C
Case Studies on Site-Specific Assessments
As discussed in Chapter 5, understanding the risk associated with a spill or a hazardous waste site requires identifying and quantifying the chemicals present, characterizing the chemical toxicity, and estimating the mixture toxicity and associated risk. This appendix provides a case study related to each element. The first case study describes approaches for refining exposure estimates for known chemicals at a hypothetical site and approaches for identifying the uncharacterized chemicals at the site. The second addresses the generation of toxicity data and exposure information on a data-poor chemical after its accidental release. The third explores a biological read-across approach for assessing mixtures at a hypothetical site.
IDENTIFYING CHEMICALS AT A SITE
For this case study, the setting is a large, historically contaminated site that comprises land and surface water near a major population center (for example, Love Canal, the Portland Harbor, or the Houston Ship Channel). Recent site characterization has produced an extensive set of environmental monitoring data for air, water, and soil at the site. The data cover multiple times and are geographically distributed throughout the site. Biomonitoring data are available from serum, urine, and hair in a representative sample of people who live and work in the area surrounding the site. The biomonitoring data are geographically distributed but in some cases limited to single times.
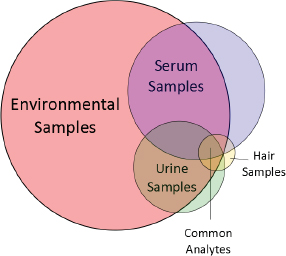
Targeted analytical chemistry produced concentration data on about 50 toxicologically well-characterized chemicals in environmental media and human blood, urine, and hair (see Table C-1). The chemicals represent four major chemical classes: polycyclic aromatic hydrocarbons, industrial chemicals and solvents, plasticizers, and pesticides. Information on metabolism and pharmacokinetics of many of the chemicals in rodents and humans is available. Assessments of external exposure of the population around the site (children, adults, and senior adults) to each chemical by the oral, dermal, and inhalation routes, where appropriate, have been conducted. Nontargeted analyses of the same environmental and biomonitoring samples revealed 5,000 unidentified substances in the environmental media, 3,000 in serum, 2,000 in urine, and 800 in hair; 300 of the unidentified analytes are common to the environmental media and all biomonitoring samples (see Figure C-1).
For this case study, the tasks become refining exposure assessment of the known chemicals, translating the external-exposure predictions into internal-exposure predictions, and identifying the unknown chemicals at the site. The following sections explore those various tasks.
Assessment of Known Chemicals and Chemical Mixtures
The initial step in this case study would be to assemble existing exposure data on the identified (known) chemicals and refine their exposure estimates for testing. The relative composition, variability, and concentration ranges of the chemicals in the various media would be assessed and quantified, taking into account the exposure routes of interest. For example, testing designed to evaluate risks associated with dermal exposures might focus on concentrations of chemicals in soil, water, and air that would come into contact with skin. Similarly, mixtures that should be evaluated for inhalation toxicity in portal-of-entry tissues (lung tissue) might best be defined by air concentrations of mixture components. Alternatively, oral exposures for toxicity testing could initially be defined by the composition and concentrations of components of soil and water or other media that might be ingested and absorbed in sufficient amounts to influence total exposure.
Once exposures have been defined, the task is to translate exposures from external measures to internal predictions to appropriate concentrations for in vitro testing with pharmacokinetic models or measurements obtained from biomonitoring. The accuracy of the model estimates will be determined partly by the amount of information available on absorption, distribution, metabolism, and excretion (ADME) processes. Cheminformatic and high-throughput systems can provide information on, for example, metabolism by hepatocytes, absorption by caco-2 cells, and binding to plasma proteins that could be used to estimate pharmacokinetic parameters (Wetmore
TABLE C-1 Site-Specific Chemicals Identified by Targeted Chemistry Analysis
| Class | Ranka | Chemical Name |
|---|---|---|
| Polycyclic aromatic hydrocarbons | 10 | Benzo(B)fluoranthene |
| 38 | Benzo(A)anthracene | |
| 80 | Naphthalene | |
| 138 | Fluoranthene | |
| 168 | Acenaphthene | |
| 185 | Dibenzofuran | |
| 255 | Pyrene | |
| High-production-volume industrial chemicals | 30 | Benzidine |
| 54 | Pentachlorophenol | |
| 84 | 2,4,6-Trichlorophenol | |
| 98 | 2,4-Dinitrotoluene | |
| 101 | 4,6-Dinitro-o-cresol | |
| 137 | 1,2,3-Trichlorobenzene | |
| 142 | 2,4,5-Trichlorophenol | |
| 172 | Cresol, para- | |
| 181 | Phenol | |
| 195 | Cresol, ortho- | |
| 206 | n-Nitrosodiphenylamine | |
| 260 | 2,6-Dinitrotoluene | |
| Plasticizers | 58 | Di-n-butyl phthalate |
| 77 | Di(2-ethylhexyl)phthalate | |
| 266 | Bis(2-ethylhexyl)adipate | |
| Pesticides | 13 | DDT, P,P’- |
| 18 | Dieldrin | |
| 25 | Aldrin | |
| 26 | DDD, P,P’- | |
| 28 | Heptachlor | |
| 34 | γ-Hexachlorocyclohexane | |
| 37 | Disulfuron | |
| 40 | Endrin | |
| 41 | Diazinon | |
| 44 | Endosulfan | |
| 47 | Heptachlor epoxide | |
| 53 | DDT, O,P’- | |
| 55 | Methoxychlor | |
| 65 | Chlorpyriphos | |
| 89 | 2,4-Dinitrophenol | |
| 99 | Ethion | |
| 103 | Dimethylarsinic acid | |
| 131 | Azinphos-methyl | |
| 144 | Dicofol | |
| 148 | Parathion | |
| 155 | Trifluralin | |
| 166 | Phorate | |
| 200 | Ethoprop | |
| 232 | Dimethoate | |
| 244 | 2,4-D Acid | |
| 246 | Butylate | |
| 250 | Diuron | |
| 269 | Metolachlor | |
| 272 | Carbaryl | |
a Rank is from the ATSDR 2015 Substance Priority List in which rank is based on frequency, toxicity, and potential for human exposure at Superfund sites.
et al. 2012, 2014). Genetic analysis of single-nucleotide polymorphisms related to human pharmacokinetics could provide information on variability in pharmacokinetic parameters in the population of concern. Ultimately, the pharmacokinetic parameters are helpful for evaluating the relationships between external and internal exposures and guiding selection of test concentrations. The data on individual chemical and mixture exposure and the related pharmacokinetic data would ideally be used to establish test concentrations or exposures for the appropriate in vivo or in vitro test systems that reflect the composition of real-world exposures at the site.
Assessment of Chemicals of Unknown Identity
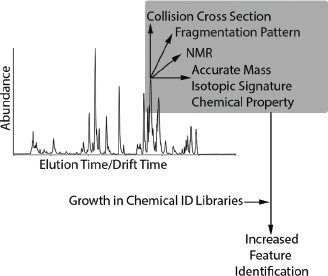
Nontargeted analyses of samples from the site revealed 5,000 unidentified chemicals in the environmental media, 3,000 in serum, 2,000 in urine, and 800 in hair (see Figure C-1). All sample types had 300 unidentified chemicals in common. One key challenge in nontargeted analysis of complex samples is to identify the unidentified chemicals accurately. Without chemical identifications, the ability to quantify exposure, conduct toxicity testing, and evaluate the plausibility of exposure–disease associations is extremely limited. To identify unknowns, standard reference materials for industrial and other chemicals and their metabolites are needed. Analytical features of the standard reference materials—such as elution time, exact mass, isotopic signature, and fragmentation pattern from, for example, gas chromatography (GC), liquid chromatography (LC), and tandem mass spectrometry (MS/MS)—can be matched to analytical features in the sample to identify the chemicals of interest. Chemical-identity libraries that contain the analytical spectra of reference standards are growing, particularly for endogenous metabolites (for example, the Human Metabolome Database, HMD), but more progress needs to be made before nontargeted analyses can become routine. The following discussion provides approaches for making progress in this field.
Two general approaches—an experimentally driven approach and another driven by cheminformatics (Horai et al. 2009; Neumann and Bocker 2010)—have been suggested to overcome the obstacles presented by the lack of chemical-identity libraries. In the experimentally driven approach, chemical-identity libraries similar to the HMD that include exact mass, elution times, isotopic signature, and mass fragmentation patterns (see Figure C-2) could be created for ToxCast and other chemicals. To support that effort, the US Environmental Protection Agency (EPA) has obtained authentic chemical standards for thousands of ToxCast chemicals and placed them in a chemical repository. Development of a complete chemical-identity library for the ToxCast chemicals (and addition of this information to such databases as the HMD) would enable measurements of these chemicals in environmental media and human biofluids. However, a major limitation in the experimental approach is the absence of standards for common environmental degradation products or metabolites that are likely to be found in biofluids. As chemical-identity libraries grow, archived GC, LC-MS, or MS/MS spectra can be searched to make new identifications.
Nuclear magnetic resonance (NMR) methods present another experimental approach to identification of unknown chemical features. The methods hold great promise because NMR analysis allows identification and quantitation of chemicals without an authentic standard. A noted limitation of the approach is its need for relatively high concentrations of target chemicals in the sample (1 µM; Bingol and Brüschweiler 2015) and its relatively low throughput. Advanced labeling techniques (Clendinen et al. 2015) and methods that involve combinations of NMR, MS, and other analytical techniques, however,
show promise for future applications (Bingol and Brüschweiler 2015).
Ion-mobility spectrometry–mass spectrometry (IMS-MS) analysis is another promising experimental approach for library-building and rapid identification of chemical features of unknowns (Ewing et al. 2016; May et al. 2016). In IMS-MS analyses, chemicals separate on the basis of their collisional cross-sectional (CCS) area during flow through a nitrogen- or helium-filled tube with a charge separation. Separation times are in the milliseconds and allow the potential for very high-throughput sample analysis. One potential advantage of IMS-MS over other analytical approaches for chemical identification for which authentic standards do not exist is that the CCS area can be calculated in silico with good accuracy (2–5% error; Paglia et al. 2014). The high throughput of the IMS-MS techniques and the possibilities of in silico library-building could produce large libraries of known chemicals, metabolites, and degradation products even if the chemical standards are not available. Those libraries could then be used to assign provisional identifications or identifications with probability statements. Furthermore, IMS-MS chemical fragmentation patterns can be matched to those in existing databases, such as the HMD, for improved chemical identification.
The other general approach is based on cheminformatics and can circumvent the challenges associated with limited chemical-identity libraries and the lack of standard reference materials. Applied in concert with emerging analytical chemistry approaches and computational methods, cheminformatics holds great potential for rapid identification or classification of unknown analytes. For example, quantitative structure–activity relationship methods that compare chromatographic behaviors of unknown analytes could be combined with other data to provide predictions about select chemical properties of the analytes. Computational approaches based on physicochemical properties have been used to predict elution times (Shah et al. 2010; Kangas et al. 2012), MS-MS fragmentation patterns (Heinonen et al. 2008; Wolf et al. 2010; Perdivara et al. 2013), and CCS area (Paglia et al. 2014). Using one or more of the analytical approaches with other cheminformatic tools for predicting metabolism and environmental degradation products (Dimitrov et al. 2010) might help to create in silico libraries that grow in breadth and accuracy and can be used to transition from nontargeted to targeted analysis.
The approaches described here represent essential methods for making the rapid transition from nontargeted to targeted analysis. For site-specific assessments with many unidentified chemicals, the approaches would provide a means of identifying analytes progressively for later hazard or risk assessment. For this case study, the committee assumed that the approaches applied to the environmental media, serum, urine, and hair samples would yield a list of 300 chemicals that are found with greatest consistency and at the highest concentrations in all samples (see Figure C-1). Chemicals that are found in environmental media and biological samples will constitute a logical choice for targeted toxicity testing because they might have a higher exposure potential than chemicals found only in environmental media.
As the number of identified chemicals increases, the data could be used to identify signatures of exposure to chemicals and mixtures. Such efforts would help to strengthen the exposure narrative and identify real-world mixtures for toxicity testing. The approaches for ranking based on hazard and bioactivity reported by Rager et al. (2016) (see Figures 2-7 and 2-8) are potentially applicable in some context of complex exposures. Other ES21 tools would then be used as needed and as described in Chapter 2 and the above section to provide better exposure char-
acterization through a more complete understanding of exposure pathways, fate and transport, and biokinetics.
CHARACTERIZING TOXICITY AFTER A CHEMICAL RELEASE
This case study considers the environmental release of a chemical that has few toxicity data and approaches for characterizing toxicity rapidly to inform decision-making. 4-Methylcyclohexanemethanol (MCHM) was the major component of a chemical mixture that was spilled into the Elk River about 1 mile upstream of a water-intake facility for the city of Charleston, West Virginia, in 2014. The immediate public-health response was a “do not drink” order, but there was not enough information to provide guidance on what types of adverse health effects might be expected from MCHM or at what exposure levels. Primarily because hazard data were sparse, an acceptable concentration of MCHM in water being consumed by the local population and the potential risks associated with exposures to it could not be easily estimated. A few models and data streams that could be used in such situations are described below. The discussion provides general guidance but is not intended to be exhaustive. For example, only the exposure scenario related to drinking of tap water is presented here. In emergency scenarios, advice would also be given on whether people, including children and infants, could bathe in the water and whether the water could be used for cooking, washing clothes, and cleaning and provided to pets. Furthermore, although the focus is on MCHM, other chemicals at concentrations of at least 1% were present in the spilled material, including 4-(methoxymethyl)cyclo-hexanemethanol (4–22%), methyl 4-methylcyclohexane carboxylate (5%), 1,4-cy-clohexanedimethanol (1–2%), and glycol phenyl ethers (propylene and dipropylene, whose concentrations were unknown).
Measured and model-predicted chemical-property information that is relevant for estimating MCHM environmental fate and toxicokinetics and for conducting an exposure assessment can be obtained from publicly available databases and software, including EPA’s EPI Suite™ program (EPA 2011), which is primarily used here to obtain chemical property, fate, and bioaccumulation information. MCHM is a relatively small (128.2 g/mol) neutral organic chemical that has a solubility limit of about 2,000 mg/L and an octanol–water partition coefficient (KOW) of about 350 (EPA 2011). It is a relatively volatile chemical (vapor pressure of about 8 Pa); however, its water solubility results in an air–water partition coefficient (KAW) of about 0.0003 (EPA 2011). Screening-level evaluative mass-balance fate models that are included in EPI Suite (EPA 2011) indicate that once released to surface water, such as a river, MCHM is not distributed significantly from water to air or sediment. The biodegradation half-life in surface water is estimated to be about 15 days (EPA 2011). Predicted bioaccumulation factor for MCHM in fish is about 20 L/kg (relatively low), and the biotransformation half-life in fish is less than 1 day (relatively short). Those screening data indicate low persistence and bioaccumulation of MCHM in the environment; chemical concentrations in the river and in possible food sources from the river would be expected to decrease relatively quickly. Long-term, chronic exposures to local residents would not be expected. More sophisticated and resource-intensive models could be used to provide more refined situation-specific calculations for the expected change in environmental concentrations over time. For example, modeling tools could be used to estimate the time that it would take for concentrations in the river at the water-intake facility to decrease. Similar tools could be applied for the water-distribution system (after intake at the treatment facility).
Measured MCHM concentrations in drinking water in the first 2 days after the spill were about 1–4 mg/L (Foreman et al. 2015; Whelton et al. 2015). To determine the safety of the water for consumption, such sensitive populations as young infants and lactating and pregnant women would need to be considered. The 95th percentile drinking-water intakes by lactating women, pregnant women, and young infants are 0.055 L/kg-day (EPA 2011), 0.043 L/kg-day (EPA 2011), and 0.24 L/kg-day (EPA 2008), respectively. Given an MCHM concentration of 2 mg/L, the estimated acute (48-hour) intake in drinking water would be 0.48 mg/kg-day for the most exposed group, young infants. Lactating women would take in 0.11 mg/kg-day. Water concentrations in Charleston tap water declined to less than 1 mg/L 5 days after the spill and continued to decline to about 0.002 mg/L 3 weeks after the spill (Foreman et al. 2015). Thus, the MCHM intake 3 weeks after the spill by the 95th percentile drinking-water consumers would have declined to 0.48 μg/kg-day in young infants and 0.11 μg/kg-day in lactating women. The predicted half-life of MCHM in humans is about 2 hours (Arnot et al. 2014), so internal concentrations are expected to decrease relatively quickly after exposure events because MCHM is not persistent or bioaccumulative in humans.
A number of symptoms were reported in the community either through emergency-room visits or in follow-up surveillance by the Centers for Disease Control and Prevention and the Kanawha Charleston Health Department. Vomiting, nausea, diarrhea, and sore throat were most associated with reported drinking of the water, whereas skin irritation and rash were associated with bathing (Whelton et al. 2015). At the time of the spill, animal data were available on acute and subacute toxicity, site-of-contact irritation, skin sensitization, and genotoxicity, but there was no information on potential developmental toxicity or long-term health effects. The information generated af-
ter the spill primarily used Tox21 tools described in Chapter 3 and provide a good example of how these tools can be used qualitatively to provide support for public-health decisions. The following discussion provides several approaches for estimating or evaluating MCMH toxicity.
A rapid approach for estimating the potential for adverse effects is chemical structural comparison with known toxicants. Published methods can be used to determine whether there are reports in the literature on chemicals that have similar structural features. Wu et al. (2013) published a decision tree for developmental toxicity that was based on a chemical structural analysis of about 900 chemicals. The decision tree contained no precedents for developmental toxicity of chemicals that had the structural features of MCMH. Although that approach does not provide a definitive answer, it is a rapid means of determining whether a chemical has a signal for developmental toxicity. It is also possible to look for structurally similar chemicals in large toxicology databases, such as those amalgamated under EPA’s Aggregated Computational Toxicology Resource program. In this case, no chemicals that had high structural similarity to MCHM were identified.
The National Toxicology Program (NTP) undertook a number of short-term assays intended to determine whether MCHM has activity against targets of concern (NTP 2016a). The testing included in vitro assays in 27 cell types, querying activity on signaling pathways relevant for development, rapid-turnaround assays in Caenorhabditis elegans and zebrafish embryos, and a 5-day toxicogenomics study in rats. No signals were generated from any in vitro assays up to relatively high concentrations (almost 100 µM) or in assays with C. elegans or zebrafish, although a minor contaminant of MCHM did have some activity in zebrafish embryos at about 100 µM. The toxicogenomics study was used to generate a biological no-observed-effect level (NOEL) for gene expression that is reported to be in the range of 6-99 mg/kg-day (the range is attributed to different methods used for data analysis). That screening-level study used six doses from 0.1 to 500 mg/kg-day (administered orally) for 5 days and evaluated gene expression in liver and kidney. A biological response was reported in liver at 6–99 mg/kg-day with no effect on kidney gene expression (NTP 2016b). The acute 95th percentile water-consumption exposure intake rates of 0.48 mg/kg-day for infants and 0.11 mg/kg-day for lactating women are lower than the NOEL for gene expression by factors of about 12–200 and 60–1,000, respectively. The committee notes that longer-term exposures were much lower. Because this example did not account for other exposure routes, which could add to ingestion exposure, the findings support the do-not-drink order issued for the entire service area (Whelton et al. 2015). Data gaps regarding other exposure routes could have been addressed by testing gene expression after administering the chemical by other relevant routes or by using physiologically based pharmacokinetic models to estimate the contribution of dermal and inhalation exposure to the total systemic concentration. Policy on interpreting the data streams will need to be created; this example is informative in describing the types of data that can be generated quickly to support risk-management decisions.
In summary, although the data differed from a standard toxicology evaluation, they were sufficient to indicate that MCHM was not structurally similar to known developmental toxicants or genotoxicants and that it did not have biological activity consistent with that of a potent developmental or systemic toxicant. A few animal studies that reported a sensitive readout (global gene expression) identified an MCHM concentration that was without biological effect, and that information supports a NOEL of about 100 mg/kg-day or somewhat lower, depending on the method of analysis. Exposure estimates derived from measurements of drinking water could be compared with the NOEL and other hazard data, and models could be used to provide initial indications of the time required for environmental concentrations to decrease to acceptable concentrations after a spill.
PREDICTING TOXICITY OF REAL-WORLD CHEMICAL MIXTURES
Once chemicals at a site or part of a spill have been identified, the first question to address is whether toxicity data on them exist. For some chemicals, there are health assessments, such as those generated by the Integrated Risk Information System program, the International Agency for Research on Cancer monographs program, and the Report on Carcinogens program. Some assessments might be out of date or have notable limitations, so there might be some benefit of using Tox21 tools and approaches described in Chapter 3 to produce additional hazard and dose–response data, perhaps focused on previously identified end points of concern or to provide missing data on variability. Many chemicals, however, will not have been assessed or not have many toxicity data, such as MCHM (described in the case study above). For those chemicals, there would be clear cost and time advantages of using Tox21 tools and approaches described in Chapter 3. For example, the potential for identified substances to pose a human health hazard can be estimated quantitatively or qualitatively by chemical structure–activity modeling (Sutter et al. 2013), by combining structural information and bioactivity profiling (Low et al. 2011, 2013), by assessing bioactivity with in vitro assays that represent a wide array of tissues and biological targets (Judson et al. 2014), by establishing appropriate points of departure followed by in vitro to in vivo extrapolation (Judson et al. 2011), and by using population-based and other in vitro models to derive chemical-specific variability estimates (Abdo et al. 2015a,b). Although the toxicity evaluation
would initially be performed on an individual-chemical basis, real-world exposures are to the chemical mixtures that have been detected in environmental samples. Adding complexity to the situation is that many of the chemicals in a mixture will not have been identified. This case study provides an approach for investigating the potential hazard posed by such mixtures.
For the toxicity assessment of complex mixtures observed in environmental samples, tissues, and biofluids, such as in the first case study described in this appendix, a biological read-across approach (Low et al. 2013; Grimm et al. 2016) that relies on bioactivity-profiling data from various in vitro toxicity assays, high-content screening assays, and possibly high-throughput genomic analyses could be used to probe potential hazards. A biological read-across might be the most expedient approach for identifying potential human health hazards posed by the uncharacterized mixtures. Heterogeneity of tissue or organ toxicity, interindividual variability, and other factors can be addressed through bioactivity profiling of real-world mixtures by using human cell models in monoculture, co-cultures of various cell types, or more complex tissue-on-a-chip models.
Figure C-3 provides an overview of the biological read-across approach. Generally, chemical representatives of various toxicant classes, such as those listed in Table C-1, should be tested in a panel of in vitro assays that will also be used to test the environmental samples to establish the range of responses. Likewise, “designed” mixtures can be created—for example, on the basis of chemical-use patterns or other exposure-based data—and tested. The testing will yield a database of the biological effects of persistent environmental pollutants from a panel of diverse in vitro assays that can be used to move the unknown mixtures into classes of known chemicals or designed mixtures and to conduct the read-across to predict potential human health hazards posed by the real-world mixtures as described further below.
The database of bioactivity readouts from representative chemicals and designed mixtures can be used as a training set for the classification models that evaluate differences between chemicals or chemical classes. The results of that activity can then be used to compare (read-across) the environmental mixtures that have unknown chemical composition with representative chemicals or designed mixtures. For example, a series of machine-learning–based models could be constructed that define biological spaces that separate one class from all others (one-vs-all) or separate a single class from another class (one-vs-one). Ultimately, a real-world environmental mixture can be profiled in the same assay battery, and the resulting bioactivity readout can be used to obtain a quantitative estimate and qualitative response related to whether the mixture behaves like a particular toxicant or toxicant class in a specific assay or assay battery.
Ultimately, high-dimensional in vitro toxicity or transcriptomic data can be used to read-across a particular mixture of unknown chemical composition to known ref-
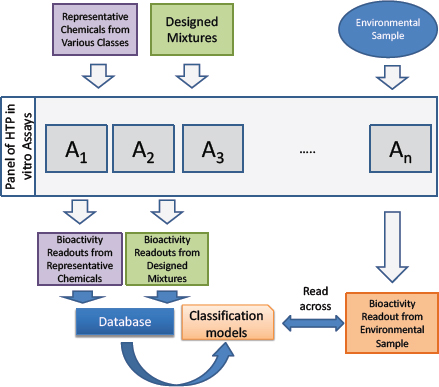
erence chemicals or chemical combinations and establish a “biological analogue” that consists of a mixture of reference chemicals for which existing toxicity benchmarks are available. If the read-across–based mixture is used as a surrogate for the original mixture, standard methods for deriving cumulative risk estimates that are based on individual chemical exposure estimates and decision benchmark methods can be applied. Although the read-across mixture might have a different chemical composition from the real-world mixture, one can assume that their biological similarity based on the in vitro toxicity testing is adequate for informing environmental decisions.
Tox21 methods of evaluating mixtures can be used to establish dose–response relationships for various bioactivity by evaluating serial dilutions of the mixture or extracts. The resulting data can be compared with the bioactivity of the samples collected at different locations at the site or adjacent areas or with the bioactivity of historical samples from the same site. A challenge in this method is similar to the one that exists for extrapolating in vitro exposure to in vivo exposure. In vitro–in vivo extrapolation (IVIVE) methods are now used to estimate the daily human oral dose, called the oral equivalent dose, necessary to achieve steady-state in vivo blood concentrations equivalent to the points of departure derived from the in vitro assays (NRC 2014). IVIVE-adjusted data from in vitro assays can be directly compared with exposure information and improve chemical priority-setting by adding a risk context to the high-throughput in vitro screening (Wetmore et al. 2013). However, IVIVE research efforts have focused on individual chemicals, not on mixtures. A study of comparative analysis of in vitro cytotoxicity of pesticide mixtures with potential human exposures is an example of computing oral equivalent doses for mixtures by using the reverse-dosimetry approach (Abdo et al. 2015a). In that study, incorporation of dosimetry with in vitro data and conversion to an oral equivalent dose of each mixture allowed a risk-relevant ranking of the mixtures that considered chemical pharmacokinetic behavior; additional exposure data were used to adjust the potencies. However, additional experimental and methodological work is needed to bridge in vitro testing data on mixtures and exposure estimates.
REFERENCES
Abdo, N., B.A. Wetmore, G.A. Chappell, D. Shea, F.A. Wright, and I. Rusyn. 2015a. In vitro screening for population variability in toxicity of pesticide-containing mixtures. Environ. Int. 85:147-155.
Abdo, N., M. Xia, C.C. Brown, O. Kosyk, R. Huang, S. Sakamuru, Y.H. Zhou, J.R. Jack, P. Gallins, K. Xia, Y. Li, W.A. Chiu, A.A. Motsinger-Reif, C.P. Austin, R.R. Tice, I. Rusyn, and F.A. Wright. 2015b. Population-based in vitro hazard and concentration-response assessment of chemicals: The 1000 genomes high-throughput screening study. Environ. Health Perspect. 123(5):458-466.
Arnot, J.A., T.N. Brown, and F. Wania. 2014. Estimating screening-level organic chemical half-lives in humans. Environ. Sci. Technol. 48(1):723-730.
Bingol, K., and R. Brüschweiler. 2015. Two elephants in the room: New hybrid nuclear magnetic resonance and mass spectrometry approaches for metabolomics. Curr. Opin. Clin. Nutr. Metab. Care 18(5):471-477.
Clendinen, C.S., G.S. Stupp, R. Ajredini, B. Lee-McMullen, C. Beecher, and A.S. Edison. 2015. An overview of methods using (13)C for improved compound identification in metabolomics and natural products. Front. Plant Sci. 6:611.
Dimitrov, S., D. Nedelcheva, N. Dimitrova, and O. Mekenyan. 2010. Development of a biodegradation model for the prediction of metabolites in soil. Sci. Total Environ. 408(18):3811-3816.
EPA (US Environmental Protection Agency). 2008. Child-Specific Exposure Factors Handbook (Final Report). EPA/600/R-06/096F. US Environmental Protection Agency, Washington, DC.
EPA (US Environmental Protection Agency). 2011. Estimation Programs Interface (EPI) Suite for Microsoft® Windows, Version 4.1. US Environmental Protection Agency, Washington, DC.
Ewing, M.A., M.S. Glover, and D.E. Clemmer. 2016. Hybrid ion mobility and mass spectrometry as a separation tool. J. Chromatogr. A. 1439:3-25.
Foreman, W.T., D.L. Rose, D.B. Chambers, A.S. Crain, L.K. Murtagh, H. Thakellapalli, and K.K. Wang. 2015. Determination of (4-methylcyclohexyl)methanol isomers by heated purge-and-trap gc/ms in water samples from the 2014 Elk River, West Virginia, chemical spill. Chemosphere 131:217-224.
Grimm, F.A., Y. Iwata, O. Sirenko, G.A. Chappell, F.A. Wright, D.M. Reif, J. Braisted, D.L. Gerhold, J.M. Yeakley, P. Shepard, B. Seligmann, T. Roy, P.J. Boogaard, H. Ketelslegers, A. Rohde, and I. Rusyn. 2016. A chemical-biological similarity-based grouping of complex substances as a prototype approach for evaluating chemical alternatives. Green Chem. 18(16):4407-4419.
Heinonen, M., A. Rantanen, T. Mielikäinen, J. Kokkonene, J. Kiuru, R.A. Ketola, and J. Rousu. 2008. FiD: A software for ab initio structural identification of product ions from tandem mass spectrometric data. Rapid Commun. Mass Spectrom. 22(19):3043-3052.
Horai, H., M. Arita, Y. Ojima, Y. Nihei, S. Kanaya, and T. Nishioka. 2009. Traceable analysis of multiple-stage mass spectra through precursor-product annotations. Pp. 173-178 in Proceedings of German Conference on Bioinformatics, September 28-29, 2009, Wittenberg, Germany, I. Grosse, S. Neumann, S. Posch, F. Schreiber, and P. Stadler, eds. Lecture Notes in Informatics-Proceedings Vol 157. Bonn: Kollen Druck+Verlag [online]. Available:http://subs.emi
s.de/LNI/Proceedings/Proceedings157/173.pdf [accessed July 27, 2016].
Judson, R.S., R.J. Kavlock, R.W. Setzer, E.A. Hubal, M.T. Martin, T.B. Knudsen, K.A. Houck, R.S. Thomas, B.A. Wetmore, and D.J. Dix. 2011. Estimating toxicity-related biological pathway altering doses for high-throughput chemical risk assessment. Chem. Res. Toxicol. 24(4):451-462.
Judson, R., K. Houck, M. Martin, T. Knudsen, R.S. Thomas, N. Sipes, I. Shah, J. Wambaugh, and K. Crofton. 2014. In vitro and modeling approaches to risk assessment from the US Environmental Protection Agency ToxCast pro-gramme. Basic Clin. Pharmacol. Toxicol. 115(1):69-76.
Kangas, L.J., T.O. Metz, G. Isaac, B.T. Schrom, B. Ginovska-Pangovska, L. Wang, L. Tan, R.R. Lewis, and J.H. Miller. 2012. In silico identification software (ISIS): A machine learning approach to tandem mass spectral identification of lipids. Bioinformatics 28(13):1705-1713.
Low, Y., T. Uehara, Y. Minowa, H. Yamada, Y. Ohno, T. Urushidani, A. Sedykh, E. Muratov, V. Kuz’min, D. Fourches, H. Zhu, I. Rusyn, and A. Tropsha. 2011. Predicting drug-induced hepatotoxicity using QSAR and toxicogenomics approaches. Chem. Res. Toxicol. 24(8):1251-1262.
Low, Y., A. Sedykh, D. Fourches, A. Golbraikh, M. Whelan, I. Rusyn, and A. Tropsha. 2013. Integrative chemical-biological read-across approach for chemical hazard classification. Chem. Res. Toxicol. 26(8):1199-1208.
May, J.C., R.L. Gant-Branum, and J.A. McLean. 2016. Targeting the untargeted in molecular phenomics with structurally-selective ion mobility-mass spectrometry. Curr. Opin. Biotechnol. 39:192-197.
Neumann, S., and S. Bocker. 2010. Computational mass spectrometry for metabolomics: Identification of metabolites and small molecules. Anal. Bioanal. Chem. 398(7-8):2779-2788.
NRC (National Research Council). 2014. A Framework to Guide Selection of Chemical Alternatives. Washington, DC: The National Academies Press.
NTP (National Toxicology Program). 2016a. West Virginia Chemical Spill: NTP Studies [online]. Available: http://ntp.niehs.nih.gov/results/areas/wvspill/studies/index.html [accessed July 27, 2016].
NTP (National Toxicology Program). 2016b. West Virginia Chemical Spill: 5-Day Rat Toxicogenomic Studies, June 2015 NTP Update [online]. Available: http://ntp.niehs.nih.gov/ntp/research/areas/wvspill/micronucleus_update_508.pdf [accessed July 27, 2016].
Paglia, G., J.P. Williams, L. Menikarachchi, J.W. Thompson, R. Tyldesley-Worster, S. Halldórsson, O. Rolfsson, A. Moseley, D. Grant, J. Langridge, B.O. Palsson, and G. Astarita. 2014. Ion mobility derived collision cross sections to support metabolomics applications. Anal. Chem. 86(8):3985-3993.
Perdivara, I., L. Perera, M. Sricholpech, M. Terajima, N. Pleshko, M. Yamauchi, and K.B. Tomer. 2013. Unusual fragmentation pathways in collagen glycopeptides. J. Am. Soc. Mass. Spectrom. 24(7):1072-1081.
Rager, J.E., M.J. Strynar, S. Liang, R.L. McMahen, A.M. Richard, C.M. Grulke, J.F. Wambaugh, K.K. Isaacs, R. Judson, A.J. Williams, and J.R. Sobus. 2016. Linking high resolution mass spectrometry data with exposure and toxicity forecasts to advance high-throughput environmental monitoring. Environ. Int. 88:269-280.
Shah, A.R., K. Agarwal, E.S. Baker, M. Singhal, A.M. Mayampurath, Y.M. Ibrahim, L.J. Kangas, M.E. Monroe, R. Zhao, M.E. Belov, G.A. Anderson, and R.D. Smith. 2010. Machine learning based prediction for peptide drift times in ion mobility spectrometry. Bioinformatics 26(13):1601-1607.
Sutter, A., A. Amberg, S. Boyer, A. Brigo, J.F. Contrera, L.L. Custer, K.L. Dobo, V. Gervais, S. Glowienke, J. van Gompel. N. Greene, W. Muster, J. Nicolette, M.V. Reddy, V. Thybaud, E. Vock, A.T. White, and L Müller. 2013. Use of in silico systems and expert knowledge for structure-based assessment of potentially mutagenic impurities. Regul. Toxicol. Pharmacol. 67(1):39-52.
Wetmore, B.A., J.F. Wambaugh, S.S. Ferguson, M.A. Sochaski, D.M. Rotroff, K. Freeman, H.J. Clewell, III, D.J. Dix, M.E. Andersen, K.A. Houck, B. Allen, R.S. Judson, R. Singh, R.J. Kavlock, A.M. Richard, and R.S. Thomas. 2012. Integration of dosimetry, exposure, and high-throughput screening data in chemical toxicity assessment. Toxicol. Sci. 125(1):157-174.
Wetmore, B.A., J.F. Wambaugh, S.S. Ferguson, L. Li, H.J. Clewell, III, R.S. Judson, K. Freeman, W. Bao, M.A. Sochaski, T.M. Chu, M.B. Black, E. Healy, B. Allen, M.E. Andersen, R.D. Wolfinger, and R.S. Thomas. 2013. Relative impact of incorporating pharmacokinetics on predicting in vivo hazard and mode of action from high-throughput in vitro toxicity assays. Toxicol. Sci. 132(2):327-346.
Wetmore, B.A., B. Allen, H.J. Clewell, III, T. Parker, J.F. Wambaugh, L.M. Almond, M.A. Sochaski, and R.S. Thomas. 2014. Incorporating population variability and susceptible subpopulations into dosimetry for high-throughput toxicity testing. Toxicol. Sci. 142(1):210-224.
Whelton, A.J., L. McMillan, M. Connell, K.M. Kelley, J.P. Gill, K.D. White, R. Gupta, R. Dey, and C. Novy. 2015. Residential tap water contamination following the Freedom Industries chemical spill: Perceptions, water quality, and health impacts. Environ Sci Technol. 49(2):813-823.
Wolf, S., S. Schmidt, M. Müller-Hannemann, and S. Neumann. 2010. In silico fragmentation for computer assisted identification of metabolite mass spectra. BMC Bioinformatics 11:148.
Wu, S., J. Fisher, J. Naciff, M. Laufersweiler, C. Lester, G. Daston, and K. Blackburn. 2013. Framework for identifying chemicals with structural features associated with the potential to act as developmental or reproductive toxicants. Chem. Res. Toxicol. 26(12):1840-1861.









