1
Introduction
Austin Burt had a question: Can an insect’s genes be manipulated to stop it from spreading disease? Burt, an evolutionary geneticist, was conducting research on site-specific selfish genetic elements, “stretches of DNA” that are certain to pass down from a parent organism to nearly all of its offspring (Burt and Trivers, 2006). Like many scientists before him, Burt wondered if the molecular mechanisms that enable selfish genetic elements to spread through a population could be harnessed to eliminate undesirable genetic traits, like the ability of some mosquitoes to carry disease-causing parasites and viruses.
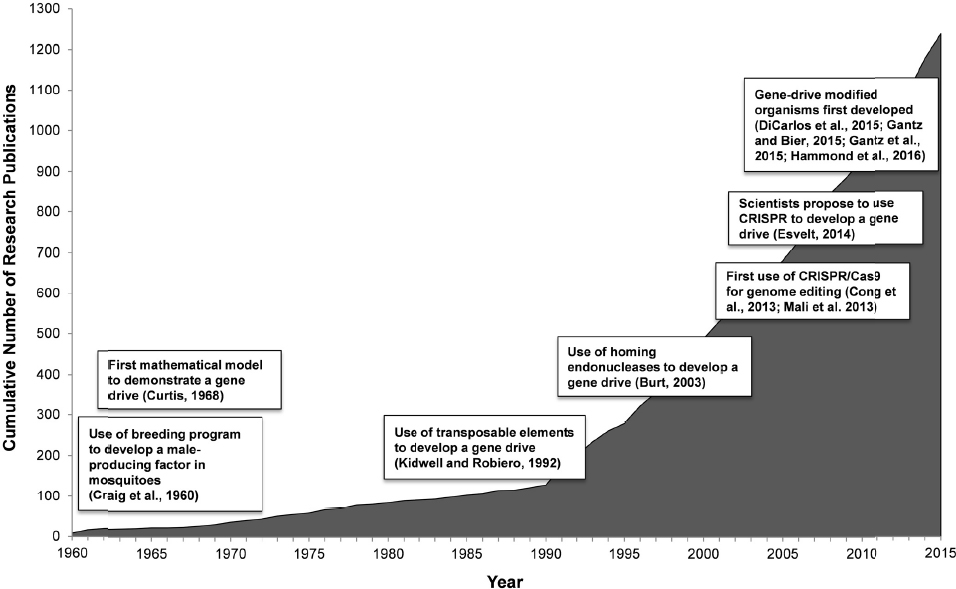
Scientists have known about selfish genetic elements since the late 1880s (Burt and Trivers, 2006). However, the idea to use selfish genetic elements as means to control natural populations did not surface until the mid-20th century. In 1960, George B. Craig, a mosquito biologist, and two of his colleagues, W.A. Hickey and R.C. Vandehey, suggested using a breeding program in which a “male-producing factor” that is naturally present in some male Aedes aegypti mosquitoes would be harnessed to control mosquito populations. When male mosquitoes with this male-producing factor breed, most of their offspring then develop as males (Craig et al., 1960). Environmental releases of male mosquitoes carrying this male-producing factor could potentially “reduce the number of females below the [population] level required for efficient disease transmission” (Craig et al., 1960). Hickey and Craig (1966a,b) later described a driving sex determining region on a chromosome in mosquitoes and its potential for population control by causing the sex ratio of the population to shift from half males and half females to an increasing proportion of males. In a related analysis of the conditions favoring the evolution of such biased sex ratios, W.D. Hamilton (1967) also realized the potential for using male bias as a mechanism to control population size. He reasoned that if “the Y chromosome could be freed from the inhibitory control of the rest of the genome,” this could be a powerful mechanism of biological control (Hamilton, 1967). Chris Curtis, a medical entomologist, then published the first mathematical model demonstrating how a naturally occurring “desirable” gene, such as a gene “to make mosquitoes non-infectible by pathogens,” could spread to fixation in a population.” In the model, the gene would always be present in enough members of the mosquito population to prevent “infectibility” from ever taking hold again (Curtis, 1968).
In the 1960s Craig, Hamilton, Curtis and the other early pioneers did not yet have the molecular tools to engineer “desirable” genes or to molecularly tie them to a biased mechanism of inheritance. More than 30 years of basic biological research in genetics and molecular biology took place before potential genetic engineering tools became available. In 1992, Margaret Kidwell, an evolutionary geneticist, and José Ribeiro, a vector biologist, proposed using transposable elements, mobile sequences of DNA, as a mechanism to drive an engineered gene into a mosquito population (Kidwell and Ribeiro, 1992). In 2003, Burt proposed using the homing endonuclease gene, a selfish gene, to drive genetic changes into a natural population (Burt, 2003). A number of geneticists were studying homing endonucleases as a potential basis for targeted gene therapy, a still-experimental approach to treat or prevent particular genetic diseases in humans. Burt extended this reasoning and wondered if homing endonucleases could also be used to drive modified genes through a mosquito population (Burt, 2003; Burt and Trivers, 2006).
Kidwell and Ribeiro (1992) and Burt (2003), in combination with advanced knowledge about genetics and more modern molecular tools, bolstered the field of inquiry into so-called gene drives. Geneticists and population biologists continued to explore how to use a variety of selfish genetic elements as the mechanistic basis for the development of a gene drive, primarily
in mosquitoes (James, 2005; Rasgon and Gould, 2005; Adelman et al., 2007; Windbichler et al., 2011). However, a precise and predictable mechanism to cause the preferential increase in an existing or engineered trait remained elusive. Then, along came CRISPR (Jinek et al., 2012; Mali et al. 2013; Cong et al., 2013).
CRISPR (Clustered regularly-interspaced short palindromic repeats) are segments of bacterial DNA that, when paired with a specific guide protein, such as Cas9 (CRISPR associated protein 9), can be used to make targeted cuts in an organism’s genome. Bacteria use the CRISPR/Cas9 union as a kind of immune system to defend themselves against foreign genetic sequences, such as those that can be inserted by viruses (Barrangou et al., 2007; Hale et al., 2009). Biologists developed a way to use CRISPR/Cas9 like a pair of scissors to make genetic changes by cutting targeted sequences so that existing DNA can be removed or new DNA sequences can be inserted. The CRISPR/Cas9 system, the newest and now most widely used gene editing technique, has rapidly led to breakthroughs in the editing the genomes of many organisms, including plants, nematodes, flies, fish, monkeys, and human cells (Basset et al., 2013; Friedland et al., 2013; Fu et al., 2013; Jiang et al., 2013; Niu et al., 2014; Gratz et al., 2015). As described by biochemists Sam Sternberg and Jennifer Doudna, “what had once been laborious and time-consuming was now facile and rapidly achievable,” because gene editing with CRISPR/Cas9 systems enabled the insertion, deletion, or replacement of specific genes in many species (Sternberg and Doudna, 2015). CRISPR/Cas9 also proved to be a perfect tool to create a gene drive. It enabled biologists to transform the idea of a gene drive into a reality.
In early 2015, 3 years after the first demonstration of CRISPR/Cas9 as a gene editing tool, a research group led by George Church created the first gene drive in yeast (DiCarlo et al., 2015). Valentino Gantz and Ethan Bier, two molecular biologists, published the first demonstration that a gene drive could be created in an insect, the fruit fly, in March 2015 (Gantz and Bier, 2015). Gantz and Bier used the term mutagenic chain reaction to describe the mechanism they developed to create a gene drive using CRISPR/Cas9. By late 2015, two independent research groups, one led by Anthony James and the other by Austin Burt and Andrea Crisanti, developed gene-drive modified mosquitoes (Gantz et al., 2015; Hammond et al., 2016). In less than 4 years, a new genetic engineering tool, CRISPR/Cas9, paired with advanced knowledge about selfish genetic elements, enabled a breakthrough in what scientists had been studying for more than 50 years (see Figure 1-1).
PURPOSE OF THE STUDY
In his seminal paper, Burt posited three reasons for conducting research on gene drives: “to motivate more rapid development of the technology; to warn of containment issues that ought to be addressed during development; and to stimulate discussions on the desirability of eradicating or genetically modifying particular species” (Burt, 2003). Development of the CRISPR/Cas9 technology has accelerated the need to address such issues and more.
Will applications of gene drives be safe? Will they be effective? Will they have unintended consequences for the environment or public health? Do we know enough to release gene-drive modified organisms into the wild? Is using a gene drive to suppress or eliminate a pest species a good idea? What can scientists do to reduce risks to humans, other organisms, and the environment? How do we decide where gene-drive modified organisms might get released? What should governments do? Who gets to decide? These and other questions prompted the National Institutes of Health (NIH) and the Foundation for the National Institutes of Health (FNIH)1,2 to ask
___________________
1A nonprofit, nongovernmental organization that is separate from the NIH.
2This study was sponsored by the National Institutes of Health and the Foundation for the National Institutes of Health, and the National Academy of Sciences Biology and Biotechnology Fund. The Defense Advanced Research Projects Agency and The Bill & Melinda Gates Foundation provided support to the NIH and the FNIH, respectively for this study.
the National Academies of Sciences, Engineering, and Medicine to convene a committee with a broad range of expertise to summarize the scientific discoveries related to gene drives and considerations for their responsible use. The committee’s task includes three primary components: (1) review the state of the science and approaches to reduce unintended harms that could potentially result from developing and using gene-drive modified organisms; (2) discuss the ethical, legal, and social considerations attendant to field release of gene drives; and (3) determine the adequacy of existing governance mechanisms and risk assessment guidelines for the environmental and public health implications of using gene drives (see Box 1-1).
To inform this task, the committee held a one-day workshop in Washington, DC, and organized 11 webinars, to gather input from experts and stakeholders. Speakers provided perspectives on science, ethics, public engagement, and governance mechanisms. Topics included the biology of the organisms that are likely initial candidates for gene drive research; evidence derived from experience with field releases of other modified organisms (e.g., use of Wolbachia) and how this evidence might inform a risk assessment process for gene drives; how gene drives do, or do not, fit into the current governance system for biotechnology, both in the United States and internationally; how specific values influence public perception of the potential deployment of organisms carrying gene drives; and how best to engage members of the public in discussions about potential benefits and harms of gene drives, particularly in low- and middle-income countries where the first deployments of gene drives to combat vector-borne diseases are likely to occur. The workshop agenda and the list of webinar topics are in Appendices A and B, respectively. The presentations from the workshop and webinars are freely available to members of the public through the project’s website.3 The committee’s deliberations led to this final consensus report, which draws on the presentations that the committee heard, the scientific and other literature, and the expertise of its members. General principles to guide responsible practices in gene drive research for the laboratory setting through to field releases are embedded as recommendations throughout the report.
WHAT ARE GENE DRIVES? AND, HOW COULD THEY BE USED?
In reviewing the history of research on what are now called selfish genetic elements, the committee noted differences in the use of terminology and definitions. Drive, gene drive, meiotic drive, driving Y chromosome, selfish gene, selfish genetic elements, and related concepts often have overlapping definitions depending on the historical period and the scientific context in which the terms are used. In this report gene drives are defined as systems of biased inheritance in which the ability of a genetic element to pass from a parent to its offspring through sexual reproduction is enhanced. Thus, the result of a gene drive is the preferential increase of a specific genotype, the genetic makeup of an organism that determines a specific phenotype (trait), from one generation to the next, and potentially throughout the population. These systems encompass the requisite molecular elements and events necessary for biased inheritance to occur. Because inheritance is biased in their favor, the genetic elements encompassed by gene drives are often called selfish genes or selfish genetic elements. Examples of selfish genetic elements include genes or their fragments, all or parts of chromosomes, or noncoding DNA (Burt and Trivers, 2006). As noted above, since the 1960’s researchers have imagined that selfish genetic elements “might serve as the basis for ‘gene drives’ capable of spreading engineered traits through wild populations” (Esvelt et al., 2014).
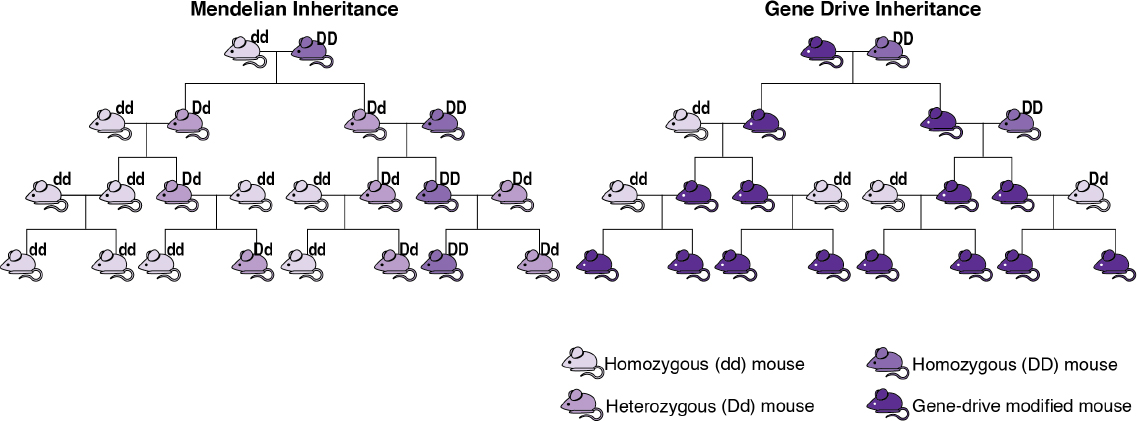
Gene drives are often described as an exception to the conventional rules of inheritance. First described in 1866 by a monk named Gregor Mendel, the conventional rules of inheritance, also known as Mendelian inheritance, dictate that offspring have on average a 50% chance of inheriting a gene from one of their parents. With Mendelian inheritance, not all offspring will inherit the gene, and so the frequency of that gene in future generations will be similar to the frequency of that gene in the parents’ generation. With gene drives, offspring have more than a
___________________
50% chance of inheriting a genetic element from a parent, and so a specific genotype will increase in the population over time. Figure 1-2 illustrates an idealized difference between Mendelian inheritance and inheritance through a gene drive. However, note that the number of generations and amount of time for a selfish genetic element to spread throughout a population will vary depending on the gene drive mechanism, the species, and a variety of environmental conditions (see Chapter 2 for additional detail).
Gene drives occur in nature through a variety of mechanisms. Researchers are studying how to harness natural mechanisms, such as transposable elements, maternal effect dominant embryonic arrest (Medea), and meiotic drive to develop gene drives in various organisms. However, the pairing of a desired trait with molecular mechanisms that will cause that trait to drive is difficult. CRISPR/Cas9 facilitates the capability to create a gene drive in laboratory populations (DiCarlo et al., 2015; Gantz and Bier, 2015; Gantz et al., 2015; Hammond et al., 2016). In the last few months of 2015 alone, two research studies demonstrated the use of CRISPR/Cas9 to create a gene drive in mosquitoes (Gantz et al., 2015; Hammond et al., 2016). These studies also demonstrated how a gene drive could be used for two key population control methods:
- Population suppression—the spread of a genetic element that causes the number of individuals in a population to decrease; and
- Population replacement—the spread of a genetic element through a population that causes a population’s genotype to change.
With the advent of CRISPR/Cas9 as an editing tool, biologists have proposed a wide range of applications for gene drives including to address public health threats, species conservation, agriculture protection, and advance basic research on evolutionary genetics (see Table 1-1). For example, gene drives could potentially be used as one component of an integrated population-control strategy for mosquitoes that transmit malaria, combined with several methods, such as applying insecticides and eliminating breeding habitats. Gene drive technologies could become an important option for addressing complex, difficult-to-solve issues, particularly those where solutions are limited or entirely lacking. For example, outbreaks of mosquito-borne diseases such as Zika virus, chikungunya, West Nile virus, and a diverse group of “neglected tropical diseases,”4 like elephantitis, can be devastating either because there are few effective treatment options or because affected individuals have limited access to such treatment, particularly in low- and middle-income countries. It is important to recognize, however, that gene drives may be challenging or not possible to develop in many organisms. The enthusiasm for proposed applications must be tempered, as discussed later in this report, by a number of scientific, ethical, legal, and social factors.
Because the field is rapidly advancing, it is reasonable to expect proofs-of-concept in organisms other than fruit flies and mosquitoes within the next few years. The fast moving nature of this field, however, is also a point of concern. For example, it may be easier and much faster to develop a population suppression drive in some organisms than to conduct research on the long term environmental effects. Regulatory systems that govern biosafety and biosecurity are outpaced by the scientific advances, which have led some scientists to propose responsible laboratory practices (Akbari et al., 2015). Gene-drive modified organisms appear to have promise if they can be responsibly developed.
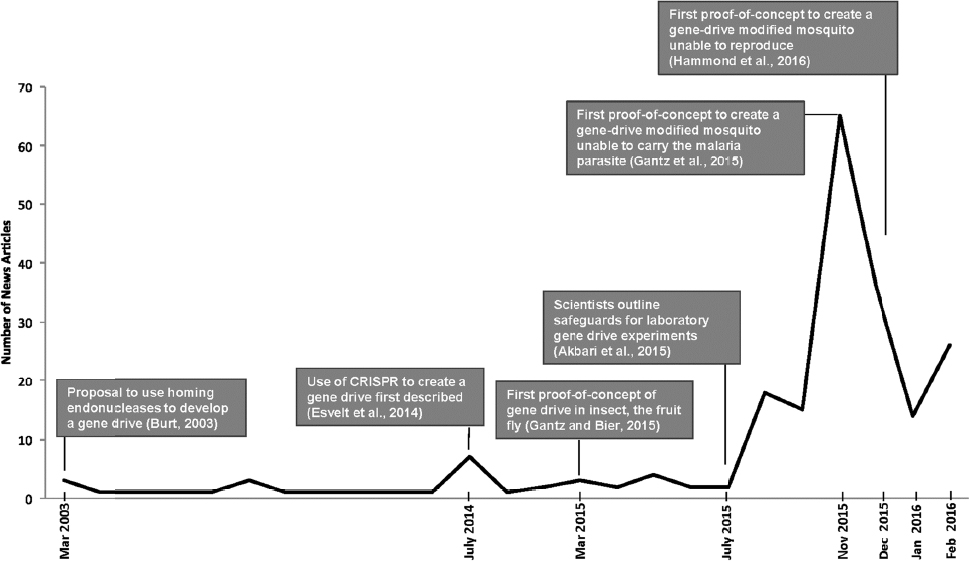
Researchers and the media have expressed concern over the potential social, environmental, and health-related impacts of the development and release of gene-drive modified organisms. Indeed, the prospect of gene drives for the control of vector-borne infectious diseases has sparked significant media interest and concern, particularly in response to the mosquito proof-of-concept publications at the end of 2015 (see Figure 1-3).
Scientific publications cite the benefits of gene drives, but also acknowledge that using gene drives “would represent an entirely new approach to ecological engineering” (Esvelt et al., 2014). Esvelt and colleagues also point out that current knowledge for containing and managing risks related to the spread of novel genes through entire populations and for evaluating ecological consequences is poor. To date, this research has mainly focused on mosquitoes and a few additional organisms for which biological control plans are in place. Similarly, Esvelt and colleagues emphasize that “given the potential for gene drives to alter entire wild populations, and therefore ecosystems, the development of this technology must include robust safeguards and methods of control” (Esvelt et al., 2014). Oye et al. (2014) argued that “studies have evaluated the possibility of releasing transgenic mosquitoes to combat the spread of malaria, dengue, and other mosquito-borne diseases, including requirements for containment, testing, controlled release, and monitoring of mosquito gene drives. This work will need to be replicated and extended for proposed gene drives seeking to alter other species. It is crucial that this rapidly developing technology continue to be evaluated before its use outside the laboratory becomes a reality.”
Laboratory research on gene drives is advancing rapidly, but the proposed applications are based on limited proof-of-concept studies. Before this technology is put forth as a safe and viable tool, further research is needed to validate laboratory studies through independent replication of results, assess the potential benefits and harms of gene-drive modified organisms for ecosystems and human health, and develop effective strategies for mitigating potential harmful outcomes.
___________________
4The World Health Organization lists 17 neglected tropic diseases, 6 of which are transmitted by insect vectors: http://www.who.int/neglected_diseases/diseases/en.
TABLE 1-1 Potential Applications for Gene Drive Research
| Public Health | |
 Aedes aegypti Image Source: US Centers for Disease Control and Prevention |
|
| Ecosystem Conservation | |
 Hemignathus munroi (‘Akiapōlā’au honeycreeper) Image Source: US Fish and Wildlife Service |
|
| Agriculture | |
 Fruit damage from spotted wing drosophila infestation Image Source: US Department of Agriculture |
|
| Basic Research | |
 DNA Double Helix Image Source: National Institutes of Health |
|
In addition, as the threat of vector-borne diseases and invasive species is not limited by political boundaries, there are many questions about potential governance and regulation of the technology, as well as ethical concerns, and a clear need to engage appropriately with communities in areas where gene drives might be tested and applied in the field.
CASE STUDIES
The committee developed a series of case studies based on the likely directions and applications of gene drive research identified in Table 1-1. The case studies are used to illustrate key considerations for gene drive research, ethics, and governance. See Chapter 3 for the detailed descriptions of the following case studies:
Case Study 1: Using Aedes aegypti and Aedes albopictus mosquitoes to manage dengue throughout the world
Case Study 2: Using Anopheles gambiae mosquitoes to combat human malaria in sub-Saharan Africa
Case Study 3: Using the Culex quinquefasciatus mosquitoes to combat avian malaria on the Hawaian islands
Case Study 4: Control populations of non-indigenous Mus musculus mice to protect native biodiversity on islands throughout the world
Case Study 5: Controlling non-indigenous Centaurea maculosa knapweeds to protect biodiversity in rangelands and forests in the United States
Case Study 6: Controlling Amaranthus palmeri (Palmer amaranth, also known as pigweed) to increase agriculture productivity in the Southern United States
Case Study 7: Developing a vertebrate model for gene drive research using Danio rerio zebrafish
KEY DEFINITIONS AND CONCEPTS
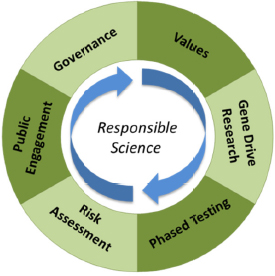
The committee was tasked to assess the science of gene drives along with the ethical, legal, and social implications of developing this basic biology as applied biotechnology. The committee chose to answer this question by focusing on elements of responsible science important to the gene
drive field as expressed in six areas of scholarship and practice: gene drive research, values, phased testing, risk assessment, engagement, and governance. In the context of this report, a responsible science approach calls for scientific habits of mind that integrate these six areas of scholarship and practice (see Figure 1-4).
Genome Editing and Gene Drives
Genome editing is a technique that allows researchers to insert, delete, or modify DNA to silence, activiate, or otherwise modify an organism’s specific genetic characteristics. There are a number of tools used to edit genomes, including the use of zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), and CRISPR. Although they share common techniques, genome editing is not necessarily designed to result in a gene drive.
A gene drive is a system of biased inheritance in which the ability of a genetic element to pass from a parent to its offspring through sexual reproduction is enhanced. Thus, the result of a gene drive is the preferential increase of a specific genotype, the genetic makeup of an organism that determines a specific phenotype (trait), from one generation to the next, and potentially throughout the population. Discussions about gene editing and gene drives, from molecular biology to population ecology, are elaborated in Chapter 2.
Values
Values are deeply held, complicated, sometimes evolving beliefs about what kinds of things—in human lives and the world at large—should be fostered, protected, or avoided, and therefore about what people should do and what they should not do. They are critical components of human identity and society. Chapter 4 describes human values that may guide our perceptions, decisions, and actions about gene drive research and its potential applications.
Phased Testing Pathway
A phased testing pathway is a step-wise approach to guide the preparation for and conduct of research in the laboratory through environmental release.
The phased testing pathway described in this report is based upon that of the World Health Organization for the testing of genetically modified mosquitoes (WHO, 2014). Chapter 5 describes the phased testing pathway and scientific approaches in each phase to identify and mitigate potential harms of gene drives.
Risk and Risk Assessment
Risk is the probability of an effect on a specific endpoint or set of endpoints due to a specific stressor or stressors. An effect can be beneficial or harmful. For example, a beneficial effect of releasing a gene-drive modified mosquito could be a reduction in the spread of avian malaria, while a harmful effect could be an increase in other types of insects that carry infectious disease.
A risk assessment is the process by which all available evidence on the probability of effects is collected, evaluated, and interpreted. Then the potential total effects are estimated from the evidence (EPA, 1984). Risk and risk assessment are discussed in greater detail in Chapter 6.
Public Engagement
Public engagement is the act of seeking and facilitating the sharing and exchange of knowledge, perspectives, and preferences between or among groups who often have differences in expertise, power, and values. Public engagement is a long-term, multidirectional, iterative process of communication. Engagement enables the exchange of information and perspectives as policy
questions are asked, refined, reconsidered, and—often only temporarily—answered. Public en-engagement is discussed in further detail in Chapter 7.
Governance
Governance is the process of exercising oversight through traditions (standards of practice) or regulations by which individuals and communities are held accountable. This includes:
- The process by which authorities are selected, monitored, and replaced;
- The capacity of governing authorities to formulate and implement sound policies; and
- The respect of governed communities for the authorities and processes that govern their economic and social interactions.
Governance in the context of scientific research includes government standard setting and regulation; education of scientists and manufacturers; systems of accreditation; public engagement; and other mechanisms for standards of behavior and controls for safety, environmental protection, and other social goods (NRC, 2015). See Chapter 8 for an in-depth discussion of governance.
CONCLUSIONS
Gene drive research is advancing rapidly, and the proposed applications will likely continue to expand as genome editing tools such as CRISPR become more refined. New scientific information and public perspectives arise almost on a monthly basis concerning the use and application of gene drive research.
The fast-moving nature of this field is both encouraging and a point of concern. Gene-drive modified organisms hold promise for addressing persistent or difficult-to-solve challenges, such as the eradication of vector-borne diseases and the conservation of threatened and endangered species. But the presumed efficiency of gene-drive modified organisms may lead to calls for their release in perceived crisis situations before there is adequate knowledge of ecological effects, and before mitigation plans for unintended harmful consequences are in place. Continuous evaluation and assessment of the social, environmental, legal, and ethical considerations of gene drives will be needed to develop this technology responsibly and adapt research and governance to the field’s complex and emerging challenges.
REFERENCES
Adelman, Z.N., N. Jasinskiene, S. Onal, J. Juhn, A. Ashikyan, M. Salampessy, T. MacCauley, and A.A. James. 2007. nanos gene control DNA mediates developmentally regulated transposition in the yellow fever mosquito Aedes aegypti. Proc. Natl. Acad. Sci. 104(24):9970-9975.
Akbari, O.S., H.J. Bellen, E. Bier, S.L. Bullock, A. Burt, G.M. Church, K.R. Cook, P. Duchek, O.R. Edwards, K.M. Esvelt, V.M. Gantz, K.G. Golic, S.J. Gratz, M.M. Harrison, K.R. Hayes, A.A. James, T.C. Kaufman, J. Knoblich, H.S. Malik, K.A. Matthews, K.M. O’Connor-Giles, A.L. Parks, N. Perrimon, E. Port, S. Russell, R. Ueda, and J. Wildonger. 2015. BIOSAFETY. Safeguarding gene drive experiments in the laboratory. Science 349(6251):927-929.
Barrangou, R., C. Fremaux, H. Deveau, M. Richards, P. Boyaval, S. Moineau, D.A. Romero, and P. Horvath. 2007. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315(5819):1709-1712.
Bassett, A.R., C. Tibbit, C.P. Ponting, and J.L. Liu. 2013. Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep. 4(1):220-228.
Burt, A. 2003. Site-specific selfish genes as tools for the control and genetic engineering of natural populations. Proc. Biol. Soc. 270(1518):921-928.
Burt, A., and R. Trivers. 2006. Genes in Conflict: The Biology of Selfish Genetic Elements. Cambridge, MA: The Belknap Press of Harvard University Press.
Cong, L., F.A. Ran, D. Cox, S. Lin, R. Barretto, N. Habib, P.D. Hsu, X. Wu, W. Jiang, L.A. Marraffini, and F. Zhang. 2013. Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819-823.
Craig, G.B., Jr., W.A. Hickey, and R.C. Vandehey. 1960. An inherited male-producing factor in Aedes aegypti. Science 132(3443):1887-1889.
Curtis, C.F. 1968. Possible use of translocations to fix desirable genes in insect pest populations. Nature 218(5139):368-369.
DiCarlo, J.E., A. Chavez, S.L. Dietz, K.M. Esvelt, and G.M. Church. 2015. Safeguarding CRISPR-Cas9 gene drives in yeast. Nat. Biotech. 33(12):1250-1257.
EPA (US Environmental Protection Agency). 1984. Risk Assessment and Management: Framework for Decision Making. EPA 600/9-85-002. US Environmental Protection Agency, December 1984.
Esvelt, K.M., A.L. Smidler, F. Catteruccia, and G.M. Church. 2014. Concerning RNA-guided gene drives for the alteration of wild populations. eLife 3:e03401.
Friedland, A.E., Y.B. Tzur, K.M. Esvelt, M.P. Colaiácovo, G.M. Church, and J.A. Calarco. 2013. Heritable genome editing in C. elegans via a CRISPR-Cas9 system. Nat. Methods 10(8):741-743.
Fu, Y., J.A. Foden, C. Khayter, M.L. Maeder, D. Reyon, J.K. Joung, and J.D. Sander. 2013. High-frequency off-target mutagenesis induced by CRISPR- Cas9 nucleases in human cells. Nat. Biotechnol. 31(9):822-826.
Gantz, V.M., and E. Bier. 2015. Genome editing. The mutagenic chain reaction: A method for converting heterozygous to homozygous mutations. Science 348(6233):442-444.
Gantz, V.M., N. Jasinskiene, O. Tatarenkova, A. Fazekas, V.M. Macias, E. Bier, and A.A. James. 2015. Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proc. Natl. Acad. Sci. 112:E6736-E6743.
Gratz, S.J., M.M. Harrison, J. Wildonger, and K.M. O’Connor-Giles. 2015. Precise genome editing of Drosophila with CRISPR RNA-guided Cas9. Methods Mol. Biol. 1311:335-348.
Hale, C.R., P. Zhao, S. Olson, M.O. Duff, B.R. Graveley, L. Wells, R.M. Terns, and M.P. Terns. 2009. RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 139(5):945-956.
Hamilton, W.D. 1967. Extraordinary sex ratios. Science 156 (3774):477-488.
Hammond, A., R. Galizi, K. Kyrou, A. Simoni, C. Siniscalchi, D. Katsanos, M. Gribble, D. Baker, E. Marois, S. Russell, A. Burt, N. Windbichler, A. Crisanti, and T. Nolan. 2016. A CRISPR-Cas9 gene drive system targeting female reproduction in the malaria mosquito vector Anopheles gambiae. Nat. Biotechnol. 34(1):78-83.
Hickey, W.A., and G.B. Craig, Jr. 1966a. Genetic distortion of sex ratio in a mosquito, Aedes aegypti. Genetics 53(6):1177-1196.
Hickey, W.A., and G.B. Craig, Jr. 1966b. Distortion of sex ratio in populations of Aedes aegypti. Can. J. Genet. Cytol. 8(2):260-278.
James, A.A. 2005. Gene drive systems in mosquitoes: Rules of the road. Trends Parasitol. 21(2):64-67.
Jiang, W., H. Zhou, H. Bi, M. Fromm, B. Yang, and D.P. Weeks. 2013. Demonstration of CRISPR/ Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res. 41(20):e188.
Jinek, M., K. Chylinski, I. Fonfara, M. Hauer, J.A. Doudna, and E. Charpentier. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816-821.
Kidwell, M.G., and J.M. Ribeiro. 1992. Can transposable elements be used to drive disease refractoriness genes into vector populations? Parasitol. Today 8(10):325-329.
Mali P., L. Yang, K.M. Esvelt, J. Aach, M. Guell, J.E. DiCarlo, J. Norville, G.M. Church. 2013. RNA-guided human genome engineering via Cas9. Science, Jan 3. 339:823-6.
Niu, Y., B. Shen, Y. Cui, Y. Chen, J. Wang, L. Wang, Y. Kang, X. Zhao, W. Si, W. Li, A.P. Xiang, J. Zhou, X. Guo, Y. Bi, C. Si, B. Hu, G. Dong, H. Wang, Z. Zhou, T. Li, T. Tan, X. Pu, F. Wang, S. Ji, Q Zhou, X. Huang, W. Ji, and J. Sha. 2014. Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156(4):836-843.
NRC (National Research Council). 2015. The Industrialization of Biology: A Roadmap to Accelerate the Advanced Manufacturing of Chemicals. Washington, DC: The National Academies Press.
Oye, K.A., K. Esvelt, E. Appleton, F. Catteruccia, G. Church, T. Kuiken, S.B. Lightfoot, J. McNamara, A. Smidler, and J.P. Collins. 2014. Biotechnology. Regulating gene drives. Science 345(6197):626-628.
Rasgon, J.L., and F. Gould. 2005. Transposable element insertion location bias and the dynamics of gene drive in mosquito populations. Insect Mol. Biol. 14(5):493-500.
Sternberg, S.H., and J.A. Doudna. 2015. Expanding the biologist’s toolkit with CRISPR/Cas9. Mol. Cell 58(4):568-574.
WHO (World Health Organization). 2014. The Guidance Framework for Testing Genetically Modified Mosquitoes. World Health Organization, Programme for Research and Training in Tropical Diseases [online]. Available at http://apps.who.int/iris/bitstream/10665/127889/1/9789241507486_eng.pdf?ua=1 [accessed April 19, 2016].
Windbichler, N., M. Menichelli, P.A. Papathanos, S.B. Thyme, H. Li, U.Y. Ulge, B.T. Hovde, D. Baker, R.J. Monnat, Jr., A. Burt, and A. Crisanti. 2011. A synthetic homing endonuclease-based gene drive system in the human malaria mosquito. Nature 473(7346):212-215.














