5
Phased Testing and Scientific Approaches to Reducing Potential Harms of Gene Drives
The acceleration of gene drive research and the increasing ease of use of the molecular technologies required to construct gene drives has generated considerable excitement about the potential use of gene-drive modified organisms to address public health, conservation, agricultural, and other challenges. However, releasing a gene-drive modified organism into the environment means that a complex molecular system will be introduced into complex ecological systems, potentially setting off a cascade of population dynamics and evolutionary processes that could have numerous reverberating effects. Thus, effective strategies to carry out laboratory and field research are needed to study each type of gene-drive modified organism, its potential benefits and harms, and approaches to reduce or mitigate the potential harms.
The preceding chapters of this report describe what is known about gene drives, key population ecology and ecosystem considerations for gene drive research, and human values that may influence whether and how gene-drive modified organisms are used. Building upon that foundation, this chapter focuses on a step-by-step pathway designed to guide research and support evidence-based decision making at each phase. In addition a range of strategies to reduce potential off-target and non-target effects are explored through the lens of this phased approach to research. Specific examples from biocontrol and existing transgenic research geared toward the suppression or replacement of populations in the wild provide additional insights and lessons learned.
THE PHASED TESTING PATHWAY
Will proposed applications of gene drives work as intended? Researchers have proved that gene drives can be developed in some laboratory populations, but to date gene-drive modified organisms have not been studied in the environment. When will gene-drive modified organisms developed in the laboratory be ready for field-based research, or release into the environment? From a research perspective, the answer to these types of questions requires careful analysis of gene drives that begins at the molecular level and continues through the population and ecosystem levels. A number of criteria must be met for gene drives to be responsibly developed. A step-by-step approach can guide research from the laboratory to the field. To help guide gene drive research, the committee adapted and expanded upon the phased testing pathway outlined by the World Health Organization (WHO) for the testing of genetically modified mosquitoes (WHO, 2014).
A phased testing pathway is a step-wise approach to guide the preparation for and conduct of research that begins in the laboratory and continues through, if applicable, environmental monitoring (see Figure 5-1). The idealized pathway for research on a gene-drive modified organism includes five steps: Research Preparation (phase 0), Laboratory-Based Research (phase 1), Field-Based Research (phase 2), Staged Environmental Release (phase 3), and Post-Release Surveillance (phase 4). Although the overall goal is for unidirectional movement from early to later phases, the pathway includes a set of feedback loops, to encourage repetition and refinement of studies based on new findings and data generated during the course of research. The phased testing pathway enables a researcher to identify milestones and decision points in regard to when the
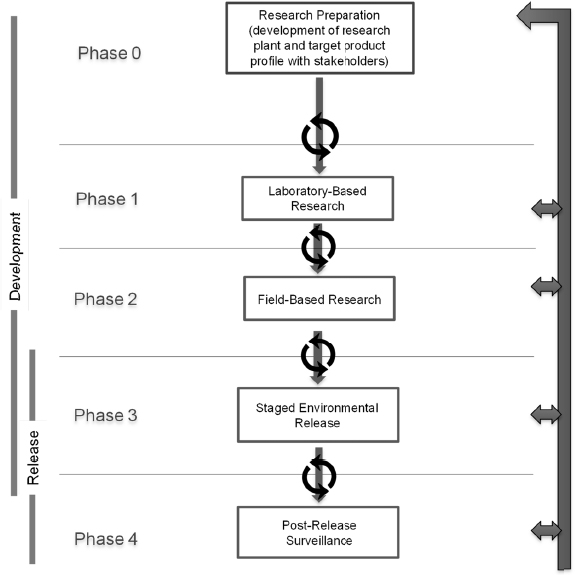
research is ready to move from one phase to the next. The decision to advance to the next phase of testing may also depend on approval from relevant publics, particularly local communities and regulatory authorities. Hence, support mechanisms for risk assessment, public engagement, and governance, are needed throughout the phased testing pathway. Considerations for public engagement and governance are discussed in Chapters 7 and 8, respectively. Some examples of activities that take place in each phase are provided in Box 5-1 and discussed in the text below.
The goal of gene drive research is to develop organisms that are viable in the environment, and that, in some cases, will persist for indeterminate periods of time. In addition, gene-drive modified organisms may potentially be able to interbreed with related, wild species. For these reasons, confinement and containment are critical considerations throughout the phased testing pathway. In reviewing the literature on different approaches to research, the committee noted that the terms confinement and containment are often used interchangeably. To provide clarity, the committee developed the following definitions derived from the 2004 National Research Council report Biological Confinement of Genetically Engineered Organisms, the World Health Organization (WHO, 2014), and the US Department of Agriculture:
- Confinement is the use of ecological conditions or biological methods to prevent unintended or uncontrolled persistence of an organism in the environment. Climatic isolation, when the surrounding environment or expected seasonal change is suboptimal for
- Containment is the use of human-made or natural physical restrictions to prevent unintended or uncontrolled release of an organism into the environment. Examples of human-made containment mechanisms include large cages, greenhouses, and aquaculture pens (NRC, 2004). Geographic isolation, such as an island setting without human inhabitants (O’Connor et al., 2012) is an example of a natural physical barrier.
an organism’s survival, is an example of ecological confinement (Adelman, 2015a; Akbari et al., 2015). An example of a biological method (sometimes called bioconfinement) is use of an organism that depends on a chemical or nutrient that is not present in the environment.
A combination of confinement and containment methods will likely be needed for each phase of gene drive research, with careful consideration for combinations that will not conflict with the purpose of the study.
Phase 0: Research Preparation
The purpose of the research preparation phase is to develop a robust plan that details the scope and goal of the study, pre-defined thresholds for success, methods of confinement and containment, and strategies to reduce the potential for unintended harms. Such a research plan can serve as the basis for funding proposals. At this stage, researchers have a working knowledge of the biology, behavior, and natural history of their target organism, as well as the environment or environments (e.g., laboratory or field contexts) in which the research will take place. Confinement, containment and biosafety mechanisms, mitigation strategies, and anticipated regulatory approvals will be developed and discussed with the relevant Institutional Biosafety Committees (IBCs), expert advisory panels, regulators and funders.
A critical component of this phase is a process for setting goals and pre-defined thresholds for success. A Target Product Profile (TPP), a strategic development tool that uses sets of criteria to pre-define ideal attributes of a candidate “product” (FDA, 2007), is one model. Although originally developed to facilitate assessment and prioritization of candidate pharmaceuticals, the TPP process has been adopted for the context of mosquito vector control product development by the WHO Vector Control and Advisory Group and by private funders such as The Bill & Melinda Gates Foundation.
A TPP can help researchers, funders, and policy makers to think through minimum standards of acceptance related to efficacy, safety, regulatory, and manufacturing endpoints for a specific application, such as the use of a gene-drive modified mouse to reduce the population of invasive wild mice on islands (Case Study 4) or the development of a gene-drive modified zebrafish as a vertebrate model for research on inheritability of off-target effects (Case Study 7). The TPP can also include specifications other than efficacy that will be important for policy decisions, such as cost comparisons of different potential courses of action, in order to weigh options and make sound decisions regarding the investment of finite resources. Table 5-1 shows a hypothetical TPP for a gene-drive modified organism.
TABLE 5-1 Hypothetical Target Product Profile (TPP) for a Gene-Drive Modified Organism
| Specification | Minimum Threshold |
| Gene drive construct uptake | >95% uptake in target species |
| Off-target effects | |
|
>98% in target species |
|
at least 5% greater than unmodified male |
| Hybridization with sympatric species | <1% over 10 generations |
| Interaction with existing applications | No change in efficacy of existing application |
| Impact | >60% reduction of target population |
| Time to impact | No greater than 1 year after release |
| Throughput | Two releases per day in target area by one technician |
| Deliverability | Delivered using existing health system |
| Training | Can be deployed by community volunteer |
| Cost at full scale deployment | No greater than current standard technology |
| Manufacturing | Meets demand |
Decisions about which specifications should be included in a TPP, including the standard endpoints to measure and minimum thresholds that should be met, are typically made by a range of stakeholders including academics, industry stakeholders, regulators, and policy makers. Some of these stakeholders can also be responsible for oversight and monitoring of research to ensure due diligence and compliance by researchers. The standards listed in the TPP are then incorporated into study designs and used to inform decisions regarding whether to move from one phase of research to the next. Another key component of phase 0 is establishing site selection criteria for proposed field-based research or staged releases of gene-drive modified organisms in the environment. The criteria are anticipated to be organism- and application-specific and reflect scientific goals, considerations for subsequent trials and ecological risk assessment, ethics, public perceptions, and regulatory requirements (Lavery et al., 2008; Brown et al., 2014; WHO, 2014). Researchers can draw from the advice of individual experts, advisory panels, personal experience, funding agency policies, and published findings to establish decision points. It is unlikely that one site will meet all of the criteria that are initially considered, and so a set of core criteria may need to be agreed upon to help with selection. WHO’s Guidance Framework for Testing of Genetically Modified Mosquitoes (WHO, 2014) suggests that the criteria for contained field trials should include spatial location (for example, an island to mitigate the movement of organisms outside of a study area). Lavery et al. (2008) identified the ability to gain access to communities and their administrative authority as a criterion. Brown et al. (2014) argued that such criteria should include the expertise of a research team in-country, a credible regulatory structure appropriate for research activities, and the presence of target wild-type species, among others. The set of reasonable potential locations may expand or shrink as more information is gathered. Where research infrastructure is lacking, for example, opportunities for capacity-building as a direct result of research funding could be considered, such as occurred with the TARGET MALARIA Project.1
Phase 1: Laboratory-Based Research
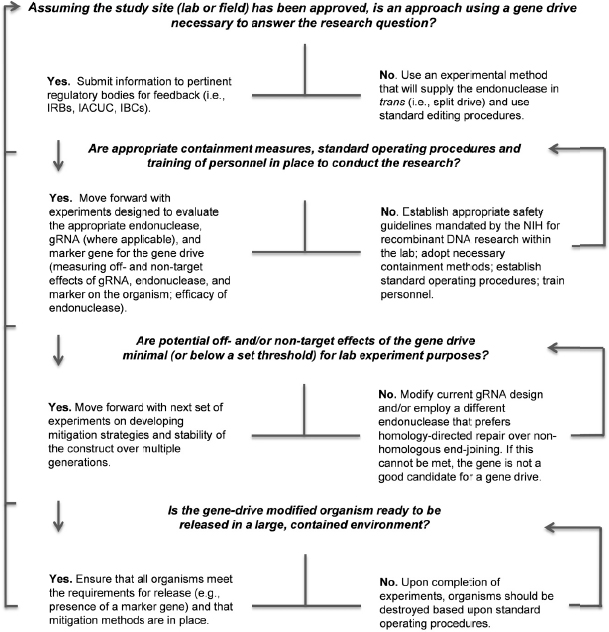
Phase 1 research on gene-drive modified organisms will be performed in the laboratory and physically contained settings under highly controlled conditions. Testing during this phase will inform researchers on the efficacy and safety of the technology in laboratory populations, including whether the gene-drive modified organism demonstrates the molecular, biological, and functional characteristics desired for the chosen application. Physically contained trials will also allow the collection of necessary behavioral data to inform future research phases. Phase 1 research will encompass incremental studies from understanding the biology of the gene-drive modified organism to testing under contained conditions. An example of a “go/no-go” decision tree to help researchers transition from one part of the research to the next is provided in Figure 5-2.
Keeping in mind future efficacy and safety, one important focus of Phase 1 is the optimization of containment settings for gene drive research. In addition to standard bench research, studies can be performed in physical containment that includes small cages, greenhouses, growth chambers, or aquaculture tanks. The choice of containment strategies will depend in part on the species in which the gene drive will be developed and on regulatory or other requirements from the research institution. Similar to good laboratory practices that include procedures to control for unintentional harm of technical staff in the laboratory, training of personnel on standard protocols for using and maintaining small-scale cages and other such facilities are required to prevent unintentional releases.
Another essential component of phase 1 research, i.e., before the organism is release into the environment, is to study bioconfinement and molecular-based strategies to mitigate harm caused by unintended release of organisms. These will be essential tools before research pro-
___________________
gresses to phase 2. Examples include the development of a reversal drive—the currently theoretical process by which the effects of a gene drive are reversed, using either the process that triggered the original gene drive or another process as yet undeveloped—to remove, replace, or restrict activity of the gene drive constructs in the modified organisms would have to be considered in phase 1.
It is also crucial to develop appropriate methods to minimize off-target effects that may reduce the ability for the gene drive to spread into the population. For instance, off-target effects could be controlled by carefully designing specific guide RNAs, optimizing endonucleases, and maximizing DNA repair mechanisms to increase the precision of editing (see further details below).
Also, phase 1 includes laboratory experiments designed to evaluate the stability of the gene drive construct (that is, whether the gene drive construct behaves in a predictable way across generations), and the fitness of the organism (its ability to survive and reproduce) in order to set the baseline population-level effects, and non-target effects. Where possible, organisms studied in phase 1 will possess a similar genetic background as the targeted wild-type organisms to help inform evaluations of the gene drive organism’s fitness and behavior, and to provide datasets that may be required as part of the TPP. Such datasets can also help anticipate interactions within the open environment, such as predator-prey relationships (Hurst et al., 2012).
Phase 2: Field-Based Research
Phase 2 (Field-Based Research) involves studies in natural settings under conditions where dispersal or persistence of the organisms outside the evaluation area is restricted. Field-based research can take place in areas with natural barriers, such as islands which constitute an ideal geographically isolated contained setting, where climatic and environmental conditions are similar from where the organism would normally thrive while physically limiting the dispersal of the organisms (O’Connor et al., 2012). Other examples of research that could be considered in phase 2 are small-scale ecologically or biologically confined field testing of gene-drive modified organisms. Confined field testing entails methods than can control the persistence of an organism in the environment. This can be done by spatial isolation, such as a controlled release occurring at a set distance away from households or in a specific environmental niche (Suwannachote et al., 2009). This can also include climatic isolation where the surrounding environment would be suboptimal for organism survival given a set threshold if unintentional release occurred (Adelman, 2015; Akbari et al., 2015) or even through the use of chemicals to alter specific biological functions in gene-drive modified organisms to reduce their viability (Phuc et al., 2007). Gene-drive modified organisms could also be “field” tested in outdoor large-scale but physically contained environments such as large screen-house facilities (Benedict et al., 2008; Ferguson et al., 2008; Facchinelli et al., 2011).
As for phase 1, phase 2 research is also intended to validate the assessment of the biological and functional activity of the gene-drive modified organisms, but under more natural conditions, and will include the measurement of consistent behaviors, population-level effects and effects of the gene drive on wild-type organisms from the same species or non-target species of specific interest (i.e., beneficial organisms, organisms that may be closely related). The considerations about what endpoints to measure are made among stakeholders prior to seeking regulatory approvals.
Physical marking of test organisms (described later in this chapter) needs to be conducted to help study staff recognize gene-drive modified organisms (Handler and Harrell, 2001). Evaluations in large outdoor cages or screen-houses could include post-test capture of test organisms using manual collection devices or traps to control for unintended release to the outdoor environment. If trials include open field releases into geographically contained or ecologically confined environments, investigators can inform community members immediately surrounding the test area so that they can report organism sightings; meanwhile, the study staff can employ appropriate methods for monitoring and collecting any escapees. Examples of measures that can be employed to control unintentional release or escape beyond the test area, or if required to ‘stop’ the trial for safety or regulatory reasons, might include manual collection techniques (such as aspirators or trapping devices), fumigation with insecticide, or treatment with rodenticides. However, it will be important for investigators to have characterized the resistance profile of test organisms if chemicals are considered as a mitigation strategy (Endersby and Hoffmann, 2013).
The decision about requirements for phase 2 testing conditions for a gene drive will be based on discussions during phase 0 regarding safety and efficacy and will be made in conjunction with the appropriate regulatory authorities (such as authorities with jurisdiction over public health, agriculture, and other areas) and local communities where the field testing will occur. Requirements for obtaining testing approvals will depend on many factors including the application of the gene drive technology and prior knowledge of the potential effects on the receiving ecosystem, and other factors that will be taken into account in risk assessment (see Chapter 6). However, the regulatory requirements for field-based research are expected to be different depending on the application of the research and the study site, since an ecologically confined field trial for gene-drive modified organisms involves intentional, although limited, release into the environment.
Phase 3: Staged Environmental Release
Phase 3 will involve a series of releases into an open environment. Initiating these larger trials and open-environment releases will require thoughtful, evidence-based decisions by a range of stakeholders applying criteria thresholds of the TPP as well as the application of relevant ethical and regulatory practices (see Table 5-1). Phase 3 trials will also include evaluating the release of the technology to inform capabilities and capacity requirements for full implementation and surveillance of the gene-drive modified organisms in phase 4.
As with phase 2, research on phase 3 may also focus on the fitness of gene-drive modified organisms under natural conditions, including elements such as climate fluctuations or variations in target-species densities that may affect the overall performance of the organism. As opposed to non-driving technologies, which can be limited by parameters such as population size or study duration, gene-drive modified organisms will likely persist in nature. Phase 3 studies will therefore help refine parameter thresholds, that once reached, that will allow the gene drive to spread throughout the wild-type population. To that end, characterization of the population structure of wild-type organisms of the same species as gene-drive modified organisms in the setting where testing will occur will be important to guide study design related to release rates (e.g., density and timing), as well as expectations of gene drive spread based on estimates of population size in the open field environment (Jeffery et al., 2009).
While measurement of effects as pre-defined in the TPP will likely remain the focus of the staged environmental release, the measurement of the impact of the gene drive on other populations within the ecosystem is also an important an component of phase 3. For example, trials requiring the demonstration of an epidemiological impact (e.g., reduced disease prevalence, population suppression, or recovery of a threatened species population) can be used to inform decisions about whether the gene-drive modified organism could be released in other countries.
Phase 4: Post-Release Surveillance
The final phase of the testing pathway encompasses surveillance and monitoring. The purpose of this phase is twofold: (1) to determine whether intended effects of the broad scale release of gene-drive modified organisms are sustained over time; and (2) to detect any changes in the organisms or the ecosystem. For example, in a release program of gene-drive modified mosquitoes unable to carry the avian malaria parasite (Case Study 3) it would be important to monitor for the presence of the mosquitoes and confirm that they continue to be unable to carry the parasite. Also, long-term surveillance of honeycreeper population size and health will be needed. As noted in WHO (2014), efficacy can change due to changes in genotype of the organisms, or due to external factors such as weather or human activities. In addition, there could be unexpected effects when the gene-drive modified organisms are released (or expand) into new areas.
Monitoring also helps to determine whether any changes are needed in management of the gene drive (e.g., the possibility that mutations in the gRNA could arise over generations, leading to other recognition sites that were not detectable in early-phase testing), the gene-drive modified organism, and the release program (e.g., coverage, frequency, and density), or other aspects of an integrated program (e.g., the use of a complimentary, alternative strategy). It will also be important to continue to assess public support through surveys and other social science research tools (Hanh et al., 2009).
Longitudinal monitoring over varying time periods may be required to build robust and informative datasets regarding the effect of seasonal changes on the biology, behavior, and species composition of wild-type organisms within the target ecosystem (see Chapter 2). Simulation modeling of existing datasets, and those generated during research, will be an important component of research using gene drive technology (Marshall and Hay, 2012; Dutra et al., 2015). Open-access data repositories and standard operating procedures will facilitate the use of such
data and models and inform standards for research design and monitoring schemes for gene drive research. In one example, Crain et al. (2013) used existing data from field research and “a modeling analysis to predict the dynamics when two Wolbachia infection types do not remain geographically isolated.”
Monitoring and surveillance are necessary to determine whether the approach continues to work over time, but these activities can be expensive and logistically challenging, particularly for low- and middle-income countries. Thus, it will be important to select the measurement tools, timeframes, and protocols that are most informative and sustainable.
CONTAINMENT, CONFINEMENT, AND MITIGATION STRATEGIES
Selecting or developing appropriate confinement and containment strategies is challenging due to the wide range of proposed gene-drive modified organisms. The case studies discussed in this report focus on mosquitoes, mice, and two species of plants. Certain mitigation measures may be an option for some types of research or certain organisms but not to others. For example, creating a gene-drive modified mosquito susceptible to insecticide might be a useful mitigation precaution, for which there would be no parallel with another type of gene-drive modified organism. Unless otherwise specified, the following sections focus on strategies that could potentially be used for any type of organism.
Two important dimensions of research carried out through the phased testing pathway are:
- Containment and confinement to reduce the potential for unintended release or persistence of gene-drive modified organisms, respectively; and
- Mitigation strategies to address potential harmful off-target and non-target effects.
Given the recent recognition by many scientists that CRISPR/Cas9 technology likely holds the greatest promise for rapidly creating gene drives in the laboratory for deployment in the field (Oye et al., 2014; Akbari et al., 2015), the considerations outlined in this pathway are primarily geared toward this technology. However, some of the same principles can be applied when using other gene drive methods described in Chapter 2.
Methods and strategies considerations for the choice of confinement and containment measures will include whether the organism will be evaluated only in the laboratory (phase 1) or in an open environment (phases 2-4).
It is important to highlight that while some effects could be harmful, some off-target and non-target effects could also potentially be beneficial, and some effects, such as cost to fitness, can be viewed as beneficial or harmful depending on the objective of the gene drive strategy. For example, regarding population suppression, a slight reduction in fitness could be considered unimportant or perhaps as a modest benefit (for example, a gene drive to reduce the population of a pest species). However, a reduction in fitness in the context of population replacement could be considered detrimental (for example, a gene drive intended to prevent a species from going extinct). Another important reason to mitigate off-target effects is that they may confound results obtained with gene drives, making it difficult to attribute phenotypes to the edited target. However, not all off-target effects are considered equal, and the number of off-target editing events may not be as important as the identity of these events (Mathews et al., 2015; Church and others personal communication, Human Gene Editing Summit, Dec. 1-3, 20152). It is crucial to consider the functional consequences of off-target effects and whether their presence is detrimental or advantageous with respect to the purpose of the gene drive.
In addition to considering off-target and non-target effects, it is important to characterize the biology and ecology of the target organism and its environment to fully understand and con-
___________________
2See http://www.nationalacademies.org/gene-editing/Gene-Edit-Summit/index.htm.
trol for these unintended effects (see Chapter 2). It will be important, for example, to support characterization studies over multiple generations to inform models of organisms’ behaviors and properties before moving to field-based studies. Such research is critical for developing effective gene drive applications in various ecological contexts and for reducing uncertainty via informed risk assessment (see Chapter 6).
This section below outlines various confinement, containment, and mitigation strategies for consideration in gene drive research, as well as mitigating other types of concerns such as “tinkering with nature” or “who gets to decide whether a gene-drive modified organism should be released” concepts highlighted in Chapter 4 (Values), Chapter 7 (Engagement), and Chapter 8 (Governance).
Containing and Confining Gene-Drive Modified Organisms
In general, confinement and containment requirements will be worked out on case-by-case basis in consultation with an IBC or equivalent institutional research oversight body. Carefully discussing containment and confinement measures during phase 0 is crucial since organisms containing a gene drive will, by essence, spread the gene drive if released in an environment that promotes their survival and reproduction. In order to prevent lab-based gene drives from escaping into wild populations, many researchers have offered suggestions for developing methods to contain gene-drive modified organisms (Esvelt et al., 2014; Oye et al., 2014). The following containment methods could be used for gene drive studies in the laboratory. These containment mechanisms are also applicable to gene drives designed for various stages of field release (see phases 2-4) and are an important component of any mitigation strategy.
A split gene drive may be equally as effective as intact gene drive methods for modifying an organism’s genome through germ line transmission (such that all cells are edited), while increasing containment capabilities. In a split gene drive, the components (Cas9 or other HEG, gRNA, and donor template; see Chapter 2, Figure 2-2) are supplied separately to the organism. With this technology, a gene drive is not actually created due to the manner in which the components for the editing are delivered to the organism. This method is particularly useful when performing standard editing techniques to alter specific genes as would have been carried out previously using more “traditional” methods. For example, one could use organisms transgenic for Cas9 (or the gRNA) and supply the other component independently, along with any donor DNA that might be required to modify the organism. This type of experiment has been successfully carried out in Drosophila (CRISPR-it; Port et al., 2015) and yeast (DiCarlo et al., 2015) and may be applicable to other organisms, especially plants and animal models of disease where transgenics are possible or in which gene editing is feasible. Although there is a small possibility that these individual components could recombine and create a gene drive, this possibility is remote and would not preclude the use of this system in the laboratory. Nonetheless, it will be important that the general considerations for gene drive usage in the laboratory, as outlined above (see Figure 5-2), be followed, particularly with respect to the choice of endonucleases, gRNAs, and measurement of off- and non-target effects, and employment of specific containment methods, standard operating procedures (SOPs), and training protocols. If the creation of an intact gene drive is required, perhaps due to limitations associated with the use of a split gene drive in a particular model system or because the ultimate goal builds toward environmental release (as in the case studies), then guidelines listed in Table 5-1 and described in detail below will also be important for researchers.
For organisms with a gene drive used exclusively in the laboratory and not intended for release, containment strategies may be minimal if appropriate mitigation strategies are employed (see next section). To this end, researchers are encouraged to follow principles of Good Laboratory Practices (GLPs), including, for example, an internal monitoring system based on IBC feedback, as well as training for staff, researchers, and students in necessary SOPs. The training
might also include specific instruction about the ecological differences between transgenic and gene-drive modified organisms.
If a specific marker can be visualized (e.g., using fluorescent proteins; see below), all personnel will need to see examples of the modified organisms to avoid confusion with other organisms without the gene drive and provided with appropriate materials, such as vials or cages, for collecting test organisms found outside of their normal area. Physical marking of adults, such as the use of fluorescent proteins, can allow for easy visualization of the research organism being studied (Hagler and Jackson, 2001). Reporting notices for the sighting of these organisms can be posted on office or laboratory doors. Keeping a form with contact numbers and sighting dates in work spaces will facilitate the ability of laboratory staff to report identification of any collected specimens to IBCs (as specified), and follow-up with resolutions to containment breaches, which includes informing surrounding laboratories of accidental releases. Because live organisms are mostly used in phase 1 testing (e.g., to identify variation in mating or other behaviors), traps are recommended in testing laboratories and rearing facilities to facilitate capture of specimens that have escaped or were released unintentionally. The US Department of Agriculture has developed guidelines for containment that are expected to apply to gene-drive modified organisms and research under laboratory testing conditions (APHIS, 2002).3
Containment and confinement measures can be categorized as being extrinsic (e.g., in the laboratory or in the ecological or geospatial environment) or intrinsic (e.g., molecular or reproductive factors) with respect to the gene-drive modified organism (Esvelt et al., 2014; Akbari et al., 2015). Gene drive research regulations will most likely fall under the Coordinated Framework for the Regulation of Biotechnology as it also regulates genetic engineering in general. As such, gene drive research would receive oversight from IBCs. This is reviewed in detail in Chapter 8 on Governance.
Extrinsic physical containment of organisms in the laboratory, as outlined in the current National Institutes of Health guidelines for organisms containing infectious agents, in the Coordinated Framework for the Regulation of Biotechnology or in the Arthropod Containment Guidelines4 can follow standard Arthropod Containment Level 2 criteria in the case of mosquitoes or other more stringent criteria depending upon the type of containment used, the organisms involved and the purpose of the experiment. These guidelines may be sufficient to conduct research with organisms containing gene drive constructs in the laboratory. Methods of physical containment may include conducting experiments in a biosafety cabinet or in a separate room with a double-door entryway; the use of appropriate directional air flow; the use of air cloths or curtains (where appropriate); storage of tubes of gene-drive modified organisms on a separate bench, refrigerator, or freezer; housing of gene-drive modified organisms in cages or tanks separate from their wild-type counterparts (and in different rooms); installation of rodent-proof doors; securing plates of gene-drive modified organisms with parafilm; and, upon completion of experiments, destruction of all materials through autoclaving, freezing or microwaving (Akbari et al., 2015). Other standard laboratory practices would also apply here, including wearing personal protective equipment such as lab coats and gloves and appropriate clothing; cleansing benches with 70% ethanol upon completion of experiment; and soaking of glassware for 24 hours in Wescodyne solution before cleaning (Akbari et al., 2015). Ecological confinement methods are also recommended to help prevent gene-drive modified organisms from mating with organisms in the native population or persisting in the context of the environmental conditions or geographical location of the laboratory. For example, this might involve working with species that do not normally survive in the region where research is being conducted; however, this might not be feasible in all instances as it could prevent research on gene drives from being conducted.
___________________
3See https://www.aphis.usda.gov/plant_health/permits/downloads/arthropod_biocontrol_containment_guidelines.pdf.
Intrinsic confinement and containment measures are also important. For example, the gene-drive modified organism could exhibit a barrier to reproduction such that it cannot mate with organisms in the wild. Additional methods of molecular containment can be explored, including the use of a split gene drive in which Cas9 is introduced separately (e.g., on a plasmid) from the gRNA (see above). Providing Cas9 (or other endonucleases) in trans has been successful in generating gene drives in yeast (DiCarlo et al., 2015) and Drosophila (Port et al., 2015), but this is likely to be species- and locus-dependent. One advantage of this method, is that less strict extrinsic confinement and containment measures would be necessary, as these organisms are considered standard transgenic animals and are thus subject to regulations already in place. Another approach is to design gene drives to be “self-limiting”, for example, by carrying both a gene that encodes for a toxin and another gene that confers immunity to the toxin. Such gene drives could self-destruct either over time or upon addition of a chemical (Gould et al., 2008; Marshal and Hay, 2012). One final intrinsic containment mechanism is to target sites for which the gene drive is only found in a laboratory organism and not in wild-type organisms.
Mitigating Potential Harms
Restoration of Wild-Type Organisms
When the intent of the gene drive is population replacement, restoration of the wild-type version of the sequence edited by the gene drive (including off-target effects) may be desired or required. One mitigation method that addresses this issue and has been demonstrated exclusively in the laboratory is the use of a reversal gene drive (DiCarlo et al., 2015). This method is based on the use of another gene drive that re-introduces the original genetic sequence into the edited organism, along with modifications to it such that it can no longer be edited in the future. This method requires a two-step modification of the organism through the use of two rounds of editing (i.e., introduction of two different CRISPR/Cas9 systems). Another mechanism proposed is an immunization drive that, when given to an organism, will prevent a second gene drive from being propagated within the organism by altering sequences targeted by the second drive. The immunization drive could be deployed so that non-target species do not inadvertently receive the gene drive. It is important to note that with either of these methods, Cas9 and the gRNA would still remain in the genome, which could cause additional undesirable effects due to persistent DNA breaks caused by Cas9. Another strategy is to adapt a new transgene system developed in Drosophila called Cas9-ablated chain termination, where possible (Wu et al., 2016), which serves as a molecular “brake” to cleave Cas9 and thus disable it in Cas9-containing organisms, thereby rendering the gene drive inactive. Finally, one could maintain a population of wild-type organisms that, upon disabling of the gene-drive modified population through any of the methods described above, could be released to re-establish the native population.
Redressing Undesirable Ecological and Evolutionary Consequences
For redressing undesirable ecological and evolutionary consequences, a strategy could include monitoring specific non-target species alongside the gene-drive modified organism. For timely recognition of undesirable ecological consequences, the best approach is to monitor the densities of species most closely linked to the target species via trophic connections (e.g., competitors whose diets overlap that of the target species or predators that might prey on the target species). One of the most likely undesirable evolutionary consequences would be the movement of the gene drive into a closely related, non-target species via reproduction between two different but related species (i.e., hybridization). Close evolutionary relatives could be monitored in the wild for the appearance of the drive unless hybridization is known to be impossible (i.e., if resulting hybrids do not produce fertile eggs). A potential example is Palmer amaranth (see Case Study 6), which has been shown to hybridize with other species. The interventions for both types
of consequences could include the re-introduction of affected species after the gene drive has been eliminated. In both cases, the speed with which gene drives can spread suggests that monitoring must be in place before the gene drive is introduced so that any unwanted effects can be recognized quickly (see description of phase 3 and phase 4 activities above). This is especially important in the context of potential ecological consequences of a suppression gene drive, because the loss of a species can, in some cases, produce effects that cascade through the ecosystem (Estes et al., 2011). These kinds of effects can be reversed (e.g., Shapiro and Wright, 1984) even by re-introduction of the lost species (Bundy and Fanning, 2005; Mumby and Steneck, 2008). Depending upon the reproductive capacity of the edited organism (e.g., its generation time), it may take some time for all organisms within the population to have the original phenotype restored. Maintenance of a copy of Cas9 and a gRNA could also have deleterious effects over time on the organism and other non-targets. Non-target effects may also be hard to control, and redressing potential undesirable ecological and evolutionary consequences of the gene drive, even when accounting for changes over time, may be difficult. These issues are discussed in detail in Chapter 2.
Optimization of gRNA Design
Off-target effects are going to occur with any gene editing methods associated with homing endonucleases that involve the creation of breaks in the DNA (e.g., Cas9, ZFNs, TALENs) as well as gRNA hybridization (CRISPR/Cas9). However the rate will likely be organism or cell type-dependent. In order to mitigate such effects for RNA-guided editing, it is critical to optimize gRNA design. To achieve high specificity, evaluation of the target DNA to identify sites for gRNA hybridization is an important step. If the target lacks specificity (i.e., if the DNA sequence resemble others in the genome) then other sequences in the genome will be targeted. Likewise, chromosomal rearrangements after imprecise repair will occur, which may trigger the activation of aberrant signaling leading to cell dysfunction (Koo et al., 2015).
To mitigate harms related to off-target effects of gRNAs, scientists have used web-based bioinformatics tools. These tools help assess the degree to which the gRNA(s) may target other sequences within the reference genome of the chosen organism and the genomes of other organisms. This is only possible to do, however, if genomic sequences of targeted and non-targeted organisms are available. Targets that have few or no closely related sequences in the genome can also be chosen.
If the gRNAs are specific (i.e., if the intended phenotype does not change over time), and if any change in fitness does not prevent the spread of the organism, then the gene drive has a chance to be successful in the wild. Re-introduction of the wild-type or original allele can also be undertaken to ensure that the phenotype in the presence of the gene drive is attributable to the editing of that allele (Bono et al., 2015). A powerful way to complement computational methods is the use of Next Generation Sequencing (NGS), which can generate a genome-wide profile of the nuclease activity. Once the putative off-target sites (i.e., sites that resemble the targeted DNA sequence) have been detected computationally, these sites are compared to the presence of nuclease activity identified at these specific sites by NGS. However this technique introduces some “observational bias” based on the assumption that off-target sites will resemble the target site, while others can exist. Other considerations include the fact that sequencing-based assays can lead to artifacts (Koo et al., 2015) that may preclude actual detection of off-target effects (Mathews et al., 2015), and the fact that the configuration of the DNA may also impact whether potential off-target sites are even accessible to the nuclease (Sander and Joung, 2014; Koo et al., 2015). To address such constraints, a new NGS method called Genome-wide Unbiased Identification of DSBs Enabled by sequencing has been developed to physically tag all potential cutting sites, including off-target sites, in an unbiased fashion (Tsai et al., 2015). When compared to computational methods, the results using this sequencing method revealed that off-target effects were observed at higher frequencies than expected. Several groups have now used such tools and
others to reveal off-target effects in various cell lines (Frock et al., 2015; Wang et al., 2015) including pluripotent human cell lines (Chan et al., 2015). Therefore, computational models to predict off-target sites and the use of NGS to profile the activities of human and animal model gRNA are necessary to maximize activity of the gRNA while minimizing potential harmful effects (Doench et al., 2016).
Optimization of Endonuclease Cutting Efficiency
Similar to the considerations for optimizing gRNAs described above, different endonucleases (e.g., Cas9 or other homing endonucleases) can vary in their ability to efficiently cut the targeted sequence. To this end, researchers have used a mutant version of Cas9, called Cas9 nickases, along with two gRNAs targeting two different sites, one on each side of the DNA strand. This endonuclease only makes breaks on one strand of the DNA as opposed to both strands (Ran et al., 2013); it also engages a higher-fidelity type of repair than the one used after a Cas9/gRNA-mediated cut is made. Other genetically engineered Cas9 variants (Anders et al., 2014; Nishimasu et al., 2014) cleave at different PAM sequences and/or with higher efficiencies and reduced off-target effects (Kleinstiver et al., 2015; Slaymaker et al., 2016). A new Cas9-like protein has now been identified, called Cpf1 (CRISPR from Prevotella and Francisella 1), that functions through the use of a single gRNA molecule; this protein generates DNA breaks in the form of overhangs (a staggered cut) instead of blunt ends, cuts at a greater distance from the PAM on the target site, and therefore does not disrupt the PAM upon cutting (Zetsche et al., 2015). These other Cas9 endonucleases have yet to be evaluated for efficacy and efficiency in living organisms. Funding for these latter experiments to address the efficacy and specificity of gene drives is critical for the future deployment of gene drives in plants and animals. Importantly, the presence of Cas9 carried in the organism will need to be evaluated to determine whether Cas9 has a harmful effect on organism fitness that would prevent the spread of the gene drive (discussed in Chapter 2), as this would raise significant questions regarding the ability of the gene-drive modified organism to function.
Optimization of Homology Directed Repair (HDR) Versus Non-Homologous End Joining (NHEJ)
When DNA cleavage occurs at the targeted site, there are two major categories of DNA repair that can restore the DNA structure: homology directed repair (HDR), which requires a homologous sequence to guide repair, and non-homologous end joining (NHEJ), which does not need a homologous template for repair and just “seals” the cut DNA ends together. Depending on the application, gene drives may require the introduction of specific genes into the target chromosome and thus would require HDR. This could be one of the biggest challenges facing gene drives, because the mechanism of repair will depend on species, cell cycle stage, cell type, and stage of development (Esvelt et al., 2014).
In order to facilitate the HDR pathway and allow the introduction of an exogenous gene, Cas9 nickases (see above) can be used, since it cuts a single strand of DNA instead of the two strands. Similarly, the nuclease Cpf1 should (theoretically) more easily allow for insertion of DNA due to the presence of compatible overhangs. Other options involve the repression of genes involved in NHEJ or the activation of genes responsible for HDR (reviewed in Esvelt et al., 2014). For instance, to optimize HDR during the development of their gene drive-modified mosquitoes, Gantz et al. (2015) and Basu et al. (2015) included dsRNAs directed to both Cas9 (on the construct) and a gene expressing a protein essential for the NHEJ activity in the mosquito. While it was not directly measured in this study, upon injection into the mosquito, this gene drive construct silenced the Cas9 protein and reduced the activity of NHEJ in favor of HDR, allowing for the insertion of the entire gene drive construct in the genome. Recently, Hammond et al. (2016) observed a bias toward the HDR repair mechanism using the CRISPR/Cas9 technology in mosquitoes without such optimization but more research would be needed to confirm such results.
Evaluating the Stability of the Gene Drive Construct Over Multiple Generations
Another challenge related to repair mechanisms is that gene drive resistant alleles may result when NHEJ repairs the break caused by homing endonucleases, leading to the loss of the cleavage site. Such alleles without the cleavage site will become resistant to the effects of the gene drive. If enough individuals contain the resistant allele, then the gene drive may become ineffective. One way to reduce the incidence of resistance would be to use multiple gRNA because resistance would require mutations at several target sites. A similar challenge stems from the fact that different DNA sequences for the same genes are found in nature (known as polymorphic sequences). This could prevent the action of a gene drive because the gRNA may not be designed to recognize such sequences outside the laboratory. If these “natural” resistance alleles are common in the wild, the gene drive may be ineffective.
The stability (or lack thereof) of a gene drive, indicated by the degree to which the modified genetic element and the driving capability are retained over multiple generations, needs to be determined on a case-by-case basis. To evaluate the gene drive’s stability and to estimate its effectiveness, it will be important to carry out a variety of experimental assays, including the use of simulation modeling to predict the spread of the gene drive over multiple generations and any population-like effects using laboratory data (phase 1). These results can be compared to field data obtained from the non-driving study (see Quantitative Approaches, below) (Esvelt et al., 2014). For example, the gene drive stability will need to be measured in a stepwise manner first in the laboratory populations and then in wild caught populations. This is because there may be no perceived instability in the laboratory population, but potentially increased opportunities for instability in the wild population. If such instability arises in the wild, then there is no reason to take this gene drive outside of the laboratory to phase 2. The exception to this is when the gene-drive modified organism is being designed for field release for use in population suppression, such that any loss of organismal fitness could be advantageous for achieving the release objectives, as long as it does not affect the spread of the gene drive.
Determining the Effects on Organismal Fitness
It is imperative to use quantitative methods to evaluate whether the expression of the homing endonuclease (for example, Cas9), the gRNA, or the cargo template (singly expressed or in various combinations) affect a gene-drive modified organism’s fitness, relative to its wild-type counterparts. This evaluation would involve a “fitness assay” that would comparatively examine fitness parameters for the engineered genotype, relative to the unaltered wild-type organisms, ideally using established empirical methods in the particular biological system or a closely related one. In general, the fitness assay approach compares the relative ability for a test genotype to produce viable offspring to that observed with wild-type organisms, and the experiment is conducted with independent empirical replicates (Chippindale et al., 2001). This repetition is necessary to provide sufficient power for a statistical analysis to detect measurable fitness differences between the genotypes. In addition, it may be useful to gauge relative survival of the engineered genotype relative to the wild-type through replicated assays of relative lifespan (Rose et al., 1992), which statistically measure whether the genome alteration negatively impacts physiological health to shorten the average lifetime of the individual. These same types of assays will also need to be conducted for an organism in which the genetic alteration has been made using a different editing method, is found naturally in the population, or is created through genome-wide mutagenesis for all comparison purposes. Although it is often assumed that genome alterations, including gene drives, will tend to negatively impact individual fitness relative to that observed in the unaltered wild-type organism (e.g., due to the addition of foreign DNA that slows replication, and/or interferes with native transcription and translation), this is an assumption that must be verified using rigorous empirical analyses. This hypothesis could be tested by performing fitness assays in the laboratory that measure the relative number and quality of viable gametes
produced by gene-drive modified and wild-type organisms, as was applied by Hammond et al. (2016) to gene drive constructs in Anopheles gambiae, and by performing survival assays that compare relative viability of altered and gene-drive modified and wild-type organisms (Isaacs et al., 2012). Additional field trials can be used to examine these fitness effects under more natural conditions. The ability to quantify these effects on organism fitness, if not masked by compensatory pathways that are up-regulated by the organism as observed previously (Rossi et al., 2015), will lead to questions regarding whether gene drives provide the best technology for editing a specific gene, and whether fitness effects are consistent with intended applications.
Using Visible Markers
Gene-drive modified organisms that possess, as part of their genetic cargo, a marker gene in order to facilitate identification can help researchers distinguish a gene-drive modified organism from wild-type or other conventional transgenic organisms. Although still under development, examples include the addition of a gene encoding a fluorescent protein that would be expressed in a region of the organism that could be easily screened/monitored (e.g., eye, skin) without requiring sequencing assays which necessitate adequate equipment and expertise, are more invasive, and may take longer to obtain results. Alternatively, the gene drive could target an additional, non-essential gene for mutation to generate a visible phenotype that could be scored. These are both examples of common genetic marking techniques that have already been employed by researchers who have constructed gene drives in Drosophila (yellow body phenotype in Gantz and Bier, 2015) and mosquitoes (white-eye phenotype and fluorescent marker in Gantz et al., 2015; fluorescent markers in Hammond et al., 2016).
The generation of unique labels for gene drive constructs, in the context of other conventional transgenic organisms possessing similar tags, could be problematic. The ability to do so will depend on the availability of specific promoter and enhancer combinations to drive marker expression in select cell types to allow for efficient and effective screening. For example, 95% of mosquito strains are labeled with only two fluorescent tags because the efficacy of expression of others is low, and there is currently a dearth of information regarding how other markers could be used in mosquitoes (M. Benedict, personal communication). This represents a significant challenge for the field. It is highly desirable to develop a consensus opinion within the community working on a particular organism with respect to how gene-drive modified organisms will be labeled and identified. Although not absolutely required, the inclusion of a visible marker is recommended when making a gene-drive modified organism.
Quantitative Approach to Evaluate Success and Impact
According to Sinkins and Gould (2006) “[m]athematical modelling can help to predict the utility of different gene drive systems, as long as realistic values for the fitness costs of the effector transgene and for the pest’s population structure are used.”
Quantitative and computational methods are vital tools for evaluating biological applications, and for advancing fundamental knowledge in biology. Often the overarching goal is to use bioinformatics, mathematical modeling and computer simulations to elucidate the dynamic properties of a biological system at one or more levels (e.g., gene, genome, population, community, and ecosystem). When this approach involves a probabilistic framework, it is possible to predict which factors are most likely to influence the success of biological applications and to reveal the variables that most influence dynamics in biological systems (Otto and Day, 2007). Such quantitative approaches can never incorporate all of the variables at play in biological systems because the mathematics quickly becomes too intractable or the simulations exceed available computing power. Nevertheless, history shows that quantitative methods can usefully identify those variables that are most important in determining dynamic properties of biological systems, especially
using an iterative process where empirical observations are employed to further refine the accuracy and predictive power of quantitative models (Kitano, 2002).
Gene drive technology is advancing quickly, and offers the possibility of an efficient tool to study fundamental questions in biology as well as a method to address problems in public health, conservation biology, agriculture, and other applications (Esvelt et al., 2014; DiCarlo et al., 2015; Gantz and Bier, 2015; Hammond et al., 2016). But the overall success and impact of gene drive technology hinges on many factors, especially when the strategy involves the release of genetically altered individuals into natural communities. The most proximate challenge is to gauge whether gene drive mechanisms such as gRNA editing are precise in altering only the target locus, versus inefficiently changing unintended (off-target) loci. If the goal of the gene drive technology is to alter genotypes for strictly laboratory-based basic research purposes, a certain level of inaccuracy may be tolerable. Still, if such experiments were intended to examine questions such as genetically inherited diseases in model organisms, any imprecision could confound assumed relationships between genotype and phenotype and thus slow the advance of knowledge. If gene drive technology inaccurately creates genotypes intended for field release, this outcome necessarily causes a disconnect between the expected introduction of individuals into the target population and the actual individuals that are placed in the wild. It may be impossible to absolutely know whether and how this proximate inaccuracy holds repercussions for overall success and environmental impact of the intended field release strategy, until the release actually occurs and the system is closely monitored. However, quantitative and computational methods should be useful in gauging the probabilities of success and possible outcomes, whether the drive is strictly laboratory-contained or intended for field release. For this reason, it is prudent for research on gene drive technology to include quantitative tools that help to refine the accuracy of their associated risk assessment frameworks.
As previously reviewed, current gene drive technologies mimic natural gene drive mechanisms (e.g., meiotic drive) that have been studied intensively, especially at the molecular and population biology levels (Jaenike, 2001). Similarly, biological control efforts are longstanding, and we possess knowledge of how released organisms can impact populations and communities (van Driesche and Bellows, 1996; Stiling and Cornelissen, 2006). Nevertheless, current gene drive technologies and their intended applications differ in several respects from naturally occurring gene drive mechanisms and prior biological control efforts. For example, if a limited number of non-driving genetically modified organisms are released into the wild, this fundamentally differs from the release of gene-drive modified organisms because only the latter case involves sustained modification of individuals across multiple generations in the target population. Therefore, it would be naïve to assume that intensive quantitative modeling and other prior efforts would suffice to predict the accuracy of gene drive manipulations and determine how these altered genotypes would affect natural communities. This possible disconnect between prior knowledge and current goals of gene drive technology offers further support for the argument that quantitative and computational tools should be developed for each gene drive study because researchers should not assume that probabilities of success and environmental impacts could be drawn conveniently from prior data in a different biological system.
Quantitative approaches offer the opportunity to efficiently examine uncertainties related to the success and impact of gene drive technology, at all stages of research. Because monetary resources for basic research and for field trials can be very limited, quantitative tools also offer the opportunity to efficiently explore whether a genetic manipulation or field release may be successful, before actually devoting funds to conduct the work. In particular, this approach can be used to evaluate key steps in the phased testing pathways described earlier in the chapter, either at individual stages or more holistically across multiple stages. In this way, scientists can gain a broader predictive framework for whether the basic research goals can be properly advanced and whether the field release may truly work when launched.
Often these modeling approaches can assess key thresholds, such as how many individuals must be released for the gene drive strategy to likely succeed in sufficiently altering the target
population. Similarly, a wide range of effects may be vital for predicting the success and impact of the gene drive technology. Considerations may include: the predicted average fitness of altered individuals relative to the genotypes in the targeted wild-type population; how quickly or slowly should the altered individuals be released to maximize (or minimize) their impact in the environment; how current sex ratio and size of the target population may influence outcomes of release; and whether geographic barriers or other effects of landscape ecology will impact the likelihood of the gene drive spreading successfully.
LEARNING FROM FIELD RESEARCH AND BIOCONTROL EFFORTS WITH OTHER TYPES OF MODIFIED ORGANISMS
Due to the expectation that organisms will disperse in the open environment during phases 3 and 4, causing the gene drive to spread and potentially impact broader human and environmental communities, mitigation in these phases offers additional challenges to those described for laboratory (phase 1) and contained releases (phase 2). Past experience with biocontrol efforts and research with modified mosquitoes, such as Release of Insects with Dominant Lethality (RIDL®) technology and infection with Wolbachia bacteria, can inform questions about population biology and ecosystem dynamics when considering mitigation strategies for research using gene drive technology.
Biocontrol Pest Species
Biological control, defined by Popovici et al. (2010) as “the release into the environment of a biological agent to control a given pest through mechanisms such as predation, parasitism, herbivory or disease” of agricultural, livestock and human pests has been undertaken successfully for centuries (Wackers et al., 2007). Examples of the range of biocontrol applications from Australia alone were reviewed by Popovici et al. 2010, and “include the release of myxoma virus to control rabbit populations (Fenner, 1983; Saunders et al., 2010), the release of Cactoblastis moths to control prickly pear (Opuntia spp) (Dodd, 1940), the introduction of dung beetles to manage cattle dung and the bush flies that breed in it (Edwards and Pavri, 2007) and the control of floating Salvinia weed (Room et al., 1981) using the beetle Cyrtobagous singularis.”
Intentional Release: Large-Scale Deployment
Sterile insect technique (SIT), “a method of pest control using area-wide inundative releases of sterile insects to reduce reproduction in a field population of the same species,”5 continues to be employed on a large-scale to control the new world screwworm, Cochliomyia hominivorax (Knipling, 1955; Vreysen et al., 2007). SIT has also been used to control the Mediterranean fruit-fly (also called the medfly) Ceratitis capitata and as part of an integrated pest management program. In addition SIT has also been employed to control the pink bollworm (Pectinophora gossypiella) since 1999, and during the cotton season, approximately 25 million sterile moths, reared at a facility in Phoenix, Arizona, are released per day.
Using the SIT approach as its foundation, the genetically engineered technique RIDL utilizes transgenic insects with a conditional, dominant, female-specific lethal gene that inhibits female offspring from developing into adults (Thomas et al., 2000). Recently, successful transformation of the diamondback moth using the piggyback transposable element prompted the development of RIDL as a control measure for diamondback moths by the biotechnology company Oxitec (Martins et al., 2012; Kelland, 2015). The RIDL approach to diamondback moth control has been evaluated in both the laboratory and contained environments; field test are un-
___________________
5FAO: http://www-naweb.iaea.org/nafa/ipc/sterile-insect-technique.html.
derway6 (Harvey-Samuel et al., 2014; Waltz, 2015). The development of RIDL approaches for the control of agricultural pests and invasive species, like the diamondback moth, represent another tool for integrated pest management programs. RIDL mosquitoes have also been released in several countries, including the Cayman Islands, Panama, Malaysia and Brazil,7 to suppress local mosquito populations for dengue control (see Case Study 2).
Another biocontrol approach is the use of Wolbachia infection. Mosquitoes infected with natural Wolbachia symbionts have been released in the United States,8 Australia, Indonesia, Vietnam and Brazil.9 The bacterial symbionts in the genus Wolbachia are widely distributed in insects (Werren et al., 1995, Werren and O’Neil, 1997; Bourtzis and Braig, 1999; Stouthamer et al., 1999) and are transmitted vertically from mother to offspring through a phenomenon known as cytoplasmic incompatibility (Ghelelovitch, 1952). Because only Wolbachia-infected females can successfully reproduce with infected males, all the offspring of infected females will carry Wolbachia, which can then spread quickly resulting in a large proportion of the local mosquito population eventually becoming infected. The use of Wolbachia infections is advantageous because it reduces the lifespan of insect hosts (Sinkins et al., 1997; Dobson et al., 2002; Ahantarig et al., 2011; Bull and Turelli, 2013) and confers resistance to infection with dengue and chikungunya viruses in Aedes aegypti (McMeniman et al., 2009; Moreira et al., 2009a; Bian et al., 2010).
This technology includes options for sustained releases similar to RIDL for population suppression; it addition, it offers the opportunity for the release of self-sustaining variants that could lead to population replacement, for example, by reducing the mosquitoes’ capacity to transmit specific pathogens.
Although these technologies have encountered hurdles during their development, protocols, strategies, and guidelines were produced in anticipation of the ultimate release of suitably engineered mosquitoes (Beech et al., 2009a,b; Mumford et al., 2009), that include sequential steps from concept to the safe and responsible release of engineered mosquitoes. These steps include development of cage (contained) trials, community engagement, and considerations of relevant ethical, social, and cultural issues. Remarkably, from what seemed like a position of insurmountable challenges, almost all of the problems have been resolved. The Gates Foundation in particular has strongly supported groups to develop recommendations and protocols related to transgenic mosquito releases (Singer et al., 2007; Lavery et al., 2008; El Zahib-Bekdash and Lavery, 2010; WHO, 2010).
The approval to deploy transgenic Aedes aegypti using RIDL technology in Brazil for dengue control demonstrates that assessment of benefits and harms based on data gathered on the biology, ecology and planned mitigation strategies can support a favorable decision (see Case Study 2). The concerns addressed are anticipated to be similar to those of gene drive technology (WHO, 2014). For example, considerations include exposure to humans, the ability of the organism to have modified competency for pathogen transmission, the possibility of gene flow to other species, the likelihood of an increase in the population of other species due to the reduction of the target organism, other environmental impacts, and an assessment of the functionality of a designed mitigation strategy to minimize unintentional harm—in this case, the requirement of tetracycline in an aquatic habitat to suppress lethal gene activation (Phuc et al., 2007).
Unintentional Release: Transboundary Movement, Hybridization, and Horizontal Transfer
Given the fact that neither dengue nor mosquitoes respect political boundaries poses important logistical considerations for the use of Wolbachia-infected mosquito releases or any other
___________________
6See http://www.oxitec.com/agriculture/our-products/diamond-back-moth.
7See www.oxitec.com.
8See www.scientificamerican.com/article/fighting-mosquitoes-with-mosquitoes.
form of biocontrol. Given the fact that Wolbachia can spread not only through mosquitoes but also through the fruit fly Drosophila, it is expected that once infected mosquitoes are released, Wolbachia would then become established and would perhaps slowly spread (i.e., an introduction in Vietnam would therefore eventually spread throughout “mainland” Asia). If Wolbachia infected organisms are detected in a neighboring country that did not approve this specific anti-dengue strategy, it could create a legal problem between the involved countries. Recognizing this issue, studies in Australia have monitored neighboring areas for the potential spread of biological agents outside the study area. While, infected larvae were only detected beyond the study cite on just three occasions, the issue of permanent establishment could not be answered with any certainty with the current data and would need further investigation.
Likewise, a key consideration for gene drive development is the possibility of horizontal transfer (for example, the transfer of a gene drive construct to a predator or humans), which could lead to unpredictable non-target effects and unintentional spread of the gene drive construct in non-target organisms. Similar concerns were raised during the development of Wolbachia-based biocontrol techniques (Popovici et al., 2010); as a result, the example offers insights that could be useful for consideration of gene drives.
Early in the Wolbachia biocontrol research process and well before release, investigators engaged the community to identify major questions that needed to be addressed. The process resulted in discussions in three major areas:
- Could Wolbachia affect/be transferred to humans via the insect saliva during blood-feeding? In order to address this, phase 1 studies were performed to detect the presence of Wolbachia in the saliva of the Aedes aegypti mosquito (Moreira et al., 2009b). DNA amplification of Wolbachia wsp genes in the mosquitos salivary glands confirmed the presence of Wolbachia in the glands but the bacteria was absent in the mosquito saliva.
- Could Wolbachia be transferred to another similar mosquito species? Whether Wolbachia could be transferred to other organisms or become established in the soil was addressed using both experimental testing and indirect evidence. The former included the attempt to transfer Wolbachia in new species such as from flies into mosquitoes. The results indicated that the horizontal transfer of Wolbachia between these species was difficult and therefore considered negligible. The latter was based on the fact that since in Australia Wolbachia has been present in Aedes Notoscriptus it could have possibly been transferred to Aedes Aegypti. However, this transfer has never occurred.
- Could Wolbachia be transferred into the environment? A number of studies were conducted to evaluate if Wolbachia could be horizontal transferred into the surrounding environments where mosquitoes would be released. Predation experiments using spiders were performed in the laboratory (phase 1). To verify that Wolbachia did not disseminate in the environment a semi-field fully enclosed outdoor greenhouse designed and constructed specifically for the project was used (i.e., phase 210). Thousands of Wolbachia-infected mosquitoes were introduced with samples of plants, soil, earthworms and millipedes (to fully represent ecosystems in which Wolbachia could have propagated). These samples were then collected from inside the enclosure and tested by PCR for the presence of the specific IS5 Wolbachia genes but none were detected, indicating that no transfer of Wolbachia to other species had occurred. Additional studies of horizontal transfer by other investigators also supported this conclusion (Hurst et al., 2012).
CONCLUSIONS AND RECOMMENDATIONS
Although the potential for gene drives to address and solve problems associated with vector-borne diseases, invasive pests, and agriculture is truly exciting, before field testing or en-
___________________
vironmental release of gene-drive modified organisms, it is crucial to establish a rich understanding of the target organism, its relationship with its environment, and potential unintended consequences, such as off-target and non-target effects.
A phased testing pathway, such as the one developed by the World Health Organization for testing genetically modified mosquitoes, can facilitate a precautionary, step-by-step approach to research on gene drives. Each step in such a pathway promotes careful study and evaluation, includes a series of checkpoints to determine whether and when research should move to the next phase before proceeding to the next step, and provides vital data and knowledge that can be used to inform and enhance the effectiveness of other phases. A phased testing framework to guide step-by-step evaluations, of genetically modified mosquitoes, can be adapted for laboratory and field research on gene-drive modified organisms.
Recommendation 5-1: Scientists conducting research on gene drives should follow a phased testing pathway, a step-by-step framework that begins with developing a research plan and continues through, if applicable, monitoring gene-drive modified organisms in the environment. Each phase in such a pathway should include pre-defined “go/no-go” decisions for determining whether to transition to the next phase based on evidence regarding harms and benefits, efficacy, and safety.
The goal of a gene drive is the rapid spread of genetic information throughout a population. This makes it especially important to minimize potential unintended consequences. Containing or mitigating unintended effects may require a combination of physical containment and biological confinement strategies. When developing biological confinement strategies, consideration will need to be given to their benefits, costs, and weaknesses or potential unintended consequences. For example, adding a visible marker to gene drive-modified organisms in some cases could have negative consequences for the organism, which will need to be weighed against the benefits of this strategy. It is particularly imperative to use caution when considering the development of a “reversal drive”—a gene drive designed to mitigate the unintended consequences of another gene drive—as it may be impossible to effectively employ this strategy without off-target effects or to fully redress ecological and environmental effects from the original gene drive.
Recommendation 5-2: Whenever possible researchers should use available datasets and models to develop and evaluate strategies to minimize the potential for harmful off-target and non-target effects throughout the phased testing pathway.
Recommendation 5-3: Whenever possible, researchers should use a split gene drive in laboratory studies to avoid issues associated with a failure of containment.
Recommendation 5-4: Whenever possible, researchers should include a gene drive that spreads a visible marker to distinguish modified organisms and facilitate research and monitoring.
Recommendation 5-5: Researchers, regulators, and other decision-makers should not rely upon a “reversal” gene drive as the sole strategy for mitigating the effects of another gene drive.
After release into the environment, a gene drive knows no political boundaries. It is desirable to expand the intellectual capital and research capacity of relevant institutions around the world to facilitate appropriate knowledge exchange and research collaborations pertaining to gene drives. In particular, this includes building long-term relationships with scientists in low- and middle-income countries where field research on gene-drive modified organisms is most likely to occur.
REFERENCES
Adelman, Z, ed. 2015a. Genetic Control of Malaria and Dengue. Boston: Academic Press.
Adelman, Z. 2015b. Gene Drives in Mosquitoes: Disease Vector Control.Webinar, October 15, 2015. Available: http://nas-sites.org/gene-drives/2015/10/02/webinar-gene-drive-research-in-different-organisms/ [accessed April 22, 2016].
Ahantarig, A., N. Chauvatcharin, T. Ruang-areerate, V. Baimai, and P. Kittayapong. 2011. Infection incidence and relative density of the bacteriophage WO-B in Aedes albopictus mosquitoes from fields in Thailand. Curr. Microbiol. 62(3):816-820.
Akbari, O.S., H.J. Bellen, E. Bier, S.L. Bullock, A. Burt, G.M. Church, K.R. Cook, P. Duchek, O.R. Edwards, K.M. Esvelt, V.M. Gantz, K.G. Golic, S.J. Gratz, M.M. Harrison, K.R. Hayes, A.A. James, T.C. Kaufman, J. Knoblich, H.S. Malik, K.A. Matthews, K.M. O’Connor-Giles, A.L. Parks, N. Perrimon, E. Port, S. Russell, R. Ueda, and J. Wildonger. 2015. BIOSAFETY. Safeguarding gene drive experiments in the laboratory. Science 349(6251):927-929.
Anders, C., O. Niewoehner, A. Duerst, and M. Jinek. 2014. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513(7519):569-573.
APHIS (Animal and Plant Health Inspection Service). 2002. Containment Guidelines for Nonindigenous, Phytophagous Arthropods and Their Parasitoids and Predators. US Department of Agriculture, Animal and Plant Health Inspection Service [online]. Available: https://www.aphis.usda.gov/plant_health/permits/downloads/arthropod_biocontrol_containment_guidelines.pdf [accessed April 25, 2016].
Basu, S., A. Aryan, J.M. Overcash, G.H. Samuel, M.A. Anderson, T.J. Dahlem, K.M. Myles, and Z.N. Adelman. 2015. Silencing of end-joining repair for efficient site-specific gene insertion after TALEN/CRISPR mutagenesis in Aedes aegypti. Proc. Natl. Acad. Sci. 112(13):4038-4043.
Beech, C.J., S.S. Vasan, M.M. Quinlan, M.L. Capurro, L. Alphey, V. Bayard, M. Bouaré, P. Kittayapong, J.V. Lavery. L.H. Lim, M.T. Marrelli, M.C. McLeod, J. Nagaraju, K. Ombongi, R.Y. Othman, V. Pillai, J.M. Ramsey, R. Reuben, R.L. Rose, B.K. Tyagi, and J. Mumford. 2009a. Deployment of innovative genetic vector control strategies: Progress on regulatory and biosafety aspects, capacity building and development of best-practice guidance. Asia Pac. J. Mol. Biol. Biotechnol. 17(3):75-85.
Beech, C.J., J. Nagaraju, S.S. Vasan, R.I. Rose, R.Y. Othman V.Pillai, and T.S. Saraswathy. 2009b. Risk analysis of a hypothetical open field release of a self-limiting transgenic Aedes aegypti mosquito strain to combat dengue. AsPac. J. Mol. Biol. Biotechnol. 17(3):99-111.
Benedict, M., P. D’Abbs, S. Dobson, M. Gottlieb, L. Harrington, S. Higgs, A. James, S. James, B. Knols, J. Lavery, S. O’Neill, T. Scott, W. Takken, and Y. Toure. 2008. Guidance for contained field trials of vector mosquitoes engineered to contain a gene drive system: Recommendations of a scientific working group. Vector Borne Zoonotic Dis. 8(2):127-166.
Bian, G.W., Y. Xu, P. Lu, Y. Xie, and Z.Y. Xi. 2010. The endosymbiotic bacterium Wolbachia induces resistance to dengue virus in Aedes aegypti. PLoS Pathog. 6(4):e1000833.
Bono, J.M., E.C. Olesnicky, and L.M. Matzkin. 2015. Connecting genotypes, phenotypes and fitness: Harnessing the power of CRISPR/Cas9 genome editing. Mol. Ecol. 24(15):3810-3822.
Bourtzis, K., and H.R. Braig. 1999. The many faces of Wolbachia. Pp. 199-219 in Rickettsiae and Rickettsia Diseases at the Turn of the Third Millennium, D. Raoult, and P. Brouqui, eds. Amsterdam: Elsevier.
Brown, D.M., L.S. Alphey, A. McKemey, C. Beech, and A.A. James. 2014. Criteria for identifying and evaluating candidate sites for open-field trials of genetically engineered mosquitoes. Vector Borne Zoonotic Dis.14(4):291-299.
Bull, J.J., and M. Turelli. 2013. Wolbachia versus dengue: Evolutionary forecasts. Evol. Med. Public Health (1):197-207.
Bundy, A., and L.P. Fanning. 2005. Can Atlantic cod (Gadus morhua) recover? Exploring trophic explanations for the non-recovery of the cod stock on the eastern Scotian Shelf, Canada. Can. J. Fish. Aquat. Sci. 62(7):1474-1489.
Chan, S., P.J. Donovan, T. Douglas, C. Gyngell, J. Harris, R. Lovell-Badge, D.J. Mathews, and A. Regenberg. 2015. Genome editing technologies and human germline genetic modification: The Hinxton group consensus statement. Am. J. Bioeth. 15(12):42-47.
Chippindale, A.K., J.R. Gibson, and W.R. Rice. 2001. Negative genetic correlation for adult fitness between sexes reveals ontogenetic conflict in Drosophila. Proc. Natl. Acad. Sci. 98(4):1671-1675.
Crain, P.R., P.H. Crowley, and S.L. Dobson. 2013. Wolbachia re-replacement without incompatibility: Potential for intended and unintended consequences. J. Med. Entomol. 50(5):1152-1158.
DiCarlo, J.E., A. Chavez, S.L. Dietz, K.M. Esvelt, and G.M. Church. 2015. Safeguarding CRISPR-Cas9 gene drives in yeast. Nat. Biotechnol. 33:1250-1257.
Dobson, S.L., C.W. Fox, and F.M. Jiggins. 2002. The effect of Wolbachia-induced cytoplasmic incompatibility on host population size in natural and manipulated systems. Proc. Biol. Sci. 269(1490):437-445.
Dodd, A.P. 1940. The biological campaign against prickly pear. Commonwealth Prickly Pear Board, Brisbane, 1-177.
Doench, J.G., N. Fusi, M. Sullender, M. Hegde, E.W. Vaimberg, K.F. Donovan, I. Smith, Z. Tothova, C. Wilen, R. Orchard, H.W. Virgin, J. Listgarten, and D.E. Root. 2016. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34:184-191.
Dutra, H.L., L.M. Dos Santos, E.P. Caragata, J.B. Silva, D.A. Villela, R. Maciel-de-Freitas, L.A. Moreira. 2015. From lab to field: The influence of urban landscapes on the invasive potential of Wolbachia in Brazilian Aedes aegypti mosquitoes. PLoS Negl. Trop. Dis. 9(4):e0003689.
Edwards PB, Pavri C. 2007. Pastures-Dung beetles. In PT Bailey, Pests of field crops and pastures: identification and control, CSIRO Publishing, Collingwood, 471-484.
El Zahabi-Bekdash, L., and J.V. Lavery. 2010. Achieving precaution through effective community engagement in research with genetically modified mosquitoes. AsPac. J. Mol. Biol. Biotechnol. 18(2):247-250.
Endersby, N.M., and A. A. Hoffmann. 2013. Effect of Wolbachia on insecticide susceptibility in lines of Aedes aegypti. Bull. Entomol. Res. 103(3):269-277.
Estes J.A., J.Terborgh, J.S. Brashares, M.E. Power, J. Berger, W.J. Bond, S.R. Carpenter, T.E. Essington, R.D. Holt, J.B.C. Jackson, R.J. Marquis, L. Oksanen, T. Oksanen, R.T. Paine, E.K. Pikitch, W.J. Ripple, S.A. Sandin, M. Scheffer, T.W. Schoener, J.B. Shurin, A.R.E. Sinclair, M.E. Soule, R. Virtanen, D.A. Wardle. 2011. Trophic downgrading of planet Earth. Science. (333):301-306.
Esvelt, K.M., A.L. Smidler, F. Catteruccia, and G.M. Church. 2014. Concerning RNA-guided gene drives for the alteration of wild populations. eLife 3:e03401.
Facchinelli, L., L. Valerio, J.G. Bond, M.R. Wise de Valdez, L.C. Harrington, J.M. Ramsey, M. Casas-Martinez, and T.W. Scott. 2011. Development of a semi-field system for contained field trials with Aedes aegypti in southern Mexico. Am. J. Trop. Med. Hyg. 85(2):248-256.
FDA (US Food and Drug Administration). 2007. Target Product Profile—A Strategic Development Process Tool. Guidance for Industry and Review Staff, March 2007 [online]. Available: http://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/ucm080593.pdf [accessed April 28, 2016].
Ferguson, H.M., K.R. Ng’habi, T. Walder, D. Kadungula, S.J. Moore, I. Lyimo, T.L. Russell, H. Urassa, H. Mshinda, G.F. Killeen, and B.G. Knols. 2008. Establishment of a large semi-field system for experimental study of African malaria vector ecology and control in Tanzania. Malaria J. 7:158.
Fenner, F. 1983. Biological control, as exemplified by smallpox eradication and myxomatosis. Proc. R. Soc. Lond. B. Biol. Sci. 218(1212):259-285.
Frock, R.L., J. Hu, R.M. Meyers, Y.J. Ho, E. Kii, and F.W. Alt. 2015. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat. Biotechnol. 33(2):179-186.
Gantz, V.M., and E. Bier. 2015. Genome editing. The mutagenic chain reaction: A method for converting heterozygous to homozygous mutations. Science 348(6233):442-444.
Gantz, V.M., N. Jasinskiene, O. Tatarenkova, A. Fazekas, V.M. Macias, E. Bier, and A.A. James. 2015. Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proc. Natl. Acad. Sci. 112(49):E6736-E6743.
Ghelelovitch, S. 1952. Genetic determinism of sterility in the cross-breeding of various strains of Culex autogenicus Roubaud [in French]. C R Hebd. Seances Acad. Sci. 234(24):2386-2388.
Gould, F., Y. Huang, M. Legros, and A.L. Lloyd. 2008. A killer-rescue system for self-limiting gene drive of anti-pathogen constructs. Proc. Biol. Sci. 275(1653):2823-2829.
Hagler, J.R., and C.G. Jackson. 2001. Methods for marking insects: Current techniques and future prospects. Annu. Rev. Entomol. 46(1):511-543.
Hammond, A., R. Galizi, K. Kyrou, A. Simoni, C. Siniscalchi, D. Katsanos, M. Gribble, D. Baker, E. Marois, S. Russell, A. Burt, N. Windbichler, A. Crisanti, and T. Nolan. 2016. A CRISPR-Cas9 gene drive system targeting female reproduction in the malaria mosquito vector Anopheles gambiae. Nat. Biotechnol. 34(1):78-83.
Handler, A.M., R.A. Harrell. 2001. Polyubiquitin-regulated DsRed marker for transgenic insects. Biotechniques. 31(4):820, 824-828.
Hanh, T.T., P.S. Hill, B.H. Kay, and T.M Quy. 2009. Development of a framework for evaluating the sustainability of community-based dengue control projects. Am. J. Trop. Med. Hyg. 80(2):312-318.
Hartl, D. 1970. Analysis of a general population genetic model of meiotic drive. Evolution 24(3):538-545.
Harvey-Samuel, T., T. Ant, H. Gong, N.I. Morrison, L. Alphey. 2014. Population-level effects of fitness costs associated with repressible female-lethal transgene insertions in two pest insects. Evol. Appl. 7(5):597-606.
Hurst, T.P., G. Pittman, S.L. O’Neill, P.A. Ryan, H.L. Nguyen, and B.H. Kay. 2012. Impacts of Wolbachia infection on predator prey relationships: evaluating survival and horizontal transfer between wMelPop infected Aedes aegypti and its predators. J. Med. Entomol. 49(3): 624-630.
Isaacs, A.T., Jasinskiene, N., Tretiakov, M., Thiery, I., Zettor, A., Bourgouin, C. and James, A.A. (2012) Transgenic Anopheles stephensi co-expressing single-chain antibodies resist Plasmodium falciparum development. Proc. Natl. Acad. Sci. USA, 109, E1922-E1930. PMID:22689959. PNAS PLUS, 109, 11070-11071.
Jaenike, J. 2001. Sex chromosome meiotic drive. Annu. Rev. Ecol. Syst. 32:25-49.
Jeffery, J.A., N. Thi Yen, V.S. Nam, T. le Nghia, A.A. Hoffman, B.H. Kay, and P.A. Ryan. 2009. Characterizing the Aedes aegypti population in a Vietnamese village in preparation for a Wolbachia-based mosquito control strategy to eliminate dengue. PLoS Negl. Trop. Dis. 3(11):e552.
Kelland, K. 2015. Genetically modified diamondback moths offers pest control hope. Reuters, July 15, 2015 [online]. Available: http://www.reuters.com/article/us-science-moths-gmo-idUSKCN0PQ00120150716 [accessed April 22, 2016].
Kitano, H. 2002. Computational systems biology. Nature 420(6912):206-210.
Kleinstiver, B.P., M.S. Prew, S.Q. Tsai, N.T. Nguyen, V.V. Topkar, Z. Zheng, and J.K. Joung. 2015. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat. Biotechnol. 33(12):1293-1298.
Knipling, E.F. 1955. Possibilities of insect control or eradication through the use of sexually sterile males. J. Econ. Entomol., 48: 902-904.
Koo, T., J. Lee, and J.S. Kim. 2015. Measuring and reducing off-target activities of programmable nucleases including CRISPR-Cas9. Mol. Cells 38(6):475-481.
Lavery, J.V., L.C. Harrington, and T.W. Scott. 2008. Ethical, social, and cultural considerations for site selection for research with genetically modified mosquitoes. Am. J. Trop. Med. Hyg. 79(3):312-318.
Marshall, J.M., and B.A. Hay. 2012. Confinement of gene drive systems to local populations: A comparative analysis. J. Theor. Biol. 294:153-171.
Martins, S., N. Naish, A.S. Walker, N.I. Morrison, S. Scaife, G. Fu, T. Dafa’alla, and L. Alphey. 2012. Germline transformation of the diamondback moth, Plutella xylostella L., using the piggyback transposable element. Insect Mol. Biol. 21(4):414-421.
Mathews, D.J., S. Chan, P.J. Donovan, T. Douglas, C. Gyngell, J. Harris, A. Regenberg, and R. Lovell-Badge. 2015. CRISPR: A path through the thicket. Nature 527(7577):159-161.
McMeniman, C.J., R.V. Lane, B.N. Cass, A.W. Fong, M. Sidhu, Y.F. Wang, and S.L. O’Neill. 2009. Stable introduction of a life-shortening Wolbachia infection into the mosquito Aedes aegypti. Science 323(5910):141-144.
Moreira, L.A., I. Iturbe-Ormaetxe, J.A. Jeffery, G.J. Lu, A.T. Pyke, L.M. Hedges, B.C. Rocha, S. Hall-Mendelin, A. Day, M. Riegler, L.E. Hugo, K.N. Johnson, B.H. Kay, E.A. McGraw, A.F. van den Hurk, P.A. Ryan, and S.L. O’Neill. 2009a. A Wolbachia symbiont in Aedes aegypti limits infection with dengue, Chikungunya, and Plasmodium. Cell 139(7):1268-12678.
Moreira, L.A., E. Saig, A.P. Turley, J.M. Ribeiro, S.L. O’Neill, and E.A. McGraw. 2009b. Human probing behavior of Aedes aegypti when infected with a life-shortening strain of Wolbachia. PLoS Negl. Trop. Dis. 3:e568.
Mumby, P.J., and R.S. Steneck. 2008. Coral reef management and conservation in light of rapidly evolving ecological paradigms. Trends Ecol. Evol. 23(10):555-563.
Mumford, J., M.M. Quinlan, C.J. Beech, L. Alphey, V. Bayard, M.L. Capurro, P. Kittayapong, J.D. Knight, M.T. Marrelli, K. Ombongi, J.M. Ramsey, and R. Reuben. 2009. MosqGuide: A project to develop best practice guidance for the deployment of innovative genetic vector control strategies for malaria and dengue. AsPac. J. Mol. Biol. Biotechnol. 17(3):93-95.
National Biosafety Technical Commission. 2014. Technical Opinion on Commercial Release of Genetically Modified Microorganism No. 3964/2014. Ministry of Science, Technology and Inovation, National Biosafety Technical Commission, Brazil.
Nishimasu, H., F.A. Ran, P.D. Hsu, S. Konermann, S.I. Shehata, N. Dohmae, R. Ishitani, F. Zhang, and O. Nureki. 2014. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156(5):935-949.
O’Connor, L., C. Plichart, A.C. Sang, C.L. Brelsfoard, H.C. Bossin, and S.L. Dobson. 2012. Open release of male mosquitoes infected with a Wolbachia biopesticide: Field performance and infection containment. PLoS Negl. Trop. Dis. 6(11): e1797.
Otto, S.P., and T. Day. 2007. A Biologist’s Guide to Mathematical Modeling in Ecology and Evolution. Princeton, NJ: Princeton University Press.
Oye, K.A., K. Esvelt, E. Appleton, F. Catteruccia, G. Church, T. Kuiken, S.B. Lightfoot, J. McNamara, A. Smidler, and J.P. Collins. 2014. Biotechnology. Regulating gene drives. Science 345(6197):626-628.
Phuc, H.K., M.H. Andreasen, R.S. Burton, C. Vass, M.J. Epton, G. Pape, G. Fu, K.C. Condon, S. Scaife, C.A. Donnelly, P.G. Coleman, H. White-Cooper, and L. Alphey. 2007. Late-acting dominant lethal genetic systems and mosquito control. BMC Biol. 5:11.
Popovici, J., L.A. Moreira, A. Poinsignon, I. Iturbe-Ormaetxe, D. McNaughton, and S.L. O’Neill. 2010. Assessing key safety concerns of a Wolbachia-based strategy to control dengue transmission by Aedes mosquitoes. Mem. Inst. Oswaldo Cruz 105(8):957-964.
Port, F., N. Muschalik, and S.L. Bullock. 2015. Systematic evaluation of Drosophila CRISPR tools reveals safe and robust alternatives to autonomous gene drives in basic research. G3(Bethesda) 5(7):1493-1502.
Ran, F.A., P.D. Hsu, C.Y. Lin, J.S. Gootenberg, S. Konermann, A.E. Trevino, D.A. Scott, A. Inoue, S. Matoba, Y. Zhang, and F. Zhang. 2013. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154(6):1380-1389.
Room, P.M., K.L.S. Harley, I.W. Forno, D.P.A. Sands. 1981. Successful biological control of the floating weed salvinia. Nature 294:78-80.
Rose, M.R., L.N. Vu, S.U. Park, and L.R. Graves, Jr. 1992. Selection on stress resistance increases longevity in Drosophila melanogaster. Exp. Gerontol. 27(2):241-250.
Rossi, A., Z. Kontarakis, C. Gerri, H. Nolte, S. Hölper, M. Krüger, and D.Y. Stainier. 2015. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 524(7564):230-233.
Sander, J.D., and J.K. Joung. 2014. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32(4):347-355.
Saunders, G., B. Cooke, K. McColl, R. Shine, and T. Peacock. 2010. Modern approaches for the biological control of vertebrate pests: An Australian perspective. Biol. Control 52(3):288-295.
Shapiro, J., and D.I. Wright. 1984. Lake restoration by biomanipulations, Round Lake, Minnesota- the first two years. In Freshwater Biol. 14:371-383.
Singer, P.A., A.D. Taylor, A.S. Daar, R.E.G. Upshur, J.A. Singh, and J.V. Lavery. 2007. Grand challenges in global health: The ethical, social and cultural program. PLoS Med. 4(9):e265.
Sinkins, S.P., and F. Gould. 2006. Gene drive systems for insect disease vectors. Nat. Rev. Genet. 7(6):427-435.
Sinkins, S.P., C.F. Curtis, and S.L. O’Neill. 1997. The potential application of inherited symbiont systems to pest control. Pp. 155-175 in Influential Passengers, S.L. O’Neill, A. Hoffman, and J. Werren, eds. Oxford: Oxford University Press.
Slaymaker, I.M., L. Gao, B. Zetsche, D.A. Scott, W.X. Yan, and F. Zhang. 2016. Rationally engineered Cas9 nucleases with improved specificity. Science 351(6268):84-88.
Stiling, P., and T. Cornelissen. 2006. What makes a successful biocontrol agent? A meta-analysis of biological control agent performance. Biol. Control 34(3):236-246.
Stouthamer, R., J.A. Breeuwer, and G.D. Hurst. 1999. Wolbachia pipientis: Microbial manipulator of arthropod reproduction. Annu. Rev. Microbiol. 53:71-102.
Suwannachote, N., J.P. Grieco, N.L. Achee, W. Suwokerd, S. Wongtong, and T. Chareonviriyaphap. 2009. Effects of environmental conditions on the movement patterns of Aedes aegypti (Diptera: Culicidae) into and out of experimental huts in Thailand. J. Vector Ecol. 34(2):267-275.
Thomas, D.D., C.A. Donnelly, R.J. Wood, and L.S. Alphey. 2000. Insect population control using a dominant, repressible, lethal genetic system. Science 287(5462):2474-2476.
Tsai, S.Q., Z. Zheng, N.T. Nguyen, M. Liebers, V.V. Topkar, V. Thapar, N. Wyvekens, C. Khayter, A.J. Iafrate, L.P. Le, M.J. Aryee, and J.K. Joung. 2015. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33(2):187-197.
van Driesche, R.D., and T.S. Bellows. 1996. Biological Control. Boston: Kluwer.
Vreysen, M. J. B., A.S. Robinson, and J. Hendrichs, eds. 2007. Area-wide Control of Insect Pests, From Research to Field Implementation. Dordrecht, The Netherlands: Springer.
Wackers, F.L., P.C.J. van Rijn, K. Winkler, D. Olson. 2007. Flower power? Potential benefits and pitfalls of using (flowering) vegetation for conservation biological control. Aspects of Applied Biology 81:135-140.
Waltz, E. 2015. Oxitec trials GM sterile moth to combat agricultural infestations. Nat. Biotechnol. 33(8):792-793.
Wang, L., Y. Shao, Y. Guan, L. Li, L. Wu, F. Chen, M. Liu, H. Chen, Y. Ma, X. Ma, M. Liu, and D. Li. 2015. Large genomic fragment deletion and functional gene cassette knock-in via Cas9 protein mediated genome editing in one-cell rodent embryos. Sci. Rep. 5:17517.
Werren, J.H., and S. O’Neil. 1997. The evolution of heritable symbionts. Pp. 1-41 in Influential Passengers: Inherited Microorganisms and Arthropod Reproduction, S. O’Neil, A.A. Hoffmann, and J.H. Werren, eds. New York: Oxford University Press.
Werren, J.H., W. Zhang, and L.R. Guo. 1995. Evolution and phylogeny of Wolbachia: Reproductive parasites of arthropods. Proc. Biol. Sci. 261(1360):55-63.
WHO (World Health Organization). 2010. Progress and Prospects for the Use of Genetically Modified Mosquitoes to Inhibit Disease Transmission [online]. Available at http://apps.who.int/iris/bitstream/10665/44297/1/9789241599238_eng.pdf [accessed April 25, 2016].
WHO. 2014. The Guidance Framework for Testing Genetically Modified Mosquitoes. World Health Organization, Programme for Research and Training in Tropical Diseases [online]. Available at http://apps.who.int/iris/bitstream/10665/127889/1/9789241507486_eng.pdf?ua=1 [accessed April 19, 2016].
Wu, B., L. Luo, and X.J. Gao. 2016. Cas9-triggered chain ablation of cas9 as a gene drive brake. Nat. Biotechnol. 34(2):137-138.
Zetsche, B., J.S. Gootenberg, O.O. Abudayyeh, I.M. Slaymaker, K.S. Makarova, P. Essletzbichler, S.E. Volz, J. Joung, J. van der Oost, A. Regev, E.V. Koonin, and F. Zhang. 2015. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163(3):759-771.


























