4
Current and Future Research Opportunities
TRANSMISSION
While the ecology of Ebola virus—including its natural host—remains largely a mystery, more is known about its close relative, Marburg virus. Jonathan Towner of the U.S. Centers for Disease Control and Prevention (CDC) discussed how Marburg virus has been linked to bats, how that knowledge is informing further studies of its transmission, and how lessons learned in that process could guide research on Ebola virus.
Bats as Reservoir Hosts for Filoviruses
Learning where Ebola and Marburg viruses reside in nature and how they are transmitted from these natural reservoirs to humans will lead to better understanding of how outbreaks start, Towner explained. A reservoir host species’ natural history must support viral persistence at the population level, he observed, as evidenced by the following characteristics: consistent infection in host populations; maintenance of viral circulation through adequate viral loads and rates of viral shedding; and host populations that are, overall, clinically healthy.
Results of two studies (Amman et al., 2012; Towner et al., 2009) conducted during an outbreak in miners in Kitaka, Uganda, and the occurrence of unrelated cases in foreign tourists in nearby Python Cave suggested that Marburg virus is horizontally transmitted among bat populations, Towner said. Infection rates generally increased as bats aged, with the exception of a spike among older juvenile bats—a phenomenon that has been noticed in other studies of viral host populations, he noted. Further analysis suggested that periods of increased active infection in juvenile bats coincided with incidences of spillover to humans (Amman
et al., 2012). He speculated that older animals inhabit preferred sites deep in the cave, but younger bats, being lower in the hierarchy, stay closer to the outside, where they are likelier to encounter humans or other primates.
Based on experimental infectious studies (Amman et al., 2015; Paweska et al., 2012) Towner stated, “We suspect that bat-to-bat transmission [of Marburg virus] is probably through biting. For bat-to-human transmission, we suspect it’s probably through infected feces, or urine containing infectious virus.” This scenario is consistent with the experience of tourists infected with the virus at Python Cave who did not actually enter the caverns, and yet were probably exposed to bat excrement.
The Experience with Ebola
Nonhuman primates are known to be susceptible to Ebola and have been confirmed as sources of human infections. The role of bats in Ebola virus transmission—although suspected since the 1976 Sudan outbreak—has yet to be confirmed; the relationships have been hypothesized from Ebola’s epidemiological features, which bear a strong resemblance to Marburg. Unfortunately, Towner pointed out, most ecological investigations of Ebola have been initiated after outbreaks were already resolved, and any knowledge of the circumstances under which the index case may have come into contact with the virus is difficult to obtain, because that person is dead and/or unknown. Nearly all of the recent Ebola outbreaks in Gabon and the Democratic Republic of Congo (DRC) have been traced to direct contact with the infected meat of nonhuman primates (which likely died of their infection), but it remains to be determined how those animals became infected with the virus.
Ecological niche models of Ebola (and other filoviruses) pursued by several research groups have reached similar general conclusions about where viral reservoirs are likely to exist, based on outbreak locations and the distribution of bat species, vegetation, and rainfall, Towner stated. “While this is interesting and informative in terms of identifying the real hosts, it is not quite granular enough,” he observed.
Ecological investigations of local bats, birds, and small vertebrates collected in Gabon and the DRC during repeated Ebola virus outbreaks in both humans and nonhuman primates found three different species of fruit bats to be positive for both immunoglobulin G (IgG) and polymerase chain reaction (PCR); however, no virus was isolated from these animals (Leroy et al., 2005). Similar studies of bats in Ghana found several IgG-positive species—including some of the same species identified in Gabon and the DRC—but none were PCR positive (Hayman et al., 2012). All three of these bat species are present in West Africa, as well, he noted.
As the West African epidemic unfolded, researchers pursued an extensive ecological investigation of the index case, a 2-year-old boy reported to have
played in a hollow tree found to be infested with the insectivorous bat Mops condylurus (Marí Saéz et al., 2014). No PCR-positive bats were discovered, nor did they find a linkage to a large mammal die-off as had occurred in several previous Ebola outbreaks in Republic of Congo, DRC, and Gabon.
Overall, the inconclusiveness of the antibody evidence linking Ebola virus to M. condylurus suggests these bats may serve as secondary hosts, Towner said. Moreover, he added, the species’ territory does not coincide with the locations of most known Ebola outbreaks. And since serological cross-reactivity has been demonstrated among Ebola viruses, a serologically positive bat found in an outbreak area is not necessarily infected with the outbreak-causing strain.
Each Ebola virus type probably resides in a different reservoir host species, Towner hypothesized. “For Marburg, we think that Rousettus [aegyptiacus]1 is it, while I hesitate to say it quite that definitively,” he explained. “There may be other factors involved, but for Ebola-Zaire it’s probably another species; for Ebola-Sudan it’s going to be a different species—same with Ebola-Côte D’Ivoire and the others. You know, those viruses perpetuate in those species in the long term.”
The restricted cave biome in which Marburg virus was detected in bats helped narrow the search for its reservoir, Towner observed—an advantage researchers seeking the Ebola virus reservoir do not have. That is especially unfortunate if—as for Marburg virus—the number of active Ebola virus infections in reservoir host populations is low, so hundreds of animals of a single species will need to be tested to find them. “When you take that into account with the fact that there are over 250 bat species in Africa, you can do the math in terms of the number of animals that you then have to test,” he said. As with Marburg virus, active infection may be seasonal, coincident with birth pulses, which are biannual in most African fruit bats,2 he added.
If Ebola virus behaves similarly to Marburg virus, it will spill over into secondary bat hosts that inhabit the same environments, Towner advised. Transmission is likely to be horizontal, and infection will not persist as it does in some rodent viral reservoir species. Viral shedding could occur through the mouth, permitting transmission to other animals by biting—and also by feeding from the same piece of fruit. Testing for oral shedding can be done by swabbing, which does minimal harm to bats, many of which are endangered species, he pointed out—a fact that also must be borne in mind if they are identified as a reservoir for Ebola, and thereby threatened with eradication.
___________________
1Rousettus aegyptiacus is a species of fruit bat found throughout Africa and the Middle East.
2 A mathematical model of Marburg virus persistence in R. aegyptiacus populations, based on the results of the previously described field and experimental studies, determined that a bat community must comprise at least 20,000 in order to sustain viral infection, given a 21-day incubation period and a biannual birth pulse (Hayman, 2015).
Genomic Surveillance of Ebola Transmission
Recent advances in high-throughput genetic sequencing allow researchers to track viruses as they spread through host populations, evolving as they go. Stephen Gire of Broad Institute of the Massachusetts Institute of Technology and Harvard University described the application of these methods to reveal how and when the Ebola virus spilled over to humans in West Africa, how many such events occurred, and how the virus evolved as the epidemic unfolded (Gire et al., 2014).
The medical team at Kenema Government Hospital in Sierra Leone, where that country’s first laboratory-confirmed Ebola case was diagnosed, acted quickly to identify more cases through contact tracing, all of whom had attended the funeral of a traditional healer who had treated Guinean Ebola patients, Gire reported. Samples from 12 of the Sierra Leonean patients were sent immediately to Gire’s team in Boston, who sequenced and analyzed them within 10 days, using a strategy they had created to track Lassa fever transmission. As the epidemic continued, the researchers continued to sequence patient samples from that hospital, publishing their initial findings (on 114 viral isolates from 78 patients) in August 2014. It was important to make this information available to the scientific community as soon as possible, Gire said, in order to quickly bring as much diverse expertise to its analysis, and thereby, to benefit efforts to quell the epidemic.
The key questions of where and how the epidemic began were quickly answered, he noted. Based on viral phylogeny, Gire and coworkers traced the origin of the West African strain to the Ebola-Zaire strain that caused the 1976 outbreak. Comparing genetic differences between the West African isolates and the sequences of other Ebola viruses, and taking into account the rate by which the virus typically undergoes sequence change—its substitution rate—they were able to determine that the specific variant of Ebola-Zaire that sparked the West African epidemic had emerged within the past 10 years in Central Africa, he noted. Further, he said, “We were able to show that it was actually one individual introduction from the animal reservoir into the human population that then was sustained in the population by human-to-human contact afterward.”
Tracing Ebola in Sierra Leone
Looking to the immediate past, the researchers identified three lineages (comprising 55 mutations) among the Sierra Leonean isolates that distinguished them from three sequences isolated several weeks earlier from Guinean patients, Gire stated. Two of those lineages were present in samples from the initial funeral event, they discovered: about half of those patients had one version of the virus, and half had the other. Perhaps, they hypothesized, the traditional healer had treated a lot of Guinean patients, and had been infected more than once, with the two different lineages; alternatively, one of the funeral attendees might have been infected with a different lineage than the healer—which seems plausible if,
as is now suspected, Ebola virus was circulating in Sierra Leone before it was detected in the first laboratory-confirmed patient.
Mapping the geographic origins of the three Sierra Leonean lineages, Gire’s group reconstructed transmission chains, allowing them to pinpoint where patients had traveled after the funeral, which aided in identifying their contacts. This information also revealed that some patients with broad social networks served as “superspreaders” of Ebola. Since the publication of their initial findings, Gire and colleagues (2014) have generated an additional 150 genomic sequences3 of Ebola virus isolates from Sierra Leone, which allowed them to continue to track viral transmission and evolution as the epidemic progressed, Gire reported.
Over the course of the epidemic, the original lineage, thought to have been imported from Guinea, died off, Gire said. Lineage 2, thought to have arisen in Sierra Leone in April 2014, gave rise to lineage 3 via a single mutation and lineage 3 quickly became dominant and widely distributed across the country. A fourth lineage then arose from lineage 3 via a single mutation and quickly became dominant, much as 3 did from 2. While lineage 2 died out in Sierra Leone, limited evidence suggests it could have reseeded Ebola in Liberia, he added.
Evolution Without Change
Overall, the researchers found that a higher rate of viral sequence substitution occurred during the West African Ebola outbreak than between previous outbreaks, according to Gire. That does not mean that mutations are accumulating within the virus, he explained; rather, the virus is “exploring” a diverse set of mutations, none of which has produced structural or functional change since 1976 (Olabode et al., 2015). This behavior, called purifying selection, tends to occur when a virus infects a naïve host population, Gire explained, describing it as a “weeding out” of deleterious mutations and a balancing force in selection. The virus is thus not gaining fitness over time, he concluded—and, he predicted, as the outbreak continues, its accumulation of mutations should continue to slow with time. In subsequent discussion, Erica Saphire of The Scripps Research Institute noted that these conclusions were not drawn from functional assay, but by sequence analysis, which is vulnerable to misinterpretation. Gire agreed. “The only way to really figure out absolutely whether there are functional implications for these mutations is to isolate each one of them and functionally follow them up in laboratories and cell culture and animal models to figure out if these actual mutations do have a functional consequence,” he stated. “That work has not been done, and that work does need to be done on these mutations that we see in this variant.”
___________________
3 At the time of this writing, 249 genomic sequences were generated. The 99 full-length and 150 partial-length sequences can be viewed at http://www.ncbi.nlm.nih.gov/bioproject/257197 (accessed November 3, 2016).
Reminding participants that the results he presented represent “a small snapshot” of the entire West African Ebola epidemic, Gire looked forward to assessing variation among Guinean and Liberian viral isolates and gaining knowledge that could reveal regional transmission chains. Additional questions to be addressed include understanding why Sierra Leone lineage 3 became dominant, given that it differed from lineage 2 by a noncoding mutation, and the possible genetic basis for decreased viral titers observed late in the epidemic.
DIAGNOSIS
Many groups and agencies have worked to improve the laboratory diagnosis of Ebola in West Africa and other low-resource settings, noted Heinz Feldmann of the National Institute of Allergy and Infectious Diseases’ Rocky Mountain Laboratories. He presented an overview of current diagnostic approaches, then used his recent experience in an Ebola treatment unit (ETU) in Monrovia, Liberia, to demonstrate challenges and opportunities associated with diagnosing Ebola.
Detecting Ebola
An acute infection, Ebola symptoms commence after an incubation period of variable length, during which the pathogen replicates. These symptoms can vary widely from case to case, thus the controversy as to whether or not the disease should be denoted a hemorrhagic fever. By the time the virus is readily detectable, the patient typically is at the tipping point between death and recovery, Feldmann observed. “Often, particularly with less sensitive assays, you do not see a very strong or any immune response in people that succumb to infection very suddenly and early during disease progression,” he said. Thus, pathogen detection assays, rather than those that target host immune response, are key to diagnosing Ebola in outbreak situations.
Standard Ebola diagnostic tests detect either the nonsegmented, negative-stranded RNA genome of the virus or its protein components, Feldmann explained; both methods were used to diagnose Ebola in West Africa, as was the presence of antibody (IgM and IgG) in survivors of the disease.4 Ebola and other filoviruses are enclosed by an envelope studded with glycoprotein trimers. The majority of the protective immune response is directed against these molecules; however, immune response to the virus’s other six structural proteins is important to its diagnosis, he noted.
___________________
4 Additional methods to detect Ebola virus include particle detection and virus isolation. However, because these techniques require stringent containment and sophisticated equipment, they are not appropriate for immediate outbreak response, according to Feldmann. Similarly, neutralization assays to detect antibodies in survivors, which, unlike enzyme-linked immunosorbent assays for IgG and IgM, are specific to virus species, are not suitable for field use because they are performed on live or surrogate viruses.
The list of commercially produced Ebola diagnostic tests has grown significantly over the course of the epidemic and continues to expand, Feldmann observed; however, the majority of field laboratories still use their own assays, many of which lack positive controls because those require biosafe containment. “I think we have to do much, much more . . . to make sure that the laboratories . . . that are doing diagnostics in certain countries actually evaluate their tests,” he urged—and better yet, to fulfill the World Health Organization’s (WHO’s) demand for rapid, sensitive, safe, and simple Ebola diagnostics. “I think we are on the way to achieving that,” he observed, but as discussed below, some obstacles remain to meeting that goal.
Putting the present in perspective, Feldmann noted that until around 2000, Ebola diagnosis took weeks to months, as samples were shipped to reference centers such as the CDC. The construction of in-country reference centers reduced that to days; with current on-site mobile support, diagnosis takes about 2 to 6 hours, he said. “The future is minutes,” he predicted, but he cautioned that methods currently capable of such speed are not sufficiently sensitive.
NIH Efforts
Michael Kurilla of the National Institute of Allergy and Infectious Disease (NIAID) gave an overview of filovirus-related projects and research under way at the National Institutes of Health (NIH), which has focused on these pathogens as targets for biodefense. (Dr. Kurilla’s contribution to this Proceedings of a Workshop can be found in Appendix B.) After the terrorist attacks of 2001, Kurilla said, Ebola and Marburg viruses became subjects of intense efforts encompassing basic research, diagnostics, vaccines, and therapeutics, as well as product development. In his presentation to the workshop, he described recent advances in each of these areas, which will be discussed in this and following sections.
To fill a shortage of research facilities of the highest biosafety level, where work could be conducted on live filoviruses and other Category A pathogens, the NIH has added three biosafety level-4 (BSL-4) laboratories since the inception of biodefense funding in 2003, Kurilla reported. These laboratories greatly increased capacity for both research and product development, and specifically have expedited work on vaccines and therapeutics, he said. NIAID personnel proficient in BSL-4 safety skills have not only conducted diagnostic work in West Africa but have also provided instruction to local health care workers, according to Kurilla. The U.S. Food and Drug Administration (FDA) granted emergency use authorization to two of the tests they employed: BioFire,5 based on film arrays, and Corgenix,6 an antigen-derived lateral flow test. Both are being pursued as potential point-of-care applications.
___________________
5 See http://www.biofiredx.com (accessed November 3, 2016).
6 See http://www.corgenix.com/news-releases/corgenix-receives-fda-authorization-and-who-listing-for-emergency-use-of-ebola-rapid-diagnostic-test (accessed November 3, 2016).
The View from Monrovia
At the time of the workshop, Feldmann’s team from the NIH and another from the CDC had been staffing the laboratory of a Médecins Sans Frontières (MSF) ETU in Monrovia, Liberia, since August 2014. “We started out in a tent structure, as almost all mobile laboratories nowadays do,” he recalled. MSF can construct such facilities within a day, allowing laboratory operations to begin in less than 48 hours, he said.
Performing diagnostic tests in the tents was challenging, according to Feldmann; nevertheless, despite incomplete physical separation of different stages of the process, there were few reports of contamination. Later the laboratory moved to a fixed structure, which improved workflow and also afforded better protection from rain and heat. A mobile plastic BSL-3 laboratory under negative pressure was also available but was not used because sample loads did not reach critical levels, he said.
The ELWA 3, an MSF Ebola management center, operated a laboratory during the West African epidemic, and employed quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) to diagnose Ebola. This method permitted detection of several different pathogens within about an hour, but it requires sophisticated equipment and trained personnel, Feldmann noted. Reverse transcription loop-mediated isothermal amplification (RT-LAMP), a faster, less-demanding method, is not as well established, but it is moving into field use, he added; however, as currently performed, it is vulnerable to detecting false negatives. An even faster method, isothermal recombinase polymerase amplification, has potential, but it has not been appropriately evaluated for field use.
Extracting viral RNA from patient samples for analysis requires an additional 30 to 60 minutes, but it can be performed easily using kits from several sources, Feldmann noted. The most common are based on chaotropic agents and can be either performed manually or automated. Although some of these processes must be adjusted in order to completely inactivate the samples, this approach has been used for more than a decade in the field and in the laboratories without incident, he said. “If you do the test in the laboratory, you have breakthrough with rest infectivity unless you add . . . ethanol. That completely inactivates the sample. Be careful, and test your procedures,” he advised.
The ELWA 3 team used a two-target approach to diagnose Ebola, performing dual PCR assays for separate genomic regions. “The positive is only a positive if both are consistent and agree on the result,” Feldman said. “Not every mobile lab does that. Some labs operate on a single target system, which, of course, is more prone to potential false negatives if you have sequence variation.” If the results of their PCR assays disagreed, they ran a third assay on a different genomic region as a tie-breaker, and used clinical parameters as well to determine their conclusion, he said. The assays, conducted in tandem by the NIH and the CDC teams, were nearly perfectly concordant, he noted—evidence of their successful collaboration.
Under optimal conditions, patients at ELWA 3 were tested as soon as they entered the facility and if positive, were isolated, Feldman reported. Surviving patients who were asymptomatic for 3 days then required two consecutive negative tests, between one and three days apart, for discharge. However, he noted, that algorithm was frequently compromised in August and September 2014, when the facility became overwhelmed with cases. “You had to make the decision between someone that you may send home who may be a false negative versus someone who is extremely ill with Ebola knocking on your door, and you can’t take that patient into your clinic,” he recalled.
Beginning in October 2014, as the crisis began to abate, the laboratory also began testing for malaria. A frequent coinfection with Ebola, malaria needs to be studied as to its possible effect on survival, Feldmann remarked. The researchers also began performing blood tests for Ebola using bedside instruments; however, he said, antigen-based “dipstick” tests (as noted by Heymann) are better prospects for future analyses, and capable of providing results within 15 minutes. Although such tests are not as sensitive as qRT-PCR, they can potentially be used successfully in an appropriately adjusted diagnostic algorithm, Feldmann suggested. “For an incoming patient, a negative [result from a rapid test] does not exclude an infection,” he emphasized. “For a survivor, a negative test does not exclude infectivity or rest infectivity.” Laboratory-based amplification approaches will still be needed to measure viral load, he added.
While unsuited to the diagnostic demands of the epidemic at its height, serology—which had been employed in previous outbreaks—may still be used to analyze samples to better understand what happened, and be developed as a basis for rapid tests, Feldmann observed. Here too, inactivating the virus is problematic, he noted: heat alone is not sufficient and must be combined with a chemotropic agent.
CONTAINMENT CARE
Health care workers responding to Ebola outbreaks have all too frequently become victims of the disease, and in disproportionate numbers to the general population. In West Africa, as of June 4, 2015, 869 health care workers had become infected over the course of the epidemic, and 507 had died (ECDC, 2015b). This situation has provoked considerable discussion of containment care for Ebola patients, noted Mark Kortepeter from the Uniformed Services University of Health Sciences. In his workshop presentation, he reviewed the science behind the development of infection containment practices, the recent history of containment technologies, and the public health policy issues surrounding efforts to reduce the risk borne by caregivers for Ebola and other emerging infectious diseases.
The Science of Containment Care
Compared with other infectious diseases, Ebola “is pretty hard to catch,” Kortepeter observed; transmission could be said to occur only in the right place at the wrong time, which unfortunately describes the fate of many health care workers.
Hospitals—the “right place”—have historically served as epicenters of Ebola spread through such unsafe practices as the reuse of contaminated needles or syringes on patients, accidental needle sticks of health care workers, exposures during surgical procedures, rubbing eyes and other mucous membranes with soiled gloves, and the splashing of infected water or body fluids, Kortepeter stated. Within households, transmission results primarily from direct physical contact with blood or body fluids of an individual who has been ill, sleeping in the same hut or mat, sharing a needle, sharing a hospital bed, or having close contact with a cadaver, he said.
The Ebola virus concentrates in the blood but is also present in other bodily fluids, Kortepeter pointed out. Yet, the extent to which such nonblood fluids pose an infection risk remains largely to be determined (Osterholm et al., 2015). In general, increasing viremia over the course of the illness means that any exposure at its peak—around 7 to 10 days after symptom onset—confers maximum risk, he noted. During this period, moreover, patients are most likely to experience high-volume diarrhea, vomiting, bleeding, and coughing. It is also unknown whether or under what circumstances asymptomatic or mildly ill patients can transmit Ebola, he added.
Evolution of Containment Care
In a 2006 consensus article (Smith et al., 2006), Ribner and colleagues defined a biocontainment patient care unit as a clinical facility designed specifically for the care of patients to prevent nosocomial transmission of highly hazardous and contagious diseases, and incorporating those engineering or safety measures that are often used in containment laboratories and have been tested over time in laboratories. Kortepeter, who worked in this type of facility at the U.S. Army Medical Research Institute of Infectious Diseases (USAMRIID), noted that their biocontainment unit was established in 1967, the same year Marburg virus was identified, and 2 years before the discovery of Lassa fever.
Additional advances in biocontainment occurred on the occasion of the first lunar landing, which—fueled in part by Michael Crichton’s best-selling novel, The Andromeda Strain—raised public concern about extraplanetary contamination. In response, the National Aeronautics and Space Administration developed a specialized negative-pressure Airstream trailer to isolate the astronauts for a period of postflight quarantine, Kortepeter said; it included a patient isolator that was never used, except on the set of the film E.T.: The Extra-Terrestrial.
These designs informed USAMRIID’s next high-level containment care unit, dubbed “The Slammer,” Kortepeter observed. It incorporated a host of features, including an external direct pass-thru entry, positive pressure suits, air intake and exhaust filtration, and a steam-sterilized sewer system. Between 1972 and 2004, 22 patients with potentially dangerous infections were admitted to The Slammer, he reported. Quoting Ribner in a recent Time magazine story (von Drehle and Baker, 2014), Kortepeter recalled that The Slammer was once viewed as a sort of “Noah’s Ark.” Today, it serves as an early example of a BSL-4 facility, he said.
Additional improvements were necessary to complete the transition from laboratory biosafety to patient care, Kortepeter explained. When experience with outbreak management demonstrated that incorporating isolation and barrier precautions into patient triage procedures significantly reduced the spread of infectious diseases to health care providers, PPE precautions changed accordingly to incorporate anterooms for donning and doffing, negative pressure where possible, and the mainstay gloves, gowns, masks, shoe covers, and protective eyewear.
However, even with those precautions in place, laboratory transmission of Ebola occurred among nonhuman primates in Reston, Kortepeter recounted. That, along with demonstrations of nonhuman primate infection by aerosol and other routes and of seroconversion to Ebola-Reston among the primates’ human caregivers, resulted in new personal protective equipment requirements for negative pressure and high-efficiency particulate air filter (HEPA) respirators that could be used by those treating patients with prominent cough, vomiting, diarrhea, or hemorrhage, he reported. In 2005, further updates in isolation recommendations stated that hospitals may choose to institute airborne precautions when caring for patients with severe pulmonary involvement or who are undergoing potentially aerosol-generating procedures, he noted.
These recommendations guided the treatment of the initial Ebola patients in the United States in 2014, Kortepeter reported. Then, after two nurses in Dallas were infected with Ebola (as described by Ribner), caregivers were instructed to wear a respirator and to cover all of their skin, Kortepeter stated.
What Next?
The aforementioned advances in containment care have not prevented shocking numbers of health care workers from becoming infected with Ebola in West Africa, Kortepeter acknowledged. Continued uncertainty about the circumstances of transmission is a problem, he noted, but the primary challenge is to protect caregivers from high concentrations of virus in bodily fluids during peak infection. There is no room for mistakes under these circumstances; therefore, as in BSL-4 laboratories, redundancy in protective measures is essential, he insisted. Such measures are best taken in designated containment care facilities capable of handling rare, dangerous infections with which typical hospital staff and clinicians are potentially unfamiliar, he insisted (Kortepeter et al., 2015). At the same
time, every hospital should evaluate its capacity to address infectious diseases, and should be competent to perform basic tasks, such as triage, specimen collection, and isolation, for patients that are best treated in containment care facilities, he said.
Even those critical of the specialized facility model can agree on basic principles of containment care, Kortepeter argued. These include the provision of deliberate care, the need to coordinate movement in treatment and laboratory facilities to prevent contamination, the importance of assessing the risk versus benefit of every procedure, the necessity of practicing key procedures, and the requirement for vigilance by care team members, who will monitor each others’ actions and point out inappropriate actions before harm is done.
Reflecting on the evolution of containment care, Kortepeter concluded that the field has come of age. That being the case, he said, addressing Ribner: if The Slammer was indeed Noah’s Ark, then perhaps it can be said that your ship has either come in, or it has set sail.
DRUG AND VACCINE DEVELOPMENT
Thomas Geisbert of the University of Texas Medical Branch in Galveston and Michael Kurilla described the range of approaches being used to develop preventive vaccines and postexposure therapeutics against Ebola and other filovirus diseases.
Vaccines
Several promising Ebola vaccine candidates existed years prior to the West African epidemic, Geisbert noted. However, because of the limited market for such products and lack of financial incentives for their development, all had stalled in animal trials; moreover, he asserted, progress toward a human vaccine was slowed by the constraints of the FDA’s Animal Rule.7 This legislation,
___________________
7 FDA Animal Rule (CFR Title 21)
- Applies to the development or testing of drugs/biologicals to reduce or prevent serious, life-threatening conditions caused by exposure to lethal or permanently disabling toxic agents (chemical, biological, radiological, or nuclear substances), where human efficacy trials are not feasible or ethical. The Animal Rule was finalized by the FDA and authorized by the U.S. Congress in 2002, following the September 2011 attacks and concerns regarding bioterrorism.
- The FDA can rely on evidence from animal studies to provide substantial evidence of product effectiveness when:
1: There is a reasonably well-understood mechanism for the toxicity of the agent and its amelioration or prevention by the product;
2: The effect is demonstrated in either:
2.1: More than one animal species expected to react with a response predictive for humans; or
2.2: One well-characterized animal species model (adequately evaluated for its responsiveness in humans) for predicting the response in humans.
spurred by biodefense concerns and intended to permit the licensure of drugs for which human efficacy trials would be unethical, nevertheless had impeded potential responses to emerging diseases (Cohen, 2014). As the epidemic spread, those barriers were overcome, and several vaccine candidates were rapidly advanced.
Geisbert briefly characterized four “fast-tracked” Ebola vaccines known to have entered clinical trials at the time of the workshop. Two had already completed phase I safety trials in humans: ChAd3, a chimpanzee adenovirus vector bearing surface glycoprotein genes from two Ebola strains, developed by GlaxoSmithKline (GSK) in collaboration with NIAID and, VSV-delta G, an attenuated vesicular stomatitis virus (a common infection of livestock) engineered to express the Ebola surface glycoprotein, produced by NewLink Genetics and Merck (Geisbert and Feldmann, 2011). These were followed by a more-attenuated “next-generation” VSV-delta G vaccine (Cooper et al., 2008), developed by Profectus Biosciences, and also by a two-step protocol that employs a modified smallpox virus to boost the efficacy of ChAd3, produced by Johnson and Johnson in collaboration with Bavarian Nordic.8
Kurilla reported that a ring vaccination trial in Guinea is under way for one of the candidates. These achievements represent a decade of focused effort, Kurilla emphasized. Nevertheless, he observed, researchers have yet to gain a firm understanding of successful immune response to the Ebola virus, in part because so few people had survived the disease. A December 2014 workshop NIAID cosponsored with the CDC, the FDA, the U.S. Department of Defense, and the Biomedical Advanced Research and Development Authority (BARDA) examined this problem, he noted.
Geisbert agreed that, despite the impressive progress achieved, important questions remain. He noted three areas with key gaps in knowledge. The first concerns the durability of the vaccine candidates. Nearly all filovirus vaccines have been evaluated in virus-challenged nonhuman primates within 8 weeks of their final vaccination—too early to predict their long-term potency. A study found that the ChAd3 vaccine alone at two different doses fails to protect monkeys adequately from Ebola as few as 10 months later (Stanley et al., 2014). “You have to boost with MVA [modified smallpox virus] to get long-term protection,” he reported—but the MVA vaccine has produced adverse effects at the subcutaneous injection site, he added.
___________________
3: The animal study endpoint is clearly related to the desired benefit in humans; and
4: Data or information on the pharmacokinetics and pharmacodynamics of the product or other relevant data or information in animals or humans is sufficiently well understood to allow selection of an effective dose in humans, and it is therefore reasonable to expect the effectiveness of the product in animals to be a reliable indicator of its effectiveness in humans. SOURCE: Geisbert slide #52.
8 ChAd3 and VSV-delta G had entered phase III trials at the time of this writing. For up-to-date information on vaccine development for Ebola see http://www.who.int/medicines/emp_ebola_q_as/en (accessed November 3, 2016).
While live attenuated viruses such as VSV have historically been durable, and a Marburg-VSV vaccine has proven sturdy (Mire et al., 2014), the Ebola construct has not been sufficiently tested to determine if it follows suit, according to Geisbert.
The potency of the virus used to create the current group of Ebola vaccines has also been called into question. New data suggest that these Ebola-Zaire virus seed stocks represented a less-pathogenic genetic variant of the wild-type virus, Geisbert remarked (Kugelman et al., 2012; Volchkova et al., 2011). While this makes a high percentage of the soluble glycoprotein that enables antigenic subversion,9 as described by Kurilla in the next section, the laboratory stock has largely lost that ability, Geisbert explained (Mohan et al., 2012). Accordingly, he added, recent evidence demonstrates that the wild-type Ebola-Zaire virus kills nonhuman primate hosts faster than the laboratory strain.
Given the possible implications of these results for Ebola vaccine efficacy, Geisbert’s group conducted a test of candidate vaccines in macaques that subsequently were infected with wild-type Ebola-Zaire virus. Unexpectedly, Geisbert reported, all the monkeys that received either of the VSV-based vaccines were completely protected and asymptomatic, while all but one monkey that received adenovirus-based vaccines became ill, and one died. “This really tells us that this is something that we have to pay attention to as we develop medical countermeasures, both vaccines and treatments for Ebola, particularly in the context of the current outbreak,” he concluded. However, he added, two promising postexposure therapeutics, ZMapp and TKM-Ebola (described below), had been tested and found effective against the wild-type virus.
In subsequent discussion, Feldmann pointed out that almost all of the laboratory-stain viruses had been grown in Vero cells, rather than in target cells derived from host animals, in which the virus retains its wild-type form. “Primary target cells in animals have a selective pressure to that wild-type virus,” he explained. “Vero cells, unfortunately, do not. That is where most of the variance is generated.” Switching to primary target cell lines would go a long way toward solving this problem, he concluded.
The third area Geisbert discussed as needing further study is the possibility of protecting against multiple species and strains of filoviruses with a single vaccine. “Long-term, we really want to make sure that we have vaccines and treatments that protect against multiple species and strains even within one species of Ebola or Marburg,” he urged. As the small sequence change underlying the capacity for antigenic subversion suggests, even strains with near-perfect sequence homology can vary in their response to the same vaccine.
___________________
9 Antigenic subversion is a novel strategy that Ebola virus uses to evade host immunity by producing large quantities of a truncated glycoprotein isoform (sGP) that is secreted into the extracellular space. This type of glycoprotein not only functions as a decoy for antibody binding but it also shifts antibody production almost exclusively toward itself.
Postexposure Treatments
Geisbert and Kurilla provided an overview of several approaches that have been taken to mitigate the effects of Ebola infection. One prominent strategy, antibody therapy to neutralize the virus, was the focus of a workshop presentation by Erica Ollmann Saphire, described in the next section. Attempts have included the treatment of patients with whole blood, serum, or plasma donated by Ebola survivors, as was first attempted during the 1995 outbreak in Kikwit. Researches have also tried to transfer whole blood among nonhuman primates that survived Ebola, but in most cases it has not worked well, Geisbert noted (Jahrling et al., 2007; Sullivan et al., 2011).
Inhibiting transcription and replication of the Ebola virus is theoretically possible, but it has not been demonstrated. The antiviral drug Favipiravir (also known as T-705 or Avigan), known to be active against several viruses, including West Nile and yellow fever, was recently used to treat Ebola patients, according to Geisbert. Considered a broad spectrum antiviral, its mechanism of action is thought to involve the selective inhibition of the RNA-dependent RNA polymerase, but that has yet to be definitively proven. At the time of the workshop, several studies of this drug were under way in nonhuman primate models of Ebola, he reported. Similarly, the antiviral drug Brincidofovir appears to inhibit replication of a variety of DNA viruses through an unknown mechanism and, surprisingly, Ebola—an RNA virus—in cell culture. Brincidofovir was used to treat the locally acquired Ebola patient in Dallas, he said; it was also intended for studies in Africa that were abruptly halted by its manufacturer, Chimerix. The results of those attempts should be instructive, he observed.
Antisense technology, in the form of small interfering RNAs (siRNAs), provides another means to inhibit Ebola virus activity by suppressing gene expression. These powerful, sequence-specific reagents naturally defend eukaryotic cells against viral pathogens, Geisbert explained. They have been used in cell-culture systems to inhibit the replication of human viruses, including HIV, hepatitis C, poliovirus, severe acute respiratory syndrome, and Marburg; however, effective delivery of siRNAs in vivo remains elusive (Xie et al., 2006).
Geisbert and co-workers attempted to use siRNAs to target three essential Ebola-Zaire virus proteins, he said. In their initial postexposure studies of guinea pigs, only about one in five animals were protected. However, when they repeated the study using a novel lipid-based delivery system for siRNA from Tekmira, every animal survived (Geisbert et al., 2010). This formulation, called TKM-Ebola, was produced by Tekmira and cleared by the FDA under compassionate use to treat several patients who contracted Ebola while serving in Liberia. Geisbert noted that a phase II trial of TKM-Ebola was planned for Africa, if enough patients could be enrolled.
In general, Geisbert observed, repatriated U.S. Ebola patients, many of whom received experimental or compassionate use drugs, fared well compared with their West African counterparts. “I don’t think we can point to any one
postexposure treatment or intervention that is the reason for that,” he concluded; rather, single drugs, combinations of drugs, and 24/7 monitoring by highly trained staff likely all supported superior outcomes.
Discoveries Informing Postexposure Treatments
Kurilla highlighted three key discoveries regarding the Ebola virus, a pathogen that, he observed, “has certainly been able to teach us quite a bit.” The Ebola virus, like many others, interferes with the innate immune system by inhibiting the production of interferon (Basler et al., 2000). The human immune system responds by generating interferon in other ways, he explained, but the Ebola virus counters this defense with viral protein 35 (VP35), a molecule that specifically blocks interferon signaling (Litterman et al., 2015). Four FDA-approved compounds shown to inhibit Ebola virus activity appear to block VP35 action and are being pursued as antiviral leads.
Many viruses also elude the immune system by producing variants that act as decoys, dividing immune defenses so as to conquer them. Ebola virus engages in a particularly insidious type of evasion by producing a variant of a key antigen, a trimer glycoprotein present on the viral surface, Kurilla explained. This variant folds as a dimer, in which state it is soluble (Mohan et al., 2012). As such, it not only functions as a decoy for antibody binding, it shifts antibody production almost exclusively toward itself, a feat its discoverers named “antigenic subversion.”
Another Ebola protein, VP40, can produce three differently folded structures from the same gene (Bornholdt et al., 2013). “How many other infectious agents are actually doing this trick?” Kurilla wondered—and might all organisms have this capacity? More specifically, he continued, this discovery suggests a reason why past Ebola vaccine candidates have failed: perhaps the expression systems used to produce the vaccine antigens failed to fold the protein to a conformation that would generate an effective immune response. This finding has significance for many kinds of infectious diseases, he noted.
Antibodies Against Ebola: A Global Collaboration
Antibody therapy for Ebola has garnered considerable attention owing to its likely safety, bioavailability, and efficacy, and because it could be administered in outbreak-prone areas in a timely manner (Saphire, 2013)—and, most recently, because of the treatment of two aid workers from the United States with the triple antibody cocktail, ZMapp. Erica Ollmann Saphire of The Scripps Research Institute provided an up-to-date account of the science supporting this therapeutic modality and outlined key questions being pursued in the quest to optimize it.
Antibody-antigen interactions The Ebola virus surface glycoprotein (GP), the virus’s sole means of attachment and invasion of host cells, is the main target of
the host immune response, Saphire stated. GP is composed of two subunits, GP1 and GP2. GP trimers stud the surface of the virus, with GP2 forming the base from which each GP1 extends. Mucin-like domains are attached to glycan caps on the tips of each GP1 lobe. GP1 attaches to the host cell, while GP2 mediates fusion with host membrane, she explained. The entire Ebola virus is engulfed by the host cell and drawn into an endosome, Saphire said; there, it is processed by human enzymes called cathepsins, which strip off the mucin-like domains, allowing GP1 to bind to its receptor.
The immune response to this scenario is predictably complex, Saphire observed. “You can actually generate a lot of antibodies against those glycosylated regions and the upper glycan caps and the mucin-like domains,” she said, but “once the virus is brought into the endosome, all of those antibody epitopes are stripped right off the virus along with the antibodies bound [to it]. The virus doesn’t care what kind of antibodies we attach to those places. It will get them cut right off.” On the other hand, antibodies targeting the receptor-binding site—which is conserved across all filoviruses—cannot recognize the entire virus, in which that site is obscured by glycosylation. This dilemma, exacerbated by antigenic subversion (the previously discussed decoy effect afforded by sGP), explains why antibodies that neutralize the Ebola virus in cell culture often do not protect experimental animals from infection, she concluded. This was true of the best neutralizing antibodies available against Ebola in 2007. By 2012, however, several laboratories had independently discovered that a mixture of individually ineffective antibodies could protect animals from Ebola infection, she reported; why that was the case, and how this effect could best be exploited, remained a mystery.
Cocktail collaboration This complex problem is currently being pursued through a novel collaboration among a broad group of researchers who are studying known Ebola antibodies combined in a vast array of cocktails. Saphire is the director of this effort, the Viral Hemorrhagic Fever Immunotherapeutic Consortium, or VIC (Saphire, 2013). “This is an open, fieldwide collaboration,” she said. “Everybody in the world with antibodies against Ebola virus” sent them to Saphire’s laboratory, where they were given code names, packed in identical boxes, and sent to every donor laboratory. Researchers compared the antibodies individually and in combination through diverse in vivo and in vitro assays and protein structure analysis. “If we are doing all of the different kinds of assays, we are able to figure out what works and why. The other idea is that if you need a cocktail, you will probably get a better cocktail if you have a bigger sample size to choose from rather than what is available in this lab or that lab,” she explained.
To achieve that better cocktail, the VIC decided to pursue two strategies in parallel, denoted “tortoise” and “hare” by Saphire. The former, a long-term definitive study of the structure and function of existing antibodies, could produce a definitive therapeutic cocktail, as well as a research benchmark. Knowing this effort would take several years, the researchers devised the “hare” strategy in the
event of a possible crisis, which then materialized in West Africa. The researchers responded by focusing on the six antibodies that constituted two different three-antibody cocktails previously found to protect nonhuman primates infected with Ebola. It was this strategy that delivered ZMapp: a three-antibody cocktail that binds three different locations on the Ebola virus, and which completely protected nonhuman primates treated five days postinfection, Saphire and Kurilla noted.
In the meantime, Saphire reported, “tortoise” studies of the first 81 antibodies have revealed that most effective neutralizing antibodies bind to the base of the GP bowl, on G2. Antibodies across a broad range of epitopes were found to be partially protective in a mouse model of Ebola, and one was completely protective. “Binding a certain epitope isn’t necessarily the answer—except for the base,” she said. “Every single one of them gives some at least partial protection, but they all bind the same place. . . . They will compete with each other . . . [and] they are all knocked out by the same point mutation as well. So one of those should probably be in the cocktail. We will pick the best,” she announced.
Saphire also noted that partially protective antibodies against other sites—that is, those that do not compete with the base binders—could also be included in the definitive cocktail. These include some nonneutralizing antibodies that bind the processed mucinlike domain or the glycan cap of GP1, and yet protect at rates as high as 50 percent, she reported. “I don’t think we can go with neutralization alone anymore when picking out antibodies to pursue,” she concluded; far better predictions of protection—such as the effectiveness of base-binding antibodies—are derived from studying the structural interactions of antibodies and the epitopes they bind. Thus, it will be important to characterize the structural biology of several protective antibodies that bind yet-unknown epitopes. The researchers are also attempting—through a series of high-throughput assays—to determine the functions of nonneutralizing, yet protective, antibodies.
Open questions Saphire ended her presentation with the following sets of intriguing questions and challenges that the VIC’s results to date have raised:
- Of the several base-binding antibodies, so far the best performers, how many should be included in the cocktail? Is it better to have the single best one, without competition, or several competitors to foil potential mutation escape by the virus?
- What exactly makes the base-binding antibodies best in class? If they compete with others, what makes them more effective? What immune effector function are they conferring if not neutralization?
- Some protective antibodies nonetheless bind “decoy” sGP; could that actually confer some benefit to the host?
- As Geisbert noted, the optimal antibody therapeutic would protect against all known Ebola and Marburg viruses. While no antibody yet identified is capable of binding to GP from all five Ebola viruses, antibodies raised
against the conserved receptor-binding site of the Marburg virus—which is exposed, in contrast to that of the Ebola virus—appear promising (Hashiguchi et al., 2015).
At the time of the workshop, several antibodies in the VIC collection had yet to be tested in vivo, and the researchers planned to extend their studies to include antibodies isolated from survivors of the West African epidemic. They also planned to examine antibodies elicited by experimental vaccines, as compared with those resulting from Ebola infection. This raises larger questions, Saphire remarked: “What kind of antibody do you want from a vaccine? Do you want a different kind of antibody preexposure than you do postexposure? Certainly, you might want a different location of antibody or a different immune effector function. Those are things we have to figure out.”
SEIZING OPPORTUNITY
Over the past 50 years, infectious disease research has progressed from “thinking we had it all fixed” to the recognition that there is much we do not know, Kurilla observed. “Those of us who have been working on Ebola countermeasures never anticipated that we would ever have the opportunity like this to be able to test vaccines and therapeutics, and we are doing our best to try to take as much advantage of [these circumstances],” he insisted. “It is hoped we never have an opportunity like this again.”
On the other hand, Kurilla noted, it is important to have plans and resources—including public health infrastructure—in place to conduct research in the event of future infectious disease outbreaks. More specifically, he encouraged the development of platform diagnostics that enable tests for the inevitable unexpected or newly emerging disease to be integrated into an existing system. While point-of-care-diagnostics could also be helpful in theory, he and Towner emphasized that existing tests had yet to prove their usefulness in low-resource settings.
Carole Heilman of NIAID reminded workshop participants that researchers not only need a platform of knowledge and technical capacity from which to launch studies as outbreaks develop, but also a “warm base” of relationships that facilitate contact tracing and sample collection. These connections enabled Gire and coworkers to obtain samples from Ebola patients in Sierra Leone, she pointed out: Gire’s colleague, Pardis Sabeti, had well-established relationships with medical scientists and policy makers in Sierra Leone through her work on Lassa fever. “When indeed there was something to do that wasn’t exactly Lassa but could be utilized within that system, she was able to turn rapidly,” Heilman observed. “These relationships that become extraordinarily important in trusting each other when an outbreak occurs, no matter what it is, are huge, in order to be able to move.”
TOWARD A GLOBAL HEALTH RISK FRAMEWORK
Recognition that the response to the West African Ebola epidemic was unacceptably slow and disorganized has prompted efforts to establish a robust mechanism for planned preparedness for future pandemic disasters. In his presentation to the workshop, Victor Dzau, President of the National Academy of Medicine (NAM) reviewed gaps in the recent public health response to Ebola and described an initiative spearheaded by the NAM to create a global health risk framework.
Although many countries and organizations ultimately responded to the crisis in West Africa, “It took a little bit long for the recognition of the severity of this issue; it took too long to get everybody together and to declare an emergency and have a response,” Dzau observed. Further, he noted, the response was plagued by problems that arose at every level of government. These included failures of disease surveillance, lack of resources for response, and the overall weakness of public health and health care systems in the affected countries, and the previously discussed breakdown in trust and communication between communities and the “outsiders” who sought to deliver care and control disease transmission.
This NAM initiative has identified gaps in four main areas of this response that can serve as focal points for the construction of a global health risk framework, Dzau said; these areas are governance, financing, health systems strengthening, and research and development. Weak governance, leadership, and coordination occurred at all levels and across and between all sectors of government, charitable organizations, private-sector entities, and other nongovernental organizations, he stated. The WHO in particular responded too slowly and failed to assume a leading role; the resulting confusion was exacerbated by disconnection between its Geneva headquarters and its regional and country offices, which presented numerous bureaucratic barriers to international participation in outbreak response, he noted. The crisis also highlighted weaknesses in the International Health Regulations (IHR), as described by Fukuda.
At the time of the workshop, the international community had committed $5 billion to the response to the West African Ebola epidemic, Dzau reported. Based on figures from the World Bank, this number pales in comparison with the cost of the crisis to the affected countries, estimated to exceed $32 billion (more than double the combined 2013 gross domestic product of Guinea, Liberia, and Sierra Leone) (World Bank, 2014). This shortfall in funding resulted from a range of deficits, he noted: no easily mobilized reserve was available to fund emergency response supplies, logistics, and personnel expenses as the epidemic flared, nor were adequate funds available for IHR implementation through the underre-sourced WHO or elsewhere. Until September 2014, the U.S. government’s funding response to the crisis was, like many other wealthy countries, quite modest. Finally, the situation was deemed too risky to finance by the reinsurance industry.
Health systems at all levels were quickly overwhelmed by the Ebola epidemic, which accelerated in part because of the delayed diagnosis and reporting of the earliest cases and the failure to recognize many more, Dzau said. A poor
understanding of community culture and traditions led to mistrust and slow uptake of disease control measures, as Jean Jacques Muyembe-Tamfum of Kinshasa University in the DRC described. The shortage of public health professionals also crippled the response, Dzau asserted. Diagnostics, supplies, and facilities were in short supply, limiting the delivery of even the most basic of clinical care, such as hydration. The allocation of scarce resources, as well as the emergency testing of unproven vaccines and drugs, created ethical and scientific challenges for health care workers.
As several presenters in the previous section pointed out, industry has had little incentive to invest in products to address Ebola, leaving responders reliant on slow and cumbersome diagnostic tests that require high levels of laboratory safety and staff expertise. The lack of multinational collaboration on financing for the research and development of Ebola diagnostics, treatments, and preventives is common to many neglected diseases, Dzau noted. Although the pharmaceutical industry is now investing in Ebola products, it will be months to years before products emerge from clinical trials, he added.
This crisis provides a clear example of the long-sounded warning that timely action is key to containing infectious diseases, Dzau observed. A swift response to an infectious threat should be informed by good planning and evidence, not fear or politics, he said. “Responders need to move as one to avoid mistrust, stigma, or miseducation of communities,” he added. To accomplish this, leaders and roles, resources, and appropriate times for responding must be established before an outbreak occurs. This is the overarching goal of the global health risk framework.
Convening a Global Conversation
At the urging of World Bank president Jim Kim, Dzau agreed to guide the NAM to lead the development of a global health risk framework—and to do it as the West African crisis was still being addressed, in order to take advantage of the “real-time learning” under way. At a December 2014 meeting at the NAM, chaired by the WHO Director-General Margaret Chan and including major stakeholders in the Ebola epidemic, sentiment was overwhelming to proceed as a group to create such a framework, Dzau recalled. On March 5, 2015, participants in a similar meeting at the NAM decided to convene an independent commission to create a detailed report that hoped to inform future United Nations (UN) and WHO policy.
Dzau reported that the Rockefeller, Gates, and Ford Foundations; the Wellcome Trust; and USAID, among others, had already provided funding to support this effort. The report will be prepared by a global commission convened by the NAM, but it will not be an NAM or a U.S. product itself; the NAM’s role is limited to that of secretariat.
For the report to have the hoped-for impact, it has to be issued from an independent international commission, Dzau explained; thus, 17 commissioners,
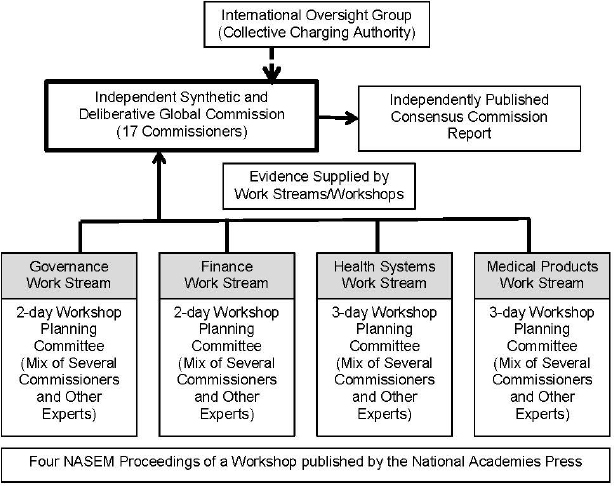
SOURCE: Dzau presentation, March 25, 2015.
reflecting a range of expertise and geopolitical representation, will oversee the report’s preparation. As shown in the organizational diagram depicted in Figure 4-1, four workstreams will address each of the previously identified gap areas, based on information gathered at four workshops scheduled between August and September 2015 and organized by the National Academies of Sciences, Engineering, and Medicine.10
Strategic dissemination will also be necessary to ensure that this report is effective and its recommendations implemented, Dzau acknowledged. “It has to start with the UN and the WHO, because they are going to be the recipients of many of the suggestions,” he observed. A financing mechanism to support distribution was under discussion with potential donors, he said; however, much more work will be needed to ensure that “it’s not just going to be a report sitting on the shelf.”11
___________________
10 The proceedings of these four evidence-gathering workshops can be accessed at https://nam.edu/initiatives/global-health-risk-framework (accessed November 3, 2016).
11 The commission’s report is titled The Neglected Dimension of Global Security: A Framework to Counter Infectious Disease Crises and can be accessed at https://nam.edu/initiatives/global-health-risk-framework (accessed November 3, 2016).






















