2
Overview and Case Studies
The first session of the workshop provided an overview of the importance of reproducibility across scientific communities. Constantine Gatsonis (Brown University, co-chair of the workshop planning committee and chair of the Committee on Applied and Theoretical Statistics) and Giovanni Parmigiani (Dana-Farber Cancer Institute, co-chair of the workshop planning committee) began by introducing the workshop. Lawrence Tabak (National Institutes of Health), Irene Qualters (National Science Foundation), Justin Esarey (Rice University and The Political Methodologist), Gianluca Setti (University of Ferrara, Italy, and Institute for Electrical and Electronics Engineers), and Joelle Lomax (Science Exchange) provided perspectives from stakeholders. Victoria Stodden (University of Illinois, Urbana-Champaign) gave an overview of statistical challenges of reproducibility, and Yoav Benjamini (Tel Aviv University) and Justin Wolfers (University of Michigan) discussed reproducibility case studies.
In addition to the references cited throughout this chapter, the workshop planning committee identified the following background references: Alogna et al. (2014); Doyen et al. (2012); Errington et al. (2014); Esarey et al. (2014); Gelman and Loken (2014); Gerber and Green (2000); Harris et al. (2013); Hayes et al. (2006); Hothorn and Leisch (2011); Imai (2005); Johnson et al. (2014); Klein et al. (2014); Molina et al. (2005); Pashler et al. (2012); Rasko and Power (2015); Simons et al. (2014); Stodden et al. (2013a); and Waldron et al. (2014).
OPENING REMARKS FROM THE WORKSHOP CO-CHAIRS
Constantine Gatsonis and Giovanni Parmigiani opened the workshop with a brief overview of the importance of examining reproducibility from a statistical perspective. Gatsonis explained that the main goal of the workshop is to address statistical challenges in assessing and fostering the reproducibility of scientific results by examining three issues: the extent of reproducibility (as measured through quantitative and qualitative approaches), the methodologic causes of reproducibility failures, and the potential methodologic remedies. The methodologic perspective emphasized throughout this workshop is unique and has not been a focus of other current reproducibility discussions, according to Gatsonis. Three overarching questions were examined throughout the workshop:
- What are appropriate metrics and study designs that can be used to quantify reproducibility of scientific results?
- How can the choice of statistical methods for study design and analysis affect the reproducibility of a scientific result?
- Are there analytical and infrastructural approaches that can enhance reproducibility within disciplines and overall?
Gatsonis noted that many researchers believe developing a new conceptual framework for reproducibility is necessary instead of simply cataloging examples in which reproducibility is weak or nonexistent. He hoped that the workshop would start the conversation about such a conceptual framework, as well as how statistical thinking broadly impacts it.
PERSPECTIVES FROM STAKEHOLDERS
Lawrence Tabak, National Institutes of Health
Lawrence Tabak discussed the issue of reproducibility in the biomedical community, specifically from the perspective of the National Institutes of Health (NIH). He stated that reproducibility is a growing challenge, as has been noted by the research community and in multiple publications. The issue crosses research areas but is especially relevant in preclinical research that uses animal models as a prelude to human research, according to Tabak. He noted that science is often viewed as self-correcting and is therefore assumed to be immune from reproducibility problems. In principle, while this remains true in the long term, reproducibility checks in the short and medium term are constrained by interrelated factors.
Insufficient reporting of methodologic approaches is also an issue for a variety of reasons, spanning from limited space within journal articles to fraudulent
claims by researchers, although the latter is the minority. Sena et al. (2007) looked at the prevalence of selected quality characteristics in literature, finding that none of the 600 papers examined provided information on sample-size calculation, very few discussed how randomization was done, and very few blinded the assessment of outcome to the researchers. Tabak emphasized that this does not mean these studies did not look at these issues, but the reader is unsure based on the published record alone.
Another challenge mentioned is the phenomenon of “p-hacking,” where data are manipulated (e.g., outliers discarded) until a desired p-value is achieved. The process of p-hacking—which many investigators may not recognize as distorting—can often lead to unsubstantiated correlations being represented as statistically significant. The lack of consideration of sex as a biological variable is also an issue, Tabak explained.
Tabak noted that there are many challenges to ensuring rigor and transparency in reporting science, including incentives to publish positive results and to aim for high-impact factors, poor training, novelty (no negative data), innovation, and grant support. The biomedical research ecosystem should have research integrity at its foundation and balance robust research training (including biostatistics, basic scientific coursework, and experimental design fundamentals) with an environment that rewards networking, mentoring, career development, and collaboration feasibility.
Tabak and NIH Director Francis Collins published a commentary describing NIH’s plans to enhance reproducibility, emphasizing that all stakeholders need to be engaged (Collins and Tabak, 2014). NIH has taken a number of steps to raise community awareness, including hosting a June 2014 workshop with journal editors to identify common opportunity areas and a July 2014 workshop with the pharmaceutical trade organization PhRMA to identify common interest with industry; obtaining input from the community on the reagent-related barriers to reproducible research; and participating in meetings with professional societies and institutions. More than 130 journals endorsed the principles discussed at the June 2014 workshop, which were broadly shared in November 2014 through editorials and other notifications.1 NIH is also engaged in several pilots to address biomedical research and funding issues:
- Evaluation of scientific premises in grant applications: new funding opportunities with additional review criteria regarding scientific premises,
- Checklist and reporting guidelines: reviewer checklists regarding reporting standards and scientific rigor,
___________________
1 The full list of journals that endorsed these principles is available at National Institutes of Health, 2014, “Endorsements—Principles and Guidelines for Reporting Preclinical Research,” http://www.nih.gov/research-training/rigor-reproducibility/principles-guidelines-reporting-preclinical-research.
- Changes to biosketch: biosketch pilot with focus on accomplishments instead of just publications,
- Approaches to reduce “perverse incentives” to publish: exploration of award options with a longer period of support for investigators,
- Training: development of materials discussing the elements of good experimental design, and
- Other efforts: use of prize challenges to encourage reproducibility of results and development of a place within PubMed Commons to share and discuss concerns.
Irene Qualters, National Science Foundation
Irene Qualters explained that the National Science Foundation (NSF) has a broad view of reproducibility. She stated that the issues of reproducibility directly impact the credibility and trust afforded to research by both the research community and the public. Research in science and technology has relied on different tools to earn credibility, such as quantification of measurements from experiments, a sustained record of success, and a willingness to retract claims that are demonstrated to be erroneous, limited, or surpassed by new information and data. Thus, progress in science and technology requires that the community acknowledge that any result is to some degree at risk, no matter how carefully it is supported. New evidence, new methods, and new tools always introduce a degree of uncertainty, and science progresses in part by rejecting findings that are disproven over time.
Software is becoming an increasingly critical component within and across all science disciplines. However, software introduces vulnerabilities that are not always appreciated and often challenging to control. Software validation is often employed, especially in areas beyond the research enterprise, such as the complex software environments of nuclear energy and some clinical trials. However, these validation approaches may not be applicable to a foundational research enterprise that often relies on dynamic community software, much of which is contributed by graduate students and postdoctoral fellows. But these communities are building research on the software contributions of others, and their findings need to be credible.
Qualters emphasized the importance of understanding how data are generated and what methodologic approaches were used, as well as what tool employed is crucial for reproducibility. There are powerful tools available to measure reliability and the confidence associated with statistical results, but they are based on assumptions about the underlying data and theory about relationships and causality. As researchers continue to strive to build software to advance science and engineering, an analogous understanding of software tools is needed in order to ensure integrity and identify and measure biases.
Justin Esarey, Rice University and The Political Methodologist
Justin Esarey discussed current efforts in political science to improve reproducibility and transparency. He explained that political science is at the forefront in improving research transparency and access to replication resources in the social sciences. For example, many journals require complete replication materials upon acceptance and have done so for years. Some journals even undertake independent verification of results before publication, although the number of journals that do this is small because of resource constraints. The Data Access and Research Transparency initiative has increased the number of journals committed to providing complete, publicly available replication materials for all published work, specifically by
- Requiring authors to ensure that cited data are available at the time of publication through a trusted digital repository (journals may specify which trusted digital repository shall be used);
- Requiring authors to delineate clearly the analytic procedures upon which their published claims rely and, where possible, to provide access to all relevant analytic materials;
- Maintaining a consistent data citation policy that increases the credit that data creators and suppliers receive for their work; and
- Ensuring that journal style guides, codes of ethics, publication manuals, and other forms of guidance are updated and expanded to include strong requirements for data access and research transparency.
The Political Methodologist, the newsletter of the Political Methodology section of the American Political Science Association, recently released an issue2 focused on reproducibility and transparency problems facing the political science community. One of the tensions identified is that standards for qualitative and quantitative methods can be hard to reconcile (e.g., How can confidentiality for interviewees be ensured? How does one provide replication data for ethnography or for process tracing?). Even defining what replication means and what qualifies as a replication is challenging. Does replication require using the exact same model with the exact same data and the exact same software to generate the same result? Or, does replication check for robustness with slightly different specifications in the same data? Or, does it require verification with independent data, which are often difficult or impossible to get for observational studies? There are also questions about how replication should be integrated into graduate teaching and training, with the
___________________
2 To read this issue, see The Political Methodologist, Volume 22, Number 1, Fall 2014, https://thepoliticalmethodologist.files.wordpress.com/2015/02/tpm_v22_n1.pdf.
warning that just requiring the materials necessary for replication materials does not prevent errors. Many other questions remain, in Esarey’s view:
- To what extent is proactive error-checking a necessary part of a plan to increase transparency and replicability in the social sciences?
- How should replication projects be rewarded? Should they be published? Should negative replications get more interest?
- What constitutes replication in the case of an analysis of a fixed observational data set for where there exists only one sample (e.g., in time series cross sections (TSCSs) of country data)?
- Where replication is not an option (e.g., in many qualitative methods or observational studies of fixed TSCS data), what would constitute a check on the quality of an empirical model?
Gianluca Setti, University of Ferrara, Italy, and
Institute of Electrical and Electronics Engineers
Gianluca Setti explained that the Institute of Electrical and Electronics Engineers (IEEE) is the world’s largest professional association dedicated to advancing technological innovation and publishes about 169 journals and magazines. While serving on the IEEE board of directors through 2014, he also chaired a committee on the Future of Information and Convening, which is chartered in part to evaluate the opportunity to promote reproducible research. He emphasized that his presentation was based on his experience and represented his personal opinion.
Setti observed that reproducibility is not necessarily based on just a published paper; it also relies on the availability of data, algorithms, codes, and details of the experimental methods (Buckheit and Donoho, 1995). Because the IEEE deals with many different disciplines, it therefore has to take into consideration many facets of reproducibility, including expectations for documenting proofs, algorithms, and experiments, which may rely on custom-designed equipment (e.g., digital/analog integrated circuit implementation, microelectromechanical systems, nanotechnology, optical devices, etc.). In spite of these challenges, Setti believes that IEEE would have advantages in pursuing improved reproducibility capabilities for collaborations and in making information more visible and directly usable. Improvements would allow researchers to advance technology more easily and practitioners to develop new products faster, and they would reduce the amount of “noise” in the research literature.
Another possible way in which improving reproducibility could benefit IEEE may depend, according to Setti, on making the review process more reliable, making it more difficult to plagiarize a paper, and making it easier to discover false results and avoid retractions. The IEEE Signal Processing Society (SPS) is probably
the most advanced of the IEEE components (Barni and Perez-Gonzalez, 2005; Vandewalle et al., 2009; Piwowar et al., 2007), with all SPS publications explicitly encouraging reproducible research by both instructing researchers to submit all relevant information and providing a digital library that can house supplemental information such as data, algorithms, and code. However, these steps have not yet made a significant impact on reproducibility, according to Setti. He suggested the following steps to improve the situation:
- Create searchable and addressable repositories for data. This would entail at least the following steps:
- A unique identifier is needed for each reference data set (perhaps analogous to the Digital Object Identifier system used for other digital information) to facilitate the process of crediting data to authors.
- Privacy for sensible data should be guaranteed, which would reduce the liability risk.
- The use of standardized sets of data for specific problems should be encouraged.
- Create a cloud-based repository for sharing of codes, algorithms, and circuits, in which each of these elements is stored with its “environment.”
- Algorithms would run in the cloud using the “original” version of the compiler/program (e.g., C, python, Matlab, Mathematica).
- A public-private partnership may be required (e.g., software companies, funding agencies, professional organizations, publishers).
- Algorithms should be run, but not revealed, to encourage companies to invest in new reproducibility tools.
- Authors could upload algorithms to be linked to the corresponding papers. Metrics need to be developed to track algorithm usage, and algorithms and corresponding papers should be cross-linked.
- Reward positive efforts of authors who contribute to the reproducibility of research.
- A well-prepared reproducibility contribution requires time and effort, which currently may be a disincentive due to “publish or perish” pressure.
- Publishers could highlight papers that display good practices with regard to replication or which attempt to reproduce earlier work.
- Awards could be given for reproducible papers only.
- The review of papers with steps that enable replication could be expedited.
Setti noted that there is evidence that papers with reproducibility measures (such as shared data) are more frequently downloaded and cited than papers without these measures. With the rise of citation databases and bibliometrics, there is a strong emphasis on achieving high visibility for publications and this may stimu-
late authors to expend additional resources to ensure the results in their papers are fully reproducible. He suggested that funding agencies might be able to help stimulate reproducibility efforts by requiring reproducibility measures for publicly funded research, which is something that may be possible in the future. Enforcing such a requirement globally, however, may be difficult. It would be beneficial for the reproducibility movement if funding agencies participated in setting up the required infrastructure.
Joelle Lomax, Science Exchange
Joelle Lomax explained that the many systems studied in science, especially in biology, are extraordinarily complex and inherently varied. A key to reproducibility is to understand what variance can be controlled and what variance will always exist in a system. Reproducibility is widely understood to be a problem in both academia (Mobley et al., 2013) and industry (Scott et al., 2008; Perrin, 2014; Steward et al., 2012; Prinz et al., 2011; Begley and Ellis, 2012). However, it is difficult to identify which factors are inhibiting reproducibility and how to go about changing them.
Many hypotheses exist for why experiments fail to replicate, including insufficient reporting of methodology, pressure to withhold negative findings, poor experimental design, biased data manipulation or interpretation, unknown factors that produce variability, and flawed statistical analysis. Lomax said that empirical evidence is needed to determine the basis for this lack of reproducibility.
She explained that Science Exchange and its partners are trying to better understand these challenges as well as how the level of reproducibility (or lack thereof) can be diagnosed. She asserted that the keys to understanding reproducibility are independent replication and transparency. Independent replication is important because biases are known to exist, especially when the person performing the experiment has emotional ties to or economic dependence on the outcome of the result. This bias can lead to flawed data analysis and p-hacking (where data are collected and analyzed in such a way as to achieve a particular p-value). Transparency, specifically openness about exactly how things are being done (e.g., documenting reagents, making all raw and transformed data openly available to other researchers), is also crucial.
Lomax offered a brief summary of Science Exchange, which is an online marketplace and centralized network for more than 1,000 expert providers (both academic core facilities and contract research institutions) offering more than 2,000 experimental services to help perform scientific replications. Science Exchange has utilized this network to undertake independent replication of preclinical research. She explained that Science Exchange has partnered with the Center for Open Science to preregister all experimental designs, protocols, and planned statistical analyses and to share all raw and analyzed data through the Center for Open
Science’s Open Science Framework (an integrated project management and data repository).
Preregistration, according to Lomax, allows experts to review key factors such as study design and planned statistical analysis approaches before the data are collected. This is essentially moving the peer-review process to the beginning of the study process to deter the researcher from making certain process adjustments that would inhibit reproducibility.
One of Science Exchange’s current projects (partnered with the Center for Open Science and sponsored by the Laura and John Arnold Foundation) is the Reproducibility Project: Cancer Biology, which aims to gather a large data set of replications of preclinical research. The project is replicating the key findings of 50 recent high-impact cancer biology papers though a detailed process, including obtaining input from the original author(s). The findings and methodologies (including protocols and calculations) of these replication studies are prepublished and peer reviewed through eLife. Expert independent laboratories that are part of the Science Exchange network are utilized to carry out the replication study, and all protocols, raw data, and results are deposited to the Open Science Framework. Lomax says that this study clarifies what is needed to perform replications of preclinical research and illustrates how difficult it can be to replicate published research.
Lomax concluded by listing some of Science Exchange’s other current projects, including partnering with the Prostate Cancer Foundation and PeerJ to look at reproducibility of prostate cancer research, participating in the Reproducibility Initiative, and partnering with reagent companies to validate antibodies.
Panel Discussion
Following their individual presentations, Lawrence Tabak, Irene Qualters, Justin Esarey, Gianluca Setti, and Joelle Lomax participated in a panel discussion with follow-up questions from the audience. The session began with a participant who asked if reproducibility issues were worse today than they were 50 years ago. Qualters noted that the complexity of modeling and simulations has increased considerably over the past 20 years, and this has led to an increase in reproducibility issues. Understanding the uncertainty associated with complex software is a challenge. Tabak noted that science today is much more interdisciplinary than it has been in the past, and this interdisciplinarity makes it more difficult for researchers to truly understand what others are doing. He also noted that the number of publications has increased rapidly, which may also be contributing to the lack of reproducibility.
Another participant noted that while the goal of independent reproducibility is admirable, it assumes that the method being used in the original study is cor-
rect; he wondered what types of approaches exist and what directions should be explored to account for variation. Lomax said that this is being observed in situations in which research plans are preregistered because reviewers of those plans can criticize the original experimental design. She explained that Science Exchange is currently exploring only direct replications to control for as many differences as possible, but additional controls are added if they were not obviously included initially. However, she emphasized that research communities want reproducibility to go even further to encompass the initial variation in results that the participant noted. Esarey agreed that this raises important statistical issues. Many models can be used to analyze a given data set, and it is hoped that a study’s results do not vary significantly if one makes minor adjustments to the analysis. Investigators need to understand any such fragility, and statisticians need to develop tools that are robust to minor adjustments, according to Esarey. However, he recognized that using the right methods is inherent in science, and eventually a preferred method is established within a community. Tabak stated that there is a level of practicality that needs to be introduced, particularly in the biomedical community. Replication efforts need to be strategic because resources are limited. NIH is emphasizing the importance of rigorous preclinical research that underlies key decisions before taking that research to human trials.
A participant wondered how free software such as R is influencing current analysis. Esarey said that these types of free open-source software make it less expensive to conduct analyses, and more studies are being carried out because of the abundance of data and analysis tools. However, this free software gives communities more tools to analyze the results from these studies, which is a possible downside because, as mentioned above, some research results can be extracted with certain methods and not others. Setti commented that it is great for research communities that there are free software packages to validate a particular data set, but that often more complex tools are needed. He said that there is an opportunity for open-source developers to partner with software companies to assist the communities.
Another participant asked how funding grants could be used to promote transparency and reproducibility. Tabak said that NIH is piloting new ways of doing grant review. One of these approaches examines the premise of the application, looking at all the work leading up to the application to ensure the research is sound. Currently, NIH only does this indirectly if the preliminary data upon which the application is based come from the principal investigator and his or her team. NIH has also been developing and supporting training modules to stimulate a conversation within the community and to help with graduate and postgraduate training. He said that NIH is considering new funding opportunities for replication studies, as well as options to assess whether preclinical findings should be replicated. Qualters said that embedding reproducibility awareness into a research culture is a big change and it has to take place over time. The many stakeholders
(including individual researchers, research institutions, funding agencies, and the public) need to come together to discuss the best solutions to their community’s issues, and this consensus cannot be built overnight. She noted that there are some good resources that discuss incentive issues generally (e.g., Wellcome Trust, 2014).
OVERVIEW OF THE STATISTICAL CHALLENGES OF REPRODUCIBILITY
Victoria Stodden, University of Illinois, Urbana-Champaign
To focus on the framework for the workshop, Victoria Stodden discussed the different ways of viewing and understanding reproducibility. She suggested that it is useful to use one of three modifiers with the word reproducibility: empirical, computational, or statistical. The problems that arise and the remedies for each of these areas are very different. She said that parsing out these differences is essential because people mean different things when they use the term reproducibility.
Empirical reproducibility refers to the traditional notion of scientific reproducibility: the ability to step through the specified physical steps, protocols, and designs as described in a publication. Hines et al. (2014) give an example of empirical reproducibility by describing the difficulty two laboratories faced when collaboratively trying to ensure that both laboratories were producing the same cell signatures in their research from the identical processes. Empirical reproducibility can entail its own special constraints. Stodden shared the example of a 2014 workshop held by the National Academies of Sciences, Engineering, and Medicine’s Institute for Laboratory Animal Research that discussed reproducibility issues in research with animals and animal models (NASEM, 2015). That workshop focused mostly on empirical reproducibility, and one of the questions that arose was how to define reproducibility when true replication may mean killing additional animals.
Computational reproducibility, Stodden explained, has only become an issue over the past 20 years. She defines it as any issue arising from having a computer involved somewhere in the work process, from researchers who do bench work and analyze their data with a spreadsheet to researchers doing work on large computing systems with an enormous amount of code and software. Traditionally, going back hundreds of years, Stodden explained, there were two branches of the scientific method: (1) deductive, including mathematics and formal logic, and (2) inductive or empirical, including statistical analysis of controlled experiments. There is now some discussion that expanded computation and the deluge of data are introducing a third (and perhaps a fourth) branch of the scientific method (Donoho et al., 2009): computational, including large-scale simulations and data-driven computational science.
Stodden noted that the scientific community has had hundreds of years to think about the first and second branches of the scientific method but only a couple
of decades to figure out how to use and assess this new technology. Standards for the use of computational modes of inquiry must mature if the technology is to reach its full potential.
This abundance of data and the technology to exploit it have revolutionized a number of scientific fields, according to Stodden. The first change comes from big data and data-driven discovery. High-dimensional data with many more variables than observations are also prevalent, and this poses new challenges of analysis (in contrast to our long history of using analysis to infer meaning from limited data). In Stodden’s view, the availability of so much data, and of data-driven discovery, change what it means to carry out inference on the data while still having the ability to reproduce results. The second change that comes from computation is that powerful machines can carry out much more analysis. Elaborate simulations of entire physical systems can be performed, and these calculations can be rerun with a range of parameters so as to explore scientific questions and obtain answers. This was not possible 50 years ago. The third big change noted by Stodden is that deep contributions to science may in some cases only be coded in software, and most of the methodology innovations within the code (e.g., R scripts or more complicated or customized types of software) largely remain inaccessible in the scholarly record.
With respect to the use of computation in conducting research and what it means to be at the standard necessary to establish an acceptable new branch of the scientific method, Stodden believes that the community needs deal with reproducibility issues. She mentioned the following excerpt from Buckheit and Donoho (1995), who paraphrase Jon Claerbout: An article about computational science in a scientific publication is not the scholarship itself, it is merely advertising of the scholarship. The actual scholarship is the complete . . . set of instructions [and data] which generated the figures.
Stodden noted that just reproducing the computational work from a particular study is not the same as replicating the experiments independently, including data collection and software implementation. She stressed that both of these steps are required in order to say that the work has been reproduced. An effort to recode the software and carry out the experiment independently will rarely result in the exact same output; what is most important is being able to tell what was different between the two that led to their distinct outputs. To be able to identify these differences, the researcher who is trying to reproduce the results will need to be able to “step through” the original work and regenerate those results to see what has been done. This is a contribution to science. Both reproducibility and replicability are needed.
A number of infrastructure responses have been created within the scientific community to address the challenges that reproducibility presents, according to Stodden:
- Dissemination platforms. These are tools that enable access to codes and data, facilitate statistical verification, and link code data with publications. Examples include ResearchCompendia.org, MLOSS.org, Open Science Framework, IPOL, thedatahub.org, Madagascar, nanoHUB.org, and RunMyCode.org.
- Workflow tracking and research environments. These are tools that make it easier for researchers to capture what is important to be communicated, both computationally and physically, from their work. Examples include Vistrails, Galaxy, Pegasus, Kepler, GenePattern, CDE, Sumatra, Jupyter, and Taverna.
- Embedded publishing. New modes of publishing go beyond being analogs of papers, with the goal of making it easier to explore and visualize information. Embedded publishing results in more active documents that allow a reader to click on a figure and see it regenerate, see the software, or go somewhere that has the version data set that generated those results. Examples include Verifiable Computational Research, College Authoring Environment, SOLE, SHARE, knitR, and Sweave.
Stodden observed that reproducibility is often a by-product of open-source software, but challenges still exist in documenting this software. Stodden noted that work to strengthen capabilities and practices of computational reproducibility is being done by members of the scientific research community, but it is piecemeal and largely unrewarded as part of their regular jobs.
Statistical reproducibility, which Stodden reminded the audience is the focus of this workshop, covers a range of ways in which statistics and statistical methods influence the degree to which science is reproducible. It includes issues such as the following:
- False discoveries, p-hacking (Simonsohn, 2012), the file drawer problem, overuse and misuse of p-values, and lack of multiple testing adjustments;
- Low power, poor experimental design, and nonrandom sampling;
- Data preparation, treatment of outliers, recombination of data sets, and insufficient reporting and tracking practices;
- Inappropriate tests or models and model misspecification;
- Model robustness to parameter changes and data perturbations; and
- Investigator bias toward previous findings, and conflicts of interest.
Stodden noted that not all of these issues are inherently bad (e.g., having small samples), but they need to be understood as part of the context of an experiment.
She commented that building the cyberinfrastructure to support reproducibility involves bringing together various stakeholders, including researchers,
funding agencies, publishers, scientific societies, and institutions to discuss these issues. In 2009, a roundtable discussion was held at Yale University to discuss pertinent issues for the community, as well as potential remedies and ambitions for the future. The outcome of this workshop was a declaration of reproducible research (Yale Roundtable Participants, 2010). The Institute for Computational and Experimental Research in Mathematics at Brown University held a 2012 workshop3 to once again bring together stakeholders and continue the discussions. In 2014, the Extreme Science and Engineering Discovery Environment (XSEDE) (an NSF-funded effort that bridges access to high-performance clusters and provides an interface that allows a much broader community to use these resources) held a workshop4 about what reproducibility means in the high-performance computing context, what the next steps might be, and how we might improve reproducibility in that context.
She summarized three broad reproducibility issues to include the following:
- Failure of traditional reporting standards to accommodate changes in the research process;
- Insufficient benchmarking, testing, and validating; and
- A lack of coordination among research incentives, universities, funding agencies, journals, scientific societies, legal experts and policy makers, internal and ethical pressures, libraries, and the public. She suggested that there are many questions yet to be resolved: What am I doing to make sure that I have a job? What are universities doing? What are promotional and tenure standards? Are people asking about code data, reproducibility, and the robustness of findings? How does funding impact reproducibility? How do legal and policy environments affect the simulations, such as whether or not researchers are generating intellectual property when they develop codes or collect data?
Stodden added that research misconduct obviously impacts reproducibility, although she hopes this is limited in scope within the research community. These situations are complex, but it is important to examine the external pressures facing researchers who are working in a system that rewards outputs without strong regard to their reproducibility.
___________________
3 Institute for Computational and Experimental Research in Mathematics, “Reproducibility in Computational Mathematics: December 10-14, 2012,” https://icerm.brown.edu/tw12-5-rcem/, accessed January 12, 2016.
4 Extreme Science and Engineering Discovery Environment, “An XSEDE Workshop,” https://www.xsede.org/web/reproducibility, accessed January 12, 2016.
She concluded by summarizing that the present workshop was designed to focus on statistical reproducibility:
- Are there metrics that can quantify the reproducibility of scientific results?
- How do statistical methods affect the reproducibility of a finding?
- How do routine statistical practices, such as hypothesis test significance thresholds, affect the reproducibility of results?
- Are there analytical and infrastructural approaches that can enhance reproducibility?
- Do we need new frameworks for assessing statistical evidence?
- Do we need the broad adoption of practices for making study protocols, data, code, and workflows openly available?
- How can this be achieved?
She encouraged workshop participants to keep the last question in mind as the discussion of reproducibility continues.
CASE STUDIES
Animal Phenotyping
Yoav Benjamini, Tel Aviv University
Yoav Benjamini began by explaining that while reproducibility and replicability have only recently come to the forefront of many scientific disciplines, they have been prevalent issues in mouse phenotyping research for several decades (Mann, 1994; Lehrer, 2010). He noted that NIH, the National Institute of Neurological Disorders and Strokes, Nature, Science, and the pharmaceutical industry have recognized that preclinical studies are prone to be nonreplicable. However, he believes there is little agreement about the cause and solution.
Animal phenotyping measures an animal’s quantitative and qualitative properties to compare gene strains and mutations and to test drugs and treatments. Animal phenotyping, according to Benjamini, is widespread across many fields of science. One example of animal phenotyping is quantifying exploratory behavior in mice. During drug development, the level of anxiety in mice is documented, often by monitoring how a mouse behaves when it enters a circular arena through a small entrance. A mouse typically explores the arena by moving a small distance into the arena around the perimeter, stopping, and then retreating to the opening. As the mouse becomes more familiar with the arena, it goes a small distance farther around the perimeter than on the first attempt, stops, and again retreats to the entrance. This behavior pattern continues until the mouse eventually makes it all the
way around the perimeter and back to the entrance. This behavior can be tracked automatically, analyzed statistically, and quantified in a number of ways (e.g., the total number of steps the mouse takes, the maximum speed, the percent of time spent in the center of the arena, etc.). The percent of time spent in the center of the arena is a popular measure of anxiety; however, there is a lack of standardization regarding the time, the duration, the arena’s size and shape, and the mouse breed. This lack of standardization may cause serious replicability problems (Nature Methods, 2012; Funio et al., 2012).
The lack of standardization is a serious problem that scientific communities should care about, Benjamini emphasized. He discussed a Crabbe et al. (1999) study where an experiment was conducted in three laboratories with strict standardization variables. This study found there were genotype and laboratory effects that may yield results that are idiosyncratic to a particular laboratory. Additional studies showed that differences between laboratories could contribute to failures to replicate results of genetic experiments (Wahlsten, 2001).
Later experiments (Richter et al., 2009, 2011; Würbel et al., 2013) compared two strains of mice (C57 and CBA) in six commercial and academic laboratories across Europe and showed random laboratory and interaction effects. He explained that the particular laboratory effect in a new laboratory is unknown but its random effect can be eliminated in this case by design by running the two strains in the same laboratory. However, the particular genotype x laboratory (GxL) mixed-model interaction cannot be eliminated by design.
Initially, the existence of significant GxL interaction was considered a lack of replicability, but Benjamini argued that the existence of GxL interaction cannot be avoided in part because the genotyping by laboratory effect is unknown. He explained that some aspects can be automated and made more uniform; however, some researchers have raised concerns about whether increased uniformity in laboratories will heighten the effects of small unimportant differences between laboratories, which could harm replicability rather than improve it.
Benjamini stated that if the GxL variability ![]() were known and could be estimated, then it could easily be corrected for by subtracting the mean of one strain of mice from the mean of another strain and dividing this by the regular t test
were known and could be estimated, then it could easily be corrected for by subtracting the mean of one strain of mice from the mean of another strain and dividing this by the regular t test ![]() plus the variability
plus the variability ![]() , making the calculation
, making the calculation
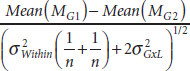
Benjamini argued that the interaction size is the correct measure to compare genetic difference when examining replicability across laboratories. He emphasized that design quality, large sample size, and transparency alone will not solve the
issue of replicability. Increasing the sample size will diminish ![]() , but it will not help with the variability. He stated that many nonreplicable results could be screened out if laboratories knew the GxL variability. However, he said the proportion of errors committed by falsely rejecting null hypotheses (known as the false discovery rate) is still high but could be approached using a false discovery rate controlling method (Benjamini and Hochberg, 1995). Therefore Benjamini recommends reporting both a p-value and a GxL-adjusted p-value for replicability, or perhaps a ratio of the two.
, but it will not help with the variability. He stated that many nonreplicable results could be screened out if laboratories knew the GxL variability. However, he said the proportion of errors committed by falsely rejecting null hypotheses (known as the false discovery rate) is still high but could be approached using a false discovery rate controlling method (Benjamini and Hochberg, 1995). Therefore Benjamini recommends reporting both a p-value and a GxL-adjusted p-value for replicability, or perhaps a ratio of the two.
The GxL variability can be estimated by using batch variability (e.g., litters of animals, days the experiment was conducted) as a surrogate for GxL variability or by purposely injecting variation into the experiment’s environmental condition when conducted in a single laboratory (Richter et al., 2009). The latter heterogenizing approach is controversial because it is the opposite of standardizing, but Benjamini believes this approach allows researchers to control the known variables. To improve estimates of GxL variability, Benjamini proposed making use of large publicly available databases of mouse phenotyping results (e.g., the International Mouse Phenotyping Consortium5).
He concluded by reiterating Victoria Stodden’s point that reproducibility of computing problems is difficult. He also noted that there is a replicability problem with two key statistical sources in animal phenotyping: using an inappropriate measure for variability (e.g., ignoring GxL interaction) and failing to adjust for selective inference. He stated that transparency, standardization, and large sample sizes do not eliminate these problems in single-laboratory experiments, but community efforts can help. He believes that replicability can move from a burden to an asset by estimating the GxL variability from multilaboratory experiments to evaluate newly suggested measurement tools and devices.
A participant asked about the infrastructure developed to address GxL interactions. Benjamini explained that the infrastructure is not well developed yet, but he and others are trying to build a database with the cooperation of the International Mouse Phenotyping Consortium, with the hopes of getting scientists to contribute results to help enrich the system.
Another participant asked if interlaboratory variability should be treated as variation in bias or whether there is something interesting about anxiety, perhaps the genetic source of anxiety in the interlaboratory variability. Moreover, the participant wondered if there is something to be learned about anxiety from knowing the ways in which laboratories are different. Benjamini said that there have been many efforts to try to identify and standardize sources of variability. However, there are often far too many sources of variability to standardize everything effectively,
___________________
5 The International Mouse Phenotyping Consortium website is http://www.mousephenotype.org, accessed January 8, 2016.
so effort is taken to standardize the variables that are believed to be important and report the other variables. This approach gives researchers an opportunity to investigate different variables in follow-up studies and demonstrates why transparency is an important issue.
A participant questioned the generalizability of the approach, specifically if the approaches to analyzing, modeling, and judging reproducibility of mice data apply to other research areas or if a different framework would need to be developed. Benjamini responded that this could be generalized by identifying the real uncertainty (Mosteller and Tukey, 1977) because the uncertainty between the animals is less relevant than the interaction between genotypes and laboratories. He stressed that insisting on the correct relative level of the uncertainty is an important issue that can be generalized to other areas and other fields.
A participant wondered if the laboratories would get the same results if they exchanged animals. Even if the mice are genetically identical, Benjamini responded, they may have diverged because of evolution in the cage, and they could be epigenetically different because they may have different methylation statuses of the DNA from the different environments. The participant noted it would be interesting to know if results were consistent when moving the mice or whether the laboratory of origin or the environment had changed it. Benjamini said that animal exchanges happen and significant variation has been shown among supposedly genetically identical mice (Kiselycznyk and Holmes, 2011), but a homogeneous set of animals is typically used for drug discovery and experimentation. He suspects this is not a large problem within the research.
Another participant commented that attempting to measure many variables about the laboratories and enter them into the model first, assuming the treatment is the same and neither bias nor fraudulent behavior is an issue, may be more efficient for recovering results and measuring the source of the variation. Benjamini responded that when assessing replicability, the concern is not what happened in the current experiment but whether another laboratory would get the same results using the described methodology. He cautioned against overspecifying properties of the laboratory because the results may end up being more particular to the laboratory. The goal is to estimate the interaction because it is the best assurance of what will happen when the next laboratory tries to replicate these results. Benjamini suggested either using a mixture of the fixed- and mixed-model analyses to capitalize on the strengths of both or simply reporting both. This means doing the best analysis possible from the two laboratories and then estimating the adjusted variability so future researchers know what to expect.
Capital Punishment
Justin Wolfers, University of Michigan
Justin Wolfers introduced his case study, which illustrates some of the challenges of doing empirical research and trying to reassess existing literature. Wolfers’s discussion was based on work he did with John Donohue, a law professor at Stanford, which examined the deterrent effect of the death penalty (Donohue and Wolfers, 2006). Wolfers noted that because the death penalty is a politically contentious issue, the challenges associated with performing objective social science research are amplified.
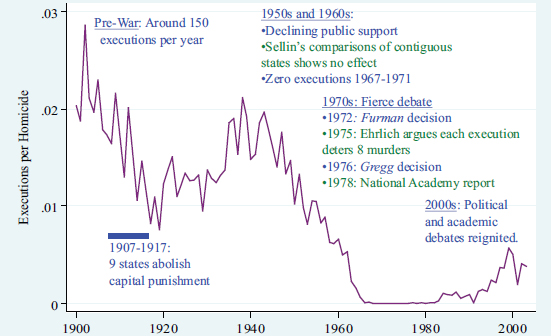
The social, legal, economic, and political histories of the death penalty in the United States are integrally wrapped up in the evolution of social science, according to Wolfers. Figure 2.1 shows the history of executions in the United States. In the early 1900s, approximately 150 people were executed each year in the United States. From 1907 to 1917, nine states abolished capital punishment, which decreased the total number of executions for the country during that time. During the 1950s and 1960s, there was further decline in public support of the death penalty. Sellin (1959) looked at what happened in a state that had the death penalty versus in an adjoining state that did not; his study suggested there was no deterrent effect of the death penalty. The laws during this time did not change, but the death penalty was rarely used; the total number of executions plummeted to zero between 1967 and 1971, despite the fact that many states still had it on the books and there were still people imprisoned for capital murder.
In 1972, the Supreme Court deemed all existing death penalty statutes in the United States unconstitutional (Furman v. Georgia), which led to a de facto moratorium on capital punishment. In contrast to Sellin’s study, Ehrlich published a paper in 1975 that analyzed the connection between executions and homicides and concluded that for each person executed, eight future homicides were prevented. The Supreme Court cited Ehrlich’s paper in its 1976 decision (Gregg v. Georgia) that allowed capital punishment to resume. In 1978, the National Research Council released a report that assessed the research cited by the Supreme Court and found that there was little evidence from social science to suggest that executions deterred homicide (NRC, 1978). In spite of the Supreme Court ruling, the death penalty fell largely into disuse with just a few executions per year.
Then in the 1980s, a few large states began to express interest in using the death penalty again, and the economic and political debates were reignited. Wolfers said that various scholars, many of whom were located in the southern United States, started to write research papers looking at the relationship between execution and homicide rates and once again claimed that higher execution rates led to lower homicide rates.
In the late 1990s and early 2000s, most of the economics literature suggested that executions deterred homicide (e.g., Dezhbakhsh and Shepherd, 2006; Dezhbakhsh et al., 2003; Shepherd, 2005; Mocan and Gittings, 2003; Zimmerman, 2004; Cloninger and Marchesini, 2001). However, Wolfers pointed out that one study, conducted by researchers who focused on prison conditions rather than the death penalty (Katz et al., 2003), controlled for what was happening with the death penalty and found that there was no deterrent factor. Indeed, they inferred that there was a slight incentive to commit murder. The deterrent effect shown in literature from this time period is outlined in Table 2.1.
Wolfers noted that the broader context is one of robust debate in the policy world. Texas decided during this time to ramp up use of executions, while California sentenced many people to death row but executed few of them. Several state governors, including those in Illinois and New Jersey, instituted moratoriums on the death penalty.
He described some of the strong claims being made, such as Emory University law professor Joanna Shepherd’s statement before Congress that there is a “strong consensus among economists that capital punishment deters crime,” “the studies are unanimous,” and “there may be people on the other side that rely on older
TABLE 2.1 The State of the Capital Punishment and Deterrence Literature, circa 2008
| Literature | Data Used | Number of Lives Saved per Execution |
|---|---|---|
| Dezhbakhsh and Shepherd (2006) | 1960-2000 state panel | 8 |
| Dezhbakhsh et al. (2003) | 1977-1996 county panel | 18 |
| Shepherd (2005) | 1977-1999 monthly state panel | 2 |
| Mocan and Gittings (2003) | 1984-1997 state panel | 4 |
| Zimmerman (2004) | 1978-1997 state panel | 14 |
| Cloninger and Marchesini (2001) | Illinois 2000-2003 moratorium | 40 |
| Texas April 1996-April 1997 moratorium | 18 | |
| Katz et al. (2003) | 1950-1990 state panel | –0.8 |
SOURCE: Courtesy of Justin Wolfers, University of Michigan, presentation to the workshop.
papers and studies that use outdated statistical techniques or older data, but all of the modern economic studies in the past decade have found a deterrent effect. So I’m not sure what other people are relying on” (Terrorist Penalties Enhancement Act of 2003). Wolfers added that this issue came up in political debates as well. In the 2000 presidential election, George W. Bush supported the use of the death penalty because it “saves other people’s lives” (Commission on Presidential Debates, 2010), while Barack Obama stated that the death penalty “does little to deter crime” (Obama, 2006).
Wolfers noted that research intending to analyze the deterrent effect of the death penalty could be approached by measuring a causal effect of an experiment with subjects. The standard error of this analysis would be large or small depending on whether the number of subjects considered was small or large, respectively. If all experiments are being reported and illustrated using a funnel chart (see Figure 2.2a), then a relationship is illustrated where the larger the standard error, the more variable the estimates across different experiments. The smaller the standard error, the less variable the findings should be, with 5 percent falling outside the 95 percent band of the funnel. If instead scholars choose to report only the results that showed statistically significant evidence of a deterrent effect of the death penalty, the literature would be limited to the analyses that fall above the 95 percent band of the funnel (see Figure 2.2b). Wolfers explained that this leaves a footprint in the data that shows a correlation between how large the estimates of the effect of the death penalty are and the standard error of the particular studies.
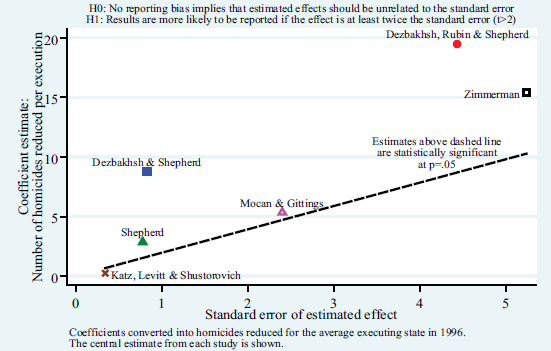
He took the key measurements from some of the studies in the death penalty literature and graphed their results, where the horizontal axis is the standard error and the vertical axis is the effect of each execution on the number of homicides.
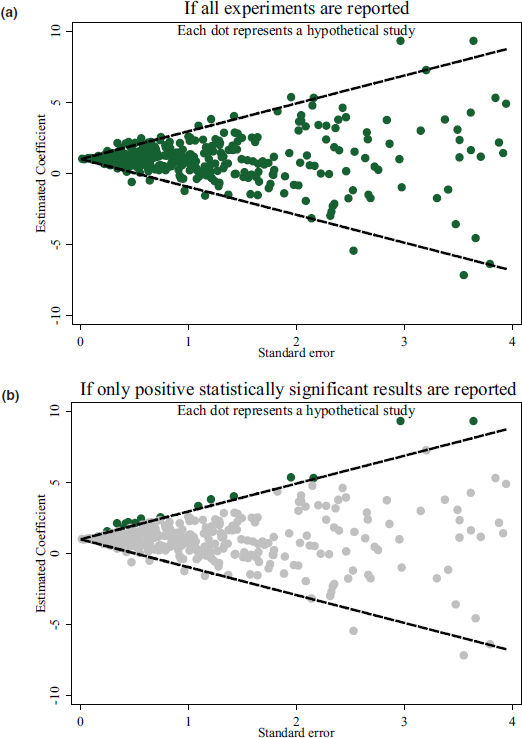
Figure 2.3 shows the central estimate from each study, which Wolfers explained showed a clear, statistically significant relationship with the six observations. He noted that imprecise results are reported when they yield extremely large effects.
In Wolfers’s view, the norm in most social science work is to subject the main estimates to a variety of sensitivity testing—for example, to see what happens if a slightly different regression specification is used, the functional form is altered, a couple of states are left out, or the sample period is changed. He explained that the literature shows a robust relationship between the size of the estimate and the size of the standard error, with the exception of the Katz et al. (2003) paper. Wolfers argued that this implies selective reporting within the literature. This could be a type of specification search or file drawer problem where the central estimates are not representative of the underlying distribution of estimates that one could find from a cleaner look at the data.
Wolfers explained that when he and Donohue replicated each of these studies, they found coding errors and what they believed were fairly substantial judgments that could not be supported. They concluded that none of the studies that showed large deterrent effects were convincing. He argued that replication research and
sensitivity testing like this work is important because it begins to fill in missing pieces and identify unconvincing evidence, essentially finding the information that had not been recorded.
Responding to Isaac Ehrlich’s study (1975) that found each execution deterred eight homicides, Wolfers commented that it is hard to find a convincing and reliable correlation using the data and statistical technique from that study. The study’s data (from 1933 to 1967) showed that both homicides and executions decreased, and Wolfers pointed out that time series analysis would thus imply that decreasing the number of executions reduces homicides. However, Ehrlich had specified the model in terms of the log number of executions and, as the number of executions falls, the log number of executions shows a positive correlation with the decrease in homicides. However, this analysis does not accurately capture the increases in homicides that occurred during some years because using the log smooths the results; if data from 1960 to the early 2000s are analyzed instead (as is done by Dezhbakhsh and Shepherd, 2006), the small decline of executions in 1960 coincides with a huge rise in homicides that lasts throughout the 1980s. Wolfers noted that this also coincides with the crack cocaine epidemic. Executions stayed low while homicides rose for the next 20 years. As executions began to rise slightly (while remaining low overall) around 1990, the data show a massive deterrent effect on the homicide rate. However, Wolfers questioned why tiny changes in executions have massive effects on homicides but massive changes in executions do not have massive effects on homicides. Because the time frame chosen for the analysis resulted in vastly different results, he concluded that simple statistical misuse (such as taking logs around zero) could dramatically alter findings.
Wolfers then discussed the natural experiment period between 1972 and 1976 when the United States no longer had the death penalty. The homicide rate rose throughout that time, implying that eliminating the death penalty was to blame. However, homicide rate data from Canada closely mirrors the pattern seen in the United States during that period (although the Canadian homicide rate is only one-third that of the United States). Similarly, Canada eliminated the death penalty in 1965, and the homicide rate there rose. However, the rate rose similarly in the United States. He also noted that when the Supreme Court eliminated the death penalty in 1973, there was obviously no change in the number of executions in the 10 states that had already abolished the death penalty. However, homicide rates also rose in those states. Wolfers stressed the importance of comparison groups when doing analyses such as this from observational data.
Dezbakhsh and Shepherd (2006) analyzed state homicide rate data 3 years before and after the death penalty was abolished in each state. They found that abolishing the death penalty caused the homicide rate to rise considerably, while its reinstatement led to a decline in the rate. However, Wolfers noted that this analysis
misses relevant comparisons during these time periods, as well as the fact that rates were similar across other states during the years being examined.
In Wolfers’s view, many of these papers on the death penalty also suffer from previously mentioned serious limitations, such as coding errors that eliminate the wrong observations (Mocan and Gittings, 2003) and judgment calls that skew the findings from the analysis (Dezhbakhsh et al., 2003).
Wolfers cautioned that unpleasant confrontations are a major disincentive to attempting to replicate others’ work. For example, when Donohue and Wolfers (2005) were published, several authors of the work that was replicated sharply criticized the new paper, though they did not provide the original data to allow a more thorough examination of their analysis. Donohue and Wolfers (2005) ultimately concluded that there are insufficient data to determine the influence of the death penalty while noting that the existing literature reflects problems including publication bias; neglect of comparison groups; coding errors; highly selected samples, functional forms, regressors, and samples; and overstatements about statistical significance.
In 2012, the National Research Council released a report on the evidence of the deterrent influence of the death penalty. That report stated, “These studies should not be used to inform judgments about the effect of the death penalty on homicide, and should not serve as a basis for policy decisions about capital punishment. . . . Fundamental flaws in the research we reviewed make it of no use in answering the question of whether the death penalty affects homicides rates” (NRC, 2012). The broader lessons of this case study, according to Wolfers, are as follows: (1) the politically charged nature of this debate may change the discussion and influence the analysis; (2) selective reporting is a pervasive problem in many attempts to look at funnel plots with respect to social science research; (3) there is a great need for sensitivity testing for replication; (4) improved norms and definitions are needed in social science research; (5) there is an important role for impartial referees; and (6) there are substantial disincentives for researchers getting involved in this research.
In response to a participant’s question, Wolfers suggested that the community work together to lay out a set of definitions about replication so it is clear what was done when someone claims to have replicated research. At present, “replication” can mean anything from simply making the code work on a different computer to thoroughly evaluating and validating the underlying analysis. He suspects one of the difficulties will be coming up with a set of definitions across disciplines.
Another participant stated that social science literature, especially when it gets into the newspapers, often has confounders and flawed results that can be attributed to poor data analysis or fitting models without thinking about the available data. This participant wondered if there are policies that could be implemented to improve the data analysis. Wolfers responded that although review is imper-
fect, a better system is not currently available. The flaws he identified could have been spotted during peer review, but mistakes happen even in reputable journals. For example, because codes are often not shared, coding errors cannot be found. Wolfers suggested that the scientific process could be improved by requiring that every published paper include the archived data, but this does not address the possibility that an archived code might fail to run or deliver the expected results. Some journals employ a research assistant to run the code to make sure that the numbers came out one time on one machine, but this is expensive. Wolfers also commented that in some top-tier newspapers, the refereeing criteria are sometimes more rigorous than those used in peer review.
Often, scientific reporting is skewed by university press offices that draft press releases in more absolute and less nuanced ways than researchers would. Andrew Gelman (2015) has suggested that researchers should be held accountable for exaggerations in press releases about their work; Wolfers agrees that this would improve the accuracy of these releases. Another participant commented that the misportrayal of science in the media is not unique to the social sciences. This participant sent a note to a top-tier journal pointing out that the results of a paper being used to guide ovarian cancer treatment were based on confounding experimental design. Stating that this letter was too technical for their readership, the journal refused to publish it; however, The New York Times published the letter two weeks later.
Wolfers noted that in economics, and in the social sciences more broadly, there is very little demand for replication work. Donohue and Wolfers (2005) overcame this by examining several major papers in the existing literature and systematically proving the results to be false. He believes that if they instead had examined only one paper it would not have been publishable. He suggested that there is actually a disincentive for journals to publish replicability research because it requires time and energy to be as careful as they need to be; good replication is incredibly valuable, but bad replication is destructive.
In response to a question by a participant regarding the benefit of occasionally calling out severe mistakes as fraud, Wolfers claimed he tends to avoid using this word because even researchers doing indefensible work can change their ways and adopt better practices in the future. In response to another participant, he clarified that having flawed analysis and making up data require a different level of response, and often the former does not rise to the level of fraud.
Another participant asked what could be done when the data cannot be shared due to data use agreement, as is often the case with social science settings (e.g., health policy). Wolfers noted that journals often confront this issue, and some have instituted a policy requiring researchers to submit all the code needed to replicate the analysis with the restricted data. Since access to the data is rarely granted, the editors have to trust that this code would actually replicate the work. This approach makes it clear how the researcher approached the analysis.
A participant posed a final question about what the scientific community could do to communicate its concerns and possible solutions to policy makers and stakeholders. Wolfers offered that in the case of the death penalty analysis, he was relieved that the Academies used its role as a highly respected independent referee while examining the literature. However, he noted that this might not be the best use of resources in all cases. He said that the social sciences are moving toward being more open through blogs that discuss the research process, which foster more public and transparent discussion of reproducibility issues. The success of this approach depends on the culture of each discipline. Institutionally, Wolfers noted that many funders are excited about developing a replicability standard, and much of the movement in the natural sciences is coming from the funders insisting that data be in the public domain.



























