8
Options for Mitigating Concerns
The study included consideration of opportunities to mitigate concerns related to the malicious use of biotechnology. The potential for mitigation was an integral part of the framework for assessing concern, as detailed in Chapter 3. As described in Chapters 4–6, considerations relevant to mitigation were included in the assessment of concern for specific potential capabilities, although these assessments did not include an in-depth analysis of current preparedness and response capabilities or speculate about the efficacy of various potential approaches. This chapter explores, from a broader perspective, some current mitigation approaches, how synthetic biology may challenge those approaches, and conversely, how synthetic biology may help address challenges or bolster mitigation approaches. A comprehensive, in-depth review of strengths and weaknesses in current U.S. or international programs was outside the scope of this study; as such, this report does not offer a full analysis of mitigation capabilities and makes no recommendations pertaining to mitigation priorities. Rather, this chapter is intended to provide useful context about fundamental mitigation concepts and approaches that arose during the course of the study, along with a brief exploration of some potential emerging challenges and opportunities.
CURRENT MITIGATION APPROACHES AND INFRASTRUCTURE
The mitigation of synthetic biology–enabled attacks essentially has two broad components: minimizing the chances of an attack and minimizing the negative outcomes once an attack has occurred. As discussed in Chapter 3, Potential for Mitigation, key elements that contribute to the potential for mitigation include deterrence and prevention capabilities, ability to recognize an attack, attribution capabilities, and consequence management capabilities. Broadly speaking, many of the same tools that are used to mitigate natural infectious disease outbreaks or exposure to chemicals (e.g., from environmental spills) are also relevant to mitigation of an intentional biological or chemical attack. In addition, the practices and rules in place to mitigate dual-use research may be relevant to some synthetic biology capabilities. The following sections provide a brief overview of selected existing mitigation approaches and infrastructures related to life sciences research, public health, emergency response, and healthcare capabilities that may be relevant to mitigating synthetic biology–enabled attacks.
Deterrence and Prevention Capabilities
Deterring or preventing the development and use of biological weapons, including those enabled by advances in synthetic biology, is of high priority for the U.S. Department of Defense (DoD) and for the nation. However, there are fundamental challenges to deterring or preventing misuse of biological advances. It has been noted that “the knowledge, materials, and technologies needed to make and use a biological weapon are readily accessible, everywhere in the world” (Gronvall, 2017). While fundamental research and clinical studies are the engines that drive public health and medical treatments, they simultaneously provide dual-use opportunities. Pathogens are ubiquitous, found in hospital and research laboratories, scientific culture collections, infected people and animals, and the environment. The skills and equipment applied to solving challenges in medicine, agriculture, and other disciplines for beneficial purposes are largely the same as those that would be used in making a biological weapon. Advances made in the age of synthetic biology add to the already-broad spectrum of biotechnologies that could be misused.
To support deterrence and prevention of misuse of biotechnology without unnecessarily hindering beneficial research, the prevailing approach has been to implement multiple overlapping tools that, when taken together, can provide greater value. These tools fall into two general categories: norms, and policies and regulations.
Norms and Self-Governance
Norms against the misuse of biology exist and are supported on many levels, from the global to the individual. The Convention on the Prohibition of the Development, Production and Stockpiling of Bacteriological (Biological) and Toxin Weapons and on Their Destruction, commonly known as the Biological Weapons Convention (BWC), is the cornerstone of international-level deterrence for biological weapons, including those created by synthetic biology (UNOG, 2017). The BWC bans such weapons, sets the standard for global norms, binds the nation-states that are party to the treaty, and defines acceptable behavior. There have been violations; for example, the Soviet Union maintained a secret bioweapons program after the treaty was ratified (Alibek, 1998; Cox and Woolf, 2002). However, no country goes against the international norm to flaunt an offensive biological weapons program; even North Korea, which openly flouts international prohibitions against nuclear testing, has denied accusations that the country is developing biological weapons (Sampathkumar, 2017). United Nations Security Council Resolution 1540, which prohibits states from assisting non–state actors in developing biological and other types of weapons, is another relevant international agreement (UN Security Council, 2004).
At the level of institutions and individuals, the scientific community has a tradition of self-governance and established norms entailing what constitutes responsible conduct in science. A landmark example is the 1975 Asilomar conference. With the advent of recombinant DNA technology, leading scientists recommended a moratorium on recombinant DNA experiments involving toxins, oncogenic viruses, and antibiotic resistance until their safety could be assessed (Berg et al., 1974). To facilitate that assessment, scientists and government officials gathered at a conference in Asilomar, California; after further research and national discussion, the moratorium was lifted in 1976, and a new guidance system was created for all recombinant DNA work funded by the U.S. government. What happened at Asilomar has become the template for scientists’ responses to scientific discoveries with social and ethical implications and a symbol of the scientific community’s capacity to self-govern.
In the decades since, this tradition of self-governance has been applied toward dual-use biotechnologies. In 2004, a National Academies report, known as the “Fink report” after the study’s chairman, geneticist Gerald R. Fink (NRC, 2004), made the case that scientists have a moral duty to avoid contributing to the advancement of biowarfare or bioterrorism and outlined types of experiments that would require consideration and review before being undertaken. These experiments—including those relevant to rendering a vaccine ineffective or conferring resistance to available therapeutics, evading detection or diagnosis methods, enhancing or creating virulence, increasing a pathogen’s transmissibility or altering its host range, or enabling weaponization—parallel the concerns considered in this report regarding uses of synthetic biology. The Fink report formed the starting point for a federal advisory committee of the U.S. Department of Health and Human Services (HHS) called the National Science Advisory Board for Biosecurity, which defined Dual Use Research of Concern (DURC) (U.S. Government, 2012)
and established the basis for a requirement that U.S. federally funded research involving certain regulated Select Agent pathogens (taken from the Federal Select Agent Program Select Agents and Toxins list; see CDC/APHIS, 2017) undergo DURC research review.
Another important area of self-governance relevant to synthetic biology is the voluntary screening of orders by vendors providing DNA synthesis services. Guided by a framework created by HHS in 2010, DNA providers are encouraged to screen orders for sequences of concern (e.g., DNA encoding Select Agents) and to screen customers to ensure that they are legitimate users of biology (HHS, 2010). Screening is intended to ensure that genetic material of regulated pathogens—including the causative agents of anthrax, smallpox, and rinderpest, for example—cannot be purchased without review and potentially consultation with government agencies. Screening is supported and facilitated by the International Gene Synthesis Consortium (IGSC), an international voluntary coalition of gene synthesis companies, which has adopted the 2010 HHS-recommended screening practices as well as even more stringent measures (IGSC, 2017; Cision PR Newswire, 2018).
Other examples of self-governance include work related to the responsible conduct of scientists (e.g., National Academies of Sciences, Engineering, and Medicine, 2017b,c), bioethics training for students, a life sciences professional code of conduct, and biosafety training for laboratory scientists. While the norms of self-governance are not going to deter or prevent a determined malicious actor from seeking to develop, obtain, or use a biological weapon (whether it is enabled by synthetic biology or not), these norms provide groundwork that could be built upon. At minimum, they offer a basis for social surveillance of unethical or malicious behavior within the scientific community.
U.S. Policies and Regulations
After the 2001 attack involving letters containing anthrax spores, the U.S. Congress strengthened several laws relevant to biosecurity and dual-use research, which resulted in the formal implementation of the Federal Select Agent Program (CDC/APHIS, 2017). In contrast to previous biosafety and containment guidance, which was geared toward equipping laboratory workers to perform experiments on dangerous pathogens without harming themselves or the public, the Select Agent program was designed to protect against unauthorized agent acquisition that might potentially result in the purposeful misuse of those specified agents and toxins deemed most harmful. The regulations require facilities handling listed pathogens to have physical security protections in place and to require individuals to undergo a security assessment before accessing agents on the list. For the most part, Select Agent regulations provide security through denial of access to pathogens, under the assumption that most bad actors would prefer the simplest method of gaining access to pathogens—stealing them from a laboratory.
Additional policies and requirements apply to researchers who receive U.S. federal funding for DURC, and these were recently reviewed by the National Academies (see National Academies of Sciences, Engineering, and Medicine, 2017b). These requirements (U.S. Government, 2012, 2014) stipulate that research using one of 15 pathogens or toxins or that falls within seven identified experimental categories is subject to additional oversight. Research proposals involving highly pathogenic avian influenza H5N1 also are subject to special evaluation by HHS. Although the government recently lifted a moratorium on gain-of-function experiments involving “pathogens of pandemic potential,” it specified additional review procedures that must be carried out before such experiments can be conducted (HHS, 2017a).
Some aspects of deterrence and prevention are based in the public health arena. For example, the availability and use of a vaccine or other countermeasure for a particular biological threat, in itself, can be a powerful deterrent—a bad actor is much less likely to use an agent for which the target population is impervious. Even in the absence of a specific medical countermeasure, a robust and healthy population, supported by strong public health infrastructure, can provide resilience against an attack. Conversely, the Ebola outbreak in Guinea, Sierra Leone, and Liberia that killed 11,310 people in 2014–2015 and impacted other countries including the United States is an example of what can happen during a natural outbreak of a serious infectious disease in the absence of a robust public health infrastructure. Kosal (2014) and others have reinforced the importance of strengthening public health infrastructure in all areas of the world as a strong deterrent to misuse of biotechnologies and as a way of enhancing international biosecurity.
Capability to Recognize and Attribute an Attack
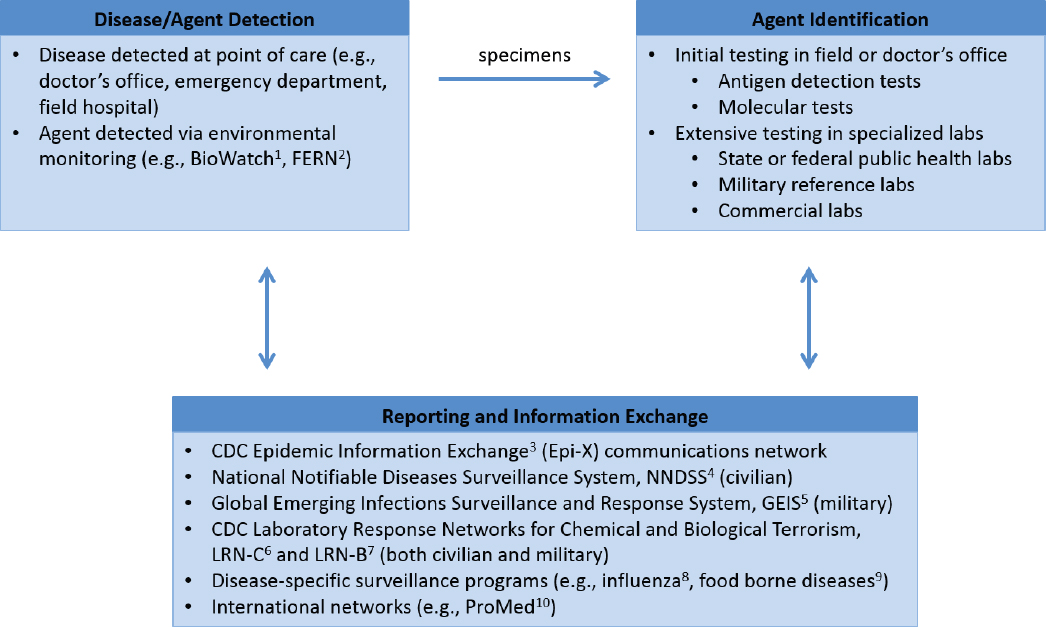
Other factors that contribute to mitigation relate to the capability to detect an emerging health threat, recognize it as a purposeful attack, and trace the attack to the actor responsible. Epidemiology, laboratory diagnostics, and environmental monitoring are essential components of systems to detect emerging health threats. Some of the procedures involved in disease surveillance and agent identification can also inform a determination of whether a health threat is the result of an intentional attack or a natural outbreak and potentially provide clues about the actor responsible. Figure 8-1 provides an overview of selected existing procedures and systems in place to identify emerging health threats affecting the U.S. public and military personnel.
In the United States, surveillance and reporting of infectious diseases occur at multiple levels and have both mandatory and voluntary components. Depending on local, state, or territorial jurisdictional requirements, healthcare providers, laboratories, hospitals, and other healthcare partners in the civilian arena must report the detection or suspicion of certain agents to their regional public health department and sometimes must submit samples for confirmatory testing at a public health laboratory. Once such a laboratory is involved, an alert is issued to support the identification of other cases of similar disease, and epidemiology becomes an essential factor in disease surveillance. In addition, the identification of certain pathogens (e.g., Select Agents) at these regional public health nodes requires notification of the U.S. Centers for Disease Control and Prevention (CDC) through the Laboratory Response Networks for Chemical and Biological Terrorism (CDC, 2014b,c). The DoD has a similar nodal system of large military reference laboratories, smaller regional laboratories, and local and point-of-contact care centers, referred to as a “soldier-provider-biosurveillance sentinel” approach. The DoD also operates a Global Emerging Infections Surveillance and Response system to monitor emerging infectious diseases (AFHSB, 2017), and DoD laboratories also participate in CDC’s Laboratory Response Networks.
To identify a pathogen, a specimen is typically compared against data available from organism banks or sequence databases, such as the Multidrug-Resistant Organism Repository and Surveillance Network (WRAIR, 2017), CDC’s MicrobeNet (CDC, 2017b), or GenBank® (NCBI, 2017). Direct antigen tests, supported by both military and civilian healthcare systems, use immunochromatographic methods to identify pathogens and can be conducted in the field or in any physician’s office. Increasingly, these tests are being replaced by newer platforms for point-of-care molecular tests, most based on polymerase chain reaction (PCR) technologies, which can rapidly detect bacteria, viruses, and parasites, and require little technical knowledge or sample handling (de Paz et al., 2014; Vidic et al., 2017). While they only target specific known and relatively common pathogens, molecular technologies can quickly rule in or rule out a known pathogen and provide more accurate and sensitive results than direct antigen tests. When tests available at the point of care are inconclusive or confirmatory testing is desired, specimens can be sent to public health, military, or commercial reference laboratories, which have a much more extensive capability based on in-house laboratory-developed tests. These tests, most based on real-time PCR or matrix-assisted laser desorption/ionization time-of-flight (MALDI-ToF) mass spectroscopy, require laboratory infrastructure and are more complex to perform and analyze, but they are capable of detecting a wider range of pathogens. The molecular identification methods used in the Laboratory Response Network laboratories and thus in the disease surveillance and reporting systems with which they interface are developed nationally and deployed via standardized methods to provide uniformity and comparability of results across each network. These efforts are also supported by extensive National Institute of Allergy and Infectious Diseases research and development efforts to advance methods for tracking, sequencing, and analyzing pathogens.
These surveillance systems support early detection and response when an emerging disease threat presents symptoms that are clearly apparent and can be linked to an identifiable pathogen or toxin. Surveillance networks in countries that have a robust public health system are also a valuable asset toward recognition of an attack, should one occur (see Kosal, 2014). However, such an attack would likely take longer to detect in less-developed countries or in war zones, which generally lack a strong public health infrastructure or for an agent that produces atypical symptoms. Another limitation is the temporal reporting delay between local and national recognition that an outbreak or attack has occurred.
To augment established surveillance and notification systems, public health authorities are exploring the use of a variety of newer networks and potential data sources. For example, the e-mail listserv ProMED-mail acted
NOTES: 1 BioWatch is a program of the U.S. Department of Homeland Security that monitors the air in public places for the presence of Select Agents (Firoved, 2016).
2 The U.S. Department of Agriculture’s Food Emergency Response Network (FERN) is responsible for detecting biological, chemical, and radiological contamination of food (FERN, 2017).
3CDC, 2017a.
4 National Notifiable Diseases Surveillance System, https://wwwn.cdc.gov/nndss. Accessed May 11, 2017.
5AFHSB, 2017.
6CDC, 2014c.
7CDC, 2014b.
8CDC, 2017c.
9CDC, 2017e.
10 The Program for Monitoring Emerging Diseases is a reporting system maintained by the International Society for Infectious Diseases, http://www.isid.org/promedmail/promedmail.shtml. Accessed January 25, 2018.
as an early warning system during the SARS (severe acute respiratory syndrome) outbreak in China in 2003 (Madoff, 2004); social media has been used to supplement traditional infectious disease surveillance tools (e.g., see Milinovich et al., 2014; Velasco et al., 2014; Charles-Smith et al., 2015; Young, 2015; Fung et al., 2016); and new data sources such as electronic medical records, search engine queries, data on pharmaceutical purchases, or longitudinal seroprevalence or biomonitoring studies (Klompas et al., 2012; Butler, 2013; Fung et al., 2015) could
potentially be mined for real-time disease surveillance purposes. Although these newer platforms are not validated data sources in surveillance and epidemiology—still requiring standards, advanced analytical capabilities, and resolution of privacy concerns (Chiolero et al., 2013; Friedman et al., 2013)—they could be valuable tools for earlier detection of natural or intentional disease events in the future.
Consequence Management Capabilities
Two key capabilities for containing and responding to a chemical or biological attack (consequence management) are the ability to limit the spread of transmissible agents and the ability to counter an agent with vaccines, therapeutics, or other tools.
Methods to Limit the Spread of Transmissible Agents
CDC provides clear definitions of classic infectious disease mitigation measures such as the isolation of infected individuals (CDC, 2014a). Isolation and quarantine, along with contact tracing and travel restrictions, were used to great effect to limit the spread of SARS during the 2003 outbreak (Anderson et al., 2004). The effectiveness of such public health measures is highly dependent on the basic reproduction number, known as R0, and the serial interval of the pathogen in question. In addition, while such measures tend to work well in a military setting, they can be more difficult to implement in a civilian setting due to poor acceptability and other social factors, as was the case in the United States during the 2015 Ebola outbreak. Other relevant measures to limit the spread of agents include personal protective equipment such as impermeable body suits, gloves, and respirators used to protect emergency workers from contamination when working in the field (FDA, 2017c).
Medical Countermeasures
Medical countermeasures include biological products, drugs, and devices approved by the U.S. Food and Drug Administration (FDA) to prevent, treat, or ameliorate illness in the event of a public health emergency caused by an infectious agent, toxin, or chemical, whether natural or manmade. These include devices such as personal protective equipment, along with vaccines, antibiotics, antivirals, antitoxins, and other drugs and therapeutics.1
HHS and the DoD share responsibilities for the development of medical countermeasures, targeted at agents on the Select Agent list, in conjunction with Material Threat Assessments provided by the U.S. Department of Homeland Security (see the Public Health Emergency Medical Countermeasures Enterprise Strategy and Implementation Plans [HHS, 2017b]). Limitations in research capacity, funding, and clinical capabilities necessitate careful decisions about which medical countermeasures can be feasibly developed, from their inception to animal testing, scale-up, clinical testing, and manufacturing. It is also difficult to engage pharmaceutical manufacturers to invest time and platforms into medicines that may not show significant return on investment. Considerations related to how these measures are manufactured (typically on an on-demand basis) and dispensed to populations are also important. Although some countermeasures are placed in the Strategic National Stockpile (maintained by CDC), which supplies state and local public health agencies with medical countermeasures in the event of a national emergency (CDC, 2017f), inventories of many countermeasures are extremely limited and are likely to be sufficient for only the first days of an outbreak situation.
MITIGATION CHALLENGES POSED BY SYNTHETIC BIOLOGY
The mitigation measures described above have strengths and weaknesses despite the advent of synthetic biology. Synthetic biology brings some of those weaknesses into sharper relief, creates new challenges, and creates opportunities for improving mitigation capabilities.
___________________
1 For further information on public health medical countermeasures, see FDA, 2017c.
Challenges to Deterrence and Prevention
Taken together, strategies such as norms and self-governance, voluntary guidance, regulations, and international bans provide numerous barriers to the misuse of biological research that are potentially larger than the sum of their individual parts. However, these strategies, many of which lack formal enforcement mechanisms, have been criticized over the years as insufficient to guard against the purposeful misuse of biology (Palmer et al., 2015). At the international level, for example, the BWC has influenced norms but has few effective enforcement mechanisms. Concerns about the weaknesses of these strategies have gained greater traction with the emergence of synthetic biology. The following sections discuss two areas in which synthetic biology has raised particular concern: the accessibility of modern biotechnology to a wider range of actors and the pitfalls of list-based screening to detect malicious activity.
Accessibility of Biotechnology
Biology today is conducted in a markedly different environment than that of the 1975 Asilomar conference, the seminal event that set the model for scientific self-governance. There is now not only an expanded array of tools available, but a far more diverse scientific community. Synthetic biology techniques are accessible to a wide variety of people, including traditional academic and commercial researchers but also amateur biologists, nonbiologist engineers, and manufacturers, not all of whom are steeped in the norms of traditional academic settings. Some have also argued that tacit knowledge is becoming less central to successful biological manipulation thanks to the increasing sophistication of information technologies (Revill and Jefferson, 2014). As noted in Chapter 2, the movement toward making biology “programmable” broadens the array of actors who may be capable of engineering biological components, although the pace and ultimate degree to which biology is and will become “programmable” is a matter of some debate.
In addition to traditional pathways for entering biotechnology—working in academic laboratories, obtaining a graduate degree, and pursuing a traditional postdoctoral fellowship—people can now enter the field through nontraditional ways. For example, do-it-yourself (DIY) models of biological experimentation have gained popularity in recent years, offering nonscientists the tools and guidance for performing biological research. As biotechnology industry analyst Rob Carlson wrote in Wired in 2005, “the era of garage biology is upon us,” noting that a person could, with a few thousand dollars of investment, get to work “hacking biology” (Carlson, 2005). The community has grown since then; in 2017, a “Global Community BioSummit” was organized at the Massachusetts Institute of Technology (MIT) Media Lab, which brought together “biohackers” and members of independent and community laboratories from dozens of countries (MIT Media Lab, 2017). Many DIY biology activities are expressly educational, fun, or tied to local community needs (e.g., testing food samples). Yet while most of these DIY projects are not sophisticated, the model does make accessible to the general public tools that can be used to do advanced work. For example, for less than $200, reagents and kits can be acquired that enable amateurs to employ gene-editing technologies such as CRISPR/Cas9, although advanced skills and additional laboratory resources would likely be required to use such kits to create a harmful agent. It is also possible that community laboratories could provide a venue for malicious actors or be implicated as misdirection in a perpetrated event.
Another example of a nontraditional group of biotechnologists is the International Genetically Engineered Machine competition (iGEM, 2017b). iGEM began in 2003 as an in-class competition at MIT in which teams of students were challenged to build synthetic biological systems from standard, interchangeable parts, called Bio-Bricks™,and operate them in living cells. Though iGEM projects are carried out by students, many of them entirely new to bioscience, some projects have been quite sophisticated. Now an annual event open to participants outside of MIT, iGEM involves students at the high school, undergraduate, and graduate levels from countries around the globe. Projects routinely involve the engineering of microbial, mammalian, and plant cells; the 2014 grand prize winner, for example, circularized proteins to make them more physically stable.
The fact that a relatively untrained individual could perform complex bioengineering has triggered concerns and mechanisms to improve the safety and knowledge of the amateur community’s activities (Kellogg, 2012; Holloway, 2013; Kolodziejczyk, 2017). A “see something, say something” campaign of the Federal Bureau of
Investigation (FBI) performs outreach to both the DIY biology community and to iGEM (Wolinsky, 2016). The FBI and the American Association for the Advancement of Science have also teamed up to increase understanding of the risks and benefits of the field (Lempinen, 2011) and explore ways to “safeguard science.”
Pitfalls of List-Based Screening
Advances in synthetic biology capabilities pose a number of challenges to list-based screening as a key tool for deterrence and prevention. In particular, the voluntary screening of orders by DNA providers, a system intended to prevent production of Select Agents, is becoming less useful (Casadevall and Relman, 2010; Carter and Friedman, 2015; DiEuliis et al., 2017b). While screening of customers is and will likely remain an important tool, recent research examples indicate that screening of the sequences ordered by those customers may become less relevant. Using lists may make it easier to implement policy, but a static list-based approach is concerning not only because many pathogens exist in nature, but because synthetic biology now allows for the creation of new pathogens and other potentially harmful biological components that are not found on such lists.
Sequence screening is based on homology to “data from all organisms on the Select Agent list, the Australia Group List, and other national lists of regulated pathogens” (IGSC, 2017), so if an agent is not on the list, it is not flagged. For example, current guidance did not prohibit a DNA provider from fulfilling an order for the genome of the extinct virus horsepox; the recent publication of the synthesis and booting of the horsepox genome (Noyce et al., 2018) raised concerns that some techniques employed to create this pox virus could be applicable to creating smallpox (DiEuliis et al., 2017a; Koblentz, 2017) because horsepox has high sequence similarity to variola virus, the causative agent for smallpox (Tulman et al., 2006). In addition, while there are processes to connect synthesis companies with U.S. law enforcement agencies in the event of a problem, DNA synthesis is performed worldwide, and it is less clear that such processes are in place in all other nations. Importantly, in addition to DNA synthesis screening, lists such as the Select Agent list also form the basis of many of the downstream mitigation tools discussed in this chapter, including detection, diagnostics, and the development and prioritization of medical countermeasures. An overreliance on the Select Agent list is a systemic weakness affecting many aspects of the United States’ current biodefense mitigation capability.
Another weakness is that DNA sequences of less than 200 base pairs (known as oligonucleotides) are not screened. This has raised concerns that a determined malicious actor could potentially obtain multiple short sequences from commercial vendors and assemble them to create full-length pathogen DNA, although such a strategy would require significant effort and skill, particularly for pathogens with large genomes. It has been argued that screening oligonucleotide orders is unworkable due to a higher expected false positive rate for any given short sequence, which would be exacerbated by the much higher volume of oligonucleotide orders (Garfinkel et al., 2007; Carter and Friedman, 2015). A counterargument has been put forth that oligonucleotide screening could be performed differently than for longer genes, such as by analyzing groups of oligonucleotides in an order (or across multiple orders) and setting sequence similarity thresholds to higher values. Another concern is that evolving trends in the life sciences enterprise may erode vendors’ incentives for screening. As DNA synthesis becomes cheaper, the somewhat fixed cost associated with screening represents an increasingly larger percentage of total costs, creating a disincentive against screening on the part of those companies (DiEuliis et al., 2017b). These costs could be especially acute if oligonucleotide screening were implemented.
Current screening approaches are primarily based on the homology of a sequence order to the sequence of a specified pathogen, as opposed to screening for sequences that confer specific pathogen characteristics. As further understanding is gained connecting sequence to function, there is an opportunity for the types of lists used to evolve. Thus, some form of list-based mitigation could continue to play a role in the deterrence and prevention toolkit, even if this strategy has limitations and will need to be part of a layered approach that includes other strategies (see Opportunities for Improving Deterrence and Prevention Capabilities, below).
Challenges to Recognizing and Attributing an Attack
In a textbook world, approaches to surveillance for disease outbreaks are based on the appearance of clear disease symptoms in a group of individuals connected in place and time and which can be attributed to a causative agent. The recent Zika outbreak in the Americas is a good example of how these “perfect conditions” are not always met. Eighty percent of Zika-infected individuals showed no signs of disease, symptoms were mild even in those who were symptomatic, and the link to microcephaly in infants born to infected women could not have been predicted. Such examples underscore remaining weaknesses in disease surveillance tools for recognizing even natural disease outbreaks; these weaknesses may create particular challenges with regard to some types of synthetic biology–enabled attacks. For example, as discussed in Chapter 6, it may be possible to develop bioweapons that alter the human host and produce health effects that are not immediately obvious as a disease outbreak or attack, such as by reducing immunity or modifying the microbiome.
Synthetic biology could also confound the ability to identify the causative agent in a biological attack. Despite the breadth and depth of available repository resources, there would not always be a reference specimen to use as comparator, particularly if the agent is markedly different from natural pathogens or toxins. Many current mitigation efforts are inherently list based (aimed at detecting Select Agents) and are heavily dependent upon the secrecy of the exact genomic regions used for the PCR primers and probes; should an adversary determine what these regions are, it could be possible to create a functional yet undetectable pathogen by altering those regions using codon-switching techniques.
In addition to challenges related to clinical surveillance, synthetic biology could also further compound weaknesses in environmental surveillance capabilities, which seek to detect agents in the environment to provide early warning before patients present in the healthcare system. For chemical threats, the Laboratory Response Network for Chemical Terrorism utilizes several forms of mass spectroscopy, which makes unbiased detection much more feasible (assuming reference standards are available) than in the biological field, where unbiased detection remains extremely challenging. Although it is feasible to utilize PCR to identify a Select Agent pathogen “needle” from the enormous environmental background “haystack,” there is no technology available today that can reliably alert us when a novel pathogen, whether natural or engineered, is present in the environmental background. These tools will not be useful in detecting unknowns, genetically engineered chimeras, or agents for which the PCR primer or probe binding site has been altered. Ultra-deep metagenomic sequencing will find vast amounts of uncharacterized sequence in any environmental sample, and sorting it all out to the point where a novel pathogen can be definitively identified is currently too costly and too lengthy a process to be useful. Bioinformatics tools provide powerful means of sifting through seas of sequences, but they rely on assumptions, for example, about what constitutes a taxonomic unit, and the incompleteness of available reference databases affects the accuracy of the results. An additional complication is that the “normal” background microbial composition is poorly characterized for many outdoor and indoor environments and can be affected by many factors (National Academies of Sciences, Engineering, and Medicine, 2017d). Given these challenges, approaches such as metagenomics and environmental surveillance are not likely to completely fulfill the need to provide early identification of agents used in synthetic biology–enabled attacks.
If current environmental surveillance methods are not capable of recognizing a novel agent, the implication is that we are dependent upon the public health system to recognize outbreaks of novel pathogens, whether natural or engineered. Relying on this reactive approach suggests that it would not be possible to act to mitigate or contain an outbreak until patients have developed symptoms that trigger a health community response; as a result of this delay, people would become ill before it is possible to know that an attack has occurred. Isolation of the novel causative agent by culturing (if possible) followed by sequencing or ultra-deep sequencing and painstaking assembly would be needed to characterize the agent and lay the groundwork for analyzing its mechanisms and origin. This initial characterization process might take a few days at best, or considerably longer if the novel agent is a highly engineered version of a normally benign microbe or is no longer present in the patient by the time symptoms are apparent. In cases in which the agent is a pathogen, PCR reagents can be developed quickly once the genome has been obtained, at which point the agent can be added to the list of agents detectable through environmental and clinical surveillance systems.
There is no magic bullet for dealing with all new routes to harm that are made possible by modern biotechnology, including synthetic biology, nor are there magic bullets for handling every natural agent that emerges, as exemplified by experiences with SARS, MERS (Middle East respiratory syndrome), West African Ebola, and other outbreaks. The 2003 SARS outbreak in particular underscored to the international public health and biosurveillance communities the need to have mechanisms in place for rapid characterization and international information sharing to respond adequately to novel and emerging threats. The types of biosecurity concerns related to synthetic biology assessed in this report provide added urgency to that message.
Consequence Management Challenges
If disease surveillance and laboratory infrastructure cannot detect, identify, and characterize the causative agent, it is also possible that current available medical countermeasures—such as vaccines and therapeutics—may be less effective or, in certain cases, ineffective. While existing medical countermeasures may be quite useful for containing or counteracting agents created with synthetic biology that are highly similar to existing pathogens of concern, not all agents may fit this model. For example, if multiple drug resistance mutations are introduced into a bacterium to produce a bioweapon, even a broad-spectrum antibiotic administered before the agent is fully characterized may be ineffective. Similarly, if a viral chimera is engineered bearing novel surface antigens, it is unlikely to be neutralized by immunoglobulin given post-exposure. In short, if the agent is not susceptible to available vaccines, drugs, or antibody-based therapeutics, existing systems are less likely to limit its spread, potentially increasing the scope of casualties. In such scenarios, developing, testing, and approving drugs and vaccines to counter the agent using traditional approaches would entail long delays and an associated likelihood of many people being affected, suggesting a need for novel approaches to rapidly manufacture and test new therapeutics. Effectively implementing such approaches would require not only technological advancement but also rapid regulatory approval processes, such as the Emergency Use Agreement mechanism used by the FDA.
POTENTIAL OPPORTUNITIES TO ADVANCE MITIGATION CAPABILITIES
Despite the challenges posed by the current and anticipated biological threat landscape, there are multiple opportunities to build upon current capabilities and fill some of the gaps. In fact, synthetic biology capabilities may themselves help advance some mitigation efforts. Providing a comprehensive list of technologies with sufficient information to judge their efficacy in dealing with novel outbreaks is outside the scope of this report. This section is intended to highlight some of the ways in which technologies currently in development could improve the ability to handle future outbreaks or attacks, including selected examples of potential opportunities for improving the capacity for deterrence, prevention, attack recognition, attribution, and consequence management.
Opportunities for Improving Deterrence and Prevention Capabilities
Engineering techniques such as abstraction, standardization, modularity, automation, and rational design are likely to enable significant advances in synthetic biology. While the degree of incorporation of computation into the synthetic biology workflow will vary, one opportunity to explore mitigating biodefense concerns, for those approaches that depend on computational engineering, is to explicitly integrate mechanisms to prevent, detect, identify, and store information about malicious activities in the computational infrastructure. This approach could be relevant to all aspects of mitigation but is perhaps most salient for prevention and attribution. Examples of types of approaches that could be further explored are discussed below. Box 8-1 outlines how such approaches might be applied to identify or prevent malicious activity at various stages of two example scenarios.
- Screening of activities with machine learning: It may be possible to develop algorithms that learn and recognize patterns, such as DNA segments or sequence transformations, material transfers, or equipment usage, that relate to the creation of a biological threat. This approach could potentially help flag suspicious activity early in the design cycle. However, developing such algorithms requires a large amount of training
-
data, and data reflecting malicious activity would be hard to come by; as a result, developing a sufficiently accurate algorithm may be infeasible.
- Systems to constrain design capabilities: Rules could potentially be encoded directly in software for engineering DNA constructs to make it difficult or impossible to create specific genetic designs, for example, by prohibiting or requiring the addition or removal of specific DNA segments, requiring specific assays, preventing the transfer of materials to specific individuals or entities, or excluding or requiring the use of specific host organisms. Although this approach could help deter or prevent some malicious activities, it would not be sufficient to prevent designs based on specific knowledge or on brute-force combinatorial testing that bypasses biological design tools and could be difficult to implement in a way that prevents user tampering.
- Maintaining registries of known expertise and materials: Database infrastructure and supporting tools could be created to track known sources of expertise and materials relevant to the capacity to produce a biological threat, such as information about laboratories, personnel, and sources of material. In addition to identifying relevant players, it could be possible to profile designs coming from them to create known “digital signatures” of the engineering designs of individuals or groups. However, obtaining access to sufficient designs to be able to profile malicious users would be difficult, as distinguishing legitimate activities would be.
- Maintaining registries of known biological threats: Despite the inherent limitations of list-based systems in light of synthetic biology capabilities, there may nonetheless be opportunities to enhance the utility of these systems by systematically connecting them to design software and to automated foundries. Furthermore, there is an opportunity for screening procedures to move from a focus on organisms to a focus on DNA functions. It has been argued that emphasizing known pathogenic functions (as opposed to whole genomes of Select Agents; see IARPA, 2017b) would allow the curation of a more meaningful registry, one drawn directly from the DNA components responsible for causing harm. For example, software used for synthetic biology could be required to periodically run “checks” against bioagent registries or to automatically add new biological threats to these registries when they are identified. For such an effort to succeed, it would need to be scalable, searchable, and resistant to hacking. Malicious users would presumably be constrained to other approaches that do not rely on design software, such as experimental approaches like DNA shuffling or mutagenesis.
- Tracking digital “signatures” in genetic designs: It may be possible to deploy information technology at key stages in the automation pipeline to identify the source and the creator of synthetic genetic material to ensure that it comes from trusted sources. Were an attack to occur, this information could also help to identify the actor responsible. However, this approach would largely be applicable to strategies employing genetic circuit design tools; attribution of synthetic materials created by other means, such as through directed evolution, would be much more difficult. Watermarks for this purpose could be “biological,” for example, if the genetic material (e.g., the DNA sequence) has additional information inserted that uniquely identifies the sample (Heider and Barnekow, 2008), or the watermarks could be “electronic,” for example, if the information is added digitally to the electronic file used to communicate the biological information (e.g., in the binary information that encodes a GenBank® file) (Cox et al., 2008). Electronic watermarks are more mature and more likely to be more useful in practice where the biological material is manipulated.
Opportunities for Improving Agent Identification and Attribution Capabilities
Because so much of the natural nucleic acid space has yet to be sequenced and characterized, it remains extremely difficult to determine if a given genetic sequence is of natural or nonnatural origin. However, current analysis methods can help identify situations in which gene sequences appear in unexpected places (e.g., identifying that the toxin gene from Clostridium botulinum has been inserted into the genome of Escherichia coli). In addition, the products of genetic circuit engineering (see Chapter 4, Figure 4-3) can clearly be recognized as nonnatural and even contain design patterns that may provide attribution clues. Additional tools that enable one to detect that a
sequence had been genetically manipulated, or tools to analyze features of a sequence or a resulting organism that contribute to actor attribution, would be valuable additions to mitigation strategies.
Although many U.S. government agencies have expertise and responsibilities relevant to preparing for, preventing, and responding to an attack involving engineered biological components, no single agency has lead
responsibility in this area. The 2001 Amerithrax letter attacks first brought focus on bioterrorism and the need for the federal government to build standardized software tools and laboratory methods to analyze engineered organisms. Several recent examples are summarized briefly below.
- Safe Genes (DARPA, 2017), a program of the Defense Advanced Research Projects Agency, focuses on developing strategies to better control genome editing activity, such as by inhibiting genome editing in cells or preventing off-target editing activity.
- Functional Genomic and Computational Analysis of Threats (Fun GCAT; IARPA, 2017b), a program of the Intelligence Advanced Research Projects Agency (IARPA), aims to facilitate the design of better tools for screening DNA synthesis orders.
- Finding Engineering-Linked Indicators (FELIX; IARPA, 2017a), another IARPA program, seeks to develop a suite of tools designed to distinguish natural organisms from animals, bacteria, insects, plants, and viruses that have been engineered to potentially cause harm.
- To help reduce risk, the U.S. Department of Homeland Security sponsors the Sequences of Interest database to bring together in a single source nucleic acid and protein data about genetic mechanisms of virulence and resistance, along with protein toxin data and nucleotide data about plasmids and artificial vectors that may signify natural or artificial bacterial genetic change (D. Shepherd, Chemical-Biological Defense Division, Department of Homeland Security, personal communication, 2018).
While these or other programs were not evaluated as part of this study, they represent examples of the kinds of investments that would increase preparedness for the types of synthetic biology–enabled capabilities discussed in this report.
As discussed in Chapters 4–6, synthetic biology techniques can be used to modify pathogens, hosts, and vector species; these agents could possibly be used in complex attacks involving multiple pathogens, hosts, or vectors. Under the public health paradigm, identifying an agent’s species and any antimicrobial resistance factors is generally sufficient to guide treatment, for example, with a particular antibiotic. However, that level of information may not be sufficient for forensics and attribution, particularly if a deliberate attack or engineering is suspected. In these cases, responsible federal agencies will want to know how similar the new sample is to strains in the sequence databases, whether it is a common laboratory strain or a strain from a different part of the world, how the new sample compares to strains found at suspected facilities, and the degree of certainty with which we can determine whether the agent is a natural strain or might have been raised in a particular type of culture media, for example. Except in cases in which leftover samples are found in the laboratory where the material was created, proving attribution in the era of synthetic biology appears to be growing increasingly difficult, particularly for complex attacks that could potentially take considerable time to achieve their intended effects. As a result, attribution in the age of synthetic biology is likely to be heavily dependent on computer-based approaches that look for molecular signatures, as well as on intelligence. It is not within the scope of this report to discuss intelligence activities, and it is recognized that highly sophisticated adversaries may be able to evade even the most elaborate attribution approaches.
One of the most significant developments for identifying agents (in the context of treatment as well as detection and attribution) is next-generation sequencing and the drastic reductions in cost and time it enables. The FBI-led analysis of the 2001 Amerithrax attack samples (which took place before the advent of next-generation sequencing) involved the sequencing of a small number of morphologically different isolates at a cost of around $100,000 each in a process taking several years. Were such samples to be analyzed using today’s tools, ultra-deep characterization of the sample (about 10 billion sequence reads from a full run on a HiSeq™ sequencing system) could be performed within 1 week with reagent costs of around $10,000. Looking to the future, it is clear that next-generation sequencing will become central to identifying synthetic biology–derived infectious agents. Box 8-2 describes some of the ways in which next-generation sequencing approaches might be used in this context.
Synthetic biology is also likely to lead to the development of new detection technologies. As an example, Pardee et al. (2014) developed a programmable diagnostic assay that is embedded in paper as a low-cost, sensitive diagnostic assay for the presence of Zika virus RNA (Hall and Macdonald, 2016). In another novel approach to diagnostics, Lu et al. (2013) describe the engineering of bacteriophages for diagnostic strategies in which phage-specific antibodies, quantitative PCR, or a reporter molecule are used to detect amplification of engineered phages when the phages encounter target bacteria. Slomovic et al. (2015) describe applications of synthetic biology in the development of both in vitro and in vivo diagnostics, including the development of sensing bacteria in which
“sentinel bacteria could reside in the guts of soldiers or aid workers and serve as short term ‘medical records’ alerting on the time and scale of contamination or pathogen infection.” These studies, while still in a research mode, suggest that synthetic biology tools can help address some of the need for alternative diagnostics that are not based on detecting a specific region of a pathogen by real-time PCR.
Opportunities for Improving Consequence Management Capabilities
Just as synthetic biology expands the types of malicious activities that may be undertaken, it also expands what is possible for beneficial applications. Synthetic biology and related advances (such as the convergent technologies discussed in Chapter 7) open the possibility of new and more systematic approaches to the development of medical countermeasures and other mitigation tools and strategies. Synthetic biology approaches such as rapid DNA synthesis, protein design tools, cell-free expression systems, and automation may significantly advance consequence management capabilities, especially with regard to the development and testing of medical countermeasures. Such approaches could, for example, provide flexibility in the control of protein expression levels, shorten the time to successful countermeasure production, and lower costs. They could potentially even enable the development of countermeasures to newly identified agents without ever culturing the agent itself; through the use of in silico characterization of an agent’s key components, antigen components for antibody development could be synthesized, potentially within hours of detection. Such approaches could represent a promising alternative to stockpiling countermeasures when the emergence of novel threats (both natural and engineered) is likely.
In addition, once bioagent and viable culturing conditions have been identified, the large-scale testing capabilities used in synthetic biology could be used to screen candidate countermeasures, for example, by surveying chemical small-molecule libraries to identify drug leads or by testing many organism-relevant phages to identify those that are potentially lethal to the bacterial strain used in an attack.
The following sections discuss ways in which synthetic biology could potentially contribute to the development of diagnostics, vaccines, and other medical countermeasures. However, the technical barriers to the development of synthetic biology–enabled vaccines or therapeutics remain steep, and it is also important to note that there must be a compelling business case for their development and a regulatory process for approval of these countermeasures before they become reality. Almost 4 years after the emergence of the Ebola virus infection in West Africa, we still lack licensed Ebola vaccines, and despite knowing the serious risk of a MERS outbreak outside of the Arabian Peninsula, we are still many years away from a licensed effective MERS vaccine. While outside the scope of this report, a comprehensive understanding of the feasibility of using synthetic biology to develop medical countermeasures would benefit from critical review of both commercial and regulatory considerations.
New “Vaccine Strains” Through Controlled Attenuation of Viruses
The replication cycle of viruses is complex, and the fitness of a given virus depends on many factors. One important factor is the particular codons incorporated into the DNA or RNA; the preferential use of particular codons (or codon pairs), termed codon bias (or codon pair bias), is thought to influence the efficiency of translation (Buchan et al., 2006). Efforts to optimize codon usage almost invariably result in attenuation of the virus, and the more the codon usage bias is disrupted, the more attenuated the resulting virus (Wimmer and Paul, 2011; Martinez et al., 2016).
Burns et al. (2006) and Coleman et al. (2008) proposed to take advantage of this attenuating phenomenon to perform genome-scale manipulation of codon pair bias in poliovirus to develop vaccines in which the degree of attenuation could be controlled by the degree of codon substitution performed. The resulting “vaccine strains” provided protective immunity in mice and, because of the hundreds of substitutions made, did not revert to virulence. Using synthetic biology tools including large-scale, low-cost construction of desired genomic sequences has been proposed as a means of making attenuated vaccines for many other RNA viruses, including influenza virus (Mueller et al., 2010; Yang et al., 2013; Fan et al., 2015), chikungunya virus (Nougairede et al., 2013), respiratory syncytial virus (Meng et al., 2014), simian immunodeficiency virus (as a model for HIV; Vabret et al., 2014), tickborne encephalitis virus (de Fabritus et al., 2015), vesicular stomatitis virus (Wang et al., 2015), and dengue virus (Shen et al., 2015).
Use of DNA Construction to Rapidly Derive Vaccine Stocks
The 2009 H1N1 pandemic made it clear that new methods of developing influenza vaccines were required to speed the response from emergence of a new virus to the development of a vaccine seed stock and production and distribution of the vaccine strain. Toward this goal, Dormitzer et al. (2013) developed a synthetic approach, constructing the hemagglutinin and neuraminidase genes with minimal errors by annealing many staggered oligonucleotides that overlapped by 30 bases with their neighbors and together covered the full length of each gene. Infectious virus was rescued from susceptible cells transfected with the synthetic hemagglutinin and neuraminidase genes and plasmid DNAs encoding viral backbone genes. In a proof-of-concept study performed in collaboration with the Biomedical Advanced Research and Development Authority, an H7N9 vaccine strain was constructed in this manner in 5.5 days; tests demonstrated the antigens expressed by the synthetic genes were immunogenic based on their reaction with ferret sera (Dormitzer et al., 2013). This example demonstrates that synthetic biology tools can facilitate the rapid derivation of vaccine strains to respond to emerging viral threats. However, the commercialization and licensure of vaccines derived in this manner is many years off; having a synthetic biology tool that can facilitate the development of a new countermeasure is a major advance, but it is far short of what is necessary to make that countermeasure safe, effective, and available.
Rapid Development mRNA Vaccines
Another approach to the development of synthetic vaccines is the use of messenger RNA (mRNA). Petsch et al. (2012) demonstrated that mRNAs of influenza hemagglutinin, neuraminidase, and nucleoprotein could be transcribed into proteins in vitro to provide protective immunity against homologous influenza virus. Hekele et al. (2013) used a synthetic self-amplifying mRNA (SAM) to create a vaccine derived from the hemagglutinin gene of the H7N9 influenza virus delivered by a nanoparticle. The vaccine, produced just 8 days after the sequence became available, was immunogenic at low doses. SAM vaccines delivered by nanoparticles have also been developed against HIV-1 (Bogers et al., 2015) and Zika virus (Pardi et al., 2017). In a further development, Richner et al. (2017) also developed a SAM vaccine against Zika virus delivered by nanoparticles but, in that case, a structural gene from the Zika virus was engineered to destroy a conserved epitope to eliminate the production of cross-reactive antibodies against dengue virus, which would exacerbate dengue disease. These examples raise the speculative possibility that self-amplifying mRNAs directly encoding antibody molecules and delivered by nanoparticles could be used as a potential therapeutic approach. However, as with the example in the prior section, because of regulatory and business factors, it would take years before this approach produces therapeutic applications for use.
Use of Synthetic Biology Tools to Develop New Therapeutics
Synthetic biology is also contributing to the development of small-molecule medical countermeasures. The development of a yeast strain capable of producing artemisinic acid, the key precursor to the antimalarial drug artemisinin, demonstrated that complex plant-based natural products can be produced via synthetic biology (Westfall et al., 2012). More recently, compounds such as opioids (Galanie et al., 2015) and penicillin (Awan et al., 2017) have similarly been produced in yeast. Development of existing and novel chemicals and materials remains a primary interest of both the academic and industrial community, making it likely that the cost and time to develop chemical production strains will improve in the future.
Krishnamurthy et al. (2016) summarized the use of synthetic biology tools in the development of new therapeutics, including approaches for the production of new antibiotics and the application of the CRISPR system in developing bacteriophages as targeted therapeutics. The enabling impact of synthetic biology in exploring the great diversity of natural products that can be used as therapeutics is reviewed by Smanski et al. (2016). Platforms for drug discovery can be envisaged using synthetic mammalian genetic circuits, and bacteria, yeasts, and plants engineered with synthetic pathways can be utilized for the large-scale production of drug and drug precursor compounds (Weber and Fussenegger, 2012).
In addition to rapid response with conventional countermeasures, such as antibodies and small-molecule drugs, synthetic biology may also enable the deployment of new types of countermeasures. For example, gene drives and other gene editing methods are being explored for the control of vector populations for illnesses such as malaria and Lyme disease (Harris et al., 2012; Esvelt et al., 2014; Hammond et al., 2016). Microbiome-based interventions for the control of gastrointestinal infections could also provide a programmable platform for combating bacterial threats. For example, Citorik et al. (2014) have described the use of CRISPR/Cas technology to create RNA-guided nucleases that act as antimicrobials by targeting specific DNA sequences. These RNA-guided nucleases enable modulation of complex bacterial populations by selective knockdown of targeted strains.
SUMMARY
The ability to respond to a disease outbreak, whether it emerges naturally or from a purposeful attack, is complex and dependent on many social, governmental, and biological factors. Recognizing that an outbreak has occurred is a vital step in this process. Then, the agent must be identified and medical countermeasures made available. The prospect that a causative agent may have been created with synthetic biology and is therefore unknown
and uncharacterized dramatically increases the complexity of these mitigation activities and underscores the need to improve the public health response system.
In light of this context, it will be vital to maintain the current systems used in the military and civilian public health infrastructure. Strengthening this infrastructure in specific areas, including broadening the current approaches to surveillance, is important to better enable the detection of an attack that does not elicit “normal” symptomology.
Although an in-depth analysis of preparedness and response capabilities was outside the scope of this report, identification and characterization of an agent derived by synthetic biology may be a significant gap in the nation’s preparedness because many current diagnostic capabilities are based on commonly seen human pathogens and on lists of pathogens designated as high risk. Untargeted approaches to detection that use multiple platforms and integrate the data obtained would be expected to be more effective at identifying and characterizing unknowns. It is also clear that while advances will need to be made in wet-bench detection technologies, computer-based interrogative and forensic methods will become more and more valuable to support prevention, agent identification, and attribution. Large-scale success of computational mitigation requires that the attack strain has been developed by rational engineering design approaches that are not yet ubiquitous; the development of agents with other approaches such as directed evolution will likely remain difficult to prevent or attribute. The difficulty of affirming attribution to the level of certainty required for counteractions or incarceration is considerable, even for “traditional,” non-engineered bioweapons.
Finally, synthetic biology is enabling advances in the rapid development and production of medical countermeasures that may be effective against synthetic biology–derived agents. However, many such efforts, which are being pursued in both industry and academia, are still in the research phase, and there remain complex barriers to widespread use of these novel approaches, including regulatory hurdles and hurdles to industry involvement. This field needs to be monitored carefully over time.




















