2
Emerging Trends and Products of Biotechnology
This chapter describes the technical, economic, and social trends that will drive the development of biotechnology products likely to emerge over the next 5–10 years. Advances in biotechnology, new actors, economic investments, and societal challenges and concerns all influence the new types of biotechnology products in development. The chapter also outlines the changes in the scope, scale, complexity, and tempo of biotechnology products, which the committee believes will lead to a profusion in the next decade of products made through the use of biotechnology. Types of products likely to be developed—and the kinds of environments in which they may be used—are reviewed.
SETTING THE STAGE: UNDERSTANDING THE KEY DRIVERS FOR FUTURE BIOTECHNOLOGY PRODUCTS
Increasing investment in the bioeconomy, complex societal challenges, the confluence of new technical drivers, and a proliferation of new actors are transforming both biotechnology products and the context in which the U.S. regulatory system operates. For this reason, it is important to track changes in multiple areas that may affect product development and penetration rates. To help set the stage about who and what is influencing the development of new biotechnology products, this section gives a brief overview of a number of these drivers and some of their possible effects on regulation of future products of biotechnology.
Technical Drivers
Several technical drivers have increased the rate at which new products can be developed and also increased the accessibility of modern tools of biotechnology, resulting in an increased number of actors who are able to create biotechnology products. Some key areas include DNA sequencing, synthesis, and editing; standardization of biological parts; and increasingly rapid design-build-test-learn cycles.
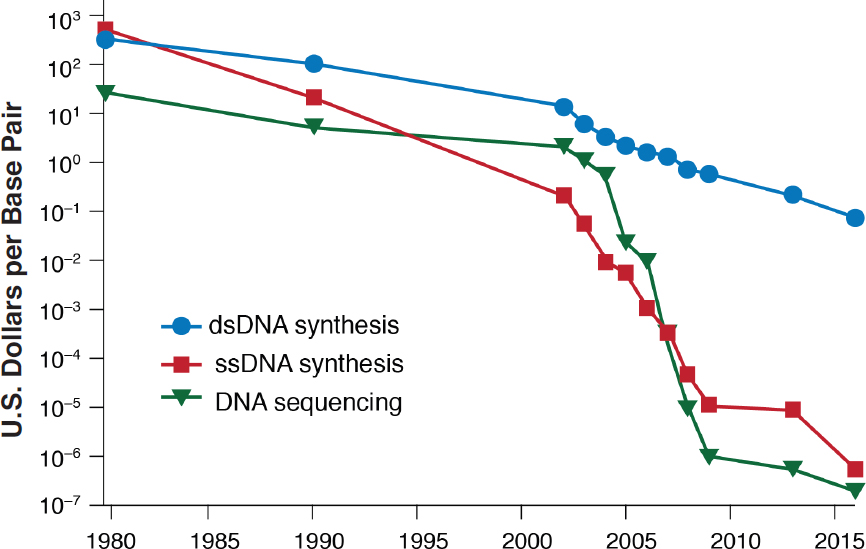
SOURCE: Adapted from Boeke et al. (2016).
New Tools for DNA Sequencing, Synthesis, and Editing
The cost of sequencing DNA dropped by seven orders of magnitude between 2002 and 2008 and has dropped by an additional order of magnitude between 2008 and 2015 (Figure 2-1), representing a rate of decrease that substantially exceeded that of Moore’s law in the 2002–2008 time frame. The price of double-stranded DNA synthesis has also decreased exponentially fast, though at a lower rate, which has enabled the creation of more complex synthetic constructs (Zhou et al., 2015; Chambers et al., 2016), including the construction of whole genomes (Juhas and Ajioka, 2017). In addition, the emergence of genome-engineering technologies (Box 2-1) has enabled targeted modification of DNA sequences—such as insertions, deletions, and site-specific replacements of DNA bases—in a variety of organisms, affording a host of applications (Gaj et al., 2013; Reardon, 2016). Similarly, advances in the understanding of how RNA interference (RNAi) silences gene expression are creating opportunities to create new products, for example for insect-pest control in agricultural crops. Taken together, these trends have made it possible to “mine” genetic data from a wide variety of organisms and then to synthesize new genetic constructs that modify the function of living organisms.
Standardized Biological Parts
The field of synthetic biology has promoted the use of open-access, standardized parts for over a decade. The Registry of Standard Biological Parts1 was established at the Massachusetts Institute
___________________
1 Registry of Standard Biological Parts. Available at http://parts.igem.org/Main_Page. Accessed September 27, 2016.
of Technology as part of the international Genetically Engineered Machine (iGEM) competition, a student synthetic-biology research competition started in 2004. In 2009, the BIOFAB: International Open Facility Advancing Biotechnology (the first biological design-build facility) placed more than 2,500 standardized, quantitatively defined, biological parts in the public domain. The biological parts are compatible, minimal DNA sequences that code for distinct biological functions, such as coding sequences that are responsible for the expression of proteins, promoters, and terminators (that cause transcription to cease). There is also a repository for scientists to share plasmids.2 In 2016, the National Institute of Standards and Technology partnered with Stanford University on a project in Silicon Valley focused on facilitating the standardization of parts relevant to biology.3
All of these efforts are intended to leverage the types of advances that occurred in mechanized production or the later information-technology revolution that relied on standardized parts with reproducible characteristics, combined with increasingly accurate measurement tools. Historically, advances in standardization have been linked to more rapid innovation, improved in-company research and development efforts, increased length and complexity of supply chains, better efficiency and quality management, and greater network effects (including trade). The use of standard parts has accelerated the assembly of complex products in other fields, such as the automotive industry, and such advances may lie ahead for biotechnology as well. Engineering of biological function will increasingly need accurate documentation of components, including description and performance characteristics, combined with incentives to make parts more modular and more predictable, with the critical goal of driving out defects in the field.
The potential importance of standardized biological parts, or other components of biotechnology, is multifold. It enables the reuse of previously engineered devices, creating the ability to design more complex systems more predictably, more rapidly, and with fewer failures. It also enables a
___________________
2 Addgene. Available at https://www.addgene.org. Accessed February 13, 2017.
3 The project is called the Joint Initiative for Metrology in Biology. Available at http://jimb.stanford.edu. Accessed September 27, 2016. The National Institute of Standards and Technology is part of the U.S. Department of Commerce.
wider variety of practitioners to make use of advances of biotechnology by packaging advances in a form that can be reused and matured by others, and it is a first step toward a biotechnology development “ecosystem” in which different companies specialize in components and subsystems, allowing others to make use of the advances across a wide variety of areas at an increased pace.
The use of standardized parts also provides an opportunity for improving regulatory analysis by incorporating safety features that enhance regulatory assessment (similar, perhaps, to the safety ratings issued by the company UL). Achieving safety in complex systems is typically not a simple process and the enormous complexity and variety of biological organisms will be challenging in this regard. Standardization of components provides a possible means to enable more rigorous safety standards and protocols for safety certification, of the sort that is seen in the automotive and civilian aerospace industry.
Increase in the Speed of the Design-Build-Test-Learn Cycle
Traditionally, biotechnology has been challenged by reproducibility issues—an engineered microbe might stop producing or the production rate could fluctuate. Scaling production from micrograms to kilograms and potentially to kilotons was an expensive, high-risk, and costly process. Predictive modeling and computer-aided design tools common in other engineering disciplines were almost nonexistent in biology.
Sun et al. (2014) noted that “[d]ecreasing the design-build-test cycle length is a fundamental challenge facing all engineering disciplines. This is acutely true in synthetic biology.” The design-build-test-learn (DBTL) cycle (Figure 2-2) is the “fundamental building block of effective and
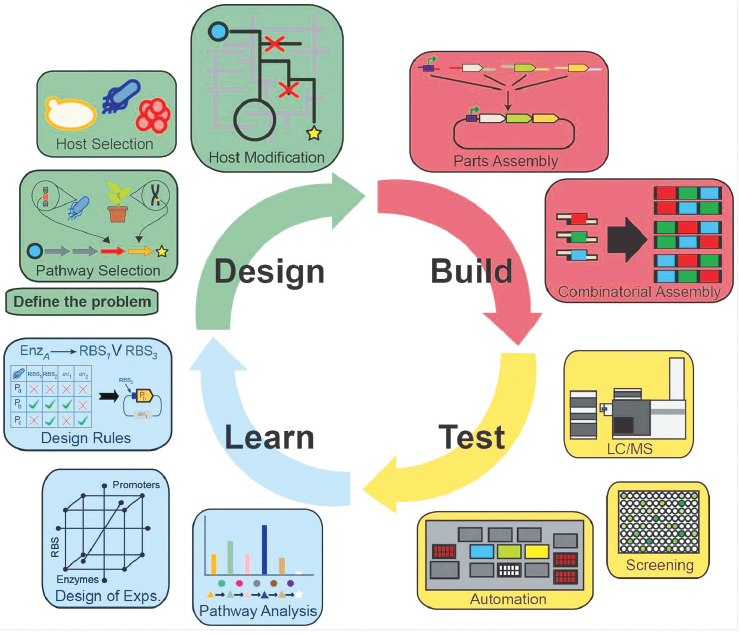
SOURCE: Petzold et al. (2015).
efficient problem solving” (Wheelwright and Clark, 1994:34). It has been adapted for the engineering of biological systems. The design component defines the problem, establishes an approach to solve the problem, and identifies the biological components needed to build or modify. The build component synthesizes, assembles, or edits (or all three) the components of the engineered biological system. The test component characterizes the different biological systems and identifies the variants with the prescribed behavior. The learn component analyzes the test data and informs subsequent iterations of the cycle. The ability to consistently shorten the DBTL cycle can reduce lead times for product development and translate to competitive advantages, especially for those firms who enter markets early with superior products and can command premium prices (Gibson et al., 2010; Gill et al., 2016; Hutchison et al., 2016). However, increases in the efficiency and speed of DBTL cycles also provide challenges and opportunities in the area of safety, where the use of well-established engineering workflows needs to be optimized to ensure that both intermediate and final designs can be developed and deployed in a safe manner.
Additional Technical Drivers
Although advances in DNA sequencing, synthesis and editing, standardization of biological parts, and increasingly rapid DBTL cycles are fundamental technical drivers, there are many other interrelated drivers that play important roles. The expansion of public and private biofoundries—centralized facilities that leverage software and automation to dramatically increase the number of organisms that can be engineered in parallel (Eisenstein, 2016)—is expected to have a substantial effect on the rate of introduction of biotechnology products to the marketplace, perhaps enabling academic laboratories and companies to complete development of multiple biotechnology products per year. The existence of “open-source” approaches to synthetic biology, such as the iGEM parts registry, are enablers for high schools, universities, and the do-it-yourself biology (DIYbio) community to develop courses that teach students and interested parties about engineering biology. Digitization, including not only the representation of DNA in digital repositories but the increased use of standard computer markup languages—such as SBML4 and SBOL5—enable sharing of information about biological models and designs. These are in turn enablers for better tools for predictive models that can be used in the design process, such as SimBiology,6 Clotho,7 and TinkerCell.8
Additional drivers that are not specific to future products of biotechnology but are nonetheless enablers of increasingly rapid product innovation are peer-to-peer sharing platforms such as Benchling,9 which provides software tools for software solution for experiment design, note taking, and molecular biology, or OpenWetWare,10 which provides an open wiki for synthetic biologists; “cloud-based” experimental platforms such as Transcriptic11 and Emerald Cloud Lab12 that provide access to advanced instrumentation and automation on a fee-for-service basis; and a variety of biotechnology incubator spaces, such as QB313 and LabCentral,14 that enable biotechnology startups to have access to advanced laboratory facilities.
___________________
4 Systems Biology Markup Language. Available at http://sbml.org. Accessed October 11, 2016.
5 Synthetic Biology Open Language. Available at http://sbolstandard.org. Accessed October 11, 2016.
6 Simbiology: Model, simulate, analyze biological systems. Available at http://www.mathworks.com/products/simbiology. Accessed October 11, 2016.
7 Clotho. Available at https://www.clothocad.org. Accessed October 11, 2016.
8 TinkerCell. Available at http://www.tinkercell.com. Accessed October 11, 2016.
9 Benchling. Available at https://benchling.com. Accessed October 11, 2016.
10 OpenWetWare. Available at http://openwetware.org. Accessed October 11, 2016.
11 Transcriptic. Available at https://www.transcriptic.com. Accessed October 11, 2016.
12 Emerald Cloud Lab. Available at http://emeraldcloudlab.com. Accessed October 11, 2016.
13 QB3. Available at http://qb3.org. Accessed October 11, 2016.
14 Lab Central. Available at http://labcentral.org. Accessed October 11, 2016.
Impact on Regulation
The combination of technical drivers described above has increased the rate at which new biotechnology products can be created, the scope and complexity of those products, and the number and type of actors who engineer new biotechnology products. In the past, many developers of biotechnology products have been established companies that have strong knowledge of the regulatory system, but when the committee was writing its report there were an increasing number of small- and medium-sized enterprises and DIYbio enthusiasts who were developing technologies and products. Handling the increased scale of products and diversity of developers will require a regulatory system that is agile enough to rapidly adapt to technological change.
Economic Drivers
A second area of rapid change is in the economic drivers that underlie the development of new biotechnology products. Although difficult to accurately determine, total domestic revenues in 2012 from biotechnology—biological, agricultural, and industrial biotechnology products derived using genetically engineered (GE) organisms—have been estimated to be at least $324 billion and to have grown at a rate of more than 5 percent of U.S. gross domestic product annually from 2007 to 2012 (Carlson, 2016).15 Interest from governments and the private sector contributes significantly to this growth.
Government Investment in Biotechnology Products
More than 40 countries, including the United States, have created national strategies or domestic priorities for developing and promoting a 21st-century bioeconomy (EC, 2012; Formas, 2012; OSTP, 2012; OECD, 2015; El-Chichakli et al., 2016). According to the Executive Office of the President’s Office of Science and Technology Policy, the bioeconomy is “research and innovation in the biological sciences [used] to create economic activity and public benefit” (OSTP, 2012:7). Governmental policies for promoting the bioeconomy seek to combine technological innovation, economic growth, ecological sustainability, and resource efficiency (GBC, 2016). For example, the Obama Administration endorsed the U.S. version of this vision in its 2012 National Bioeconomy Blueprint, which set forth broad-based advances in biotechnology, including biobased chemicals, biofuels, and new tools to address challenges and next-generation opportunities in agriculture and manufacturing (OSTP, 2012). The rapid global growth of the bioeconomy is expected to accelerate and increase the demand for biotechnology products (Carlson, 2016).
Private-Sector Investment and Diversification of Sources of Capital
In 2015, U.S. biotechnology companies raised more than $61 billion,16 32 percent more than the previous record set in 2014 (EY, 2016; Figure 2-3). Roughly half of that funding went to companies with revenues of less than $500 million (EY, 2016). Venture capital investment in biotechnology companies increased more than $3 billion from 2014 to 2015, reaching $9.4 billion (EY, 2016). Although much of this money went to companies developing human drugs and medical devices, funding for nonhuman biotechnology was also strong; for example, many of the people who built the dot.com economy, from Jerry Yang of Yahoo to Eric Schmidt of Google and Peter
___________________
15 This estimate includes biotechnology-based drugs and medical devices for human use, which were not part of the committee’s statement of task.
16 Biotechnology companies producing human drugs and medical devices are included in this total.
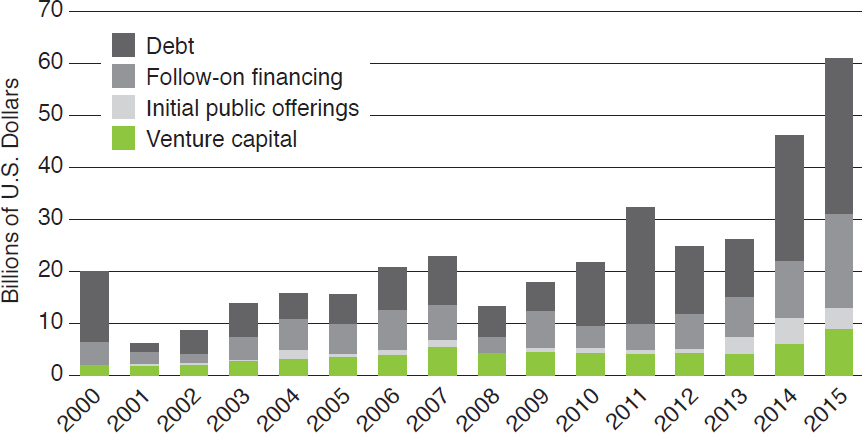
NOTE: Biotechnology companies in the figure include those producing human drugs and medical devices, which were not within the committee’s statement of task.
SOURCE: EY (2016).
Thiel of PayPal, invested heavily in synthetic-biology startup companies between 2009 and 2015 (Hayden, 2015).
Increasingly, capital for biotechnology research is not limited to governmental sources and private-sector funding. Crowdsourcing websites like Indiegogo and Kickstarter have been a source of funding for entrepreneurs and small companies for biotechnology products (see examples in Table 2-1) since they launched in 2008 and 2009, respectively, though Kickstarter announced in 2013 that it would cease funding campaigns that offered a GE organism as a reward following backlash to a campaign for a bioengineered glowing plant (Luzar, 2013). Changes in 2016 to the U.S. Securities and Exchange Commission codes affecting crowdfunding have given rise to a number of equity sites, such as WeFunder.org, which provide another option for biotech startups and entrepreneurs (SEC, 2016). Accelerator organizations like Y Combinator and IndieBio also have provided seed money for biotechnology-product developers.17 The diversification of capital sources has played a role in the diversification of biotechnology-product developers (Box 2-2).
Impact on Regulation
The combination of new investments in biotechnology research, diversified sources of capital, and new players in the development of biotechnology products has the potential to create many new challenges for the biotechnology regulatory system. Crowdfunding and other new financing mechanisms for research and product development may place these activities outside the reach of traditional Coordinated Framework research and biosafety oversight mechanisms such as the National Institutes of Health Guidelines (see Chapter 3). New players and lower barriers to entry
___________________
17 See Y Combinator: Biotech, available at https://www.ycombinator.com/biotech, and Indie Bio: Companies, available at http://sf.indiebio.co/companies-2. Accessed January 8, 2017.
TABLE 2-1 Examples of Crowdfunded Biotechnology Products
| Crowdfunding Site | Project | Year | Amount Raised | |
|---|---|---|---|---|
| Glowing Plant | Kickstarter | Glow-in-the-dark plant | 2013 | $484,013 |
| Real Vegan Cheese | Indiegogo | Milk protein from yeast | 2014 | $37,369 |
| Purdue iGEM | Experiment.com | Waste-clearing E. coli | 2016 | $3,090 |
SOURCES: Information from Kickstarter.com, Indiegogo.com, and Experiment.com cited in Regalado (2016).
may alter the number and types of regulated entities whose activities the regulatory agencies are tasked with overseeing and may blur key jurisdictional concepts such as who is the “product sponsor,” “product developer,” or “manufacturer” that the agencies can regulate. These trends include DIYbio community laboratories, at-home and direct-to-consumer biotechnology developers, crowd-sourced funding and idea generation, and smaller-scale and decentralized manufacturing.
Societal Drivers
In addition to technical and economic drivers, there are a large variety of societal drivers that come into play in the context of both current and future products of biotechnology. The extent to which these societal drivers are directly part of the regulatory system depends on the specifics of the agency and the authority, but they often set the stage for the discussion regarding the evaluation, oversight, and usage of a specific biotechnology product or class of products. The committee reviewed some of the background and context for these societal drivers, with a view toward the types of changes that future biotechnology products may play.
Potential Societal Benefits from Future Biotechnology Products
Biotechnology innovations may have the ability to simultaneously address societal challenges and produce economic benefits. Some biotechnology products are envisaged as tools to help address food security and climate change and to promote “green growth” and environmental sustainability. For example, a U.S. government study predicts that new biotechnology products associated with biomass production could cut U.S. greenhouse gas emissions by 400 million tons per year, or 8 percent (Biomass R&D Board, 2016). Biotechnology products that are successful at addressing societal challenges may have associated economic benefits such as increased productivity and new job creation (including jobs for higher skilled labor).
Nontraditional Players in the Development of Biotechnology
Once the purview of PhD-level researchers, biotechnology is taught in some high school science classes, and, since 2009, a growing number of DIYbio community laboratories in the United States and Europe teach basic biotechnology to nonexperts through formal classes and informal education approaches. In 2013, a survey of the DIYbio community, estimated to be between 3,000 and 4,000 people worldwide, found that the majority of the 359 respondents (82 percent) were in the United States, 10 percent were in Europe, 4 percent in Canada, 1 percent in China, and 2 percent from elsewhere. The community respondents were mostly adult males (75 percent), and few of them (less than 10 percent) work solitarily, that is, outside of community laboratory spaces where technical expertise and equipment are concentrated (Grushkin et al., 2014). Projects sup-
ported at community laboratories involve bacteria, fungi, and plants and largely fall into educational, artistic, and commercial categories. Importantly, U.S. and European community laboratories independently develop and operate under similar codes of conduct that include shared themes of transparency, safety, open access, and education (Kuiken, 2016). DIYbio community laboratories have already developed and adopted safety protocols and provide access to biosafety professionals via a Web-based portal.18 In addition to access to community laboratories, equipment, and biosafety professionals, the DIYbio community has increasing access to funding for their work through crowdfunding platforms, such as Kickstarter and Indiegogo, that allow public donations to support interesting projects.
The iGEM Foundation also encourages nontraditional players through its annual iGEM competition. Student teams compete during the summer to create biotechnology systems, using standardized biological parts, with the objective of making positive contributions to their communities and the world. The number of iGEM teams has grown from 5 U.S. teams in 2004 to more than 300 teams, including 40 high school teams, from 30 countries, in 2016. The distribution of iGEM teams
___________________
18 Ask a biosafety professional your question. Available at http://ask.diybio.org. Accessed January 23, 2017.
from around the world differs from that of the DIYbio community survey; in 2015, the majority of iGEM teams (37 percent) were from Asia, 28 percent from North America, and 25 percent from Europe. The iGEM teams are a growing source of creative ideas and prototypes for future biotechnology products in a large number of global challenge “tracks” from a broad range of applications including therapeutics, energy, environment, food and nutrition, information processing, art and design, hardware, and software. The difference in geographical distribution between 2013 DIYbio community adult survey respondents and 2015 iGEM student teams may foreshadow an increase in future biotechnology products with origins outside the United States. A large percentage of these products could be developed with the intention of export to the United States, which would further increase the number of products U.S. regulatory agencies would have to assess in the next 5–10 years.
Researchers in universities and government laboratories are also active in researching and developing new biotechnology approaches and products, including in specialty crops where markets may be too small to appeal to big agricultural companies. For example, in the 1980s, university researchers began developing a papaya cultivar with resistance to papaya ringspot virus, a devastating disease that threatened to end papaya production in Hawaii. With the GE resistance to the virus,
papaya production has continued on Hawaii; as of 2009, the resistant cultivar was planted on more than 75 percent of papaya-producing acres (USDA–NASS, 2009).
Patent expirations also provide an opportunity for nontraditional players to contribute to biotechnology advances. After the patent expired for the first commercialized soybean with GE resistance to the herbicide glyphosate, researchers at the University of Arkansas’s Crop Variety Improvement Program spent a number of years mating plants with the off-patent resistance gene into a soybean that was in development at the university, essentially creating a GE generic product that offers growers a lower-priced alternative that can be planted in subsequent years without paying a technology fee that is required with patented seeds (Miller, 2014).
Societal Views of Biotechnology
Societal views about biotechnology differ widely among different regions of the world, including within the United States. Although some sectors of U.S. society see biotechnology as a way to solve the great challenges facing humanity today (for example, increase food production efficiency, reduce carbon footprint, or develop more humane farming systems), other sectors of society perceive biotechnology as having both negative and positive aspects or as a threat. The nature of the concern varies but generally pertains to categories that include physical threats to human health, the environment, biodiversity, and resource accessibility as well as other threats such as ownership of biomaterials, technological systems, agriculture or the environment, or genetic resources, and power and voice in decision making about technological choices.
The root of several of the views about biotechnology stems from differing world views about how uncertainty should be treated in decision making, what types of risks should be considered in oversight, the role of technology in addressing problems of society, and who should have power, voice, and choice. Some groups argue for the use of the precautionary principle, adopted by several international treaties such as the Cartagena Protocol on Biodiversity19 and the Convention on Biological Diversity20 as well as the European Union, which argues that decisions should be made and actions taken that err on the side of protecting health and the environment in situations characterized by scientific uncertainty.
Examples of differing world views are most easily cited from experiences with agricultural biotechnology. Although commercially deployed GE crops have generally had favorable economic outcomes for adopters of these crops, outcomes for farmers are heterogeneous because the social and economic effects depend not only on the fit of the crop variety to the environment, but also on the institutional support available to the farmer, such as access to credit, affordable inputs, extension services, and markets (NASEM, 2016b). With regard to access to affordable inputs and to markets, some people argue that the industrialization of agriculture through biotechnology may reduce the number of agents with economic access to agriculture (for example, inability of small farmers to compete with transnational enterprises) as well as decrease genetic diversity that can be achieved through plant-breeding programs and seed sharing at the grower level (Shiva et al., 2011; Vidal, 2011). In the context of biodiversity, some GE herbicide-resistant crops have been found to adversely affect populations of birds that feed on weed seeds due to such high levels of weed control (Gibbons et al., 2006).
Through an in-depth societal impacts analysis, social scientists have found that economic impacts of GE crops for different groups of farmers are mixed; that the political and regulatory context has significant impact on the ability of different groups to benefit; and that current private-sector control of GE crops, which is reinforced by the intellectual property system, reduces the ben-
___________________
19 Cartagena Protocol on Biosafety. Available at https://bch.cbd.int/protocol/text. Accessed January 30, 2017.
20 Convention on Biological Diversity. Available at https://www.cbd.int/convention/text. Accessed January 30, 2017.
efits of GE crops for poor farmers due to high seed costs and distributional constraints (Fischer et al., 2015). On a related note, some are concerned that industrial deployment of certain seed varieties over others may reduce the biodiversity of the food supply (that is, reduction in the seed varieties to be planted and cultivated worldwide) (Jacobsen et al., 2013). Concentration of the global transgenic seed market has been rapidly increasing, and food and agriculture are increasingly controlled by just a few companies which focus on profitable GE crop products (Bonny, 2014).
Another concern is the deployment of resources. Some argue that funding should be devoted to policy rather than technical solutions. For example, using data and historical analysis of GE crops and their effects, some scientists have argued that research funding currently available for the development of GE crops would be better spent in other areas (such as funding for nutrition, policy research, governance, and solutions originating closer to the local level) in order to sustainably provide sufficient food for the world’s growing population (for example, IAASTD, 2009; Jacobsen et al., 2013). Scientists have also found that, so far, genetic engineering has not increased the yield potential of crops, though the technology has been used to reduce yield losses due to pests and research to improve nutrient use and increase the efficiency of photosynthesis is ongoing (NASEM, 2016b).
How important concerns about biotechnology are in comparison to the benefits provided depends not only on the interpretation of evidence, but also on an individual’s and social group’s perception of risk and technologies. Social science offers tools for understanding societal values and provides context for how disruptive technologies are viewed by different subgroups. Risk-perception theory points to different factors and cultural predispositions as to why people perceive risks differently including trust, risk and benefit distributions, controllability, familiarity, and world views (Slovic, 1987; Kahan et al., 2007; Kahan, 2012). Across multiple technological domains these factors affect risk perceptions in people, experts, and citizens alike.
Cultural groups that have been historically marginalized and do not hold as much power in society, such as women and underrepresented minorities, tend to rate risks higher and take more precautious attitudes toward technologies and risk than white men in the United States, even when education, income, and age are accounted for (Finucane et al., 2000; Kahan et al., 2007). An in-depth look at gender differences in response to environmental concerns also found that gender differences in risk perception seem to account for gender differences in worry about health-related environmental problems (McCright and Xiao, 2014). Cultural-cognition theory has been criticized, however, for its limitation to studies in Western, industrialized cultures, for its failings to account for more moderate positions, and for its tendency to blame the individual for their perceptions rather than to focus on risk reduction (Abel, 1985; Marris et al., 1998; van der Linden, 2016).
Other factors that contribute to perceptions of risks and benefits for technologies and their products have been studied and interpreted to form different theories and frameworks. For example, the psychometric paradigm focuses on identifying aspects of the technologies and the risks associated with them, such as whether or not these technologies and risks are dreaded, catastrophic, uncertain, voluntary, and novel, and how these factors affect risk perception and attitudes toward technologies (Fischhoff et al., 1978; Slovic, 1987). This theory suggests that risk perception would become more negative due to anxiety-provoking factors associated with biotechnology products such as uncertainty, involuntary exposure, unfamiliarity, uncontrollability, and catastrophic risk; this has been shown to be true in some studies that include genetic engineering in comparison to other technologies (Slovic, 1987; Marris et al., 1998). Controllability and familiarity have also decreased expert ratings of risks of potential future synthetic-biology products (Cummings and Kuzma, 2017).
There are also several sociological and cultural frameworks that emphasize the role of social factors in consumer attitudes toward products. Trust and confidence in social networks (for example, social groups, communities, extended families, and friends) and societal systems (that is, the market, the political system, the regulatory system, and news media) play an important role in
perceptions of risk for products, especially when those risks are new, uncertain, or ambiguous (Rohrmann and Renn, 2000). They also influence people’s reactions or behaviors in response to risk; for example, lack of trust in industry’s ability to handle risk is associated with greater levels of political activism (Rohrmann and Renn, 2000). Returning to the example of agricultural biotechnology, a national public-perception study found that “trust of government to manage technology” was an important factor for influencing views about the balance of risks versus benefits of GE foods, which in turn affected decisions about acceptance and purchasing (Yue et al., 2015).
The committee notes that most of these theories and factors associated with risk perception are not unique to products of biotechnology. Similar factors can be observed in the perception of climate change by the general public in the United States, for example (Hansen et al., 2003; Kahan et al., 2012). The “deficit model,” often promoted by natural scientists, presumes that there is a knowledge deficit in the public that can be corrected by giving more information and that, if members of the public are given the facts, they will support new technologies (Hansen et al., 2003). To the contrary, the field of public understanding of science has shown that, even with increased knowledge and information, a complex set of societal, political, individual, and cultural factors comes into play in people’s perception of technologies and risk (Hansen et al., 2003; Kahan et al., 2012). According to the fields of public perception and risk communication, education is not likely to change public attitudes; scholars in these fields instead promote the idea of public deliberation, engagement, and communication to help increase technological understanding and mitigate unwarranted perceptions of risk deriving from social amplification and information asymmetries (for example, Thompson, 2011).
Impact on Regulation
Although societal benefits and societal values are not necessarily part of the process of assessing the technological risk associated with biotechnology products, they play an important role in the governance and oversight of biotechnology products, and the laws that society passes reflect societal values. As outlined in the next section, future biotechnology products have the capability to be much more complex than current products, and it is likely that these new products will have the promise of enhanced social benefits at the same time as being more controversial in terms of their use. The role of nontraditional developers may also play a strong role because many developers may not be as aware of the biotechnology regulatory system as current industrial players. The concerns around genome editing that have surrounded the advances in CRISPR-Cas9 are a preview of the scope and complexity of societal discourse that may surround future products of biotechnology (Baltimore et al., 2015; Ledford, 2016; NASEM, 2016b).
Finally, it is important to note that regulation is not the only means of governance and oversight. Codes of conduct, such as those developed in the DIYbio community (Kuiken, 2016), can also play an important role. Community and industry agreement on appropriate oversight frameworks and standards, even in the absence of explicit regulation, will be important for addressing how new products of biotechnology are evaluated.
FUTURE BIOTECHNOLOGY PRODUCTS
As a key element of the committee’s charge, this section describes the future products of biotechnology and how their scope, scale, complexity, and tempo are accelerating. Examples of products that illustrate that change are given. As was stated in Chapter 1, for the purposes of the committee, biotechnology products are defined as products developed through genetic engineering or genome engineering (including products where the engineered DNA molecule is itself the “product,” as in an engineered molecule used as a DNA information-storage medium) or the targeted or
in vitro manipulation of genetic information of organisms, including plants, animals, and microbes. The committee also included products produced by such plants, animals, microbes, and cell-free systems or products derived from all of the above.
Increasing Scope, Scale, Complexity, and Tempo of Products
Many of the invited presentations the committee heard focused on the potential for increased scope, scale, complexity, and tempo of future products of biotechnology. These were also recurring themes throughout the committee’s deliberations.
“Increased scope” means new types of biotechnology products that have not yet been handled by the U.S. regulatory system. Input from companies at the committee’s information-gathering meetings and surveys conducted by the Woodrow Wilson Center for International Scholars revealed that future products of biotechnology are quite diverse and make use of a wide variety of host organisms—bacteria, fungi, plants, animals, and humans—to serve a large number of markets such as health, energy, environment, food, and personal care (Munnelly, 2016; Peck, 2016; Reed, 2016; Sewalt, 2016; Stanton, 2016).21 Work in advanced academic laboratories and an industry report indicate growing interest in in vitro technologies (BIO, 2016). Plants that glow, yogurts that harbor biosensors, pigs that develop twice as much muscle, and microbial communities that may protect honey bees from parasitic mites are just a few possible future products of biotechnology in development.
“Increased scale” refers to the vast number of products (and variants thereof) that will enter the system as a consequence of advances in knowledge and technology. As the technologies for modifying genomes have expanded, so have the number of variants of prospective products. The increase in speed of the DBTL cycle (Gibson et al., 2010; Gill et al., 2016; Hutchison et al., 2016) has lowered the cost of creating variants of genes and pathways such that more prospective products can be screened for product viability.
The U.S. Environmental Protection Agency (EPA) has already noted an increase in the number of biotechnology products that are being submitted to it for regulatory purposes under the Toxic Substances Control Act (TSCA) (Figure 2-5). The number of microbial commercial activity notice submissions to EPA doubled from 2012 to 2013 and grew sharply in subsequent years. In addition, EPA identified that the newer submissions from algal-strain developers were from “companies that have had little or no experience with new substance review under TSCA” (EPA, 2015), which is in line with the observation that more actors with limited experience in the regulatory sphere are working on biotechnology-product development (see Box 2-2).
The number of products being submitted to the U.S. Department of Agriculture’s Animal and Plant Inspection Service (USDA–APHIS) for field release has dropped from its peak in the late 1990s and early 2000s; however, the number of gene constructs22 tested in those field releases is increasing. There has also been an increase since 2009 in the number of acres on which those releases are conducted (Table 2-2).
“Increased complexity” refers to a movement away from single-gene or single-pathway engineering using recombinant-DNA (rDNA) technology to the use of genome engineering to create multiplexed pathways and, at the extreme, engineered microbial communities for release in environments ranging from animal guts to large ecosystems. In considering biotechnology products, two biological “systems” are relevant: the host—the organism into which new material is introduced—and the source organism of the genetic material being introduced. Generally speaking, the majority
___________________
21 See also Synthetic Biology Project. Available at http://www.synbioproject.org. Accessed October 11, 2016.
22 A gene construct is the name used for a functional unit of DNA necessary for the transfer or the expression of a gene of interest. It includes the gene or genes of interest, a marker gene (to facilitate detection inside the organism), and appropriate control sequences as a single package.
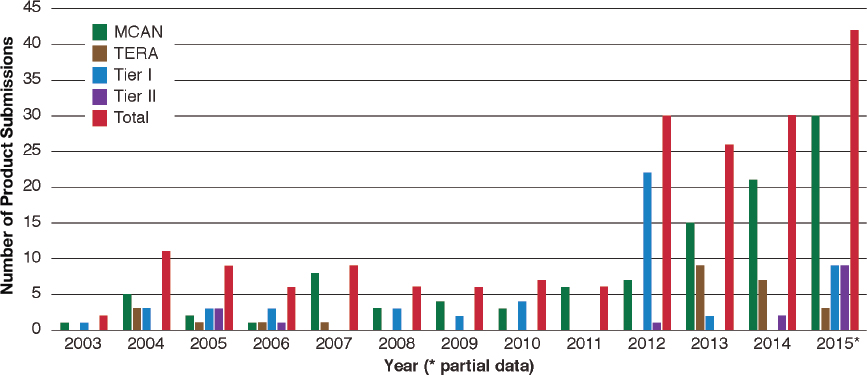
NOTES: MCAN = Microbial commercial activity notice; TERA = TSCA experimental release application. A Tier I exemption requires certain certifications and recordkeeping. A Tier II exemption requires certain certifications and a notification to EPA and EPA review of specific physical containment and control technologies.
SOURCE: EPA (2015).
of biotechnology products in commerce as of 2016—such as crops genetically engineered to resist herbicides or insects—were the result of the transformation of a well-characterized host organism, such as corn or soybean, with a few genes from another source organism that code for a desired trait, such as herbicide resistance along with a selectable marker gene to permit selection for transformed plants (Figure 2-6, column A). Such organisms are easily compared against their nontransformed (that is, nonbiotechnology) counterparts in risk assessments. As biotechnologies have matured over time, new types of products are being developed that allow for the transformation of less well-characterized hosts. For example, new genome-editing technologies allow developers to make changes in genomes of nearly any host organism for which there is a genome sequence available, from microbes to insects to mammals (Figure 2-6, columns B and C; Reardon, 2016). Advances in biotechnology also allow the introduction of novel, synthetic gene sequences and the creation of consortia—a collection of genes derived from multiple unrelated sources—which can be inserted into a host (Figure 2-6, column D). The products that fall in the second, third, and fourth columns are not as easily compared against a nontransformed (nonbiotechnology) counterpart. The challenge this presents to the regulatory system is discussed in Chapter 4.
“Tempo” refers to the groups of similar new products that will predictably follow similar paths through the regulatory system following regulatory decisions about first-of-their-kind products. For example, in 2010 and 2011, USDA–APHIS reviewed a GE herbicide-resistant grass species transformed with biolistics instead of Agrobacterium; the agency decided that because the transformation did not occur through the use of a plant pest and the grass species was not classified as a plant pest, the grass did not qualify as a regulated article.23 Because the grass was not intended for food and did not contain a pesticide, it did not fall under the regulatory purview of the U.S. Food and
___________________
23 See letter from Michael C. Gregoire, Deputy Administrator, U.S. Department of Agriculture–Animal and Plant Health Inspection Service to Richard Shank, Senior Vice President, Scotts Miracle-Gro Company concerning confirmation of regulatory status of Kentucky bluegrass (July 1, 2011). Available at https://www.aphis.usda.gov/brs/aphisdocs/scotts_kbg_resp.pdf. Accessed January 9, 2017.
TABLE 2-2 Number of Releases, Gene Constructs, and Acres Authorized by the U.S. Department of Agriculture’s Animal and Plant Health Inspection Service (USDA–APHIS) for Evaluation, 1987–2012
| Releases | Authorized Gene Constructs | Acresa | |
|---|---|---|---|
| 1987 | 11 | 5 | — |
| 1988 | 16 | 16 | — |
| 1989 | 30 | 30 | — |
| 1990 | 51 | 50 | — |
| 1991 | 90 | 89 | — |
| 1992 | 160 | 160 | — |
| 1993 | 301 | 306 | 948 |
| 1994 | 579 | 585 | 8,117 |
| 1995 | 711 | 710 | 62,394 |
| 1996 | 612 | 604 | 7,084 |
| 1997 | 763 | 761 | 23,817 |
| 1998 | 1,071 | 1,075 | 89,620 |
| 1999 | 983 | 1,005 | 56,959 |
| 2000 | 925 | 904 | 40,199 |
| 2001 | 1,083 | 1,083 | 54,195 |
| 2002 | 1,194 | 1,191 | 139,023 |
| 2003 | 813 | 810 | 24,713 |
| 2004 | 893 | 891 | 58,809 |
| 2005 | 955 | 956 | 99,510 |
| 2006 | 865 | 2,149 | 84,061 |
| 2007 | 932 | 4,920 | 45,931 |
| 2008 | 871 | 8,581 | 182,964 |
| 2009 | 751 | 16,650 | 166,315 |
| 2010 | 660 | 30,770 | 139,517 |
| 2011 | 792 | 35,186 | 235,226 |
| 2012 | 665 | 38,795 | 374,338 |
| 2013 | 602 | 50,963 | 368,384 |
| 2014 | 557 | 39,382 | 365,089 |
| 2015 | 467 | 46,214 | 447,631 |
aRecords of the authorized planting acreages prior to 1993 incomplete.
SOURCE: USDA–APHIS (2017).
Drug Administration (FDA) or EPA either.24 Between that decision in 2011 and December 2016, more than 40 GE plant products had been submitted to USDA–APHIS to determine if the product would fall outside the definition of a regulated article. Most have been determined to be outside the scope of USDA–APHIS’s plant-pest authorities; however, the committee does not know which products have entered consumer markets.25
___________________
24 See Chapter 3 for more discussion of the roles and responsibilities of the regulatory agencies and Appendix D for the Federal Food, Drug, and Cosmetic Act definition of food.
25 Regulated Article Letters of Inquiry. Available at https://www.aphis.usda.gov/aphis/ourfocus/biotechnology/am-i-regulated/Regulated+Article+Letters+of+Inquiry. Accessed January 9, 2017.
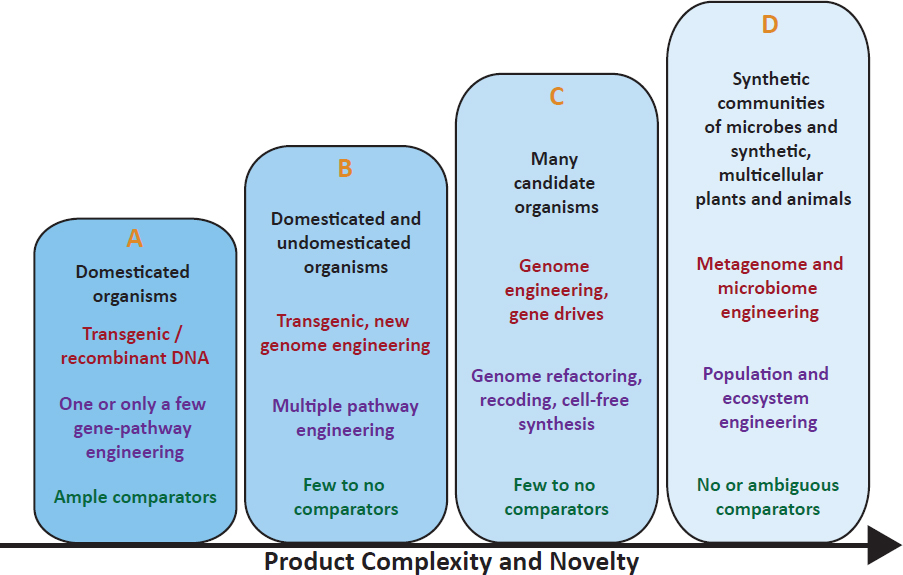
NOTE: Products of biotechnology can be conceptualized as fitting into the depicted columns with the indicated characteristics, moving toward column D as a product increases in complexity and likelihood of providing new challenges for risk assessment.
The committee anticipated that a similar trend will be seen once the first genome-editing animal enters commerce. Advances in CRISPR-mediated genome editing have given rise to predictions of a wide range of precisely engineered animals including monkeys, mosquitoes, pigs, bees, cows, carp, dogs, ferrets, shrews, and chickens for an array of purposes including disease models, drug production, disease control, pets, food production, vector control, and behavioral studies (Reardon, 2016). Although it may be too soon to determine how many of these precisely engineered animals will enter commerce, developers who are planning to create such products of biotechnology are likely to be closely observing the regulatory path followed by the first successfully marketed animal.
Biotechnology Product Classes
The committee scanned the horizon for products emerging in the biotechnology space in the next 5–10 years. Horizon scanning is “a technique for detecting early signs of potentially important developments through a systematic examination of potential threats and opportunities, with emphasis on new technology and its effects on the issue at hand.”26 It is done worldwide to identify and understand the effects of new technologies; a well-established example is in the area of health technologies (Douw et al., 2003). The committee conducted its horizon-scanning exercise by inviting
___________________
26Horizon scanning in Overview of Methodologies. Organisation for Economic Co-operation and Development. Available at http://www.oecd.org/site/schoolingfortomorrowknowledgebase/futuresthinking/overviewofmethodologies.htm. Accessed December 18, 2016.
product developers to speak at the various data-gathering sessions; reviewing submitted public comments; reading scientific literature, popular press reports, and patents; consulting previous reports by the National Academies; searching publicly available iGEM projects; and checking information available on agency websites and crowdfunding websites. It also made use of the Synthetic Biology Database27 curated by the Woodrow Wilson Center, which is focused on a subset of biotechnology products derived through the use of synthetic biology. After careful review of the products, the committee classified the products in order to better manage the task of describing them. All products within the scope of this report were grouped into three major classes: open-release products, contained products, and platforms. The following sections describe the qualities of products in each class and provide examples for products that regulators should expect to confront in the future.
Open-Release Products
This class includes all plants, animals, and microbes that have been engineered (either via rDNA techniques or genome engineering) that will be deliberately released in an open environment (Figure 2-7, Table 2-3). Anticipated future products include logical extensions of these products but also shift to products consisting of organisms whose genome could be largely synthetic, including both organisms with advanced genetic delimitation and those engineered to be capable of sustaining themselves in the environment. Additionally, it includes organisms that have gone extinct (or are close to being extinct) and may be revived (that is, deextinction).
The ability to sustain existence in the environment is a key change between existing products of biotechnology and some of the future ones anticipated in this class. As of 2016, most biotechnology products designed for open release into the environment were introduced into managed systems. For example, GE crops are grown in agricultural fields that are regularly tended and periodically harvested. Only a few deregulated GE crops (such as glyphosate-resistant alfalfa and virus-resistant papaya) are cultivated over more than one growing season, and these exceptions decline in productivity after a few years and are not designed to persist in the environment. However, some biotechnology products in development are being engineered to survive and persist in open environments with minimal or no management.
Furthermore, the types of environments in which a product may persist are likely to become more diverse. Plants and insects may be designed to continue in low-management systems such as forests, pastures, and cityscapes; microbes may be developed to persist in those environments as well as in mines, waterways, and animal guts. The committee anticipates that open-release products created to survive in a wide variety of environments will become more common in the next 5–10 years.
In Figure 2-7 and Table 2-3, biotechnology products are organized around their time to market (horizontal axis) and the family of the host organism, or lack thereof in the case of synthetic products (vertical axis). On the basis of its information-gathering efforts, the committee found that plant hosts will continue to be a dominant area for biotechnology-product development. At the time the committee was writing its report, genetically engineered traits were being introduced into crops other than just corn, soybean, and cotton (which were the most commonly engineered host plants in the 1996–2016 period), and traits besides insect resistance (conferred through the insertion of genes from Bacillus thuringiensis) and herbicide resistance were being engineered (NASEM, 2016b). Additionally, more techniques were being used along with or in place of rDNA technology. Genome editing was being used to knock out genes to create new traits, such as reducing browning of flesh in fruits and vegetables when exposed to oxygen, as was demonstrated in mushrooms (Waltz, 2016). Genome editing was also being used to introduce genes that improve disease resistance, for example
___________________
27 Synthetic Biology Products and Applications Inventory. Available at http://www.synbioproject.org/cpi. Accessed October 11, 2016.

NOTES: This figure diagrams market status of various open-release biotechnology products. Entries in bold are examples of products that the committee has identified as areas with high growth potential. Arrows indicate that the committee anticipated that similar products or products using the same transformation technology (for example genome editing or gene drives) are likely to be developed. a“On Market” is equivalent to “In Use”; thus, products that have received regulatory approval but are not in use were not considered by the committee to be “On Market.” b“Under development” spans products from the prototype stage to field trials.
in wheat (Wang et al., 2014). Drought tolerance in corn (Shi et al., 2017) and more healthful oil quality in soybean (Haun et al., 2014) were being demonstrated through genome editing as well.28 RNAi technology had already been used to introduce traits, including the reduction of browning in the flesh of apples and potatoes (NASEM, 2016b), and such products had cleared U.S. regulatory requirements. Scientists were using RNAi to create virus resistance in cassava, a staple crop in many African countries (Taylor et al., 2012). RNAi was also being used as a way different from rDNA--
___________________
28 At the time the committee was writing its report, a canola variety with herbicide resistance had been commercialized by the company Cibus, which described the variety as developed using genome editing (Gocal, 2015). The resistance arose from a single nucleotide mutation in the BnAHAS1C gene selected for during an oligonucleotide-mediated genome-editing approach. However, Canadian regulatory documents note that, although the variety was developed using a genome-editing approach, Cibus “hypothesized that the single nucleotide mutation was the result of spontaneous somaclonal variation” rather than directly from the oligonucleotide-mediated editing. See Novel Food Information–Cibus Canola Event 5715 (Imidazolinone and Sulfonylurea Herbicide Tolerant). Available at http://www.hc-sc.gc.ca/fn-an/gmf-agm/appro/canola-5715-eng.php. Accessed March 31, 2017.
TABLE 2-3 Market Status of Products Designed for Open Release in the Environmenta
| Product Description | On Marketb | Under Developmentc | Early-Stage Concept | |
|---|---|---|---|---|
| Plants and Plant Products | Bt crops with recombinant DNA (rDNA) | ✓ | ||
| Herbicide-resistant crops with rDNA | ✓ | ✓ | ||
| Disease-resistant crops with rDNA | ✓ | ✓ | ||
| RNAi modified crops | ✓ | ✓✓✓ | ✓✓✓ | |
| Fragrant moss | ✓ | |||
| Do-it-yourself glowing plants | ✓ | |||
| Genome-edited crops | ✓✓✓ | ✓✓✓ | ||
| Crops with CRISPR knockouts | ✓✓✓ | ✓✓✓ | ||
| Grasses for phytoremediation | ✓ | |||
| Plants as sentinels | ✓ | |||
| Crops with increased photosynthesis efficiency | ✓ | |||
| Ever-blooming plants | ✓ | |||
| Nitrogen-fixing nonleguminous plants | ✓ | |||
| Bioluminescent trees | ✓ | |||
| Plants with gene drives for conservation purposes | ✓ | |||
| Plants with gene drives for agricultural purposes | ✓ | |||
| Animals and Animal Products | Fluorescent zebra fish | ✓ | ||
| Sterile insects | ✓ | |||
| Genome-edited animals (e.g., polled cattle) | ✓ | ✓ | ||
| Reduced-allergen goat’s milk | ✓ | |||
| Landmine-detecting mice | ✓ | |||
| Animals revived from near extinction or extinction | ✓ | |||
| Animals with gene drives for control of invasive mammals | ✓ | |||
| Animals with gene drives for control of insect pests | ✓ | |||
| Microbes and Microbial Products | Biosensors/bioreporters | ✓ | ||
| Bioremediation | ✓ | |||
| Engineered algal strains | ✓✓✓ | |||
| Nitrogen-fixing symbionts | ✓ | |||
| Probiotics | ✓ | |||
| Genomically engineered microbial communities | ✓✓✓ | |||
| Biomining/bioleaching | ✓✓✓ | |||
| Synthetic Organisms/Nucleic Acids | Cell-free products | ✓ | ||
| DNA barcodes to track products | ✓ | ✓ | ||
| RNA-based spray for insect-pest control | ✓ | |||
| Genomically recoded organisms | ✓ | |||
| Biological/mechanical hybrid biosensors | ✓ | ✓ |
✓✓✓ = an area the committee has identified as having high growth potential.
aThe table reflects the market status of products at the time the committee was writing the report.
b“On Market” is equivalent to “in use”; thus, products that have received regulatory approval but are not in use were not considered by the committee to be “On Market.”
c“Under development” spans products from the prototype stage to field trials.
mediated toxin expression to introduce insect resistance into corn; this product was deregulated by USDA–APHIS in 2015.29
___________________
29 See Petitions for Determination of Nonregulated Status. U.S. Department of Agriculture–Animal and Plant Health Inspection Service. Available at https://www.aphis.usda.gov/aphis/ourfocus/biotechnology/permits-notifications-petitions/petitions/petition-status. Accessed December 18, 2016.
Further out on the horizon, research was under way to reengineer processes in crops, such as photosynthesis (NASEM, 2016b). In 2016, tobacco was engineered to serve as a proof of concept for photosynthesis improvement. The engineered change increased the speed at which tobacco recovered from overexposure to sunlight (Kromdijk et al., 2016), which in turn increased leaf carbon dioxide uptake and plant dry matter productivity by 15 percent.
The committee anticipated that, in addition to crops, plants engineered for nonagricultural purposes would become more common and that many of these would be designed to persist in the environment under low and no management conditions. For example, a GE American chestnut contained an introduced enzyme, oxalate oxidase, which was extracted from wheat to confer resistance to a blight that has killed about 4 billion chestnut trees in North America since the early 1900s (Zhang et al., 2013). Many years of research had already been undertaken on this tree, and the committee thought it likely that the product would be submitted for regulatory approval within the next 5 years. Given the pressures that U.S. forests face from insect and disease infestation, invasive species, and the effects of climate change (Potter and Conkling, 2016), the committee thought it likely that more tree species will be engineered to resist such stressors.
Plants may also be engineered for biosecurity purposes. Scientists have engineered switchgrass and creeping bentgrass to degrade toxic munitions compounds from the soil in live fire–training ranges to prevent the toxins from leaching into groundwater (Zhang et al., 2017). Plants were also being engineered to serve as sentinels of environmental contamination (Kovalchuk et al., 1998; Kovalchuk and Kovalchuk, 2008).
Plants engineered for biosecurity purposes and trees represent examples of plants that will often be released into environments in which there is little or no management. The committee concluded that agricultural crops will continue to comprise the bulk of biotechnology plants but that more plants designed for little or no continual management will be developed than had been the case before 2016.
One other general type of open-release plant that the committee thought would become more common is one that is not designed for an agricultural or environmental purpose. Instead, the point is to appeal to consumers. The glowing plant mentioned earlier in the chapter is one such example. The committee heard that the product developer of that plant was also working on developing plants that are always in bloom, caffeinated apples, fragrant moss, and mosquito-repelling ivy (Evans, 2016).
As of 2016, few animals had been engineered for open release into the environment, but the committee anticipated that more such products would be developed in the next 5–10 years. At the time the committee was writing its report, U.S. regulators were already seeing insects transformed with rDNA technology that were created for open release. Two insect species had been engineered thus far, using two different approaches for controlling insect populations. A mosquito species (Aedes aegypti) was engineered to prevent the survival of all offspring and released-engineered adults without specialized treatment in laboratory conditions (Oxitec, 2016). The use of this strategy for biocontrol requires the repeated release of engineered adult male mosquitoes that have been reared in a laboratory to serve as breeding stock for wild female mosquitoes. Because the male mosquitoes (and subsequent offspring) are not designed to persist in the environment, it is anticipated that this intervention will have a limited environmental footprint beyond reducing the population of the Aedes aegypti species. A similar concept was being applied to control the population of diamondback moths; however, in this approach, only engineered males (and any resulting male offspring) can survive to adulthood. Over time, the balance of males to females shifts to the point that the population of moths would decline (Harvey-Samuel et al., 2015). As of February 2017, engineered diamondback moth and Aedes aegypti mosquito had completed contained trials
and were being readied for environmental release in field tests in the United States.30 Regulators could expect to see in the future variations of these biocontrol concepts applied to other insect lines, and possibly to mammals (such as invasive rodents) (Campbell et al., 2015).
Changes will also be introduced into livestock, which live in an open environment, though typically under conditions with regular human management and intervention. One example is an introduced trait that makes horned animals hornless. The trait has been demonstrated in cattle. Via TALENs, a naturally occurring polled allele has been isolated from hornless variants that are common in beef breeds and added to embryos from dairy breeds; the research resulted in two hornless calves (Carlson et al., 2016). Another biotechnological change that will likely be made to livestock animals is one that reduces the presence of allergens. As an example, scientists in China have reportedly modified goats to produce allergen-free milk (Zhu et al., 2016) by knocking out the whey protein that is the most common allergen for humans and knocking in a whey protein more similar in composition to human milk.
As with plants, animals may be modified for biosecurity purposes. Small mammals with an acute sense of smell can sniff out landmines without detonating them; giant African pouched rats have already been trained for this purpose. Mice have been engineered to have an odorant receptor that is particularly sensitive to explosives (D’Hulst et al., 2012), and they may transition from laboratory experiments to active identifiers of landmines in the next 5–10 years. Also as with plants, it is possible that more animals will be engineered to appeal to consumers. Engineered fluorescent zebra fish have been on the market since 2003. These fish were initially created for use in a research setting and then marketed to the general public (Nagare et al., 2009). The committee would expect other such novelty products to be developed, particularly in the pet market.
Biotechnology may be used to reintroduce extinct animals, or at least animals that are genetically similar to those that have gone extinct. Research is under way, for example, to use CRISPR genome editing to engineer elephant cells with mammoth versions of genes potentially involved in cold tolerance as a possible preamble to resurrecting the mammoth or creating an Asian elephant able to survive in cold temperatures (Callaway, 2015; Shapiro, 2015). Efforts are also ongoing for deextincting the passenger pigeon (Biello, 2014). The committee presumed that such animals, once approved by regulatory agencies, would be introduced into the environment under minimal or no management conditions.
Plants and animals with gene drives is a subclass of organisms that the committee also thought would be an area of growth in the biotechnology-product space in the next 5–10 years. A gene drive is a system of biased inheritance in which the ability of a genetic element to pass from a parent to its offspring through sexual reproduction is enhanced. Thus, the result of a gene drive is the preferential increase of a specific genotype that determines a specific phenotype from one generation to the next with the intention to spread throughout a population (NASEM, 2016a). Gene-drive mechanisms that have been explored for use in plants are for the control of knapweed for conservation purposes and for the control of pigweed in agricultural fields (NASEM, 2016a). In terms of animal applications, gene-drive mechanisms are being developed to control populations of the mosquito species Culex quinquefasciatus (which is a vector for avian malaria) and populations of non-native mice on islands (which negatively affect the habitats and ecosystems necessary for native species to thrive) (NASEM, 2016a). A future potential application is the use of a gene drive to spread disease resistance through a population of snails to prevent the continued transmission of schistosomiasis (Tennessen et al., 2015). Application of this technology to other invasive species
___________________
30 Sterilized pink bollworm with a genetically engineered fluorescent marker has been field tested in Arizona since 2006. The marker allows for easy identification that the insect is sterile. Sterility in the insect has been achieved by irradiation, not genetic engineering.
has also been discussed in the popular press (Langin, 2014), yet it is unclear how many of these suggestions are being further developed.
As with animals, few microbes engineered for open release into the environment had been developed and successfully approved by regulatory agencies as of 2016. However, efforts have been under way for many years to genetically engineer microbes destined for the environment for a number of applications, including bioremediation (Cases and de Lorenzo, 2005) and as environmental biosensors (Xu et al., 2013). Such products were envisioned as future products of biotechnology in the 1986 Coordinated Framework for Regulation of Biotechnology. In the 1980s and early 1990s, limited field trials occurred with live GE organisms for such purposes as frost prevention and pest control. Recombinant biopesticides briefly formed a niche market in the early 1990s prior to development of insect-resistant transgenic crops. The failure of product advancement to the marketplace for engineered microbes may be attributed in some cases to a lack of performance of the product against expectations rather than evidence of failed safety tests (Wozniak et al., 2012). Other views posit alternative explanations for the lack of advancement of engineered microorganisms to commerce, including sentiment against GE organisms bringing the field of bioremediation to a standstill (de Lorenzo et al., 2016) and the inability to patent “non-novel” bioreporter technologies creating a disincentive to private-sector investment (Xu et al., 2013). An engineered bacterium, Pseudomonas fluorescens HK44, was the first GE microorganism to be field released for subsurface soil bioremediation of polycyclic aromatic hydrocarbons such as naphthalene and salicylate using bioluminescence. Despite promising results under a range of conditions, few applications of Pseudomonas fluorescens HK44 in relevant ecosystems have been implemented, “primarily due to legislative restrictions encompassing the use of genetically engineered microorganisms and their environmental release” (Trögl et al., 2012).
However, a Pseudomonas putida strain genetically engineered for aerobic bioremediation of 1,2,3-trichloropropane, a recalcitrant chlorinated hydrocarbon used as an industrial solvent, paint remover, and cleaning agent among other uses, was recently created as an attractive option for groundwater decontamination (Samin et al., 2014). Although the engineered strain performed well in a bioreactor with 1,2,3-trichloropropane as the only organic carbon source, its prospects for use as continuous bioremediation of 1,2,3-trichloropropane in contaminated environments remain untested. Because single-strain bioremediation approaches may be vulnerable to slow growth or high decay rates caused by reactive side products, the use of microbial consortia that could stimulate growth rates by cross-feeding or remove reactive dead-end metabolites is envisioned as a possible strategy to mitigate these vulnerabilities (Samin et al., 2014; Jia et al., 2016; Lindemann et al., 2016). Such anticipated environmental experiments of large-scale, self-propagating bioremediation approaches aimed to reduce the impacts of human-made pollution may be supported by the notion that “assuming a reasonable risk is preferable to the sure disastrous effect of inaction,” given the prospect of increasing environmental and ecosystem degradation (de Lorenzo et al., 2016).
Synthetic biology holds a great deal of potential for microbes in open environments, an area that the committee sees as gaining momentum. Despite the historical challenges discussed above, this area is very active in current research (OSTP, 2012) and includes research to characterize and manipulate the microbiomes of essentially any life form or environment of interest. Prototype engineered biosensors already exist to traverse mammalian guts and “record” events of interest to which the microbe was exposed (Kotula et al., 2014). Similar systems are following in areas such as pollinator health (Kwong and Moran, 2016). Many researchers (Fredrickson, 2015; Jia et al., 2016) and a number of iGEM teams (iGEM, 2012, 2013, 2015a,b) have worked to establish stable synthetic consortia of microorganisms—and the biological principles behind their establishment and maintenance—that could be used as the bases of a wide variety of future applications. At the time the committee was writing its report, product developers were working to create engineered
consortia of microorganisms to market as potential new products for open release for a broad range of markets including mining and human and plant nutrition.
Biomining involves the use of microorganisms to extract rare and base metals from minerals and ore. For example, the bacterium Acidithiobacillus ferrooxidans and relatives are able to assist with bio-oxidation and bioleaching of many types of minerals for mining. Biotechnology is being applied to enhance these processes, and research has been conducted to engineer microbes for increased redox potential and leaching rates (Brune and Bayer, 2012). Researchers and companies, such as Universal BioMining, are working on synthetic-biology techniques to create inoculants containing extremophiles with targeted genetic alterations designed for metal extraction (DaCunha, 2016). When deployed at scale in the field, these inoculants could greatly improve the capture of valuable metals such as gold and copper from the increasing supply of low-grade ore, while simultaneously reducing the environmental effects caused by traditional mining practices.
An invited speaker (Cumbers, 2016) described to the committee a series of small company efforts where open-release biotechnology products containing synthetically engineered microorganisms for the human gut are envisioned for enriched foods, medical purposes, and lifespan elongation. Microbial products that are genetically engineered are also under development for plant microbiomes. The clearest example is the manipulation of nitrogen fixation in heterologous prokaryotes (Smanski et al., 2014). One product concept is to implement the cluster in plant-associated microbes or, conversely, in microbes that the plant will selectively internalize. (The concept could also be implemented as a transgenic manipulation of the plant or a specific compartment in the plant.) Other applications currently under development are engineered bacterial strains that secrete double-stranded RNA (dsRNA) that can be applied to crop plants topically. The dsRNA produced by the bacteria is designed to serve as a crop protection agent, causing harm to pest insects that consume the treated plant (Killiny et al., 2014). Future microbial-consortium products of biotechnology, whether for bioremediation, biomining, or nutrition, present substantial challenges to regulators given their complexity, lack of comparators to nonbiotechnology products, and lack of predictive risk-assessment pathways available to evaluate their impacts and safety (see Chapter 4).
Advances in DNA synthesis and assembly technologies have created the possibility of engineering organisms whose genome is substantially altered and may consist largely of DNA sequences that have been chemically synthesized (Boeke et al., 2016; Hutchison et al., 2016). For example, work from the research groups of Farren Isaacs, George Church, and others have produced a family of novel prokaryotes (Isaacs et al., 2011; Lajoie et al., 2013; Mandell et al., 2015; Napolitano et al., 2016; Ostrov et al., 2016). Through advanced genome-engineering tools, an organism’s genome has been recoded to change its organization (Isaacs et al., 2011; Lajoie et al., 2013) or to alter fundamentally how it codes and decodes information. Some variations repurpose codon usage without adding a requirement for new amino acids (Isaacs et al., 2011; Lajoie et al., 2013; Rovner et al., 2015; Napolitano et al., 2016; Ostrov et al., 2016), and other strategies fundamentally change codon usage and add new amino acid requirements (Mandell et al., 2015; Rovner et al., 2015). Each of these options results in genomically recoded organisms (GROs) with increased genetic isolation from other prokaryotes in the environment. While GROs are still in the early stages of development for research purposes, the committee can imagine open-release applications of such organisms for agricultural, bioremediation, and nutritional (probiotic) purposes. Their genomic isolation would prevent meaningful gene flow to or from the organism (Lajoie et al., 2013; Ma and Isaacs, 2016) and some of these recoding operations can be further modified to create highly effective kill switches, which can be leveraged to allow for tightly controlled environmental releases. No open-release products are being currently developed (or at least not yet disclosed), but one could anticipate the movement of these into regulatory purview in the future. It is much more likely that regulators will first see GROs for use as a contained product.
Other examples of synthetic products currently under development are synthetic RNAi to be used as pesticides and DNA barcodes to track products through the manufacturer pipeline. RNAi sprays, produced via a cell-free expression system to protect crops from insect pests, are under development and being tested in small greenhouse trials, with one demonstration showing the protection of potato plants from the Colorado potato beetle for up to 28 days (San Miguel and Scott, 2016). DNA barcodes have just begun to be utilized by the U.S. military to track small mechanical parts, such as bolts, to counter their rising concern over counterfeit parts of low quality (Mizokami, 2016). In this application, a DNA sequence is applied to a mechanical part using an epoxy ink. The novelty of such a tracking device is that it can be applied to small components of a system without impeding their function.
Contained Products
A second major class of products is those that are largely contained, that is, used in industrial fermentation or produced in other sealed environments such as laboratories or ponds. Organisms of many genera are used in fermenters to produce commodity chemicals, fuels, specialty chemicals or intermediates, enzymes, polymers, food additives, and flavors. When considering the laboratory as a contained environment, then many examples of transgenic animals from vendors are widely used today for research and development. Because performing biotechnology in contained environments allows higher control over the choice of host organism, systems with advanced molecular toolboxes are already in high use. As above, possible future biotechnology products captured in Figure 2-8 and Table 2-4 are organized around their time to market (horizontal axis) and the family of the host organism, or lack thereof in the case of synthetic products (vertical axis).
On the basis of its information-gathering efforts, the committee concluded that future biotechnology products that are produced in contained environments are more likely to be microbial based or synthetically based rather than based on an animal or plant host. However, the committee did identify a few animal and plant products, and they or variants thereof may become more common in the next 5–10 years. The CRISPR-edited mushroom, described in the open-release section above, can also be cultivated as a contained product in a laboratory or greenhouse setting (Waltz, 2016). An animal example approved by the regulatory agencies when the committee was writing its report was the GE salmon, which contains a gene insertion that speeds the pace at which the fish grows to market size. GE salmon are considered to be “contained” as a condition of regulatory approval because they are restricted to growth in specific land-based facilities and are prohibited from being grown in ocean net pens.31 Another example of an existing animal product is laboratory animals, many of which are designed to have genes knocked in or out for experimental purposes, such as mini-swine (F. Li et al., 2014) or dogs (Zou et al., 2015) engineered for research purposes.32 It is not clear how many follow-on product concepts are planned in this space by developers.
Other products that are not on the market yet, but that the committee thought could be commercialized in the near future, are polymers produced by plants for industrial use—for example, silk and collagen (reviewed by Van Beilen and Poirier, 2008)—and animal products derived from animal cells rather than from animals themselves. The committee heard from product developers working to create hamburgers by editing and expanding cultures of muscle cells in the laboratory (Datar, 2016; Shigeta, 2016). Leather from animal proteins expressed in skin cells has also been cre-
___________________
31 U.S. Food and Drug Administration. AquAdvantage Salmon Fact Sheet. Available at http://www.fda.gov/AnimalVeterinary/DevelopmentApprovalProcess/GeneticEngineering/GeneticallyEngineeredAnimals/ucm473238.htm. Accessed December 19, 2016.
32 It is possible that such animals could one day be released into open environments for use as pets. It was reported in 2015 that an institute in China was considering selling as pets pigs that had been genome-edited for use as models for human disease (Cyranoski, 2015).
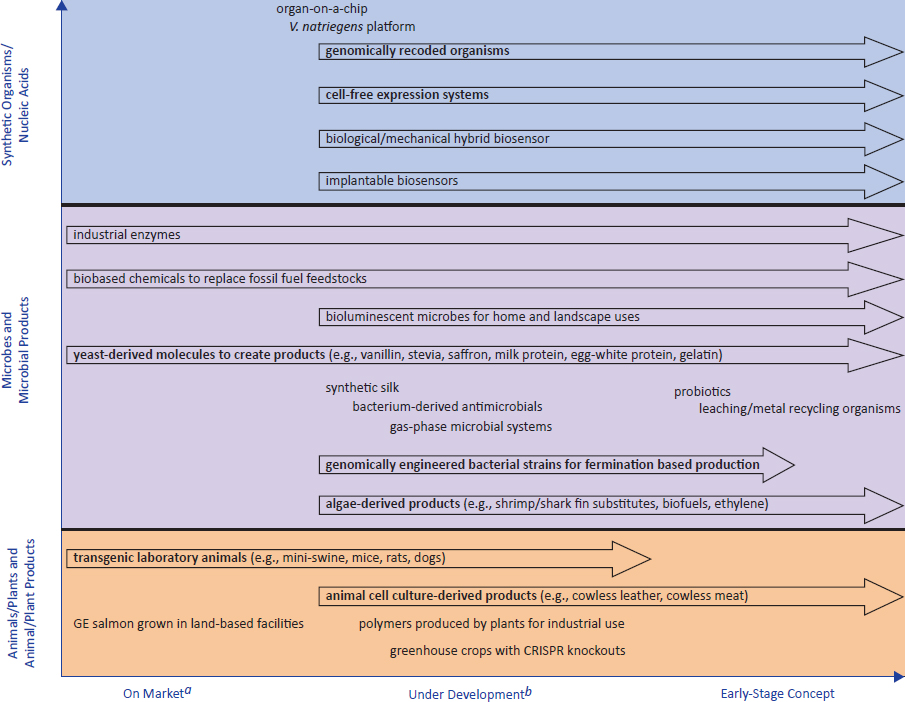

NOTES: This figure diagrams the type of technology and market status of various contained biotechnology products. Entries in bold are examples of products that the committee has identified as areas with high growth potential. Arrows indicate that the committee anticipated that similar products or products using the same transformation technology (for example genome editing or gene drives) are likely to be developed. a“On Market” is equivalent to “In Use”; thus, products that have received regulatory approval but are not in use were not considered by the committee to be “On Market.” b“Under development” spans products from the prototype stage to field trials.
ated, and companies in this area are expanding and preparing to bring their products to the market in the near future (Shontell, 2016).
The committee also anticipated that replacements for products traditionally sourced from animals will be more and more likely to come from microbes such as yeasts or algae. In presentations by product developers, the committee heard about yeast strains developed to produce various products using traditional fermentation methods (Datar, 2016; Shigeta, 2016). A popular area of development is the creation of food products such as milk proteins to be combined with sugars and oil for the creation of vegan milk and cheese that are biologically the same as the animal sources of these products (Gertz, 2014; Bowler, 2016; Datar, 2016). A few other examples of food or flavor products being made in this way are egg whites (Datar, 2016; Shigeta, 2016), stevia,33 gelatin (Duan et al.,
___________________
33 In early 2016, Cargill, Inc., submitted a generally-recognized-as-safe exemption claim to FDA for steviol glycosides from Saccharomyces cerevisiae expressing steviol glycoside biosynthesis pathway (Cargill, 2016). In May 2016, FDA responded that it had no questions about Cargill’s conclusion that the steviol glycosides were generally recognized as safe, based on information provided by Cargill and other information available to the agency (FDA, 2016).
TABLE 2-4 Market Status of Contained Productsa
| Product Description | On Marketb | Under Developmentc | Early-Stage Concept | |
|---|---|---|---|---|
| Animals/Plants and Animal/Plant Products | Transgenic laboratory animals (mini-swine, mice, rats, dogs) | ✓ | ✓✓✓ | |
| Genetically engineered salmon grown in land-based facilities | ✓ | |||
| Animal cell culture–derived products (e.g., cowless leather and cowless meat) | ✓✓✓ | ✓✓✓ | ||
| Polymers produced by plants for industrial use | ✓ | |||
| Greenhouse crops with CRISPR knockouts | ✓ | |||
| Microbes and Microbial Products | Industrial enzymes | ✓ | ✓ | ✓ |
| Biobased chemicals to replace fossil fuel feedstocks | ✓ | ✓ | ✓ | |
| Bioluminescent microbes for home and landscape uses | ✓ | ✓ | ||
| Yeast-derived molecules to create products (e.g., vanillin, stevia, saffron, egg whites, milk protein, gelatin) | ✓ | ✓✓✓ | ✓✓✓ | |
| Synthetic silk | ✓ | |||
| Bacterium-derived antimicrobials | ✓ | |||
| Genomically engineered bacterial strains for fermentation-based products | ✓✓✓ | |||
| Gas-phase microbial systems | ✓ | |||
| Algae-derived products (e.g., substitute for shark fins and shrimp, biofuels, ethylene) | ✓✓✓ | ✓✓✓ | ||
| Probiotics | ✓ | |||
| Leaching/metal recycling organisms | ✓ | |||
| Synthetic Organisms/Nucleic Acids | Organ-on-a-chip | ✓ | ||
| V. natriegens platform | ✓ | ✓ | ||
| Genomically recoded organisms | ✓✓✓ | ✓✓✓ | ||
| Cell-free expression systems | ✓✓✓ | ✓✓✓ | ||
| Biological–mechanical hybrid biosensor | ✓ | ✓ | ||
| Implantable biosensors | ✓ | ✓ |
✓✓✓ = an area the committee has identified as having high growth potential.
aThe table reflects the market status of products at the time the committee was writing the report.
b“On Market” is equivalent to “in use”; thus, products that have received regulatory approval but are not in use were not considered by the committee to be “On Market.”
c“Under development” spans products from the prototype stage to field trials.
2011), and vanillin (Hansen et al., 2009). The committee expects to see growth in the number and market acceptance of such food products as they are being marketed as more sustainable and cruelty-free products.34 Other applications include the production of materials, such a silk (Tokareva et al., 2013), which some developers are working to optimize to create improved versions of this material (Seltenrich, 2015). Similarly, strains of algae have been engineered for use in fermentation processes to produce food products and other important industrial chemicals. Products such as vegan shrimp or shark fins made from collagen expressed by GE algae and combined with other ingredients are under development (Davis, 2015; Bryce, 2016; Shigeta, 2016). Other GE algae strains are being utilized
___________________
34 See, for example, Memphis Meats, About Us: Better Meat, Better World, available at http://www.memphismeats.com/about-us, and Clara Foods: Egg Whites Without Hens at http://www.new-harvest.org/clara_foods. Accessed January 8, 2017.
to produce ethylene (Xiong et al., 2015), for use in making downstream products such as plastic, polyester, and PVC pipes, and for biofuel production (Radakovits et al., 2010).
The committee also expects to see microbial organisms with bioluminescent capabilities engineered to glow brighter for human use as novelty items in the home (Lafrance, 2015; Lombardo, 2015) or as greener light sources in urban settings (Marcellin, 2016). These products are designed for use in contained settings and not intended for consumers to release into the environment. Other microbial products are bacterial strains engineered to extract metal compounds and improve recovery from mining ores in a contained reactor (Schippers et al., 2014) or to produce a wide variety of chemical compounds such as antimicrobials or industrial enzymes. The regulators have long seen such products, with the classic example being insulin for medical purposes. A more recent, synthetic biology example is being deployed against traditional gas-phase fermentation organisms (already in use at pilot or demonstration scale for ethanol production from CO or CO2) to expand the chemical targets beyond ethanol, acetate, and butanediol (Liew et al., 2016). Engineering bacterial strains to improve their production abilities is an area that the committee anticipates will grow. Many developers are creating production strains for their own production use or for selling to partners (see Table 2-5 for examples of such companies). There are numerous sources for production hosts, but one specific and recently deployed example to accelerate the DBTL cycle is Vibrio natriegens, a marine bacterium with a generation time of 10 minutes under optimal conditions (Lee et al., 2016; Weinstock et al., 2016). This generation time is much shorter than other production hosts and will allow developers to rapidly scale up new products and increase their yield. Future variations of this strain, or other bacteria species, should be expected by the regulators in the coming years.
On the far end of the spectrum of chassis modifications are the GROs mentioned in the previous section, which have engineered features that allow for tightly controlled release applications. However, it seems more likely that their initial exposure to the regulatory agencies will be in contained operations where the product value can justify the novel hosts or the production of novel classes of bioproducts (for example, novel sequence-defined synthetic polymers or materials comprising many instances of nonstandard amino acids), or where intellectual-property security or biocontainment in the event of unintended release is desired to be high and enhanced barriers for genetic isolation are preferred. Further research and development of GROs will provide developers the ability to better predict how adding in sequences for the production of novel products will be received by the chassis organism and likely contribute to an accelerated DBTL cycle (Way et al., 2014). It would seem that an anticipatory opportunity for agencies and primary technology developers would lie in establishing rubrics for demonstrating the basic biosafety profiles of novel production strains. Doing this might have two benefits, one in clarifying data packages or dossiers (that is, the studies that form the knowledge base to inform a risk assessment) for entities bringing the initial submission of a novel host to a regulatory agency, but also establishing data reuse protocols or other considerations which would incentivize others to follow in adoption of new host chassis.
There appears to be growing interest in cell-free expression systems, but it is not clear at this juncture what challenges would be present at an agency review.35 Such systems can be utilized to produce research tools such as proteins for microarrays (Zarate and Galbraith, 2014) as well as RNA (J. Li et al., 2014). Other research entities are working on a number of biotechnology projects involving new materials, highly engineered biosensors (though these are likely to be used as medical devices and are outside the purview of this report), and engineered microbial consortia that will eventually leave the research setting, potentially making their way to the regulatory agencies.
An example of a biomolecular robotic sensing device funded by the U.S. government is Cyberplasm (Roberts et al., 2015). Cyberplasm integrates engineered bacteria, yeast, mammalian
___________________
35 The update to the Coordinated Framework notes that “nucleic acids produced via cell-free synthesis are used for pesticidal purposes, these products are regulated by EPA/OPP” (EOP, 2017:Table 2).
TABLE 2-5 Examples of Companies Developing Engineered Microbial Strains
| Company | Product |
|---|---|
| Aequor, Inc. | Engineered marine microbe for antifouling and antibiofilm |
| Caribou Bioscience | Precision cell engineering |
| Chain Biotechnology Ltd. | Microbial hosts (chassis) for engineering anaerobic bacteria |
| DNA 2.0 | Gene synthesis—tools provider |
| Enevolv, Inc. | Engineered microbes: bacteria, yeast, algae |
| Gen9a | Gene synthesis |
| Ginkgo Bioworks | Engineered microorganisms |
| Greenlight Biosciences | Cell-free bioprocessing technology |
| Molecular Assemblies | DNA synthesis |
| Muse Biotechnologies, Inc. | Strain engineering |
| Oligos Biotech | Engineered fungus |
| Pareto Biotechnologies | Polyketide pathways |
| Syngulon | Bacteriocin engineering |
| Synpromics | Synthetic promotors for gene expression |
| Synthetic Genomics | Advanced genomic engineering: microbial cell lines, DHA Omega-3, Astaxanthin |
| Teselagen | Combinatorial gene design and editing |
| Twist Bioscience | Gene synthesis on silicon |
| Zymergen | Strain improvement |
aIn January 2017, Ginkgo Bioworks acquired Gen9.
SOURCE: BIO (2016).
cells, and cell parts to undertake device-like functions capable of sensing and treating pathogens or chemicals within plants and animals or for other functions involving environmental sensing and remediation (for example, Ayers et al., 2010). Cell parts for detection, signaling, motion, and delivery would be integrated into a biogel matrix to mimic the movement of the sea lamprey. Another example of a combination of mechanical and living cells is organ-on-a-chip systems for research purposes (Bhatia and Ingber, 2014).
Biotechnology “Platforms”
A final class of products of biotechnology includes those that are used as platforms in the creation of other biotechnology products. New tools are always in the development pipeline and often represent the fastest-to-market products of biotechnology. Tools include products that are traditionally characterized as “wet lab,” such as DNA/RNA, enzymes, vectors, cloning kits, cells, library prep kits, and sequencing prep kits, and products that are “dry lab,” such as vector drawing software, computer-aided design software, primer calculation software, and informatics tools. These two categories continue to meld as newer approaches are published or commercialized. For instance, cloning from native hosts can be replaced by automated systems taking genomic sequence and creating libraries of genetic elements by blending DNA synthesis, assembly protocols, and automation workflows. Developers can currently access these as complete packages offered by third parties or à la carte by making a customized flow in house. Some of these services are available as cloud-based laboratory resources.
One dimension of this is the scale on which it is taking place. For instance, Twist Bioscience and Ginkgo Bioworks disclosed a commitment to transact 100 million bases of DNA, reportedly representing about 10 percent of the global synthesis market (Segran, 2015). Companies specializing in new software tools to manage such workflows are now squarely in this part of the biotechnology value chain.
Another dimension of change is the customer base of such product offerings. Such kits, software, and even hardware no longer reside exclusively in academic or industrial settings, as discussed previously in this chapter. The growing collegiate competition community, iGEM, actively consumes and contributes to a growing body of “parts” and kits. Likewise, the DIYbio community is increasingly interested in tools for genome-scale engineering, and more economical hardware and automation is being developed in maker spaces. Community laboratories are being used more and more by the DIYbio community and for science education (for instance, Building with Biology from the National Science Foundation). Amino Labs is producing a benchtop biolab for home use, and similar products can be expected to follow suit in the coming years.
SUMMARY AND CONCLUSIONS
This chapter describes some of the technological, economic, and social trends that will likely drive the development of future products of biotechnology; outlines the changes in the scope, scale, complexity, and tempo of biotechnology products; and provides a detailed account of new products that are likely to emerge in the next 5–10 years. The committee reached the following broad conclusions regarding emerging trends and products of biotechnology.
Conclusion 2-1: The U.S. bioeconomy is growing rapidly; the scope, scale, complexity, and tempo of biotechnology products are increasing.
Factors contributing to this increase include advances in basic understanding of biological processes that enable biotechnology products to be designed for a wider range of applications, more efficient and robust technologies such as genome-editing techniques, the “standardization” of bioengineering components, and the decreasing costs of genome sequencing and gene synthesis (which in turn decrease the costs of DBTL cycles and enable more startup companies to enter this area) and increasing amounts and sources of funding such as crowdfunding. Indeed, these factors have fostered a thriving DIYbio community in the United States and student competitions to design biotechnology products. With so many new actors, tools, and resources involved, future products of biotechnology are poised to permeate all aspects of human endeavor.
Conclusion 2-2: Societal factors will continue to play an important role in the public debate regarding the safe and effective use of biotechnology products.
The committee notes that there are many competing interests, risks, and benefits regarding future biotechnology products. The United States and international regulatory systems will need to achieve a balance among these competing aspects when considering how to manage the development and use of new products of biotechnology. Many parts of society have concerns over the safety and ethics of various biotechnologies, while others see prospects for biotechnology to address challenging social and environmental issues. Biotechnology products that are on the horizon are likely to generate substantial public debate. For example, gene-drive technology, for which there have already been numerous studies and reports regarding its use, is a type of technological advance that will increase the amount of public debate and for which society will have to take a balanced approach among the interested and affected parties, developers, and scientists.
Conclusion 2-3: Many future biotechnology products will be similar to existing biotechnology products, but they may be created through new processes.
Biotechnology products that have become familiar—such as insect-resistant crops and products made with bacteria in fermentation processes—will continue to be developed, though the method of genetic transformation will likely change. New forms of genetic transformation that are faster and more specific than recombinant-DNA technology, such as CRISPR, will likely allow product developers to design, build, test, and learn from experiments and product development more quickly in the next 10 years than has been the case in the last two decades. Increases in the speed and efficiency of DBTL cycles, along with an increase in the number of actors in the biotechnology space, will translate into the availability of more biotechnology products that are similar to the biotechnology products of 2016 but are transformed through processes other than recombinant DNA.
Conclusion 2-4: Some future products of biotechnology may be wholly unlike products that existed in 2016.
The increased capabilities to transform genomes afforded by advances in genomic engineering allow product developers to expand the number and kinds of modifications in future biotechnology products. The committee anticipates growth in the genetic transformation of microbes (such as yeast, algae, and bacteria) in contained systems to produce products such as chemicals and biofuels. It also expects the development of communities of microbes that are created from synthetic DNA and microbial communities formed from the combination of DNA from a number of different microbes; such communities may be designed for release in open environments to enhance nitrogen fixation by plants or for bioremediation use at contaminated sites. A much broader array of host organisms targeted for genetic transformation is also likely.
Conclusion 2-5: Novel biotechnology platforms will contribute to an increase in biotechnology products.
Biotechnology platforms—such as computational tools to improve efficiency, novel biotechnology kits to provide new actors the tools to become developers, improved capability to synthesize DNA and RNA, and increasingly automated systems—will proliferate and add to the number of and speed at which novel biotechnology products are developed.
REFERENCES
Abel, R.L. 1985. Blaming victims. Law & Social Inquiry 10(2):401–417.
Amiram, M., A.D. Haimovich, C. Fan, Y.S. Wang, H.R. Aerni, I. Ntai, D.W. Moonan, N.J. Ma, A.J. Rovner, S.H. Hong, N.L. Kelleher, A.L. Goodman, M.C. Jewett, D. Soll, J. Rinehart, and F.J. Isaacs. 2015. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nature Biotechnology 33:1272–1279.
Ayers, J., N. Rulkov, D. Knudsen, Y.-B. Kim, A. Volkovskii, and A. Selverston. 2010. Controlling underwater robots with electronic nervous systems. Applied Bionics and Biomechanics 7(1):57–67.
Baltimore, D., P. Berg, M. Botchan, D. Carroll, R.A. Charo, G. Church, J.E. Corn, G.Q. Daley, J.A. Doudna, M. Fenner, H.T. Greely, M. Jinek, G.S. Martin, E. Penhoet, J. Puck, S.H. Sternberg, J.S. Weissman, and K.R. Yamamoto. 2015. A prudent path forward for genomic engineering and germline gene modification. Science 348(6230):36–38.
Bhatia, S.N., and D.E. Ingber. 2014. Microfluidic organs-on-chips. Nature Biotechnology 32(8):760–772.
Biello, D. August 29, 2014. Ancient DNA could return passenger pigeons to the sky. Scientific American. Available at https://www.scientificamerican.com/article/ancient-dna-could-return-passenger-pigeons-to-the-sky. Accessed January 8, 2017.
BIO (Biotechnology Innovation Organization). 2016. Advancing the Biobased Economy: Renewable Chemical Biorefinery Commercialization, Progress, and Market Opportunities, 2016 and Beyond. Available at https://www.bio.org/sites/default/files/BIO_Advancing_the_Biobased_Economy_2016.pdf. Accessed October 11, 2016.
Biomass R&D Board. 2016. Federal Activities Report on the Bioeconomy. Available at http://www.biomassboard.gov/pdfs/farb_2_18_16.pdf. Accessed August 10, 2016.
Boeke, J.D., G. Church, A. Hessel, N.J. Kelley, A. Arkin, Y. Cai, R. Carlson, A. Chakravarti, V.W. Cornish, L. Holt, F.J. Isaacs, T. Kuiken, M. Lajoie, T. Lessor, J. Lunshof, M.T. Maurano, L.A. Mitchell, J. Rine, S. Rosser, N.E. Sanjana, P.A. Silver, D. Valle, H. Wang, J.C. Way, and L. Yang. 2016. The genome project–write. Science 353:126–127.
Bonny, S. 2014. Taking stock of the genetically modified seed sector worldwide: Market, stakeholders, and prices. Food Security 6(4):525–540.
Bowler, J. August 29, 2016. This new dairy alternative is made from real milk proteins—but with no cows required. Science Alert. Available at http://www.sciencealert.com/this-new-dairy-alternative-is-made-from-real-milk-proteins-but-with-no-cows-required. Accessed January 8, 2017.
Brune, K., and T. Bayer. 2012. Engineering microbial consortia to enhance biomining and bioremediation. Frontiers in Microbiology 3:203.
Bryce, E. August 26, 2016. Synthetic prawns: A bid to make ‘seafood’ that’s sustainable and slavery-free. The Guardian. Available at https://www.theguardian.com/environment/world-on-a-plate/2016/aug/26/synthetic-prawns-a-bid-to-make-seafood-thats-sustainable-and-slavery-free. Accessed January 8, 2017.
Callaway, E. 2015. Mammoth genomes hold recipe for Arctic elephants. Nature 521(7550):18–19.
Campbell, K.J., J. Beek, C.T. Eason, A.S. Glen, J. Godwin, F. Gould, N.D. Holmes, G.R. Howald, F.M. Madden, J.B. Ponder, D.W. Threadgill, A.S. Wegmann, and G.S. Baxter. 2015. The next generation of rodent eradications: Innovative technologies and tools to improve species specificity and increase their feasibility on islands. Biological Conservation 185:47–58.
Cargill. 2016. GRAS Exemption Claim for Steviol Glycosides from Saccharomyces cerevisiae Expressing Steviol Glycoside Biosynthesis Pathway. February 1. Available at http://www.fda.gov/downloads/Food/IngredientsPackagingLabeling/GRAS/NoticeInventory/ucm505151.pdf. Accessed January 8, 2017.
Carlson, D.F., C.A. Lancto, B. Zang, E.-S. Kim, M. Walton, D. Oldeshulte, C. Seabury, T.S. Sonstegard, and S.C. Fahrenkrug. 2016. Production of hornless dairy cattle from genome-edited cell lines. Nature Biotechnology 34(5):479–481.
Carlson, R. 2016. Estimating the biotech sector’s contribution to the US economy. Nature Biotechnology 34(3):247–255.
Cases, I., and V. de Lorenzo. 2005. Genetically modified organisms for the environment: Stories of success and failure and what we have learned from them. International Microbiology 8(3):213–222.
Chambers, S., R. Kitney, and P. Freemont. 2016. The Foundry: The DNA synthesis and construction foundry at Imperial College. Biochemical Society Transactions 44:687–688.
Cohen, S.N., A.C.Y. Chang, H. Boyer, and R.B. Helling. 1973. Construction of biologically functional bacterial plasmids in vitro. Proceedings of the National Academy of Sciences of the United States of America 70:3240–3244.
Cong, L., F.A. Ran, D. Cox, S. Lin, R. Barretto, N. Habib, P.D. Hsu, X. Wu, W. Jiang, L.A. Marraffini, and F. Zhang. 2013. Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823.
Cumbers, J. 2016. Future Emerging Open Release Products. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 1, Washington, DC.
Cummings, C., and J. Kuzma. 2017. Societal risk evaluation scheme (SRES): Scenario-based multi-criteria evaluation of synthetic biology applications. PLOS ONE 12:e0168564.
Cyranoski, D. September 29, 2015. Gene-edited ‘micropigs’ to be sold as pets at Chinese institute. Nature 526(7571):18.
DaCunha, C. 2016. Universal Bio Mining: Transforming Mining with Synthetic Biology. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 1, Washington, DC.
Datar, I. 2016. Small Business Perspectives from the 501(c)(3) Research Institute Advancing Cellular Agriculture. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 27, San Francisco, CA.
Davis, N. November 14, 2015. Biotech bid to take shark off the menu and cut the fin trade. The Guardian. Available at https://www.theguardian.com/environment/2015/nov/15/shark-fin-soup-lab-grown-conservation-finning. Accessed January 8, 2017.
de Lorenzo, V., P. Marliere, and R. Solé. 2016. Bioremediation at a global scale: From the test tube to planet Earth. Microbial Biotechnology 9(5):618–625.
D’Hulst, C., M. Saparauskaite, and P. Feinstein. 2012. Generating a biosensor for the detection of landmines. Program Poster No. 815.09/FFF73. 2012 Neuroscience Meeting Planner. New Orleans, LA: Society for Neuroscience. Available at http://www.abstractsonline.com/Plan/ViewAbstract.aspx?sKey=5d17d281-d287-4859-8092-77a84caf35c9&cKey=7f76a7f8-fee2-4d64-925d-b88be0b87d90&mKey=%7b70007181-01C9-4DE9-A0A2-EEBFA14CD9F1%7d. Accessed December 18, 2016.
Doudna, J.A., and E. Charpentier. 2014. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346(6213):1258096.
Douw, K., H. Vondeling, D. Eskildsen, and S. Simpson. 2003. Use of the Internet in scanning the horizon for new and emerging health technologies: A survey of agencies involved in horizon scanning. Journal of Medical Internet Research 5(1):e6.
Duan, H., S. Umar, R. Xiong, and J. Chen. 2011. New strategy for expression of recombinant hydroxylated human-derived gelatin in Pichia pastoris KM71. Journal of Agricultural and Food Chemistry 59(13):7127–7134.
EC (European Commission). 2012. The European Bioeconomy in 2030: Delivering Sustainable Growth by Addressing the Grand Societal Challenges. Available at http://www.epsoweb.org/file/560. Accessed August 10, 2016.
Eisenstein, M. 2016. Living factories of the future. Nature 531(7594):401–403.
El-Chichakli, B., J. von Braun, C. Lang, D. Barben, and J. Philp. 2016. Policy: Five cornerstones of a global bioeconomy. Nature 535(7611):221–223.
EOP (Executive Office of the President). 2017. Modernizing the Regulatory System for Biotechnology Products: An Update to the Coordinated Framework for the Regulation of Biotechnology. Available at https://obamawhitehouse.archives.gov/sites/default/files/microsites/ostp/2017_coordinated_framework_update.pdf. Accessed January 30, 2017.
EPA (U.S. Environmental Protection Agency). 2015. Biotechnology Algae Project. August 5. Available at https://www.epa.gov/sites/production/files/2015-09/documents/biotechnology_algae_project.pdf. Accessed September 14, 2016.
Evans, A. 2016. TAXA. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 27, San Francisco, CA.
EY. 2016. Beyond Borders 2016: Biotech Financing. Bountiful Harvest Leaves Biotech Well Prepared for Financial Winter. Available at http://www.ey.com/Publication/vwLUAssets/ey-beyond-borders-2016-biotech-financing/$FILE/ey-beyond-borders-2016-biotech-financing.pdf. Accessed September 27, 2016.
FDA (U.S. Food and Drug Administration). 2016. Agency Response Letter GRAS Notice No. GRN 000626. May 27. Available at http://www.fda.gov/Food/IngredientsPackagingLabeling/GRAS/NoticeInventory/ucm512620.htm. Accessed January 8, 2017.
Finucane, M.L., P. Slovic, C.K. Mertz, J. Flynn, and T.A. Satterfield. 2000. Gender, race, and perceived risk: The “white male” effect. Health, Risk & Society 2:159–172.
Fischer, K., E. Ekener-Petersen, L. Rydhmer, and K.E. Björnberg. 2015. Social impacts of GM crops in agriculture: A systematic literature review. Sustainability 7(7):8598–8620.
Fischhoff, B., P. Slovic, S. Lichtenstein, S. Read, and B. Combs. 1978. How safe is safe enough? A psychometric study of attitudes towards technological risks and benefits. Policy Sciences 9(2):127–152.
Formas (Swedish Research Council for Environment, Agricultural Sciences and Spatial Planning). 2012. Swedish Research and Innovation Strategy for a Bio-based Economy. Available at http://www.formas.se/PageFiles/5074/Strategy_Biobased_Ekonomy_hela.pdf. Accessed August 10, 2016.
Fredrickson, J.K. 2015. Ecological communities by design. Science 348(6242):1425–1427.
Gaj, T., C.A. Gerbach, and C.F. Barbas. 2013. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends in Biotechnology 31:397–405.
GBC (German Bioeconomy Council). 2016. Bioeconomy Policy (Part I): Synopsis and Analysis of Strategies in the G7. Available at http://bioeconomia.agripa.org/download-doc/64046. Accessed August 10, 2016.
Gertz, E. August 6, 2014. Can biohackers succeed at making “real vegan cheese”? Popular Science. Available at http://www.popsci.com/article/science/can-biohackers-succeed-making-real-vegan-cheese. Accessed January 8, 2017.
Gibbons, D.W., D.A. Bohan, P. Rothery, R.C. Stuart, A.J. Haughton, R.J. Scott, J.D. Wilson, J.N. Perry, S.J. Clark, R.J.G. Dawson, and L.G. Firbank. 2006. Weed seed resources for birds in fields with contrasting conventional and genetically modified herbicide-tolerant crops. Proceedings of the Royal Society B 273(1596):1921–1928.
Gibson, D.G., J.I. Glass, C. Lartigue, V.N. Noskov, R.-Y. Chuang, M.A. Algire, G.A. Benders, M.G. Montague, L. Ma, M.M. Moodie, C. Merryman, S. Vashee, R. Krishnakumar, N. Assad-Garcia, C. Andrews-Pfannkoch, E.A. Denisova, L. Young, Z.-Q. Qi, T.H. Segall-Shapiro, C.H. Calvey, P.P. Parmar, C.A. Hutchison, H.O. Smith, and J.C. Venter. 2010. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329(5987):52–56.
Gill, R.T., A.L. Halweg-Edwards, A. Clauset, and S.F. Way. 2016. Synthesis aided design: The biological design-build-test engineering paradigm? Biotechnology and Bioengineering 113(1):7–10.
Gocal, G. 2015. Non-transgenic trait development in crop plants using oligo-directed mutagenesis: Cibus’ Rapid Trait Development System. Pp. 97–106 in NABC Report 26 New DNA-Editing Approaches: Methods, Applications, & Policy for Agriculture, A. Eaglesham and R.W.F. Hardy, eds. Ithaca, NY: North American Agricultural Biotechnology Council.
Grushkin, D., T. Kuiken, and P. Millet. 2014. Seven Myths & Realities about Do-It-Yourself Biology. Washington, DC: Woodrow Wilson Center for International Scholars.
Hansen, E.H., B.L. Møller, G.R. Kock, C.M. Bünner, C. Kristensen, O.R. Jensen, F.T. Okkels, C.E. Olsen, M.S. Motawia, and J. Hansen. 2009. De novo biosynthesis of vanillin in fission yeast (Schizosaccharomyces pombe) and baker’s yeast (Saccharomyces cerevisiae). Applied and Environmental Microbiology 75(9):2765–2774.
Hansen, J., L. Holm, L. Frewer, P. Robinson, and P. Sandøe. 2003. Beyond the knowledge deficit: Recent research into lay and expert attitudes to food risks. Appetite 41(2):111–121.
Harvey-Samuel, T., N.I. Morrison, A.S. Walker, T. Marubbi, J. Yao, H.L. Collins, and L. Alphey. 2015. Pest control and resistance management through release of insects carrying a male-selecting transgene. BMC Biology 13(1):49.
Haun, W., A. Coffman, B.M. Clasen, Z.L. Demorest, A. Lowy, E. Ray, A. Retterath, T. Stoddard, A. Juillerat, F. Cedrone, L. Mathis, D.F. Voytas, and F. Zhang. 2014. Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family. Plant Biotechnology Journal 12(7):934–940.
Hayden, E.C. 2015. Tech investors bet on synthetic biology. Nature 527(7576):19.
Hutchison, C.A., III, R.Y. Chuang, V.N. Noskov, N. Assad-Garcia, T.J. Deerinck, M.H. Ellisman, J. Gill, K. Kannan, B.J. Karas, L. Ma, J.F. Pelletier, Z.Q. Qi, R.A. Richter, E.A. Strychalski, L. Sun, Y. Suzuki, B. Tsvetanova, K.S. Wise, H.O. Smith, J.I. Glass, C. Merryman, D.G. Gibson, and J.C. Venter. 2016. Design and synthesis of a minimal bacterial genome. Science 351(6280):aad6253.
IAASTD (International Assessment of Agricultural Knowledge, Science and Technology for Development). 2009. Agriculture at a Crossroads: Global Report. Washington, DC: Island Press.
iGEM (The International Genetically Engineered Machine). 2012. Modeling Microbial Consortia: The Auxotroph Approach, University of British Columbia. Available at http://2012.igem.org/Team:British_Columbia/Consortia. Accessed October 11, 2016.
iGEM. 2013. Engineering a Synthetic Microbial Consortium, Technische Universität Braunschweig. Available at http://2013.igem.org/Team:Braunschweig. Accessed October 11, 2016.
iGEM. 2015a. Synenergene, Team Amsterdam. Available at https://www.synenergene.eu/content/igem-2015-team-amsterdam. Accessed October 11, 2016.
iGEM. 2015b. Massachusetts Institute of Technology. Human Practices: State of the Art in Consolidated Bioprocessing. Available at http://2015.igem.org/Team:MIT/CBP. Accessed October 11, 2016.
Isaacs, F.J., P.A. Carr, H.H. Wang, M.J. Lajoie, B. Sterling, L. Kraal, A.C. Tolonen, T.A. Gianoulis, D.B. Goodman, N.B. Reappas, C.J. Emig, D. Bang, S.J. Hwang, M.C. Jewett, J.M. Jacobson, and G.M. Church. 2011. Precise manipulation of chromosomes in vivo enables genome-wide codon replacement. Science 333(6040):348–353.
Jacobsen, S.E., M. Sørensen, S.M. Pedersen, and J. Weiner. 2013. Feeding the world: Genetically modified crops versus agricultural biodiversity. Agronomy for Sustainable Development 33(4):651–662.
Jia, X., C. Liu, H. Song, M. Ding, J. Du, Q. Ma, and Y. Yuan. 2016. Design, analysis and application of synthetic microbial consortia. Synthetic and Systems Biotechnology 1(2):109–117.
Jinek, M., A. East, A. Cheng, S. Lin, E. Ma, and J. Doudna. 2013. RNA-programmed genome editing in human cells. Elife 2:e00471.
Juhas, M., and J.W. Ajioka. 2017. High molecular weight DNA assembly in vivo for synthetic biology applications. Critical Reviews in Biotechnology 37(3):277–286.
Kahan, D.M. 2012. Cultural cognition as a conception of the cultural theory of risk. Pp. 725–759 in Handbook of Risk Theory, S. Roeser, R. Hillerbrand, P. Sandin, and M. Peterson, eds. Dordrecht, Netherlands: Springer.
Kahan, D.M., D. Braman, J. Gastil, P. Slovic, and C. Mertz. 2007. Culture and identity-protective cognition: Explaining the white-male effect in risk perception. Journal of Empirical Legal Studies 4:465–505.
Kahan, D.M., E. Peters, M. Wittlin, P. Slovic, L.L. Ouellette, D. Braman, and G. Mandel. 2012. The polarizing impact of science literacy and numeracy on perceived climate change risks. Nature Climate Change 2(10):732–735.
Killiny, N., S. Hajeri, S. Tiwari, S. Gowda, and L.L. Stelinski. 2014. Double-stranded RNA uptake through topical application, mediates silencing of five CYP4 genes and suppresses insecticide resistance in Diaphorina citri. PLOS ONE 9(10):e110536.
Klein, T.M., E.D. Wolf, R. Wu, and J.C. Sanford. 1987. High velocity microprojectiles for delivering nucleic acids into living cells. Nature 327(6117):70–73.
Kosuri, S., and G.M. Church. 2014. Last-scale de novo DNA synthesis: Technologies and applications. Nature Methods 11:499–507.
Kotula, J.W., S.J. Kerns, L.A. Shaket, L. Siraj, J.J. Collins, J.C. Way, and P.A. Silver. 2014. Programmable bacteria detect and record an environmental signal in the mammalian gut. Proceedings of the National Academy of Sciences of the United States of America 111:4838–4843.
Kovalchuk, I., and O. Kovalchuk. 2008. Transgenic plants as sensors of environmental pollution genotoxicity. Sensors 8(3):1539–1558.
Kovalchuk, I., O. Kovalchuk, A. Arkhipov, and B. Hohn. 1998. Transgenic plants are sensitive bioindicators of nuclear pollution caused by the Chernobyl accident. Nature Biotechnology 16(11):1054–1059.
Kromdijk, J., K. Głowacka, L. Leonelli, S.T. Gabilly, M. Iwai, K.K. Niyogi, and S.P. Long. 2016. Improving photosynthesis and crop productivity by accelerating recovery from photoprotection. Science 354:857–861.
Kuiken, T. 2016. Our Collective Biology: Enabling Public Science to Build an Ecosystem of Makers in Biology. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 1, Washington, DC.
Kwong, W.K., and N.A. Moran. 2016. Gut microbial communities of social bees. Nature Reviews Microbiology 14(6): 374–384.
Lafrance, A. September 24, 2015. Bioluminescence in a bottle. The Atlantic. Available at http://www.theatlantic.com/technology/archive/2015/09/bioluminescence-in-a-bottle/407152/. Accessed January 8, 2017.
Lajoie, M.J., A.J. Rovner, D.B. Goodman, H.-R. Aerni, A.D. Haimovich, G. Kuznetsov, J.A. Mercer, H.H. Wang, P.A. Carr, J.A. Mosberg, N. Rohland, P.G. Schultz, J.M. Jacobson, J. Rinehart, G.M. Church, and F.J. Isaacs. 2013. Genomically recoded organisms expand biological functions. Science 342(6156):357–360.
Langin, K. July 18, 2014. Genetic engineering to the rescue against invasive species? National Geographic. Available at http://news.nationalgeographic.com/news/2014/07/140717-gene-drives-invasive-species-insects-disease-science-environment/. Accessed January 8, 2017.
Ledford, H. 2016. UK bioethicists eye designer babies and CRISPR cows. Nature 538(7623):17.
Lee, H.H., N. Ostrov, B.G. Wong, M.A. Gold, A. Khalil, and G.M. Church. 2016. Vibrio natriegens, a new genomic powerhouse. bioRxiv:058487.
Li, F., Y. Li, H. Liu, H.H. Zhang, C.X. Liu, X.J. Zhang, H.W. Dou, W.X. Yang, and Y.T. Du. 2014. Production of GHR double-allelic knockout Bama pig by TALENs and handmade cloning. Hereditas 36(9):903–911.
Li, J., L. Gu, J. Aach, and G.M. Church. 2014. Improved cell-free RNA and protein synthesis system. PLOS ONE 9(9):e106232.
Liew, F., M.E. Martin, R.C. Tappel, B.D. Heijstra, C. Mihalcea, and M. Köpke. 2016. Gas fermentation—A flexible platform for commercial scale production of low-carbon-fuels and chemicals from waste and renewable feedstocks. Frontiers in Microbiology 7:694.
Lindemann, S.R., H.C. Bernstein, H.-S. Song, J.K. Fredrickson, M.W. Fields, W. Shou, D.R. Johnson, and A.S. Beliaev. 2016. Engineering microbial consortia for controllable outputs. The ISME Journal 10:2077–2084.
Lombardo, T. October 11, 2015. Bioluminescent lamp: Oddity, novelty, engineering challenge. Available at http://www.engineering.com/ElectronicsDesign/ElectronicsDesignArticles/ArticleID/10799/Bioluminescent-Lamp-Oddity-Novelty-Engineering-Challenge.aspx. Accessed January 8, 2017.
Luzar, C. August 3, 2013. Kickstarter bans GMOs in wake of glowing plant campaign. Crowdfund Insider. Available at http://www.crowdfundinsider.com/2013/08/20031-kickstarter-bans-gmos-in-wake-of-glowing-plant-fiasco. Accessed September 27, 2016.
Ma, N.J., and F.J. Isaacs. 2016. Genomic recoding broadly obstructs the propagation of horizontally transferred genetic elements. Cell Systems 3(2):199–207.
Mali, P., L. Yang, K.M. Esvelt, J. Aach, M. Guell, J.E. DiCarlo, J.E. Norville, and G.M. Church. 2013. RNA-guided human genome engineering via Cas9. Science 339(6121):823–826.
Mandell, D.J., M.J. Lajoie, M.T. Lee, R. Takeuchi, G. Kuznetsov, J.E. Norville, C.J. Gregg, B.L. Stoddard, and G.M. Church. 2015. Biocontainment of genetically modified organisms by synthetic protein design. Nature 518(7537):55–60.
Marcellin, F. February, 26, 2016. Glow-in-the-dark bacterial lights could illuminate shop windows. New Scientist. Available at https://www.newscientist.com/article/2078921-glow-in-the-dark-bacterial-lights-could-illuminate-shop-windows. Accessed January 8, 2017.
Marris, C., I.H. Langford, and T. O’Riordan. 1998. A quantitative test of the cultural theory of risk perceptions: Comparison with the psychometric paradigm. Risk Analysis 18(5):635–647.
McCright, A.M., and C. Xiao. 2014. Gender and environmental concern: Insights from recent work and for future research. Society & Natural Resources 27(10):1109–1113.
Miller, F. December 3, 2014. Arkansas releases first Roundup Ready soybean. University of Arkansas News. Available at http://arkansasagnews.uark.edu/8273.htm. Accessed November 27, 2016.
Mizokami, K. November 21, 2016. The Pentagon uses plant DNA to catch counterfeit parts. Popular Mechanics. Available at http://www.popularmechanics.com/military/research/a23988/plant-dna-pentagon-counterfeit/. Accessed January 8, 2017.
Munnelly, K. 2016. Advancements in DNA Construction Tools and Technology. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 1, Washington, DC.
Nagare, P., B.A. Aglave, and M.O. Lokhande. 2009. Genetically engineered zebra fish—fluorescent beauties with practical applications. The Asian Journal of Animal Science 4(1):126–129.
Napolitano, M.G., M. Landon, C.J. Gregg, M.J. Lajoie, L. Govindarajan, J.A. Mosberg, G. Kuznetsov, D.B. Goodman, O. Vargas-Rodriguez, F.J. Isaacs, D. Söll, and G.M. Church. 2016. Emergent rules for codon choice elucidated by editing rare arginine codons in Escherichia coli. Proceedings of the National Academy of Sciences of the United States of America 113:E5588–E5597.
NASEM (National Academies of Sciences, Engineering, and Medicine). 2016a. Gene Drives on the Horizon: Advancing Science, Navigating Uncertainty, and Aligning Research with Public Values. Washington, DC: The National Academies Press.
NASEM. 2016b. Genetically Engineered Crops: Experiences and Prospects. Washington, DC: The National Academies Press.
OECD (Organisation for Economic Co-operation and Development). 2015. Building a Bioeconomy: How Nations Approach Capacity Building in Industrial Biotechnology. Paris: OECD.
OSTP (Office of Science and Technology Policy). 2012. National Bioeconomy Blueprint. Washington, DC: The White House.
Ostrov, N., M. Landon, M. Guell, G. Kuznetsov, J. Teramoto, N. Cervantes, M. Zhou, K. Singh, M.G. Napolitano, and M. Moosburner. 2016. Design, synthesis, and testing toward a 57-codon genome. Science 353(6301):819–822.
Oxitec. 2016. Draft Environment Assessment for Investigational Use of Aedes aegypti OX513A. February. Oxfordshire, UK: Oxitec, Ltd. Available at http://www.fda.gov/downloads/AnimalVeterinary/DevelopmentApprovalProcess/GeneticEngineering/GeneticallyEngineeredAnimals/UCM487377.pdf. Accessed September 30, 2016.
Peck, B. 2016. Reimagine SequenceSpace. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 1, Washington, DC.
Petzold, C.J., L.J.G. Chan, M. Nhan, and P.D. Adams. 2015. Analytics for metabolic engineering. Frontiers in Bioengineering and Biotechnology 3:135.
Potter, K.M., and B.L. Conkling, eds. 2016. Forest Health Monitoring: National Status, Trends, and Analysis 2015. Ashe-ville, NC: U.S. Forest Service–Southern Research Station.
Radakovits, R., R.E. Jinkerson, A. Darzins, and M.C. Posewitz. 2010. Genetic engineering of algae for enhanced biofuel production. Eukaryotic Cell 9(4):486–501.
Reardon, S. 2016. Welcome to the CRISPR zoo. Nature 431(7593):160–163.
Reed, T. 2016. Intrexon. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 1, Washington, DC.
Regalado, A. July 15, 2016. Why Kickstarter’s glowing plant left backers in the dark. MIT Technology Review. Available at https://www.technologyreview.com/s/601884/why-kickstarters-glowing-plant-left-backers-in-the-dark. Accessed September 27, 2016.
Roberts, J.P., S. Stauffer, C. Cummings, and J. Kuzma. 2015. Synthetic Biology Governance: Delphi Study Workshop Report. GES Center Report No. 2015.2. Available at https://research.ncsu.edu/ges/files/2014/04/Sloan-Workshop-Report-final-ss-081315-1.pdf. Accessed October 10, 2016.
Rohrmann, B., and O. Renn. 2000. Risk perception research. Pp. 11–53 in Cross-Cultural Risk Perception: A Survey of Empirical Studies, O. Renn and B. Rohrmann, eds. New York: Springer.
Rovner, A.J., A.D. Haimovich, S.R. Katz, Z. Li, M.W. Grome, B.M. Gassaway, M. Amiram, J.R. Patel, R.R. Gallagher, J. Rinehart, and F.J. Isaacs. 2015. Recoded organisms engineered to depend on synthetic amino acids. Nature 518(7537):89–93.
Samin, G., M. Pavlova, M.I. Arif, C.P. Postema, J. Damborsky, and D.B. Janssen. 2014. A Pseudomonas putida strain genetically engineered for 1,2,3-trichloropropane bioremediation. Applied and Environmental Microbiology 80(17): 5467–5476.
San Miguel, K., and J.G. Scott. 2016. The next generation of insecticides: dsRNA is stable as a foliar-applied insecticide. Pest Management Science 72(4):801–809.
Schippers, A., S. Hedrich, J. Vasters, M. Drobe, W. Sand, and S. Willscher. 2014. Biomining: Metal recovery from ores with microorganisms. Advances in Biochemical Engineering/Biotechnology 141:1–47.
SEC (U.S. Securities and Exchange Commission). 2016. Investor Bulletin: Crowdfunding for Investors. February 16. Available at https://www.sec.gov/oiea/investor-alerts-bulletins/ib_crowdfunding-.html. Accessed September 27, 2016.
Segran, E. November 4, 2015. Twist Bioscience inks deal to sell 100 million base pairs of synthetic DNA. Fast Company. Online. Available at https://www.fastcompany.com/3053065/most-creative-people/twist-bioscience-inks-deal-to-sell-100-million-pairs-of-synthetic-dna. Accessed October 11, 2016.
Seltenrich, N. July 23, 2015. Improving the work of silkworms and spiders, with yeast. Berkeley Engineering. Available at http://engineering.berkeley.edu/2015/07/improving-work-silkworms-and-spiders-yeast. Accessed January 8, 2017.
Sewalt, V. 2016. Managing Risks Associated with Biotechnology Used in Containment. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 27, San Francisco, CA.
Shapiro, B. 2015. How to Clone a Mammoth: The Science of De-Extinction. Princeton, NJ: Princeton University Press.
Shi, J., H. Gao, H. Wang, H.R. Lafitte, R.L. Archibald, M. Yang, S.M. Hakimi, H. Mo, and J.E. Habben. 2017. ARGOS8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnology Journal 15(2):207–216.
Shigeta, R. 2016. Indie Bio: New Opportunities, a New Generation of Biotech Entrepreneurs. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 27, San Francisco, CA.
Shiva, V., D. Barker, and C. Lockhart. 2011. The GMO Emperor Has No Clothes: A Global Citizens Report on the State of GMOs, Synthesis Report. Florence: Navdanya International.
Shontell, A. June 28, 2016. A Brooklyn startup that’s armed with $40 million is growing real leather in a lab without hurting a single animal. Business Insider. Available at http://www.businessinsider.com/modern-meadow-lab-grown-leather-2016-6. Accessed January 8, 2017.
Slovic, P. 1987. Perception of risk. Science 236:280–290.
Smanski, M.J., S. Bhatia, D. Zhao, Y. Park, L.B.A. Woodruff, G. Giannoukos, D. Ciulla, M. Busby, J. Calderon, R. Nicol, D.B. Gordon, D. Densmore, and C.A. Voigt. 2014. Functional optimization of gene clusters by combinatorial design and assembly. Nature Biotechnology 32:1241–1249.
Stanton, B. 2016. Ginkgo Bioworks. Presentation to the National Academies of Sciences, Engineering, and Medicine Committee on Future Biotechnology Products and Opportunities to Enhance Capabilities of the Biotechnology Regulatory System, June 1, Washington, DC.
Sun, Z.Z., V. Noireaux, and R.M. Murray. 2014. Accelerating the design-build-test cycle of synthetic biological circuits in E. coli using S30 TX-TL cell-free systems, linear DNA, and modular assembly. General submission in the Conference Proceedings of the Synthetic Biology: Engineering, Evolution & Design Conference, July 14–17, Manhattan Beach, CA. Available at http://www3.aiche.org/proceedings/Abstract.aspx?PaperID=392699. Accessed September 27, 2016.
Taylor, N., E. Gaitán-Solís, T. Moll, B. Trauterman, T. Jones, A. Pranjal, C. Trembley, V. Abernathy, D. Corbin, and C. Fauquet. 2012. A high-throughput platform for the production and analysis of transgenic cassava (Manihot esculenta) plants. Tropical Plant Biology 5(1):127–139.
Tennessen, J.A., A. Théron, M. Marine, J.-Y. Yeh, A. Rognon, and M.S. Blouin. 2015. Hyperdiverse gene cluster in snail host conveys resistance to human schistosome parasites. PLOS Genetics 11(3):e1005067.
Thomke, S.H. 2003. Experimentation Matters: Unlocking the Potential of New Technologies for Innovation. Boston: Harvard Business Press.
Thompson, P. 2011. Understanding and coping with social risk in emerging technology risk assessment. Pp. 1–16 in Biotechnology and Nanotechnology Risk Assessment: Minding and Managing the Potential Threats around Us, S. Ripp and T.B. Henry, eds. Washington, DC: American Chemical Society.
Tokareva, O., V.A. Michalczechen-Lacerda, E.L. Rech, and D.L. Kaplan. 2013. Recombinant DNA production of spider silk proteins. Microbial Biotechnology 6(6):651–663.
Trögl, J., A. Chauhan, S. Ripp, A.C. Layton, G. Kuncová, and G.S. Sayler. 2012. Pseudomonas fluorescens HK44: Lessons learned from a model whole-cell bioreporter with a broad application history. Sensors 12:1544–1571.
USDA–APHIS (U.S. Department of Agriculture–Animal and Plant Health Inspection Service). 2017. Regulatory Impact Analysis & Initial Regulatory Flexibility Analysis, Proposed Rule, APHIS 2015-0057, RIN 0579-AE15; Importation, Interstate Movement, and Environmental Release of Organisms Produced through Genetic Engineering (7 CFR part 340). Available at https://www.regulations.gov/document?D=APHIS-2015-0057-0002. Accessed March 31, 2017.
USDA–NASS (U.S. Department of Agriculture–National Agricultural Statistics Service). 2009. Hawaii Papayas. Available at https://www.nass.usda.gov/Statistics_by_State/Hawaii/Publications/Fruits_and_Nuts/papaya.pdf. Accessed December 3, 2016.
Van Beilen, J.B., and Y. Poirier. 2008. Production of renewable polymers from crop plants. The Plant Journal 54(4):684–701.
van der Linden, S. 2016. A conceptual critique of the cultural cognition thesis. Science Communication 38(1):128–138.
Vidal, J. October 19, 2011. GM foods: A “biotech revolution”? The Guardian. Available at https://www.theguardian.com/environment/2011/oct/19/gm-foods-a-biotech-revolution. Accessed January 30, 2017.
Voytas, D.F., and C. Gao. 2014. Precision genome engineering and agriculture: Opportunities and regulatory challenges. PLOS Biology 12:e1001877.
Waltz, E. 2016. Gene-edited CRISPR mushroom escapes US regulation. Nature 532(7599):293.
Wang, H.H., F. J. Isaacs, P.A. Carr, Z.Z. Sun, G. Xu, C.R. Forest, and G.M. Church. 2009. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460(7257):894–898.
Wang, Y., X. Cheng, Q. Shan, Y. Zhang, J. Liu, C. Gao, and J.L. Qiu. 2014. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nature Biotechnology 32:947–951.
Way, J.C., J.J. Collins, J.D. Keasling, and P.A. Silver. 2014. Integrating biological redesign: Where synthetic biology came from and where it needs to go. Cell 157(1):151–161.
Weinstock, M.T., E.D. Hesek, C.M. Wilson, and D.G. Gibson. 2016. Vibrio natriegens as a fast-growing host for molecular biology. Nature Methods 13:849–851.
Wheelwright, S.C., and K.B. Clark. 1994. Accelerating the design-build-test cycle for effective product development. International Marketing Review 11:32–46.
Wozniak, C.A., G. McClung, J. Gagliardi, M. Segal, and K. Matthews. 2012. Regulation of genetically engineered microorganisms under FIFRA, FFDCA and TSCA. Pp. 57–94 in Regulation of Agricultural Biotechnology: The United States and Canada, C. Wozniak and A. McHughen, eds. Heidelberg, Germany: Springer.
Xiong, W., J.A. Morgan, J. Ungerer, B. Wang, P.-C. Maness, and J. Yu. 2015. The plasticity of cyanobacterial metabolism supports direct CO2 conversion to ethylene. Nature Plants 1:15053.
Xu, T., D.M. Close, G.S. Sayler, and S. Ripp. 2013. Genetically modified whole-cell bioreporters for environmental assessment. Ecological Indicators 28:125–141.
Yue, C., S. Zhao, C. Cummings, and J. Kuzma. 2015. Investigating factors influencing consumer willingness to buy GM food and nano-food. Journal of Nanoparticle Research 17(7):283.
Zarate, X., and D.W. Galbraith. 2014. A cell-free expression platform for production of protein microarrays. Methods in Molecular Biology 1118:297–307.
Zetsche, B., J.S. Gootenberg, O.O. Abudayyeh, I.M. Slaymaker, K.S. Makarova, P. Essletzbichler, S.E. Volz, J. Joung, J. van der Oost, A. Regev, and E.V. Koonin. 2015. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163(3):759–771.
Zhang, B., A.D. Oakes, A.E. Newhouse, K.M. Baier, C.A. Maynard, and W.A. Powell. 2013. A threshold level of oxalate oxidase transgene expression reduces Cryphonectria parasitica-induced necrosis in a transgenic American chestnut (Castanea dentata) leaf bioassay. Transgenic Research 22(5):973–982.
Zhang, L., R. Routsong, Q. Nguyen, E.L. Rylott, N.C. Bruce, and S.E. Strand. 2017. Expression in grasses of multiple trans-genes for degradation of munitions compounds on live fire training ranges. Plant Biotechnology Journal 15(5):624–633.
Zhou, H., B. Vonk, J.A. Roubos, R.A.L. Bovenberg, and C.A. Voigt. 2015. Algorithmic co-optimization of genetic constructs and growth conditions: Application to 6-ACA, a potential nylon-6 precursor. Nucleic Acids Research 43(2):10560–10570.
Zhu, H., L. Hu, J. Liu, H. Chen, C. Cui, Y. Song, Y. Jin, and Y. Zhang. 2016. Generation of β-lactoglobulin-modified transgenic goats by homologous recombination. FEBS Journal 283(24):4600–4613.
Zou, Q., X. Wang, Y. Liu, Z. Ouyang, H. Long, S. Wei, J. Xin, B. Zhao, S. Lai, J. Shen, Q. Ni, H. Yang, H. Zhong, L. Li, M. Hu, Q. Zhang, Z. Zhou, J. He, Q. Yan, N. Fan, Y. Zhao, Z. Liu, L. Guo, J. Huang, G. Zhang, J. Ying, L. Lai, and X. Gao. 2015. Generation of gene-target dogs using CRISPR/Cas9 system. Journal of Molecular Cell Biology 7(6):580–583.
This page intentionally left blank.








































