4
Advancing Our Understanding of the Chemistry of Soil Microbiomes
Trent R. Northen,a,b,* Zheyun Zhang,aJian Gao,aTami Swenson,band Yasuo Yoshikunia,b
INTRODUCTION
Soil health is the foundation of civilization and integral to life on the Earth, yet the processes of building or maintaining soil fertility are poorly understood (Amundson et al., 2015). This is not surprising, given that soils are perhaps the most complex environments on the Earth with vast microbial diversity. It is estimated that there are some 1030 microbes on the Earth, a large fraction of which live in soils (Whitman et al., 1998). Collectively, these microbes govern organic soil compounds, which have long been associated with soil health and microbial diversity. In fact, soils are the largest terrestrial organic carbon reservoir (~1,500 gigatons [GT] to a depth of 1 m) (Paustian et al., 2016). The organic contents in soils can be thought of as a long-term chemical balance between deposition and mineralization of small organic metabolites derived from plant and microbial inputs (Schmidt et al., 2011).
In soil systems, plant biomass and root exudates are two main carbon inputs as substrates for soil microbiomes (Lange et al., 2015). Microbes decompose plant biopolymers using exogenous enzymes to produce metabolites like hexose, pentose, and lignols that they take up to support their growth and metabolism. Plants exude up to 50% of their photosynthate through roots (Rellán-Álvarez et al., 2015; Grossmann et al., 2011), presumably to pay beneficial microbes for activities ranging from soil-borne pathogen exclusion, nitrogen fixation, nutrient mobilization, and plant growth promotion through special secondary metabolites. Therefore, soil chemistry is a strong unifying component for soil ecosystems (Bradford et al., 2016). Understanding the relationships between microbes and soil metabolites has tremendous potential to enable us to predict and harness microbiomes to address critical challenges to soil health (Falkowski et al., 2008; see Figure 4-1).
Soils are presently facing major threats at a time when it is projected that agricultural production must increase 60% by 2050 to accommodate a growing population (Blaser et al., 2016). Degradation and erosion have been drastically accelerated by anthropogenic disturbances, like cultivation, at an alarming rate. Soil degradation exceeds natural soil formation (Amundson et al., 2015; Blaser et al., 2016), and approximately 50% of agricultural soil carbon has been lost as a result of extractive agricultural and land management practices. Depletion of soil organic carbon leads to a downward spiral of declining biodiversity, decreased water infiltration, and loss of nutrient and water retention, ultimately resulting in marginal soils with less than 1% organic carbon, which are
___________________
a Joint Genome Institute, U.S. Department of Energy.
b Environmental Genomics and System Biology Division, Lawrence Berkeley National Laboratory.
* Corresponding Author: trnorthen@lbl.gov.
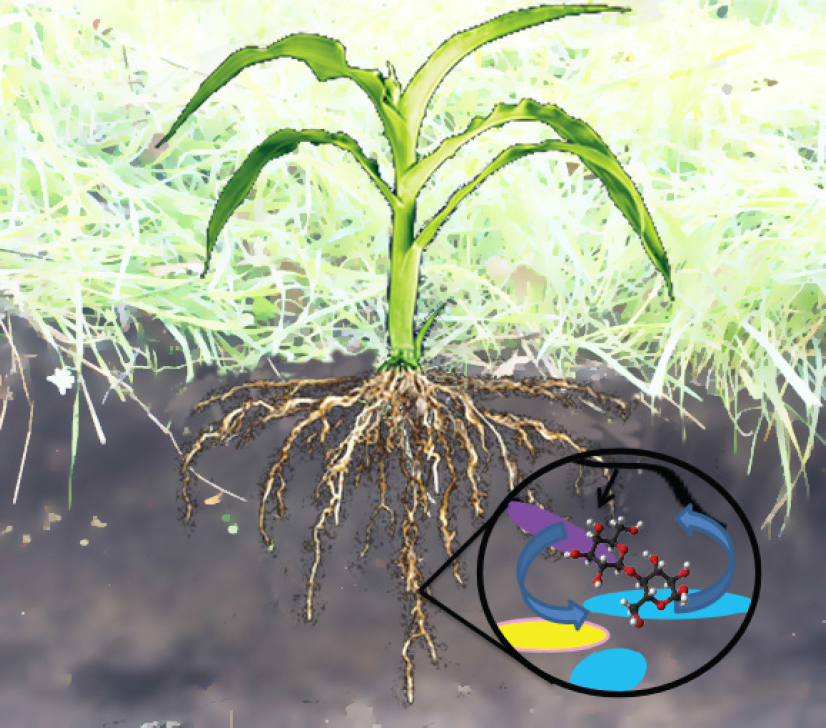

NOTE: Colored shapes in the circle represent diverse soil microbes exchanging metabolites with a black plant root hair.
largely unsuitable for conventional food crops growth (Lal, 2004). Because microbiomes play essential roles in soil carbon cycling, a better understanding of their activities and control over soil chemistry is essential to protect and build the healthy soils that are needed to feed and fuel humanity, while maintaining ecosystems.
Advancing our understanding of the mechanisms by which plant–soil microbiomes control soil chemistry has the potential to identify approaches that build soil organic carbon in marginal soils. This is highly desirable not only because it could decrease atmospheric CO2, but also because it would create an upward cycle of increased biodiversity, and improved water and nutrient retention. Ultimately, we can imagine approaches for increasing the low-input productivity of degraded soils by developing bioenergy crops with custom microbiomes that build soil carbon. A recent report suggests that a further 0.4-1.2 GT of carbon per year can be stored in world soils, helping offset 5-15% of global fossil-fuel emissions (Lal, 2004). Therefore, advancing our understanding of the chemistry of soil microbiomes is critical for the stewardship of soil ecosystems, protecting and enhancing the global carbon sink, and provisioning our growing population.
SOIL CHEMISTRY: EMERGING VIEW OF SOIL ORGANIC MATTER AS MICROBIAL METABOLITES
Environmental metagenomics now allows us to directly measure genetic potential of microbiomes in situ with the capability of linking soil chemistry to soil biology (Tringe and Rubin, 2005; Vogel et al., 2010; Lewis et al., 2012; Eloe-Fadrosh et al., 2016). Yet, we lack vital knowledge to link these soil microbiomes to their in situ activities. This is because community-level phenotypes of microbial consortia vary significantly due to the presence of genes, their expression, localization, and population size of microbes in various biogeochemical conditions. Complementing genetic analysis with direct biochemical observations presents an opportunity to establish direct association between genes and functions (Phelan et al., 2012). However, this requires a precise understanding of soil organic matter composition, concentration, and accessibility to microbes. Since these processes occur within the three-dimensional architecture of the soil environment, the spatial distribution of microbes and metabolites represents another important factor in linking metagenomes to soil chemistry.
Up to now, our understanding of soil organics has been very coarse. Traditionally, soil carbon was assumed to be composed of recalcitrant macromolecules, like humic substances, formed via in situ polymerization and other processes. However, recent spectroscopic analyses have led to the emerging view that soil carbon is largely composed of small molecular weight microbial metabolites associated, with varying affinity, to soil minerals (Schmidt et al., 2011). Therefore, to decipher the underlying processes of soil chemistry, the definition of soil carbon as total organic carbon and more recently as labile and recalcitrant carbon is not enough to link soil metabolites to microbial genomics (Lehmann and Kleber, 2015). Fortunately, soil metabolomics methods are being developed to directly characterize the small molecule metabolites within soils, enabling determination of the critical factors governing soil carbon cycling such as plant biomass deconstruction, metabolite partitioning into the microbiome, and metabolite-mineral sorption (Swenson et al., 2015a,b).
EXOMETABOLOMICS: COUPLING SOIL MICROBIOMES TO SOIL CHEMISTRY
Direct characterization of the metabolic potential and chemistry of soil microbiomes can provide correlations between soil microbes and specific metabolites (Tringe and Rubin, 2005; Woyke et al., 2006; Pati et al., 2010; see Figure 4-2). Moving to metabolic reality is much more challenging because of the gap between genotype and
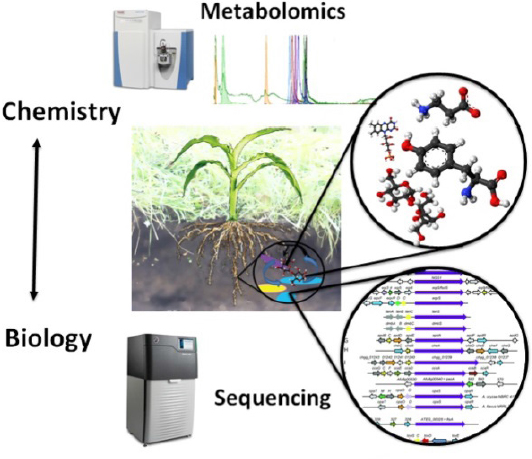
phenotype. Laboratory-based studies of extracellular metabolites or exometabolomics provide a strong approach to coupling the metabolic activities of soil microbiomes to soil chemistry, mixtures of hundreds of soil metabolites with relevant composition and concentrations (Baran et al., 2009). Specifically, by culturing isolates on environmentally relevant media, we can determine the range of substrates they can produce and mineralize. This can provide causal data to help understand the mechanisms by which plant–soil microbiomes control soil chemistry and inform the development of predictive ecosystem models of carbon and nutrient cycling (Rinehart et al., 2014).
Exometabolomics also has the capacity to define the uptake and release of a broad spectrum of metabolites in realistic environments; delineate microbial exometabolite niche patterns among sympatric soil consortia; couple soil chemical diversity to microbial diversity; annotate gene/microbe function; and improve our understanding of soil metabolic webs and nutrient cycling. For example, based on exometabolomics data, Baran et al. hypothesized “exometabolite niche partitioning with high levels of microbial substrate specialization” as a critical strategy by which microbial diversity is coupled to metabolite diversity in desert biocrusts and mesophilic soils (Baran et al., 2015).
Improving the understanding of metabolites’ spatial distributions within soils will be extremely challenging. Mass spectrometry imaging (MSI) is a rapidly emerging technique that enables direct measurement of the spatial locations of biomolecules. We anticipate that this approach, in conjunction with exometabolomics, will provide important new insights into in situ microbial metabolite production to localize metabolites within soil systems (Yang et al., 2009; Watrous and Dorrestein, 2011; Silva and Northen, 2015). For example, combining MSI with the introduction of stable labeling isotopes can lead to discovery of hotspots of microbial activities for detailed examination using systems biology approaches, ultimately generating hypotheses that can be tested in ecosystem studies (Watrous et al., 2012).
LABORATORY ECOSYSTEMS: DETERMINING CHEMICAL MECHANISMS OF MICROBIOMES
Directly testing the metabolic webs of microbes using exometabolomics in field studies is greatly complicated by soil heterogeneity, irreproducibility, lack of control, and inability to use many standard reductionist tools like comparing mutants. Hence, there is an urgent need for reproducible fabricated laboratory plant–soil–microbe ecosystems to test the hypothesis of metabolic interactions within microbiomes (see www.eco-fab.org). These systems would need to be rooted and routinely validated against native ecosystems to ensure their relevance. They could range from complex plant–native soil microbiomes within devices enabling careful control and observations to highly simplified ecosystems that enable testing of specific plant–microbe interactions. Once relevance is established, laboratory ecosystems that enable genetic control over the plant and the microbiome as well as the chemistry of the system would be a powerful tool for the scientific community, ideally enabling scientists around the world to reproduce and build on each other’s work.
One great advantage of model laboratory ecosystems is the ability to establish causal mechanisms between specific genes; microbes and plants; metabolites; and abiotic factors, for example, to discover biotic and abiotic factors driving soil carbon accumulation. There are several existing approaches that enable discovery and testing of the impact of specific genes on the biochemical ecology of soil microbiomes, which simply are currently not possible to utilize in natural ecosystems. For example, mutant fitness profiling provides a rapid tool for discovering functional annotation of uncharacterized genes that are responsible for important ecological processes. This approach has been used to examine genes mediating electron transfer in syntrophic co-cultures, and to evaluate gene regulation in relation to metabolic needs (Baran et al., 2013; Wetmore et al., 2015; Kosina et al., 2016). Synthetic biology techniques like CRISPR (clustered regularly interspaced short palindromic repeats)/Cas9 (CRISPR-associated protein 9)/sgRNA (single guide RNA) and recombinase-assisted genome engineering provide other powerful tools for testing gene and pathway function (Qi et al., 2013; Santos et al., 2013; Doench et al., 2014; Shalem et al., 2015). These approaches have been used to construct and manipulate artificial microbial communities with lower biosynthetic cost. There are also numerous reports of using these tools to construct microbes that are chemiluminescent or express fluorescent proteins in response to specific biotic or abiotic factors (Grossmann et al., 2011; Rállan-Álvarez et al., 2015). We believe these approaches could
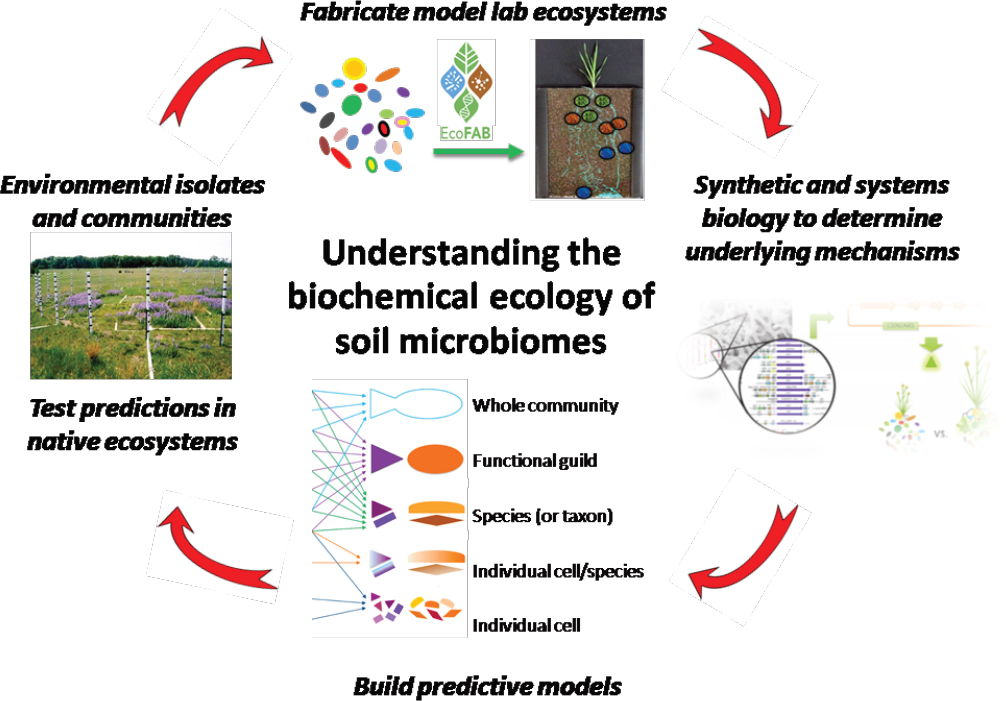
be especially powerful in localizing in situ processes within laboratory ecosystems for subsequent investigation using systems biology tools and MSI. Finally, the resulting data and mechanisms would enable development of computational models, including stoichiometric metabolic and functional gene-centric models, for developing soil microbiomes that can then be tested in field studies (see Figure 4-3), which are required to develop effective approaches for restoring soil carbon, promoting low-input agriculture, mitigating global climate change, and ultimately understanding the chemistry of soil microbiomes (Tyson et al., 2004; Treseder et al., 2012; Bordbar et al., 2014; Wang et al., 2016).
ACKNOWLEDGMENTS
This work was funded through the Laboratory Directed Research and Development Program of Lawrence Berkeley National Laboratory supported by the U.S. Department of Energy Office of Science and the U.S. Department of Energy Office of Biological and Environmental Research Early Career Program (awarded to Trent R. Northen) under Contract No. DE-AC02-05CH11231.
REFERENCES
Amundson, R., A. A. Berhe, J. W. Hopmans, C. Olson, A. E. Sztein, and D. L. Sparks. 2015. Soil science: Soil and human security in the 21st century. Science 348:1261071.
Baran, R., W. Reindl, and T. R. Northen. 2009. Mass spectrometry based metabolomics and enzymatic assays for functional genomics. Curr Opin Microbiol 12:547-552.
Baran, R., B. P. Bowen, M. N. Price, A. P. Arkin, A. M. Deutschbauer, and T. R. Northen. 2013. Metabolic footprinting of mutant libraries to map metabolite utilization to genotype. ACS Chem Biol 8:189-199.
Baran, R., E. L. Brodie, J. Mayberry-Lewis, E. Hummel, U. N. Da Rocha, R. Chakraborty, B. P. Bowen, U. Karaoz, H. Cadillo-Quiroz, F. Garcia-Pichel, and T. R. Northen. 2015. Exometabolite niche partitioning among sympatric soil bacteria. Nat Commun 6:8289.
Blaser, M. J., Z. G. Cardon, M. K. Cho, J. L. Dangl, T. J. Donohue, J. L. Green, R. Knight, M. E. Maxon, T. R. Northen, K. S. Pollard, and E. L. Brodie. 2016. Toward a predictive understanding of Earth’s microbiomes to address 21st century challenges. mBIO 7(3):e00714-16.
Bordbar, A., J. M. Monk, Z. A. King, and B. O. Palsson. 2014. Constraint-based models predict metabolic and associated cellular functions. Nat Rev Genet 15:107-120.
Bradford, M. A., W. R. Wieder, G. B. Bonan, N. Fierer, P. A. Raymond, and T. W. Crowther. 2016. Managing uncertainty in soil carbon feedbacks to climate change. Nat Clim Change 6:751-758.
Doench, J. G., E. Hartenian, D. B. Graham, Z. Tothova, M. Hegde, I. Smith, M. Sullender, B. L. Ebert, R. J. Xavier, and D. E. Root. 2014. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat Biotechnol 32:1262-1267.
Eloe-Fadrosh, E. A., N. N. Ivanova, T. Woyke, and N. C. Kyrpides. 2016. Metagenomics uncovers gaps in amplicon-based detection of microbial diversity. Nat Microbiol 1:15032.
Falkowski, P. G., T. Fenchel, and E. F. Delong. 2008. The microbial engines that drive Earth’s biogeochemical cycles. Science 320:1034-1039.
Grossmann, G., W. J. Guo, D. W. Ehrhardt, W. B. Frommer, R. V. Sit, S. R. Quake, and M. Meier. 2011. The RootChip: An integrated microfluidic chip for plant science. Plant Cell 23:4234-4240.
Kosina, S. M., M. A. Danielewicz, M. Mohammed, J. Ray, Y. Suh, S. Yilmaz, A. K. Singh, A. P. Arkin, A. M. Deutschbauer, and T. R. Northen. 2016. Exometabolomics assisted design and validation of synthetic obligate mutualism. ACS Synth Biol 5:569-576.
Lal, R. 2004. Soil carbon sequestration impacts on global climate change and food security. Science 304:1623-1627.
Lange, M., N. Eisenhauer, C. A. Sierra, H. Bessler, C. Engels, R. I. Griffiths, P. G. Mellado-Vázquez, A. A. Malik, J. Roy, S. Scheu, S. Steinbeiss, B. C. Thomson, S. E. Trumbore, and G. Gleixner. 2015. Plant diversity increases soil microbial activity and soil carbon storage. Nat Commun 6:6707.
Lehmann, J., and M. Kleber. 2015. The contentious nature of soil organic matter. Nature 528:60-68.
Lewis, N. E., H. Nagarajan, and B. Ø. Palsson. 2012. Constraining the metabolic genotype-phenotype relationship using a phylogeny of in silico methods. Nat Rev Microbiol 10:291-305.
Pati, A., N. N. Ivanova, N. Mikhailova, G. Ovchinnikova, S. D. Hooper, A. Lykidis, and N. C. Kyrpides. 2010. GenePRIMP: A gene prediction improvement pipeline for prokaryotic genomes. Nat Methods 7:455-457.
Paustian, K., J. Lehmann, S. Ogle, D. Reay, G. P. Robertson, and P. Smith. 2016. Climate-smart soils. Nature 532:49-57.
Phelan, V. V., W. T. Liu, K. Pogliano, and P. C. Dorrestein. 2012. Microbial metabolic exchange—the chemotype-to-phenotype link. Nat Chem Biol 8:26-35.
Qi, L. S., M. H. Larson, L. A. Gilbert, J. A. Doudna, J. S. Weissman, A. P. Arkin, and W. A. Lim. 2013. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152:1173-1183.
Rellán-Álvarez, R., G. Lobet, H. Lindner, P. L. Pradier, J. Sebastian, M. C. Yee, Y. Geng, C. Trontin, T. LaRue, A. Schrager-Lavelle, C. H. Haney, R. Nieu, J. Maloof, J. P. Vogel, and J. R. Dinneny. 2015. GLO-Roots: An imaging platform enabling multidimensional characterization of soil-grown root systems. Elife 4.
Rinehart, D., C. H. Johnson, T. Nguyen, J. Ivanisevic, H. P. Benton, J. Lloyd, A. P. Arkin, A. M. Deutschbauer, G. J. Patti, and G. Siuzdak. 2014. Metabolomic data streaming for biology-dependent data acquisition. Nat Biotechnol 32:524-527.
Santos, C. N. S., D. D. Regitsky, and Y. Yoshikuni. 2013. Implementation of stable and complex biological systems through recombinase-assisted genome engineering. Nat Commun 4.
Schmidt, M. W. I., M. S. Torn, S. Abiven, T. Dittmar, G. Guggenberger, I. A. Janssens, M. Kleber, I. Kögel-Knabner, J. Lehmann, D. A. Manning, P. Nannipieri, D. P. Rasse, S. Weiner, and S. E. Trumbore. 2011. Persistence of soil organic matter as an ecosystem property. Nature 478:49-56.
Shalem, O., N. E. Sanjana, and F. Zhang. 2015. High-throughput functional genomics using CRISPR-Cas9. Nat Rev Genet 16:299-311.
Silva, L. P., and T. R. Northen. 2015. Exometabolomics and MSI: Deconstructing how cells interact to transform their small molecule environment. Curr Opin Biotechnol 34:209-216.
Swenson, T. L., B. P. Bowen, P. S. Nico, and T. R. Northen. 2015a. Competitive sorption of microbial metabolites on an iron oxide mineral. Soil Biol Biochem 90:34-41.
Swenson, T. L., S. Jenkins, B. P. Bowen, and T. R. Northen. 2015b. Untargeted soil metabolomics methods for analysis of extractable organic matter. Soil Biol Biochem 80:189-198.
Treseder, K. K., T. C. Balser, M. A. Bradford, E. L. Brodie, E. A. Dubinsky, V. T. Eviner, K. S. Hofmockel, J. T. Lennon, U. Y. Levine, B. J. MacGregor, J. Pett-Ridge, and M. P. Waldrop. 2012. Integrating microbial ecology into ecosystem models: Challenges and priorities. Biogeochemistry 109:7-18.
Tringe, S. G., and E. M. Rubin. 2005. Metagenomics: DNA sequencing of environmental samples. Nat Rev Genet 6:805-814.
Tyson, G. W., J. Chapman, P. Hugenholtz, E. E. Allen, R. J. Ram, P. M. Richardson, V. V. Solovyev, E. M. Rubin, D. S. Rokhsar, and J. F. Banfield. 2004. Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428:37-43.
Vogel, J. P., D. Garvin, T. C. Mockler, J. Schmutz, D. Rokhsar, and M. Bevan. 2010. Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 463:763-768.
Wang, M. X., J. J. Carver, V. V. Phelan, L. M. Sanchez, N. Garg, Y. Peng, D. D. Nguyen, J. Watrous, C. A. Kapono, T. Luzzatto-Knaan, C. Porto, A. Bouslimani, A. V. Melnik, M. J. Meehan, W. T. Liu, M. Crüsemann, P. D. Boudreau, E. Esquenazi, M. Sandoval-Calderón, R. D. Kersten, L. A. Pace, R. A. Quinn, K. R. Duncan, C. C. Hsu, D. J. Floros, R. G. Gavilan, K. Kleigrewe, T. Northen, R. J. Dutton, D. Parrot, E. E. Carlson, B. Aigle, C. F. Michelsen, L. Jelsbak, C. Sohlenkamp, P. Pevzner, A. Edlund, J. McLean, J. Piel, B. T. Murphy, L. Gerwick, C. C. Liaw, Y. L. Yang, H. U. Humpf, M. Maansson, R. A. Keyzers, A. C. Sims, A. R. Johnson, A. M. Sidebottom, B. E. Sedio, A. Klitgaard, C. B. Larson, P. CA. Boya, D. Torres-Mendoza, D. J. Gonzalez, D. B. Silva, L. M. Marques, D. P. Demarque, E. Pociute, E. C. O’Neill, E. Briand, E. J. Helfrich, E. A. Granatosky, E. Glukhov, F. Ryffel, H. Houson, H. Mohimani, J. J. Kharbush, Y. Zeng, J. A. Vorholt, K. L. Kurita, P. Charusanti, K. L. McPhail, K. F. Nielsen, L. Vuong, M. Elfeki, M. F. Traxler, N. Engene, N. Koyama, O. B. Vining, R. Baric, R. R. Silva, S. J. Mascuch, S. Tomasi, S. Jenkins, V. Macherla, T. Hoffman, V. Agarwal, P. G. Williams, J. Dai, R. Neupane, J. Gurr, A. M. Rodríguez, A. Lamsa, C. Zhang, K. Dorrestein, B. M. Duggan, J. Almaliti, P. M. Allard, P. Phapale, L. F. Nothias, T. Alexandrov, M. Litaudon, J. L. Wolfender, J. E. Kyle, T. O. Metz, T. Peryea, D. T. Nguyen, D. VanLeer, P. Shinn, A. Jadhav, R. Müller, K. M. Waters, W. Shi, X. Liu, L. Zhang, R. Knight, P. R. Jensen, B. Ø. Palsson, K. Pogliano, R. G. Linington, M. Gutiérrez, N. P. Lopes, W. H. Gerwick, B. S. Moore, P. C. Dorrestein, and N. Bandeira. 2016. Sharing and community curation of mass spectrometry data with global natural products social molecular networking. Nat Biotechnol 34:828-837.
Watrous, J. D., and P. C. Dorrestein. 2011. Imaging mass spectrometry in microbiology. Nat Rev Microbiol 9:683-694.
Watrous, J., P. Roach, T. Alexandrov, B. S. Heath, J. Y. Yang, R. D. Kersten, M. van der Voort, K. Pogliano, H. Gross, J. M. Raaijmakers, B. S. Moore, J. Laskin, N. Bandeira, and P. C. Dorrestein. 2012. Mass spectral molecular networking of living microbial colonies. Proc Natl Acad Sci USA 109:E1743-E1752.
Wetmore, K. M., M. N. Price, R. J. Waters, J. S. Lamson, J. He, C. A. Hoover, M. J. Blow, J. Bristow, G. Butland, A. P. Arkin, and A. Deutschbauer. 2015. Rapid quantification of mutant fitness in diverse bacteria by sequencing randomly bar-coded transposons. mBIO 6(3):e00306-15.
Whitman, W. B., D. C. Coleman, and W. J. Wiebe. 1998. Prokaryotes: The unseen majority. Proc Natl Acad Sci USA 95:6578-6583.
Woyke, T., H. Teeling, N. N. Ivanova, M. Huntemann, M. Richter, F. O. Gloeckner, D. Boffelli, I. J. Anderson, K. W. Barry, H. J. Shapiro, E. Szeto, N. C. Kyrpides, M. Mussmann, R. Amann, C. Bergin, C. Ruehland, E. M. Rubin, and N. Dubilier. 2006. Symbiosis insights through metagenomic analysis of a microbial consortium. Nature 443:950-955.
Yang, Y. L., Y. Q. Xu, P. Straight, and P. C. Dorrestein. 2009. Translating metabolic exchange with imaging mass spectrometry. Nat Chem Biol 5:885-887.
This page intentionally left blank.








