2
Presentation Summaries
Workshop presenters covered a range of topics relevant to the scientific needs, technological capabilities, funding structures, and system design requirements involved in considerations around convergence.
SCIENCE DRIVERS AND NEEDS
Alexander Szalay (Johns Hopkins University), Eliu Huerta (University of Illinois at Urbana-Champaign), Saurabh Sinha (University of Illinois at Urbana-Champaign), and Shyue Ping Ong (University of California, San Diego) spoke to the following questions relevant to the needs of the scientific community:
- What are the trends and frontiers in simulation and data-driven science?
- What is the distribution of needs for both across the science community?
- How well are current needs being met by today’s facilities and services?
- What new workloads are emerging (e.g., deep learning or graph analytics), and what architectures can (best) support them?
Numerical Laboratories on Exascale
Alexander Szalay, Johns Hopkins University
Cosmologist and computer scientist Alexander Szalay discussed trends in advanced scientific computing and highlighted examples of approaches being employed in physics, cosmology, and oceanography. He made two overarching observations. First, simulations are becoming first-tier scientific instruments in their own right. Second, at exascale, everything becomes a big data problem.
Most scientists increasingly rely on large computer simulations, so it is vital to be able to compare simulations and observational data. As a result, we have to pay attention to how the simulation outputs can be shared more broadly for smaller-scale analyses to answer such questions, now and in the future.
“Numerical laboratories” allow domain scientists to access the results of the best and latest simulations on petabyte scales. In many fields, large experimental facilities such as the Large Hadron Collider are producing enormous amounts of data. They apply in-line triggers to keep only 1 out of 10 million events—the ones that have the
highest probability to contain new discoveries. Even after this enormous down-selection, they still produce many petabytes of data every year. Numerical simulations should follow this example, and should aim to find similar trade-offs in generating their outputs from the most scientifically valuable subsets of the data.
Similarly, numerical laboratories can be used to smaller-scale studies through the use of streaming algorithms that avoid the need for petabytes of local storage. This approach has been demonstrated in various fields of study including turbulence,1 oceanography,2 and cosmology.3
Also, Szalay emphasized the need to shift from capturing complete simulation snapshots to triggering on interesting events in the simulation output. Machine learning can be used to identify these interesting events. The outcome of this approach is a much sparser output covering a wider parameter space that can be stored and made available for offline analysis by a broader community.
As we are on a path toward exascale, simulation outputs grow faster than our ability to store them, and we face trade-offs, such as which data points to store and how much can be computed in situ versus offline. While no scientist will turn down more data, what they actually need is more data that is relevant to the science that they are doing—not more noise.
Frontiers at the Interface of High-Performance Computing, Big Data Analytics, Deep Learning, and Multimessenger Astrophysics
Eliu Huerta, University of Illinois at Urbana-Champaign
Eliu Huerta, an astrophysicist at the National Center for Supercomputing Applications (NCSA), discussed trends and needs in simulation and data-driven science and described two recent demonstrations of convergence. He posited that deep learning should be seen not as an alternative for established algorithms but as a preferred choice to accelerate scientific discovery.
The past decade has brought a new vision for advanced computation in which big data analytics and high-performance computing have equal status; in 2015, the U.S. National Strategic Computing Initiative emphasized the convergence of big data and the high-performance computing ecosystem. Developments in big data and artificial intelligence (AI), made possible by the availability of high-quality labeled data sets and the widespread availability of graphics processing units (GPUs) for parallel computing, have led to numerous commercial applications with substantial economic value. These developments are now percolating into academic research, of which Huerta highlighted two examples.
The first is a novel computational framework connecting NCSA’s Blue Waters supercomputer to the Laser Interferometer Gravitational-Wave Observatory (LIGO) Data Grid. This was accomplished by using software developed for the Open Science Grid as a universal adapter and Shifter, a tool for containerizing analysis software, to seamlessly run high-throughput LIGO workflows concurrently with highly parallel numerical relativity simulations on Blue Waters.4 The framework was used at scale for the gravitational wave detection of two colliding neutron stars by the LIGO and Virgo scientific collaborations.5 To date, Huerta said, this demonstration is the most complex example of convergence on a leadership class computer and was achieved much earlier than had been expected.
A second example illustrates the fusion of high-performance computing with AI. Using GPU-accelerated deep learning using the results of astrophysics simulations as training data, NCSA scientists have enabled faster
___________________
1 For example, Open Numerical Turbulence Laboratory, http://turbulence.pha.jhu.edu.
2 “Oceanography,” SciServer, http://www.sciserver.org/integration/oceanography/, the 2PB is a pending National Science Foundation proposal.
3 For example, Virgo-Millennium Database, http://gavo.mpa-garching.mpg.de/Millennium.
4 E.A. Huerta, R. Haas, E. Fajardo, D.S. Katz, S. Anderson, P. Couvares, J. Willis, T. Bouvet, J. Enos, W.T.C. Kramer, H.W. Leong, and D. Wheeler, 2017, “BOSS-LDG: A Novel Computational Framework that Brings Together Blue Waters, Open Science Grid, and the LIGO Data Grid to Accelerte Gravitational Wave Discovery,” 2017 IEEE 13th International Conference on e-Science (eScience), doi:10.1109/ eScience.2017.47.
5 B.P. Abbot, R. Abbott, T.D. Abbott, M.R. Abernathy, F. Acernese, K. Ackley, C. Adams, et al., 2016, Observation of gravitational waves from a binary black hole merger, Physical Review Letters 119:161101.
real-time detection of gravitational wave signals in raw LIGO data.6 The method has been shown to recover the signals detected by the LIGO scientific collaboration’s standard detection system and been used to identify new classes of signals that are beyond the scope of established LIGO detection algorithms. The approach requires minimal computational resources (a single GPU as opposed to large-scale computer clusters). The researchers use scientific visualizations of the model’s neural network to facilitate interpretability.
Looking forward, Huerta noted that machines like Summit, which will be deployed at Oak Ridge National Laboratory in 2018, will provide thousands of sophisticated GPUs, allowing the creation of deeper and bigger deep learning models. This fusion at scale of high-performance computing and AI promises to transform the practice of science by enabling the analysis of large data sets that would otherwise remain intractable.
Data Science for Genomics
Saurabh Sinha, University of Illinois at Urbana-Champaign
Saurabh Sinha co-directs the National Institutes of Health BD2K Center of Excellence at the University of Illinois at Urbana-Champaign and Mayo Clinic (known as the “KnowEnG Center”). He spoke to the ways in which big data is generated and used in the biosciences, noted gaps in the use of tools and simulations, and highlighted examples of how deep learning and graph mining are being applied in this space.
Like many fields, biology has seen a data explosion, particularly in the area of genomics. One extrapolation estimates that this data will reach tens of exabytes by 2025. Genomic data is held in standard repositories, such as GenBank, as well as in an array of specialized repositories such as those focused on specific diseases or organisms. Efforts to make the most of this data focus largely on increasing access to genomic data, advancing tools for analysis, and improving interoperability among various databases and tools. Although there are efforts to bring data and tools together, these efforts are in their early stages and data and tools remain largely disconnected, Sinha said.
Because many biological systems are under-specified, with the components and interactions incompletely understood, the use of simulations to model biological phenomena is fairly limited. Most such simulations are small scale and focus on areas such as systems biology, networks, and pathways. Far more common in biology are a different type of simulation—statistical simulations that are used to assess the significance of patterns extracted from complex, heterogeneous data. These simulations use various kinds of bootstrap sampling and permutation tests to assess statistical significance, akin to simulating the null distribution.
Sinha offered examples of how deep learning and graph analytics approaches are being applied to advance genomic research. The KnowEnG Center is working to develop a cloud-based portal for genomics data mining that would allow scientists to upload their data and then select one of many different functionalities, primarily machine learning operations, to apply to it. These functionalities will incorporate prior knowledge from a “Knowledge Network” of heterogeneous data aggregated from existing annotated databases that describe genes and their relationships.
The system is hosted on Amazon Web Services (AWS), and its functionalities are containerized, orchestrated by a combination of mesos and chronos. The KnowEnG team is working to support interoperability between different cloud-based systems such that a scientist could use tools from one cloud system, data from another, and the deep learning functionalities in the KnowEnG cloud seamlessly.
___________________
6 D. George and E.A. Huerta, 2018, Deep neural networks to enable real-time multimessenger astrophysics, Physical Review D 97:044039.; D. George and E.A. Huerta, 2018, Deep learning for real-time gravitational wave detection and parameter estimation: Results with Advanced LIGO data, Physics Letters B 778:64-70.
Data-Driven Materials Science
Shyue Ping Ong, University of California, San Diego
Materials scientist Shyue Ping Ong described how data and simulations are used in materials science, a field that has embraced a big data perspective and machine learning techniques more recently than fields addressed by other workshop presenters, such as physics, astronomy, and genomics, and that involves comparatively smaller data sets.
Data-driven materials science is propelled by four major approaches: high-throughput calculations (in use for the past decade), combinatorial experiments (a newer approach), data mining, and machine learning (both of which have been adopted in response to the increasingly large data sets being generated). Although researchers have not yet successfully combined all four techniques for true data-driven materials design, various subsets of these approaches have been effectively combined to make new discoveries.
For example, data mining of the results of high-throughput calculations has been used to predict new materials with certain combinations of characteristics. One database facilitating such activities is the Materials Project.7 This database, used by about 45,000 researchers, is quite extensive; in the area of crystals, for example, it contains about 70,000 known and predicted crystals, which users can explore for desired properties such as energy, elastic constants, and band gap. Researchers developed a completely new phosphor by using this database to identify unexplored chemistries, applying machine learning to predict structures of crystals in that space, using high-throughput calculations to calculate descriptives, and ultimately synthesizing the designed crystal in the laboratory.
Scientists also have used machine learning on the results of high-throughput calculations to build models for material properties and to model ways to bridge the gap between accuracy and scale, a persistent challenge in materials science. In addition, data mining and machine learning have been applied to combinatorial experiments to carry out up to tens of thousands of experiments in high-throughput mode. This approach was used to discover a new photo-anode that outperforms previous ones and is being applied to explore possible new compounds for rechargeable lithium batteries.
Ong emphasized that many of the needs of the materials science community are well served by high-throughput, modest-scale computational resources, such as the San Diego Supercomputer Center’s Comet system. As machine learning models mature, it may be useful to have access to exascale computers, he said, but in general the system scale requirements of the materials science community are modest. He added that GPU resources are particularly useful for deep learning, which is nascent in materials science but holds the promise of performing better than traditional architectures for data-driven materials science.
ENVISIONING A CYBERINFRASTRUCTURE ECOSYSTEM FOR AN ERA OF EXTREME COMPUTE AND BIG DATA
Manish Parashar (National Science Foundation) offered an overview of how NSF views the role of cyberinfrastructure in scientific advancement, recent rapid changes in this space, and ways in which cyberinfrastructure must evolve in response.
Manish Parashar, National Science Foundation
Manish Parashar, director of NSF’s Office of Advanced Cyberinfrastructure, provided an overview of NSF’s investments in cyberinfrastructure and discussed how the agency plans to evolve its approach in response to changing needs and opportunities.
___________________
7 The Materials Project, https://materialsproject.org/.
NSF’s Role in Cyberinfrastructure
Noting that cyberinfrastructure is central to all aspects of NSF’s mission, Parashar said his office’s mission is to foster the cyberinfrastructure ecosystem to transform computational and data-intensive research across all of science and engineering, which includes supporting research cyberinfrastructure as well as cyberinfrastructure research and innovation.
NSF leads and participates in many programs that support computing, data, software, networking and cybersecurity, and learning and workforce development. NSF also invests in leadership class computing resources, as well as an array of smaller systems, and supports these with Open Science Grid, a distributed platform that coordinates distributed resources. Parashar noted that the fiscal year (FY) 2018 budget gives NSF a $300 million increase over the FY2017 budget, the second largest increase in the NSF research budget in 15 years. This increase is centered around NSF’s “Big Ideas” that are driving the agency’s long-term research agenda and processes, one of which is midscale research infrastructure.
Current and Future Developments
Parashar recognized a number of developments that are changing the way science is done and, as a result, changing NSF’s view on how it can best support research cyberinfrastructure. NSF’s challenge is to determine how to allocate limited funding such that its investments add value and complement investments from universities and other agencies and enterprises.
One development is the rapid increase in data, both from simulations and from instruments and observatories. Simulations are now producing so much data that it may make sense to include analytics and data processing along with simulations in scientific workflows to reduce the costs of processing and moving all that data. There is also an increasing need to feed collected data into the simulations to improve modeling. A result of this, is that it is becoming more important for different resources (observatories and high-performance computers, for example) to seamlessly communicate with each other, both from a technological perspective and in terms of human operators.
Scientists are also changing the way they use computational resources. While some rely on very large systems, the “long tail” of science is growing wider and thicker, Parashar said, with smaller-scale demands (using 1,000 cores or less) now dominating in terms of total units of computing used. Scientists are also increasing their use of cloud and edge services, and nontraditional software stacks such as Spark, Kafka, and Jupyter, which existing high-performance computing resources were not built to accommodate. Use of disruptive technologies and accelerators has also advanced rapidly, now including not only GPUs but application-specific integrated circuits for machine learning such as Google’s tensor processing units (TPUs), which will become more critical if deep learning continues to become a bigger part of the science workflow.
A request for information NSF issued in 20178 provides insights into the scientific community’s expected cyberinfrastructure needs over the next decade and beyond. Responses reflect a growing need for computing as well as continued changes in how computing infrastructure is used, with an increasing emphasis on on-demand computation, rapid data processing, comparisons between simulations and observations, data management, machine learning, big data techniques, and streaming data from the Internet of Things, large instruments, and experimental facilities. Discoverability, accessibility, and reproducibility are key concerns, and software training and workforce development will be needed to keep pace with capabilities.
Another key challenge is anticipating and responding to the cyberinfrastructure needs of Major Research Equipment and Facilities Construction (MREFC) projects. Because these large facilities can take a decade from conceptualization to deployment, there are always changes in cyberinfrastructure requirements as well as the technology landscape; for example, the amount of data involved, the technologies and instrumentation that can be leveraged, and the way data is delivered and used are not accounted for in the architectures.
There are a number of emerging cyberinfrastructure challenges rated to MREFC projects. How can data, computing, and applications be better connected? What’s the best way to support on-demand processing and on-
___________________
8 For full results, see National Science Foundation, “NSF CI 2030 Overview,” https://www.nsf.gov/cise/oac/ci2030.
demand analytics for large amounts of data? How can we move from existing silos to a more integrated system that seamlessly links data-generating facilities with high-performance computing resources?
Guiding Principles Going Forward
Going forward, Parashar outlined three guiding principles for NSF. The first is to approach cyberinfrastructure more holistically to support scientists’ needs, rather than approaching it in terms of separate silos for software, data, networking, and computing. The second is the need to support translational and foundational innovation while at the same time enabling the sustainable services that support the scientific needs, a balancing act that requires sustained funding. Third is the need to more closely couple the cycles of innovation and discovery on the science side and the cyberinfrastructure side to enable greater translational impact.
ARCHITECTURAL LANDSCAPE AND TRENDS
Pete Beckman (Argonne National Laboratory), David Konerding (Google, Inc.), and Sanjay Padhi (AWS) addressed the following questions relevant to the evolution of architectures for high-performance computing and data-centric computing:
- How have high-performance computing and data-centric computing architectures been evolving and where are they heading? What science and non-science applications are driving their evolution? Where are the points of convergence and divergence?
- Are the memory needs the same for simulation and data-intensive workloads?
- What are appropriate file systems that could support both high-performance computing and data-centric computing? Are there lessons from the high-performance computing side that could inform the data side?
- How are the associated software stacks evolving? To what extent are they converging?
Evolving Toward Convergence
Pete Beckman, Argonne National Laboratory and University of Chicago
Computer scientist Pete Beckman launched the workshop session on evolving architectures toward convergence of high-performance computing and data-centric computing.
Under the traditional paradigm, high-performance computing infrastructure is designed around achieving impressive levels of computation, with far less emphasis on establishing workable systems for giving users access to the system, managing files, and connecting with data-collecting instruments, Beckman said. However, the limitations of that classic model are becoming increasingly apparent as science becomes more data-driven and as demand for access to computing resources expands to a broader swath of the scientific community. The Big Data and Extreme-Scale Computing workshops and report9 articulate the need for a strategy to better align the high-performance computing community and the data science community so each can better leverage the other’s work and resources. While Beckman acknowledged that bringing these divided communities together may be an uphill task, he emphasized that it is nonetheless worthwhile.
Outlining the need for convergence, Beckman emphasized the central role of computing in modern research and described the diverse ecosystem of resources involved. Computing resources and data generators are arrayed along a continuum from a large number of small, inexpensive Internet of Things devices to a small number of large, expensive supercomputers, clouds, and instruments. A new project, which sits roughly in the middle of that continuum, is the Array of Things,10 which is deploying hundreds of nodes in U.S. cities that include sensors and
___________________
9 “The Basis for BDEC,” Big Data and Extreme-Scale Computing, last updated 2018 http://www.exascale.org/bdec/
10 Array of Things, https://arrayofthings.github.io/.
cameras, as well as 12 cores and a GPU. These nodes can perform computations onsite to monitor and study urban problems such as traffic and pedestrian safety.
Compared to the resources of industry leaders like Amazon and Microsoft, Beckman said that research computing infrastructure is lagging far behind, in large part because of the high-performance computing community’s historic singular focus on floating point operations per second, latency, and bandwidth. While those factors remain important, he posited that it is vital to invest in building flexibility into the software stack and even in the hardware, an approach Beckman calls fluid high-performance computing.
To achieve the needed flexibility and make this infrastructure easier to use, Beckman proposed a three-pronged approach: replace the classic HPC file system with flexible storage services that use token-based authentication, tweak high-performance computing interconnects to allow software-defined networking, and make it possible to clear a node quickly and immediately make it available to another user. Such updates would make this infrastructure more competitive with commercial cloud services, which are currently far easier to use than research cyberinfrastructure, and also better position it to integrate AI hardware that is coming down the pipeline. But achieving these changes, Beckman noted, will require the community to overcome its reluctance to tweak hardware and move beyond a “curmudgeon” culture that is too closely tied to past assumptions and ways of operating.
Google’s Tools for Convergence
David Konerding, Google, Inc.
David Konerding is a software engineer at Google, Inc., who works on production infrastructure for high-performance computing. He has a background in computational biology and bioinformatics.
Google has reorganized its architecture around machine learning, investing heavily in hardware and software to support machine learning systems. Its newer systems, based on the flagship technology TensorFlow, challenge many long-running assumptions in ways that are relevant to the drive toward convergence in the research community. Echoing Beckman’s emphasis on flexibility, Konerding said the advantage of TensorFlow stems from its ability to help the underlying system determine where to run, optimizing the flow of data from storage to compute elements and between machines.
Google’s previous architecture,11 designed primarily to support search and advertising, had high aggregate throughput, low individual throughput, high latency, high machine failure rates, and high stability maintenance requirements. By contrast, its new architecture, designed for a wider range of internal customers, is driven by machine learning. It provides much more flexible configuration of GPUs, TPUs, memory, disk storage, and the cluster network. The cache, PCIe and TLB are still fairly inflexible, however, making these parts of the system the most common source of contention among jobs running on the same host.
TensorFlow represents convergence between Google architecture and high-performance computing, Konerding said. Available as an open source software product that can run on any system, TensorFlow is essentially a tool for configuring computation graphs that is able to execute those graphs with large amounts of data flowing through them and optimize the graphs with compiler techniques.
TPUs, which represent a significant increase in total computational power, are especially useful for machine learning workloads. Google’s file systems (in which Colossus has largely replaced Google File System) also support high-throughput computing. Google’s Spanner database service addresses many problems with pre-existing NoSQL and SQL systems, scaling to very large data sizes and query rates. The NVLink communications protocol allows GPUs to trade machine learning weight data with minimal host CPU involvement. Support for these new architectures, as well as the message passing interface (MPI) long used in high-performance computers, make TensorFlow attractive to those wanting to run scientific simulations.
___________________
11 Documented in L.A. Barroso, J. Clidaras, and U. Hölzle, 2013, “The Datacenter as a Computer: An Introduction to the Design of Warehouse-Scale Machines,” Google, https://ai.google/research/pubs/pub41606.
Konerding suggested the future will bring increased convergence between the architectures and software used by Google and the supercomputing community noting that there are numerous opportunities for industry-academic-government collaboration to improve architectures for peak simulation performance.
Architectures at Amazon Web Services
Sanjay Padhi, Amazon Web Services
Research and technical computing expert Sanjay Padhi presented examples illustrating how AWS and its architectures support various fields, including AI, machine learning, big data analytics, and high-performance computing to help advancements in scientific computing. Dr. Padhi also discussed the AWS Research Initiative,12 which fosters innovation in research in collaboration with NSF.
Computer architectures are evolving at a rapid pace, mainly in response to the changing needs and capabilities, also driven by the use of machine learning for applications such as natural language processing, convolutional neural nets in image and video recognition, autonomous vehicles, and reinforcement learning in robotics, as well as high-performance computing for studies in areas that include computational fluid dynamics, financial and data analytics, weather simulation, high-energy physics, and computational chemistry. Padhi highlighted ways that AWS provides capabilities that support these ever-changing needs.
Computing resources have become more heterogeneous; nowadays, different architectures are needed for application specific workflows such as compute, storage, memory optimized instances. For example, Padhi emphasized in particular the rapid development of accelerated computing that facilitate dramatic improvements in performance for deep learning applications. GPUs and other resources support visualization and three-dimensional (3D) rendering at a large scale. Also, increasingly important are trigger-based serverless-computing, container orchestrations, networking on-instance, internode communications (including NVLink fabrics), and elastic file systems.
Accelerated computing can not only support machine learning but standard analytics, as well. Field programmable gate arrays (FPGAs) can play an important role in scientific research. AWS has developed various methods to help users program FPGAs and also shares pre-built accelerators through the AWS Marketplace.
Padhi pointed to several case studies to illustrate the value AWS provides to science and engineering research:
- The ATLAS and CMS experiments at the Large Hadron Collider make use of AWS’s elasticity and scalability.13
- Clemson University researchers while studying natural language processing used more than 1,100,000 vCPUs on Amazon EC2 Spot Instances running in a single AWS region. Spot instances allow users to draw spare compute capacity from any of the AWS regions around the world to quickly process high-throughput applications at lower cost.14
- DigitalGlobe, a major provider of high-resolution Earth imagery, uses a raster data access framework to extract specific areas from archived data sets and Amazon SageMaker to handle machine learning at any scale. This completely archive-based method allows access to 100 petabytes of data without having to move them to live object-based storage.
- The Rice Genome Project started as a small database to allow users to find the best rice strains given soil and weather parameters; it has since grown to include more than 3,000 rice varieties from 89 countries.15 The story underscores the value of building community around resources of common interest.
___________________
12 Amazon Web Services, AWS Research Initiative (ARI), https://aws.amazon.com/government-education/research-and-technical-computing/nsf-aribd/.
13 For example, J. Barr, 2016, “Experiment that Discovered the Higgs Boson Uses AWS to Probe Nature,” AWS News Blog, March 30, https://aws.amazon.com/blogs/aws/experiment-that-discovered-the-higgs-boson-uses-aws-to-probe-nature/.
14 For further discussion, see J. Barr, 2017, “Natural Language Processingat Clemson University—1.1 Million vCPUs and EC2 Spot Instances, AWS News Blog, September 28, https://aws.amazon.com/blogs/aws/natural-language-processing-at-clemson-university-1-1-million-vcpus-ec2-spot-instances/.
15 Amazon Web Services, “3000 Rice Genomes Project,” https://registry.opendata.aws/3kricegenome/.
SERVICE, USAGE MODELS, AND ECONOMICS
Roger Barga (AWS), Thomas Furlani (University at Buffalo), and Vani Mandava (Microsoft) addressed questions relevant to the implications of different usage and service models:
- What are the characteristics and implications of different usage models, such as remote access, co-located computing, and community compute, over data as a whole?
- What are the characteristics and implications of different service models, such as allocated time on high-performance computing systems and purchased on-demand cloud services?
- What metrics should be used (e.g., cost/performance, throughput, researcher productivity) to evaluate different usage and service models?
Stream Processing and Simulation at Amazon Web Services
Roger Barga, Amazon Web Services
Roger Barga is general manager and director of development at AWS. He described Amazon’s efforts to advance two main fronts: stream processing and simulation. Noting that the types of computing problems being grappled with in scientific research are exactly the problems many private-sector businesses and enterprises aim to tackle in the next 5 to 10 years, Barga said there are opportunities for AWS advancements to help researchers and for the science community to help inform industry roadmaps.
Stream processing becomes important when data is being generated at such high volumes that it would be impossible to store it all for later processing. Stream processing will become increasingly critical as more and more continuous data generators, such as Internet of Things devices, come online. Although existing open-source software may be good at ingesting streaming data, Barga said this software falls short when it comes to actually processing the data to perform real-time analytics capable of triggering actions and alerts.
The Robust Random Cut Forest concept16 offers one solution to process such data streams and extract meaningful information on the fly. One reason the traditional machine learning paradigm does not work well with high-volume heterogeneous streaming data is because it can be hard to acquire suitable training data to iteratively develop decision trees. The Robust Random Cut Forest approach continually refreshes the ensemble of trees with new data as the stream evolves in time, keeping a little history as it goes forward—essentially creating a sketching library for streaming data.
The approach has proved effective for detecting anomalies, and developers are experimenting with the ability to use it to show directionality (what makes a data point an anomaly and the direction in which it is changing), as well as to identify hotspots. With these capabilities, developers aim to enhance real-time, actionable insights and forecasting. One specific goal is to reduce false alarms by verifying whether or not an anomaly is meaningful using a semi-supervised approach that makes guesses and then incorporates user feedback.
On the simulation front, Barga said AWS is well positioned to offer developers a scalable distributed laboratory in which to run simulations to verify their apps and products. The cloud is an attractive option for simulations because it allows massively parallel, multiuser, serverless computation drawing on data that is stored cheaply. Clouds save companies the capital cost of clusters and are also more flexible than investing in hardware, since a given cluster might be optimized for some simulations but unsuitable for others.
For today’s business needs, simulation is not a single, one-cut deal but a data and compute-intensive exercise done periodically. Barga pointed to Tesla, iRobot, and other companies that are using extensive simulations on AWS to test software releases before they are deployed. These tests involve many steps to verify that the instructions are being executed as intended, incorporate the physics or chemistry of various interactions, perform ensemble testing, pick specific metrics and run simulations in batches, and finally evaluate whether the product achieves
___________________
16 For full description, see S. Guha, N. Mishra, G. Ory, and O. Schrijvers, 2016, “Robust Random Cut Forest Based Anomaly Detection on Streams,” Proceedings of the 33rd International Conference on Machine Learning, New York, NY, http://proceedings.mlr.press/v48/guha16.pdf.
what it sets out to achieve. The simulations involved in these tests are built on a wide variety of elements, from mathematics to real-time streaming data. Reinforcement learning to tune the simulation is very high value because there may be limited opportunity for follow-up refinements once the product is deployed.
Campus-Based Systems and the National Cyberinfrastructure Ecosystem
Thomas Furlani, University at Buffalo
Thomas Furlani directs the Center for Computational Research (CCR) at the University at Buffalo. In addition to managing a campus high-performance computing system, CCR has an OpenStack-based cloud system to support research and education and is collaborating with Cornell University and the University of California, Santa Barbara, on an NSF Data Infrastructure Building Blocks (DIBBs) award to federate local clouds with the capability to burst to public clouds.
Furlani presented an analysis of the workloads supported by CCR (an example of a campus-based system), Blue Waters (a national capability class system), and the NSF Innovative High Performance Computing Program resources (national capacity class systems whose resources are allocated by XSEDE) (see Figure 2.1) to offer insights on the usage patterns, benefits, and drawbacks of different service models. He argued that the current NSF model of supporting campus-based systems, national capability class systems, and national capacity class systems provides a diverse cyberinfrastructure ecosystem to support academic research needs and that, given the critical role that simulation science and data-driven science play in science and engineering as well as the U.S. economy, additional funding is needed to support acquisition at all of these levels.
No matter the service model employed (cloud, national systems, or campus systems) to support simulation and data-driven science, Furlani emphasized that local staff with expertise in computational science and data analytics are needed to work with faculty and students in order for them to fully leverage existing resources. This is especially true for researchers in domain sciences without a long history of utilizing high-performance computing.
Furlani outlined four main reasons why moderate sized campus based systems are an important part of the cyberinfrastructure ecosystem. The first is that the needs of many academic researchers are concentrated in the small to moderate scale. Furthermore, many scientific application codes do not scale well beyond a few hundred
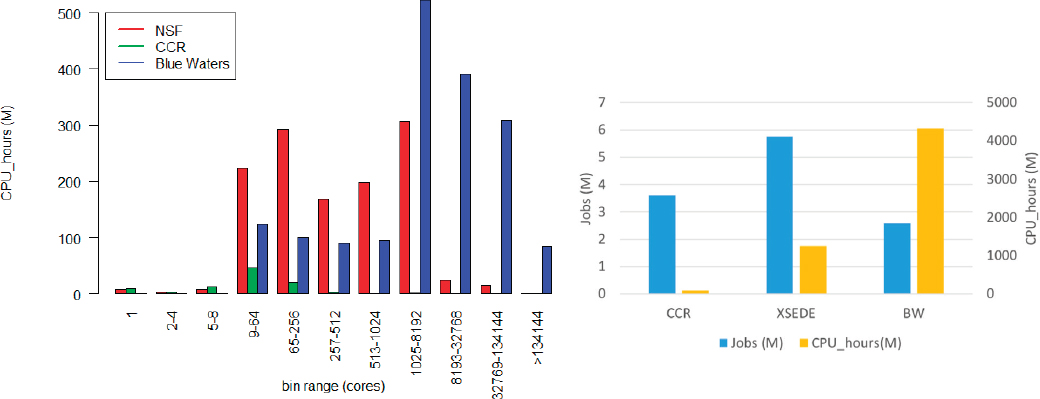
cores, and some problems being addressed by researchers are not large enough for the application to require more large numbers of cores. As a result, a significant portion of the traditional high-performance computing workload on campus-based systems is for jobs that run on 32 cores or less. Thanks to increased processing power, this means that many jobs can now be run on a single node. However, at the same time, data storage and sharing needs are growing rapidly, making cloud systems an increasingly attractive option for universities.
Second, many large national facilities are optimized for the most demanding computational problems with expensive interconnects and file systems designed for large I/O rates, and not necessarily configured to efficiently support smaller jobs (tens to hundreds of processors).
Third, the operating costs (power, cooling, and staffing) of campus systems are typically borne by the host institution, thereby maximizing the return on NSF’s investment. Finally, campus cyberinfrastructure plays a crucial role in the training of a next generation to create and support new tools and methodologies.
For these reasons, Furlani argued that the best way to continue to support academic research needs is to continue to cultivate a diverse cyberinfrastructure ecosystem with a range of usage models and service levels.
Cloud Computing with Microsoft Azure
Vani Mandava, Microsoft Research
Vani Mandava, director of data science at Microsoft Research, described partnerships with the academic community aimed at helping researchers adopt Microsoft’s cloud computing service, Azure. Lessons learned from those experiences will help inform how Microsoft provides cloud services to researchers in the future.
In a period of just over 3 years, Microsoft issued more than 1,600 cloud awards (generally worth $5,000 to $45,000) to more than 800 universities in 74 countries. Roughly half of the projects were computer science focused and half were driven by other science applications. Microsoft also targeted the data science community as part of this effort by donating cloud services to the NSF Big Data Innovation Hubs.
One particularly powerful benefit of using cloud services for research is the flexibility in terms of the types of virtual machines that can be utilized. Over the years, Azure has provided more than 100 options for virtual machines, from general purpose to computer-optimized, memory-optimized, GPU, FPGA, and high-performance computing. Although some researchers often express the belief that it is difficult to use the cloud, Mandava said most were ultimately successful finding the right virtual machine for their research needs.
Universities used their cloud awards for a wide variety of research tasks, including conducting scientific simulations and analyzing observational or experimental data. (They also used the awards for non-research purposes such as to deploy smart water management infrastructure on campus or to provide computing resources for student projects.) The most common way of using Azure was simply to take the same simulation or project a researcher had running onsite and shift it to the cloud—an approach that is particularly beneficial when a researcher’s needs exceed the number of cores available to them onsite. This ability to tap computing capacity on demand significantly improved research productivity, allowing many researchers to go from submitting a proposal to generating tangible results in about a year. Others used higher-level Azure services such as IoT Hub, Spark, HDInsight, and Apache Storm to perform real-time processing and analytics. While many used CPU and GPU based virtual machines, multinode compute clusters, and blob storage, the specific services used varied by scientific domain.
There are also challenges to adopting cloud services for academic research. Procurement can be particularly vexing because it is difficult to sustain pay-as-you-go data storage in the long run when funding comes from fragmented, temporary sources or one-time donations. It is also important to consider costs beyond those associated with the service itself, including costs around supporting users and data hosting, sharing, and long-term retention; however, these same considerations are also important with non-cloud or on-campus cloud systems and may in some ways be easier to address with commercial cloud infrastructure. Also as with onsite systems, it is vital for universities to educate their users about how to make use of cloud services system and allocate and monitor their usage.
DATA STORAGE AND RETENTION
Robert Ross (Argonne National Laboratory) and Robert Grossman (University of Chicago) spoke to the following questions related to the implications of different data storage and retention models:
- What are characteristics and implications of different data usage models (e.g., continuous online versus episodic access, local versus remote storage)?
- What are the implications in terms of cost and future infrastructure needs?
- How should researchers, funders, and institutions make decisions about what data to retain? What are the implications for future platforms?
Data and Convergence at the Department of Energy
Robert Ross, Argonne National Laboratory
As a computer scientist at Argonne National Laboratory and director of the RAPIDS Institute for Computer Science and Data, Robert Ross drew upon Department of Energy (DOE) programs and experiences to highlight insights about data usage and retention in the context of convergence. While DOE’s high-performance programs and systems have historically been largely focused on simulation, they increasingly support a wide range of data analysis and machine learning applications, reflective of the convergence happening across scientific research.
To frame how we think about data storage, Ross stressed that it is becoming increasingly important to differentiate among different classes of data, recognizing that the right data storage solution often depends on how long the data will need to be stored. For example, it may be best to store temporary data in the system, store longer-term “campaign” data in an external parallel file system, and use tape to store data indefinitely. Performance analysis, which provides insights on how system components are working to identify problems and tune performance, is also useful. Performance analysis capabilities are enhanced by newer learning approaches and automated root cause analysis.
In terms of implications of data storage needs for cost and infrastructure, Ross underscored the likely need for a great deal of solid-state storage, complemented by spinning disks and tape. How much data needs to be stored on these various components varies from facility to facility, depending on how long the data needs to be stored and how much data comes from events or experiments that cannot be repeated. For example, a supernova only explodes once, so data capturing that event would need to be stored for a much longer period than data generated for a specific short-term project.
There are also important implications for the software side. While historically storage options have been limited to a single, facility-specific file system, there is a growing trend toward drawing on different services to meet different storage needs. Alternative, flexible approaches are emerging in which data is moved in bulk back and forth between in-house and external systems.
Determining what data to retain remains a challenge. Noting that many facilities have weak systems for cataloguing data resources, Ross suggested steps toward smarter data retention systems could include indexing capabilities or even applying a graph approach to file system metadata. Going further, some decisions about what to retain could be automated based on an understanding of the relationships between jobs and data products and among multiple data products, enabling an assessment of the implications of losing a given data product. On the data generation end, researchers are moving toward in situ data analysis, which can help to identify the most important data to keep and reduce storage needs.
Ross concluded with three thoughts: (1) high-performance computing systems are already seeing a mix of simulation, analysis, and learning applications, and this is already having an impact on storage and I/O requirements; (2) software supporting data requirements is changing in response to technology changes and application requirements, with services moving into the compute fabric and new classes of services augmenting and/or replacing file systems; and (3) before we can make good decisions on what data to retain, we first need to consider how to better understand what data has been created, where it came from, and how it relates to other data products.
To Keep, or Not To Keep, That Is the Question: And Whether Convergence, Clouds, or Commons Is Better
Robert Grossman, University of Chicago and Open Commons Consortium
Robert Grossman discussed different architectures to support data science. He began by observing that there are two Branscomb pyramids17—one for simulation science and one for data science. Possible architectures for data science include converged architectures, architectures leveraging large-scale cloud computing, such as data commons, and what are sometimes called narrow middle architectures for data ecosystems.
Data commons co-locate data with cloud computing infrastructure and commonly used software services, tools and applications for managing, analyzing, and sharing data to create an interoperable resource for a research community. Data commons are one of the architectures used to support data science. Data ecosystems integrate multiple data commons, cloud computing resources, and various services, applications, and resources to support data science for a research discipline or disciplines.
In Grossman’s view, a data ecosystem consists of multiple integrated resources and software services that support a data science research discipline or domain, including resources for managing data, processing data, analyzing data, exploring data, collaborating with data, and sharing data. Software services in a data ecosystem include services for authentication, authorization, digital IDs, data access, metadata, and task and workflow execution services.
Grossman emphasized that the goal of data science is to make discoveries using data, not just to support computation using data. From this point of view, there are two critical architecture decisions and two corresponding trade-offs. The first decision is the architecture of the system elements that manage and analyze data. Here the trade-offs are the typical ones associated with designing a system for a fixed cost. They include how to split investments among computing resources such as CPUs, GPUs, and TPUs; storage including memory, disk, remote disk, and so on; and network communications.
The second decision, according to Grossman, is defining the architecture of the data ecosystem as a whole. Here the trade-offs include the following: How much should be invested in generating the data, adding the data to the ecosystem, managing the data, and supporting analysis of the data to make discoveries? It is sometimes useful to view these trade-offs over the lifetime of the data: there are costs for the generation, processing, and analysis of the data by the first project that uses the data; costs for curating and harmonizing the data so that it can be used by other projects within the same discipline; and further costs so that the data can be re-used by other projects in other disciplines or domains. Key to these trade-offs are decisions about when to discard data.
CONVERGENCE OPPORTUNITIES AND LIMITS
Daniel Reed (University of Utah), Michela Taufer (University of Tennessee, Knoxville), and Douglas Kothe (Oak Ridge National Laboratory) examined the opportunities—and limits—of convergence, with attention to the following questions:
- What are the trade-offs between more specialized and more general architectures?
- How can lessons learned in the high-performance computing and data-centric worlds be used to inform development in the other?
- How much leverage does the scientific community have to drive future architectures?
- To what extent do architectural decisions affect what science you can do?
- What are some plausible directions for convergence?
___________________
17 The Branscomb pyramid was first used in a 1993 committee report to the National Science Board and was named after Lewis Branscomb, the committee chair. It relates computational power (y-axis, more is up), the number of systems acquired (also y-axis, fewer is up), and number of users supported (x-axis). See https://www.nap.edu/read/21886/chapter/6#67.
Convergence Lessons: Future Infrastructure
Dan Reed, University of Utah
Dan Reed is senior vice president for academic affairs at the University of Utah; until recently, he served as vice president for research and economic development at the University of Iowa. He explored how big data challenges social and technical expectations and outlined strategies for forging a path forward.
We are shifting from a world where data was rare, precious, and expensive to one where it is ubiquitous, commonplace, and inexpensive. Massive streams of digital data from scientific instruments and Internet of Things devices, combined with deep learning and inexpensive hardware accelerators, are bringing new data-driven approaches, challenging some long held scientific computing beliefs, and illuminating old questions in new ways.
Even as the underlying hardware continues to diversify due to the end of Dennard scaling and increasing focus on accelerators and FPGAs, Reed asserted that the traditional scientific computing software ecosystem and the new data analytics and deep learning ecosystems are not converging. Today, the rapid developments are concentrated at the two extremes: the very small (edge computing and sensors) and the very large (clouds, exascale, and big data). This has profound technical, economic, and cultural implications for future cyberinfrastructure.
Reed highlighted key considerations and offered several strategies for dealing with these implications. The explosion of data, he suggested, points to a need to develop robust content distribution networks for scientific data, expand edge networks for sensor data capture, and develop continuum technical computing models that integrate the two extremes. Economically, there is a need to develop data curation marketplaces that allow the community to triage data and decide what data should be shared and retained and for how long. Likewise, it is important to be clear about the balance of investments in on-campus versus public cloud infrastructure. Finally, it is important to balance the need for long-term, strategic plans and infrastructure investments against the rapidly changing landscape of technical capabilities.
Reed noted that computing is not alone in its struggle to balance the expanding and diverse needs of its research community. Other domains are also struggling to balance investments in high-end infrastructure deployment and smaller, day-to-day investments that are the lifeblood of the scientific enterprise. In developing future cyberinfrastructure for science, Reed concluded, we must consider the needs of high-end users as well as the long tail, the fusion of data analytics and computing, the nature of public-private partnerships, and both special and general purpose infrastructure, while grounding all decisions in long-term plans and economic realism.
Opportunities for Overcoming Data Bottlenecks
Michela Taufer, University of Tennessee, Knoxville
Michela Taufer is a professor and leader of the Global Computing Lab at the University of Tennessee, Knoxville; until recently she was an Association for Computing Machinery Distinguished Scientist and a J.P. Morgan Chase Scholar at the University of Delaware. She outlined some of the key challenges to achieving convergence.
Taufer stressed that I/O and other data movement will become an increasingly important bottleneck; although floating-point operations have become ever cheaper, data movement remains expensive. Likewise, with the move to exascale computing, computing performance will further increase but the speed at which data can be moved down the memory hierarchy—from registers to main memory to local storage and ultimately to a parallel file system—will stagnate.
Burst buffers—a layer of high-performance storage positioned between compute nodes and back-end storage systems—are one solution to this I/O problem. While burst buffers are a useful step, they are no silver bullet, Taufer explained. Not only do burst buffers themselves have capacity limits, they allow data to be written out quickly but do not help when reading data back from storage. This is an important limitation because many applications require that information produced by the simulation be streamed back into the simulation.
Taufer suggested that one of the next advances in dealing with the I/O bottleneck will be to use in situ and in-transit analytics. By moving the analytics close to the node itself as the data are generated, knowledge to inform decisions can be extracted and fed back into the simulation immediately, rather than having to wait for a simulation result to be computed, stored, and then analyzed.
Finally, Taufer described how stream processing and in-memory processing—techniques for analyzing high-rate, real-world data—will become increasingly important tool for data-centric science. One application of stream processing is in digital twinning, where one links a simulation of a system with the real-world data collected from that system.
Modeling and Simulation (ModSim) Convergence in the Exascale Computing Project
Douglas Kothe, Oak Ridge National Laboratory
Douglas Kothe directs the Exascale Computing Project (ECP) at DOE’s Oak Ridge National Laboratory.
Although simulation has traditionally been a main driver of DOE’s computational research, the ECP incorporates a number of elements relevant to advancing convergence with data-driven science. Kothe noted that one area of work within ECP has been to enable convergence, in particular around workflow challenges, and emphasized that important gains have been made toward more dynamic interaction between analysis and simulation.
ECP aims to advance science in a wide variety of application areas, including cosmology, weather and climate, molecular dynamics, metagenomics, precision medicine, and the power grid. Projects in these areas frequently involve both big data and simulation.
To facilitate this research, ECP is incorporating and advancing technologies for AI, in situ analytics, machine learning, new approaches to steering simulations, predictive analytics, and graph analytics, among other areas. For much of this work, the ultimate goal is to bring the time required for experimental analysis down from weeks to minutes and ultimately to make it possible to interpret data in real time. Toward this goal, Kothe stressed the need for reduced precision wherever possible, making compression techniques an essential component not only for in situ analytics but deterministic simulations, as well.
ECP is also exploring opportunities for a broader AI co-design center. AI or machine learning can be valuable for developing surrogate models, which are particularly important for uncertainty quantification for complex, multi-physics applications. AI can also be used to improve data quality by helping to curate and label simulation data and reduce noise. However, Kothe emphasized that verification and validation is critical if AI is used in systems that influence decisions with national security or major economic implications.
Summit, coming online in 2018, will replace Titan as DOE’s leadership supercomputer. Summit’s increased compute power and memory, faster interconnects, higher bandwidth between CPUs and GPUs, and larger and faster file system mean that it is poised to significantly advance machine learning capabilities, making it a true “machine that thinks.”















