2
Biotechnology in the Age of Synthetic Biology
To frame and guide the study, the relationship of synthetic biology to other areas of biotechnology was explored along with the context in which synthetic biology tools and applications are being pursued. This chapter describes, in the context of this study, what it means to be in “the age of synthetic biology” and introduces key concepts, approaches, and tools that were considered.
WHAT IS SYNTHETIC BIOLOGY?
Biotechnology is a broad term encompassing the application of biological components or processes to advance human purposes. Although the term itself is thought to have been in use for only about a century, humans have used various forms of biotechnology for millennia. Synthetic biology refers to a set of concepts, approaches, and tools within biotechnology that enable the modification or creation of biological organisms. While there remains no universally agreed-upon definition of synthetic biology (with some defining it more narrowly and others more broadly; see, e.g., Benner and Sismour, 2005; Endy, 2005; Dhar and Weiss, 2007), one distillation is that synthetic biology “aims to improve the process of genetic engineering” (Voigt, 2012). By way of backdrop for this statement, it is useful to note that some of the concepts and approaches now associated with synthetic biology have roots going back to the early days of genetic engineering in the 1970s and the improvements and achievements that were envisaged then. In 1974, for example, the molecular biologist Walter Szybalski set the stage for some key synthetic biology concepts and presaged activities that have now been demonstrated.1 An inflection point for the field occurred around the year 2000, after which synthetic biology gained significant attention and momentum. Two publications often identified with the field’s acceleration are by Elowitz and Leibler (2000) and Gardner et al. (2000). Although genetic engineering was occurring—and improving—prior to 2000, and the principles espoused by synthetic biologists were already noted and in use to varying extents (see, e.g., Toman et al., 1985; and Ptashne, 1986), that year marked a shift toward the adoption of approaches more typical of engineering disciplines, but which had previously been given only modest attention in the biological sciences.
___________________
1 “Up to now we are working on the descriptive phase of molecular biology. . . . But the real challenge will start when we enter the synthetic biology phase of research in our field. We will then devise new control elements and add these new modules to the existing genomes or build up wholly new genomes. This would be a field with the unlimited expansion potential and hardly any limitations to building ‘new better control circuits’ and . . . finally other ‘synthetic’ organisms, like a ‘new better mouse’. . . . I am not concerned that we will run out [of] exciting and novel ideas . . . in the synthetic biology, in general” (Szybalski, 1974).
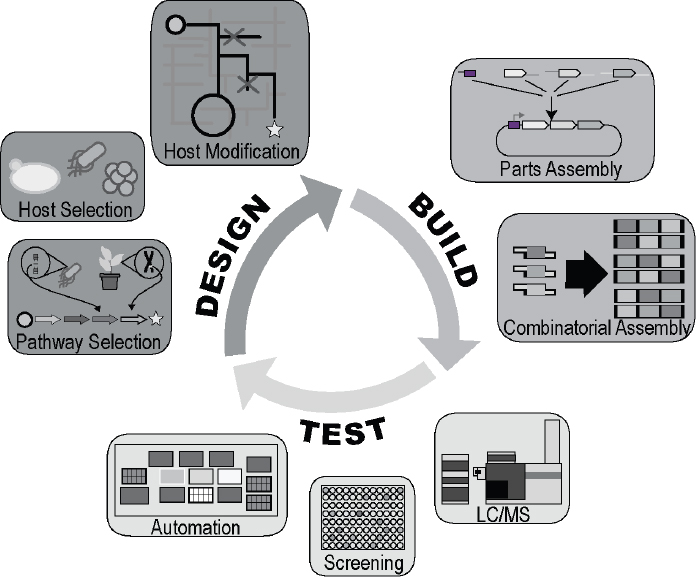
NOTE: LC/MS = liquid chromatography–mass spectrometry.
SOURCE: Modified from Petzold et al., 2015.
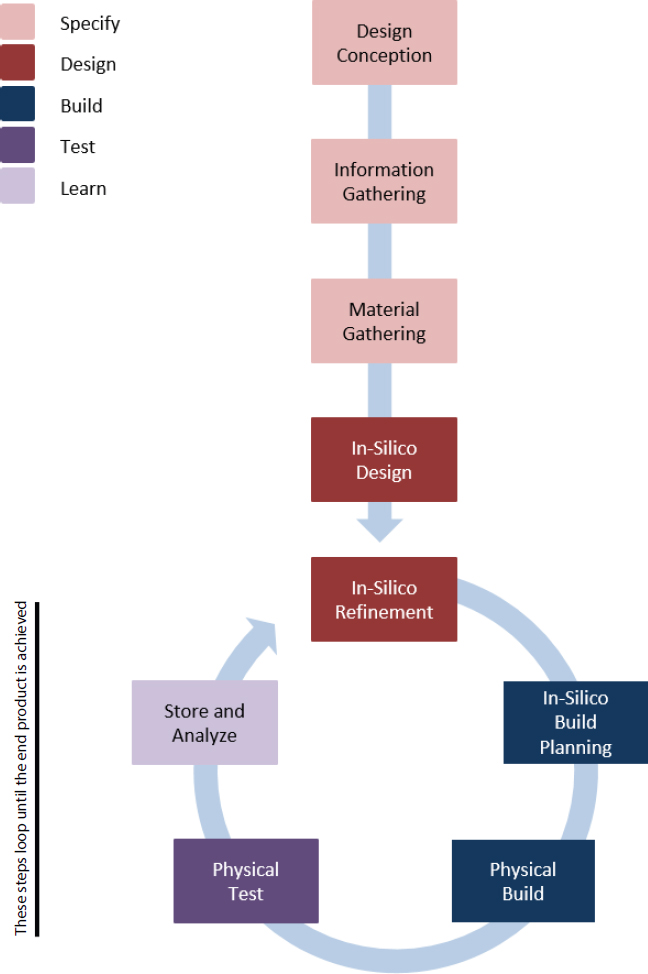
In improving the process of genetic engineering, synthetic biology places special emphasis on the Design-Build-Test (DBT) cycle2 (see Figures 2-1 and 2-2), the iterative process of designing a prototype, building a physical instantiation, testing the functionality of the design, learning from its flaws, and feeding that information back into the creation of a new, improved design. Developments such as enhanced computing power, laboratory automation, cost-effective DNA synthesis and sequencing technologies, and other powerful techniques to manipulate DNA have made it possible for biological engineers to rapidly repeat the DBT cycle to refine designs and products for a desired purpose. Key developments exemplifying these approaches include the establishment of standardized genetic parts registries, intensive use of models and other quantitative tools to simulate biological designs before building them, the availability of open-source DNA assembly methods, and the ability to create rationally designed genetic “circuits”—systems of DNA-encoded biological components designed to perform specific functions (Elowitz and Leibler, 2000; Gardner et al., 2000; Knight, 2003; iGEM, 2017a).
The age of synthetic biology is marked by the broad adoption and consolidation of these concepts, approaches, and tools within the DBT cycle to accelerate the engineering of living organisms. The concepts, approaches, and tools developed to advance synthetic biology will continue to be integrated more broadly into the life sciences toolkit and applied toward many biological research and biotechnology activities. As a result, this report does not draw a precise distinction between synthetic biology and other aspects of advancing biological sciences, but considers synthetic biology a crucial contributor to the spectrum of activities within biology and biotechnology more broadly.
___________________
2 Sometimes referred to as a Specify-Design-Build-Test-Learn cycle or other variations.
The age of synthetic biology is ushering in not only novel technologies, but the application of engineering paradigms to biological contexts. The general intent to manipulate biological systems and to apply engineering paradigms from other disciplines is not new; from the introduction of recombinant DNA technologies in the 1970s to the present, there has been a concerted effort to manipulate genetic material and biological organisms. What has changed is the increased power of particular technologies that enable engineering paradigms to be applied to biological materials. Assessing new technologies and platforms that may enable the creative or destructive manipulation of biological materials, systems, and organisms will be important for identifying potential security opportunities and vulnerabilities.
IMPLICATIONS OF THE AGE OF SYNTHETIC BIOLOGY
Synthetic biology is enabled by tools and techniques from a variety of scientific disciplines, from electrical engineering to computation to biology to chemistry. For example, the exponential improvements in DNA sequencing capabilities, initially developed to further our understanding of the human genome but soon applied to characterize many other organisms, have provided crucial raw material for synthetic biology and fueled innovation over the past decade. More recently, genome editing tools such as CRISPR/Cas9 (“clustered regularly interspaced short palindromic repeats”) (Jinek et al., 2012; Cong et al., 2013) have been adopted for synthetic biology techniques such as the regulation of gene circuits and the development of gene drives (genetic elements for which inheritance is favorably biased; see National Academies of Sciences, Engineering, and Medicine, 2016). Scientific progress in domains relevant to synthetic biology has been remarkably rapid; CRISPR/Cas9, for example, was extended from mammalian cell culture (in the United States) to primates (in China) in a single year (Cong et al., 2013; Jinek et al., 2013; Mali et al., 2013; Niu et al., 2014).
Two somewhat dichotomous phenomena are increasing the pace and progress of engineering of biological systems. The first is that bioengineering can be more theoretical, due to increased predictability of biological systems and evolving standards for biological performance. Biological engineering approaches make it possible to separate the design of a biological material or organism from its manufacture, and standards are evolving to facilitate a theoretical approach to biological design. Biological knowledge may thus be captured and applied in the design stage. The second phenomenon is the ability to try many different designs, often in parallel, and to potentially use directed evolution (see Appendix A) in living systems to perfect the design (see Box 2-1). The inexpensive technologies involved in designing and creating new DNA constructs to test make it easier to proceed without a hypothesis of how the design will work; in other words, it is “cheaper to make than to think.”3 However, the level of underlying biological knowledge still affects the degree to which these biological engineering techniques can be successfully applied; for example, adjusting well-understood pathways to increase ethanol production is fundamentally easier than increasing the virulence of Francisella tularensis, whose virulence mechanisms remain largely unknown.
These advances have real-world consequences for the development of new biotechnologies as well as their accessibility to actors of all types. On the positive side, it is expected that these technologies will enable a wider range of therapeutics, a wider range of biological detection and diagnostic methods, and opportunities to detect biological anomalies. However, these developments also potentially increase the power of even less-resourced malicious actors to produce a harmful biological agent. In this context, it is useful to consider the technologies that enable synthetic biology and how these developments may drive paradigm shifts in the practice of bioengineering.
Enabling Technologies for Synthetic Biology
Synthetic biology is enabled by numerous technologies that enhance success rates and facilitate experimentation, particularly in the DBT cycle. The development of these technologies to some extent defines the transition to the current age of synthetic biology. These include technologies specifically created for synthetic biology, as well as technologies developed for general molecular biology and biotechnology that are being exploited by synthetic biologists. These enabling technologies serve as the tools that facilitate the specification of biological designs and constructions. Key enabling technology areas, examples of which are described in more detail in Appendix A and below (see Specific Synthetic Biology Technologies and Applications), include the following:
- DNA synthesis and assembly. The heart of synthetic biology is the ability to make DNA constructs quickly and efficiently. Improvements in synthesis technology have followed a “Moore’s Law–like” curve for both
___________________
3 For example, researchers recently synthesized and tested more than 7,000 genes to identify diverse homologs capable of complementing the deletion of two essential Escherichia coli genes. While the function of those 7,000 genes could be inferred by sequence similarity, it was more tractable to prove their function via synthesis and testing rather than developing a model of their function from first principles. In practice, these large-scale efforts are synergistic with modeling techniques because they provide systematic data that can strengthen models for predicting biological functions (Plesa et al., 2018).
- Genome engineering. Although in the past it has proven possible to engineer organismal or viral genomes via painstaking mutational methods, the ability to synthesize DNA quickly, coupled with improvements in transformation technologies and “booting” (the steps needed to go from DNA to a viable organism), has led to an acceleration in the ability to make mutations, including multiple mutations in parallel (e.g., Wang et al., 2009). In particular, the ongoing CRISPR revolution (Doudna and Charpentier, 2014) has led to the ability to introduce site-specific changes into a wide variety of organisms that may have previously been refractory to such techniques.
- Improved computational modeling. With new approaches to modeling biological systems and improved computing power, more complex biomolecular designs and system behaviors can be explored. This allows for larger areas of the theoretical “design space” in biology to be explored and tested in parallel, leading to better working systems in less time. Modeling advances are abetted by new computational advances
reductions in costs and increases in the length of constructs that are attainable. These trends are likely to continue.
-
in machine learning and big data that have allowed the results of past experiments (both successes and failures) to inform the next round of design and experimentation. In the future, the creation of “rules” from the machine learning process should greatly improve the specification of future successful designs.
- Genetic logic. A key development in the field that meshes with improvements in modeling is the development of genetic logic circuits (Moon et al., 2012; Kotula et al., 2014) that allow living systems to make basic “decisions” based on both current inputs to the system (combinational logic) and the history of inputs (memory or sequential logic). The inherently programmable nature of genetic logic circuits is expected to mesh with advanced modeling approaches to improve the DBT cycle. An example of the use of genetic logic is plants that have been modified to act as radiation sensors capable of indicating when large amounts of gamma radiation have been detected (Peng et al., 2014).
- Directed evolution. While directed evolution methods are not new, their application has been accelerated by recent advances in DNA synthesis and genome engineering and are thus addressed in this report under the umbrella of biotechnology in the age of synthetic biology. Directed evolution methods stand both as an alternative to design-based models and as a supplement to them, in that they can return enormous amounts of data on fitness landscapes that can further improve computational modeling approaches. Additionally, the combination of design and selection moves constructs well beyond the bounds of what nature would attempt while still allowing the facile repair of unintended unnatural or less-fit deficiencies and interactions. A somewhat notorious example of the use of directed evolution was the introduction of an engineered version of a more virulent strain of influenza virus into ferrets, where it rapidly evolved to become airborne-transmissible (Fouchier, 2015). While this research was done for reasons some argue were appropriate, it also provided a blueprint for potential misuse.
Engineering Paradigms for Synthetic Biology
Enabling technologies have allowed synthetic biologists to make genetic changes in organisms with greater ease, precision, and scale. As a maturing engineering discipline, synthetic biology is also being advanced by engineering paradigms that allow these tools to be used with greater predictability of result. Engineering paradigms are methods of adapting enabling technologies to abstraction, standards, computing, workflow optimization, and other engineering principles. If enabling technologies provide options for what tools will be used in synthetic biology, engineering paradigms describe how these technologies will be used. In other words, these paradigms encompass the processes and decisions followed in designing, building, and testing biological constructs. The following engineering concepts and paradigms are particularly relevant to the context of this study:
- Specify-Design-Build-Test-Learn cycle. The Specify-Design-Build-Test-Learn cycle refers to an iterative process that requires a formal description of the desired biological behavior or function (Specify), the planned modification of an organism in silico or via rational design principles to realize that behavior (Design), the physical assembly of the biological material representing those designs (Build), the testing of the material to determine if it functions as specified (Test), and formally capturing and storing information about the entire process to inform the next revision or subsequent design (Learn). The boundaries between the cycle stages are fluid, and for the purposes of this report, the cycle is simplified to Design-Build-Test, with other stages implicitly included in these core elements. For example, Specify is incorporated into Design, and Learn is incorporated in the analytical steps of Test. Additional elements that are pertinent to biodefense considerations, such as Scale and Delivery, are also included.
- Combinatorial approaches. Although not an engineering paradigm per se, it is a fundamental shift that in many cases, it is now often “cheaper to make than to think.” It is becoming increasingly common to use combinatorial approaches—approaches in which a large number of genetic variants are created and then tested. Variants can be created by using a technique in which a large number of DNA variants are incorporated systematically to synthesize multiple variants (i.e., combinatorial assembly). The concept is that one can generate a large number of variants with limited knowledge of sequence-function relationships.
-
These approaches enable many design options to be explored, even in the absence of predictive tools to model the performance of those designs. Directed evolution is a related concept, discussed in Appendix A.
- High-throughput data acquisition. The speed of the DBT cycle has been greatly increased by the raft of enabling technologies such as combinatorial assembly (Smanski et al., 2014; Carbonell et al., 2016), CRISPR/Cas-based editing methods (Black et al., 2017; Schmidt and Platt, 2017; Mendoza and Trinh, 2018), and directed evolution (Cobb et al., 2013; Tizei et al., 2016). By synergizing with advances in analytical chemistry and biology, such as microfluidics and high-throughput sequencing, these technologies may allow the functional assessment of millions of constructs in parallel, hence providing particularly robust feedback for the next iteration of design.
- Separation of design and manufacturing. Specifying and designing a system can now be done in one location (e.g., an academic environment) while the manufacturing process (the Build step in the DBT cycle) is done in another location (e.g., a remotely operated facility or “cloud laboratory”). The increasing physical and virtual separation of design and manufacturing not only further increases the accessibility of synthetic biology but also creates potential security concerns where designs cannot necessarily be explicitly connected to manufacturing locations and vice versa.
- Standards. Standards have emerged that make DNA assembly easier and parts more “sharable” (e.g., Gibson and modular cloning assembly methods). Data standards such as Synthetic Biology Open Language4 have made the sharing, analysis, and software ecosystem of synthetic biology increasingly sophisticated. Such standards may ultimately allow engineers to focus on raising the level of abstraction in designs since lower-level mechanisms have been well defined and vetted.
SPECIFIC SYNTHETIC BIOLOGY TECHNOLOGIES AND APPLICATIONS
The technologies and engineering paradigms described above have led to a number of applications that drive synthetic biology development because they provide unique ways to take advantage of what synthetic biology offers. They are not all unique to synthetic biology, nor are they all routinely used to explore synthetic biology designs. For example, all synthetic biologists use software to store and analyze DNA sequences and use some form of computation in specifying designs (e.g., using biophysical models or algorithms to design ribosome binding sites, to check folding energies of DNA primers used for amplification and assembly, or to refactor the DNA sequence encoding a protein to increase protein production, a technique known as “codon optimization”). However, far fewer have the requisite library of DNA parts and accompanying software tools to achieve a level of abstraction that would allow the researcher to query, for example, a logic gate that accepts glucose concentration as input and activates transcription of a tethered reporter when a specific concentration is achieved. In other words, there are approaches and tools that are continuing to develop and gain traction within synthetic biology but which have not necessarily reached their full technical potential or user adoption.
Although the technologies used in each of the component phases of the DBT cycle may evolve over time or be replaced by new technologies, the fundamental concepts of the DBT cycle will stand. Thus, it is useful to consider current technologies and anticipated future developments in terms of the ways in which they enable the DBT cycle. However, it is important to recognize that the component phases of the DBT cycle are not strictly separate. It is possible, even probable, that some technologies or approaches will have impacts across multiple phases of the DBT cycle; one such example may be directed evolution, where repeated passage in a model host or in cell cultures under stress permits nature to Design, Build, and Test new phenotypes. There are also likely areas in which advances in synthetic biology capabilities relevant to biodefense would arise from synergies or convergence among technologies relevant to different phases. For example, it is important to consider potential synergies between Design technologies and Build technologies, because a malicious actor would need both Design and Build capabilities to carry out an attack. Similarly, synergies may arise if large-scale Test technologies are developed to match the enormous output of certain Build technologies, thus helping those Build technologies reach their full potential.
___________________
4 See http://sbolstandard.org. Accessed November 9, 2017.
TABLE 2-1 Synthetic Biology Concepts, Approaches, and Tools That Enable the DBT Cycle
| Key Synthetic Biology Concepts, Approaches, and Tools | Design | Build | Test |
|---|---|---|---|
| Automated biological design | |||
| Metabolic engineering | |||
| Phenotype engineering | |||
| Horizontal transfer and transmissibility | |||
| Xenobiology | |||
| Human modulation | |||
| DNA construction | |||
| Editing of genes or genomes | |||
| Library construction | |||
| Booting of engineered constructs | |||
| High-throughput screening | |||
| Directed evolution |
NOTE: Shading indicates which phase of the DBT cycle each example aligns with most closely. See Appendix A for full descriptions.
Appendix A describes a core set of current synthetic biology concepts, approaches, and tools that enable each step of the DBT cycle, focusing particularly on areas in which advances in biotechnology may raise the potential for malicious acts that were less feasible before the age of synthetic biology. Although the examples presented are intentionally quite broad and somewhat arbitrary—and do not represent an exhaustive list of all technologies or all possible applications of synthetic biology—they provide useful context for understanding how specific tools or approaches might enable the potential capabilities analyzed in Chapters 4–6 and can be adapted to assess new areas of concern as the biotechnology landscape continues to evolve. In addition, although Appendix A captures the main known technologies at the time of writing, this list will need to be updated and modified to stay relevant as the science advances.
Table 2-1 summarizes how the concepts, approaches, and tools described in Appendix A map to the phases of the DBT cycle. Going forward, it will be useful to consider how each phase of the DBT cycle may be further enabled by future developments in technology and knowledge, particularly in areas where a current bottleneck may be overcome. Appendix A also indicates the relative degree of maturity of specific techniques discussed (see Figure A-1).








