3
Life in High-Temperature Environments: Modern-Day Analogs of Early Earth Still Relevant Today
INTRODUCTION
The discovery of new single-celled organism lineages has been remarkable since the adoption of molecular genetics and the discovery of the domain Archaea (Woese et al., 1990). Prior to molecular techniques that initially emphasized the sequences of ribosomal genes (e.g., 16S rRNA), the discipline of microbiology relied nearly entirely on cultivation and the ability to grow a specific microorganism in pure culture under defined conditions. This meant that only microorganisms that grew easily under laboratory conditions were cultivated, and, in many cases, these often rapidly growing organisms do not correspond to the more numerous and relevant microbes that actually inhabit different microbiomes. We now appreciate that many of the microorganisms easily grown under laboratory conditions are often related to their more abundant and important relatives found in situ, but they generally do not exhibit the same functional attributes as numerically relevant microorganisms. Moreover, in the current -omics era, sequencing technologies and the analysis of proteins and metabolites in different environments provide detailed information regarding the specific microbes present, and the pathways employed to carry out different chemical transformations. The ability to analyze different microbiomes using coupled metagenomics, proteomics, metatranscriptomics, and metabolomics provides exciting opportunities for understanding details regarding the individual function of specific microorganisms, but also how they might interact with other community members. The fact that these analyses can be performed without the need to obtain pure cultures of microorganisms represents a major paradigm shift in environmental microbiology. It is now common to study details of numerous specific microbial populations in the same community or habitat using molecular methods which can provide details on community composition, the metabolism of specific community members, and the biochemical interactions occurring between organisms.
High-temperature geothermal or hydrothermal systems are often defined by geochemical extremes across observable spatial scales, and can result in reduced biological complexity that is metabolically focused around a specific set of micro-environmental conditions. These advantages provide opportunities for understanding the function of individual organisms in situ as well as their respective roles in community interactions that ultimately impact prediction in natural and/or managed ecosystems and our ability to optimally utilize these principles in engineering design. Examples of new discoveries and fundamental principles gained from studying geothermal
___________________
a Thermal Biology Institute, Montana State University.
* Corresponding Author: binskeep@montana.edu.
systems have been remarkable, and since the advent of molecular techniques, we have discovered several new lineages of high-temperature microorganisms living in Yellowstone National Park (YNP), and have a much clearer appreciation for how communities are organized, both in terms of composition and the expression of specific functional properties in time and space.
A detailed appreciation and understanding of the diversity of microorganisms that inhabit different Earth microbiomes has improved dramatically in the past 20 years (Anantharaman et al., 2016; Hug et al., 2016). The application of genomic approaches in environmental microbiology has provided data on the actual composition and functional capabilities of numerous different microbial communities. It is now possible to determine their specific genetic repertoire and map other -omics datasets to this genomic foundation. For example, transcriptomes, proteomes, and metabolomes require adequate reference genome sequence for interpretation, and it is becoming more routine to develop an inventory of relevant genome sequence for analyzing multiple -omics datasets for a specific habitat type (see Figure 3-1).
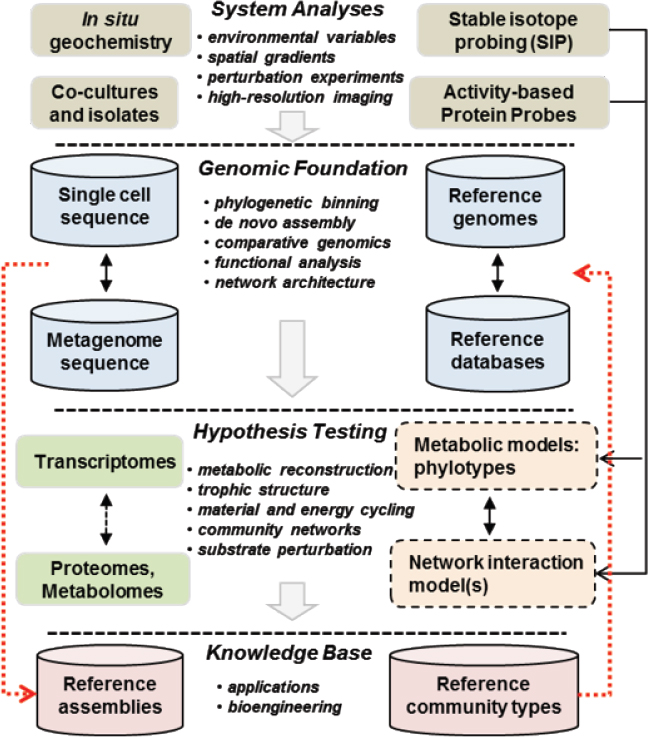
NOTE: Geochemical, isotopic, and imaging technologies can be linked to a genomics platform where sequencing and assembly of appropriate reference populations provides tools necessary for adequate interpretation of activity-based measurements in transcriptomics, proteomics, and metabolism. Deatiled community network modeling and feedback analysis can be based on informed -omic assignments coupled with an accurate model of in situ spatial and temporal realities obtained from site characterization and analysis. Appropriate references can be archived for further study and linked to databases for specific community types.
Ultimately, we need to know the diversity of microorganisms that exist in nature, the biomes they inhabit, and the interactions that occur between microbiomes and other environmental processes. Moreover, the diverse genetic capabilities found in different microorganisms are not only important for contributing to stability and resilience in natural microbiomes, but are also useful in genetic engineering and custom design. A fuller realization of the metabolic transformations and specific biochemical pathways employed by microorganisms will result in a greater ability to predict responses to environmental change in natural communities and improve application and use of microorganisms in industrial processes. Research on the chemistry of the Earth microbiomes informs different academic and industrial goals including an understanding of evolution and its history, a predictive understanding of the Earth’s elemental cycles, and response to environmental change, as well as the utilization of microbial capabilities in custom pathways and product synthesis for biological engineering.
Numerous lines of evidence suggest that life on Earth may have originated in high-temperature environments (Nisbet and Sleep, 2001; Stetter, 2006). One potential scenario for the origin of our moon is that the Earth was struck by another inner planet circa 4.5 Gya, resulting in the tilt and spin of the Earth. These conditions are thought to have created surface temperatures of near 100°C and, coupled with frequent meteorite impacts, would have created extremely hot oceans. Early Mars may have also been habitable, and life could have been transferred among the early planets. The carbon isotopic record suggests that reduced carbon was formed from photosynthetic organisms as early as 3.5 Gya, and microfossils with similar ages have been discovered in the rock record. Phylogenetic analysis of single-celled organisms generally suggests that thermophilicity was an early trait and that hyperthermophiles may have been the only life-forms to have survived bottlenecks where the Earth’s ocean temperatures may have easily reached 100°C. Although a nonthermophilic origin is also possible, current phylogenetic analyses generally place thermophilic organisms near the roots of their respective lineages. From a practical standpoint, this reality suggests that most organisms evolving from thermophiles share a significant fraction of the basic housekeeping proteins and biochemical attributes necessary for life on Earth. It follows that thermophiles are excellent models for understanding the potential function of specific proteins also found in nonthermophilic organisms, especially considering that thermal environments are often less diverse and, hence, more tractable for interpreting fundamental processes operative in microbial communities. An understanding of gene and protein function in thermophiles can often lead to a better understanding of the evolutionary history and function of genes in mesophilic organisms and/or eukarya, which are common in more complex systems.
Thermodynamic favorability is a requirement for life, but life has evolved around only a subset of favorable possibilities due to the evolution of proteins that define specific pathways and mechanisms of chemical conversion. The transfer of electrons from reduced to more oxidized chemical species provides the energy necessary for microbial metabolism as well as the fixation of inorganic carbon into microbial biomass. The capture of electrons by microorganisms from inorganic elements through processes like chemolithotrophy represents an important and fundamental linkage between the geo- and biospheres, one which has shaped the evolutionary history of Earth. Although the fixation of inorganic carbon into biomass is thought to be an important trait among early microbial life, and the importance of heterotrophy in numerous deeply rooted thermophilic lineages cannot be overstated. Nearly all archaeal lineages, except for methanogens, are chemoorganoheterotrophs capable of utilizing organic compounds for energy as well as for a primary carbon source, and this metabolic attribute defines a significant majority of all known biological diversity. Early sources of reduced carbon on the Earth may have been interplanetary, in addition to that fixed by chemo- and photoautotrophs. These primary producers are often the early colonizers of new environments, and supply a diverse array of organic compounds to other heterotrophs through the fixation of CO2. The genomics era provides tools to map specific biochemical processes and interactions occurring among different populations within a community. Progress in applying these tools to numerous different community types will provide a new infusion of knowledge about how communities function and their fundamental role in carbon and multielement cycling.
CASE STUDIES IN YELLOWSTONE NATIONAL PARK: A GEOCHEMISTRY WORKBENCH
Elements and compounds ubiquitous in the Earth’s crust and/or atmosphere are key constituents necessary for energy capture and carbon acquisition in high-temperature environments, and include iron, sulfur, arsenic, carbon
dioxide, methane, and oxygen (see Figure 3-2). YNP provides a natural laboratory for studying microbiological responses to different geochemical conditions, where several major environmental state variables, such as pH and the concentrations of electron donors and acceptors, vary across geothermal systems. Geochemical differences are also coupled with hydrogeological properties and may be expressed within a microbial mat as spatial and/or temporal chemical gradients. It is possible to utilize the variation in attributes of geothermal springs to develop an understanding of metabolic attribute distributions as a function of key environmental variables (see Figure 3-3). This information can address questions pertaining to the evolution of aerobic life, but also those regarding optimum conditions and opportunities for using microorganisms and/or their biochemistry in custom design or synthesis strategies.
The fundamental linkage between protein function and environmental circumstance is a guiding principle for the evolution of life. Examples of this principle discovered in the high-temperature systems of YNP include numerous chemolithoautotrophic pathways such as the oxidation of sulfur, arsenic, and iron. The lineages that contain proteins necessary for mediating this type of energy transfer are distributed in environmental circumstances where the function of these proteins is optimized. For example, all three major lineages of Aquificales in YNP are capable of oxidizing arsenite to arsenate through the action of a dimethyl sulfoxide (DMSO)-molybdopterin arsenite oxidase, and are likely capturing energy from this reaction (Inskeep et al., 2007; Hamamura et al., 2009). Similar mechanisms of thiosulfate, sulfide, and elemental sulfur oxidation by members of the Aquificales (Bacteria) and Sulfolobales (Archaea) suggest that the last universal common ancestor of bacteria and archaea had these capabilities. DMSO-molybdopterins responsible for the reduction of sulfur and arsenate are specifically important to members of the Thermoproteales (Jay et al., 2016). The aerobic oxidation of ferrous iron by specific members of the Thermoplasmatales (Euryarchaeota), Sulfolobales (Crenarchaeota), and acidophilic iron-oxidizing bacteria
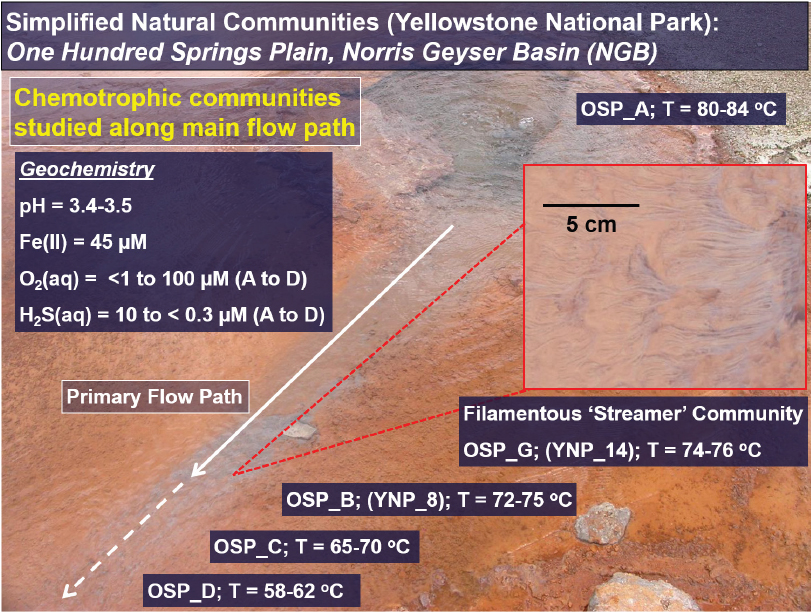
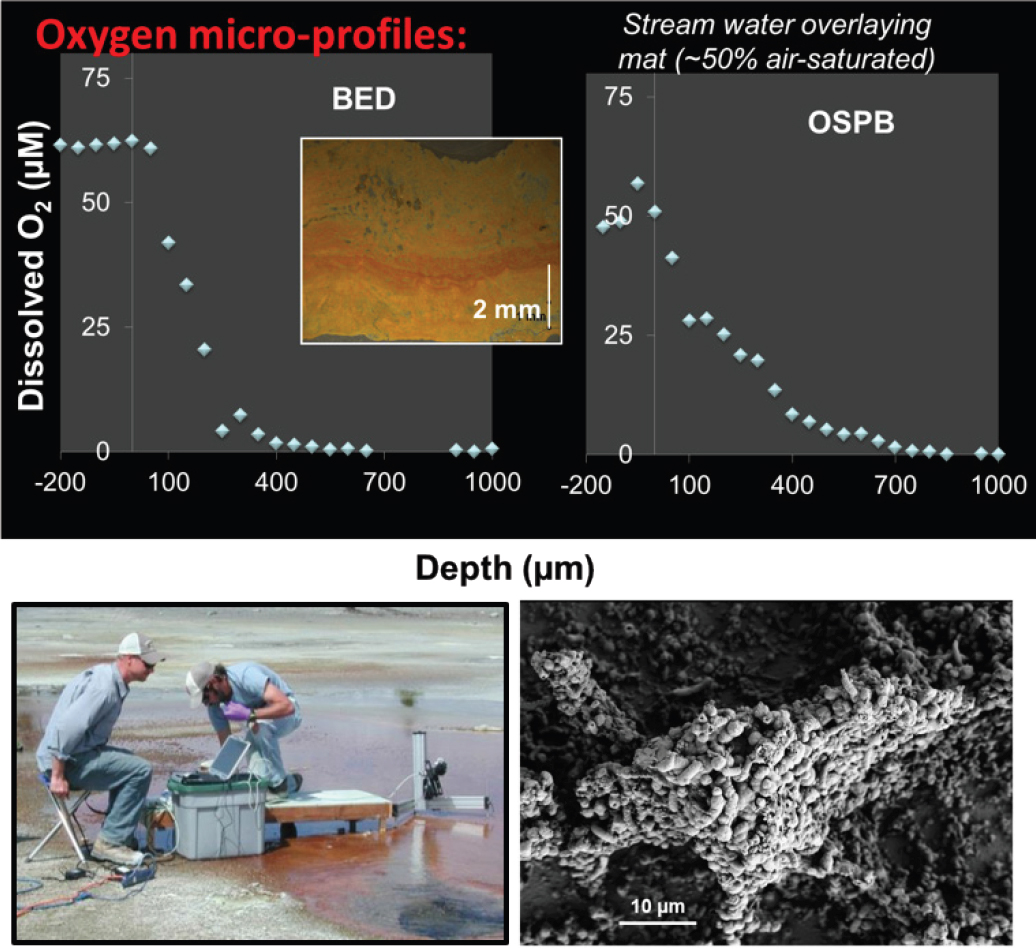
NOTE: In situ characterization of spatial gradients in oxygen inform factors responsible for functional partitioning of Fe(II)oxidation and microbial mat growth in thermo-acidic springs of Norris Geyser Basin (Yellowstone National Park, Wyoming, United States). Biological oxygen consumption rates determined with microelectrodes (H. Bernstein, J. Beam) are limited by oxygen diffusion through the mat and support the formation of microterracettes (lower right) comprised of individual cells and Fe(III)-oxides containing high contents of adsorbed arsenate.
like Acidimicrobium ferroxidans reveal similar biochemical mechanisms that may have been especially important in early respiratory processes. Details of the respiratory process in Metallosphaera yellowstonensis reveal the importance of small blue copper proteins and a novel cytochrome (FoxC; see Figure 3-4) that likely receives electrons from Fe(II) (Kozubal et al., 2011). This unique group of Fox proteins imparts functional attributes that explain the distribution of highly similar Metallosphaera populations in acidic ferric oxide mats throughout YNP (Kozubal et al., 2011, 2012).
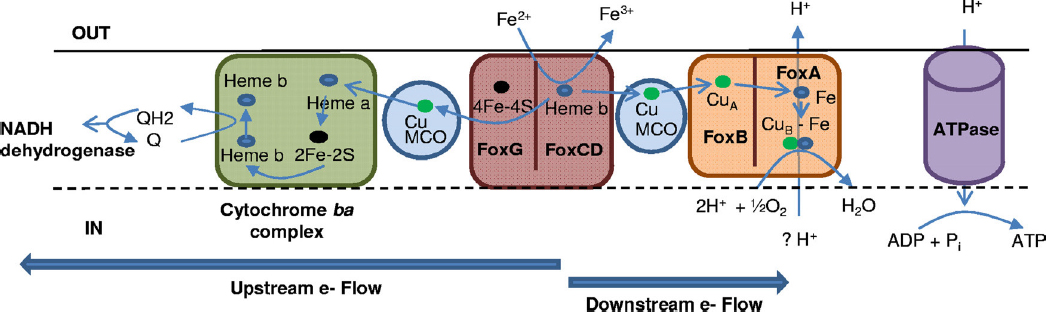
NOTE: Blue arrows indicate paths of electrons. Q, quinone; QH2, hydroquinone.
SOURCE: Kozubal et al., 2011.
Several new phyla of Archaea and Bacteria have been discovered in specific habitat types of YNP, and phylogenomic analyses reveal their importance as deeply rooted lineages in specific habitat types. For example, two new lineages of thermophilic Archaea have been described in thermoacidic ferric oxide microbial mats: Members of the Geoarchaeota and novel archaeal group 2 are dominant heterotrophs in slightly acidic iron mats (Kozbual et al., 2012, 2013; Beam et al., 2016). New bacterial lineages including members of the Pyropristinus and Calescamantes (Colman et al., 2016) have been described in higher-pH, filamentous-streamer communities containing Thermocrinis spp. (Aquificales) as a major autotrophic community member. These newly described thermophiles are all aerobic chemoorganotrophs. Consequently, thermal systems support active communities containing deeply rooted heterotrophic archaea and bacteria, which suggest early linkages between the aerobic production of autotrophic and heterotrophic biomass.
FUTURE OPPORTUNITIES IN HIGH-TEMPERATURE BIOLOGY
The genomics platform can be applied to dissect the organisms, pathways, and molecules involved in microbial community networks (Taffs et al., 2009; Hunt et al., 2016). As these tools become more mainstream and successfully define more environmental circumstances, progress will emerge toward understanding the dynamics of regulation, and modes of community interaction. This level of understanding will lead to further insight regarding microbial community response, resilience, control, and modification. And, although the genomics era has provided the tools for characterizing numerous new phylotypes (Anantharaman et al., 2016; Hug et al., 2016), we still need to make more significant strides in understanding and quantifying in situ physiological activities, modes of microbial interactions, the dynamics of microbial response to environmental change, and rates of genetic change in natural communities. A fully integrated -omics platform with greater knowledge of temporal and spatial changes will enable a more predictive understanding of naturally occurring microbial communities and their role in other major biological cycles responding to global change. Moreover, the details of biochemical transformation employed in natural and engineered communities can be integrated more readily into custom product synthesis. The advantages of thermal stability in enzymology are recognized and implied in the very tools that have empowered the molecular revolution via high-throughput, high-fidelity polymerases. High temperature can be advantageous for custom product synthesis for numerous reasons, and thermal stable enzymes involved in replication, repair, macromolecular synthesis, and carbon cycling reactions represent a source of biotechnical information that is extremely useful to the future of applied chemical engineering.
ACKNOWLEDGMENTS
William P. Inskeep acknowledges support from the U.S. Department of Energy (DOE)–Pacific Northwest National Laboratory (Richland, Washington), the DOE–Joint Genome Institute (Walnut Creek, California), the National Science Foundation Integrative Graduate Education and Research Traineeship, the Yellowstone Center for Resources (National Park Service, Mammoth, Wyoming), and the Montana Agricultural Experiment Station (Project 911300).
REFERENCES
Anantharaman, K., C. T. Brown, L. A. Hug, I. Sharon, C. J. Castelle, A. J. Probst, B. C. Thomas, A. Singh, M. J. Wilkins, U. Karaoz, E. L. Brodie, K. H. Williams, S. S. Hubbard, and J. F. Banfield. 2016. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat Commun 7:13219.
Beam, J. P., H. C. Bernstein, Z. J. Jay, M. A. Kozubal, R. deM. Jennings, S. G. Tringe, and W. P. Inskeep. 2016. Assembly and succession of iron oxide microbial mat communities in acidic geothermal springs. Front Microbiol 7:25.
Colman, D. R., Z. J. Jay, W. P. Inskeep, R. deM. Jennings, K. R. Maas, D. B. Rusch, and C. D. Takacs-Vesbach. 2016. Novel, deep-branching heterotrophic bacterial populations recovered from thermal spring metagenomes. Front Microbiol 7:304.
Hamamura, N., R. E. Macur, S. Korf, G. Ackerman, W. P. Taylor, M. Kozubal, A-L. Reysenbach, and W. P. Inskeep. 2009. Linking microbial oxidation of arsenic with detection and phylogenetic analysis of arsenite-oxidase genes in diverse geothermal environments. Environ Microbiol 11:421-431.
Hug, L. A., B. J. Baker, K. Anantharaman, C. T. Brown, A. J. Probst, C. J. Castelle, C. N. Butterfield, A. W. Hernsdorf, Y. Amano, K. Ise, Y. Suzuki, N. Dudek, D. A. Relman, K. M. Finstad, R. Amundson, B. C. Thomas, and J. F. Banfield. 2016. A new view of the tree of life. Nature Microbiol 1:16048.
Hunt, K., R. deM. Jennings, W. P. Inskeep, and R. P. Carlson. 2016. Stoichiometric modeling of assimilatory and dissimilatory biomass utilization in a microbial community. Environ Microbiol 18(12):4946-4960.
Inskeep, W. P., R. E. Macur, N. Hamamura, T. P. Warelow, S. A. Ward, and J. M. Santini. 2007. Detection, diversity and expression of aerobic bacterial arsenite oxidase genes. Environ Microbiol 9:934-943.
Jay, Z. J., J. P. Beam, M. A. Kozubal, R. deM. Jennings, D. Rusch, and W. P. Inskeep. 2016. The distribution, diversity and function of predominant Thermoproteales in high-temperature environments of Yellowstone National Park. Environ Microbiol 18:4755-4769.
Kozubal, M., R. E. Macur, M. Dlakic, and W. P. Inskeep. 2011. Terminal oxidase diversity and function in Metallosphaera yellowstonensis: Gene expression and protein modeling suggest mechanisms of Fe(II) oxidation in the Sulfolobales. Appl Environ Microbiol 77:1844-1853.
Kozubal, M. A., R. E. Macur, Z. J. Jay, J. P. Beam, S. A. Malfatti, S. G. Tringe, B. D. Kocar, T. Borch, and W. P. Inskeep. 2012. Microbial iron cycling in acidic geothermal springs of Yellowstone National Park: Integrating molecular surveys, geochemical processes, and isolation of novel Fe-active microorganisms. Front Microbiol 3:109.
Kozubal, M. A., M. Romine, R. deM. Jennings, Z. J. Jay, S. G. Tringe, D. B. Rusch, J. P. Beam, L. A. McCue, and W. P. Inskeep. 2013. Geoarchaeota: A new candidate phylum in the Archaea from high-temperature acidic iron mats in Yellowstone. ISME J 7:622-634.
Nisbet, E. G., and N. H. Sleep. 2001. The habitat and nature of early life. Nature 409:1083-1091.
Stetter, K. O. 2006. Hyperthermophiles in the history of life. Phil Trans R Soc B 361(1474):1837-1843.
Taffs, R., J. E. Aston, K. Brileya, Z. Jay, C. G. Klatt, S. McGlynn, N. Mallette, S. Montross, R. Gerlach, W. P. Inskeep, D. M. Ward, and R. P. Carlson. 2009. In silico approaches to study mass and energy flows in microbial consortia: A syntrophic case study. BMC Systems Biology 3:114.
Woese, C. R., O. Kandler, and M. L. Wheelis. 1990. Towards a natural system of organisms: Proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 87:4576-4579.
This page intentionally left blank.








