Proceedings of a Workshop
| IN BRIEF | |
|
May 2018 |
The Promise of Genome Editing Tools to Advance Environmental Health Research
Proceedings of a Workshop—in Brief
Advances in genome editing—the process for making precise additions, deletions, and alterations of DNA and RNA—have opened the door for studying biological mechanisms of health and disease. In particular, the 2012 development of the CRISPR/Cas9 system1 as a genome editing tool has enabled genome editing to become more rapid, efficient, and flexible than ever before. In 2014, MIT Technology Review magazine called CRISPR, “the biggest biotech discovery of the century.” The potential to use genome editing to cure human disease or stop the spread of invasive species has led to substantial investment in biomedical and public health applications. Genome editing tools such as CRISPR/ Cas9, TALENs,2 and zinc finger nucleases are emerging as a means to use more human-relevant in vitro assays (based on both human and animal cells) to screen the toxicity of environmental contaminants and better identify the health implications of environmental exposures. Even more recently, researchers are exploring the use of CRISPR/Cas9 and piRNAs for epigenome editing. However, genome editing tools have yet to be widely used in environmental health and toxicology research.
On January 10–11, 2018, the National Academies of Sciences, Engineering, and Medicine’s (the National Academies’) Standing Committee on Emerging Science for Environmental Health Decisions held a 2-day workshop to explore what role genome and epigenome editing tools could play in advancing environmental health research and decision-making. The workshop, sponsored by the National Institute of Environmental Health Sciences (NIEHS), brought together experts in molecular biology, toxicology, public health, and risk assessment to explore the topic. Lesa Aylward of Summit Toxicology, one of the workshop’s organizers, briefly outlined three workshop goals:
- broaden scientists’ familiarity with how these tools can move environmental health science research forward,
- aid in the understanding of environmental health issues, and
- inform regulatory decision-making.
Aylward pointed to the recent publication of a suite of National Academies’ publications on genome editing, including Human Genome Editing: Science, Ethics, and Governance (2017); Gene Drives on the Horizon: Advancing Science, Navigating Uncertainty, and Aligning Research with Public Values (2016); Genetically Engineered Crops: Experiences and Prospects (2016); Gain-of-Function Research: Summary of the Second Symposium (2016); and Preparing for Future Products of Biotechnology (2017). These publications, especially the first two, include extensive considerations of the ethical issues linked to the use of these new tools, therefore, this workshop’s organizers elected not to devote specific time to those issues, Aylward said. The motivation for this workshop was to ensure coverage of new ground by focusing on the applications of genome and epigenome editing for environmental health research, she emphasized.
________________
1 The pairing of segments of bacterial DNA known as Clustered regularly-interspaced short palindromic repeats with the RNA-guided DNA endonuclease enzyme, CRISPR associated protein 9.
2 TALENs stands for transcription activator-like effector nucleases.
![]()
GENOME EDITING 101
Discoveries in the early 1990s set the stage for the development of gene editing tools, explained Fyodor Urnov of the Altius Institute for Biomedical Sciences and the University of California, Berkeley. The first breakthrough was the discovery of zinc finger DNA-binding proteins by Aaron Klug and Carl Pabo in 1991. Zinc fingers are unusual among DNA binding proteins because they can be linked linearly to recognize nucleic acid sequences of varying lengths, explained Urnov. Then, in 1994, Marie Jasnin set the stage for gene editing with her recognition that double-stranded DNA breaks in mammalian cells tend to cause the genes to recombine. Dana Carroll built on those discoveries in work with fruit flies and Xenopus frogs from 2001–2003 to show how the zinc finger nuclease can be engineered to induce a double-stranded DNA break. This led to the discovery of other nucleases capable of inducing double-stranded breaks, including TAL (transcription activator-like) effectors in 2009 by Ulla Bonas, Urnov explained. Finally, in a paper published in 2012, Jennifer Doudna and Emmanuel Charpentier reported the RNA-guided nature of the Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and the CRISPR-associated genes (Cas, including the Cas9 nuclease).3
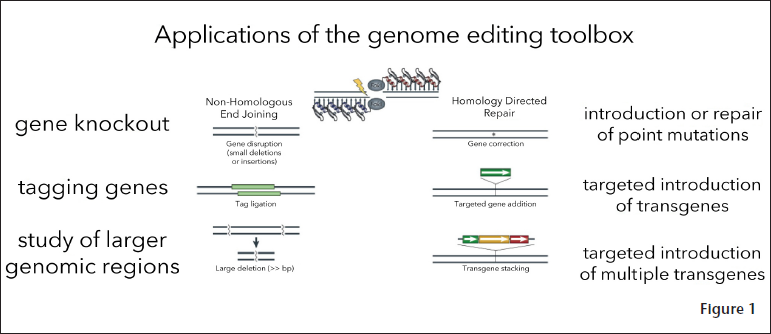
CRISPR and Cas are essential for adaptive immunity in selected bacteria, such as E. coli. They enable bacteria to capture snippets of DNA from invading viruses so that subsequent generations of bacteria can repel these pathogens more effectively. When a virus infects a bacterium, the bacterium integrates fragments of DNA from the invading phage into its own genome.4 The snippets of phage DNA are a CRISPR locus—a “molecular barcode” that helps the bacterium identify future viral invaders, explained Urnov. If a virus returns, Cas9 “grabs” the molecular barcode to determine whether it matches the DNA of the invading virus “as if it were somebody on the FBI’s most wanted list,” said Urnov. Doudna and Charpentier were among the first to recognize that they could harness the capabilities to achieve site-specific DNA recognition and cleavage, said Urnov. However, Urnov stressed that the tools for genome editing that are best for a given research project are entirely dependent on the project’s goals. He said that asking whether one genome editing tool is better than another is like “asking whether the iPhone is better than the Google Pixel—they’re both wonderful smartphones that do amazing things, it’s a matter of nuance.” Urnov also underscored that the “tool box” of genome editing applications is universal, even though the approach to targeting DNA varies among the different types of genome editing tools (see Figure 1). For environmental health scientists, one useful value of genome editing is its potential to allow researchers to evaluate cases where they suspect that a given compound has an effect on a cell that is mediated by a specific gene, Urnov said. Genome editing tools allow researchers to remove that gene and see what happens.
Zinc finger proteins, TAL effectors, CRISPR/Cas9, and other tools can also be epigenome editors. In the late 1990s, scientists learned that engineered DNA-binding domains can be used to target specific transcriptional outcomes, either by activating or repressing endogenous genes.5 Urnov described epigenome editing as a
________________
3 Jinek, M., K. Chylinski, I. Fonfara, M. Hauer, J. A. Doudna, and E. Charpentier. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821. doi: 10.1126/science.1225829.
4 Jiang, F., and J. A. Doudna. 2015. The structural biology of CRISPR-Cas systems. Current Opinion in Structural Biology 30:100–111. doi: 10.1016/j.sbi.2015.02.002.
5 Thakore, P. I., J. B. Black, I. B. Hilton, and C. A. Gersbach. 2016. Editing the epigenome: Technologies for programmable transcriptional modulation and epigenetic regulation. Nature Methods 13(2):127–137. doi: 10.1038/nmeth.3733.
“complimentary technology” to genome editing. Using targeted double-strand breaks to change the transcriptional state of a gene can “create really interesting cell phenotypes,” said Urnov.
Vikram Pattanayak of the Massachusetts General Hospital discussed some of the latest research on the precision of genome editing. His research helped to establish how well CRISPR/Cas9 is able to target a specific part of the human genome without causing off-target effects. The human genome includes approximately 3 billion base pairs. “What is the chance of finding any given 22 base pair target site?” Pattanayak asked. Because the genome is a random collection of adenine, cytosine, guanine, and thymine nucleic acid base pairs, there is a one in four chance of putting each nucleic acid at each of 22 positions. This sequence size would be statistically unique in a genome of 422 (approximately 1013) base pairs, a genome far larger than the human genome, he said.
Zinc fingers and CRISPR/Cas9 can be engineered to clinical grade nuclease specificity. However, a critical question for using genome editing tools in the clinic, as well as for research, is how likely can the tools alter segments of DNA that are not the intended target for editing, i.e., off-target effects. Initial studies in human cells demonstrated that CRISPR/Cas modified genes without significant toxicity and some single base pair mismatch tolerance. Research findings on off-target effects indicate that Cas9 is specific enough to edit desired sites in the human genome and suggest approaches for minimizing off-target effects. Later work began showing more off-target effects using a technique called GUIDE-seq. The number of detectable off-targets ranges from zero to hundreds depending on the target site, Pattanayak said. He added that the detection of off-target effects in engineered Cas9 variants can be challenging and is an active area of ongoing research. However, Pattanayak noted that existing off-target prediction programs can serve as good tools to evaluate different potential guide RNAs (gRNAs) and their potential off-target effects.
Pattanayak emphasized that the level of concern about off-target effects depends on the purpose of the research. If CRISPR/Cas9 is part of a clinical therapy in humans, precision is likely a much more prominent factor than it is for laboratory research with no clinical applications, he noted. For most research applications, Pattanayak said that the specificity possible with CRISPR/Cas9 is good enough. Researchers can use multiple gRNAs targeted to the same gene to try to minimize off-target effects. If the sequences of the gRNAs are sufficiently different, the off-target profiles will be different. When research conducted with different guides produces the same results, researchers can be pretty confident about them, he said.
Another issue to keep in mind, according to David Taylor of The University of Texas at Austin, who conducted postdoctoral research in Doudna’s lab, is what we know and do not know about how the Cas9 enzyme works to unwind DNA and induce double-stranded breaks. Understanding “the basic mechanism of Cas9 is as important as using it for genome engineering applications,” he argued. Scientists still are not quite sure of all of the specifics about how the enzyme itself works. There are different types of Cas9 from different bacteria, such as SpyCas9 from Streptococcus pyogenes. These enzymes can search through billions of base pairs to find the protospacer adjacent motif that causes the Cas9 to unwind into an R-loop structure, significantly bending the DNA in the process. His lab has been working out the kinetics of how that takes place. Taylor pointed out that researchers now know that more Cas9-like enzymes exist and he thinks that other genome engineering guides may also be discovered.
GENOME EDITING IN ENVIRONMENTAL HEALTH
Is genome editing just a clever way to conduct environmental health research? Or do the new tools offer something more? Workshop speakers presented evidence that the new tools can aid environmental health researchers in moving toward the approaches for investigating toxicity outlined in the National Research Council’s 2007 report Toxicity Testing in the 21st Century: A Vision and a Strategy.
Richard Woychik, Deputy Director of the NIEHS, pointed out that the genomics era has brought a more mechanistic focus to environmental health research as scientists identify the genes and pathways implicated in an individual’s response to an environmental exposure. Genome editing improves researchers’ abilities to investigate mechanisms of toxicity and understand the genes and the pathways that are implicated in how an organism responds to environmental perturbations. Researchers are already taking advantage of inherent, naturally occurring genetic variability within extant strains of mice, rats, and other organisms using approaches that eventually could be applied to look at genetic variability in humans. The new tools hold promise for enabling researchers to pinpoint much more quickly which genes are actually responsible for toxicity by assessing the role of numerous genes that have been shown to be involved in responses to toxic exposures, he said.
Woychik described several areas of research that could benefit from the use of genome editing tools to understand environmental mechanisms of toxicity. One of those areas involves using the existing genetic variability within new sets of reference populations of mice, called the Collaborative Cross and the Diversity Outbred. The use
of Collaborative Cross and Diversity Outbred mice would involve entirely different population-based strategies on how mice are used for experiments in toxicology. He explained that most toxicities are likely to be influenced by the genetic background of individuals (humans or animals) and that these new population-based strategies will likely be able to more accurately identify toxicities that appear in a genetically diverse population of humans. He continued by explaining how utilizing Collaborative Cross and Diversity Outbred animals could greatly enhance the ability to identify variations in genes that are responsible for the differential toxicity between individuals. Additionally, he went on to describe how genome editing tools such as CRISPR/Cas9 could be used to identify modifier genes. He explained that it would be enormously valuable to be able to understand how the sequence variations within genes across the genome influence a toxicity phenotype generated by the knock-out of a gene. He described how an “in vitro genetics” approach utilizing panels of embryonic stem cells derived from different genetic backgrounds could be used for this purpose. Woychik and Keith Houck of the U.S. Environmental Protection Agency (EPA) agreed that this can greatly speed up the efforts to identify such modifier genes. Identifying the location of a gene of interest from any species—including humans, mice, and others—within a genomic interval is relatively simple, but the intervals often have numerous single-nucleotide polymorphisms (SNPs) with effector genes that contribute to the phenotype. Several workshop attendees remarked that CRISPR/Cas9 could be a powerful tool to identify the causal variants.
Luoping Zhang of the University of California, Berkeley’s Superfund Research Program is at the vanguard of efforts to apply genome editing tools to environmental health. Zhang emphasized that there is considerable opportunity to use CRISPR in environmental health research, particularly for toxicity testing. However, when she examined the literature from the past 10 years, Zhang found fewer than 50 environmental health publications that involve CRISPR. Two of the more notable publications included an assessment of the impact of the antimicrobial triclosan exposure on human liver cells6 and the identification of pathways associated with cell death induced by exposure to the herbicide paraquat.7 Zhang has been using CRISPR for genome-wide editing as part of a project to identify genes in human cells associated with increased susceptibility to chemical exposures. Her team employs Cas9 variants, including CRISPR interference and CRISPR activation. These allow the team to decrease or increase gene expression, rather than completely knock-out gene expression, and assess the effects of more subtle modulations in gene expression on important cellular attributes including growth, death, gene expression, cell membranes, and mitochondrial function.
In collaboration with Quan Lu of Harvard University’s T.H. Chan School of Public Health and Chris Vulpe of the University of Florida, Zhang is working on a project called ToxCRISPR. The project includes 3,675 genes responsive to environmental exposures, based on ones identified by the Tox21 program (which is run by the NIEHS, the National Toxicology Program, EPA, and the U.S. Food and Drug Administration [FDA]), as well as genes identified through the Environmental Genome Project and other efforts. Combined, these genes make up a targeted CRISPR library for probing toxicology mechanisms. The use of a targeted library reduces the number of cells needed to carry out screening studies (7.5 million cells for 500× coverage) in comparison to genome-wide libraries (30 million cells). The ToxCRISPR library contains multiple sgRNA for each gene, as is standard in screening libraries, in order to compensate for potential off-target effects and increase statistical confidence in research results.
Zhang says that the approach works both for screening as well as targeted validation. The researchers are collaborating with biostatisticians and bioinformaticists to identify the gene linked to responses to environmental exposures shown through their testing. They are also attempting to analyze the pathways involved in the response to try to discover the underlying mechanism of action. They are using in vitro testing and in vivo testing to validate their findings. The approach holds promise for enabling more rapid screening of chemicals with more doses, Vulpe and Zhang agreed. “Although any one screen isn’t going to tell you everything about a process, we think that it helps you fill some holes in our knowledge to work out adverse outcome pathways,” Vulpe said.
Genome editing can be used to confirm mechanisms of environment–pathway–disease relationships found in epidemiologic studies said Rebecca Fry of the University of North Carolina at Chapel Hill. For risk assessors, the tools can be used to identify hazards and develop and optimize adverse outcome pathways (AOPs), Zhang said. AOPs encompass events that initiate detectable changes on the molecular level, as well as later key events that connect the molecular-level initiating events to effects at higher levels of biological organization, and eventually result in adverse (e.g., disease) outcomes for risk assessment. In all of these areas of environmental health research, technology enables researchers to ask bigger questions and answer them more thoroughly, said Gary Miller of Emory University.
Adding to the appeal of genome editing tools is their promise for use in untargeted toxicology and environmental health studies that can shed light on previously unsuspected linkages between environmental exposures
________________
6 Xia, P., X. Zhang, Y. Xie, M. Guan, D. L. Villeneuve, and H. Yu. 2016. Functional toxicogenomics assessment of triclosan in human HepG2 Cells using genome-wide CRISPR-Cas9 screening. Environmental Science & Technology 50(19):10682–10692.
7 Reczek, C. R., K. Birsoy, H. Kong, I. Martínez-Reyes, T. Wang, P. Gao, D. M. Sabatini, and N. S. Chandel. 2017. A CRISPR screen identifies a pathway required for paraquat-induced cell death. Nature Chemical Biology 13(12):1274–1279. doi: 10.1038/nchembio.2499.
and health outcomes, said Shruthi Mahalingaiah of Boston University’s School of Medicine. Scientists affiliated with the NIEHS are investigating how to use the tools to address the agency’s research priorities. The fact that genome editing tools are available “off the shelf” and can be purchased at a reasonable cost adds significantly to their appeal, observed Norbert Kaminski of Michigan State University.
EMERGING USES OF GENOME EDITING TOOLS
The projects under way that provide insights into how genome editing tools can aid environmental health research include work by David H. Sherr at Boston University. He is using funds from the NIEHS Superfund Research Program to track the activation of the aryl hydrocarbon receptor (AHR). The AHR is known to be activated by environmental chemicals, including polychlorinated biphenyls, polyaromatic hydrocarbons, and dioxin-like compounds, and it can be overexpressed and hyperactive in cancer. Because AHR activation is linked to cells’ tendencies to be invasive and to migrate, as well as metastasis and the tendency to act like stem cells, improving the understanding of the AHR’s role in cancer holds the potential for treatment. “What you want to do is take the AHR away, find out what kind of molecular and biologic outcomes happen downstream, and then design drugs to attack the AHR,” he explained.
Sherr’s group previously used techniques to “knock-down” the AHR response to chemical exposures. With CRISPR/Cas9, the team can completely “knock-out” the AHR gene. Doing this has allowed the team to definitively show that removing it can halt the response initiated when cells are exposed to the chemicals known to trigger the receptor. This also enabled them to pinpoint other gene sequences implicated in the response, something that many researchers had previously tried to do using other techniques without complete success. Looking further, Sherr’s team was able to identify leukocyte white blood cells associated with the AHR’s immune response that they targeted. They designed a drug that mimics the action of what they have discovered.
CRISPR “has the ability to revolutionize toxicology by enabling rapid functional assessment of the role of a targeted gene in toxicity,” said Vulpe, who is using genome-wide CRISPR (see Figure 2) to study two prototypical toxicants, acetaldehyde and arsenic. Both are carcinogens that have been exhaustively studied for a long time, but their mechanisms of toxicity remain poorly understood because there are so many different mechanisms that researchers have not been able to determine which ones are important. His group is using genome-wide screening with CRISPR
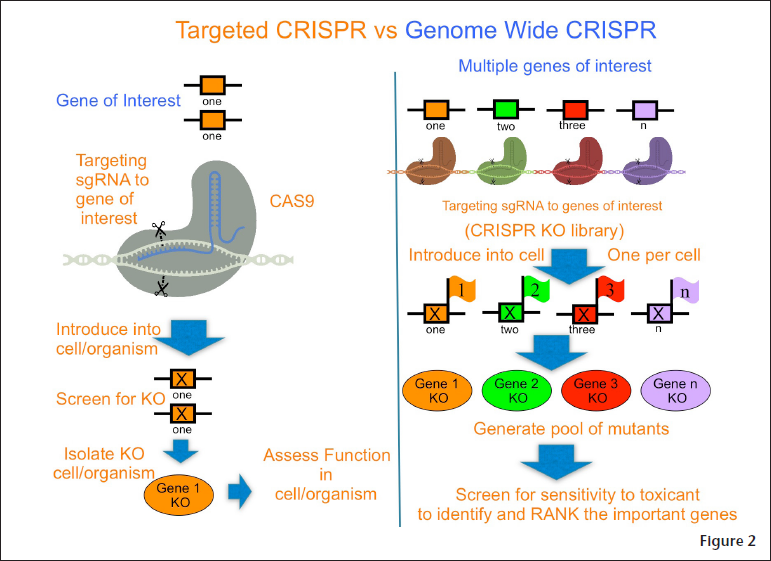
as a way to rank these mechanisms and gain insight into what is happening and the relative importance of the mechanisms involved. The approach involves systematically silencing a mammalian cell’s genes and exposing the modified cells to a toxicant to identify which genes are important in the response to the toxicant. This allows the team to assess the function of all genes simultaneously, he said. It helps fill in “some of the holes in our knowledge” and work out more of the details in our understanding of AOPs, he added.
The approach involves using a pool of individual mutant cells, each containing a knock-out of a single gene, and altogether representing knock-outs in all targeted genes. Each cell is tagged with a unique single gRNA, which is like a “barcode,” enabling the cell to be tracked. The system screens for sensitivity or resistance of the modified cells containing gene knock-outs to a toxicant. Then, the important genes associated with the response to the exposure are identified and ranked. The test generally measures whether the modified cells’ growth improves or declines in response to toxic exposures.
Vulpe’s team used the genome-wide CRISPR screening approach with the human GeCKO (Genome-scale CRISPR Knock-Out) lentiviral pooled libraries to study arsenic trioxide. Arsenic is the number one substance on the 2017 Agency for Toxic Substances and Disease Registry priority list. The screen identified 100 candidate genes linked to sensitivity or resistance to arsenic trioxide exposure and the team verified the findings with a confirmatory CRISPR screen. The results confirmed previous research showing that arsenic trioxide exposure increases cellular content of reactive oxygen species. Vulpe also found that the loss of any gene associated with incorporating selenocysteine into proteins led to resistance to arsenic trioxide. The resistance effect was unexpected, said Vulpe. Although it is known that arsenic binds selenium, the relationship between arsenic and selenium in cells was unclear.
Lu has used a similar genome-wide CRISPR screening approach to establish a cell-based model for endoplasmic reticulum (ER) stress response induced by inorganic arsenic, which is ubiquitous in the environment. Approximately 150 million people throughout the globe are exposed to high levels of arsenic—including 2 million people in North Carolina with private wells containing elevated levels of up to 800 parts per billion (ppb); EPA’s maximum contaminant level for arsenic in public water supplies is 10 ppb. Chronic exposure to arsenic is associated with skin lesions, cancers, neuropathy, cardiovascular disease, and type 2 diabetes, as well as birth defects. The CRISPR-based screen identified new gene suppressors of ER stress response associated with crucial developmental and physiological processes in the cell and found SNPs associated with neurodevelopment and arsenic exposure in children. Lu and his colleagues plan to follow up by looking for SNPs in humans exposed to arsenic and they are optimistic that what they are learning will eventually help to protect children from arsenic toxicity.
David Gerhold, who works for the National Institutes of Health’s (NIH’s) National Center for Advancing Translational Sciences’ Genomic Toxicology division, said that his research group has a few projects using CRISPR. Although the team has just begun to use the genome editing tool, it has already changed how they work such that the bottleneck is no longer addressing the region of the genome that the research team wants to target. Instead, the bottleneck is now in introducing DNA effectively and efficiently to the cell type of interest. His team is working with stem cells and he says that coaxing them to grow into colonies from single cells is currently a limiting factor in his work.
Mark Hahn of the Woods Hole Oceanographic Institution discussed the new tools’ promise for shedding light on the developmental origins of health and disease. Chemical exposure during early life stages can have delayed and long-lasting consequences, affecting susceptibility to a variety of diseases later in life that can even be passed along trans-generationally. For obvious reasons, this has been challenging to study.
Hahn’s investigations using genome editing to flesh out the understanding of molecular mechanisms of chemical sensitivity associated with developmental toxicity involve work with the AHR using fish models. “Many fish are exposed to high concentrations of chemicals in the environment, so they can serve as good models for population-level impacts,” he said. Fish are also different than mammals in that they have extra copies of the genes for the AHR. One of Hahn’s projects involves work with a small estuarine fish, the Atlantic killifish Fundulus heteroclitus. Several populations of this fish have evolved tolerance or resistance to aromatic hydrocarbons after long-term exposure; the tolerant populations are 300–8,000-fold less sensitive than sensitive populations.
To identify genes that might be involved in resistance, Hahn sequenced the genomes of fish from both tolerant and sensitive populations. His research team looked for parts of the genome under natural selection and identified some strong signals that were shared among tolerant populations, including AHR1a/2a and AHR1b/2b, cytochrome P450 1a (CYP1A), and AHR-interacting protein (AIP). He said a key finding was the fact that proteins other than the AHR were involved in the pathway, and the identification of the AIP was surprising. The results raise the “intriguing possibility that resistance may involve epistatic interactions between proteins,” he said. From there, Hahn and his team used genome editing approaches with zinc fingers and CRISPR/Cas9 to generate AHR2a and AHR2b gene mutations in
killifish embryos. They successfully induced mutations in both and detected no off-target effects.8 During the project, they identified several SNPs, including two present in sensitive killifish as well as mammals, but not the tolerant killifish. The researchers are currently introducing the individual SNPs to test their function in zebrafish.
Anne Kwitek of The University of Iowa noted that the rat was the first mammalian model in which zinc fingers were used to knock-out a gene. TALENs and CRISPR have also been used in rats and a significant number of transgenic and gene-targeted models exist. The Gene Editing Rat Resource Center at the Medical College of Wisconsin provides information about rats with gene knock-outs related to heart, lung, and blood-type characteristics.
Because genome editing can facilitate identifying the genes related to complex diseases, Kwitek is using it to validate some of her previous genetic research into hypertension and the metabolic syndrome using rat models. She and her group have been working with Lyon rats from an outbred population, which includes rats that can serve as a genetic model of metabolic syndrome because they are spontaneously hypertensive, obese, and have abnormally high lipid levels. After mapping these rats’ genomes, the group identified a candidate gene, C17h6orf52, associated with metabolic syndrome. Working with the Gene Editing Rat Resource Center, they generated two mutant lines with deletions in the gene using CRISPR/Cas9. The blood pressure of the knock-out rats (rats without C17h6orf52) increased when the rats were fed high salt diets. In other words, the gene seemed to have a protective role. “When you knock it out, the susceptibility [to hypertension] actually gets worse,” Kwitek explained. However, when they started breeding the animals, they saw that both the wild type and the knock-outs had lower body weights. Kwitek said that she and her colleagues are still trying to determine what is going on, but the findings suggest that the gene does not regulate body weight. In turn, this suggests that the genome editing in the experiment may have had an off-target impact on other genes. Still, Kwitek stressed that the method is sound.
Agouti mice, which can be genetically identical but look very different due to a single epigenetic change that confers obesity, are often used to illustrate why environmental health researchers are interested in epigenetics. To improve understanding of how epigenetic modifications influence environmentally mediated changes in gene expression is one of the NIEHS’s current areas of focus. Dana Dolinoy of the University of Michigan is using Piwiinteracting RNA (or piRNA), small non-coding RNA molecules that act as guardians of the genome, in what she characterizes as an alternative approach to genome editing. piRNA molecules are capable of editing a portion of the epigenome known as the DNA methylome in a unidirectional fashion by inducing DNA methylation at particular sites. Exposing agouti mice to toxicants including lead, bisphenol A, and phthalates can cause them to become obese. Dolinoy is involved in an ongoing project using piRNAs to protect agouti mice exposed to an agent known to shift their epigenome so that they do not become obese. She pointed out that the approach’s appeal includes the fact that it takes advantage of the body’s epigenetic maintenance machinery.
As knowledge moves forward, genome editing may enable researchers to address questions that they never before knew to ask, or were unable to ask because the approach or tools needed to answer them did not (yet) exist. Harris Wang of Columbia University spoke about one new potentially game-changing application, the use of CRISPR/ Cas technologies to record biological signals. Biological signals are difficult to record in complex environments and over long time scales, he explained. Wang and his research team discovered that by modifying the DNA of E. coli, they could transform the bacteria into a living, microscopic “tape recorder.”9 This work couples recent research on the use of DNA as a synthetic information storage medium with the natural CRISPR-based bacterial immune response to acquire and store snippets of DNA from viral invaders. As a result, Wang and his team were able to convert an input signal into DNA abundance and systematically incorporate the captured DNA elements into CRISPR arrays. The researchers were also able to develop and “barcode” different types of E. coli recorders, one to sense copper, one to sense trehalose, and another to senses fucose, and then pull them all together to do simultaneous recording. The new tool can also be used to record a variety of different endogenous, temporal, or transcriptomic signals, said Wang. He suggested that a living biological recorder may enable environmental health scientists to record otherwise invisible changes linked to chemical exposures without disrupting its surroundings.
Miller said the recorder “shows how CRISPR is the gift that keeps on giving,” in that many new and innovative uses of the technology are emerging. “I think it’s a great application of how we can sense environmental perturbations and use genome editing approaches to record that,” he said. Hahn expressed optimism that the recorder may help researchers interested in studying the mechanisms underlying how chemical exposures early in development can have lasting impacts.
________________
8 Aluru, N., S. I. Karchner, D. G. Franks, D. Nacci, D. Champlin, and M. E. Hahn. 2015. Targeted mutagenesis of aryl hydrocarbon receptor 2a and 2b genes in Atlantic killifish (Fundulus heteroclitus). Aquatic Toxicology 158:192–201. doi: 10.1016/j.aquatox.2014.11.016.
9 Sheth, R. U., S. S. Yim, F. L. Wu, and H. H. Wang. 2017. Multiplex recording of cellular events over time on CRISPR biological tape. Science 358(6369):1457–1461. doi: 10.1126/science.aao0958.
NEXT STEPS FOR THE USE OF GENOME EDITING TOOLS
Speakers discussed a variety of ways the use of genome editing tools could advance environmental health research, including new research platforms and the expansion of knowledge about the biology of environmentally induced disease. Speakers also commented on known limitations and bottlenecks with the use of genome editing tools.
Hahn emphasized that genome editing methods have opened up the world of experimental organisms so that any organism can become a “model organism.” Houck pointed out that the use of genome editing may enable researchers to more rapidly identify animal models with sensitivity and resistance to specific modes of action that are more comparable to humans. He foresees that the tool will help with the generation of sensitive population models both in vivo and in vitro, including single genetic mutation disease models and models with engineered absorption, distribution, metabolism, and excretion (ADME) genetic variability. Houck predicted that the use of genome editing in organisms and cells would help researchers modulate critical ADME parameters that can be part of the modes of action. Several workshop attendees agreed that a broader array of model organisms could lead to significant advances in environmental health and facilitate research with human cells.
Creating synthetic genomes to study gene function is another area where the genome editing technologies can be used for experiments in the environmental health sciences. Woychik said that genome editing tools can be used to complete studies that take much longer and are much more expensive with the traditional homologous recombination techniques. This includes quickly creating synthetic genomes to study the function of single genes, as well as to produce and study single nucleotide variations within promoter, coding, and non-coding regions. Similarly, the tools could also be used for the creation of structural alterations like translocations and deletions. NIEHS researchers and grantees are using genome editing techniques to create mutations in cells and model organisms that were previously difficult or refractory to genetic modification. “Doing targeted mutagenesis in rats, zebrafish, and other organisms and even in various cell lines is now possible,” Woychik said.
Researchers are contemplating how the ability to more easily manipulate a variety of biological systems at various levels of organization with these tools may help broaden the understanding of targets for high throughput testing, Houck said. However, he is not aware of anyone actually doing this yet. However, genome editing is already helping to move discovery in environmental health away from apical endpoints, such as death and gross adverse effects, to upstream events that lead to those adverse effects, observed Stanley Barone of EPA’s Office of Chemical Safety and Pollution Prevention. However, Kaminski pointed out that unlike with drug development, environmental health researchers often do not know what the target or targets are of the chemicals they study. Houck acknowledged that most chemicals have apparent polypharmacology in that they can trigger numerous adverse pathways. This can make the mechanism of action hard to pin down.
While understanding the events that initiate the hundreds of known pathways that are the end-stage actors in toxicity may not be necessary for toxicity screening, the knowledge would be important for advancing understanding of how these agents act, suggested Kim Boekelheide of Brown University. Lu responded that genome editing tools could help in identifying such initiating events. Combining CRISPR with chemoproteomics tools that record direct interaction between a toxicant and the proteome could help identify which hits are important and generate a list of potential molecular initiating events, Vulpe said. Dolinoy shared ideas for how genome editing could be used to assess the biological significance of epigenetic regulation. For example, it might be possible to “knock-in” DNA methylation to assess the consequences of methylation and/or environmental exposures. Another possibility would be to follow an epigenome-wide association study with epigenome editing to figure out which “hits” are most influential. Genome editing may help scientists to improve their understanding of AOPs in terms of variation in the human population, noted Gary Ginsberg of the Connecticut Department of Public Health.
Workshop speakers and participants also discussed how genome editing could help the NIEHS further its current focus area of studying how environmentally mediated changes in the function of cells’ mitochondria organelle impact the biology of organisms. “A lot of environmental toxicants have their deleterious effects on the organism by hitting the mitochondria, so if we better understand how the mitochondria communicates with the nucleus, we’ll have a better understanding of how mitochondrial toxicants are actually influencing the biology of the cell and the organism,” Woychik said. The limited data available thus far show that changing mitochondrial function can change the epigenetic regulation of gene expression in the nucleus. For example, if the mitochondrial DNA is completely eliminated, there is a dramatic impact on the epigenome in the nucleus. Woychik suggested that in the future it would be useful to have the genome editing tools to make subtle changes in the mitochondrial DNA in order to study the function of specific mitochondrial genes. “Having the ability to go in and make changes to the mitochondrial genome is going to have a huge impact on better understanding of how the mitochondria can influence the biology of the rest of the cell,” emphasized Woychik.
Fry pointed out that genome editing could facilitate ongoing environmental health research. For example, her team determined the role that the glucocorticoid receptor (GR) plays in developmental toxicity using a chick embryo model by blocking the GR receptor. They inhibited the receptor chemically, but she pointed out that genome editing with a tool like CRISPR should enable researchers to achieve the same result—and to conduct similar experiments. She also spoke about an experiment her team conducted to gain insights into cadmium’s links to preeclampsia based on data collected about cadmium in placentas from an epidemiological study involving pregnant women. They used a chemical approach to block the transforming growth factor beta (TGF-ß) pathway to show that it plays a role in cadmium-induced preeclampsia, but Fry pointed out that genome editing might also be used to achieve the same end.
Lynae Brayboy of Brown University pointed out that an idea for a target could be gonadotoxic agents that harm human fertility, one of her research interests. She also spoke about her preliminary research suggesting that multi-drug resistance (MDR) genes may have a link to mitochondrial function that could be protective. This is of importance to her as a physician researcher because of her interest in finding ways to help protect the fertility and ovarian function of women who undergo chemotherapy. She has been using genome editing to investigate the role that MDR genes have on the ovary. Her work to date suggests that the genes confer a protective effect that may safeguard both the ovary and the oocytes it contains from exposure to everyday toxicants.
Vulpe said that more and more validated libraries of sgRNA, the guide that tells Cas9 which gene should be targeted, are becoming available. He expects this to improve the utility of genome editing of techniques for identifying which sequences are important for functions of interest.
A number of the workshop participants, including workshop planning committee member Melissa Perry of The George Washington University, brought up the idea of using the tools to evaluate thorny problems such as exposure to mixtures of environmental toxicants and synergistic responses to environmental exposures.
Houck said he is very interested in studying mixtures, but it is a huge combinatorial problem. However, he said it would be possible to set up potential models where scientists expect to see synergistic effects and then use gene editing tools to validate the ideas. Eventually this could lead to the creation of a test system to look for synergies.
Workshop attendees also grappled with how to deal with new and unexpected information that results from these tools, such as revealing compensatory mechanisms of which scientists were previously not aware. Hahn commented that researchers need to keep in mind that compensation will lead to some false negatives, which may ultimately provide key information to biologists about how cells work. “Embrace the unexpected result, because it may be telling you something important,” Hahn said. Hahn also cautioned that the results of genome editing research can be difficult to interpret, especially when there are discrepancies between previous results from knockdown studies and new results from CRISPR/Cas9 knock-out studies of the same genes. He explained that “genetic compensation,” a natural capability of animals or cells to overcome the loss of a single gene, makes the interpretation of negative results from knock-out studies a particular challenge.
Hahn and Kaminski agreed that a combination of all of the complementary approaches to understanding the effects of chemical exposures are likely to lead to deepened understanding. High throughput bioassays, transcriptomics analysis, focused target validation and mechanistic research, and genome-wide CRISPR screening will all complement one another. “CRISPR won’t replace them,” Hahn stressed. Houck added that leveraging animal toxicology data may also help scientists work out some of these details brought to light by the new technology. Fry pointed out that organic compounds that are well represented in databases could be used to create a Venn diagram of what is being seen with Tox21 versus CRISPR in toxicology-relevant enriched gene sets. As research progresses it may help remediate or at least counter-balance genome editing weaknesses. The approach is currently limited to identifying genes that lead to a reduction in cell viability, Houck pointed out. The problems inherent to CRISPR technology include unintended mutations and the fact that pathway analysis is not necessarily definitive, he said. Other weaknesses, Houck and Vulpe agreed, mirror the issues associated with using in vitro approaches and high throughput testing: many of the cells are from immortalized lines and the metabolism and toxicokinetics are not reflective of how living cells function in multicellular beings.
Challenges in minimizing Cas9 off-target effects include contending with the reality that genome-wide assays use cell lines that are basically testing only a single reference, Pattanayak pointed out. If the cell population being studied has naturally occurring genetic variation, it can cause problems. Another potential issue is that applications of genome editing potentially involve the modification of billions of cells, but the current limit of detection for off-target effects is ~1/10,000, he said.
Using the tools to do single nucleotide substitutions is still a challenge, Kaminski said. Their applicability for epigenome editing is still on the horizon, he added. The fact that the action of genome editing needs to be validated by polymerase chain reaction sequencing is also a bottleneck.
WILL GENOME EDITING REQUIRE MODIFIED RESEARCH POLICIES?
Workshop speakers discussed key resources or guidance needed to better incorporate the use of genome editing tools in environmental health research. Whether or not the NIEHS should provide guidance on how gene editing is used by its grantees remains an open question. The fact that genome editing knock-outs are different from knock-downs may give prospective users pause, commented Reza Rasoulpour of Dow AgroSciences.
Fry also noted that guidelines, guidance, quality controls, and probably a set of minimum standards are needed for using these tools in any field. Similarly, she pointed out the need for information on delivery to different tissue types, as well as methodological input for standardization and optimization. The existence of improved gRNA design algorithms and improved prediction of off-target effects could also help environmental health research move forward, Hahn said. Houck suggested that more proof-of-concept and pilot studies be completed with well-characterized chemicals. He also articulated the value of developing assay technologies that facilitate the expansion of functional endpoints beyond cell viability.
Urnov stated that the field would benefit from having the federal government step in to enable the creation of a larger-scale resource for gene editing data. Woychik agreed that he could see a role for the federal government and the NIEHS, but he also noted that ensuring good quality control can be expensive. Margaret Karagas of Dartmouth University pointed out that forming centers may help move the field forward. Woychik responded that the NIEHS’s Children’s Health Exposure Analysis Resource program might be a potential model.
Getting federal support for genome editing research may be challenging. Miller and Hahn agreed that NIH study sections10 tend to be conservative and most supportive of mechanistically focused research. Hahn suggested that NIH’s Request for Applications process, which generally involves narrowly defined areas for which one or more of the agency’s institutes have set aside funds for awarding grants, may be a way to facilitate the embrace of genome editing tools. The field would benefit from “a change in philosophy … and culture” to accept findings generated by new technologies, Fry commented.
DESIGNING RESEARCH TO INFORM DECISIONS
The final portion of the workshop focused on the application of genome editing tools to informing public policy decisions. Houck pointed out that EPA is responsible for assessing the safety of tens of thousands of chemicals and little publicly available data exist on many of these compounds. Cost, time, and ethical considerations preclude the use of traditional toxicology methods to assess these chemicals, he said. He is researching new, more efficient ways to assess chemicals for potential risk to human health and the environment at EPA’s National Center for Computational Toxicology’s Computational Toxicology Center. The center’s ToxCast program uses high throughput in vitro technologies for hazard screening to generate information on data-poor chemicals. It is part of the Tox21 collaboration among NIH, EPA, and FDA that has now screened more than 3,000 chemicals through more than 800 bioassays, cellular and biochemical assays. The center has also screened about 8,000 chemicals, through about 80 different, mostly cellular, assays focused on stress and nuclear receptor responses. All of those data are in a database that can be analyzed and used to build models predictive of toxicity.
By analyzing the data, EPA has shown that about 80 percent of the tested chemicals are active (causing “perturbations” in gene expression) in ten or more bioassays. EPA scientists have recognized for a long time that “this large suite of in vitro assays really begins to approach the potential range of biology that might be affected by the chemicals that we’re testing,” although there is “still lots of unexplored biological space,” Houck said.
Functional genomics, including genome editing, may help researchers to pinpoint the critical mechanism of action in cases of tested chemicals that are active in many bioassays. Functional genomics also can allow scientists to bridge between the genotype, the set of genes in DNA that is responsible for a particular trait, and phenotype, the physical expression, or characteristics of that trait. CRISPR opens the door for higher throughput applications using mammalian cells in cases where yeast cells were used previously, Houck said.
Houck is also optimistic about the potential for using functional genomics for untargeted toxicology and environmental health studies where the biology primarily drives the focus. He said that he believes these kinds of studies “are really valuable, in particular, for the kind of work that we’re doing with so many unknown chemicals.”
Assuming that risk assessment continues to integrate high throughput testing, more proof-of-concept and pilot studies with well-characterized chemicals will help environmental risk assessors integrate functional genomics, including genome editing, Houck said. He would like to see the development of assay technologies that facilitate the expansion of functional endpoints beyond cell viability, for example, cell line engineering of pathway reporters that drive lethality, sorting on reporter gene expression, and single cell RNA sequencing could all generate useful data.
________________
10 Study sections are discipline-specific groups that review research proposals submitted to NIH.
The development of in vivo assays in organisms suited to high throughput testing, such as roundworms (C. elegans) or zebrafish (Danio rerio), would also be beneficial, he said.
Houck predicted that genome editing will prove useful for validating high throughput transcriptomic signatures. The tools can help with validation of AOPs to show what the effect of removing or activating individual genes has on molecular initiating events and other key events related to the adverse endpoint. He added that the tools may help with the development of quantitative AOPs.
Houck and Vulpe are involved in a pilot project to test the feasibility of using CRISPR/Cas9 genome editing in human cells for screening environmental chemicals in a functional genomics toxicology format. Their research teams are trying to identify putative initiating events, as well as mechanisms of action and potential modulating responses that could be used in identifying sensitive populations, Houck said. Other uses of genome editing of potential interest to environmental health, Houck added, include the validation of high throughput transcriptomic signatures and single genetic mutation disease models.
In the discussion following Houck’s talk, Hahn observed that “setting exposure limits used to be easy when all we were worrying about is preventing people from dying from a chemical exposure. Now we’re worried about more subtle effects that generally occur at lower levels of exposure. It’s a hard problem. Adding to the difficulty of the problem is that exposure and effects are often separated widely in time.” In this content, he argued that anything that enhances mechanistic understanding will improve decisions related to policy and regulations regarding environmental health.
“We have been modifying the genome in humans for a long time,” observed Carolyn Wilson of FDA’s Center for Biologics Evaluation and Research. FDA has been regulating gene therapy in general since the first retroviral vector mediated gene therapy trial in 1989, she said. The agency started regulating clinical trials involving various forms of gene editing, including zinc fingers and TALENs, in 2008, generally for very serious or life-threatening conditions. “Regulatory decision-making always comes down to benefit-risk decisions,” she said. The center has been reaching out to the research community about the need for improved algorithms to predict off-target effect of gRNAs and improved methods to detect inadvertent germline editing or other off-target effects in the genome, including insertions, deletions, and chromosomal instability and unwanted immunogenicity that might be conferred.
Treye Thomas of the U.S. Consumer Product Safety Commission observed that genome editing tools’ potential to shed light on the drivers of individual susceptibility in response to chemical exposures might in time allow scientists to identify susceptible populations and perhaps even help them to resist the development of disease. Similarly, Linda Wennerberg of NASA said that in addition to identifying potential harms that can result from environmental exposures, genome editing tools may bring to light details about protective measures, such as ones involving nutrition, that can be taken to make people less vulnerable.
Barone commented that gene editing tools may help risk assessors collect data on susceptible populations to improve the safety factors used to protect susceptible individuals. Currently a factor of 10 is used by default in the absence of better data. “The more we know about susceptible populations or certain life stages where exposure should be limited, the better risk communication we can do. More regulation or better information about personal protection may result,” he said.
Determining how to validate the tools will be very important, Thomas pointed out. A suggestion for validating genome editing techniques by Aylward involved identifying sets of candidate complex pathways from which scientists have already elucidated an understanding and chemicals that researchers already think they know a lot about through ToxCast, Tox21, and high throughput testing. Finding activity patterns and chemicals that activate them may be a step for cross-validation across different approaches, she said.
Fry proposed an experiment whereby genome editing tools could enable research that helps determine human-relevant exposure limits. The goal would be to test how genetically diverse cell populations respond to toxic substances. A first step would be to generate a targeted knock-out in cells, then screen for sensitivity in responses to an exposure. Next would be to establish the genes underlying the responses and use knock-ins to examine SNP function. The final step would be to examine SNPs in human populations.
Wennerberg noted that the decisions risk managers and regulators make are very important because they can inform consumer choices having the potential to impact their health.
For environmental health researchers, this an exciting time, Rasoulpour said at the end of the workshop. The new tools enable research that could never have been conducted before, he said. “If you can think about something that you want to do, chances are that someone is working on a tool to help you do that…. As an optimistic scientist, I think that we actually have the opportunity to take this tool—along with others we have at our disposal—and really make a difference with regard to human health and public health,” Rasoulpour concluded.
Disclaimer: This Proceedings of a Workshop—in Brief was prepared by Kellyn Betts as a factual summary of what occurred at the workshop. The planning committee’s role was limited to planning the workshop. The statements made are those of the rapporteur or individual meeting participants and do not necessarily represent the views of all meeting participants, the planning committee, or the National Academies of Sciences, Engineering, and Medicine.
Reviewers: The Proceedings of a Workshop—in Brief was reviewed in draft form by Rebecca Fry, University of North Carolina at Chapel Hill; Mark Hahn, Woods Hole Oceanographic Insitution; Shaun D. McCullough, U.S. Environmental Protection Agency; and Chris Vulpe, University of Florida to ensure that it meets institutional standards for quality and objectivity. The review comments and draft manuscript remain confidential to protect the integrity of the process.
Organizing Committee on Emerging Genome Editing Tools to Advance Environmental Health Research: From Cells to Populations: A Workshop: Lesa Aylward, Summit Toxicology; David Gerhold, National Center for Advancing Translational Sciences; Norbert Kaminski, Michigan State University, Shruthi Mahalingaiah, Boston University School of Medicine; Gary Miller, Emory University, Reza Rasoulpour, Dow AgroSciences, Treye Thomas, U.S. Consumer Products Safety Commission, Chris Vulpe, University of Florida; Luoping Zhang, University of California, Berkeley
Sponsor: This workshop was supported by the National Institute of Environmental Health Sciences.
About the Standing Committee on Emerging Science for Environmental Health Decisions The Standing Committee on Emerging Science for Environmental Health Decisions is sponsored by the National Institute of Environmental Health Sciences to examine, explore, and consider issues on the use of emerging science for environmental health decisions. The Standing Committee’s workshops provide a public venue for communication among government, industry, environmental groups, and the academic community about scientific advances in methods and approaches that can be used in the identification, quantification, and control of environmental impacts on human health. Presentations and proceedings such as this one are made broadly available, including at http://nas-sites.org/emergingscience.
Suggested citation: National Academies of Sciences, Engineering, and Medicine. 2018. The Promise of Genome Editing Tools to Advance Environmental Health Research: Proceedings of a Workshop—in Brief. Washington, DC: The National Academies Press. doi: https://doi.org/10.17226/25136.
Division on Earth and Life Sciences

Copyright 2018 by the National Academy of Sciences. All rights reserved.












