4
Case Studies in Big Data and Analysis
In the two morning sessions, the workshop participants learned about some of opportunities that big data holds for infectious disease surveillance and research and about the challenges that need to be addressed in order to take full advantage of those opportunities in a way that benefits public health. The afternoon sessions presented more in-depth examples of how big data are being applied today in real-world settings to tackle problems related to infectious diseases as well as in areas outside of public health. Assaf Anyama, an associate research scientist at the National Aeronautics and Space Administration’s (NASA’s) Goddard Space Flight Center, described how satellite data can inform vector-borne disease research. Luciana Borio, the Food and Drug Administration’s (FDA’s) acting chief scientist, discussed several ways in which the FDA is already leveraging big data to address disease surveillance, drug resistance, and clinical trial improvement. William DuMouchel, the chief statistical scientist at Oracle Health Sciences, then spoke about the use of Bayesian statistics to analyze FDA’s database of spontaneous adverse drug events to spot potential adverse drug–drug interactions. Chicago’s Chief Data Officer Tom Schenk described the city of Chicago’s program to mine its databases in order to identify public health problems, and William So, a policy and program specialist with the Federal Bureau of Investigation’s (FBI’s) Weapons of Mass Destruction Directorate, discussed the FBI’s efforts to secure medical information from cyberattacks. Dale Griffin, an environmental public health microbiologist with the U.S. Geological Survey (USGS), provided examples of how big data and geographical information systems (GISs) can provide information about the natural environmental background distribution of pathogens, and David
Attaway, a solution engineer at ESRI,1 discussed another approach that uses big data and GIS to map vector-borne diseases. Adam Sadilek, a senior scientist at Google, then discussed his work using artificial intelligence with big data associated with social media in order to model the spread of disease.
SATELLITE DATA IN VECTOR-BORNE DISEASE RESEARCH
Data generation is one of NASA’s primary functions, and the agency has several big climate- and weather-related datasets that can be applied to research on vector-borne diseases, Anyambar said. These datasets include 35 years of sea surface temperatures and vegetation patterns, 37 years of precipitation amounts, and 16 years of land surface temperatures. These datasets are useful for disease research because, for example, sea surface temperatures affect precipitation, which in turn affects land surface temperatures and vegetation, creating conditions under which different disease vectors emerge and are able to propagate and spread disease. In particular, Anyamba explained, long-term datasets such as these are valuable because they enable the detection of anomalies, which by themselves are not important but their persistence over time is. Long periods of abnormally wet or dry conditions affect vegetation and are important for creating conditions in which vectors flourish, Anyamba said. These periods are also important metrics used in his modeling work.
Rift Valley Fever
As an example of how these datasets are used to track vector-borne disease outbreaks, Anyamba discussed the modeling work he and his colleagues have done for Rift Valley Fever, a mosquito-borne viral disease originally discovered in Kenya in 1930 which causes severe economic and nutritional impacts on humans through illness and livestock loss (Linthicum et al., 2016). The key climate feature tied to Rift Valley Fever outbreaks is the El Niño/Southern Oscillation cycle that warms the waters of the eastern Pacific and Indian Ocean and increases cloudiness in the eastern Pacific and eastern Africa. Anyamba said that almost all outbreaks of Rift Valley Fever in sub-Saharan Africa have been associated with the El Niño cycle.
In addition to climate information, another important component of the model is its representation of the population dynamics of the species of mosquitos that transmit the virus. Aedes mosquitos, the primary vectors that infect livestock and other animals, appear within 5 to 10 days of heavy rains that cause rivers to flood, Anyamba said, while Culex mosquitos, which spread the virus from livestock and other animals into humans, appear in large numbers from 5
___________________
1 A GIS software company formerly called Environmental Systems Research Institute.
to 30 days after flooding. These population spikes occur in concert with grasses sprouting after heavy rains.
The model calculates anomalies in rainfall and vegetation over a predetermined threshold value that persist over the course of moving 3-month periods for sub-Saharan Africa, East Africa, and Southern Africa (Anyamba et al., 2010). Anyamba explained that these regions were selected based on a literature survey and historical records of climate indicating areas that are conducive to Rift Valley Fever outbreaks. The result is a risk map (see Figure 4-1) that changes over time. A retrospective analysis of the model’s performance showed that it would have successfully predicted 70 percent of the outbreaks in East Africa and about half of the outbreaks in Sudan, but only 30 percent of the outbreaks in Southern Africa. Anyamba said that the poor performance of the model in Southern Africa is largely a result of the model being created using mosquito dynamics studies from Kenya. He is now collaborating with researchers in Southern Africa to develop better information on vector population dynamics in Southern Africa in order to improve the model’s performance. He also pointed out that outbreaks have occurred in regions that had not previously been classified as an epizootic area. “This may be telling us that our epizootic area map is not well defined or that these are new areas where Rift Valley Fever is expanding,” Anyamba said.
Sharing the Results
This project was funded by the U.S. Department of Defense, whose interest is in protecting troops deployed to high-risk areas. Modeling results are also shared with the U.S. Department of Agriculture (USDA) for the purpose of screening commodities imported from high-risk areas and also with the U.S. Department of State, which alerts the resident ambassadors of potentially affected countries that may need help dealing with an outbreak. Information is also provided to the World Health Organization, the Food and Agriculture Organization, the World Organisation for Animal Health, and other international and U.S. government agencies, including an interagency group that focuses on pandemic prevention. All told, Anyamba said, some 25 organizations and agencies receive the model’s output, which includes detailed Google maps pinpointing where early surveillance efforts should focus. USDA publishes a monthly report based on the model’s output on the Agricultural Research Service website.2
Early warning provided by this model has had a positive impact with regard to putting preventive measures in place, Anyamba said. In 2007, for example, the model gave an early warning some 3 months prior to an outbreak in Kenya and 5 months before Rift Valley Fever appeared in Tanzania. The result was that health officials were able to mount a response approximately 2 months earlier than in the epidemics that occurred in 1997 and 1998. Anyamba said that for
___________________
2 See http://www.ars.usda.gov/Business/docs.htm?docid=23464 (accessed October 31, 2016).
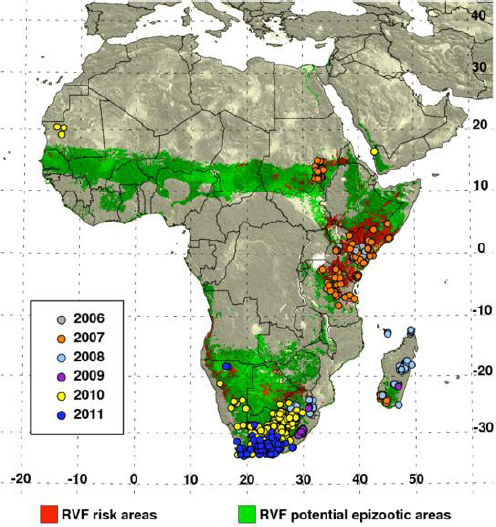
NOTE: “Potential epizootic areas” are those with higher likelihood of flooding events resulting in mass hatching of infected Aedes eggs and subsequent Culex mosquitoes.
SOURCE: Anyamba presentation.
2016 the model warned of potential high-risk areas 1 year in advance, and early mitigation activities have resulted in no reports of Rift Valley Fever activity in the epizootic regions.
Anyamba and his colleagues are now working to apply the same techniques to modeling other vector-borne diseases, including West Nile virus, dengue, Murray Valley encephalitis, and Zika. He predicted that new data sources, including
higher-resolution imaging of cloud cover, will enable more detailed interrogation and modeling of areas prone to outbreaks. He also said that there is still a need to invest in early ground surveillance and rapid field diagnostic capabilities for vector identification and virus isolation. In addition, he stressed that geo-referenced outbreak information—as opposed to country reports—would be important for improving models or building new models.
BIG DATA AT THE FOOD AND DRUG ADMINISTRATION
The FDA’s soul is data, Borio said, and the bigger the dataset, the better, as long as the data meet a standard for quality and integrity. In the infectious disease space, the FDA and the Centers for Disease Control and Prevention (CDC) have partnered to create the GenomeTrakr program, which uses big data for tracing foodborne pathogens back to their sources. This network of whole-genome-sequencing activities by state, federal, and commercial partners has compiled sequence data from more than 51,000 isolates and comprises 17 terabytes of data (see Figure 4-2). According to Borio, this network, which sequences more than 1,000 isolates per month, has enabled a true paradigm shift in the way food-borne outbreaks are identified and investigated. In 2014 case, this database enabled the FDA to trace a multi-state outbreak of Salmonella food poisoning to one nut butter production facility, halt production by the facility, and stop the outbreak (CDC, 2014).
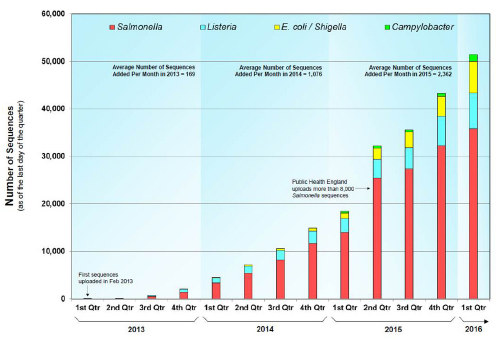
SOURCE: Borio presentation.
Drug Resistance
The FDA is also leveraging big data from next-generation sequencing to analyze drug resistance when it reviews new drug applications for antiviral drugs. Next-generation sequencing, Borio explained, allows the agency to identify lower-frequency mutations in viral genomes that would not be detected using more traditional sequencing methods. In one case, the FDA’s scientists used these data to characterize potential pathways by which hepatitis C virus could become resistant to a novel antiviral agent. Although these mutations were relatively rare, they have the potential to reduce the effectiveness of the drug and limit future treatment options, should resistance develop. This information is now included in the drug’s product label and is used by clinicians to inform optimal treatment regimens. To support this type of data analysis, Borio said, the FDA’s Division of Antiviral Products built a new data architecture and infrastructure using the type of process Catherine Ordun, deputy project manager for data science and health surveillance at Booz Allen Hamilton, had described earlier in the workshop.3
In the area of bacterial resistance, the FDA uses big data to help set what are known as breakpoints, the concentrations of an antibacterial drug at which bacterial species become resistant to a drug. Breakpoints are used to assess the likelihood of treatment success and to inform the care of patients, and the FDA and other organizations update breakpoints to keep pace as new resistance mechanisms appear in bacterial populations. To inform its efforts, the FDA supports a project at Johns Hopkins Medical Institution to create a database with medical information from more than 5,000 patients at multiple hospitals who are being treated for bacterial infections.4 The data collected include the characteristics of the isolated bacteria, the antibacterial drugs used to treat the infection, patient risk factors, and clinical outcomes. Borio said that these data will allow the FDA to use real-world clinical data to establish and update breakpoints in a clinically meaningful manner.
Designing Better Clinical Trials
One area that Borio singled out as a prime candidate for big data applications is improving the design of clinical trials for antibacterial drugs. Typically, she noted, clinical trials to test the efficacy of antibacterial drugs for serious acute infections are difficult and expensive, and this is particularly true for hospital-acquired and ventilator-associated pneumonia (HAP-VAP). The FDA is funding a large observational study by Duke University’s Clinical Trials Transformation Initiative to identify risk factors for patients who develop HAP-VAP in order to
___________________
3 For more information, please reference the “Big Data Architecture and Analytics” section in Chapter 3.
4 More information is available at http://www.fda.gov/ScienceResearch/SpecialTopics/RegulatoryScience/ucm454721.htm (accessed October 31, 2016).
identify those patients who are at high risk and then pre-consent them to participate in a clinical trial when they first enter the hospital. This will make it possible to enroll these patients in a clinical trial at a much earlier time-point in the course of their illness, Borio explained. This approach will also enable trial sponsors to adjust the inclusion and exclusion criteria for participation in a trial, which she predicts will increase enrollment. “The ability to improve the efficiency of clinical trials is one of the conditions that will be necessary to support the pipeline of new antibacterial drugs,” she said.
MODELING ADVERSE DRUG REACTIONS
Until the last decade or so, statisticians were reluctant to apply formal analysis models to the enormous databases that might yield associations between prescription drugs and adverse outcomes, DuMouchel said. Instead, medical reviewers would analyze spontaneous reports of adverse events and try to determine if there was a problem needing further attention. The reason for this reluctance, he explained, was that statisticians then were not interested in working with noisy data. Datasets containing adverse drug event information are inherently noisy because sample sizes are small—which is a good thing, DuMouchel said, because that means that there are not many safety hazards associated with approved drugs. Furthermore, DuMouchel said, the number of potential safety issues is large, making it difficult to decide which ones to focus on in a statistical analysis using older techniques that require deciding ahead of time what to look for at a pre-specified threshold.
The FDA database of spontaneous reports contains some 7 million records of patient reports of one or more adverse events classified according to a fine-grained vocabulary of some 10,000 different terms for approximately 5,000 drugs. From the resulting 50 million drug–event combinations, the expectation would be thousands of “significant results” simply by chance alone, DuMouchel said, and each of these would require follow-up. Traditional statistical techniques would calculate disproportionalities for every drug–event pair, generating millions of comparisons. However, a crucial insight from Bayesian statistics is that it is possible to consider these combinations as ensembles of similar drugs and similar events. While the statistical details incorporated in what are called Bayesian hierarchical models may be complicated, DuMouchel said, the result is that these models will detect changes quickly without too many false positives.
Bayesian hierarchical models can also isolate problems arising from a drug taken commonly with many other drugs, as is the case for many patients with diabetes, heart failure, or AIDS. These models are also useful for the early detection of adverse events in more structured data, such as from clinical trials, and from unstructured data such as Web search logs (White et al., 2016).
BIG DATA TO IMPROVE DELIVERY OF PUBLIC SERVICES
Chicago, Illinois, like many cities, makes a wealth of data about the city available for download by the public via its website.5 In Chicago’s case, the city posts nearly 600 datasets, including the results of testing for the West Nile virus in mosquitos trapped in the city’s 100,000 catch basins distributed across 235 square miles. According to Schenk, Chicago uses WindyGrid, an open-source system that brings together over a dozen data sources in real time into a single application in order to improve operational efficiency. Developed by Chicago, WindyGrid will be released for public use as an open-source project in 2016.
Getting Ahead of Rodents
The advanced analytics unit in Schenk’s office is working on a number of predictive applications, including several related to infectious diseases. One project developed analytics that enables the city to more efficiently focus its limited resources on the control of rodents, an important disease vector for both humans and pets. Using citizen complaints, these analytics generate a heat map showing rodent hotpots as they develop across the city. Working with researchers from Carnegie Mellon University’s Event and Pattern Detection Laboratory, Schenk’s team developed a spatiotemporal model to identify correlations among 350 different factors and spikes in rodent complaints across the city. The model identified 31 factors that predict when and where rodent complaints are most like to occur over the subsequent week.
The resulting Web application now informs a city manager who dispatches city sanitation employees to bait and look for rodents. As an aside, Schenk noted that tests showed that the city manager, who had been at her job for 20 years, was just as good at predicting where rodent outbreaks would occur, but doing so took her 1 to 2 days per week of planning. Today, the Web application handles the planning at night, and she handles dispatch duties each morning. One interesting byproduct of this project is that it identified areas of the city where citizens were not reporting rodent infestations, either because there were language barriers or because the residents did not want the city in their back alleys. “Whatever the reason, we are able to provide better service without being asked for it,” Schenk said, “which we think is a good measure of improving customer service.”
Maximizing Limited Resources
Another project, conducted in partnership with Allstate Insurance’s data science team and the Civic Consulting Alliance, was aimed at improving the effectiveness of restaurant inspections and reducing incidents of food-borne illness. As Schenk explained, Chicago has some 15,000 restaurants and 32 inspectors
___________________
5 See http://data.cityofchicago.org (accessed October 31, 2016).
who are supposed to inspect every restaurant at least once per year. He called this a classic queuing theory problem in which the object is to assign inspectors to those restaurants that most need inspecting. The project team used over a dozen publicly available data sources to develop a model that predicts the likelihood of a food establishment having a critical violation, that is, one that is most likely to lead to food-borne illnesses. Ultimately, 10 variables proved to be useful predictors of critical violations.
When the city went live with this program, it conducted an experiment comparing the performance of the inspectors without the program’s guidance with the expected performance if they had been given the prioritized list generated by the analytics. This 2-month experiment found that the data-driven approach would have to led to the average violation being discovered 7 days earlier, which would have reduced the public’s exposure time to food-borne illness. These analytics, as well as the underlying statistical model and a technical white paper explaining how the model and associated computer code were developed, have been posted online and are available as an open-source application.6
Sharing for the Benefit of Others
Schenk said that, in addition to being an example of governments sharing with each other by leveraging open-source applications, posting these models and the underlying code provides the opportunity for others to improve on the model. Schenk stressed how interesting it has been to collaborate with others on open-source projects to improve public services and said he hopes that Chicago’s experience serves as an example for other cities to follow. He added that the city ran a competition through the Kaggle data science website7 offering a $20,000 first prize for the best model to predict where West Nile would be spotted using data available through the city’s data portal. The city declared three entrants as winners, and his team is now blending the three models to produce a new statistical model that the city hopes to use in 2017. Schenk noted, too, that citizens have created their own applications using city data, including a program that drivers whose cars have been towed can use to find their car. In another instance, a group of citizen scientists took an existing model that had been developed by the federal government and State of Michigan to predict Escherichia coli levels in Lake Michigan and improved the model by using Chicago’s publicly available data. Chicago will pilot the new model as an aid for deciding whether to close the city’s beaches.
When asked what it is about Chicago that enables Schenk’s office to overcome the barriers that many public health officials deal with to get access to data, Schenk said that Chicago’s Mayor, Rahm Emanuel, created the chief data officer
___________________
6 See http://github.com/Chicago/food-inspections-evaluation (accessed October 31, 2016).
7 See https://www.kaggle.com (accessed October 31, 2016).
role when he first came into office and put his office in charge of maintaining and controlling all of the data generated by the city’s different administrative offices. He also said, in response to a question about privacy concerns and pushback from citizens and city employees, that his office is careful about publishing some details in the data, such as the exact address where a crime occurred. There have been instances of pushback, but he characterized them as minor.
SECURITY AND HEALTH CARE SYSTEMS
The health care sector is not immune to cyberattacks or attempts to illegally acquire data or analytics, So said. A report issued in 2016 (Independent Security Evaluators, 2016) based on a 2-year study of 12 health care facilities, two health care data facilities, two medical device companies, and two Web applications (such as the one operated by Chicago but for information on health and health care) identified a number of potential adversaries who were targeting both patient health and patient records. This study also described the potential consequences of a successful attack, including impacts on patient health, the loss of patient records and intellectual property, and even, in the event of multiple or consecutive attacks, damage to the nation’s health care infrastructure.
In 2014, the FBI warned health care providers to guard against cyberattacks after a large U.S. hospital operator reported that Chinese hackers had broken into its computer network and stolen the personal records of 4.5 million patients. According to So, the FBI has found that electronic health records with personally identifying information are worth $50 each on the black market, more than a cyber thief can get for stolen credit card information. Gaining an individual’s DNA sequence could someday enable an adversary to engineer a specific virus to target that individual, a scenario first outlined in 2012 (Hessel et al., 2012) and becoming more feasible as science progresses, So said. In fact, one of the authors of that scenario has started a company that claims to be able to engineer an individual-specific virus within 2 weeks to treat cancer. “What are the security implications if this does come to fruition?” So asked.
Personalized Medicine and Data Security
This question is particularly germane given the Precision Medicine Initiative’s goal of getting high-fidelity gene-sequencing data for at least 1 million Americans.8 This goal also raises additional questions about data sharing and privacy concerns. In addition, So pointed out that there could be national security implications if the resulting data show some genetic vulnerability in specific ethnic groups.
___________________
8 See https://www.whitehouse.gov/precision-medicine (accessed October 31, 2016).
Wearable technology presents other security concerns, given that it provides detailed information on individual activities and location and even the mass movement of individuals. “How does that affect our national security posture,” So asked, “and what is a new paradigm for protecting U.S. interests and the U.S. population? Those are some of the questions that the FBI is trying to better understand.” In closing, he noted that these concerns and others related to big data, life sciences, and national security are detailed in a report produced by the FBI, the American Association for the Advancement of Science, and the United Nations Interregional Crime and Justice Research Institute (Berger, 2014).
INVESTIGATING THE NATURAL BACKGROUND DISTRIBUTION OF PATHOGENS
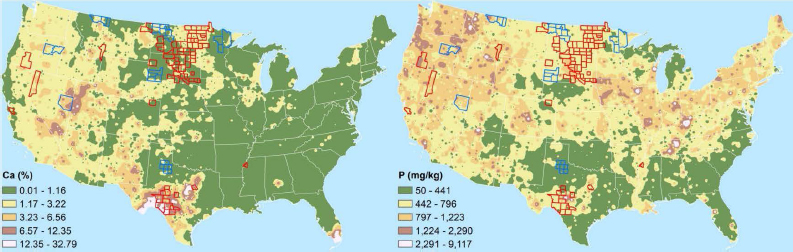
One of the biggest needs in environmental microbiology when investigating a disease outbreak is an understanding of the natural background distribution of pathogens, Griffin said. “The data do not exist to tell us what the natural background distribution in the lower 48 states is for any given pathogen,” he said. “You might be able to tell me Ebola is not here, but you cannot tell me what is here.” Developing such a database, Griffin said, would help determine the risk to any given human, livestock, or wildlife population to exposure to various pathogens and provide insights into the roles that geochemistry and climate play in controlling those organisms.
The process of creating a GIS-based model of pathogen distribution started with the USGS Geochemical Landscapes project, which ran from 2004 to 2010 and determined the geochemistry of soils across the North America from more than 5,000 sample sites (Smith et al., 2013). This survey generated heat maps for 44 different elements and a variety of minerals.9 From the perspective of a microbiologist, these heat maps can serve as a surrogate petri dish containing a specific type of culture media that some organisms will grow on and others will not.
In a pilot study, Griffin and his colleagues combined the maps from the geochemical landscape project with a map of the outbreaks of anthrax in livestock reported since 2000 to see if there were geochemical differences associated with areas that experienced outbreaks and those that did not. This analysis identified a number of chemical elements whose levels were elevated in counties that had experienced outbreaks compared to outbreak-free counties (see Figure 4-3) as well as elements for which higher concentration levels were not associated with outbreaks. Griffin and his collaborators are now developing an application that will enable users to highlight a county and click on each element of the periodic table to search for patterns that may be illustrative for laboratory-based studies.
___________________
9 More information and examples of the heat maps are available at http://mrdata.usgs.gov/soilgeo chemistry (accessed October 31, 2016).
USGS has a wealth of GIS-based models for the distribution of a wide range of climate and environmental variables that Griffin’s team is now combining with the results of polymer chain reaction–based assays for particular microorganisms, including anthrax, Bacillus species, and Naegleria fowleri, a highly lethal amoeba that consumes brain tissue. For the latter organism, low levels of copper and high levels of zinc appear to correlate to where cases of infection with the amoeba have been reported. These models were scheduled to be available on the USGS website for public use in late 2016.
In response to a question, Griffin said it is important to combine knowledge about microbial biology with the information gleaned from the model. For example, Griffin’s analysis identified strontium as an element present in soils where anthrax was found, and when an anthrax researcher questioned him about this, he was able to remind the researcher that strontium is critical to anthrax spore formation.
GIS AND VECTOR-BORNE DISEASES
By combining published information on a variety of climate and geographical data with outbreaks of various infectious disease and known locations of the vectors that transmit the infectious organism and by using a tool called similarity search, Attaway has been able to generate maps that relate environmental and climate conditions to the likelihood of future outbreaks (see Figure 4-4). A similarity search, which relies on a statistical application known as cosine similarity, makes it possible to identify the features or candidates that are most similar or dissimilar to specific features or attributes in much the same way that a consumer application such as Yelp makes recommendations based on a customer criteria.
ArcGIS is another tool that ESRI has developed for predictive analysis, Attaway said. This free tool provides the ability to look at suitable locations for outbreaks and offers other applications as well, such as threat detection, drug use, and urban planning, based on historical data. ArcGIS uses a process called pattern-of-life analysis that enables hypothesis testing and retesting over multiple iterations and produces predictive maps. For example, Attaway and his colleagues used ArcGIS to analyze temperature, precipitation, elevation, land cover, population density, and other variables available from public sources to identify locations suitable for year-round Aedes mosquito activity (see Figure 4-5). In response to a question, he acknowledged that while the analysis itself can be done in minutes, it depends on the availability of data collected over months and even years. ArcGIS’s strength, he said, is its ability to pull data together from a variety of sources, analyze it, and produce actionable insights.
MODELING THE SPREAD OF DISEASE AT SCALE
The challenge that Sadilek is attempting to address involves using artificial intelligence or machine learning in combination with online data to enable the
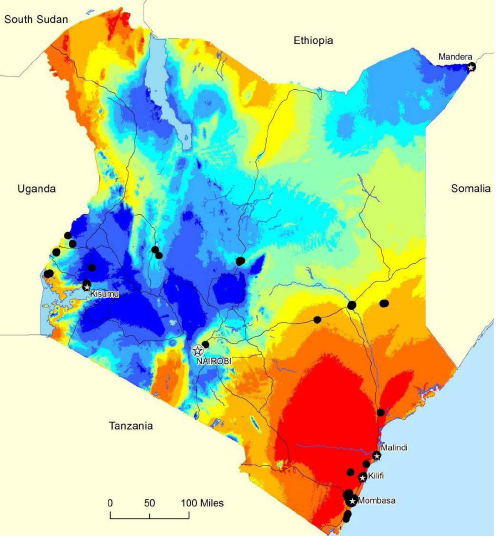
SOURCE: Attaway presentation.
development of new tools for fighting disease. Some 65 percent of adults now use online social media, and much of that activity is via smartphone, which provides real-time, in-the-moment, location-aware data. Sadilek calls this an organic sensor network, and he believes it should be possible to mine the data generated by such a network and derive value from it. Because many of these data have a location component, it could be possible to draw inferences about related events from the data and use them to make predictions.
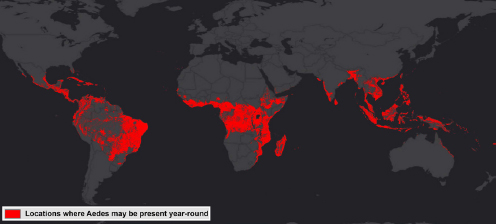
SOURCE: Attaway presentation.
Although the technology is new, this approach is not. In the mid-1800s, John Snow went door to door in London asking people how they felt, and he plotted the results to create a heat map of cholera. His brilliant idea, according to Sadilek, was noting where public wells were located in relationship to clusters of cholera and drawing the conclusion that cholera spreads through contaminated water. “I think we can do this at scale now without going door to door in London, but rather by listening to the data we already have that is generated by people running around and tweeting about their environment, acting as sensors,” Sadilek said.
Following the Spread of Influenza in a City
The key challenge in using tweets, he explained, is to extract useful information from these public messages using some form of natural language processing. Simple algorithms that look for a word such as sick will not work because “I am sick of work” and “I feel sick” cannot both be interpreted as having something to do with illness. One approach would be to hire physicians to examine every tweet with the word “sick” and judge whether the person has the flu or is tweeting about something unrelated. Sadilek and colleagues at the University of Rochester took this approach for a preliminary evaluation, but this type of analysis would never scale. To create something that would scale, they developed a statistical word N-gram model, a learning application that takes human-annotated messages representing illness and others that do not and learns from those examples.
With that model in hand, it is possible to analyze all of the tweets published, identify which ones are likely coming from someone with the flu, and then track
all of the other individuals who happen to come in close contact with those individuals, as identified by the geolocation of their tweets. The result, shown graphically (see Figure 4-6), predicts that someone is exponentially more likely to get sick as the number of sick individuals that he or she encounters increases. This type of analysis also predicts that having a sick friend increases one’s chances of getting sick.
The value of this model, Sadilek said, is that it provides the ability to infer which specific individuals are going to get sick, something that was not possible before. In fact, one evaluation method that he and his colleagues used was to contact a small number of individuals that the model predicted were at risk, invite them to get tested for influenza, and confirm the predictions. Another approach is to aggregate the data to a zip code or county level and see if the inferred numbers agree with official CDC statistics. Such an evaluation found that the inferences correlated well with the real statistics (Sadilek et al., 2012).
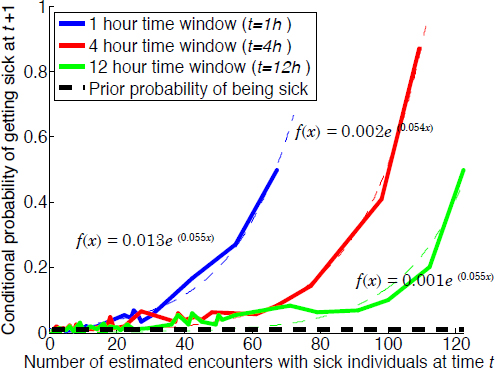
SOURCE: Sadilek presentation.
Tracking Food-Borne Illnesses
Using the same language-processing technique, Sadilek and his collaborators were able to mine tweets to identify restaurants associated with cases of food poisoning in New York City. The output of this model correlated strongly with official health department reports, and Sadilek believes it can serve as a real-time tool to inform restaurant inspections. In fact, a 3-month pilot program conducted with Las Vegas, Nevada, that selected inspections based on an analysis of Twitter feeds and compared the results with a traditional method of assigning restaurants for inspection identified 50 percent more problem restaurants and resulted in 70 percent more closures (Sadilek et al., 2016). Sadilek estimates that over the course of 1 year, adaptive inspections would prevent some 9,000 cases of food poisoning and more than 500 hospitalizations in Las Vegas alone. He also noted that the adaptive inspection list uncovered unpermitted venues, identified a chef with the flu, and resulted in half the normal complaints from consumers about food poisoning that would be otherwise expected. While he has not used this approach to mine data on Zika virus cases, Sadilek said he thought there was an opportunity to do so.
When asked if the growing use of private, encrypted, and closed social media systems posed a threat to this type of application, Sadilek said that while those forms of communication are growing, public venues such as Facebook and Twitter are still seeing growth. He also said that there is more information than just static location in tweets; there is motion data that reflects an individual’s levels of physical activity and driving. In fact, Sadilek has mined mobility patterns for signals of depression and found that individuals whose tweets suggest they are depressed move less and go to restaurants less frequently.
One challenge that these types of efforts face, Sadilek said, is modeling the demographic shifts associated with specific forms of communication. He noted, though, that he and his colleagues were able to generalize their Twitter model for use with Instagram, and he added that the demographics of each particular social media platform are known. Sadilek also addressed the issue of privacy and the potential intrusiveness of this type of analysis and acknowledged that there is a tradeoff between the benefits that these models can produce and the privacy that people will have to give up to reap those benefits.
This page intentionally left blank.