSTEM FOR BIOMARKER TESTS FOR MOLECULARLY TARGETED THERAPIES
The IOM’s Best Care report identifies three imperatives for achieving a continuously learning health system: managing rapidly increasing complexity; achieving greater value in health care; and capturing opportunities from technology, industry, and policy (IOM, 2013a, p. 8). These three imperatives also provide the rationale for the development of the
rapid learning system envisioned by the committee. Further justification for the development of such a system is the central role of biomarker tests in the evolving field of precision medicine; the need to capture large amounts of genomic information being generated, understand the uses for the data, and translate it into clinically useful knowledge; the necessity to accelerate learning in the field and disseminate knowledge across different clinical practice settings, patient populations, and geographic regions; and the urgency of reaching consensus on common evidentiary standards of clinical utility for biomarker tests for molecularly targeted therapies. Finally, the overarching rationale for such an approach lies in the potential to significantly improve patient care, management, and treatment outcomes.
The heightened awareness of the importance of the genetic basis of disease and the ways in which genetic factors influence patients’ varying responses to treatment is of particular relevance for the development of a rapid learning system for biomarker tests for molecularly targeted therapies. The NRC’s Toward Precision Medicine report called for an update to the International Classification of Diseases codes to incorporate the impact of genetic factors on disease (IOM, 2011). A rapid learning system would facilitate and accelerate the analysis of genetic data to guide treatment decisions (Etheredge, 2014).
In crafting the framework for a rapid learning system for biomarker tests for molecularly targeted therapies, the committee drew on two sets of guiding principles. First, the committee took into account the six aims of high-quality care conceptualized in the IOM’s landmark 2001 report Crossing the Quality Chasm—namely that care should be safe, effective, patient-centered, timely, efficient, and equitable (IOM, 2001). Second, the IOM’s Best Care report articulated key characteristics of a learning health care system (IOM, 2013a), which served to inform the committee’s vision of a rapid learning system for biomarker tests for molecularly targeted therapies (see Box 2-2).
Consistent with the characterization of a learning health care system as the framework to enable the melding of policy, process, and technology (Friedman et al., 2015), the committee’s vision of a rapid learning system for biomarker tests for molecularly targeted therapies encompasses three key components: supportive policy environment, supporting data infrastructure, and processes to improve patient care. An illustrative representation of such a rapid learning system is shown in Figure 2-1. The set of double arrows serves to highlight the process through which capturing and translating information generated by clinical research and patient care can create closed feedback loops among data, research, policy, and clinical practice to facilitate continuous learning. In this way, the process of developing new knowledge would be hard wired into the health care
delivery system, with the ability and expectations for continuous learning and updating of best evidence and clinical practices (IOM, 2013c), creating what may be characterized as “swift bidirectional learning” in which practice is informed by evidence and vice versa (Greene et al., 2012). A rapid learning system as envisioned by the committee aims to transcend
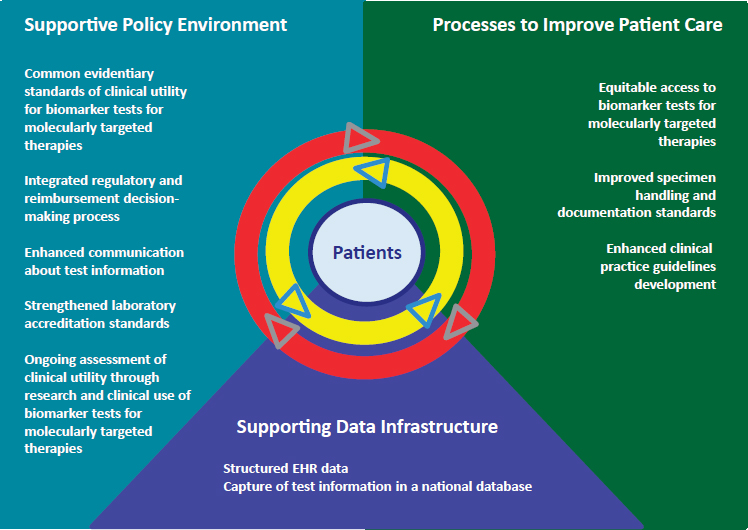
NOTE: EHR = electronic health record.
traditional barriers between research and clinical practice to support the generation of new knowledge to improve patient management and outcomes. The rapid learning system serves as the conceptual framework for the committee’s recommended approaches to addressing key clinical, regulatory and reimbursement issues facing biomarker tests for molecularly targeted therapies. The issue areas targeted by the committee are shown within each of the respective components of the committee’s envisioned rapid learning system in Figure 2-1.
Supportive Policy Environment
The committee examined a number of key regulatory and reimbursement policy challenges that influence the development and use of biomarker tests for molecularly targeted therapies. These challenges, highlighted briefly below, are discussed further in Chapter 3.
First, the lack of common evidentiary standards of clinical utility for biomarker tests is a significant challenge to the adoption of effective biomarker tests for molecularly targeted therapies into mainstream clinical practice. The lack of consensus on evidentiary standards influences health care providers’ use of tests as well as payers’ willingness to pay for the tests; both of these clearly affect patient access to tests. At the same time, some biomarker tests with limited evidence of clinical utility may be used to direct patient treatment, raising concerns about potential patient harm. The concept of the gradual evolution of evidence of clinical utility of a biomarker test for molecularly targeted therapies through clinical use over time is critical, but is not always considered. The committee’s envisioned rapid learning system represents one approach to generating evidence of clinical utility of biomarker tests for molecularly targeted therapies, and would systematically collect and analyze data on biomarker tests, molecularly targeted therapies, and patient management and outcomes. The system would support continuous learning about biomarker tests for molecularly targeted therapies by integrating data from both retrospective and prospective studies as well as incorporating “real-world” clinical data captured in EHRs, generating new evidence, and subsequently applying the knowledge gained to improve patient care.
Second, the processes for making regulatory and payment decisions for biomarker tests for molecularly targeted therapies currently are not in alignment. FDA and CMS have distinct statutory mandates. Moreover, the types of evidence required for regulatory and reimbursement decisions are inconsistent. Reimbursement decisions typically require evidence of clinical utility, which is more difficult to demonstrate than the analytic and clinical validity evidence required for regulatory approval of a biomarker test; thus, tests may be clinically available without reimbursement by CMS and other payers. To advance the adoption of appropriate, accurate, and reliable biomarker tests for molecularly targeted therapies more broadly into clinical use, the processes for regulatory and reimbursement decisions should be more closely coordinated to create a more streamlined process, discussed further in Chapter 3.
Third, the proliferation of biomarker tests for molecularly targeted therapies in the absence of transparent communication about the tests’ performance characteristics and intended uses creates uncertainty for health care professionals regarding the appropriate selection and use of tests. Clear, consistent, easy-to-understand information is needed for all biomarker tests used to direct molecularly targeted therapy, to enable health care providers to determine which test to order and to support patient engagement in the decision-making process.
Fourth, CMS regulates all clinical laboratories through the Clinical Laboratory Improvement Amendments (CLIA). CLIA oversight of labo-
ratories is not up to date and is widely viewed as insufficient for the oversight of increasingly complex biomarker tests for molecularly targeted therapies. Changes are needed to strengthen the oversight and accreditation of laboratories providing biomarker tests for molecularly targeted therapies.
Finally, it can be quite difficult to generate strong evidence of biomarker test clinical utility prior to clinical use, so in many cases, promising biomarker tests for molecularly targeted therapies may be implemented in clinical practice without sufficient data to support coverage and reimbursement decisions. As noted above, evidence evolves over time, thus continuous data collection is needed to confirm the impact of the test on longer-term patient outcomes and clinical management, or its clinical utility. Innovative reimbursement policy is needed to promote and support the ongoing assessment of clinical utility of biomarker tests. A rapid learning system as envisioned by the committee would facilitate the generation of clinical utility through collection and analysis of data on biomarker tests for molecularly targeted therapies.
The committee’s recommended policy measures to address these challenges are presented in Chapter 3. Implementation of these measures will create a supportive policy environment for the assessment and clinical implementation of biomarker tests for molecularly targeted therapies, one of the three components of the committee’s vision of a rapid learning system.
Supporting Data Infrastructure
Ideally, a learning health system uses data and technology to “learn” from clinical experience. This would be accomplished by systematically collecting various types of patient data, including laboratory test results, genomic information, treatments, and clinical outcomes; analyzing the captured data; and subsequently translating the knowledge gained from these analyses into clinical practice. To ensure continuous learning and improve the effectiveness of care, treatments, and patient outcomes need to be included in the rapid learning system database and evaluated over time, generating new hypotheses to implement and assess in clinical care (Abernethy et al., 2010; IOM, 2013b, 2015; Krumholz, 2014).
Though such cycles of continuous learning are ideal, in many cases, the opportunity to learn from clinical patient data is unrealized (Krumholz, 2014). EHRs are an important source of information to improve quality of care and generate real-world evidence, but the challenge is to ensure that patient data are structured appropriately so that information on biomarker test-directed treatment and patient outcomes can be linked. An effective learning health system should include the development and use
of EHRs and related tools that support the clinical use of biomarker tests for molecularly targeted therapies by facilitating the capture of structured data on biomarker test use, therapeutic decisions, and patient outcomes for continuous learning, research, and the development of clinical decision support tools.
A second data-related challenge is that while much genetic/genomic, treatment, and outcomes patient data are available, such data remain siloed in separate institutions and organizations and thus are not available to all for continuous learning. Data sharing is critical in order to accrue the large sample sizes needed to study increasingly subdivided, biomarker-defined patient populations. Aggregating shared data across clinical practices into a single national repository will enable research using real-world patient data to support the use of biomarker tests for molecularly targeted therapies.
Addressing these two data-related challenges is critical as “the promise of massive data assets lies not merely in their size, but the way they are used. . . . Adequately utilized, these reservoirs of data can be a practically inexhaustible source of knowledge to fuel a learning health care system” (Krumholz, 2014, p. 1169), though much remains to be learned about the uses of many types of genomic data. Thus, the committee proposes a series of measures related to data use for biomarker tests for molecularly targeted therapies, discussed further in Chapter 4. Implementing these measures will result in the creation of a supporting data infrastructure, the second key component of the committee’s vision for a rapid learning system.
Processes to Improve Patient Care
The third and final element of the committee’s vision of a rapid learning system involves processes to improve patient care related to the effective use of biomarker tests for molecularly targeted therapies. In considering approaches to implement the process component of the rapid learning system for biomarker tests for molecularly targeted therapies, the committee identified challenges in three key areas.
First, in the context of precision medicine, patients of particular economic, ethnic, cultural, and geographic backgrounds may face challenges in accessing care. These challenges may include the lack of awareness about advances in precision medicine on the part of patients and/or health care providers, as well as the ability of patients to access biomarker testing and to receive treatment with targeted therapies if appropriate. Equitable access to testing for all patients requires that health care professionals possess the expertise to properly order tests, interpret test results, and determine optimal therapy selection, in spite of the challenges posed
by the rapid pace of biomedical advances. A rapid learning system could facilitate research into potential barriers to equity and access.
Second, the reliability of biomarker test results depends on the adequacy and quality of the specimen collected. If the amount of specimen from a patient is inadequate for the tests that need to be conducted, repeat biopsy procedures may be required, exposing the patient to unnecessary risk. Uniform standards are needed regarding the handling and subsequent documentation in the EHR and/or laboratory information system of specimens for biomarker tests for molecularly targeted therapies.
Finally, consistent with the committee’s vision of a rapid learning system, clinical practice guidelines (CPGs) serve an important role in the practice of medicine and a critical educational purpose—in terms of clinical decision making as well as input for biomarker test and associated drug labeling. The IOM has recommended that for CPGs to be considered trustworthy, they should be based on a systematic review of the evidence; be transparently developed by a knowledgeable and multidisciplinary panel in conjunction with patients and reflect patient preferences; provide ratings of evidence and strength of recommendations; and be updated regularly (IOM, 2011). Although some CPGs focus on biomarker tests for molecularly targeted therapies, they may not be updated frequently, may not be user-friendly, or may conflict with other guidelines on similar topics. Increasingly, a broader base of interdisciplinary expertise is needed to generate trustworthy and consistent guidelines related to biomarker tests for molecularly targeted therapies.
A detailed examination of these challenges related to processes of patient care is presented in Chapter 5 of this report. The committee’s proposed approach to these challenges represents the third component of the committee’s vision of a rapid learning system for biomarker tests for molecularly targeted therapies.
Implementation Challenges
The committee recognizes the complexity and challenging nature of implementing such a rapid learning health system, which will require first and foremost collaboration among multiple public and private agencies, health care providers, patients, insurers, researchers, members of the health information community, health policy makers, and test and drug developers and manufacturers (IOM, 2013b). For such a system to operate effectively, all stakeholders need to value and support continuous learning.
The development of a rapid learning system will require investments of time, funding, and expertise. The federal government, particularly HHS, would be expected to play a key role in supporting a rapid learning
system for biomarker tests for molecularly targeted therapies. NIH and the National Cancer Institute (NCI) with their existing efforts detailed above, as well as FDA would be closely involved in the development of a rapid learning system for biomarker tests for molecularly targeted therapies. Moreover, CMS and other payers would be expected to provide incentives for health care researchers, providers, and manufacturers to participate in a rapid learning system.
While HHS would be expected to take a central role, many other stakeholders such as researchers, health care delivery organizations and institutions, health care providers, payers, manufacturers, and health information technology vendors would all need to contribute to—and be active participants in—the rapid learning system through openly sharing data and information for continuous learning and knowledge generation. Just as the establishment of the learning health care system requires broad-based participation, the system, if properly designed and implemented, will yield benefits that will extend across diverse groups of stakeholders: health care researchers, providers, manufacturers, and most critically, to patients who could ultimately benefit from improved patient care and outcomes.
As noted earlier, a rapid learning system represents one approach to generating evidence of clinical utility for biomarker tests for molecularly targeted therapies. Improved data collection and tracking will enable health care providers to focus on the clinical use of the most effective biomarker tests for molecularly targeted therapies. Currently health care resources are spent on some ineffective tests that carry the potential harm to patients of under- or overtreatment (FDA, 2015). The investments required for the system’s development and maintenance of a rapid learning system for biomarker tests for molecularly targeted therapies should be viewed within the context of the potential for broader health system and societal benefits to accrue from its implementation.
Other implementation challenges include the difficulties of EHRs to transmit and receive structured data successfully. The lack of such interoperability represents a significant, but not insurmountable obstacle to broad-based data-sharing initiatives as do privacy concerns as discussed further in Chapter 4 of this report. Addressing such challenges is critical for the effective operation of a rapid learning system for biomarker tests for molecularly targeted therapies as envisioned by the committee.
Many foundational elements and models exist to guide the development of such a rapid learning system, which is required to harness the new knowledge from gene-based discoveries in biology and translate it into clinical practice at a rate commensurate with its potential to direct therapy. The imperative of improved patient care underlies the necessity and urgency of building on these models to develop a rapid learning sys-
tem for biomarker tests for molecularly targeted therapies. Such a system offers profound opportunities to improve patient care and outcomes.
The three components of the committee’s vision of a rapid learning system—policy environment, data infrastructure, and processes of care—are explored in greater detail in the three subsequent chapters of this report. The rapid learning system serves as the framework for the committee’s 10 recommendations, presented in the chapters that follow.
REFERENCES
Abernethy, A. 2015. Written input to the Committee on Policy Issues in the Clinical Development and Use of Biomarkers for Molecularly Targeted Therapies. May 26, 2015.
Abernethy, A. P., L. M. Etheredge, P. A. Ganz, P. Wallace, R. R. German, C. Neti, P. B. Bach, and S. B. Murphy. 2010. Rapid-learning system for cancer care. Journal of Clinical Oncology 28(27):4268-4274.
Curtis, L. H., J. Brown, and R. Platt. 2014. Four health data networks illustrate the potential for a shared national multipurpose big-data network. Health Affairs 33(7):1178-1186.
EDM (Electronic Data Methods) Forum. 2015. EDM forum review. http://www.edm-forum.org/review (accessed November 10, 2015).
Etheredge, L. M. 2007. A rapid-learning health system. Health Affairs 26(2):w107-w118.
Etheredge, L. M. 2014. Rapid learning: A breakthrough agenda. Health Affairs 33(7):1155-1162.
FDA (Food and Drug Administration). 2015. The public health evidence for FDA oversight of laboratory developed tests: 20 case studies. Silver Spring, MD: Office of Public Health Strategy and Analysis.
Fleurence, R. L., A. C. Beal, S. E. Sheridan, L. B. Johnson, and J. V. Selby. 2014. Patient-powered research networks aim to improve patient care and health research. Health Affairs 33(7):1212-1219.
Forrest, C. B., P. Margolis, M. Seid, and R. B. Colletti. 2014. PEDSnet: How a prototype pediatric learning health system is being expanded into a national network. Health Affairs 33(7):1171-1177.
Friedman, C., J. Rubin, J. Brown, M. Buntin, M. Corn, L. Etheredge, C. Gunter, M. Musen, R. Platt, W. Stead, K. Sullivan, and D. Van Houweling. 2015. Toward a science of learning systems: A research agenda for the high-functioning learning health system. Journal of the American Medical Informatics Association 22(1):43-50.
Ginsburg, G. 2014. Medical genomics: Gather and use genetic data in health care. Nature 508(7497):451-453.
Ginsburg, G. S., and N. M. Kuderer. 2012. Comparative effectiveness research, genomics-enabled personalized medicine, and rapid learning health care: A common bond. Journal of Clinical Oncology 30(34):4233-4242.
Ginsburg, G. S., J. Staples, and A. P. Abernethy. 2011. Academic medical centers: Ripe for rapid-learning personalized health care. Science Translational Medicine 3(101):101cm127.
Greene, S. M., R. J. Reid, and E. B. Larson. 2012. Implementing the learning health system: From concept to action. Annals of Internal Medicine 157(3):207-210.
Health Affairs. 2015. Health policy briefs: The FDA’s Sentinel Initiative. http://www.healthaffairs.org/healthpolicybriefs/brief.php?brief_id=139 (accessed June 15, 2015).
Heger, M. 2015. Geisinger begins returning clinically actionable exome sequencing results to patients. https://www.genomeweb.com/sequencing-technology/geisinger-begins-returning-clinically-actionable-exome-sequencing-results (accessed December 5, 2015).
IOM (Institute of Medicine). 2001. Crossing the quality chasm: A new health system for the 21st century. Washington, DC: National Academy Press.
IOM. 2007. The learning healthcare system: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2010. A foundation for evidence-driven practice: A rapid learning system for cancer care: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2011. Clinical practice guidelines we can trust. Washington, DC: The National Academies Press.
IOM. 2013a. Best care at lower cost: The path to continuous learning health care in America. Washington, DC: The National Academies Press.
IOM. 2013b. Delivering affordable cancer care in the 21st century: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2013c. Delivering high-quality cancer care: Charting a new course for a system in crisis. Washington, DC: The National Academies Press.
IOM. 2015. Genomics-enabled learning health care systems: Gathering and using genomic information to improve patient care and research: Workshop summary. Washington, DC: The National Academies Press.
Kean, M. A., A. P. Abernethy, A. M. Clark, W. S. Dalton, B. H. Pollock, L. N. Shulman, and S. B. Murphy. 2012. Achieving data liquidity in the cancer community: Proposal for a coalition of stakeholders. http://nam.edu/wp-content/uploads/2015/06/NCPF-AchievingData-Liquidity.pdf (accessed August 22, 2015).
Krumholz, H. M. 2014. Big data and new knowledge in medicine: The thinking, training, and tools needed for a learning health system. Health Affairs 33(7):1163-1170.
Lawler, M., L. L. Siu, H. L. Rehm, S. J. Chanock, G. Alterovitz, J. Burn, F. Calvo, D. Lacombe, B. T. Teh, K. N. North, C. L. Sawyers, and Clinical Working Group of the Global Alliance for Genomics and Health. 2015. All the world’s a stage: Facilitating discovery science and improved cancer care through the Global Alliance for Genomics and Health. Cancer Discovery 5(11):1133-1136.
NRC (National Research Council). 2011. Toward precision medicine: Building a knowledge network for biomedical research and a new taxonomy of disease. Washington, DC: The National Academies Press.
ONC (Office of the National Coordinator for Health Information Technology). 2014. Connecting health and care for the nation: A 10-year vision to achieve an interoperable health IT infrastructure. https://www.healthit.gov/sites/default/files/ONC10yearInteroperabilityConceptPaper.pdf (accessed November 20, 2015).
Psek, W. A., R. A. Stametz, L. D. Bailey-Davis, D. Davis, J. Darer, W. A. Faucett, D. L. Henninger, D. C. Sellers, and G. Gerrity. 2015. Operationalizing the learning health care system in an integrated delivery system. EGEMS (Wash DC) 3(1):1122.
Schilsky, R. L., D. L. Michels, A. H. Kearbey, P. P. Yu, and C. A. Hudis. 2014. Building a rapid learning health care system for oncology: The regulatory framework of CancerLinQ. Journal of Clinical Oncology 32(22):2373-2379.
Senge, P. M. 1990. The fifth discipline: The art and practice of the learning organization. New York: Doubleday.
Sledge, G. W., C. A. Hudis, S. M. Swain, P. M. Yu, J. T. Mann, R. S. Hauser, and A. S. Lichter. 2013. ASCO’s approach to a learning health care system in oncology. Journal of Oncology Practice 9(3):145-148.
Slutsky, J. R. 2007. Moving closer to a rapid-learning health care system. Health Affairs 26(2):w122-w124.
Stead, W. 2015. The future of diagnostics for clinical care—a challenge of integration. Presentation to the Committee on Policy Issues in the Clinical Development and Use of Biomarkers for Molecularly Targeted Therapies. April 1, 2015, Washington, DC.
Tunis, S. R., T. V. Carino, R. D. Williams, and P. B. Bach. 2007. Federal initiatives to support rapid learning about new technologies. Health Affairs 26(2):w140-w149.
Van Driest, S. L., Q. S. Wells, S. Stallings, W. S. Bush, A. Gordon, D. A. Nickerson, J. H. Kim, D. R. Crosslin, G. P. Jarvik, D. S. Carrell, J. D. Ralston, E. B. Larson, S. J. Bielinski, J. E. Olson, Z. Ye, I. J. Kullo, N. S. Abul-Husn, S. A. Scott, E. Bottinger, B. Almoguera, J. Connolly, R. Chiavacci, H. Hakonarson, L. J. Rasmussen-Torvik, V. Pan, S. D. Persell, M. Smith, R. L. Chisholm, T. E. Kitchner, M. M. He, M. H. Brilliant, J. R. Wallace, K. F. Doheny, M. B. Shoemaker, R. Li, T. A. Manolio, T. E. Callis, D. Macaya, M. S. Williams, D. Carey, J. D. Kapplinger, M. J. Ackerman, M. D. Ritchie, J. C. Denny, and D. M. Roden. 2016. Association of arrhythmia-related genetic variants with phenotypes documented in electronic medical records. Journal of the American Medical Association 315(1):47-57.
Wallace, P. J., N. D. Shah, T. Dennen, P. A. Bleicher, and W. H. Crown. 2014. Optum labs: Building a novel node in the learning health care system. Health Affairs 33(7):1187-1194.
Weil, A. R. 2014. Big data in health: A new era for research and patient care. Health Affairs 33(7):1110.
Williams, M. S. 2015. Clinical practice issues. Presentation to the IOM Committee on Policy Issues in the Clinical Development and Use of Biomarkers for Molecularly Targeted Therapies. April 1, 2015, Washington, DC.
Yu, P. P. 2015. Knowledge bases, clinical decision support systems, and rapid learning in oncology. Journal of Oncology Practice 11(2):e206-e211.






















